Abstract
Itaconate, a C5 unsaturated dicarboxylic acid, is an important chemical building block that is used in manufacturing high-value products, such as latex and superabsorbent polymers. Itaconate is produced by fermentation of sugars by the filamentous fungus Aspergillus terreus. However, fermentation by A. terreus involves a long fermentation period and the formation of various byproducts, resulting in high production costs. E. coli has been developed as an alternative for producing itaconate. However, fermentation of glucose gives low conversion yields and low productivity. Here, we report the whole-cell bioconversion of citrate to itaconate with enhanced aconitase and cis-aconitate decarboxylase activities by controlling the expression of multiple cadA genes. In addition, this bioconversion system does not require the use of buffers, which reduces the production cost and the byproducts released during purification. Using this whole-cell bioconversion system, we were able to catalyze the conversion of 319.8 mM of itaconate (41.6 g/L) from 500 mM citrate without any buffer system or additional cofactors, with 64.0% conversion in 19 h and a productivity of 2.19 g/L/h. Our bioconversion system suggests very high productivity for itaconate production.
Similar content being viewed by others
Introduction
Itaconate is a C5 unsaturated dicarboxylic acid that is produced from biomass and can be used in a number of high-value bio-based chemicals1. It is used industrially in the production of polymers such as synthetic latex, unsaturated polyester resins (UPR) and super absorbent polymers (SAP) and as a substitute for acrylic acid2. The total market demand for itaconate is increasing annually. In 2014, the global market for itaconate was valued at 126.4 million USD and is expected to increase to 204.6 million USD by 20233.
Biosynthesis of itaconate using Aspergillus itaconicus was discovered by Kinoshita in 19314. Since then, there have been many attempts to increase the titer of itaconate (Table 1). In 1945, Kane et al. reported itaconate production by A. terreus, which reached a titer of 27 g/L5. Apart from Aspergillus sp., other species have also been used to produce itaconate. Tabuchi et al. reported that Candida sp. can produce 35 g/L itaconate6. Ustilago sp. and Pseudozyma sp. were also shown to produce itaconate7,8,9. Currently, itaconate is produced using A. terreus with sugars as the substrate, and the titer and productivity reach 129 g/L and 1.15 g/L/h, respectively10. However, this titer is still lower than the expected theoretical maximum titer of 240 g/L by A. niger11, and the production cost is high due to high consumption of sugars, long duration of fermentation and purification of uncontrollable byproducts such as maleic acid, α-ketoglutaric acid, oxalic acid and other unidentified secondary metabolites10,12. Furthermore, the filamentous growth of fungi blocks the supply of oxygen during fermentation, which in turn hinders production of itaconate13. To address the oxygen supply issue and create a stable continuous system, bioconversion of citrate to itaconate using A. terreus in membrane bioreactor14, and fermentation using immobilized A. terreus has been also used for itaconate production15. However, the titer and productivity were uncompetitive compared to other conventional fermentation methods. In addition, A. niger has been developed as itaconate producer. However, the titer reached was only 26.2 g/L16. Although using a bacterial host for production has several advantages, such as rapid growth and easy controllability, E. coli does not have cis-aconitate decarboxylase (cadA), which converts cis-aconitate into itaconate. Since the discovery of CadA as the key enzyme in itaconate production13,17, strain of E. coli with heterologous expression of cadA gene was developed that can produce itaconate11. Vuoristo et al. produced itaconate using an E. coli strain expressing heterologous cadA, achieving a titer of 690 mg/L18. Another E. coli strain with random synonymous codon substitutions produced 7.23 g/L of itaconate19. Model-based metabolic engineering of E. coli increased itaconate production to 32 g/L20. However, E. coli fermentation is still not economically competitive with A. terreus fermentation in terms of titer and productivity. Thus, there is a need to develop additional strategies20.
