Abstract
To investigate soil microbial community dynamics in sediment microbial fuel cells (MFCs), this study applied nonhydric (D) and hydric (S) soils to single-chamber and mediator-free MFCs. Glucose was also used to enrich microorganisms in the soils. The voltage outputs of both the D and S sediment MFCs increased over time but differed from each other. The initial open circuit potentials were 345 and 264 mV for the D and S MFCs. The voltage output reached a maximum of 503 and 604 mV for D and S on days 125 and 131, respectively. The maximum power densities of the D and S MFCs were 2.74 and 2.12 mW m−2, analyzed on day 50. Clustering results revealed that the two groups did not cluster after glucose supplementation and 126 days of MFC function. The change in Geobacter abundance was consistent with the voltage output, indicating that these bacteria may act as the main exoelectrogens on the anode. Spearman correlation analysis demonstrated that, in the D soils, Geobacter was positively correlated with Dialister and negatively correlated with Bradyrhizobium, Kaistobacter, Pedomicrobium, and Phascolarctobacterium; in the S soils, Geobacter was positively correlated with Shewanella and negatively correlated with Blautia. The results suggested that different soil sources in the MFCs and the addition of glucose as a nutrient produced diverse microbial communities with varying voltage output efficiencies.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Microbial fuel cells (MFCs), in which microorganisms oxidize organic matter to produce electrical energy, have emerged as a new technology for electric energy generation. Electrons are supplied from reduced substrates that are metabolically oxidized by the bacteria and transferred to the anode through the following three mechanisms: direct transfer via conductive pili; direct transfer via redox-active proteins; and indirect transfer via electron shutters. The electrons are then transferred to the cathode via a resistor. Protons derive from the oxidation of organic matter, move to the cathode, and combine with electrons and oxygen to form water on the surface of the cathode. This dual-chamber device is separated by a proton exchange membrane between the cathode and anode to prevent oxygen from diffusing to the anode [1]. MFCs can be operated with marine sediments, sludge, wastewater, or other aquatic sediments. The organic matter provided by sediments or wastewater supplies nutrients for the growth of microorganisms, achieving the biological decontamination of the environment and production of energy [1]. Studies have focused on sustainable or enhanced electricity generation in, for example, the use of microalgae Spirulina platensis acting as the substrate and use of a silver nanoparticle–activated carbon composite serving as the cathode catalyst to sustain the power output [2], which are supplemented with iron powder in rice paddy MFCs to enhance electricity generation [3].
Exoelectrogenic bacteria, referred to as exoelectrogens, are microorganisms that can transfer electrons extracellularly. For example, Shewanella oneidensis, Geobacter metallireducens, and G. sulfurreducens are exoelectrogens that use cytochromes as the electron transfer proteins or oxidoreductive enzymes to catalyze the reduction of reactive substrates; such species are responsible for the power produced in MFCs [4, 5]. The electron transfer pathways in the anode occur in these exoelectrogenic bacteria, which supply electrons to the anode through direct electron transfer via cytochromes, direct electron transfer via nanowires, and mediated electron transfer via soluble electron shuttles [1]. However, if a complex mixture of bacteria are applied to MFCs, electron transfer could occur through quorum-sensing chemicals, minerals (which serve as mediators), cell–cell communication; for example, the oxidation of methane by anaerobic methanotrophic archaea is linked to sulfate reduction by sulfate-reducing bacteria [4, 5]. In a previous study, methane as a substrate for MFCs was analyzed through the syntrophic association between Geobacter and methanotrophs [6]. For sediment-derived MFCs or other environmentally derived mixed-culture MFCs, the power density could be reduced by nonexoelectrogenic bacteria or nonactive cells that disrupt the electrical conductivity of the biofilm [4].
