Abstract
Chloroplast genome organization, gene order, and content are highly conserved among land plants. We sequenced the chloroplast genome of Trachelium caeruleum L. (Campanulaceae), a member of an angiosperm family known for highly rearranged genomes. The total genome size is 162,321 bp, with an inverted repeat (IR) of 27,273 bp, large single-copy (LSC) region of 100,114 bp, and small single-copy (SSC) region of 7,661 bp. The genome encodes 112 different genes, with 17 duplicated in the IR, a tRNA gene (trnI-cau) duplicated once in the LSC region, and a protein-coding gene (psbJ) with two duplicate copies, for a total of 132 putatively intact genes. ndhK may be a pseudogene with internal stop codons, and clpP, ycf1, and ycf2 are so highly diverged that they also may be pseudogenes. ycf15, rpl23, infA, and accD are truncated and likely nonfunctional. The most conspicuous feature of the Trachelium genome is the presence of 18 internally unrearranged blocks of genes inverted or relocated within the genome relative to the ancestral gene order of angiosperm chloroplast genomes. Recombination between repeats or tRNA genes has been suggested as a mechanism of chloroplast genome rearrangements. The Trachelium chloroplast genome shares with Pelargonium and Jasminum both a higher number of repeats and larger repeated sequences in comparison to eight other angiosperm chloroplast genomes, and these are concentrated near rearrangement endpoints. Genes for tRNAs occur at many but not all inversion endpoints, so some combination of repeats and tRNA genes may have mediated these rearrangements.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Land plant chloroplast genomes are highly conserved in organization, gene order, and content. Typically, these circular genomes range in size from about 115 to 165 kb (Palmer 1991; Raubeson and Jansen 2005) and are organized into large (LSC) and small (SSC) single-copy regions, separated by an inverted repeat (IR). The chloroplast genomes of ferns, the gymnosperm Ginkgo, and most angiosperms are nearly colinear, reflecting the conserved gene order in lineages that diverged from lycopsids and the ancestral chloroplast gene order over 350 million years ago (Raubeson and Jansen 1992). Likewise, the gene content of land plant chloroplast genomes is highly consistent across taxa. Earlier mapping studies identified a number of distantly related families including Fabaceae, Geraniaceae, and the closely related Campanulaceae and Lobeliaceae, in which at least several rearrangements have occurred (reviewed by Raubeson and Jansen 2005). Of these more complex genomes, the only one that has been completely sequenced to date is the highly rearranged chloroplast genome of Pelargonium (Geraniaceae [Chumley et al. 2006]). Many of the rearrangements and gene duplications found in Pelargonium are attributed to the massive expansion of the inverted repeat regions into the LSC and SSC regions, as well as a series of inversions of blocks of genes (Palmer et al. 1987; Chumley et al. 2006).
Gene mapping studies of representatives of the Campanulaceae (Cosner 1993; Cosner et al. 1997, 2004) and Lobeliaceae (Knox et al. 1993; Knox and Palmer 1999) identified large inversions, contraction and expansion of the inverted repeats, and several insertions and deletions in the chloroplast DNAs (cpDNAs) of these closely related families. Detailed restriction site and gene mapping of the chloroplast genome of Trachelium caeruleum (Campanulaceae) identified 7 to 10 large inversions, five families of repeats associated with rearrangements, possible transpositions, and even the disruption of operons (Cosner et al. 1997). Seventeen other members of the Campanulaceae were mapped and exhibit many additional rearrangements (Cosner et al. 2004). The cause of rearrangements in this group is unclear based on the limited resolution available with mapping techniques, but several mechanisms have been proposed: recombination between repeats, transposition, and temporary instability due to loss of the inverted repeat (Cosner et al. 1997). Sequencing whole chloroplast genomes within the Campanulaceae offers a unique opportunity to examine both the extent and the mechanisms of rearrangements within a phylogenetic framework.
We report here the first complete chloroplast genome sequence of a member of the Campanulaceae, Trachelium caeruleum. This work will serve as a benchmark for subsequent sequencing and comparative analysis of other members of this family and close relatives, with the goal of further understanding chloroplast genome evolution. We confirmed features previously identified through mapping, and discovered many additional structural changes, including several partial to entire gene duplications, deterioration of at least four normally conserved chloroplast genes into gene fragments, and the nature and position of numerous repeat elements at or near inversion endpoints.
The focus of our study was on characterizing sequences at or near major rearrangements in Trachelium caeruleum. Inversions are believed to occur in chloroplast genomes due to the presence of repeat elements subject to homologous recombination (Palmer 1991; Knox et al. 1993). Repeats may facilitate inversions or other genome rearrangements (Achaz et al. 2003), and higher incidences of repeats have been correlated with greater numbers of rearrangements (Rocha 2003; Pombert et al. 2006; Chumley et al. 2006). Alternatively, repeats may proliferate within a genome as a result of DNA strand repair mechanisms following a rearrangement event such as an inversion. Gene mapping studies previously identified five families of dispersed repeats in Trachelium at or near inversion endpoints (Cosner et al. 1997). Here we examined the sequences of these repeats and identify, map, and characterize numerous additional repeats within the genome. We compared the number and size of repeats in other angiosperm chloroplast genomes to what we found in the highly rearranged chloroplast genome of Trachelium. The Trachelium chloroplast genome has the highest number of and the largest repeats, along with Pelargonium (Chumley et al. 2006) and the less rearranged chloroplast genome of Jasminum (Oleaceae [Lee et al. 2007]). In Trachelium, these repeats are generally clustered at or near rearrangement endpoints and they are of diverse origins: partial or entire chloroplast gene duplications, noncoding chloroplast sequences, or novel DNA with no clear sequence identity to any existing chloroplast DNA sequences. Trachelium has one of the most highly rearranged chloroplast genomes of land plants and its bizarre organization is clearly associated with the high incidence of dispersed repetitive DNA.
