Abstract
The complete plastome sequencing is an efficient option for increasing phylogenetic resolution and evolutionary studies, as well as may greatly facilitate the use of plastid DNA markers in plant population genetic studies. Merostachys and Guadua stand out as the most common and the highest potential utilization bamboos indigenous of Brazil. Here, we sequenced the complete plastome sequences of the Brazilian Guadua chacoensis and Merostachys sp. to perform full plastome phylogeny and characterize the occurrence, type, and distribution of SRRs using 20 Bambuseae species. The determined plastome sequence of Merostachys sp. and G. chacoensis is 136,334 and 135,403 bp in size, respectively, with an identical gene content and typical quadripartite structure consisting of a pair of IRs separated by the LSC and SSC regions. The Maximum Likelihood and Bayesian Inference analyses produced phylogenomic trees identical in topology. These trees supported monophyly of Paleotropical and Neotropical Bamboos clades. The Neotropical bamboos segregated into three well-supported lineages, Chusqueinae, Guaduinae, and Arthrostylidiinae, with the last two forming a well-supported sister relationship. Paleotropical bamboos segregated into two well-supported lineages, Hickeliinae and Bambusinae + Melocanninae. We identified 141.8 cpSSR in Bambuseae plastomes and an inferior value (38.15) for plastome coding sequences. Among them, we identified 16 polymorphic SSR loci, with number of alleles varying from 3 to 10. These 16 polymorphic cpSSR loci in Bambuseae plastome can be assessed for the intraspecific level of polymorphism, leading to innovative highly sensitive phylogeographic and population genetics studies for this tribe.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Bamboo Phylogeny Group (BPG 2006, 2012) suggested that a taxonomic reorganization of bamboos by means of a reclassification strongly supported by estimate of phylogeny was necessary, which can be addressed with plastid genome (plastome) markers (Kelchner and Group 2013). The advancement of next-generation sequencing technologies has enabled the rapid acquisition of whole plastome sequences at low cost when compared with traditional sequencing approaches. Thus, the increasing number of plastome sequences in the public databases is now an efficient option for increasing phylogenetic resolution and evolutionary studies between and within different plant groups, families, genus and species (Zhang et al. 2011; Wysocki et al. 2015; Rogalski et al. 2015).
Besides phylogenetic studies, the low evolving rate, and the predominantly nonrecombinant, uniparentally inherited nature of plastome may greatly facilitate the use of plastid DNA markers in plant population genetic studies (Powell et al. 1995; Provan et al. 2001; Rogalski et al. 2015). By comparing nuclear and plastid markers, inferences about the relative contributions of seed and pollen flow to the genetic structure of natural populations are possible (Provan et al. 2001; Delplancke et al. 2012; Roullier et al. 2011; Khadivi-Khub et al. 2013). Powell et al. (1995) reported the presence of simple sequence repeats in plastids (cpSSR), consisting of DNA sequences in tandemly repeated motifs of 1–6 base pairs (bp). This cpSSR became widely studied due to their ability to generate highly informative DNA markers. The development and application of these plastid molecular markers was demonstrated by several studies (see review by Rogalski et al. 2015). Plastid genome sequences are also useful tools utilized to study plastid gene transfer to nucleus (Huang et al. 2003, 2005; Stegemann et al. 2003; Bock 2006; Stegemann and Bock 2006), and recently, evidence for horizontal transfer of mitochondrial DNA to the plastid genome in Pariana Aubl., a bamboo genus was found (Ma et al. 2015).
Bamboos are indigenous of all continents, except Antarctica and Europe, and occupy a broad range of habitat types, especially forests, from temperate to tropical climate zones (Clark et al. 2015). Brazil holds a high diversity of bamboo species, of which we can highlight Merostachys Spreng. (43 native species—41 endemic species), Chusquea Kunth (45 native species—41 endemic species) as the most common genera, and Guadua Kunth (18 native species—5 endemic species) as the highest important for use in construction, furniture, and handcraft (Greco et al. 2015; List of Species of the Brazilian Flora 2015).
Merostachys s.l. is the largest genus of the subtribe Arthrostylidiinae Soderstr. and R.P. Ellis (Lizarazu et al. 2011). This genus spreads from Mexico to Southern Brazil in forest and forest margin habitats, but it is particularly diverse in Brazil, where remains many undescribed species (Judziewicz et al. 1999; Lizarazu et al. 2011; Santos-Gonçalves et al. 2012; Viana et al. 2013; Viana and Filgueiras 2014; Greco et al. 2015). Its taxonomic identification has been demonstrated to be very complicated due to their long periods of vegetative development, ca. 25–30 years (Lizarazu et al. 2011).