In this study, microbial communities in MFCs inoculated with sediment or sludge were investigated. The convergent adaptation of anodic microbial communities was the likely phenomenon through which Geobacteraceae [7] and Desulfobulbaceae [8] predominantly interact in anodic biofilms and was positively correlated with electricity generation efficiency. This syntrophic interaction suggested that Geobacter may work with other microorganisms to generate electricity; for example, Sporomusa could convert methanol into acetate, which is then utilized by Geobacter [9, 10]. Alcaligenes monasteriensis, Comamonas denitrificans, and Dechloromonas sp. are also potential exoelectrogenic bacteria from anaerobic sludge [11]. Different microorganisms have been proposed to adapt and generate electricity in MFCs [12]. An anodic biofilm community could be altered in response to different nutrients [13], and the surface potential and carbon sources can also affect microbial communities [14]. Therefore, different sediments may contain different exoelectrogens, with other microorganisms providing supporting material, such as oxidized-form mediators, oxygen-removing species, or nutrients for the microbial community to adapt to the power output. Thus far, applied sediments have originated from marine sediment, sludge, or rice paddy fields, all of which can be classified as hydric soils [15]. However, studies on the application of nonhydric soils, such as nonflooded or nonsaturated soil, in sediment MFCs to modulate the dynamics of microbial communities have been limited.
In this study, we applied nonhydric (D) and hydric (S) soils in single-chamber and mediator-free sediment MFCs for microbial community analysis. The voltage was recorded, and the soil was collected for chromosomal DNA preparation. The 16S rDNA sequence was then analyzed, followed by a microbe comparison and biodiversity analysis. This study investigated how the microbial communities and electricity outputs were affected by different soil sources and the effects from the addition of glucose as a nutrient.
Material and Methods
Sediment MFC Construction and Operation
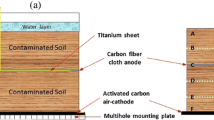
The nonhydric and hydric soils used in this study were collected from land and from the ecological pool of a university campus (24°00′N, 120°36′E) in Dacun Township, Changhua County, Taiwan. A single-chamber, mediator-free sediment MFC device was designed. The soil was placed in a transparent plastic container (23 × 14 × 12 cm3) at a depth of 5 cm; the soil volume was 1680 mL. A phosphate buffer solution with pH 7.2 containing 0.137 M NaCl, 0.0027 M KCl, 0.01 M Na2HPO4, and 0.0018 M KH2PO4 was added to a depth of 10 cm. Two woven carbon fiber sheets (13.5 × 13.5 cm2; Skyline Carbon, New Taipei, Taiwan) were used as the anode (−) and cathode (+). The anode was buried in the soil (2 cm depth), and the cathode was floated under the buffer solution. To sustain MFC operation, 20 mL of glucose solution (0.1 g/mL−1) was applied once, and it acted as a nutrient when the voltage output decreased to almost 0 on day 90. The voltage and current changes were recorded using a Prova 803 dual-channel digital meter (TES Electrical Electronic, Taipei, Taiwan), and the data were downloaded using a RS-232 connection to a computer. The performance of the MFCs, corresponding to the changes in voltage and current, was analyzed at the following resistance sequence: 1 MΩ, 50 kΩ, 10 kΩ, 1 kΩ, 300 Ω, 150 Ω, and 82 Ω.
DNA Extraction, Polymerase Chain Reaction (PCR), and Sequencing
On days 14, 90, 105, and 126, soil samples from the same location were collected: samples D2, D3, D4, and D5 denote the nonhydric MFCs, and S2, S3, S4, and S5 denote hydric MFCs. D1 and S1 represent the initial soil samples. The water in the samples was first removed and placed in a beaker, a spatula was then used to move the soil close to the anode, and the water was finally readded. The soils were stored at −20 °C. Bacterial genomic DNA (gDNA) was extracted from 0.5 g of soil using an UltraClean soil DNA isolation kit (MO BIO Laboratories, Carlsbad, CA, USA) according to the manufacturer's instructions, and DNA sequencing was performed at a facility owned by BIOTOOLS (New Taipei, Taiwan). For the 16S rRNA gene sequencing, the V3–V4 region was amplified with a specific primer set (319F: 5′-CCTACGGGNGGCWGCAG-3′, 806R: 5′-GACTACHVGGGTATCTAATCC-3′, according to the 16S Metagenomic Sequencing Library preparation procedure; Illumina, San Diego, CA, USA). In brief, 12.5 ng of gDNA was used for a polymerase chain reaction (PCR), performed with KAPA HiFi HotStart ReadyMix (Roche, Basel, Switzerland) under the following PCR conditions: 95 °C for 3 min; 25 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s; 72 °C for 5 min; it was held at 4 °C. The PCR products were monitored while on 1.5% agarose gel. Samples with bright main bands of approximately 500 bp were chosen and purified using AMPure XP beads for subsequent library preparation.
