Abstract
Cold-active lipases are presently employed extensively in the detergent, chemical intermediate, fine chemical, food, and pharmaceutical industries. Seven cold-adaptive bacteria were isolated from the Mediterranean Sea near Alexandria, Egypt, and tested for their ability to produce cold-active lipase, with the highest activity at 10 °C. The most potent isolate was Pseudomonas sp. A6. To determine the most important variables, the bacterium was exposed to a necessary medium component and environmental factor screening using a single factor-at-a-time approach, followed by a multifactorial Plackett-Burman design strategy. After purification and characterization, the optimal activity levels for the cold-active lipase were figured out. Inoculation of Pseudomonas A6 under near optimum conditions using medium consisting of (g/L) peptone 7.14; soybean oil 7.5% (v/v); K2HPO4, 0.4; MgSO4, 0.1; glucose 2; pH 8; and temperature 10 °C led to a maximum lipase activity anticipated to be 23.36 U/mL. Purified lipase showed the best activity and thermal stability at a pH of 8 and a temperature of 10 °C. The Pseudomonas A6 lipase tolerated the monovalent ions, while greater valence ions did not.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The marine environment is rich in psychrotolerant bacteria. These bacteria have developed numerous adaptation mechanisms to help them withstand the severe impacts of such conditions [1, 2]. Lipases are hydrolytic enzymes that belong to the triacylglycerol hydrolases (EC: 3.1.1.3) family [3]. Even though lipases have been extensively investigated, cold-active lipases have not received much attention [4, 5]. Bacteria that can live at low temperatures generally have cold-active lipases [6]. Cold-adapted lipases are attractive biocatalysts in biotechnology because they are used as food additives or in laundry detergents to enable efficient washing at low temperatures [7]. They are also gaining popularity as a tool for producing extremely unstable compounds at low temperatures in the organic synthesis of chemical intermediates [6]. Moreover, in the manufacturing of fine chemicals, as well as in the food and pharmaceutical industries [8]. Cold-adapted was successful in catalyze the production butyl and oleic esters synthesis which has a bright future in biofuel and food industries [9, 10]. Microbial lipases had a $400 million market in 2017, and it is expected to increase to $590 million by 2023 [8]. Cold-active lipases are primarily produced extracellularly, making them suitable for fermentation and downstream purification processes. However, the physiology of the producing strain, environmental factors, and nutritional components such as carbon and nitrogen sources, inducer presence or absence, and so on are affecting the enzyme production [11, 12]. For instance, the type and concentration of carbon and nitrogen sources, as well as the aeration and pH value of the growth medium, are all factors that influence the lipase production [13]. The Plackett-Burman design was used before for screening of the most crucial factors in a biological process [14]. The isolation source is crucial to isolate psychrophilic microbes and is usually isolated from high altitude or from cold environments. However, the distribution of these microbes is not limited to such environments. Due to the change in the global weather, Alexandria city, Egypt, has experienced unusual cold weather in the winter during the past few years. This might have influenced the microbial structure in this geographical area. In this study, we successfully isolated a cold-adaptive Pseudomonas strain from the Mediterranean Sea. The growth requirements and production of cold-active lipase were examined using a multifactorial approach to figure out the major components that govern the lipase production. Furthermore, the cold-active lipase produced was purified and characterized for maximum effectiveness.
Materials and methods
Isolation of psychrotolerant bacteria and screening for cold-active lipase production
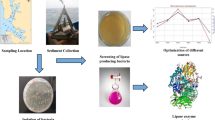
Seawater samples were collected following the method described by Arayes et al. [15] from different depths in clean sterile screw cap bottles. Bottles with a capacity of 250 mL were opened 15 cm and 1 m below the water’s surface in the Mediterranean Sea in Alexandria, Egypt, beside the National Institute of Oceanography and Fisheries (31°12′44.7″N 29°53′07.2″E). The water temperature was 22 °C. Within 4 h of the collection, samples were transported to the laboratory at 4 °C and processed. For isolation, seawater medium (SWM) was used, which included the following ingredients expressed in g/L: peptone 5; yeast extract 2.5; glucose 1; K2HPO4, 0.2; MgSO4, 0.05; and agar-agar 15. The components were dissolved in 75% of seawater, pH 7.2, and an antifungal agent, Mycostatin (1 mL/L), was added. SWM plates were surface inoculated with 1 ml of water samples and incubated at 10 °C for 3 to 7 days [16]. Colonies that grew on agar plates were picked up with a bacteriological needle and purified on the same medium using the traditional spatial streaking method. The pure bacterial isolates were then tested for their ability to grow in liquid SWM at pH 7 for 2 days while shaking at different temperatures (5 °C, 10 °C, 20 °C, and 30 °C). The isolates that showed high growth at 10 °C were selected for screening of lipase production. The screening was carried on SWM having 0.2% tributyrin, 1% Arabic gum, and 2% (w/v) agar [17]. After 7 days of incubation, a clear zone around the growth was considered a positive result.
