Abstract
Organic agriculture is a farming method that provides healthy food and is friendly to the environment, and it is developing rapidly worldwide. This study compared microbial communities in organic farming (Or) paddy fields to those in nonorganic farming (Nr) paddy fields based on 16S rDNA sequencing and analysis. The predominant microorganisms in both soils were Proteobacteria, Chloroflexi, Acidobacteria, Actinobacteria, and Nitrospirota. The alpha diversity of the paddy soil microbial communities was not different between the nonorganic and organic farming systems. The beta diversity of nonmetric multidimensional scaling (NMDS) revealed that the two groups were significantly separated. Distance-based redundancy analysis (db-RDA) suggested that soil pH and electrical conductivity (EC) had a positive relationship with the microbes in organic paddy soils. There were 23 amplicon sequence variants (ASVs) that showed differential abundance. Among them, g_B1-7BS (Proteobacteria), s_Sulfuricaulis limicola (Proteobacteria), g_GAL15 (p_GAL15), c_Thermodesulfovibrionia (Nitrospirota), two of f_Anaerolineaceae (Chloroflexi), and two of g_S085 (Chloroflexi) showed that they were more abundant in organic soils, whereas g_11-24 (Acidobacteriota), g__Subgroup_7 (Acidobacteriota), and g_Bacillus (Firmicutes) showed differential abundance in nonorganic paddy soils. Functional prediction of microbial communities in paddy soils showed that functions related to carbohydrate metabolism could be the major metabolic activities. Our work indicates that organic farming differs from nonorganic farming in terms of microbial composition in paddy soils and provides specific microbes that might be helpful for understanding soil fertility.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
At present, there are an increasing number of trends in the analysis of soil microbiota by overall genomics or overall transcriptomics, such as the application of soil microbiota in Arctic peat [1], grassland soil microbiota [2], dry soil microbiota in Antarctica, and changes in water and organic matter microbiota [3]. The study of the bacterial flora of wetland sedimentary soil shows that Proteobacteria is the main bacterial phylum, and the next most prominent phyla are Acidobacteria, Bacteroidetes, Actinobacteria, and Verrucomicrobia. In Proteobacteria, Gamma-, Delta-, and Betaproteobacteria are the representative classes [4]. Comparing different soil management practices, such as forest and agricultural soil, the results show that the bacteria in forest soil are more abundant than those in agricultural soil [5]. It is possible that land preparation, fertilization, and spraying of pesticides in agricultural management may cause the bacterial phase of the soil to tend to be monotonous.
The addition of organic fertilizers (alfalfa straw, glucose, compost manure, and wood biochar) helps to promote the growth of Acidobacteria and Gemmatimonadetes bacteria, whereas synthetic fertilizers may promote an increase in Actinobacteria and Proteobacteria bacteria in rocket (Eruca sativa) cultivation [6]. In paddy soil, the application of inorganic fertilizers increased the abundance of bacteria Bacteroidetes and Acidobacteria, but the application of organic fertilizers (composted pig manure and rice straw) increased the abundance of eutrophic bacteria (such as Proteobacteria) [7]. Moreover, studies on the long-term effects of organic and inorganic fertilizers on the functioning of soil microbial communities have shown that the application of nitrogen fertilizer alone reduces microbial diversity, while farmyard manure application alone or in combination with chemical fertilizers improves soil fertility [8]. However, farmyard manure application might not be the major cause, but balanced fertilizer containing nitrogen, phosphorus, and potassium would result in the highest microbial diversity in paddy soil [9]. Indeed, paddy soil fertility is affected by microbial communities, and high-yield soil microbes are likely more active in modulating soil fertility for rice production [10]. The positive correlation of specific soil bacteria with grape-growing regions in the United States shows that the identification of specific microbes is helpful for understanding viticulture and flavor [11]. Overall, it is not yet clear the microbial communities between organic and nonorganic farming that certain bacteria could be applied as indicators to distinguish organic paddy soil.
In this study, we collected soils from organic and nonorganic farming of paddy fields in Changhua County, Taiwan. Soil microbial DNA was prepared, and 16S rDNA was sequenced, followed by microbe comparison and biodiversity analysis. This study aims to use next-generation sequencing to comprehensively compare microbial communities between organic and nonorganic paddy fields and further apply bioinformatics tools for functional analysis.
Material and methods
Soil sample collection
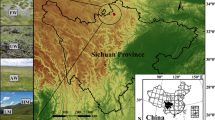
The soils used in this study were collected from six paddy fields with organic farming (Or1-5, Or6-8, Or9-11, and Or12-14) and nonorganic farming (Nr1-5, and Nr6-8) in Changhua County, Taiwan (Fig. 1). The soil collection periods for Or1-5 and Nr1-5 samples were during Oct. 2018, and for Or1-5, Or6-8, Or9-11, and Nr6-8 samples were during Oct. 2019. Five or three soil samples collected from each paddy field were applied as the replicates. The soil was collected with a soil sampler at a depth of 10–20 cm. After collection, samples were placed in labeled sterile polyethylene centrifuge tubes, brought to the laboratory, and stored at − 20 °C. The study information includes samples, farming types, collection date, collection location, soil pH, and electrical conductivity (EC), which are listed in Table 1.