Guadua s.l. is one of the five genera placed in the subtribe Guaduinae Soderstr. and R.P. Ellis. This genus occurs throughout tropical America, from Mexico to Brazil and northern Argentina (Londoño and Peterson 1992). Guadua chacoensis (Rojas) Londoño & P.M. Peterson is a native species from Brazil, but also occurs in northern Argentina, southeastern Bolivia, and southern Paraguay, and is one of the three southeasternmost species of the genus (Londoño and Peterson 1992).
Here, we sequenced and analyzed the complete plastome sequences of two Brazilian native species of tribe Bambuseae: G. chacoensis and Merostachys sp. We also performed a full plastome phylogeny using 20 Bambuseae species with 2 newly sequenced and 18 existing plastomes, and characterized the occurrence, type, and distribution of SRRs in the Bambuseae. Merostachys sp. is the first species of subtribe Arthrostylidiinae to have its plastome sequenced.
Materials and methods
Sequencing and assembly of the bamboo plastomes
Fresh leaf material of Merostachys sp. (voucher Th. Greco 18—FLOR_16/09/2011) were collected from a natural population, and Guadua chacoensis (voucher Th. Greco 159—FLOR_15/02/2013) were collected from cultivated plants in Florianópolis, Santa Catarina—Brazil (27°35′49″S48°32′56″W). Both species were located on a private property and were collected with the owner permission. The chloroplast isolation and plastid DNA extraction were carried out using leaves from a single plant following the protocol described by Vieira et al. (2014). A total of 1 ng of plastid DNA was used to prepare sequencing libraries with Nextera XT DNA Sample Prep Kit (Illumina Inc., San Diego, California, USA) according to the manufacturer’s instructions. Libraries were sequenced using MiSeq Reagent Kit v3 (600 cycles) on Illumina MiSeq Sequencer (Illumina Inc., San Diego, California, USA).
The obtained paired-end reads (2 × 300 bp) were used for de novo assembly performed by CLC Genomics Workbench 8.0v. Initial annotation of the obtained plastome sequences was performed using Dual Organellar GenoMe Annotator (DOGMA) (Wyman et al. 2004). From this initial annotation, putative starts, stops, and intron positions were determined based on comparisons to homologous genes in other plastomes. The tRNA genes were further verified by using tRNAscan-SE (Schattner et al. 2005). The physical map of the circular plastomes was drawn using OrganellarGenomeDRAW (OGDRAW) (Lohse et al. 2013). REPuter (Kurtz et al. 2001) was used to identify the IRs in both plastomes sequenced in this work and those used for phylogeny estimation by forward versus reverse complement (palindromic) alignment. The minimal repeat size was set to 30 bp and the identity of repeats ≥90 %. The complete nucleotide sequence of Merostachys sp. and G. chacoensis plastome were deposited in the GenBank database under accession number KT373815 and KT373814, respectively.
Phylogeny estimation
The phylogeny estimation was done according to Wysocki et al. (2015). The IRA was omitted to prevent over representation of the IR sequences. G. chacoensis and Merostachys sp. plastomes were aligned using MAFFT program (Katoh and Standley 2013) along with 18 previously published Bambuseae plastomes (Table 1) and Lolium perenne (Poaceae: Pooideae; NC_009950) plastome was used as outgroup. Nucleotide positions that contained one or more gaps introduced by the alignments were omitted from the matrix. The substitution model was selected using jModeltest (number of substitution schemes = 5). The General Time Reversible model of substitution, incorporating invariant sites and a gamma distribution (GTR + I + G), was used in subsequent plastome analyses. Maximum likelihood (ML) analysis was performed using the RAxML v 7.2.8 (Stamatakis 2006) with 1000 non-parametric bootstrap replicates. MrBayes 3.2.2 (Ronquist and Huelsenbeck 2003) was used to perform a Bayesian inference (BI) analysis. The Markov chain Monte Carlo (MCMC) analysis was run for 2,000,000 generations. Average standard deviation of split frequencies remained below 0.001 after the 25 % burn-in. Resulting trees were represented and edited using FigTree v1.4.1.