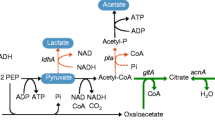
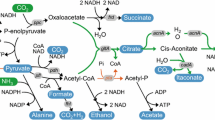
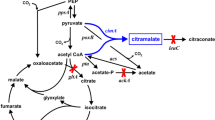
Here, we suggest an efficient whole-cell bioconversion system for the production of itaconate instead of fermentation (Fig. 1). In our bioconversion system, citrate is converted into itaconate by recombinant E. coli cells expressing aconitase and cis-aconitate decarboxylase. Compared to conventional fermentation, this whole-cell bioconversion system can decrease the cost and time of process because E. coli is a fast-growing species and the conversion reaction is rapid. Additionally, bioconversion produces fewer byproducts, which is an advantage in the purification process. Furthermore, citrate, which is the substrate for our conversion system, is readily available. Therefore, we investigated the feasibility of a high-yield conversion process using an E. coli whole-cell biocatalyst and optimized the reaction conditions to establish an efficient and competitive itaconate production bioprocess that would improve yield and productivity.
(a) E. coli whole-cell bioconversion strategy and (b) itaconate synthesis pathway in A. terreus (fermentation). In the E. coli whole-cell bioconversion strategy, two enzymes involved in this pathway, aconitase and cis-aconitate decarboxylase, were overexpressed. Unlike A. terreus, E. coli has no organelles, so the reaction proceeds spontaneously without the need for intracellular transport. In A. terreus fermentation, glucose is metabolized through glycolysis and the TCA cycle. Citric acid generated by the TCA cycle is converted to cis-aconitic acid by aconitase in the cytosol. Cis-aconitic acid is transferred to the mitochondria and converted to itaconic acid by cis-aconitate decarboxylase.
Results
Construction of an E. coli whole-cell biocatalyst expressing acn and cadA
To construct the E. coli whole-cell biocatalyst, acn from Corynebacterium glutamicum ATCC 13032 and a synthetic, codon-optimized version of the cadA (cis-aconitate decarboxylase) gene of A. terreus were cloned into pCDF-duet1 (pHMS01). acn from Corynebacterium glutamicum ATCC 13032 was selected as it has the lowest Km value to citrate among other prokaryotic species21. The E. coli strain BL21(DE3), which is typically used for protein production, was transformed with pHMS01. The expression of each gene was tested by running an SDS-PAGE and testing the whole-cell conversion reaction. Initially, BL21 containing pHMS01 converted only 3.4 mM of 100 mM citrate to itaconate (3.4%). SDS-PAGE analysis data showed that a portion of the CadA formed an inclusion body (data not shown). As shown in Fig. 2, changes in induction time and temperature influenced the expression of cadA. Conversion of citrate by BL21 containing pHMS01 increased 2.1-fold compared to the previous condition (7.2%). Although changing induction conditions increased the activities of the whole-cell, conversion was still low. Since CadA is the key enzyme for itaconate production13, the number of cadA gene copies was increased to overexpress cis-aconitate decarboxylase. The cadA gene was cloned into other duet vectors. pACYC-duet1 and pRSF-duet1 were tested for additional cadA expression. Increasing the number of cadA gene copies also improved the conversion of citrate. The addition of pHMS02 (pACYC-duet1::cadA) and pHMS03 (pRSF-duet1::cadA) (Fig. 2), each of which contained one cadA gene, improved citrate conversion compared to the pHMS01 single-vector system alone. The pHMS02 plasmid increased conversion from 7.2% to 10.2% and pHMS03 increased it to 15.9%. Furthermore, pHMS04 (pACYC-duet1::cadA::cadA) and pHMS05 (pRSF-duet1::cadA::cadA), which contain two cadA gene copies, improved conversion to a much greater degree. The pHMS04 plasmid increased conversion from 7.2% to 16.7%, while pHMS05 increased it to 25.7% (Fig. 2). All tested plasmids showed better conversion than the pHMS01 single-vector system. Comparing pHMS02 with pHMS03 and pHMS04 with pHMS05, pRSF-duet1 showed better expression than pACYC-duet1, as the former contains 100 copies and the latter fewer than 12 copies22. BL21 containing pHMS01 and pHMS05 (JY001) converted 25.7 mM of citrate to itaconate, resulting in a 7.6-fold increase which is equivalent to 25.7%. Thus, JY001 was used as a biocatalyst in further experiments.