Soil pH and EC analysis
For soil pH analysis, 5 g of air-dried soil was put into a 15 mL sterile polyethylene centrifuge tube, followed by the addition of 5 mL deionized water (sample: water = 1:1), and placed on a reciprocal shaker for 30 min at 140 rpm [12]. For soil EC analysis, 5 g of air-dried soil was put into a 50 mL sterile polyethylene centrifuge tube, mixed with 25 mL of deionized water (sample: water = 1:5), and placed on a reciprocal shaker for 60 min at 140 rpm, followed by standing for 30 min [13]. The solution or supernatant was recorded for the soil pH or EC, respectively, on a multifunctional water detector meter, WA-2017SD (LUTRON ELECTRONIC ENTERPRISE CO., LTD., Taipei, Taiwan).
DNA extraction, polymerase chain reaction (PCR), and sequencing
Bacterial genomic DNA (gDNA) was extracted from 0.5 g of soil by using a NucleoSpin Soil DNA isolation kit (MACHEREY–NAGEL, Dueren Germany) according to the manufacturer’s instructions, and DNA sequencing was performed at BIOTOOLS Co., Ltd. (New Taipei, Taiwan). For 16S rRNA gene sequencing, the V3-V4 region was amplified by a specific primer set (319F: 5′ CCTACGGGNGGCWGCAG 3′, 806R: 5′ GACTACHVGGGTATCTAATCC 3′—according to the 16S Metagenomic Sequencing Library preparation procedure (Illumina, San Diego, CA, USA)). PCR and sequencing were performed as described [14]. Nucleotide sequences determined in the study have been deposited into the NCBI Sequence Read Archive database, and the BioProject accession number is PRJNA997222.
Processing and analysis of sequence data
The demultiplexed 16S rDNA sequences were primers removed and denoised with DADA2 [15] to obtain the amplicon sequence variants (ASVs), and the derived table indicated the number of times each ASV was observed in each sample by QIIME 2 [16]. For DADA2 denoising steps, the qiime dada2 denoise-paired command was applied, and the parameters were –p-trim-left-f 10, –p-trim-left-r 10, –p-trunc-len-f 240, and –p-trunc-len-r 200 based on the quality profile and amplicon length. The quick phylogeny with fast tree, rarefaction curve generation, and taxonomy assignment were performed with QIIME 2 [16]. The Silva 138 ribosomal RNA database was assigned for taxonomy [17]. The phyloseq project was built for microbial community analysis [18] as described [19] in R version 4.3.0 [20].
Hierarchical clustering was calculated using the MicrobiotaProcess package [21] with the get_clust function in methods, Jaccard (distance method for dissimilarity), Hellinger (distance method with standardization) and average (average distance between two clusters), and plotted with the ggclust function. Alpha diversity was generated with the plot_richness function, and beta diversity of nonmetric multidimensional scaling (NMDS) was calculated with Jaccard distance, followed by the analysis of similarities (ANOSIM) test, and generated with the plot_ordination function [19]. Distance-based redundancy analysis (db-RDA) was performed using the Microeco package [22] in the Jaccard method with cal_ordination, trans_ordination, and plot_ordination functions.
Functional prediction of microbial communities was performed with the Functional Annotation of Prokaryotic Taxa (FAPROTAX) [23], Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt2) [24], and Tax4Fun2 [25] as described previously [26].
Statistical analysis
The statistical analysis of metagenomic profiles (STAMP) was applied to compare the nonorganic and organic framing groups (Table 1) using White’s nonparametric t test (p < 0.05) with difference between proportions parameter (0.1) for FAPROTAX and Tax4Fun2 analyses and difference between proportion (0.05) for PICRUSt2 [27]. For ASV analysis, Storey false discovery rate (FDR) correction was applied with a q-value < 0.05, implying that the false-positive rate was 5% and that the difference between proportions was 0.2%. The White’s nonparametric t test is a permutation method for non-normal distributions. For features with statistical significance, we further applied the difference between proportions filter to present widely differing features for comparison. The selection of parameters will vary according to different methods and is subjective.
Results
Phylogenetic and diversity analysis
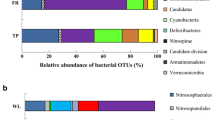
All organic rice fields were certified by the agency, and mixed organic fertilizer containing peanut meal, rapeseed meal, castor meal, sesame meal, corn meal, red bran, palm ash, soybean meal, rice bran, fish meal, bone powder, plant ash, and weathered lignite was applied (personal communication). We collected soils from paddy fields of organic (Or) and nonorganic (Nr) farming, and the microbial communities were examined. Figure 2 compares two different farms based on the percent composition of different bacterial varieties. Both Or and Nr soils showed the same top five dominant microorganisms, but the order of relative abundance was different, which were Proteobacteria (22.5%), Chloroflexi (14.8%), Acidobacteria (12.8%), Nitrospirota (4.7%), and Actinobacteria (4.5%) in Or soils and Proteobacteria (19.8%), Acidobacteria (17.6%), Chloroflexi (15.2%), Actinobacteria (5.6%), and Nitrospirota (4.6%) in Nr soils. After comparing the community richness (Chao1 and Observed_features) and diversity (Shannon and Simpson), we found that there was no significant difference between Or and Nr soils (Fig. 3).
Microbial community analysis
NMDS, a nonparametric ordination analysis based on Jaccard distance, revealed two separated groups of microbial communities from Or and Nr soils (Fig. 4A), and the ANOSIM test showed a statistically significant difference (p < 0.001), suggesting that both microbial communities of paddy fields were distinct after organic and nonoganic farming. Hierarchical clustering of the soil microbial communities showed that the microbial communities in Or12, Or13, and Or14 soils were grouped together with the microbial communities in Nr6, Nr7, and Nr8 soils (Fig. 4B). The possible reason is that these soils were collected in proximity. However, the microbial communities in organic farming soils were different from those in nonorganic farming soils.