The sequencing library was prepared according to the 16S Metagenomic Sequencing Library preparation procedure (Illumina). In brief, secondary PCR was performed using the 16S rRNA V3–V4 region PCR amplicon and Nextera XT Index Kit with dual indices and sequencing adapters (Illumina). The indexed PCR product quality was assessed on a Qubit 4 fluorometer (Thermo Scientific, Waltham, MA, USA) and Qsep100TM system. Equal amounts of the indexed PCR product were mixed to generate the sequencing library. Finally, the library was sequenced on an Illumina MiSeq platform, and paired 300‐bp reads were generated.
Processing and Analysis of Sequence Data
The processing and analysis of sequence data were performed by BIOTOOLS (New Taipei, Taiwan), as previously described [16]. For each representative sequence, the Greengenes Database [17] was processed by the RDP classifier (version 2.2) [18] algorithm to annotate taxonomic information.
Alpha diversity was applied to analyze the complexity of species diversity for each sample through five indices, namely the observed species and Chao1 and the Shannon, Simpson, and accumulated cyclone energy indices. Data on these indices were obtained with QIIME (version 1.7.0) and visualized using R software (version 2.15.3). A heatmap of dominant genera was generated in R (version 3.4.1) with the ComplexHeatmap function. Hierarchical clustering was conducted using the Spearman correlation [19, 20]. Principal coordinate analysis (PCoA) was performed to obtain the principal coordinates and visualize complex, multidimensional data. A distance matrix of weighted or unweighted UniFrac distances among previously obtained samples was transformed to a new set of orthogonal axes, in which the maximum variation factor is represented by the first principal coordinate, the second maximum variation factor is represented by the second principal coordinate, and so on. PCoA analysis was conducted per a previously described method [16]. The Spearman correlation of dominant genera was generated in R (version 3.4.1) using the recorr and corrplot functions [20]. Nucleotide sequences determined in the study were entered into the National Center for Biotechnology Information Sequence Read Archive database under the accession numbers SRR11992844, SRR11992843, SRR11992842, SRR11992841, SRR11992840, SRR11992839, SRR11992838, SRR11992837, SRR11992836, SRR11992835, and SRR11992835 for samples D1, D2, D3, D4, D5, S1, S2, S3, S4, and S5, respectively.
Results
Electricity Generation
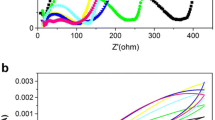
In this study, the D and S soil samples collected on campus were applied as a source of microorganisms and nutrients for sediment MFCs. Figure 1 illustrates the daily voltage output fluctuation results under a loading of 1 kΩ external resistance. The voltage output from the MFCs containing the D and S soils both increased in the first month, and the voltage output of S was higher than that of D. In the initial stage, the microorganisms in the D samples encountered environmental changes, which induced flooding stress; thus, oxygen became the limiting factor. Gradually, the voltage output of the MFC with D soils was higher than that of the MFC with S soils after 30 days, which was likely caused by the microbial community changes driven by electrochemical activity. After the voltage output consistently declined for another 30 days, on the 90th day, 20 mL of glucose solution (0.1 g mL−1) was applied. Notably, the voltage outputs of the two types of MFCs were reversed; the voltage output increased in the MFC with S soils but decreased in the MFC with D soils, where the water turned black and turbid (data not shown). This phenomenon persisted for nearly a month, after which, although fluctuations were present in both MFCs, the voltage outputs did not drop to zero before the data ceased to be recorded on day 376. Polarization analysis (Fig. 2) on day 50 indicated that the MFC of D soils exhibited a slightly higher power density than did the MFC of S soils. The maximum power densities of the MFCs with D and S soils were 2.74 and 2.12 mW m−2, respectively. A higher internal resistance, the lack of a platinum catalyst, and the use of a one-chamber device in our simple system were likely responsible for the low power density output and contributed to the instability in voltage output. Thus, nonhydric soil can be applied in a sediment MFC, and it performs more effectively than hydric soil does in a sediment MFC.