Selection of induction media and substrate concentration
To determine the optimal reaction conditions, several factors were evaluated. Induction medium and induction timing were selected to maximize the expression of acn and cadA and increase the mass of the biocatalyst. Since TB and 2x YT broth contained higher concentrations of nitrogen source and improved the yields of the plasmid over that of LB broth23, yields from those three media were compared to determine the highest protein expression and cell growth. Cells cultured in LB medium showed a higher production yield than cell cultured in the two other types of media. One milligram of whole-cell biocatalyst in LB broth converted 1.05 mM of citrate to itaconate per hour; the equivalent value for TB was 0.69 mM and for 2xYT, it was 0.52 mM (Fig. 3a). However, the dry cell weight (DCW) from TB was the highest, at 5.05 g/L, which was 7.2-fold higher than that of LB, and that of 2x YT was 1.71 g/L, which was 2.4-fold higher than that of LB (Fig. 3a). These properties conferred a longer and more stable growth following induction and showed the highest DCW in the TB medium24. Therefore, TB medium was selected as the induction medium, as it showed a higher conversion rate compared to the other two media tested (3.48 mM/h). Regarding the timing of induction, the addition of IPTG at the initial point of induction resulted in the highest production. Regarding the optimal citrate concentration, 81.9 mM itaconate was produced from 500 mM of citrate and conversion was 16.3% (Fig. 3b). Although conversion at this concentration was lower than at other concentrations, it produced the highest amount of itaconate compared to other citrate concentrations tested, and thus, 500 mM citrate was selected for use in further experiment.
(a) Selection of induction medium. Enzyme expression was higher in LB. However, cell mass was higher in Terrific broth. Itaconate/DCW/Hour ( ); DCW (
); DCW ( ). (b) Determination of the optimal citrate concentration; 500 mM citrate produced the highest amount of itaconate. The reaction was conducted at pH 7 and 35 °C for 24 h. Itaconate concentration (bar); Conversion (⚫).
). (b) Determination of the optimal citrate concentration; 500 mM citrate produced the highest amount of itaconate. The reaction was conducted at pH 7 and 35 °C for 24 h. Itaconate concentration (bar); Conversion (⚫).
Effects of pH, temperature and permeability of whole cells
The optimal pH for Acn and CadA have been reported as 7.5–7.75 and 6.2, respectively21,25,26. However, the optimal pH for whole-cell conversion reaction was evaluated since overall reaction was mainly affected by initial extracellular pH in our previous study27. The optimal pH for the whole-cell reaction was pH 5.5, which is different from the optimal pH for each individual enzyme (Fig. 4a). The optimal temperatures for Acn and CadA were previously reported as 50 °C and 37 °C, respectively21,25,26. However, in our system, itaconate production was maximal at 35 °C (Fig. 4b). Under optimum pH and temperature, 239 mM itaconate was produced from 500 mM citrate (47.8%). Apart from pH and temperature, cofactors and FeSO4 also determined aconitase activity28. However, no significant effect was found in whole-cell system (Supplement Fig. 1). Fe2+ is a metal ion that positively influences aconitase activity28 but also inhibits cis-aconitate decarboxylase29. As a result, we did not observe any remarkable effects of FeSO4 on itaconate production.
Permeability is another factors that is important in the whole-cell reaction since the substrate and product have to be transferred through the cell membrane30. Rapid uptake of citrate and rapid excretion of itaconate would significantly increase the conversion rate. To increase E. coli membrane permeability, the citrate carrier protein and dicarboxylic acid transporter citT was overexpressed in JY002, and its effects on itaconate production were evaluated (Supplement Fig. 2); however, no significant increase in productivity was observed. Surfactant were also applied in an attempt to increase cell permeability (Supplement Fig. 3). There have been reports of using surfactants to increase production by whole-cell biocatalysts31,32. Two surfactants, polyoxyethylene sorbitan monooleate (Tween 80), which is a non-ionic surfactant, and sodium dodecyl sulfate (SDS) an anionic surfactant, were evaluated. SDS at a concentration of 0.01% had no significant effect, and >0.1% SDS resulted in no conversion activity due to cell lysis and protein destruction. In contrast, Tween 80 at a concentration of 0.1% resulted in a slight increase in conversion activity (Supplement Fig. 3). The optimum concentration of Tween 80 was found to be 0.5%, which resulted in the production of 260.5 mM itaconate (52.1%) (Fig. 5).