DNA extraction and molecular identification
The genomic DNA of the chosen isolate was extracted from a 2 mL of overnight bacterial culture using the technique outlined by Sambrook et al. [18]. The 16s rDNA was amplified using universal primers for 16s rRNA gene. The primers used: F27 (5′-AGAGTTTGATCMTGGCTCAG-3′) and 1492R (5′-TACGGYTACCTTGTTACGACTT-3′) [19]. In 50 μL PCR reaction buffer, 30 picomoles of each primer, 10 μL of chromosomal DNA, 200 mg dNTPs, and 2.5 units of Taq polymerase were mixed. The PCR was carried out for 30 cycles at 94 °C for 1 min, 55 °C for 1 min, and 72 °C for 2 min. The product was purified using a QIAquick (Qiagen) PCR purification kit following the manufacturer’s instructions, after the size of the PCR product was confirmed using 1% agarose gel electrophoresis. DNA sequences were obtained using an ABI PRISM 377 DNA Sequencer. Unipro UGENE integrated bioinformatics software was used to edit the obtained sequences [20]. The sequence was BLASTed against the NCBI non-redundant nucleotide database and the phylogenetic relationships were constructed using MEGA X [21].
Culture conditions for lipase production
Seed culture (5 mL OD600=0.8–1.0) was inoculated into a 250-mL Erlenmeyer flask having 50 mL of SWM broth (pH 7) and incubated at 10 °C for 3 days under shaking conditions (160 rpm). For growth curve monitoring, samples were taken every 2 h to measure the optical density at 600 nm. After centrifugation at 10,000 g for 10 min, the cell-free culture supernatant was used to measure lipase activity.
Lipase activity assay
The breakdown of p-nitrophenyl laurate and liberation of p-nitrophenol was measured spectrophotometrically at 420 nm [22]. Briefly, 0.1 mM phosphate buffer and p-nitrophenyl laurate were freshly prepared in ethanol. Then, 700 μL phosphate buffer and 100 μL of p-nitrophenyl laurate solution were added to 50 μL of the cell-free extract. After 30 min at 10 °C, 250 μL of Na2CO3 was added and centrifuged for 20 min at 13,000 rpm. The optical density of the resulting supernatant was measured at 420 nm against a blank. The blank was prepared using distilled water instead of cell-free supernatant. A standard curve was prepared using standard solutions based on the weight of pure p-nitrophenol ester. Under test conditions, one unit of lipase activity was defined as the quantity of enzyme that released one micromole of p-nitrophenol per min per milliliter.
Factors affecting lipase production
The effects of several organic nitrogen sources (beef extract, yeast extract, malt extract, and peptone) on lipase synthesis were investigated either individually single-factor-at-a-time approach or in combination. The potential nitrogen source(s) were then tested for lipase production along with different oil types. The different oils used include coconut, soybean, sesame, castor, mustard, flaxseed, sunflower, olive, or deep-frying waste supplemented to the screening medium in 1.5% v/v. Over and above, glucose, fructose, and sucrose were studied as the only carbon source. In the optimization stage, the best carbon and nitrogen sources with the greatest lipase activity were used.
Statistical experimental design
The Plackett-Burman experimental design of seven independent variables established the importance of medium components. The matrix in Table 1 shows that the seven independent factors resulted in 8 distinct combinations in 8 separate trials. Besides, the basal level at trial number 9 (Table 2). Matrix was created using the statistical software Statistica v6.0 (StatSoft Inc., 2001, USA). All trials were performed in triplicates, and the arithmetic mean of the triplicates was calculated as the response. The following equation was used to calculate the main effect of each variable:
R(H) is the response parameter that holds a higher quantity of a given component. R(L) is the response parameter that holds a lower quantity of a given component. N is the number of combinations divided by 2. The t-test was used to calculate t-values, p-values, and confidence-level percentages for the experimental variables using Microsoft Excel.