Materials and Methods
Sample Acquisition, cpDNA Isolation, and DNA Sequencing
Trachelium caeruleum plants were purchased from a local nursery and grown in the UT-Austin research greenhouses. Plants were placed in the dark for 24 h prior to harvesting of leaves, and a voucher specimen (RCHaberle154) is deposited at TEX. A chloroplast DNA-enriched sample was isolated from living material using the sucrose gradient method (Palmer 1986; Jansen et al. 2005). The DNA was sheared into ∼3-kb fragments using a Hydroshear device (Genemachines, San Carlos, CA, USA), then these were enzymatically repaired to blunt ends, gel purified, and ligated into pUC18 plasmid vector, which was then introduced into competent E. coli by electroporation to create a random shotgun library for sequencing. Colonies were picked randomly into 384-well plates with bacterial media and these were grown overnight without shaking or aeration. An aliquot was processed using rolling circle amplification and one sequencing read was made from each end of each clone using BigDye terminators (Applied Biosystems). Sequencing reads were processed using PHRED, assembled using PHRAP (Ewing and Green 1998), and visualized using CONSED (Gordon et al. 1998) and Sequencher (Gene Codes Corp., Ann Arbor, MI). The draft sequence had 8–10 × coverage but included multiple areas of low coverage as well as gaps between contigs. We developed primers that amplified chloroplast-enriched DNA from the original isolation to augment areas of low coverage and to fill in gaps with at least two reads with a PHRED quality score (Q value) of at least 20 (Jansen et al. 2005). Using Sequencher, we manually reconstructed part of the second copy of the inverted repeat (IR), as automated PHRAP assembly cannot distinguish between reads that belong in one or the other copy. This allowed us to produce a complete circular genome with both copies of the IR for annotation and analysis.
Genome Annotation and Analysis
Genes were annotated using DOGMA (Dual Organellar GenoMe Annotator [Wyman et al. 2004]; http://www.evogen.jgi-psf.org/dogma) based on the similarity of their nucleotide or inferred amino acid sequences to a curated database of 20 previously published chloroplast genomes. Genes for tRNAs and rRNAs were located by BlastN searches of the same database. Relative gene content and sequence divergence between Trachelium and 10 other angiosperms (Amborella trichopoda [NC_005086], Arabidopsis thaliana [NC_000932], Calycanthus floridus var. glaucus [NC_004993], Jasminum nudiflorum [NC_008407], Lotus corniculatus var. japonicus [NC_002694], Nicotiana tabacum [NC_001879], Nymphaea alba [NC_006050], Pelargonium x hortorum [NC_008454], Spinacia oleracea [NC_002202], and Zea mays [NC_001666]) were visualized using MultiPipMaker (Schwartz et al. 2003).
Sizes and locations of direct and inverted repeats in the Trachelium chloroplast genome were determined by running REPuter (Kutrz et al. 2001) at a repeat length ≥30 bp with a Hamming distance of 3. REPuter was run using the entire genome in order to map repeats in both copies of the IR, but numbers of repeats were based on results from only one IR copy. Repeats were mapped onto the Trachelium chloroplast genome, and those located at or near inversion endpoints and other sites of rearrangement were characterized by BlastN searches in GenBank. We ran the same REPuter analyses against the 10 angiosperm chloroplast genomes that were used for MultiPipMaker to assess the relative number of repeats in chloroplast genomes. BlastN searches of intergenic regions between blocks of inverted gene sequences in Trachelium were performed against GenBank.
Results
Organization of the Trachelium Chloroplast Genome
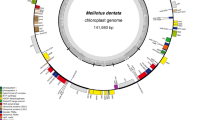
The complete chloroplast genome sequence of Trachelium caeruleum (GenBank: EU_090187) is 162,321 bp, with an IR of 27,273 bp separating a LSC region of 100,114 bp and a SSC region of 7661 bp (Fig. 1). We confirmed the contraction of the IR boundary with the LSC region and expansion of the IR into the SSC region, previously described by Cosner et al. (1997). The G + C content is 38.3%; within coding regions it is 40.59%, and in noncoding regions it is 35.6%. Coding regions comprise 59.67% of the genome.
Map of the complete chloroplast genome of Trachelium caeruleum represented as a circular molecule. Outer circle of numbered arrows indicates conserved blocks of genes relative to an unrearranged genome such as Nicotiana. In Nicotiana genes are numbered consecutively 1–116, but in Trachelium they have been rearranged in location and/or orientation. Genes shown on the inside of the middle circle are transcribed clockwise, while genes on the outside are transcribed counterclockwise. Asterisks by gene names indicate truncation. Small arrows perpendicular to gene block arrows show locations of tRNA genes in relation to rearrangements
The Trachelium chloroplast genome includes 132 genes, and their relative locations are mapped in Fig. 1. These include 17 that are duplicated in the IR, plus 1 (trnI-cau; gene 87) duplicated once in the LSC region and another (psbJ; gene 55) with two additional copies in the LSC region. Expansion of the IR into the SSC region caused the duplication of ndhE, ndhG, ndhI, ndhA, ndhH, rps15, and ycf1. The conserved open reading frame ycf2 that normally occurs in the IR is single copy in Trachelium and located in the LSC region, due to contraction of the IR. Trachelium has 71 different intact protein-coding genes of known function, 4 ycfs, 4 rRNAs, and 30 tRNAs; unlike most land-plant chloroplast genomes it has a number of partial or entire gene duplications and several truncated or otherwise altered genes that are likely pseudogenes. Seventeen genes contain introns; the intron commonly present in rps16 is absent in Trachelium. Whole-genome alignment of the Trachelium chloroplast genome with 10 other angiosperms shows high conservation of many coding regions as well as marked divergence in others (Fig. 2). In Trachelium, four genes or ycfs are abbreviated and presumably nonfunctional: (1) ycf15 is truncated to include only 191 bp of the 5’ end; (2) only 50 bp of the 3’ end of rpl23 exists, and this occurs at an inversion endpoint in the LSC region, with a 34-bp repeat of part of this fragment at another inversion endpoint within the LSC region; (3) infA is reduced to a fragment consisting of 191 bp of the middle of the gene, lacking both the 5’ and the 3’ ends; and (4) only a 290-bp fragment of accD exists, embedded in the highly diverged ycf1 gene in the IR in the vicinity of a number of other rearrangements. ndhK may be a pseudogene, containing multiple internal stop codons generated by a single deletion causing a frameshift and several additional indels. Multiple genome alignment shows that three other genes, clpP, ycf1, and ycf2, have diverged greatly (percentage similarity shown in Fig. 3) from most other angiosperms examined, especially those with genomes that are not rearranged (all except Jasminum, Pelargonium, and Zea). These eight reduced or altered genes align with intact copies of these genes in other angiosperm chloroplast genomes.