Time-dependent monitoring of itaconate production
We conducted itaconate production using the whole-cell conversion system under the optimum conditions outlined above (pH 5.5, 500 mM citrate, 0.5% Tween 80, JY001 cells at 35 °C). The conversion rate increased constantly until 6 h and decreased after 6 h. The reaction was almost saturated after 13 h and was completed at 19 h. The pH value also increased constantly to 7.9 in a manner similar to the conversion pattern (Fig. 6b). To control pH value constantly, buffers were applied. However, no significant effect was observed (Supplement Fig. 4). The concentration of cis-aconitate, an intermediate in the two-step reaction, increased in 3 h to 23.6 mM due to aconitase activity. After 3 h, the cis-aconitate level decreased steadily to 12.7 mM as it was converted to itaconate by cis-aconitate decarboxylase. After 19 h, there was no other byproduct produced and the efficiency of citrate conversion to itaconate was revealed as 100% since concentration of consumed citrate/isocitrate and produced itaconate was equal. At the end of the reaction, the itaconate concentration was 319.8 mM (41.6 g/L), equivalent to 64.0% conversion (Fig. 6a). Overall productivity was 2.19 g/L/h, and maximum productivity was 5.43 g/L/h within a 4 to 6 h period.
Discussion
Many studies have reported improved itaconate production over several decades. Most recent studies have been based on a fermentation strategy using fungus A. terreus or E. coli. However, conventional fermentation requires long fermentation periods, high sugar consumption and unavoidable byproducts, which reduce productivity and increase production costs.
Therefore, we suggested bioconversion of itaconate from citrate using recombinant E. coli as a possible alternative method. Considering that E. coli is typically used for whole-cell reactions32,33 and that the speed of reaction by bioconversion is much faster than the conventional method (less than 19 h), resulting in high productivity, our bioconversion system could be comparable to conventional fermentation systems that use A. terreus or E. coli. Furthermore, our system uses citrate as the substrate, and the production of citrate is well established in the industry. A. niger produces 360 g/L of citric acid34, resulting in a cheap source of our starting material35,36.
To maximize the efficiency of our bioconversion system, we improved acn and cadA expression because aconitase and cis-aconitate decarboxylase is a critical enzyme for itaconate production13. Increasing cadA protein expression by increasing its gene copy number enhanced conversion yield by 18.5% (Fig. 2). However, increasing acn protein expression did not show significant effect on conversion (data not shown). This result shows that high expression of the cadA gene is more critical for this system. This is a reasonable proposition because aconitase and cis-aconitate decarboxylase compete for the key precursor, cis-aconitate (Fig. 1). Other studies have indicated the importance of cadA expression for itaconate production19,20. E. coli BL21(DE3) containing acn and three cadA genes yielded a much higher conversion (64.0%).
One of the important factors in itaconate production by bioconversion is the reaction conditions. Since itaconate production is the result of a two-step reaction, finding an optimal reaction condition was important. The optimal pH and temperature of the whole-cell biocatalyst were determined to be pH5.5 and 35 °C. The conversion was enhanced by adding 0.5% Tween 80, which increased the permeability of the cell membrane. The final concentration of itaconate obtained was 319.8 mM (41.6 g/L), a 64.0% conversion. This is still lower than that of other whole-cell reactions used in the production of other chemicals, such as cadaverine and acrylamide27,37. There are various possibilities on incomplete citrate conversion. It may be due to ceased utilization of citrate by pH or isocitrate production as a byproduct. However, considering that the overall productivity of a A. terreus fermentation is only 1.15 g/L/h10, the bioconversion of citrate to itaconate could be an alternative for itaconate production, as it shows an average productivity of 2.19 g/L/h and a maximum productivity of 5.43 g/L/h (4~6 h). This bioconversion system can be used in the production of itaconate without buffer as a whole-cell system. This represents a further advantage in that no purification is needed and this decreases the production cost.
In conclusion, we have developed the first E. coli whole-cell bioconversion system for the production of itaconate. The presence of multiple copies of cadA and optimization of the reaction conditions resulted in itaconate production at a level comparable to that achieved in traditional methods.
Materials and Methods
Reagents
Sodium citrate dihydrate (Dae-Jung, Korea) and citric acid (Duk-San Pure Chemical Co., Korea) were used to prepare a 1 M citrate solution. Citric acid, itaconic acid (Sigma-Aldrich, USA) and cis-aconitic acid (Alfa Aesar, USA) were used for in the preparation of a standard curve. PCR was performed with nPfu-Forte polymerase (Enzynomics, Daejeon, Korea). Amplified DNA and plasmid were purified with MG Plasmid SV Miniprep Kit (Doctor Protein, Seoul, Korea). All endonucleases (NdeI, EcoRV, BamHI, SacI, XhoI, and NotI) were purchased from New England Biolabs (USA).