Purification of extracellular lipase
To purify lipase, various saturation levels of ammonium sulfate were added to the culture filtrate at 10 °C to precipitate the crude enzyme from cell-free supernatant. The added concentration was increased in ten percent increments from 20 to 100% saturation, then centrifuged for 15 min at 10,000 g at 10 °C. The precipitate was dialyzed against sterile distilled water after being dissolved in a 5 mM phosphate buffer at a pH of 7.0. The dialyzed protein was then purified using Sephadex G-100 column gel filtration chromatography. At a flow rate of 1 mL/min, after the column purification was run, the fractions were collected in a 3-mL quantity. The lipase activity was measured quantitatively in each fraction [23].
Characterization of purified lipase
The pH of pure lipase was profiled at various pH values (from pH 5 to pH 10). The enzyme (50 μL) was added to a reaction mixture having p-nitrophenyl laurate as substrate and phosphate buffer (50 mM, 700 μL) at each pH level. The reaction mixture was incubated at 10 °C for 10 min before the lipase activity was estimated. The reaction mixture was incubated at various temperatures (10 to 50 °C for 10 min) to determine the optimal temperature for lipase activity. The thermostability of lipase was examined by pre-incubating the enzyme at different temperatures [24]. The effect of varying substrate concentrations on the enzyme activity was also investigated by increasing the concentration of the substrate in the reaction mixture from 0 to 2 mg/mL. The influence of several cations on enzyme activity (Fe3+, Cu2+, Na+, K+, Mn2+, and NH4+) was investigated. The cations were added to the reaction mixture individually at a concentration of 1% w/v [25]. The lipase’s molecular mass was figured out by mixing 10–20 μg of the pure enzyme with 2x SDS-loading buffer, denatured for 5 min at 95 °C, cooled on ice, and then loaded into a 12% SDS-PAGE with a protein ladder. The gel was run in glycine buffer for 10 min at 10 mV before being raised to 25 mV for 1 h. After staining, the gel was removed from the glass plates and de-staining was performed.
Results
Cold-active lipase production by psychrotolerant bacteria
Seven bacterial isolates capable of growing at 10 °C were isolated directly from seawater. Only a strain named A6 displayed the greatest growth (OD600 ~ 1.3) at 10 °C after all recovered isolates were examined for their best growing temperature. The ability of A6 to produce lipase enzyme was assessed qualitatively using the clear zone formation method (Supplementary Material Fig. S1). As a result, this isolate was chosen for future research. The best growth was seen at 10 °C and pH 7–8, with lesser growth in the alkaline range (pH 9 and 10). The bacteria grew well in various NaCl concentrations (0–30%), with 15% providing the best results.
Molecular identification
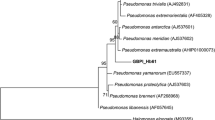
Based on phylogenetic relatedness analysis (Fig. 1), strain A6 was recognized as a novel member of the genus Pseudomonas. It is given the name Pseudomonas sp. A6. The nucleotide sequences were submitted to GenBank with the accession number KC417345. Blast revealed that it had a 100% similarity to the genus Pseudomonas.
Factors affecting lipase production
The greatest lipase activity (15 U/mL) was obtained when peptone was the only N source, whereas a combination of yeast extract and peptone yielded only 10U/mL (Fig. 2). The yeast extract, on the other hand, produced minimal activity (2 U/mL). In Fig. 3, glucose was the best carbon source among the simple sugars tested, with a lipase activity of 15 U/mL. Soybean oil, on the other hand, exhibited activity of 17 U/mL, which was greater than that of glucose among the oils examined.
Plackett-Burman design for main effect determination
The main effect of the evaluated factors on lipase activity is estimated and visually shown in Fig. 4. The highly influential variables were MgSO4, glucose, and medium pH. The t-values, p-values, and confidence-level percentages for the experimental variables are shown in Table 3.
The key factors boosting lipase synthesis, according to the main effect calculation for the factors under research and the confidence levels, were pH, glucose, and MgSO4. From here, a near optimum medium containing the following components (g/L): peptone 7.14; soybean oil 7.5% (v/v); K2HPO4, 0.4; MgSO4, 0.1; glucose 2; pH 8; temperature 10 °C; and incubation time 36 h should be about optimum. The basal, near optimal, and anti-optimum media were inoculated for verification, with the optimum medium showing a 1.5-fold increase in lipase activity to reach 23.36 U/mL. Lipase synthesis was completely suppressed in the anti-optimum experiments (Fig. 5).