Phylogenetic and Diversity Analysis
We collected the soils near the anodes, and the microbial communities were examined. Figure 3 presents a comparison between the two MFCs on the basis of the percent composition of different bacterial varieties. The predominant microorganisms in the original D soil (D1) were Proteobacteria (45.4%), Firmicutes (21.5%), Acidobacteria (7.7%), Bacteroidetes (5.2%), and Actinobacteria (3.8%), whereas those in the original S soil (S1) were Euryarchaeota (26.8%), Proteobacteria (26.7%), Firmicutes (21.9%), Acidobacteria (4.4%), and Nitrospirae (3.5%). After 126 days of sediment MFC operation, the predominant microorganisms in the D5 soil were Firmicutes (41.8%), Proteobacteria (25.1%), Euryarchaeota (6.5%), Acidobacteria (6.2%), and Bacteroidetes (4.2%). By contrast, the predominant microorganisms in the S5 soil were Proteobacteria (53.6%), Firmicutes (16.6%), Acidobacteria (8.4%), Bacteroidetes (3.1%), and Nitrospirae (3.0%). The abundances of several microorganisms in the D and S soils changed after MFC manipulation. For example, Euryarchaeota and Firmicutes abundances increased in the D soils but decreased in the S soils. Conversely, Alphaproteobacteria and Gammaproteobacteria abundances decreased in the D soils but increased in S the soils. Based on the microbial community comparison for hydric samples S4 and S3, the abundances of Bacteroidetes, Deltaproteobacteria, Gammaproteobacteria, and Firmicutes were computed to be 150.2%, 116.2%, 139.2%, and 114.8%, respectively (Fig. 3); their higher abundance in S4 than in S3 suggested that these three bacteria were activated by the glucose nutrient. For the nonhydric samples, the abundances of Acidobacteria, Betaproteobacteria, Deltaproteobacteria, Euryarchaeota, and Gammaproteobacteria were 170.6%, 207.6%, 162.8%, 384.6%, and 146.5%, respectively; they were higher in D4 than in D3. However, Alphaproteobacteria, Bacteroidetes, and Firmicutes abundances decreased by 42.9%, 52.8%, and 63%, respectively (Fig. 3), after the addition of glucose. These differences in the microbial community between the D and S samples may account for the voltage drop in the D4 MFC after glucose addition.
The Shannon diversity index of the soil microbial communities increased after glucose supplementation in the D3 (5.256) and D4 (6.405) nonhydric samples (Table 1); however, in the S3 and S4 hydric samples, the Shannon index decreased from 6.458 to 6.438, respectively. After comparing the community richness (Chao1) and diversity (Shannon), we determined that the Chao1 index increased but the Shannon index decreased from D1 to D5. However, the microbial communities in the S samples exhibited different patterns wherein both the Chao1 and Shannon indices increased (Table 1).
Microbial Community Analysis
PCoA based on the weighted and unweighted UniFrac distances revealed two separated groups of microbial communities from the D and S soils; the within-group samples clustered together (Fig. 4), suggesting the diversity of both microbial communities after the MFC operation. No convergent dynamic changes in soil microbial communities were recorded. Based on the unweighted UniFrac distance, the last sample (D5) of D soil was located at the same coordinate as the first sample (S1) of S soil (Fig. 4), suggesting that these microbial communities were similar. The microbial community in the D soil may have changed to become a community similar to that in S soil.
