Abstract
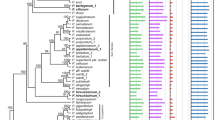
Minute inversions (4 bp in length), associated with probable hairpin secondary structures, were inferred from comparative analysis of rpl16 intron sequences from the chloroplast genomes of Chusquea species and related bamboos (Poaceae). The inverted sequences, which appear to have arisen independently on several occasions, comprise entire loops of the putative hairpins. The process of inversion seems dependent upon the stem length of the hairpin and its estimated free energy of formation. A similar inversion was uncovered for other plants in a previously published data set for a different non-coding region of the chloroplast genome, suggesting that the inversional process may be a common feature of non-coding DNA evolution. Several implications for phylogenetic analysis are noted.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 5 March 1996 / 17 April 1996
Rights and permissions
About this article
Cite this article
Kelchner, S., Wendel, J. Hairpins create minute inversions in non-coding regions of chloroplast DNA. Curr Genet 30, 259–262 (1996). https://doi.org/10.1007/s002940050130
Issue Date:
DOI: https://doi.org/10.1007/s002940050130