Cold-active lipase purification
Purification of an enzyme is a key step in isolating the target protein and eliminating unwanted proteins. Extracellular lipase from Pseudomonas sp. A6 was isolated using a series of techniques. The purification profile of cold-active lipase is shown in Table 4. With a minor reduction of activity, the restored lipase had an activity of 17.8 U/mL. The recovered enzyme activity from each fraction is displayed in Fig. 6.
Cold-active lipase characterization
Effect of temperature and thermal stability
Lipase activity of the Pseudomonas sp. A6 purified enzyme was figured out by incubating the reaction mixture (p-nitrophenyl laurate), purified lipase enzyme, and phosphate buffer, pH 7 at different temperatures (10 °C, 20 °C, 37 °C, and 50 °C). The maximum lipase activity was seen at 10 °C. Additionally, at 20°C, the cold-active lipase showed relatively high lipase activity (75% of its activity at 10 °C). It can also tolerate temperatures up to 30 °C with a considerable reduction in activity (Fig. 7a). The activity decreased abruptly with a further rise in temperature and was almost lost at 60 °C. The purified lipase was partially stable up to 50 °C but maximum stability was determined at 10 °C.
Effect of pH
At acidic pH (pH 6), the pure enzyme showed no activity. At pH 6.5, the activity was low, but it steadily rose with increasing pH, peaking at pH 9 at 10 °C. At pH 8.5–9, Pseudomonas sp. A6 lipase activity was at maximum (19.5 U/mL, 1 U/mg specific activity) (Fig. 7b).
Effect of incubation time
Pure lipase enzyme activity was measured by incubating the reaction mixture (p-nitrophenyl laurate, purified lipase enzyme, and phosphate buffer pH 9) at 10 °C for various durations of time (10 to 60 min). As shown in Fig. 7c, after 30 min of incubation, the maximum activity (19.7 U/mL and 1 U/mg) was detected. A longer incubation period, on the other hand, resulted in a considerable decrease in lipase activity.
Effect of substrate concentration
As showed in Fig. 7d, Pseudomonas sp. A6 lipase activity increased as substrate (p-nitrophenyl laurate) concentration increased up to a point of 0.0010 g/mL, after which the activity remained constant. The Michaelis constant (Km) was figured out from the Lineweaver and Burk plot (Fig. 8), by dividing the slope of the line by the intercept. Vmax was figured out as the reciprocal of the intercept. The enzyme was found to have a Km of 6.6 * 10−4 m mol L−1 and a Vmax of 256.4 m mol L−1min−1.
Effect of enzyme concentration
The optimal enzyme concentration for maximal lipase activity (19.8 U/mL, 1.1 U/mg) by Pseudomonas sp. A6 was 1.4 mg/mL.
Effect of salts and metal ions
Metal ions can either promote or hinder the development of microbial enzymes. As shown in Table 5, Na+ ion was the most tolerable metal ion for lipase activity (19.5 U/mL, 1 U/mg), followed by K+ ion (16.6 U/mL, 0.85 U/mg), and NH4+ ion (15 U/mL). Alternatively, Cu2+ and Mn2+ inhibited the lipase activity.
Molecular weight determination of Pseudomonas sp. A6 cold lipase
As shown in Fig. 9, the crude enzyme produced several bands, while the protein precipitated with 60% ammonium sulfate produced five bands. The molecular weight of lipase was 65 kDa in the purified fraction by column chromatography (Sephadex G-100).