A Nonparametric multidimensional scaling (NMDS) analysis with Jaccard distance on bacterial communities of paddy soil. Each point represents the bacterial community of a given sample. B Hierarchical cluster analysis of the soil microbial communities by the Jaccard, Hellinger, and average methods. Nr indicates nonorganic farming. Or indicates organic farming
We applied db-RDA to test the relationship between the environmental factors (pH and EC) and microbial community structures, and the results showed that the angle between the pH and EC variables was smaller than 90°, suggesting that both are positively correlated (Fig. 5A). Most nonorganic soils were located in the opposite direction of the EC arrow, suggesting a negative relationship between the EC variable and the microbes, and the EC of organic paddy soils appeared to be higher than that of nonorganic paddy soils (Fig. 5B). The microbes of organic paddy fields had a positive relationship with the pH variable, and the pH of organic paddy soils was higher than the pH of nonorganic paddy soils (Fig. 5B). The results highly suggested that the microbial community composition in organic paddy soil was associated with higher pH and EC environment compared to the microbial community in nonorganic paddy soil. The RDA at the phylum level showed that Acidobacteria bacteria were positively correlated with pH, and Proteobacteria, Chloroflexi, Actinobacteria, Nitrospirota, and Firmicutes bacteria were negatively correlated with pH. For EC, bacteria Chloroflexi, Actinobacteria, and Firmicutes showed a positive correlation, and bacteria Acidobacteria showed a negative correlation (Fig. S1).
In order to analyze the significant ASVs in different soils, a differential abundance test of ASVs was applied by the STAMP software with White’s nonparametric t test to organic and nonorganic farming groups, followed by the Story FDR correction with the parameters of q-value < 0.05 and the difference between proportions of 0.2%. Twenty-three of 23937 ASVs showed differential abundance with statistical significance (Fig. 6A). There were 13 ASVs from Or soils that showed higher abundance than the ASVs in Nr soils. Among them, 5 ASVs were assigned to Chloroflexi bacteria, and 4 were assigned to Proteobacteria bacteria. The other 10 ASVs showed a higher abundance in Nr soils, 3 were Acidobacteria bacteria, and 3 were Nitrospirota bacteria (Fig. 6B). In the heatmap (Fig. S2) and bar plot (Fig. S3) analyses, the differential abundances of ASVs were used to create a graphical representation for individual soil samples. If the uniformity of ASV abundance differences is considered, rather than being limited to only a few samples, these ASVs, g_B1-7BS (Proteobacteria), s_Sulfuricaulis limicola (Proteobacteria), g_GAL15 (p_GAL15), c_Thermodesulfovibrionia (Nitrospirota), two of f_Anaerolineaceae (Chloroflexi), and two of g_S085 (Chloroflexi), show that they are indeed more abundant in organic soils (Fig. S3A), whereas these ASVs, g_11-24 (Acidobacteriota), g__Subgroup_7 (Acidobacteriota), and g_Bacillus (Firmicutes), show differential abundance in nonorganic paddy soils (Fig. S3B). These significant ASVs can be used as the potential quality parameters to differentiate the two systems.
Differential abundance analysis of amplicon sequence variants (ASVs) in paddy soils. A Significant ASVs in organic and nonorganic farming groups were calculated by the statistical analysis of metagenomic profiles (STAMP) software with White’s nonparametric t test, followed by the Storey false discovery rate (FDR) correction with the parameters of q-value < 0.05 and the difference between proportions of 0.2%. The ASVs are shown in the assigned taxon. B The numbers of ASVs assigned to phyla
Functional feature prediction
There were 3, 17, and 17 functional features shown to be statistically significant from the programs FAPROTAX, PICRUSt2, and Tax4Fun2, respectively, and the differential abundance of the functional features is shown in Fig. 7A, B, C. According to functional characteristics, we referred to the KEGG pathway maps to classify manually and compare these functional features (Table S1). Among these features, functions related to carbohydrate metabolism were the only functions included in all three calculated programs, and the sum of the percentage was 62.8%. Subsequently, the functions of energy metabolism were included in the two programs, and the sum percentages were 72.6% (Fig. 7D). Cellulolysis, bifidobacterium shunt, and pyruvate fermentation to isobutanol (engineered) of the functions relative to carbohydrate metabolism were upregulated in nonorganic paddy soils, whereas glycolysis II (from fructose 6-phosphate), butanoate metabolism, and amino sugar and nucleotide sugar metabolism were upregulated in organic paddy soils, suggesting carbon nutrient levels in both soils (Fig. 7A, B, C). Again, methanol_oxidation, nitrite_respiration and oxidative phosphorylation of the functions relative to energy metabolism were upregulated in organic soils, suggesting that the bacteria involved in these metabolic pathways are differentially abundant in organic soils (Fig. 7A, C). Among the metabolic pathways of the functional features, 8 pathway categories showed a higher differential abundance in Or soils, and 5 pathway categories showed a higher differential abundance in Nr soils (Table S1).