Bacterial strains, plasmids, primers and media
E. coli strains and plasmids used in this study are listed in Table 2. All the E. coli strains were pre-cultured in 5 mL Luria-Bertani (LB) medium (10 g/L tryptone, 5 g/L yeast extract and 5 g/L NaCl) containing appropriate antibiotics (kanamycin 50 μg/mL, spectinomycin 100 μg/mL, and chloramphenicol 25 μg/mL) for 16 h at 37 °C. A total of 2% of the pre-cultured cells were inoculated in 50 mL of induction medium. LB medium, Terrific Broth (TB) medium (12 g/L tryptone, 24 g/L yeast extract, 72 mM K2HPO4, 17 mM KH2PO4 and 0.4%v/v glycerol) and 2x YT Broth medium (16 g/L tryptone, 10 g/L yeast extract, 5 g/L NaCl) were compared as induction media. To maintain the plasmids, appropriate antibiotics were added to the media. Initially, the cells were induced under 30 °C for 24 h with 0.5 mM of isopropyl β-D-1-thiogalactopyranoside (IPTG) at initial point. To prevent the formation of enzyme inclusion bodies, the induction temperature and time were adjusted to 25 °C for 48 h. The induction culture was carried out in a 250 mL baffled Erlenmeyer flask with a shaking incubator (HB-201SL, Han-Baek, Korea) at 25 °C, 200 rpm.
Genetic methods
A general molecular biology method was used for gene cloning38. Based on the sequence of cis-aconitate decarboxylase, originally from A. terreus, cadA was codon-optimized and synthesized by Cosmogenetech. (Seoul, Korea). The full sequence of the synthesized cadA gene is provided in the supplementary material. The acn gene encodes aconitase originally from Corynebacterium glutamicum ATCC 13032, and citT encodes a citrate carrier protein originally from E. coli K12. The acn, cadA and citT sequences were amplified by PCR using the primers listed in Table 2. The amplified DNA was purified and digested with endonucleases. acn was digested with NdeI and EcoRV, cadA at MCS1 with BamHI and SacI, and at MCS2 with NdeI and XhoI, and citT was digested with SacI and NotI. The digested PCR products were ligated to vectors that had been digested with the same endonuclease. Ligated plasmids were transformed into E. coli DH5α competent cells using the heat-shock method39. Constructed plasmids (Fig. 2) were confirmed using sequencing (Cosmogenetech, Seoul, Korea) and were used for further experiments.
The expression of each enzymes was tested by conversion reactions and SDS-PAGE (Sodium Dodecyl Sulfate PolyAcrylamide Gel Electrophoresis). A total of 1 mL of culture broth was centrifuged for cell harvest and washed with 50 mM of phosphate-buffed saline (PBS) pH 6.8. Harvested cells were treated with BugbusterTM (Novagen, Madison, WI, USA) under the conditions recommended by the manufacturer. After centrifugation at 15,184 × g for 3 min, the supernatant was sampled as the soluble fraction. The pellet of the insoluble fraction was washed 3 times and re-suspended in the same volume of PBS. SDS-PAGE was performed at 120 V for 2 h.
Whole-cell reaction
The activity of Acn and CadA as single whole-cell biocatalysts was determined. Initially, the reaction was carried out at 35 °C and pH 7 for 24 h. For optimization, the total reaction volume was 500 μL, containing 500 mM citrate and 43.5 mg of cells27. For time-dependent monitoring of the conversion reaction, the total volume was 20 mL with optimized conditions, and the reaction solution was mixed intermittently for sampling every hour. The cover of the reactor was lifted whenever a sample was obtained. The reaction was stopped by heating at 90 °C for 5 mins. The reaction solution was then diluted at 1:50 and analyzed by HPLC. The optimal pH was determined using 1 M trisodium citrate dihydrate (pH 8.7) and 1 M citric acid (pH 1.5) stocks. Activity of each gene was examined by production of itaconate and presented in supplementary Table 2.