Discussion
Cold-adapted proteins and enzymes are found in psychrotolerant bacteria, which allow them to keep metabolic activity at low temperatures. Recently, many publications reported the production of cold-active lipase from Pseudomonas strains, for instance, Pseudomonas fluorescence KE38, Pseudomonas sp. LSK25, and Pseudomonas sp. CRBC14 [4, 26, 27]. An extracellular cold-adapted lipase enzyme from the marine psychrotolerant Pseudomonas sp. A6 that grew best at 10 °C was isolated and described in this work. Pseudomonas sp. A6 produced a novel cold-active lipase that was most active at 10 °C. A lipase with an optimal temperature and pH of 15 °C and 8, respectively, was isolated from the psychrotolerant Pseudomonas sp. AKM-L5 [28]. The lipase production in the current study showed maximum enzyme activity after 36 h of incubation during the stationary phase. Enzyme production is greatly dependent on the culture conditions like pH, temperature, and type of substrate. The highest growth for the organism was seen to be in the pH range of 7–8. Similar findings were seen for lipase production in neutral to alkaline media conditions by a psychrotrophic bacterium that was isolated from alpine regions [4, 29]. Moreover, lipases are produced by a variety of extremophiles, such as haloalkalitolerant [15]. A cold-active lipase was also discovered in another strain, Pseudomonas sp. AKM-L5 isolated from a soil sample in India. It has a molecular mass of 57 kDa as measured by SDS-PAGE, and a comparable optimal temperature of 10 °C to the lipase given, but pH 7 [29]. The optimum temperature and pH determined for lipase from Pseudomonas sp. KE38 were 25 °C and pH 8.5, respectively [27], which is considered high temperature and not a true cold-active lipase. On the other hand, and in accordance to our results, the lipase from Pseudomonas sp. LSK25 showed the exact optimum temperature of the reported lipase at 10 °C but lower pH value ranged from 7 to 7.5 [4].
Oil varieties used as lipase production inducers can affect enzyme production; soybean oil was produced the most, followed by mustard oil. Extracellular lipase synthesis is influenced by the kind of nitrogen supply as well as the carbon source. Peptone, an organic form of nitrogen, is extremely important since it supplies and serves as a precursor to the manufacture of vital amino acids, which are necessary for the formation of proteins, enzymes, and other cellular components [30]. The peptone and soybean were found to be the best nitrogen supply and oil substrate for generating cold-active lipase from Pseudomonas sp. A6. Their influence was not as strong or significant as the predicted main effect from the Plackett-Burman design. It is shown that MgSO4 and glucose concentration in the medium, in addition to pH value, was significant. The oil source is crucial as an inducer of lipase synthesis. In our study, soybean oil had the highest lipase activity compared to the other evaluated oil types.
In earlier research, olive oil was the best oil source for producing lipase from the fungus Curvularia sp. DHE 5 [31]. Using the Plackett-Burman optimization approach, MgSO4 was revealed to be essential to produce cold-active lipase from the Bacillus cereus HSS strain [12], which is consistent with our findings. In rodents and in vitro investigations, magnesium is important for lipase activity [32, 33]. Furthermore, pH 8 was identified as being truly relevant for lipase synthesis, which might be linked to the natural pH of seawater (pH 8.1) [34], from which the isolated strain was recovered and adapted to live in the marine environment. The best pH for cold-active lipase was in the alkaline range between pH 8 and 9.
Finally, the addition of glucose not only increased lipase synthesis in the presence of an oil substrate but also played a substantial role in the optimization experiment. It was reported that in the optimization of cold-active lipase from B. cereus HSS, the facile absorption of glucose to form the precursor necessary for microbial lipase may account for the favorable influence on lipase synthesis [12]; nevertheless, in a recent investigation, the presence of 20 mM glucose hindered lipase activity, which contradicts our findings [35].
Ammonium sulfate precipitation and gel filtering were used to efficiently purify Pseudomonas sp. A6 crude enzyme to homogeneity. Numerous investigations on the multi-step purification of cold-active lipase from various psychrophilic and psychrotrophic bacteria have been done. Ammonium salt precipitation is initially performed, followed by a succession of chromatographic phases [5, 11, 36].
SDS-PAGE revealed that the isolated lipase had a molecular weight of 65 kDa. The purified lipase’s biochemical evaluation revealed that it was active in the pH range of 7 to 9, with a best activity at pH 9. This shows that the purified lipase is alkaline. This form of alkaline active lipase has been shown in various microorganisms. Another alkaline lipase with activity in a pH range of 8.0–10.5 with maximum activity at pH 8.5 [37]. Such enzymes benefit from alkali tolerance since they are cold-active and actively stable in alkaline environments.
Conclusion
The current study looked for cold-active lipase-producing bacteria from saltwater in the Alexandrian Mediterranean Sea. Pseudomonas sp. A6 is found to produce cold-active lipase. The Plackett-Burman statistical experimental approach was used to study the primary determinants regulating cold-active lipase. The optimum medium component found to be peptone 7.14 g/L; soybean oil 7.5% (v/v); K2HPO4, 0.4 g/L; MgSO4, 0.1 g/L; glucose 2 g/L; pH 8; and temperature 10 °C. The lipase produced under optimal culture conditions increased activity 1.5-fold when compared to the un-optimized medium. The features of the pure lipase enzyme generated have been carefully investigated, making it a practical option for various applications that need lipase activity at low temperatures.