Simple sequence repeats analysis
Simple sequence repeats (SSRs) were detected using MISA perl script, available at (http://pgrc.ipk-gatersleben.de/misa/), with threshold of eight repeat units for mononucleotide SSRs, four repeat units for di- and tri-nucleotide SSRs, and three repeat units for tetra-, penta- and hexa-nucleotide SSRs. For the SSRs identification, we used the plastid genome sequence of all species described in Table 1 with one IR region removed, and in their coding sequences (CDS). Additionally, we compared the identified SSRs in order to find a potential set of microsatellite markers identified for all Bambusea species described in Table 1. Primers were designed for 16 interspecific polymorphic SSR with the software PRIMER3 (http://bioinfo.ut.ee/primer3-0.4.0/) by setting product size ranges from 100 to 300 bp, primer size from 18 to 25 bp, and GC content from 40 to 60, with 1 °C as the maximum difference between the melting temperatures of the left and right primers.
Results
Plastome assembly and content
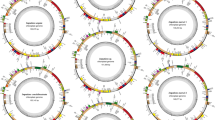
The Illumina MiSeq reads obtained and submitted to de novo assembly resulted in high genome coverage for both G. chacoensis (~140×) and Merostachys sp. (~460×) plastomes (Table 2). The determined complete plastome sequence of Merostachys sp. and G. chacoensis is 136,334 and 135,403 bp in size, respectively, with a GC content determined of 38.81 % for Merostachys sp. and 38.76 % for G. chacoensis. The Guadua genus presents the smaller plastome size in comparison with other species belonging to tribe Bambuseae (135,324–135,403 bp), ~1000 bp smaller than Merostachys sp., but with an identical gene content (Tables 1, 3). Both plastomes exhibit the general quadripartite structure typical of angiosperms, consisting of a pair of IRs (20,258–19,773 bp) separated by the LSC (82,859–82,877 bp) and SSC (12,960–12,980 bp) regions (Table 1). The IR/SSC boundary is just within the coding sequence of ndhH, creating a short ndhH fragment in IRA. They encode an identical set of 132 genes and 4 pseudogenes with the same gene order and gene clusters, of which 90 genes were single copy and 21 genes were duplicated (Fig. 1; Table 3). The following genes were identified and are listed in Fig. 1 and Table 3: 4 duplicated ribosomal RNA genes, 21 unique and 9 duplicated transfer RNA genes, 15 unique and 6 duplicated genes encoding large and small ribosomal subunits, 1 translational initiation factor, 4 genes encoding DNA-dependent RNA polymerases, 45 unique and 1 duplicated genes encoding photosynthesis-related proteins, 4 unique and 1 duplicated genes encoding other proteins, including the unknown function gene ycf68.
Gene map of Guadua chacoensis plastid genome. Genes drawn inside the circle are transcribed clockwise, and genes drawn outside are counterclockwise. Genes belonging to different functional groups are color-coded. The darker gray in the inner circle corresponds to GC content, and the lighter gray to AT content. Gene map of Merostachys sp. is equal to G. chacoensis in gene order and content
Phylogeny estimation
The matrix of the 21 completely aligned Bambusoideae plastomes was 122,950 nucleotide positions in length, and the exclusion of gaps reduced this matrix to 105,270 sites. ML (−lnL = −206179.38) and BI (−lnL = −206212.43) analyses produced phylogenomic trees that were identical in topology, which is represented by Fig. 2. The ML bootstrap values between 68 and 98 % for 4 nodes and 100 % for the rest, and the BI posterior probability values between 0.95 and 0.99 for 3 nodes and 1.0 for the rest (Fig. 2). These trees supported monophyly of Paleotropical and Neotropical Bamboos clades. The Neotropical bamboos segregated into three well-supported lineages, Chusqueinae, Guaduinae, and Arthrostylidiinae, with Guaduinae forming a well-supported sister relationship with Arthrostylidiinae. All Neotropical bamboos genera (Chusquea, Guadua, Olmeca, Otatea, Merostachys) were resolved as monophyletic with 100 % ML bootstrap support (Fig. 2). The Paleotropical bamboos segregated into two well-supported lineages, Hickeliinae and Bambusinae + Melocanninae. The genus Bambusa was resolved as monophyletic with 100 % ML bootstrap support with very short branches and two internal nodes with 81 and 98 % ML bootstrap support (Fig. 2). Dendrocalamus latiflorus (Bambusinae) and Neohouzeaua sp. (Melocanninae) formed a well-supported node, with 100 % ML bootstrap support. These two species are nowadays classified as belonging to different subtribes (Clark et al. 2015). Greslania sp. and Neololeba atra form a well-supported sister relationship.