Differential abundance analyses of functional features calculated by A FAPROTAX, B PICRUSt2, and C Tax4Fun2 in nonorganic and organic paddy soils. Statistical analyses were performed with STAMP using White’s nonparametric t test (p < 0.05) with a difference between proportions parameter (0.1) for FAPROTAX and Tax4Fun2 analyses and a difference between proportion (0.05) for PICRUSt2 without correction
Discussion
Bacteria Proteobacteria (33.0%), Acidobacteria (12.0%), Actinobacteria (5.8%), Bacteroidetes (5.4%), Firmicutes (3.9%), and Verrucomicrobia (3.6%) have been shown to be the dominant phyla in paddy soils [28]. The relative abundances of bacterial phyla in a 47-year-old long-term fertilizer experiment under a rice‒rice cropping system were Proteobacteria (43.3%), Acidobacteria (18.8%), Actinobacteria (7.9%), Chloroflexi (6.3%), and Firmicutes (3.9%) in nitrogen (N) + phosphorus (P) + potassium (K) fertilizer, whereas in farmyard manure, as the organic fertilizer, they were Proteobacteria (36.4%), Acidobacteria (24.5%), Actinobacteria (7.6%), Chloroflexi (9.9%), and Firmicutes (3%) [8]. Furthermore, with the addition of composted pig manure and rice straw, the bacteria Acidobacteria, Verrucomicrobia, and Latescibacteria were significantly decreased, but Actinobacteria and Firmicutes were significantly increased in paddy soils [29]. Nitrospira bacteria have been shown to increase in nitrogen fertilizer application alone over 47 years [8], suggesting that the presence of Nitrospira is related to nitrogen nutrients in soils. Biochar application in paddy soils significantly reduced Nitrospirae bacteria by 25%, thus depressing nitrification [30]. Our results showed the top five microorganisms were Proteobacteria (22.5%), Chloroflexi (14.8%), Acidobacteria (12.8%), Nitrospirota (4.7%), and Actinobacteria (4.5%) in Or soils and Proteobacteria (19.8%), Acidobacteria (17.6%), Chloroflexi (15.2%), Actinobacteria (5.6%), and Nitrospirota (4.6%) in Nr soils. Therefore, our results are consistent with previous studies, that is, the dominant bacterial phyla in rice fields are Proteobacteria, Acidobacteria, and Actinobacteria, and the distribution of Nitrospirota could be related to nitrogen fertilizer in the soil. Interestingly, a study on soil bacterial communities in paddy fields under organic and conventional farming conditions reported similar results: Proteobacteria (25–30%), Chloroflexi (20%), Acidobacteria (15%), and Actinobacteria (10%) were the predominant bacteria in all treatments, and rice bran and rapeseed oil cake were used as the organic fertilizers [31]. Because different organic fertilizers were applied, such as farmyard manure, compost pig manure, green manure, rice straw, rice bran, rapeseed oil cake, biochar, and mixed organic fertilizer, the different nutrients drove soil microbes toward diverted adaptation.
The promising results have been shown to improve wildlife diversity in organic farming. Research on the impact of organic farming on wildlife in rice fields has shown that the number and richness of wildlife have increased, and the number and richness of waterbird have a positive correlation with organic farming [32]. Moreover, organic farming in paddy fields can improve terrestrial arthropod diversity but not underground soil eukaryotic diversity [33]. However, there were no consistent results for microbial diversity in paddy soil under organic farming. For example, organic fertilizers could increase bacterial alpha diversity [6, 8], while no significant differences were observed [7, 29, 31]. For beta diversity, organic fertilizers seem to produce distinct clusters separated from the microbial communities with chemical fertilizers [7, 29]. Therefore, organic fertilizer application should provide different nutrients to influence the growth of certain microbes. Furthermore, cultivation periods, such as elongation, booting, grain filling, or ripening stages, also affect soil microbial community composition [31, 34].
Our db-RDA showed that the microbial community composition in organic paddy soil was positively correlated with higher pH and EC compared to the microbial community in nonorganic paddy soil (Fig. 5A, B). Organic amendments, such as alfalfa straw, compost manure, or wood biochar, significantly increase soil pH and decrease soil EC compared to mineral applications [6]. A similar result was presented in which biochar application could elevate soil pH in paddy soil, and Nitrospirae bacteria were also found to decrease pH by 25% [30]. A previous study showed that the relative abundances of the Proteobacteria and Bacteroidetes phyla were positively correlated with soil pH, while the abundances of Actinobacteria, Firmicutes, and Chloroflexi were negatively correlated with soil pH [35]. Cesarano et al. [6] also reported that Actinobacteria bacteria were significantly negatively correlated with soil pH, whereas Acidobacteria bacteria were negatively correlated, and Actinobacteria bacteria were significantly positively correlated with soil EC. The soil pH indeed affects the composition and diversity of soil bacteria [8]. Furthermore, soil EC plays roles in the changes in soil texture, cation exchange capacity, and percentage of humic matter, presenting the specific relationships between EC and nutrient concentrations [36]. Further studies are needed to clarify the relationships between certain bacteria and soil pH and EC and the relationships between certain organic fertilizer applications and soil pH and EC.