Principal coordinate analysis (PCoA) of sequence data. The PCoA plot was generated from the weighted and unweighted UniFrac analysis. The x- and y-axes indicate the first and second coordinates, respectively, and the percentage on each axis represents the contribution of discrepancy among samples. Black and red symbols represent D and S microbial communities, respectively
In the heatmap and clustering analysis, the microbial genera and quantities were used to create a graphical clustering representation (Fig. S1). The results demonstrated that the microbial communities in the D and S soils clustered separately, suggesting that the disparate electricity generation could be attributed to the differences in the microbial communities. Geobacter exhibited increasing trends in both the D and S samples consistent with the power output, except for sample D4 (Fig. S1). Both Lachnospira and Roseburia exhibited contrasting distributions after glucose addition but were not synchronized with the voltage output. Anaeromyxobacter, Azospirillum, Clostridium, and Faecalibacterium were dominant, but the change in abundance for these genera was not consistent with the voltage output (Fig. S1). Thus, Geobacter could be responsible for the electricity generation.
To analyze the relationships between Geobacter and other bacteria, we chose the dominant 50 genera from the D and S samples for Spearman rank correlation analysis (P < 0.05). The results suggested that in the D samples, Geobacter was positively correlated with Dialister (rs = 0.9) and negatively correlated with Bradyrhizobium (rs = − 0.9), Kaistobacter (rs = − 0.9), Phascolarctobacterium (rs = − 0.9), and Pedomicrobium (rs = − 1.0; Fig. S2A). In the S samples, Geobacter was positively correlated with Shewanella (rs = 0.9) and negatively correlated with Blautia (rs = − 0.9; Fig. S2B).
Discussion
A study on rice paddy field-rhizosphere sediment MFCs with graphite felt anodes and cathodes with a platinum catalyst loaded on the cathode had a maximum power density of 19 ± 3.2 mW m−2 [9]. In a combined plant and sediment MFC two-chamber device with a graphite anode and cathode, the power density reached 100 mW m−2 and it operated for 119 days [21]. Citrus waste was explored as the feedstock for MFC operation, with the power density reaching 71.1 mW m−2 [22]. Raw algae biomass with acetate serving as the cosubstrate would enhance the power density to 410 mW m−2, in contrast to the 230 mW m−2 for MFCs without acetate [23]. In addition to the supplementation of a catalyst, a two-chamber design improves the power output from our sediment MFCs and the optimal nutrients serving as substrates enhance the electricity generation.
Glucose and glutamate were applied to enrich electrochemically active bacteria, and the results demonstrated that Gammaproteobacteria (36.5%), Firmicutes (27%), and Deltaproteobacteria (15%) were the significant populations at the electrode [24]. In glucose-fed MFCs, Deltaproteobacteria (27.7%), Gammaproteobacteria (14.5%), and Firmicutes (6.9%) are the main bacteria in graphite felt anodes [9]. Consistent with the results of other studies, our experimental results obtained after glucose addition indicated that Deltaproteobacteria and Gammaproteobacteria increased in the D and S samples and that Firmicutes increased only in the S soil (Fig. 3). Jung and Regan [25] applied acetate, lactate, and glucose as electron donors and compared the anode bacterial communities and performance in MFCs, reporting that Firmicutes was identified only in glucose-fed MFCs. Thus, certain nutrients in soil affect the microbial community. Soil nutrients and carbon availability affect the microbial community in response to glucose amendment, resulting in distinct variations in glucose-utilizing microbes [26]. Furthermore, high glucose amendment enhanced soil biofilm formation, and the Shannon diversity of the soil biofilm communities was significantly improved by 18.2% [27]. The diversity results of this study (Table 1) suggested that the species richness increased in both types of soil after glucose enrichment in the sediment MFCs. However, in the D samples, the abundances of specific bacteria varied to different extents, resulting in an overall decrease in diversity. In the S samples, the microbial diversity increased.