Whole-genome alignment of the Trachelium chloroplast genome with 10 other angiosperm chloroplast genomes using Nicotiana as the reference genome, generated by MultiPipMaker (Schwartz et al. 2003). The top line indicates genes in Nicotiana, and their relative position in the Nicotiana sequence is indicated by the scale at the bottom. Sequence identity of both genes and intergenic spacers to other genomes is shown by dark gray (75–100%), light gray (50–75%), and white (<50%). Arrows show relative locations of selected divergent or altered genes in Trachelium
Closeup of MultiPipMaker alignment of selected regions discussed in the text. Nicotiana serves as the reference chloroplast genome, and heavy black bars represent its exons. Aligned regions are illustrated with horizontal lines drawn indicating percentage similarity between 50% and 100% (scale to right of Amborella panel)
We compared the gene order of Nicotiana, which has the ancestral angiosperm gene order (Raubeson et al. 2007), to that of Trachelium. We found 18 conserved blocks of genes in Trachelium in which the genes in each block are in the same gene order as Nicotiana but the blocks have been rearranged relative to their order in the Nicotiana chloroplast genome (Fig. 1). These blocks of genes ranged in size from 4 to 17 kb. The gene order in Trachelium is further altered by the insertion of entire genes or gene fragments from other parts of the genome between and within a number of these otherwise conserved blocks of genes.
Location of tRNA Genes in Relation to Rearrangements
tRNA genes are associated with rearrangements in 10 locations (Fig. 1; arrows). They occur at the ends of conserved gene blocks at four locations in the LSC region (trnT-ugu, trnM-cau, trnC-gca, and one copy of trnI-cau) and two locations in the IR and are hence duplicated (trnL-caa and trnN-guu). In two other cases, a tRNA gene has been relocated to a position between two conserved gene blocks. The second copy of trnI-cau (gene 87) occurs in the LSC region between conserved blocks 39–46 and 35–20; trnV-gac (gene 94) is moved from its normal IR location to the LSC region between gene blocks 86–69 and 66–55.
Repeats in the Trachelium Chloroplast Genome
All 11 genomes analyzed have multiple repeats, many of which are mono- or dinucleotide strings (Fig. 4). The highly rearranged genomes of Trachelium and Pelargonium and the moderately rearranged Jasminum chloroplast genome have the highest number of repeats and the largest repeats among all genomes compared, suggesting a positive correlation between the number of repeats and genomic rearrangements in these genomes.
Direct and inverted repeat size and frequency in Trachelium and 10 other angiosperm chloroplast genomes identified with REPuter (Kurtz et al. 2001) at a repeat length ≥30 bp with a Hamming distance of 3. Vertical bars represent repeats clustered in classes of 30–39, 40–49, 50–75, 76–200, and 201–3500
In Trachelium, many repeat elements were found at some but not all inversion endpoints and at or near other rearrangements, such as gene duplications. The length, orientation, and coordinates for these repeats were pinpointed and mapped (Fig. 5; middle circle). A total of 767 direct and inverted repeats ≥30 bp was identified in Trachelium, of which 483 were direct and 284 were inverted repeats. Three hundred three repeats occurred either as parts of genes or in intergenic spacers within conserved blocks of genes. Four hundred sixty-four repeats occurred either between inverted gene blocks or near other rearrangements, suggesting a strong association between repeats and rearrangements. BlastN searches against GenBank showed that repeat elements at or near inversion endpoints are derived from protein-coding regions within the chloroplast genome (i.e., partial to entire gene duplications; discussed below), a tRNA gene (trnI-cau), noncoding cpDNA, and novel DNA not previously identified as being chloroplast in origin (Table 1).
Trachelium chloroplast genome map showing location of repeats ≥30 bp with a Hamming distance of 3, identified by REPuter, in relation to blocks of rearranged genes. The outer circle of numbered arrows identifies conserved gene blocks and their orientation in relation to Nicotiana. The circle of hashmarks indicates the location of 767 direct and inverted repeats reported by REPuter. Numbered arrows, r1–r12, identify selected large repeats of diverse origin, characterized in Table 1. Shaded areas A, B, and C show rearrangement hotspots
Inversion Endpoints as Hotspots for Rearrangements and Repeats
Multiple rearrangements and repeats of diverse origin are concentrated between inverted blocks of genes in the Trachelium chloroplast genome. For example, between conserved blocks of genes 86–69 and 66–55 in the LSC region (Fig. 5; shaded area A), a 2.7-kb segment of normally unassociated sequence is found together in the space between psbB (gene 69) and rpl20 (gene 66) where genes 68 (clpP) and 67 (5’rps12) would typically be found. trnV-gac (gene 94), which is normally found within the IR, has moved into this endpoint in the LSC region. Additionally, a duplicate, presumably functional copy of psbJ (gene 55) is also inserted into this area (Fig. 5; r6). Finally, repeats of noncoding cpDNA sequences from different areas of the genome are located within this hotspot between psbB and rpl20, flanking trnV-gac and psbJ (Fig. 5; r4, r7).
Another inversion endpoint with high sequence complexity occurs between gene blocks 66–55 and 39–46 (Fig. 5; shaded area B). The second copies of r4 and r5 (Fig. 5) occur here along with the copy of psbJ (Fig. 5, r6), which is the original copy occurring in its operon with psbF, psbE, and psbL. A 105-bp repeat of noncoding chloroplast DNA sequence (Fig. 5; r7) is shared with the second copy of psbJ between gene blocks 86–69 and 66–55 but not with the third copy of psbJ located within block 35–20. Finally, a small repeat of part of the clpP exon 1 is located here (Fig. 5; r8). The entire, presumably functional copy of clpP is located in the IR (see below).