The ASVs assigned to g_B1-7BS (Proteobacteria), s_Sulfuricaulis limicola (Proteobacteria), g_GAL15 (p_GAL15), c_Thermodesulfovibrionia (Nitrospirota), two of f_Anaerolineaceae (Chloroflexi), and two of g_S085 (Chloroflexi) are specifically present in organic soils (Fig. 6A, S3A). B1-7BS showed a relatively higher dispersion degree among the anaerobic, anoxic, and oxic zone in a carrousel oxidization ditch system for domestic wastewater treatment [37]. In a study of Deoxynivalenol (DON) degradation by microbes, B1-7BS was also reported to be abundant in wheat fields [38]. Sulfuricaulis limicola could oxidize thiosulfate, tetrathionate, and elemental sulfur as electron donors to support autotrophic growth [39]. Phylum GAL15 has been shown to increase in abundance with increasing soil depth [40,41,42] and might be related to wetlands, such as riparian soils [42] and floodplain soils [41]. The genus Thermodesulfovibrio, a thermophilic, gram-negative bacterium, can reduce sulfate, thiosulfate, and sulfite as electron acceptors for growth [43]. Thermodesulfovibrio was found to be frequently present in environments with sulfur, nitrogen, and methane cycling [44], suggesting that its abundance in soil is related to soil fertility, responsible for the energy metabolism pathway. Interestingly, we found one ASV assigned to c_Thermodesulfovibrionia specific to organic paddy soil and two ASVs specific to nonorganic paddy soil (Fig. 6A). The genus Anaerolinea contains gram-negative, nonmotile, and filamentous bacteria that strictly grow in anaerobic conditions [45]. Anaerolineaceae could syntrophically cooperate with Methanosaeta archaea in the process of methanogenic degradation of alkanes [45, 46]. Furthermore, Thermodesulfovibrio spp. have been shown to be associated with Anaerolineaceae and Methanoculleus spp., and Anaerolineaceae initiates the activation of long-chain n-alkane biodegradation, followed by Thermodesulfovibrio metabolizing the intermidiates [47], implying in Or soils the potential cooperation between Anaerolineaceae and Thermodesulfovibrio. Genus S085 belongs to class Dehalococcoidia, which reduces dechlorinates tetrachloroethene to ethene [48]. S085 has also been shown to be present significantly in paddy soils with high arsenic levels compared to soils with low arsenic levels [49]. It is difficult to rationalize the bioremediation of S085 bacteria in Or soils.
The ASVs assigned to g_11-24 (Acidobacteriota), g__Subgroup_7 (Acidobacteriota), and g_Bacillus (Firmicutes) showed differential abundances in nonorganic paddy soils (Fig. 6A, S3B). Genus 11–24 belongs to class Blastocatellia. Blastocatellia, a gram-negative, spherical to rod-shaped and nonmotile bacterium, shows a broad tolerance range of pH for growth and is usually isolated from the soil environment [50]. A study on the prokaryotic communities from a lava tube cave on La Palma Island (Spain) reported 11–24 bacteria in the ochre stalactite [51]. In addition, 11–24 bacteria have also been shown to be the key player in the biofilm community regarding biological waste and wastewater treatment [52]. Genus Subgroup_7 belongs to class Holophaga. Holophaga, a gram-negative, nonspore-forming, obligate anaerobe, can ferment several aromatic compounds to acetate [53]. Subgroup_7 bacteria have a positive correlation with soil pH (p < 0.05), and the average abundance is 2% among 26 subgroups in 87 soils examined [54]. Subgroup_7 bacteria have also been shown to have a positive correlation with nutrient availability and a negative correlation with soil acidity [55]. Bacillus spp. isolated from the rice rhizosphere could promote plant traits and protect rice from bacterial blight disease [56].
These specific ASV sequences originate from the V3-V4 segment of the 16S rRNA gene. After taxonomy assignment, the taxonomic annotation cannot fully match to the species level; therefore, their metabolic functions in soil can only be inferred and discussed based on closely related microbes. The ASVs showed differential abundance with statistical significance (Fig. 6A), suggesting different agricultural farming, such as fertilizers, pesticides, or herbicides cause the differences. With the development of metagenomics, the taxonomic annotation of these ASVs might be clarified to better explain their potential roles in Or and Nr soils, and thus would aid in the evaluation and modulation of soil fertility in paddy fields.
We used FAPROTAX, PICRUSt2, and Tax4Fun2 tools to predict the functions of microbial communities based on 16S rDNA sequences. The carbohydrate metabolism features are the only function included in all three calculated programs. They are cellulolysis (FAPROTAX, Fig. 7A), bifidobacterium shunt, glycolysis II (from fructose 6-phosphate), pyruvate fermentation to isobutanol (engineered) (PICRUSt2, Fig. 7B), butanoate metabolism, and amino sugar and nucleotide sugar metabolism (Tax4Fun2, Fig. 7C), suggesting that the carbon nutrients in both soils might be abundant. Interestingly, degradation of aromatic compounds, benzoate degradation, and aminobenzoate degradation, metabolisms related to aromatic compound degradation, showed differential abundance in nonorganic paddy soils from Tax4Fun2 analysis (Fig. 7C). The benzoate degradation pathway is related to folate synthesis, tyrosine metabolism, phenylalanine metabolism, aminobenzoate degradation, and chlorobenzene degradation [57, 58], suggesting that bacteria that degrade aromatic compounds might be capable of metabolizing aromatic pollutants [59]. At present, we do not have results to show that aromatic pollutants are present in nonorganic soils.
For functional prediction, the major concern is the limited reference databases for soil 16S rDNA, resulting in the low number of taxa assigned. The second problem is the different tools that would perform and derive different functional features [60]. We found that 79.63% of the ASVs were not assigned to any group in FAPROTAX. In PICRUSt2, the weighted nearest-sequenced taxon index (weighted NSTI) was used to evaluate the average distance for the ASVs in a given sample to a reference bacterial genome [24], and the weighted NSTI scores of this study were between 0.21 and 0.35 (> 0.15), suggesting few related references with low prediction quality (Table S2). Tax4Fun2 provides the fraction of taxonomic units that were unused (FTU) and the fraction of sequences unused (FSU) indices as quality indicators [25]. In this study, the FTU scores of all samples were larger than 0.90, whereas the FSU scores of 17 samples, among 22 samples, were larger than 0.90 (Table S2). Thus, although we used three tools to compromise the bias, the functional predictions were based on few sequences.