Geobacteraceae [7, 8, 13, 14] and Desulfobulbaceae [8] are predominantly present in anodic biofilms and are positively correlated with electricity generation efficiency; convergent adaptation is the likely mechanism in anodic microbial communities. However, this study demonstrated the diversity of microbial community formation in the D and S soils after 126 days of MFC function and glucose supplementation. Other sediments [28] or inoculums [29] in MFCs could yield divergent microbial communities because of the diverse microbial community structures of the source materials. The microbial community dynamics could also be affected by the nutrient supply [14]. A study on soil bio-electrochemical remediation determined that microbial community dynamics could be affected by MFC operation and that Geobacter and Thermincola bacteria could be responsible for bioelectricity generation [30]. Thus, the original soil microbial community and nutrients play roles in the microbial diversity of sediment MFCs in a competitive or syntrophic relationship. Microorganisms that can respire at the most negative anode potential outcompete other strains that are unable to adjust to the negative potential [4]. Syntrophic interactions have been discussed in a study on anode biofilms from glucose- and acetate-fed MFCs, in which distinct microbial communities were prevalent [9, 10]. Glucose addition enhances voltage output and power density, suggesting that the presence of glucose accelerates the acclimation of the electrochemically active microbial community [31]. These studies have proposed that the nutrient content in soils affects the microbial community, which maintains a dynamic equilibrium. In this study, glucose addition and power output were two variables for two different soils, D and S.
Although the change in abundance matches the voltage output, more MFC replicates are necessary to compare the performance of hydric and nonhydric soils and to determine the effect of glucose addition to validate Geobacter as the major exoelectrogen in the MFC. For mixed communities, syntrophic and competitive interactions are the likely forces driving electrochemical reactions. Correlation analysis could reveal the relationships between Geobacter and other microorganisms. In D soils, Geobacter was positively correlated with Dialister and negatively correlated with Bradyrhizobium, Kaistobacter, Phascolarctobacterium, and Pedomicrobium. Dialister is a gram-negative and anaerobic or microaerophilic bacterium whose metabolic end-products comprise small amounts of acetic, propionic, and lactic acids [32]; in addition, Geobacter assimilates propionate [33]. In the S soils, Geobacter was positively correlated with Shewanella and negatively correlated with Blautia. Shewanella can accumulate around insoluble electron acceptors; its ability to accumulate on a mineral surface significantly strongly correlates with the presence of critical extracellular electron transport genes [34]. The microbial interactions in this mixed community are complex and unpredictable, undergoing rapid dynamic change. High-throughput sequencing enabled microbial analysis on a macro scale. However, a functional metatranscriptomic analysis would identify soil microorganisms and protein activity during the sampling period for a more comprehensive interpretation of regulatory mechanisms [35].
In conclusion, we demonstrated that nonhydric and hydric soils can be microbial sources for sediment MFCs. Soils with distinct microbial populations after MFC operation produce diverse microbial communities with different voltage output efficiencies. A spearman correlation analysis suggested that Geobacter was positively correlated with Dialister and Shewanella. Therefore, if the environment is suitable for the formation of microbial communities, they may generate power sustainably and optimally.