Effect of additives on bioconversion reaction
FeSO4 as cofactor, and polyoxyethylene sorbitan monooleate (Tween 80) and Sodium Dodecyl Sulfate (SDS) (as cell membrane weakening agents) were tested. FeSO4 was prepared at a 10 mM concentration, and Tween 80 and SDS were used at 10%. Each reagent was added to the reaction solution. The final concentration of FeSO4 was 0, 5, 10, 15, or 20 μM, and those of Tween 80 and SDS were 0.01, 0.1, or 1%. Since Tween 80 caused increased conversion, the optimal concentrations of Tween 80 were tested from at concentrations ranging from 0.1 to 0.5%.
Analytical methods
Citrate, isocitrate, itaconate, and cis-aconitate were quantified by high-performance liquid chromatography (HPLC). The reaction solution was heated to 90 °C for 5 min to stop the reaction and to aggregate proteins and then centrifuged at 15,314 × g to eliminate cell debris. The supernatant was diluted to 1:50 and filtered through a Whatman syringe filter (0.2 μm, PVDF, Sigma-Aldrich, St. Louis, USA). The filtrate was analyzed by HPLC (Prominence-i LC-2030, Shimadzu, Japan) equipped with an Aminex HPX-87H column (300 × 7.8 mm, 9 μm, Bio-Rad, Hercules, California, USA). The solution was eluted using 0.004 M H2SO4 as the mobile phase with flow rate of 0.6 mL/min flow rate. The oven temperature was set at 60 °C, and a UV detector was used to monitor the analytes at 210 nm40,41. Concentrations of each organic acid were calculated with standard curve and retention time of citrate, isocitrate, itaconate, and cis-aconitate were 7.6, 7.7, 12.0, and 6.8, respectively.
Additional Information
How to cite this article: Kim, J. et al. Production of itaconate by whole-cell bioconversion of citrate mediated by expression of multiple cis-aconitate decarboxylase (cadA) genes in Escherichia coli. Sci. Rep. 7, 39768; doi: 10.1038/srep39768 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Werpy, T. et al. Top value added chemicals from biomass. Volume 1-Results of screening for potential candidates from sugars and synthesis gas. (DTIC Document, 2004).
Willke, T. & Vorlop, K. D. Biotechnological production of itaconic acid. Appl. Microbiol. Biotechnol. 56, 289–295 (2001).
Sudip, S. Itaconic acid market is expectd to reach US$ 204.6 Mn by 2023. Transparency market researchhttp://www.transparencymarketresearch.com/pressrelease/itaconic-acid-market.htm. (Date of access:21/10/2016) (2015).
Kinoshita, K. Production of itaconic acid and mannitol by a new mold, Aspergillus itaconicus. Acta Phytochim 5, 271–287 (1931).
Kane, J. H., Finlay, A. C. & Amann, P. F. Inventors; Pfizer & Co C., assignee. Production of itaconic acid. United States patent US 2,385,283. 1945 Sep 18.
Tabuchi, T., Sugisawa, T., Ishidori, T., Nakahara, T. & Sugiyama, J. Itaconic acid fermentation by a yeast belonging to the genus Candida. Agric. Biol. Chem. 45, 475–479 (1981).
Guevarra, E. D. & Tabuchi, T. Accumulation of itaconic, 2-hydroxyparaconic, itatartaric, and malic acids by strains of the genus Ustilago. Agric. Biol. Chem. 54, 2353–2358 (1990).
Voll, A., Klement, T., Gerhards, G., Büchs, J. & Marquardt, W. Metabolic modelling of itaconic acid fermentation with Ustilago maydis. Chem. Eng. Trans. 27, 367–372 (2012).
Levinson, W. E., Kurtzman, C. P. & Kuo, T. M. Production of itaconic acid by Pseudozyma antarctica NRRL Y-7808 under nitrogen-limited growth conditions. Enzyme. Microb. Technol. 39, 824–827 (2006).
Hevekerl, A., Kuenz, A. & Vorlop, K.-D. Influence of the pH on the itaconic acid production with Aspergillus terreus. Appl. Microbiol. Biotechnol. 98, 10005–10012 (2014).
Li, A. et al. A clone-based transcriptomics approach for the identification of genes relevant for itaconic acid production in Aspergillus. Fungal. Genet. Biol. 48, 602–611 (2011).