The most complex rearrangements in the Trachelium chloroplast genome occur within the IR in association with multiple repeats (Fig. 5; shaded area C). A 4.6-kb portion of sequence normally found in the LSC region as well as several smaller duplicated sequences from the LSC region and the IR are inserted into one heterogeneous area between gene blocks 95–102 and 116–110 (Fig. 6). The first two genes of the clpP operon, clpP and 5’rps12 (genes 68 and 67, respectively), were moved here in their entirety, and a 99.7% identical 1014-bp repeat of sequence normally found adjacent to the start of this operon was duplicated and moved as well (r3). This repeat includes an identical copy of the first 300 bp of psbB (gene 69) and an intergenic spacer between the functional copy of psbB and trnV-gac at the 86–69/66–55 inversion endpoint in the LSC region. Exon 1 of clpP contains a large insertion. Found within the first intron in clpP is repeat r4 (Fig. 5), which is also found in the LSC region in two places (Fig. 5; shaded areas A and B). clpP and 5’rps12 are separated by insertions of part of the third exon of ycf3 (r10) and a 457-bp repeat of noncoding sequence (r11) from the vicinity of the functional copy of ycf3 within the LSC region. Immediately adjacent to this area is a very divergent copy of ycf1 (gene 116), into which a 290-bp vestige of accD (gene 50) is inserted. A 99.6% identical 487-bp repeat of the 5’ end of the 23S rrn gene (r12) is found between ycf1 and rps15 (gene 115).
Detailed view of shaded area C in Fig. 5, in inverted repeat (IR). Asterisks indicate gene fragments. Genes numbered 69, 36, 50, and 98 reflect partial duplications of genes; entire, functional copies of genes 69 (psbB) and 36 (ycf3) are located in the large single-copy (LSC) region. accD (gene 50) is normally found between rbcL and psaI in the LSC region
Discussion
Genome Organization
The complete Trachelium chloroplast genome sequence is far more complex than originally described based on restriction site and gene mapping (Cosner et al. 1997). Although other genomes have been identified as having multiple rearrangements, the Trachelium genome shows a unique combination and number of genome rearrangements, including partial to entire gene duplications, several gene reductions, intron loss, numerous large inversions, and a concentration of repeats and tRNA genes at or near inversion endpoints.
Gene duplications are infrequently reported for chloroplast genomes. The psbA duplication in some ferns (Stein et al. 1992) and numerous duplications of normally single copy genes in Pelargonium (Palmer et al. 1987; Chumley et al. 2006) have been attributed primarily to expansion of the IR. Wolfe (1988) suggested that the partial duplications of rbcL and psbA in Pisum are associated with loss of one copy of the IR; this was recently supported by the findings of Saski et al. (2005) as having occurred simultaneous to the loss of the IR in the entire clade of legumes that includes Pisum. The duplication of psaM and several tRNA genes in black pine may be due to the inherent instability caused by severe reduction of the IR (Wakasugi et al. 1994; Hipkins et al. 1995). The presence of three complete copies of psbJ in the LSC region of Trachelium is distinctive and unlikely to be explained solely by inversions or expansion/contraction of the IR. Although one of the psbJ duplications occurs within an inversion endpoint between conserved blocks of genes (shaded area A; Fig. 1), the other duplicated copy is found within an otherwise unrearranged block of genes (block 35–20). This suggests that some mechanism other than inversion or IR boundary changes may be responsible, perhaps a duplicative transposition, which has been suggested in the generation of dispersed repeats in conifers (Tsai and Strauss 1989) and subclover (Milligan et al. 1989). There is no direct evidence of transposable elements within the Trachelium genome, although they may have been present transiently. Whatever the mechanism, the two duplications must have occurred relatively recently, as they have 100% sequence identity to the original copy.
Duplications of tRNA genes have recently been reported in otherwise relatively unrearranged chloroplast genomes, for example, in those of Jasminum and close relatives in the Oleaceae (Lee et al. 2007) and Arabidopsis and other Brassicaceae (Koch et al. 2005). Partial duplications of tRNA genes have been reported in taxa known for rearranged chloroplast genomes, for example, grasses (Hiratsuka et al. 1989) and conifers (Tsai and Strauss 1989). In the second case of gene duplication in Trachelium, the extra copy of trnI-cau occurs between two conserved gene blocks inverted in relation to each other. One explanation for the duplication is that strand repair following a series of inversions may have generated the duplicate copy of trnI-cau, which was later moved to its present location. An alternative hypothesis for the duplication in Trachelium entails generation of a tandem repeat of trnI-cau by expansion and contraction of the IR that was subsequently moved during the course of inversions. Whether the duplication is responsible for the inversion due to nonhomolgous recombination between one of the adjacent tRNA genes and the original copy of trnI-cau or is the result of an error in the repair of a double strand break cannot be determined from these data alone. There is only a single-base pair difference between the copies.
Another striking feature of the Trachelium genome is the partial loss of four genes: ycf15, rpl23, infA, and accD. These four genes have been lost or altered in other chloroplast genomes but not all four in the same genome. The sequence of ycf15 has been shown to be variable among angiosperm chloroplast genomes, with conserved motifs at the 5’ and 3’ ends and an intervening 250 bp in some taxa that renders it a pseudogene (Schmitz-Linneweber et al. 2001; Steane 2005). A comparative study of ycf15 transcripts in taxa with or without the insertion suggests that this may not be a functional protein-coding gene even when intact (Schmitz-Linneweber et al. 2001), and recent comparisons of sequence evolution of ycf15 among angiosperms confirm this observation (Raubeson et al. 2007). rpl23 is a pseudogene in spinach (Thomas et al. 1988; Schmitz-Linneweber et al. 2001) and a pseudogene copy persists in grasses in the LSC region, with an intact copy in the IR situated in the site of the lost accD gene between rbcL and psaI (Morton and Clegg 1993). A very similar duplication occurs in certain members of Jasminum, and is inserted in the same region in the genome as in Poaceae (Lee et al. 2007). In Trachelium, rpl23 is neatly severed after 50 bp of well-conserved sequence; the truncation may have occurred as the result of an inversion and/or in a process involving recombination between repeats, as there is a partial repeat of this fragment elsewhere in the LSC region in the vicinity of rbcL and psaI that are now separated by multiple inversions. Millen et al. (2001) found 24 independent losses or reduction of infA in a survey of 308 angiosperms, including Campanula, Trachelium, and Platycodon (Campanulaceae) and 2 members of their sister family, Lobeliaceae, but the gene is present in other members of the Asterales. This indicates that within the Asterales, the loss or reduction of infA occurred in the recent common ancestor of the Campanulaceae/Lobeliaceae clade. Our sequence of infA in Trachelium confirms the earlier evidence found in Southern hybridization data that the gene is reduced in size. Earlier mapping studies of Trachelium and other members of the Campanulaceae and Lobeliaceae reported that accD is absent in both families, and this is a synapamorphy supporting their sister relationship (Downie and Palmer 1992; Cosner et al. 1997; Knox and Palmer 1999); however, we discovered a vestige of this gene in the Trachelium genome sequence. accD is also lost in the Poaceae and close relatives (Hiratasuka et al.1989; Downie and Palmer 1992; Maier et al. 1995; Katayama and Ogihara 1996; Ogihara et al. 2002) from a hotspot between rbcL and psaI into which an rpl23 pseudogene has been inserted.