Bayesian phylogeny of 20 complete plastomes sequences of Bambuseae tribe species and the outgroup Lolium perenne (Poaceae: Pooideae). The numbers above the branches are maximum likelihood bootstrap values/Bayesian posterior probabilities. The branch length is proportional to the inferred divergence level, the scale bar indicating the number of inferred nucleic acids substitutions per site
Simple sequence repeats analysis
We analyzed the occurrence, type, and distribution of SRRs in the Bambuseae species listed in Table 1. The number and types of SSR identified for all 20 Bambuseae species were similar (Fig. 3). In the complete plastome sequence, the average of the total identified SSR in all Bambuseae species were 141.8 (1 SSR unit every 823.14 bp), while for coding sequences a lower value was found (38.15; 1 SSR unit every 1466.60 bp). Among them, mono- and di-nucleotide repeats were the most common, whereas tri- and tetra-nucleotide repeats occurred with lower frequency (Fig. 3). Penta- and hexa-nucleotide repeats occurred only in complete plastome sequence, no penta- or hexa-nucleotide repeats were identified in coding sequences for any analyzed species.
Although we observed that the number and type of SSR identified in Bambuseae species were quite similar, the same was not true for their presence across all species. Less than half of SSR (di-, tri-, tetra-, and penta-nucleotide) were common across all species (Table 4). In addition, all these SSR cited in Table 4 were assessed for interspecific polymorphism, and except for (TTCTA)6–7, common to G. weberbaueri and G. chacoensis, respectively, no other SSR showed interspecific polymorphism. For mononucleotide repeats, it was possible to identify 16 polymorphic SSR loci with conserved down- and up-stream regions (Table 5). For these regions specific primers were designed, and the polymorphisms for these 16 loci ranged from 3 to 10 alleles, 15 in intergenic regions and intron 1 (Table 5).
The most mononucleotide repeats identified were constituted by A/T sequences (96.61 %), and for the dipolymers, 56.85 % were also constituted by multiple A and T bases, no dinucleotide repeats constituted by multiple C and G bases were found (Fig. 3). Similarly, for trinucleotide repeats, 31.92 % were constituted by multiple A and T bases (AAT/ATT), 48.17 % by AAG/CTT and 19.92 % by AGC/CTG. We also identified 11 tetranucleotide repeats, with AAAT/ATTT being the most common (33.86 %). Interestingly, we also did not identified any tri- of tetra-nucleotide unit size constituted by only C or G bases.
Discussion
Plastome assembly and content
The plastome size in Neotropical bamboos was typically shorter than in Paleotropical bamboos, with determined plastomes measuring between 135,324–138,257 and 138,276–139,606 bp, respectively (Table 1). This increase in Paleotropical plastome sizes seems to be distributed by LSC and IR regions, which ranges between 80,743–82,859 and 81,925–83,145 bp for LSC regions in Netropical and Paleotropical bamboos, respectively, and between 19,773–21,804 and 21,755–21,852 bp for IR regions, also respectively. Differently, for the SSC regions, little length variation was found (12,671–12,980 bp for Neotropical and 12,743–12,980 bp for Paleotropical bamboos), suggesting that this region is more conserved in size than LSC and IR regions for Bambuseae. The IR/SSC boundary was conserved in Bambuseae tribe, in G. chacoensis, Merostachys sp. and all Bambuseae plastomes analyzed in this study (Table 1) the IR/SSC boundary was just within the coding sequence of ndhH, with a short ndhH fragment in IRA (Wu et al. 2009; Zhang et al. 2011; Burke et al. 2012). This feature is similar to the results found in other Poaceae species (Ogihara et al. 2002; Saski et al. 2007), but differ from the Olyreae species, which were shown to present the IR/SSC boundary within ndhF coding sequence, an suggested synapomorphy for the tribe (Burke et al. 2012, 2014). The gene content and gene order of the G. chacoensis and Merostachys sp. plastomes were identical to other sequenced Bambuseae plastomes, including the loss of introns in clpP and rpoC1, the loss of accD, ycf1 and ycf2 genes, and the IR expansion to include rps19 (Wu et al. 2009; Zhang et al. 2011; Burke et al. 2012).
Phylogeny estimation
Plastid genomes are very evolutionarily conserved, presenting rates of substitutions extremely low as compared to nuclear genomes (Palmer 1985). In addition, different plastome regions evolve at different rates, allowing measuring evolutionary distance at many taxonomy levels (Palmer, 1985; Shaw et al. 2005, 2007). Phylogenetic inferences based on whole plastome sequences have been used to address evolutionary features in Bambusoideae (Wu et al. 2009; Zhang et al. 2011; Burke et al. 2012, 2014; Wysocki et al. 2015). These full plastome analyzes provide enough information to resolve difficult interspecific relationships, an issue to woody bamboos that generally hybridize readily and exhibit very long generation times (Wysocki et al. 2015).