References
Slate AJ, Whitehead KA, Brownson DA, Banks CE (2019) Microbial fuel cells: an overview of current technology. Renew Sustain Energy Rev 101:22
Sallam ER, Khairy HM, Elnouby MS, Fetouh HA (2021) Sustainable electricity production from seawater using Spirulina platensis microbial fuel cell catalyzed by silver nanoparticles-activated carbon composite prepared by a new modified photolysis method. Biomass Bioenergy 148:106038. https://doi.org/10.1016/j.biombioe.2021.106038
Matsumoto A, Nagoya M, Tsuchiya M, Suga K, Inohana Y, Hirose A, Yamada S, Hirano S, Ito Y, Tanaka S, Kouzuma A, Watanabe K (2020) Enhanced electricity generation in rice paddy-field microbial fuel cells supplemented with iron powders. Bioelectrochemistry 136:107625. https://doi.org/10.1016/j.bioelechem.2020.107625
Logan BE (2009) Exoelectrogenic bacteria that power microbial fuel cells. Nat Rev Microbiol 7:375–381. https://doi.org/10.1038/nrmicro2113
Shi L, Dong H, Reguera G, Beyenal H, Lu A, Liu J, Yu HQ, Fredrickson JK (2016) Extracellular electron transfer mechanisms between microorganisms and minerals. Nat Rev Microbiol 14:651–662. https://doi.org/10.1038/nrmicro.2016.93
Kondaveeti S, Mohanakrishna G, Lee J-K, Kalia VC (2019) Methane as a substrate for energy generation using microbial fuel cells. Indian J Microbiol 59:121–124
Bond DR, Holmes DE, Tender LM, Lovley DR (2002) Electrode-reducing microorganisms that harvest energy from marine sediments. Science 295:483–485. https://doi.org/10.1126/science.1066771
Holmes DE, Bond DR, O’Neil RA, Reimers CE, Tender LR, Lovley DR (2004) Microbial communities associated with electrodes harvesting electricity from a variety of aquatic sediments. Microb Ecol 48:178–190. https://doi.org/10.1007/s00248-003-0004-4
Kouzuma A, Kasai T, Nakagawa G, Yamamuro A, Abe T, Watanabe K (2013) Comparative metagenomics of anode-associated microbiomes developed in rice paddy-field microbial fuel cells. PLoS ONE 8:e77443. https://doi.org/10.1371/journal.pone.0077443
Yamamuro A, Kouzuma A, Abe T, Watanabe K (2014) Metagenomic analyses reveal the involvement of syntrophic consortia in methanol/electricity conversion in microbial fuel cells. PLoS ONE 9:e98425. https://doi.org/10.1371/journal.pone.0098425
Sun Y, Zuo J, Cui L, Deng Q, Dang Y (2010) Diversity of microbes and potential exoelectrogenic bacteria on anode surface in microbial fuel cells. J Gen Appl Microbiol 56:19–29. https://doi.org/10.2323/jgam.56.19
Beecroft NJ, Zhao F, Varcoe JR, Slade RC, Thumser AE, Avignone-Rossa C (2012) Dynamic changes in the microbial community composition in microbial fuel cells fed with sucrose. Appl Microbiol Biotechnol 93:423–437. https://doi.org/10.1007/s00253-011-3590-y
Zhang H, Chen X, Braithwaite D, He Z (2014) Phylogenetic and metagenomic analyses of substrate-dependent bacterial temporal dynamics in microbial fuel cells. PLOS ONE 9:e107460. https://doi.org/10.1371/journal.pone.0107460
Ishii S, Suzuki S, Norden-Krichmar TM, Phan T, Wanger G, Nealson KH, Sekiguchi Y, Gorby YA, Bretschger O (2014) Microbial population and functional dynamics associated with surface potential and carbon metabolism. ISME J 8:963–978. https://doi.org/10.1038/ismej.2013.217
Inglett PW, Reddy KR, Corstanje R (2005) Anaerobic soils. In: Hillel D (eds) Encyclopedia of soils in the environment. Elsevier, pp 72–78
Yang SF, Huang HD, Fan WL, Jong YJ, Chen MK, Huang CN, Chuang CY, Kuo YL, Chung WH, Su SC (2018) Compositional and functional variations of oral microbiota associated with the mutational changes in oral cancer. Oral Oncol 77:1–8. https://doi.org/10.1016/j.oraloncology.2017.12.005
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072. https://doi.org/10.1128/AEM.03006-05
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Gu Z, Eils R, Schlesner M (2016) Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32:2847–2849. https://doi.org/10.1093/bioinformatics/btw313