RNA editing has been reported in NADH dehydrogenase (ndh) genes in a number of angiosperm chloroplast genomes (Maier et al. 1995; Hirose et al. 1999; Fiebig et al. 2004). In Trachelium a single-base pair deletion in ndhK causes a frameshift. Although insertion/deletion editing has been detected in mitochondrial genomes (e.g., Simpson et al. 2003), we are not aware of any evidence for this process in chloroplast genomes. Other cases of lost functional ndh genes in chloroplast genomes include Pinus thunbergii (Wakasugi et al. 1994), Epifagus virginiana (Wolfe et al. 1992), and Phalaenopsis aphrodite (Chang et al. 2006).
Large Inversions and the Evolutionary Influence of Repeats and tRNA Genes
In Trachelium, the most conspicuous alterations in the chloroplast genome are its large (>4 kb), multiple inversions and relocation of blocks of genes. Trachelium also has more and larger repeats than most other angiosperm chloroplast genomes (Fig. 4). It has a concentration of repeats of diverse origin at or near these inversion endpoints and other rearrangements, such as the cluster of rearrangements within the IR. With few exceptions, the repeats are direct repeats, not the inverted repeats one would expect to be associated with inversions (Palmer 1991). It is possible that short inverted repeats were responsible for some inversions in the Trachelium genome, but were subsequently reoriented to direct repeats as a result of additional inversions, or have diverged or been eliminated over time. Our parameters for repeat searches were quite stringent at ≥30 bp and a Hamming distance of 3. Less stringent searches yield many more repeats: with a 20-bp window and a Hamming distance of 4, we found >30,000 repeats (data not shown).
Comparison of the size and number of repeats in Trachelium to those in 10 other angiosperm chloroplast genomes reveals that there is a modest background of repeats even in unrearranged chloroplast genomes. Polymorphic, simple sequence repeats (SSRs) <15 bp have been identified in many chloroplast sequences and in all completely sequenced land plant chloroplast genomes (Marshall et al. 2001; Provan et al. 2001; Raubeson et al. 2007). Short dispersed repeats have been associated with inversion endpoints and occur in a number of taxa (in Pelargonium [Palmer et al.1987; Chumley et al. 2006], wheat [Howe 1985; Quigley and Weil 1985; Bowman and Dyer 1986; Bowman et al. 1988; Ogihara et al. 1988], rice [Shimada and Sugiura 1989], subclover [Milligan et al. 1989], Douglas fir [Tsai and Strauss 1989], Asteraceae [Kim et al. 2005; Timme et al. 2007], Oleaceae [Lee et al. 2007]). A recent comparison of four chlorophyte algal chloroplast genomes showed a strong correlation between the number of repeats in the chloroplast genome and the degree of rearrangement (Pombert et al. 2005, 2006). The most highly rearranged green algal chloroplast genome is Chlamydomonas reinhardtii (Maul et al. 2002), which also has the greatest number of repeats in its lineage (Pombert et al. 2005).
Even in unrearranged chloroplast genomes, small inversions regularly occur in intergenic areas, caused by short (11- to 24-bp) inverted repeats forming hairpins that can easily flip-flop the orientation of the intervening sequences (Kelchner and Wendel 1996; Kelchner 2000;, Kim and Lee 2005). Larger (>200-bp) inversions are found in some angiosperm chloroplast genomes, but generally not more than a few within a genome. A number of possible mechanisms have been proposed for these events. Inversions may occur in a specific location due to the presence of short repeat elements subject to homologous recombination (Palmer 1991; Knox et al. 1993). In the grasses, with three large inversions, repeats flank the borders of a 28-kb inversion and may have facilitated the inversion or nonhomologous recombination between tRNA genes with high sequence similarity may have caused the rearrangement (Hiratsuka et al. 1989; Sugiura 1989). A 54-kb inversion in Oenothera elata has a series of small inverted repeats at each end (Hupfer et al. 2000). In the Ranunculaceae, Hoot and Palmer (1994) found up to six inversions in certain taxa, ranging in size from 5.6 to 53.6 kb. They proposed that some inversions might have positioned repeat sequences in a way that would cause subsequent inversions. They also noted the similarity of inversion endpoints in Anenome to those reported by Knox et al. (1993) in other rearranged chloroplast genomes of lineages distantly related to the Ranunculaceae, including the Lobeliaceae, which is sister to the Campanulaceae.
Chloroplast genome inversions have also been attributed to nonhomologous recombination between different tRNA genes (Knox et al. 1993; Hoot and Palmer 1994). A 20-kb inversion in rice and the generation of a tRNA pseudogene were attributed to recombination between two different tRNA genes (Hiratsuka et al. 1989). tRNA genes are present at or near 12 of 17 inversion endpoints in the highly rearranged chloroplast genome of the charophyte Chaetospheridium globosum, and it was suggested that inversions may be due to the presence of short direct repeats within or near the tRNA genes (Turmel et al. 2002). In Trachelium, tRNA genes may be implicated in some inversions because there are tRNA genes at the ends of 10 of 18 rearranged blocks of genes (Fig. 1).