In Bambuseae, the well supported monophyletic lineages that represent Neotropical and Paleotropical woody bamboos retrieved here were previously reported in phylogenetic studies using combined analysis of plastid DNA regions (Sungkaew et al. 2009) and complete plastome sequences (Wysocki et al. 2015). In Wysocki et al. (2015), the segregation of Neotropical bamboos into two well-supported lineages, Chusqueinae and Guaduinae, was reported.
In this work, we present for the first time a phylogenetic tree based on complete plastome sequence including an Arthrostylidiinae species, providing the phylogenetic tree with well supported monophyly of subtribes and with a sister relationship between Guaduinae and Arthrostylidiinae. Previous analyses based on matK and/or ndhF sequence data also reported this sister relationship (Guala et al. 2000; Zhang and Clark 2000; Soreng et al. 2015), and a study based on morphology and rpl16 intron sequence data indicate that Guaduinae may be derived from within Arthrostylidiinae (Clark et al. 2007). In Chusqueinae, especially Chusquea spectabilis, there was a high substitution rate in both ML and BI analyses. This feature has been reported by Wysocki et al. (2015) and associated with the extremely different flower intervals in Chusqueinae species, and the shorter flowering intervals observed in C. spectabilis. In addition, C. spectabilis long branches may be associated with the fact that this species belongs to subgenus Magnifoliae, one of the two earliest-diverging clades in genus Chusquea, and C. circinata (subgenus Rettbergia) is known to be a sister of the large Euchusquea clade, represented in this analysis by C. liebmannii (Fisher et al. 2014).
Low support (82 % ML bootstrap) between the Paleotropical bamboos subtribes, Hickeliinae and Bambusinae were reported by Wysocki et al. (2015), corroborating our results with low support between Hickeliinae and Bambusinae + Melocanninae (74 % ML bootstrap support). Wysocki et al. (2015) also reported the well supported clade Neololeba atra + Greslania sp., as well as Bambusa spp. monophyly. However, the inclusion of Neohouzeaua sp. (Melocanninae), resulted in its unexpected sister relationship to D. latiflorus (Bambusinae), and consequently, the non-monophyly of Bambusinae.
Simple sequence repeats analysis
Powell et al. (1995) introduced the cpSSR as a readily usable chloroplast marker, exhibiting length variation and polymorphism. Even though in plastomes the occurrence of di-, tri-, tetra-, penta-, and hexa-nucleotide repeats is less common (George et al. 2015), it became a widely used molecular marker, which have aroused considerable interest due to their ability to generate highly informative DNA markers (Provan et al. 2001). These regions may be used for both intraspecific and interspecific variability analyses, with practical value for monitoring gene flow, population differentiation and cytoplasmic diversity (Powell et al. 1995), in both basic plant sciences and applied agricultural. In native species, cpSSRs have been used most frequently in population genetics studies, but also for understanding uniparental genetic structure (e.g. seed or pollen dispersal) and in studies of hybridization (see review by Wheeler et al. 2014). Given that not all chloroplast loci are likely to be equally diverse across plants, it is important to consider a more targeted approach to developing cpSSR loci specific to a study system rather than relying on universal primers (Wheeler et al. 2014).
The cpSSRs may be identified in completely sequenced plant chloroplast genomes by simple database searches, followed by primers designed to screen for polymorphism even in non-model species, and is becoming a common pathway for developing variable markers (Wheeler et al. 2012; Li et al. 2014; Nazareno et al. 2015). George et al. (2015) analyzed the abundance and distribution of simple and compound SSR in 164 sequenced plastomes from wide range of plants. Corroborating our results in Bambuseae, George et al. (2015) described that mononucleotide repeats occurs in higher number as compared to di-, tri-, tetra-, penta-, and hexa-nucleotide repeats, longer SSRs are excluded from coding regions, AT/TA followed by CT/TC was the most common dinucleotide repeat motif observed, and GC/CG was rarely found and even absent from few plastomes.