References
Kavitha M, Shanthi C (2013) Isolation and characterization of cold active lipase producing Pseudomonas sp. 4 from marine samples of Tamilnadu Coast. Res J Biotechnol 8:57–62
Abd-Elnaby HM, Abou-Elela GM, Hussein H, Ghozlan HA, Sabry SA (2019) Characterization and bioremediation potential of marine psychrotolerant Pseudomonas spp. isolated from the Mediterranean Sea, Egypt. Egypt J Aquat Biol Fish 23(4):669–683. https://doi.org/10.21608/ejabf.2019.63537
Gupta R, Gupta N, Rathi P (2004) Bacterial lipases: an overview of production, purification and biochemical properties. Appl Microbiol Biotechnol 64(6):763–781. https://doi.org/10.1007/s00253-004-1568-8
Salwoom L, Raja Abd Rahman R, Salleh A, Mohd Shariff F, Convey P, Pearce D, Mohamad Ali M (2019) Isolation, characterisation, and lipase production of a cold-adapted bacterial strain Pseudomonas sp. LSK25 isolated from Signy Island, Antarctica. Molecules 24(4):715. https://doi.org/10.3390/molecules24040715
Kumar A, Mukhia S, Kumar N, Acharya V, Kumar S, Kumar R (2020) A broad temperature active lipase purified from a psychrotrophic bacterium of Sikkim Himalaya with potential application in detergent formulation. Front Bioeng Biotechnol 8. https://doi.org/10.3389/fbioe.2020.00642
Kavitha M (2016) Cold active lipases – an update. Front Life Sci 9(3):226–238. https://doi.org/10.1080/21553769.2016.1209134
Sarmiento F, Peralta R, Blamey JM (2015) Cold and hot extremozymes: industrial relevance and current trends. Front Bioeng Biotechnol 3. https://doi.org/10.3389/fbioe.2015.00148
Mhetras N, Mapare V, Gokhale D (2021) Cold active lipases: biocatalytic tools for greener technology. Appl Biochem Biotechnol 193(7):2245–2266. https://doi.org/10.1007/s12010-021-03516-w
Le LTHL, Yoo W, Jeon S, Lee C, Kim KK, Lee JH, Kim TD (2020) Biodiesel and flavor compound production using a novel promiscuous cold-adapted SGNH-type lipase (HaSGNH1) from the psychrophilic bacterium Halocynthiibacter arcticus. Biotechnol Biofuels 13(1):55. https://doi.org/10.1186/s13068-020-01696-x
Mahmoud NH, Elsherbiny BA, Moffit SM, Mohamed JH, Abouelkheir SS, Abdella B (2022) Cell biology and microbial interactions in algal cells. In: El-Sheekh MM, Abdullah N, Ahmad I (eds) Handbook of research on algae as a sustainable solution for food, energy, and the environment. IGI, pp 84–108. https://doi.org/10.4018/978-1-6684-2438-4.ch004
Wang Q, Zhang C, Hou Y, Lin X, Shen J, Guan X (2013) Optimization of cold-active lipase production from psychrophilic bacterium Moritella sp. 2-5-10-1 by statistical experimental methods. Biosci Biotechnol Biochem 77(1):17–21. https://doi.org/10.1271/bbb.120104
Hassan SWM, Abd El Latif HH, Ali SM (2018) Production of cold-active lipase by free and immobilized marine Bacillus cereus HSS: application in wastewater treatment. Front Microbiol 9. https://doi.org/10.3389/fmicb.2018.02377
Pirghorbani M, Ebrahimi M (2021) Statistical optimization of alkaline lipase production by extreme halophilic archean Natrialba asiatica. J Microbiol Biotechnol Food Sci 11(2):1–6. https://doi.org/10.15414/jmbfs.1164
Arayes MA, Mabrouk MEM, Sabry SA, Abdella B (2022) Exopolysaccharide production from Alkalibacillus sp. w3: statistical optimization and biological activity. Biologia 78(1):229–240. https://doi.org/10.1007/s11756-022-01233-1
Arayes MA, Mabrouk MEM, Sabry SA, Abdella B (2021) Diversity and characterization of culturable haloalkaliphilic bacteria from two distinct hypersaline lakes in northern Egypt. Biologia 76(2):751–761. https://doi.org/10.2478/s11756-020-00609-5