The most rearranged region in the Trachelium genome occurs in the IR, where within only 12.5 kb, there are two partial gene duplications of genes that remain intact in the LSC region, a partial duplication of 23Srrn, and relocation of clpP and 5’rps12 from the LSC region, with a remnant of accD nearby within the highly divergent gene ycf1. Although a series of inversions might explain the relocation of these coding sequences into the IR, selection is believed to constrain against inversions between the LSC region and the IR, and within operons and genes (Palmer 1991). Loss of one copy of the IR in an ancestor to Trachelium would have eliminated the constraint against LSC/IR inversions, but gene mapping data of other Campanulaceae show this to be unlikely (Cosner 1993; Cosner et al. 1997). Unlike Trachelium, several other Campanulaceae lack the clpP-5’rps12 relocation to the IR. These taxa have the IR/SSC region boundary that characterizes the Campanulaceae/Lobeliaceae clade, and are basal to Trachelium. Together, this suggests that the transfer of these genes into the IR occurred after establishment of the IR/SSC region boundary in the most recent common ancestor to Campanulaceae/Lobeliaceae. Transposition may be the most parsimonious explanation for how these genes and gene fragments became concentrated into this one area. The sole known example of a transposon in a chloroplast genome occurs in the highly rearranged cpDNA of the green alga Chlamydomonas reinhardtii, which had two copies of a disabled transposable element, Wendy (Fan et al. 1995). Although no transposon-like element was found in the Trachelium chloroplast genome through BlastN searches, one may have been present in a Campanulaceae ancestor, generated genome instability, and been expunged.
These explanations for complex genome rearrangements are predicated on the idea of a circular genome, which replicates in a manner that maintains the integrity of the genome (Kolodner and Tewari 1975). Recent fluorescent microscopy studies show that chloroplast genomes may exist at least part of the time as multigenomic, branched structures or as linear strands (Oldenburg and Bendich 2004a, b). The presence of even transient single strands or dimers would increase the possibility of inter- and intramolecular recombination, but how chloroplast genomes persist as stable and conservatively evolving units is unclear, if this scenario is accurate.
The evidence for the mechanisms responsible for structural rearrangements in the Trachelium chloroplast genome may have been lost over evolutionary time. The genome has such a high incidence of rearrangements and an accumulation of repeats that something must have happened within this lineage that has made it susceptible to instability. Earlier mapping studies found at least 42 inversions in 18 Campanulaceae chloroplast genomes, at least 8 possible transpositions, and multiple different IR expansions/contractions (Cosner 1993; Cosner et al. 2004). Preliminary analysis of the draft sequences of seven other Campanulaceae chloroplast genomes (R. Haberle and R. Jansen, unpublished) suggests that there are many other rearrangements in these genomes and that many of these are also associated with repeats.
Trachelium’s unique combination of inversions, the proliferation of repeats, and multiple cases of gene loss or deterioration suggest an inherent instability in this genome. Completion of the Trachelium chloroplast genome sequence raises many questions that can be best addressed through comparative analysis with complete genome sequences of other, closely related taxa. Rates of structural or nucleotide substitutions in this group may be faster than in groups with unrearranged chloroplast genomes, and they may have an accelerated rate of transfer of genes to the nucleus. If they share partial loss or deterioration of the same genes, likely there are stages of these changes apparent among these relatives that may help clarify the order and nature of these events. If the repeats in other members of the Campanulaceae are associated with the same repeats and rearrangements as in Trachelium, this may support the role of repeats in contributing to certain rearrangements. Using tools of comparative chloroplast genomics may reveal clues to the underlying causes of structural evolution in this unusual group that would be obscured over time in more distantly related taxa.
References
Achaz G, Coissac E, Netter P, Rocha EPC (2003) Associations between inverted repeats and the structural evolution of bacterial genomes. Genetics 164:1279–1289
Bowman CM, Barker RF, Dyer TA (1988) In wheat ctDNA, segments of ribosomal-protein genes are dispersed repeats, probably conserved by nonreciprocal recombination. Curr Genet 14:127–136
Bowman CM, Dyer TA (1986) The location and possible evolutionary significance of small dispersed repeats in wheat ctDNA. Curr Genet 10:931–941
Chang CC, Lin HC, Lin IP, Chow TY, Chen HH, Chen WH, Cheng CH, Lin CY, Lieu SM, Chang CC Chaw SM (2006) The chloroplast genome of Phalaenopsis aphrodite Orchidaceae: comparative analysis of evolutionary rate with that of grasses and its phylogenetic implications. Mol Biol Evol 23:279–291
Chumley TW, Palmer JD, Mower JP, Fourcade HM, Callie PJ, Boore JL, Jansen RK (2006) The complete chloroplast genome sequence of Pelargonium × hortorum: organization and evolution of the largest and most highly rearranged chloroplast genome of land plants. Mol Biol Evol 23:2175–2190
Cosner ME (1993) Phylogenetic and molecular evolutionary studies of chloroplast DNA variation in the Campanulaceae. Ph.D. thesis. The Ohio State University, Columbus
Cosner ME, Jansen RK, Palmer JD, Downie SR (1997) The highly rearranged chloroplast genome of Trachelium caeruleum Campanulaceae: multiple inversions, inverted repeat expansion and contraction, transposition, insertions/deletions, and several repeat families. Curr Genet 31:419–429
Cosner ME, Raubeson LA, Jansen RK (2004) Chloroplast DNA rearrangements in Campanulaceae: phylogenetic utility of highly rearranged genomes. BMC Evol Biol 4:27