In conclusion, the beta diversity of NMDS revealed that microbial communities in organic and nonorganic paddy soils were significantly separated. The db-RDA analysis showed the microbial community composition in organic paddy soil was associated with higher pH and EC environment compared to the microbial community in nonorganic paddy soil. There were 23 of 23937 ASVs that showed differential abundance. Among them, g_B1-7BS (Proteobacteria), s_Sulfuricaulis limicola (Proteobacteria), g_GAL15 (p_GAL15), c_Thermodesulfovibrionia (Nitrospirota), two of f_Anaerolineaceae (Chloroflexi), and two of g_S085 (Chloroflexi) show that they are more abundant in organic soils, whereas g_11-24 (Acidobacteriota), g__Subgroup_7 (Acidobacteriota), and g_Bacillus (Firmicutes) show differential abundance in nonorganic paddy soils. These specific microbes are likely associated with soil nutrition, plants, and environmental factors. Therefore, understanding the metabolic physiology of these microbes will contribute to assessing the impact of organic farming on the microbial community in paddy soil. The existence of these microbes may also aid in the evaluation and modulation of soil fertility in paddy fields.
Data availability
The data and materials are available according to request.
References
Tveit AT, Urich T, Svenning MM (2014) Metatranscriptomic analysis of arctic peat soil microbiota. Appl Environ Microbiol 80:5761–5772. https://doi.org/10.1128/AEM.01030-14
Garoutte A, Cardenas E, Tiedje J, Howe A (2016) Methodologies for probing the metatranscriptome of grassland soil. J Microbiol Methods 131:122–129. https://doi.org/10.1016/j.mimet.2016.10.018
Buelow HN, Winter AS, Van Horn DJ, Barrett JE, Gooseff MN, Schwartz E, Takacs-Vesbach CD (2016) Microbial community responses to increased water and organic matter in the arid soils of the McMurdo Dry Valleys, Antarctica. Front Microbiol 7:1040. https://doi.org/10.3389/fmicb.2016.01040
Ligi T, Oopkaup K, Truu M, Preem JK, Nolvak H, Mitsch WJ, Mander U, Truu J (2014) Characterization of bacterial communities in soil and sediment of a created riverine wetland complex using high-throughput 16S rRNA amplicon sequencing. Ecol Eng 72:11. https://doi.org/10.1016/j.ecoleng.2013.09.007
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AK, Kent AD et al (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1:283–290. https://doi.org/10.1038/ismej.2007.53
Cesarano G, De Filippis F, La Storia A, Scala F, Bonanomi G (2017) Organic amendment type and application frequency affect crop yields, soil fertility and microbiome composition. Appl Soil Ecol 120:254–264. https://doi.org/10.1016/j.apsoil.2017.08.017
Wang JC, Song Y, Ma TF, Raza W, Li J, Howland JG et al (2017) Impacts of inorganic and organic fertilization treatments on bacterial and fungal communities in a paddy soil. Appl Soil Ecol 112:42–50. https://doi.org/10.1016/j.apsoil.2017.01.005
Kumar U, Nayak AK, Shahid M, Gupta VVSR, Panneerselvam P, Mohanty S et al (2018) Continuous application of inorganic and organic fertilizers over 47 years in paddy soil alters the bacterial community structure and its influence on rice production. Agr Ecosyst Environ 262:65–75. https://doi.org/10.1016/j.agee.2018.04.016
Nayak PK, Nayak AK, Panda BB, Lal B, Gautam P, Poonam A et al (2018) Ecological mechanism and diversity in rice based integrated farming system. Ecol Indic 91:359–375. https://doi.org/10.1016/j.ecolind.2018.04.025
Luo XS, Fu XQ, Yang Y, Cai P, Peng SB, Chen WL, Huang QY (2016) Microbial communities play important roles in modulating paddy soil fertility. Sci Rep 6:20326. https://doi.org/10.1038/srep20326
Burns KN, Kluepfel DA, Strauss SL, Bokulich NA, Cantu D, Steenwerth KL (2015) Vineyard soil bacterial diversity and composition revealed by 16S rRNA genes: differentiation by geographic features. Soil Biol Biochem 91:232–247. https://doi.org/10.1016/j.soilbio.2015.09.002
SOP (2021) SOP: Soil pH. Soil lab. University of Illinois Urbana-Champaign., Urbana, IL. https://margenot.cropsciences.illinois.edu/methods-sops. Accessed 20 June 2023
FAO (2021) Standard operating procedure for soil electrical conductivity, soil/water, 1:5. Food and Agriculture Organization of the United Nations (FAO), Rome, Italy
Kuo J, Liu D, Wang SH, Lin CH (2021) Dynamic changes in soil microbial communities with glucose enrichment in sediment microbial fuel cells. Indian J Microbiol 61:497–505. https://doi.org/10.1007/s12088-021-00959-x
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/Nmeth.3869
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590-596. https://doi.org/10.1093/nar/gks1219
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Hui Y (2021) Tutorial for microbiome analysis in R. https://www.yanh.org/2021/01/01/microbiome-r/ Accessed 20 June 2023
R Core Team R (2019) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. Accessed 20 June 20023