In the present study, we determined the complete plastome sequence of Merostachys sp. and Guadua chacoensis, which were identical in gene content and order, also with identical gene content as compared to other Bambuseae plastomes. These results enabled the report for the first time of a phylogenetic tree based on complete plastome sequences including an Arthrostylidiinae species, the Merostachys sp. The ML and BI trees supported monophyly of Paleotropical and Neotropical Bamboos clades, with all Neotropical bamboos genera resolved as monophyletic. The Paleotropical bamboos segregated into two well-supported lineages, Hickeliinae and Bambusinae + Melocanninae. We also identified several SSR loci, whose distribution and types are highly similar between sequenced plastomes. Among them, we identified 16 polymorphic SSR loci suitable for interspecific population genetic analysis, with number of alleles varying from 3 to 10. These 16 polymorphic cpSSR loci in Bambuseae plastome can be assessed for the intraspecific level of polymorphism, leading to innovative highly sensitive phylogeographic and population genetics studies for this tribe. Merostachys sp. and G. chacoensis are both native species from the Atlantic Forest Biome, which is considered one of the 25 biodiversity hotspots of the world (Myers et al. 2000), increasing the interest in population genetics studies for these species.
Availability of supporting data
All nucleotide sequences were deposited in the NCBI Genbank repository. Genbank accessions numbers can be found in Table 1.
References
Bamboo Phylogeny Group (2006) The bamboo phylogeny project. Bamboo 27:11–14
Bamboo Phylogeny Group (2012) An updated tribal and subtribal classification for the Bambusoideae (Poaceae). In: Gielis J, Potters G (eds) Proceedings of the 9th World Bamboo Congress. Antwerp, Belgium: World Bamboo Organization, pp 3–27
Bock R (2006) Extranuclear inheritance: gene transfer out of plastids. Prog Bot 67:75–100. doi:10.1007/3-540-27998-9_4
Burke SV, Colin PG, Melvin RD (2012) Plastome sequences of two New World bamboos—Arundinaria gigantea and Cryptochloa strictiflora (Poaceae)—extend phylogenomic understanding of Bambusoideae. Am J Bot 99:1951–1961. doi:10.3732/ajb.1200365
Burke SV, Clark LG, Triplett JK, Grennan CP, Duvall MR (2014) Biogeography and phylogenomics of new world Bambusoideae (Poaceae), revisited. Am J Bot 101:886–891. doi:10.3732/ajb.1400063
Clark LG, Dransfield S, Triplett JK, Sánchez-Ken JG (2007) Phylogenetic relationships among the one-flowered, determinate genera of Bambuseae (Poaceae: Bambusoideae). Aliso 23:315–332
Clark LG, Londoño X, Ruiz-Sanchez E (2015) Bamboo taxonomy and habitat. In: Liese W, Kohl M (eds) Bamboo, Tropical forestry. Springer International Publishing, Switzerland, pp 1–30
Delplancke M, Alvarez N, Espíndola A, Joly H, Benoit L, Brouck E, Arrigo N (2012) Gene flow among wild and domesticated almond species: insights from chloroplast and nuclear markers. Evol Appl 5:317–329. doi:10.1111/j.1752-4571.2011.00223.x
Fisher AE, Clark LG, Kelchner SA (2014) Molecular phylogeny estimation of the bamboo genus Chusquea (Poaceae: Bambusoideae: Bambuseae) and description of two new subgenera. Syst Bot 39:829–844. doi:10.1600/036364414X681554
George B, Bhatt BS, Awasthi M, George B, Singh AK (2015) Comparative analysis of microsatellites in chloroplast genomes of lower and higher plants. Curr Genet. doi:10.1007/s00294-015-0495-9
Greco TM, Pinto MM, Tombolato FC, Xia N (2015) Diversity of bamboo in Brazil. J Trop Subtrop Botany 23:1–16. doi:10.11926/j.issn.1005-3395.2015.01.001
Guala G, Bogler D, Sadle J, Francisco-Ortega J (2000) Molecular evidence for polyphyly in the genus Apoclada (Poaceae: Bambusoideae). Bamboo Sci Cult 14:15–20
Huang CY, Ayliffe MA, Timmis JN (2003) Direct measurement of the transfer rate of chloroplast DNA into the nucleus. Nature 422:72–76. doi:10.1038/nature01435
Huang CY, Grunheit N, Ahmadinejad N, Timmis JN, Martin W (2005) Mutational decay and age of chloroplast and mitochondrial genomes transferred recently to angiosperm nuclear chromosomes. Plant Physiol 138:1723–1733. doi:10.1104/pp.105.060327