Downie SR, Palmer JD (1992) Use of chloroplast DNA rearrangements in reconstructing plant phylogeny. In: Soltis PS, Soltis DE, Doyle JJ (eds) Molecular systematics of plants. Chapman and Hall, London, UK, pp 14–35
Ewing B, Green P (1998) Base-calling of automated sequencer traces using phred. II. error probabilities. Genome Res 8:186–194
Fan WH, Woelfle MA, Mosig G (1995) Two copies of a DNA element, Wendy, in the chloroplast chromosome of Chlamydomonas-reinhardtii between rearranged gene clusters. Plant Mol Biol 29:63–80
Fiebig A, Stegemann S, Bock R (2004) Rapid evolution of RNA editing sites in a small non-essential plastid gene. Nucleic Acids Res 32:3615–3622
Gordon D, Abajian C, Green P (1998) Consed: a graphical tool for sequence finishing. Genome Res 8:195–202
Hipkins VD, Marshall KA, Neale DB, Rottmann WH, Strauss SH (1995) A mutation hotspot in the chloroplast genome of a conifer Douglas-fir, Pseudotsuga is caused by variability in the number of direct repeats derived from a partially duplicated transfer-RNA gene. Curr Genet 27:572–579
Hiratsuka J, Shimada H, Whittier R, Ishibashi T, Sakamoto M, Mori M, Kondo C, Honji Y, Sun CR, Meng BY, Li YQ, Kanno A, Nishizawa Y, Hirai A, Shinozaki K, Sugiura M (1989) The complete sequence of the rice Oryza sativa chloroplast genome—intermolecular recombination between distinct transfer-RNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol Gen Genet 217:185–194
Hirose T, Kusemegi T, Tsudzuki T, Sugiura M (1999) RNA editing sites in tobacco chloroplast transcripts: editing as a possible regulator of chloroplast RNA polymerase activity. Mol Gen Genet 262:452–467
Hoot SB, Palmer JD (1994) Structural rearrangements, including parallel inversions, within the chloroplast genome of Anemone and related genera. J Mol Evol 383:274–281
Howe CJ (1985) The endpoints of an inversion in wheat chloroplast DNA are associated with short repeated sequences containing homology to att-lambda. Curr Genet 102:139–145
Hupfer H, Swiatek M, Hornung S, Herrmann RG, Maier RM, Chiu WL, Sears B (2000) Complete nucleotide sequence of the Oenothera elata plastid chromosome, representing plastome I of the five distinguishable Euoenothera plastomes. Mol Gen Genet 263:581–585
Jansen RK, Raubeson LA, Boore JL, dePamphilis CW, Chumley TW, Haberle RC, Wyman SK, Alverson A, Peery R, Herman SJ, Fourcade HM, Kuehl JV, McNeal JR, Leebens-Mack J, Cui L (2005) Methods for obtaining and analyzing whole chloroplast genome sequences. Molecular evolution: producing the biochemical data, part B. Methods Enzymol 395:348–384
Katayama H, Ogihara Y (1996) Phylogenetic affinities of the grasses to other monocots as revealed by molecular analysis of chloroplast DNA. Curr Genet 296:572–581
Kelchner SA (2000) The evolution of non-coding chloroplast DNA and its application in plant systematics. Ann Mo Bot Gard 87:482–498
Kelchner SA, Wendel JF (1996) Hairpins create minute inversions in non-coding regions of chloroplast DNA. Curr Genet 303:259–262
Kim K-J, Choi KS, Jansen RK (2005) Two chloroplast DNA inversions originated simultaneously during early evolution in the sunflower family. Mol Biol Evol 22:1783–1792
Kim K-J, Lee HL (2005) Widespread occurrence of small inversions in the chloroplast genomes of land plants. Mol Cells 19:104–113
Knox EB, Palmer JD (1999) The chloroplast genome arrangement of Lobelia thuliniana Lobeliaceae: expansion of the inverted repeat in an ancestor of the Campanulales. Plant Syst Evol 214:49–64
Knox EB, Downie SR, Palmer JD (1993) Chloroplast genome rearrangements and the evolution of giant lobelias from herbaceous ancestors. Mol Biol Evol 10:414–430
Koch MA, Dobes C, Matschinger M, Bleeker W, Vogel J, Kiefer J, Mitchell-Olds T (2005) Evolution of the trnF-gaa gene in Arabidopsis relatives and the Brassicaceae family: monophyletic origin and subsequent diversification of a plastid pseudogene. Mol Biol Evol 22:1032–1043
Kolodner R, Tewari KK (1975) Chloroplast DNA from higher-plants replicates by both cairns and rolling circle mechanism. Nature 256:708–711
Kurtz S, Choudhuri JV, Ohlebusch E, Schleiermacher C, Stoye J, Giegerich R (2001) REPuter: the manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res 29:4633–4642
Lee H-L, Jansen RK, Chumley TW, Kim K-J (2007) Gene relocations within chloroplast genomes of Jasminum and Menodora (Oleaceae) are due to multiple, overlapping inversions. Mol Biol Evol 24:1161–1180
Maier RM, Neckermann K, Igloi GL, Kossel H (1995) Complete sequence of the maize chloroplast genome-gene content, hotspots of divergence and fine-tuning of genetic information by transcript editing. J Mol Biol 251:614–628
Marshall HD, Newton C, Ritland K (2001) Sequence-repeat polymorphisms exhibit the signature of recombination in lodgepole pine chloroplast DNA. Mol Biol Evol 11:2136–2138
Maul JE, Lilly JW, Cui L, dePamphilis CW, Miller W, Harris EH, Stern DB (2002) The Chlamydomonas reinhardtii plastid chromosome: islands of genes in a sea of repeats. Plant Cell 14:2659–2679
Millen RS, Olmstead RG, Adams KL, Palmer JD, Lao NT, Heggie L, Kavanagh TA, Hibberd JM, Giray JC, Morden CW, Calie PJ, Jermiin LS, Wolfe KH (2001) Many parallel losses of infA from chloroplast DNA during angiosperm evolution with multiple independent transfers to the nucleus. Plant Cell 13:645–658
Milligan BG, Hampton JN, Palmer JD (1989) Dispersed repeats and structural reorganization in subclover chloroplast DNA. Mol Biol Evol 6:355–368
Morton BR, Clegg MT (1993) A chloroplast DNA mutational hotspot and gene conversion in a non-coding region near rbcL in the grass family Poaceae. Curr Genet 244:357–365
Ogihara Y, Isono K, Kojima T, Endo A, Hanaoka M, Shiina T, Terachi T, Utsugi S, Murata M, Mori N, Takumi S, Ikeo K, Gojobori T, Murai R, Murai K, Matsuoka Y, Ohnishi Y, Tajiri H, Tsunewaki K (2002) Structural features of a wheat plastome as revealed by complete sequencing of chloroplast DNA. Mol Gen Genom 266:740–746
Ogihara Y, Terachi T, Sasakuma T (1988) Intramolecular recombination of chloroplast genome mediated by short direct-repeat sequences in wheat species. Proc Natl Acad Sci USA 85:8573–8577