Xu SB, Zhan L, Tang WL, Wang QW, Dai ZH, Zhou L, et al. (2023) MicrobiotaProcess: a comprehensive R package for deep mining microbiome. The Innovation 4(2):00388. https://doi.org/10.1016/j.xinn.2023.100388
Liu C, Cui YM, Li XZ, Yao MJ (2021) microeco: an R package for data mining in microbial community ecology. FEMS Microbiol Ecol 97:fiaa255. https://doi.org/10.1093/femsec/fiaa255
Louca S, Parfrey LW, Doebeli M (2016) Decoupling function and taxonomy in the global ocean microbiome. Science 353:1272–1277. https://doi.org/10.1126/science.aaf4507
Douglas GM, Maffei VJ, Zaneveld JR, Yurgel SN, Brown JR, Taylor CM et al (2020) PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 38:685–688. https://doi.org/10.1038/s41587-020-0548-6
Wemheuer F, Taylor JA, Daniel R, Johnston E, Meinicke P, Thomas T, Wemheuer B (2020) Tax4Fun2: prediction of habitat-specific functional profiles and functional redundancy based on 16S rRNA gene sequences. Environ Microbiome 15:1–12. https://doi.org/10.1186/s40793-020-00358-7
Kuo J, Liu D, Lin CH (2023) Functional prediction of microbial communities in sediment microbial fuel cells. Bioeng-Basel 10:199. https://doi.org/10.3390/bioengineering10020199
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Liu C, Ding N, Fu Q, Brookes PC, Xu J, Guo B, Lin Y, Li H, Li N (2016) The influence of soil properties on the size and structure of bacterial and fungal communities along a paddy soil chronosequence. Eur J Soil Biol 76:8. https://doi.org/10.1016/j.ejsobi.2016.06.002
Ma TF, He XH, Chen SG, Li YJ, Huang QW, Xue C, Shen QR (2022) Long-term organic-inorganic fertilization regimes alter bacterial and fungal communities and rice yields in paddy soil. Front Microbiol 13:890712. https://doi.org/10.3389/fmicb.2022.890712
Yin XH, Chen JA, Fan L, Tao Z, Huang M, Zou YB (2020) Nitrospira bacteria in paddy soil reduced by biochar application. Agrosys Geosci Env 3:e20009. https://doi.org/10.1002/agg2.20009
Suzuki K, Takemura M, Miki T, Nonaka M, Harada N (2019) Differences in soil bacterial community compositions in paddy fields under organic and conventional farming conditions. Microbes Environ 34:108–111. https://doi.org/10.1264/jsme2.ME18101
Katayama N, Osada Y, Mashiko M, Baba YG, Tanaka K, Kusumoto Y et al (2019) Organic farming and associated management practices benefit multiple wildlife taxa: a large-scale field study in rice paddy landscapes. J Appl Ecol 56:1970–1981. https://doi.org/10.1111/1365-2664.13446
Gong SX, Zhou XS, Zhu XM, Huo JL, Faghihinia M, Li BL, Zou Y (2023) Organic rice cultivation enhances the diversity of above-ground arthropods but not below-ground soil eukaryotes. Agr Ecosyst Environ 347:108390. https://doi.org/10.1016/j.agee.2023.108390
Tang ZQ, Zhang LY, He N, Gong DK, Gao H, Ma ZB et al (2021) Soil bacterial community as impacted by addition of rice straw and biochar. Sci Rep-Uk 11:22185. https://doi.org/10.1038/s41598-021-99001-9
Li R, Khafipour E, Krause DO, Entz MH, de Kievit TR, Fernando WGD (2012) Pyrosequencing reveals the influence of organic and conventional farming systems on bacterial communities. Plos One 7:e51897. https://doi.org/10.1371/journal.pone.0051897
Heiniger RW, McBride RG, Clay DE (2003) Using soil electrical conductivity to improve nutrient management. Agron J 95:508–519. https://doi.org/10.2134/agronj2003.0508
Xu D, Liu ST, Chen Q, Ni JR (2017) Microbial community compositions in different functional zones of Carrousel oxidation ditch system for domestic wastewater treatment. Amb Express 7:1–13. https://doi.org/10.1186/s13568-017-0336-y
Cai CG, Zhao MM, Yao F, Zhu RY, Cai HY, Shao SQ et al (2022) Deoxynivalenol degradation by various microbial communities and its impacts on different bacterial flora. Toxins 14:537. https://doi.org/10.3390/toxins14080537
Kojima H, Watanabe T, Fukui M (2016) Sulfuricaulis limicola gen. nov., sp nov., a sulfur oxidizer isolated from a lake. Int J Syst Evol Micr 66:266–270. https://doi.org/10.1099/ijsem.0.000709
Brewer TE, Aronson EL, Arogyaswamy K, Billings SA, Botthoff JK, Campbell AN et al (2019) Ecological and genomic attributes of novel bacterial taxa that thrive in subsurface soil horizons. Mbio 10:10–1128. https://doi.org/10.1128/mBio.01318-19
Steger K, Kim AT, Ganzert L, Grossart HP, Smart DR (2019) Floodplain soil and its bacterial composition are strongly affected by depth. FEMS Microbiol Ecol 95:fiz014. https://doi.org/10.1093/femsec/fiz014
Li WH, Siddique MS, Liu MJ, Graham N, Yu WZ (2022) The migration and microbiological degradation of dissolved organic matter in riparian soils. Water Res 224:119080. https://doi.org/10.1016/j.watres.2022.119080
Henry EA, Devereux R, Maki JS, Gilmour CC, Woese CR, Mandelco L et al (1994) Characterization of a new thermophilic sulfate-reducing bacterium - Thermodesulfovibrio yellowstonii, gen-nov and sp-nov - its phylogenetic relationship to thermodesulfobacterium commune and their origins deep within the bacterial domain. Arch Microbiol 161:62–69. https://doi.org/10.1007/Bf00248894