Judziewicz EJ, Clark LG, Londono X, Stern M (1999) American bamboos. Smithsonian Institution Press, Washington DC
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. doi:10.1093/molbev/mst010
Kelchner SA, Group BP (2013) Higher level phylogenetic relationships within the bamboos (Poaceae: Bambusoideae) based on five plastid markers. Mol phylogenetics Evol 67:404–413. doi:10.1016/j.ympev.2013.02.005
Khadivi-Khub A, Zamani Z, Fattahi R, Wünsch A (2013) Genetic variation in wild Prunus L. subgen. Cerasus germplasm from Iran characterized by nuclear and chloroplast SSR markers. Trees 28:471–485. doi:10.1007/s00468-013-0964-z
Kurtz S, Choudhuri JV, Ohlebusch E, Schleiermacher C, Stoye J, Giegerich R (2001) REPuter: the manifold applications of repeat analysis on a genomic scale. Nucl Acids Res 29:4633–4642
Li W, Feng Y, Sun H, Deng Y, Yu H, Chen H (2014) Analysis of simple sequence repeats in the Gaeumannomyces graminis var. tritici genome and the development of microsatellite markers. Curr Genet 60:237–245. doi:10.1007/s00294-014-0428-z
List of Species of the Brazilian Flora (2015) Rio de Janeiro Botanical Garden. http://floradobrasil.jbrj.gov.br. Accessed 04 July 2015
Lizarazu MA, Agrasar ZER, Vega AS (2011) A new species of Merostachys (Poaceae, Bambusoideae, Bambuseae) and synopsis of the genus in Argentina and neighboring regions. Sist Bot 36:896–906. doi:10.1600/036364411X604912
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucl Acids Res. doi:10.1093/nar/gkt289
Londoño X, Peterson PM (1992) Guadua chacoensis (Poaceae: Bambuseae), its taxonomic identity, morphology, and affinities. Novon 2:41–47
Ma PF, Zhang YX, Guo ZH, Li DZ (2015) Evidence for horizontal transfer of mitochondrial DNA to the plastid genome in a bamboo genus. Sci Rep 5:11608. doi:10.1038/srep11608
Myers N, Mittermeier RA, Mittermeier CG, Fonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858. doi:10.1038/35002501
Nazareno AG, Carlsen M, Lohmann LG (2015) Complete Chloroplast Genome of Tanaecium tetragonolobum: The First Bignoniaceae Plastome. PLoS One 10(6):e0129930. doi:10.1371/journal.pone.0129930
Ogihara Y, Isono K, Kojima T, Endo A, Hanaoka M, Shiina T, Terachi T, Utsugi S, Murata M, Mori N, Takumi S, Ikeo K, Gojobori T, Murai R, Murai K, Matsuoka Y, Ohnishi Y, Tajiri H, Tsunewaki K (2002) Structural features of a wheat plastome as revealed by complete sequencing of chloroplast DNA. Mol Genet Genomics 266:740–746. doi:10.1007/s00438-001-0606-9
Palmer JD (1985) Chloroplast DNA and molecular phylogeny. BioEssays 2:263–267
Powell W, Morgantet M, Andre C, McNicol JW, Machray GC, Doyle JJ, Tingey SV, Rafalski JA (1995) Hypervariable microsatellites provide a general source of polymorphic DNA markers for the chloroplast genome. Curr Biol 5:1023–1029
Provan J, Powell W, Hollingsworth PM (2001) Chloroplast microsatellites: new tools for studies in plant ecology and evolution. Trends Ecol Evol 16:142–147. doi:10.1016/s0169-5347(00)02097-8
Rogalski M, Vieira LN, Fraga HPF, Guerra MP (2015) Plastid genomics in horticultural species: importance and applications for plant population genetics, evolution, and biotechnology. Front Plant Sci 6:586. doi:10.3389/fpls.2015.00586
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinform 19:1572–1574. doi:10.1093/bioinformatics/btg180
Roullier C, Rossel G, Tay D, McKey D, Lebot V (2011) Combining chloroplast and nuclear microsatellites to investigate origin and dispersal of New World sweet potato landraces. Mol Ecol 20:3963–3977. doi:10.1111/j.1365-294X.2011.05229.x
Santos-Gonçalves APS, Carvalho-Okano RMD, Filgueiras TS (2012) A new species of Merostachys (Poaceae: Bambusoideae) from southeastern Brazil. Syst Bot 37:938–940. doi:10.1600/036364412X656400
Saski C, Lee SB, Fjellheim S, Guda C, Jansen RK, Luo H, Tomkins J, Rognli OA, Daniell H, Clarke JL (2007) Complete chloroplast genome sequences of Hordeum vulgare, Sorghum bicolor and Agrostis stolonifera, and comparative analyses with other grass genomes. Theor Appl Genet 115:571–590. doi:10.1007/s00122-007-0567-4