Oldenburg DJ, Bendich AJ (2004a) Most chloroplast DNA of maize seedlings in linear molecules with defined ends and branched forms. J Mol Biol 335:953–970
Oldenburg DJ, Bendich AJ (2004b) Changes in the structure of DNA molecules and the amount of DNA per plastid during chloroplast development in maize. J Mol Biol 344:1311–1330
Palmer JD (1986) Isolation and structural analysis of chloroplast DNA. Methods Enzymol 118:167–186
Palmer JD (1991) Plastid chromosomes: structure and evolution. In: Bogorad L (ed) Molecular biology of plastids. Academic Press, San Diego, CA, pp 5–53
Palmer JD, Nugent JM, Herbon LA (1987) Unusual structure of geranium chloroplast DNA—a triple-sized inverted repeat, extensive gene duplications, multiple inversions, and two repeat families. Proc Natl Acad Sci USA 843:769–773
Pombert J-F, Lemieux C, Turmel M (2006) The complete chloroplast DNA sequence of the green alga Oltmannsiellopsis viridis reveals a distinctive quadripartite architecture in the chloroplast genome of early diverging ulvophytes. BMC Biol 4:3
Pombert J-F, Otis C, Lemieux C, Turmel M (2005) The chloroplast genome sequence of the green alga Pseudendoclonium akinetum Ulvophyceae reveals unusual structural features and new insights into the branching order of chlorophyte lineages. Mol Biol Evol 22:1903–1918
Provan J, Powell W, Hollingsworth PM (2001) Chloroplast microsatellites: new tools for studies in plant ecology and evolution. Trends Ecol Evol 16:142–147
Quigley F, Weil JH (1985) Organization and sequence of 5 transfer-RNA genes and of an unidentified reading frame in the wheat chloroplast genome—evidence for gene rearrangements during the evolution of chloroplast genomes. Curr Genet 96:495–503
Raubeson LA, Jansen RK (1992) Chloroplast DNA evidence on the ancient evolutionary split in vascular land plants. Science 255:1697–1699
Raubeson LA, Jansen RK (2005) Chloroplast genomes of plants. In: Henry RJ (ed) Plant diversity and evolution: genotypic and phenotypic variation in higher plants. CABI, Cambridge, MA, pp 45–68
Raubeson LA, Peery R, Chumley T, Dziubek C, Fourcade HM, Boore JL, Jansen RK (2007) Comparative chloroplast genomics: Analyses including new sequences from the angiosperms Nuphar advena and Ranunculus macranthus. BMC Genom 8:174
Rocha EPC (2003) DNA repeats lead to the accelerated loss of gene order in bacteria. Trends Genet 19:600–603
Saski C, Lee S-B, Daniell H, Wood TC, Tomkins J, Kim H-G, Jansen RK (2005) Complete chloroplast genome sequence of Glycine max and comparative analyses with other legume genomes. Plant Mol Biol 59:309–322
Schmitz-Linneweber C, Maier RM, Alcaraz J-P, Cottet A, Herrmann RG, Mache R (2001) The plastid chromosome of spinach Spinacia oleracea: complete nucleotide sequence and gene organization. Plant Mol Biol 45:307–315
Schwartz S, Elnitski L, Li M, Weirauch M, Riemer C, Smit A, Program NCS, Green ED, Hardison RC, Miller W (2003) MultiPipMaker and supporting tools: alignments and analysis of multiple genomic DNA sequences. Nucleic Acids Res 31:3518–3524
Shimada H, Sugiura M (1989) Pseudogenes and short repeated sequences in the rice chloroplast genome. Curr Genet 164:293–301
Simpson L, Sbicego S, Aphsizhev R (2003) Uridine insertion/deletion RNA editing in trypanosome mitochondria: a complex business. RNA 9:265–276
Steane DA (2005) Complete nucleotide sequence of the chloroplast genome from the Tasmanian blue gum, Eucalyptus globules Myrtacaceae. DNA Res 12:215–220
Stein DB, Conant DS, Ahearn ME, Jordan ET, Kirch SA, Hasebe M, Iwatsuki K, Tan MK, Thomson JA (1992) Structural rearrangements of the chloroplast genome provide an important phylogenetic link in ferns. Proc Natl Acad Sci USA 89:1856–1860
Sugiura M (1989) The chloroplast chromosomes in land plants. Annu Rev Cell Biol 5:51–70
Thomas F, Massenet O, Dorne AM, Briat JF, Mache R (1988) Expression of the rp123, rp12 and rps19 genes in spinach chloroplasts. Nucleic Acids Res 16:2461–2472
Timme RE, Kuehl JV, Boore JL, Jansen RK (2007) A comparison of the first two sequenced chloroplast genomes in Asteraceae: lettuce and sunflower. Am J Bot 94:302–312
Tsai CH, Strauss SH (1989) Dispersed repetitive sequences in the chloroplast genome of Douglas-fir. Curr Genet 163:211–218
Turmel M, Otis C, Lemieux C (2002) The chloroplast and mitochondrial genome sequences of the charophyte Chaetosphaeridium globosum: insights into the timing of the events that restructured organelle DNAs within the green algal lineage that led to land plants. Proc Natl Acad Sci USA 99:11275–11280
Wakasugi T, Tsudzuki J, Ito S, Nakashima K, Tsudzuki T, Sugiura M (1994) Loss of all ndh genes as determined by sequencing the entire chloroplast genome of the black pine Pinus thunbergii. Proc Natl Acad Sci USA 91:9794–9798
Wolfe KH (1988) The site of deletion of the inverted repeat in pea chloroplast DNA contains duplicated gene fragments. Curr Genet 131:97–99
Wolfe KH, Morden CW, Palmer JD (1992) Function and evolution of a minimal plastid from a nonphotosynthetic parasitic plant. Proc Natl Acad Sci USA 89:10648–10652
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255
Acknowledgments
Funding for this project was provided by a grant from the NSF (DEB 0120709) to R.K.J. and J.L.B. Part of this work was performed under the auspices of the U.S. Department of Energy, Office of Biological and Environmental Research, by the University of California, Lawrence Berkeley National Laboratory, under contract No. DE-AC02-05CH11231. The authors thank Stacia Wyman for computational assistance and Tim Chumley and Gwen Gage for technical assistance in generating figures. We also thank Andrew Alverson, Katie Hansen, Paul Wolf, and Elizabeth Ruck for their helpful comments and suggestions on an early version of the manuscript. This paper represents a portion of R.C.H.’s Ph.D. thesis in botany at the University of Texas at Austin.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Haberle, R.C., Fourcade, H.M., Boore, J.L. et al. Extensive Rearrangements in the Chloroplast Genome of Trachelium caeruleum Are Associated with Repeats and tRNA Genes. J Mol Evol 66, 350–361 (2008). https://doi.org/10.1007/s00239-008-9086-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-008-9086-4