R Core Team (2017) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Timmers RA, Strik DP, Hamelers HV, Buisman CJ (2010) Long-term performance of a plant microbial fuel cell with Spartina anglica. Appl Microbiol Biotechnol 86:973–981. https://doi.org/10.1007/s00253-010-2440-7
Kondaveeti S, Mohanakrishna G, Kumar A, Lai C, Lee J-K, Kalia VC (2019) Exploitation of citrus peel extract as a feedstock for power generation in microbial fuel cell (MFC). Indian J Microbiol 59:476–481. https://doi.org/10.1007/s12088-019-00829-7
Kondaveeti S, Mohanakrishna G, Pagolu R, Kim I-W, Kalia VC, Lee J-K (2019) Bioelectrogenesis from raw algal biomass through microbial fuel cells: effect of acetate as co-substrate. Indian J Microbiol 59:22–26. https://doi.org/10.1007/s12088-018-0769-2
Choo YF, Lee J, Chang IS, Kim BH (2006) Bacterial communities in microbial fuel cells enriched with high concenrations of glucose and glutamate. J Microbiol Biotechnol 16:4
Jung S, Regan JM (2007) Comparison of anode bacterial communities and performance in microbial fuel cells with different electron donors. Appl Microbiol Biotechnol 77:393–402. https://doi.org/10.1007/s00253-007-1162-y
Arcand MM, Levy-Booth DJ, Helgason BL (2017) Resource legacies of organic and conventional management differentiate soil microbial carbon use. Front Microbiol 8:2293. https://doi.org/10.3389/fmicb.2017.02293
Wu Y, Cai P, Jing X, Niu X, Ji D, Ashry NM, Gao C, Huang Q (2019) Soil biofilm formation enhances microbial community diversity and metabolic activity. Environ Int 132:105116. https://doi.org/10.1016/j.envint.2019.105116
Song N, Jiang H, Yan Z (2019) Contrasting effects of sediment microbial fuel cells (SMFCs) on the degradation of macrophyte litter in sediments from different areas of a shallow eutrophic lake. Appl Sci 9:13
Yang J, Cheng S (2019) Effects of using anode biofilm and cathode biofilm bacteria as inoculum on the start-up, electricity generation, and microbial community of air-cathode single-chamber microbial fuel cells. Pol J Environ Stud 28:8
Li X, Li Y, Zhang X, Zhao X, Chen X, Li Y (2020) The metolachlor degradation kinetics and bacterial community evolution in the soil bioelectrochemical remediation. Chemosphere 248:125915. https://doi.org/10.1016/j.chemosphere.2020.125915
Li X, Wang X, Wan L, Zhang Y, Li N, Li D, Zhou Q (2016) Enhanced biodegradation of aged petroleum hydrocarbons in soils by glucose addition in microbial fuel cells. J Chem Technol Biotechnol 91:9
Jumas-Bilak E, Jean-Pierre H, Carlier JP, Teyssier C, Bernard K, Gay B, Campos J, Morio F, Marchandin H (2005) Dialister micraerophilus sp. nov. and dialister propionicifaciens sp. nov., isolated from human clinical samples. Int J Syst Evol Microbiol 55:2471–2478. https://doi.org/10.1099/ijs.0.63715-0
Gan Y, Qiu Q, Liu P, Rui J, Lu Y (2012) Syntrophic oxidation of propionate in rice field soil at 15 and 30 °C under methanogenic conditions. Appl Environ Microbiol 78:4923–4932. https://doi.org/10.1128/AEM.00688-12
Harris HW, Sanchez-Andrea I, McLean JS, Salas EC, Tran W, El-Naggar MY, Nealson KH (2017) Redox sensing within the genus Shewanella. Front Microbiol 8:2568. https://doi.org/10.3389/fmicb.2017.02568
Carvalhais LC, Dennis PG, Tyson GW, Schenk PM (2012) Application of metatranscriptomics to soil environments. J Microbiol Methods 91:246–251. https://doi.org/10.1016/j.mimet.2012.08.011
Acknowledgements
The authors would like to thank Mr. Cheng-Yu Lin for his hard work and Mrs. Glory Chen (C.-H.L.’s wife) for her correction on English writing. This manuscript was edited by Wallace Academic Editing. This research was supported partly by Grant ORD-104074 from Da-Yeh University to C.-H. L.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kuo, J., Liu, D., Wang, SH. et al. Dynamic Changes in Soil Microbial Communities with Glucose Enrichment in Sediment Microbial Fuel Cells. Indian J Microbiol 61, 497–505 (2021). https://doi.org/10.1007/s12088-021-00959-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-021-00959-x