Li, A. et al. Reduced by-product formation and modified oxygen availability improve itaconic acid production in Aspergillus niger. Appl. Microbiol. Biotechnol. 97, 3901–3911 (2013).
Klement, T. & Büchs, J. Itaconic acid–a biotechnological process in change. Bioresour. Technol. 135, 422–431 (2013).
Bressler, E. & Braun, S. Conversion of citric acid to itaconic acid in a novel liquid membrane bioreactor. J. Chem. Technol. Biotechnol. 75, 66–72 (2000).
Kautola, H., Rymowicz, W., Linko, Y.-Y. & Linko, P. Itaconic acid production by immobilized Aspergillus terreus with varied metal additions. Appl. Microbiol. Biotechnol. 35, 154–158 (1991).
Hossain, A. H. et al. Rewiring a secondary metabolite pathway towards itaconic acid production in Aspergillus niger. Microb. Cell. Fact. 15, 130 (2016).
Kanamasa, S., Dwiarti, L., Okabe, M. & Park, E. Y. Cloning and functional characterization of the cis-aconitic acid decarboxylase (CAD) gene from Aspergillus terreus. Appl. Microbiol. Biotechnol. 80, 223–229 (2008).
Vuoristo, K. S. et al. Metabolic engineering of itaconate production in Escherichia coli. Appl. Microbiol. Biotechnol. 99, 221–228 (2015).
Jeon, H. G., Cheong, D. E., Han, Y., Song, J. J. & Choi, J. H. Itaconic acid production from glycerol using Escherichia coli harboring a random synonymous codon-substituted 5′-coding region variant of the cadA gene. Biotechnol. Bioeng (2015).
Harder, B.-J., Bettenbrock, K. & Klamt, S. Model-Based Metabolic Engineering Enables High Yield Itaconic Acid Production by Escherichia coli. Metab. Eng. (2016).
Baumgart, M. & Bott, M. Biochemical characterisation of aconitase from Corynebacterium glutamicum. J. Biotechnol. 154, 163–170 (2011).
Held, D., Yaeger, K. & Novy, R. New coexpression vectors for expanded compatibilities in E. coli. InNovations 18, 4–6 (2003).
Tartof, K. D. & Hobbs, C. A. Improved media for growing plasmid and cosmid clones. Focus 9, 12 (1987).
Byrne, J. Under what circumstances and why is terrific broth preferred to LB broth for E. coli growth? Quorahttps://www.quora.com/Under-what-circumstances-and-why-is-terrific-broth-preferred-to-LB-broth-for-E-coli-growth. (Date of access: 21/10/2016) (2011).
Steiger, M. G., Blumhoff, M. L., Mattanovich, D. & Sauer, M. Biochemistry of microbial itaconic acid production. Front. Microbiol. 4, 23 (2013).
Dwiarti, L., Yamane, K., Yamatani, H., Kahar, P. & Okabe, M. Purification and characterization of cis-aconitic acid decarboxylase from Aspergillus terreus TN484-M1. J. Biosci. Bioeng. 94, 29–33 (2002).
Kim, H. J. et al. Optimization of direct lysine decarboxylase biotransformation for cadaverine production with whole cell biocatalysts at high substrate concentration. J. Microbiol. Biotechnol. 25, 1108–1113 (2015).
Tsuchiya, D., Shimizu, N. & Tomita, M. Cooperativity of two active sites in bacterial homodimeric aconitases. Biochem. Biophys. Res. Commun. 379, 485–488 (2009).
Bentley, R. & Thiessen, C. P. Biosynthesis of itaconic acid in Aspergillus terreus III. The properties and reaction mechanism of cis-aconitic acid decarboxylase. J. Biol. Chem. 226, 703–720 (1957).
Chen, R. R. Permeability issues in whole-cell bioprocesses and cellular membrane engineering. Appl. Microbiol. Biotechnol. 74, 730–738 (2007).
Matsushima, Y., Hirasawa, T. & Shimizu, H. Enhancement of 1,5-diaminopentane production in a recombinant strain of Corynebacterium glutamicum by Tween 40 addition. J. Gen. Appl. Microbiol. 62, 42–45 (2016).