Arshad A, Martins PD, Frank J, Jetten MSM, den Camp HJMO, Welte CU (2017) Mimicking microbial interactions under nitrate-reducing conditions in an anoxic bioreactor: enrichment of novel Nitrospirae bacteria distantly related to Thermodesulfovibrio. Environ Microbiol 19:4965–4977. https://doi.org/10.1111/1462-2920.13977
Sekiguchi Y, Yamada T, Hanada S, Ohashi A, Harada H, Kamagata Y (2003) Anaerolinea thermophila gen. nov., sp nov and Caldilinea aerophila gen. nov., sp nov., novel filamentous thermophiles that represent a previously uncultured lineage of the domain Bacteria at the subphylum level. Int J Syst Evol Micr 53:1843–1851. https://doi.org/10.1099/ijs.0.02699-0
Liang B, Wang LY, Mbadinga SM, Liu JF, Yang SZ, Gu JD, Mu BZ (2015) Anaerolineaceae and Methanosaeta turned to be the dominant microorganisms in alkanes-dependent methanogenic culture after long-term of incubation. Amb Express 5:1–13. https://doi.org/10.1186/s13568-015-0117-4
Liang B, Wang LY, Zhou ZC, Mbadinga SM, Zhou L, Liu JF et al (2016) High frequency of Thermodesulfovibrio spp. and Anaerolineaceae in association with Methanoculleus spp. in a long-term incubation of n-alkanes-degrading methanogenic enrichment culture. Front Microbiol 7:1431. https://doi.org/10.3389/fmicb.2016.01431
MaymoGatell X, Chien YT, Gossett JM, Zinder SH (1997) Isolation of a bacterium that reductively dechlorinates tetrachloroethene to ethene. Science 276:1568–1571. https://doi.org/10.1126/science.276.5318.1568
Zhang SY, Xiao X, Chen SC, Zhu YG, Sun GX, Konstantinidis KT (2021) High arsenic levels increase activity rather than diversity or abundance of arsenic metabolism genes in paddy soils. Appl Environ Microb 87:e01383-21. https://doi.org/10.1128/AEM.01383-21
Huber KJ, Geppert AM, Gross U, Luckner M, Wanner G, Cooper P et al (2017) Aridibacter nitratireducens sp nov., a member of the family Blastocatellaceae, class Blastocatellia, isolated from an African soil. Int J Syst Evol Micr 67:4487–4493. https://doi.org/10.1099/ijsem.0.002318
Gonzalez-Pimentel JL, Martin-Pozas T, Jurado V, Miller AZ, Caldeira AT, Fernandez-Lorenzo O et al (2021) Prokaryotic communities from a lava tube cave in La Palma Island (Spain) are involved in the biogeochemical cycle of major elements. Peerj 9:e11386. https://doi.org/10.7717/peerj.11386
Nguyen QA (2022) Microbial community analysis using next-generation sequencing and bioinformatics tools to better understand biological waste and wastewater treatment. Dissertation, University of Technology Sydney
Fukunaga Y, Ichikawa N (2014) The class holophagaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The Prokaryotes. Springer, Berlin, Heidelberg. pp 683–687. https://doi.org/10.1007/978-3-642-38954-2_161
Jones RT, Robeson MS, Lauber CL, Hamady M, Knight R, Fierer N (2009) A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J 3:442–453. https://doi.org/10.1038/ismej.2008.127
Navarrete AA, Venturini AM, Meyer KM, Klein AM, Tiedje JM, Bohannan BJM et al (2015) Differential response of acidobacteria subgroups to forest-to-pasture conversion and their biogeographic patterns in the Western Brazilian Amazon. Front Microbiol 6:1443. https://doi.org/10.3389/fmicb.2015.01443
Rajer FU, Samma MK, Ali Q, Rajar WA, Wu HJ, Raza W et al (2022) Bacillus spp.-mediated growth promotion of rice seedlings and suppression of bacterial blight disease under greenhouse conditions. Pathogens 11:1251. https://doi.org/10.3390/pathogens11111251
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M (2012) KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40:D109-114. https://doi.org/10.1093/nar/gkr988
Caspi R, Billington R, Fulcher CA, Keseler IM, Kothari A, Krummenacker M et al (2018) The MetaCyc database of metabolic pathways and enzymes. Nucleic Acids Res 46:D633–D639. https://doi.org/10.1093/nar/gkx935
Uhlik O, Wald J, Strejcek M, Musilova L, Ridl J, Hroudova M et al (2012) Identification of bacteria utilizing biphenyl, benzoate, and naphthalene in long-term contaminated soil. PLoS One 7:e40653. https://doi.org/10.1371/journal.pone.0040653
Djemiel C, Maron PA, Terrat S, Dequiedt S, Cottin A, Ranjard L (2022) Inferring microbiota functions from taxonomic genes: a review. Gigascience 11:giab090. https://doi.org/10.1093/gigascience/giab090
Acknowledgements
The authors would like to thank Chien-Ni Wei for her correction on English writing.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by W.H.W., C.Y.C., W.C., Y.W.W., F.-T.L., and C.-H.L. The first draft of the manuscript was written by J.K., D.L., and C.-H.L. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Enderson Ferreira
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kuo, J., Liu, D., Wen, W.H. et al. Different microbial communities in paddy soils under organic and nonorganic farming. Braz J Microbiol 55, 777–788 (2024). https://doi.org/10.1007/s42770-023-01218-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01218-5