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:W686–W689. doi:10.1093/nar/gki366
Shaw J, Lickey EB, Beck JT, Farmer SB, Liu W, Miller J, Siripun KC, Winder CT, Schilling EE, Small RL (2005) The tortoise and the hare II: relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am J Bot 92:142–166. doi:10.3732/ajb.92.1.142
Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Am J Bot 94:275–288. doi:10.3732/ajb.94.3.275
Soreng RJ, Peterson PM, Romaschenko K, Davidse G, Zuloaga FO, Judziewicz EJ, Filgueiras TS, Davis JI, Morrone O (2015) A worldwide phylogenetic classification of the Poaceae (Gramineae). J Syst Evol 53:117–137. doi:10.1111/jse.12150
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinform 22:2688–2690. doi:10.1093/bioinformatics/btl446
Stegemann S, Bock R (2006) Experimental reconstruction of functional gene transfer from the tobacco plastid genome to the nucleus. Plant Cell 18:2869–2878. doi:10.1105/tpc.106.046466
Stegemann S, Hartmann S, Ruf S, Bock R (2003) High-frequency gene transfer from the chloroplast genome to the nucleus. Proc Natl Acad Sci USA 100:8828–8833. doi:10.1073/pnas.1430924100
Sungkaew S, Stapleton CM, Salamin N, Hodkinson TR (2009) Non-monophyly of the woody bamboos (Bambuseae; Poaceae): a multi-gene region phylogenetic analysis of Bambusoideae ss. J Plant Res 122:95–108. doi:10.1007/s10265-008-0192-6
Viana PL, Filgueiras TS (2014) Three new species of Aulonemia (Poaceae: Bambusoideae) from the Brazilian Atlantic rainforest. Phytotaxa 156:235–249. doi:10.11646/phytotaxa.156.4.6
Viana PL, Filgueiras TS, Clark LG (2013) Cambajuva (Poaceae: Bambusoideae: Bambuseae: Arthrostylidiinae), a new woody bamboo genus from southern Brazil. Syst Bot 38:97–103. doi:10.1600/036364413X662097
Vieira LN, Faoro H, Fraga HPF, Rogalski M, Souza EM et al (2014) An improved protocol for intact chloroplasts and cpDNA isolation in conifers. PLoS One 9(1):e84792. doi:10.1371/journal.pone.0084792
Wheeler GL, McGlaughlin ME, Wallace LE (2012) Variable length chloroplast markers for population genetic studies in Acmispon (Fabaceae). Am J Bot 99:e408–e410. doi:10.3732/ajb.1200129
Wheeler GL, Dorman HE, Buchanan A, Challagundla L, Wallace LE (2014) A review of the prevalence, utility, and caveats of using chloroplast simple sequence repeats for studies of plant biology. Appl Plant Sci 2:1400059. doi:10.3732/apps.1400059
Wu FH, Kan DP, Lee SB, Daniell H, Lee YW, Lin CC, Lin NS, Lin CS (2009) Complete nucleotide sequence of Dendrocalamus latiflorus and Bambusa oldhamii chloroplast genomes. Tree Physiol tpp015:1–10. doi:10.1093/treephys/tpp015
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255. doi:10.1093/bioinformatics/bth352
Wysocki WP, Clark LG, Attigala L, Ruiz-Sanchez E, Duvall MR (2015) Evolution of the bamboos (Bambusoideae; Poaceae): a full plastome phylogenomic analysis. BMC Evol Biol 15:50. doi:10.1186/s12862-015-0321-5
Zhang W-P, Clark LG (2000) Phylogeny and classification of the bamboos (Poaceae: Bambusoideae). In: Jacobs SWL, Everett J (eds) Grasses: systematics and evolution. CSIRO Publishing, Collingwood, Victoria, pp 35–42
Zhang Y-J, Ma P-F, Li D-Z (2011) High-throughput sequencing of six bamboo chloroplast genomes: phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS One 6(5):e20596. doi:10.1371/journal.pone.0020596
Acknowledgments
This work was supported by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq Proc. 457726/2013-0, and 306126/2013-3). The authors thank to Instituto Çarakura for kindly providing plant material.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Vieira, L.d., dos Anjos, K.G., Faoro, H. et al. Phylogenetic inference and SSR characterization of tropical woody bamboos tribe Bambuseae (Poaceae: Bambusoideae) based on complete plastid genome sequences. Curr Genet 62, 443–453 (2016). https://doi.org/10.1007/s00294-015-0549-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-015-0549-z