Gao, B. et al. Development of recombinant Escherichia coli whole-cell biocatalyst expressing a novel alkaline lipase-coding gene from Proteus sp. for biodiesel production. J. Biotechnol. 139, 169–175 (2009).
de Carvalho, C. C. C. R. Enzymatic and whole cell catalysis: finding new strategies for old processes. Biotechnol. Adv. 29, 75–83 (2011).
Tsao, G. T., Cao, N. J., Du, J. & Gong, C. S. In Recent progress in bioconversion of lignocellulosics 243–280 (Springer, 1999).
Vieira, G. S., Cavalcanti, R. N., Meireles, M. A. A. & Hubinger, M. D. Chemical and economic evaluation of natural antioxidant extracts obtained by ultrasound-assisted and agitated bed extraction from jussara pulp (Euterpe edulis). J. Food Eng. 119, 196–204 (2013).
Mielcarek, A., Rodziewicz, J., Janczukowicz, W. & Thornton, A. The feasibility of citric acid as external carbon source for biological phosphorus removal in a sequencing batch biofilm reactor (SBBR). Biochem. Eng. J. 93, 102–107 (2015).
Kim, B.-Y. & Hyun, H.-H. Production of acrylamide using immobilized cells of Rhodococcus rhodochrous M33. Biotechnol. Bioprocess Eng. 7, 194–200 (2002).
Sambrook, J. & Russell, D. W. Molecular cloning: a laboratory manual 3rd edition (Coldspring-Harbour Laboratory Press, UK, 2001).
Inoue, H., Nojima, H. & Okayama, H. High efficiency transformation of Escherichia coli with plasmids. Gene 96, 23–28 (1990).
Zhang, D. et al. Consolidated pretreatment and hydrolysis of plant biomass expressing cell wall degrading enzymes. Bioenergy Res. 4, 276–286 (2011).
Kautola, H., Vahvaselkä, M., Linko, Y. Y. & Linko, P. Itaconic acid production by immobilizedAspergillus terreus from xylose and glucose. Biotechnol. Lett. 7, 167–172 (1985).
Taylor, R. G., Walker, D. C. & McInnes, R. R. E. coli host strains significantly affect the quality of small scale plasmid DNA preparations used for sequencing. Nucleic Acids Res. 21, 1677–1678 (1993).
Acknowledgements
This study was supported by the National Research Foundation of Korea ((NRF-2015R1A2A2A04006014, NRF-2016R1D1A1B03932301, NRF-2015M1A5A1037196) and by the Korea Institute of Energy Technology Evaluation and Planning (KETEP) and the Ministry of Trade, Industry & Energy (MOTIE) of the Republic of Korea (No. 20163010092150). Consulting service from the Microbial Carbohydrate Resource Bank (MCRB, Seoul, Korea) was kindly appreciated.
Author information
Authors and Affiliations
Contributions
Y.H.Y. supervised the study and revised the manuscript; J.K. performed the experiments and wrote the paper; H.M.S. performed plasmid construction; J.H.K. performed the whole-cell reactions; J.M.J. provided useful help in reaction optimization; H.S.S. performed the HPLC experiments; S.K.B. designed the experiments; K.Y.C. contributed to the study design; Y.G.K. conceived the study; W.K. analyzed the data; and J.J.Y. reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kim, J., Seo, HM., Bhatia, S. et al. Production of itaconate by whole-cell bioconversion of citrate mediated by expression of multiple cis-aconitate decarboxylase (cadA) genes in Escherichia coli. Sci Rep 7, 39768 (2017). https://doi.org/10.1038/srep39768
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep39768
- Springer Nature Limited
This article is cited by
-
Efficient production of itaconic acid from the single-carbon substrate methanol with engineered Komagataella phaffii
Biotechnology for Biofuels and Bioproducts (2024)
-
Construction of cell factory through combinatorial metabolic engineering for efficient production of itaconic acid
Microbial Cell Factories (2022)
-
Synergistic effects on itaconic acid production in engineered Aspergillus niger expressing the two distinct biosynthesis clusters from Aspergillus terreus and Ustilago maydis
Microbial Cell Factories (2022)
-
Inhibition of Cyclopropane Fatty Acid Synthesis in the Membrane of Halophilic Halomonas socia CKY01 by Kanamycin
Biotechnology and Bioprocess Engineering (2022)
-
Improvement of cadaverine production in whole cell system with baker’s yeast for cofactor regeneration
Bioprocess and Biosystems Engineering (2021)