Romanenko LA, Schumann P, Rohde M, Lysenko AM, Mikhailov VV, Stackebrandt E (2002) Psychrobacter submarinus sp. nov. and Psychrobacter marincola sp. nov., psychrophilic halophiles from marine environments. Int J Syst Evol Microbiol 52(4):1291–1297. https://doi.org/10.1099/00207713-52-4-1291
Amoozegar M, Malekzadeh F, Malik KA (2003) Production of amylase by newly isolated moderate halophile, Halobacillus sp. strain MA-2. J Microbiol Methods 52(3):353–359. https://doi.org/10.1016/S0167-7012(02)00191-4
Sambrook J, Fritsch ER, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, New York, NY USA
Abdella B, El-Wazzan E, El-Sersy NA, Sabry SA, El-Helow ER (2017) Pathogenicity and antibiotic susceptibility of two bacterial pathogens associated with the clam Tapes decussatus in some Egyptian fisheries. Ege J Fish Aquat Sci 34(4):383–389. https://doi.org/10.12714/egejfas.2017.34.4.04
Okonechnikov K, Golosova O, Fursov M, Varlamov A, Vaskin Y, Efremov I, German Grehov OG, Kandrov D, Rasputin K, Syabro M, Tleukenov T (2012) Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics 28(8):1166–1167. https://doi.org/10.1093/bioinformatics/bts091
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Yuan BH, Cai YJ, Liao XR, Yun LH, Zhang F, Zhang DB (2010) Isolation and identification of a cold-adapted lipase producing strain from decayed seeds of Ginkgo biloba L. and characterization of the lipase. Afr J Biotechnol 9(18):2661–2667
Shaheen M, Ullah I, Rafiq M, Maqsood Ur Rehman M, Shah AA, Hasan F (2020) Purification and characterization of lipase from psychrophilic bacteria Pseudomonas mandelii htb2 from Batura Glacier, Pakistan. Appl Ecol Environ Res 18(3):4103–4114. https://doi.org/10.15666/aeer/1803_41034114
Chakraborty K, Raj RP (2008) An extra-cellular alkaline metallolipase from Bacillus licheniformisMTCC 6824: purification and biochemical characterization. Food Chem 109(4):727–736. https://doi.org/10.1016/j.foodchem.2008.01.026
Joshi GK, Kumar S, Tripathi BN, Sharma V (2006) Production of alkaline lipase by Corynebacterium paurometabolum, MTCC 6841 isolated from Lake Naukuchiatal, Uttaranchal State, India. Curr Microbiol 52(5):354–358. https://doi.org/10.1007/s00284-005-0224-6
Farooq S, Ganai SA, Ganai BA, Mohan S, Uqab B, Nazir R (2022) Molecular characterization of lipase from a psychrotrophic bacterium Pseudomonas sp CRBC14. Curr Gen 68(2):243–251. https://doi.org/10.1007/s00294-021-01224-w
Karakaş F, Arslanoğlu A (2020) Gene cloning, heterologous expression, and partial characterization of a novel cold-adapted subfamily I.3 lipase from Pseudomonas fluorescence KE38. Sci Rep 10(1):22063. https://doi.org/10.1038/s41598-020-79199-w
Maharana AK, Ray P (2014) Application of Plackett-Burman design for improved cold temperature production of lipase by psychrotolerant Pseudomonas sp. AKM-L5. Int J Curr Microbiol App Sci 3(4):269–282
Maharana A, Ray P (2015) A novel cold-active lipase from psychrotolerant Pseudomonas sp. AKM-L5 showed organic solvent resistant and suitable for detergent formulation. J Mol Catal B: Enzym 120:173–178. https://doi.org/10.1016/j.molcatb.2015.07.005
Divya K, Padma PN (2016) Screening of diverse organic and inorganic nitrogen sources for cold-active polygalacturonase and amylase production by Geotrichum sp using Plackett-Burman design. Int J Technol Res Eng 3(5):875–878
El-ghonemy DH, El-gamal MS, Tantawy AE (2017) Extracellular alkaline lipase from a novel fungus Curvularia sp . DHE 5 : optimisation of physicochemical parameters , partial purification and characterisation. Food Technol Biotechnol 55(2):206–217. https://doi.org/10.17113/ftb.55.02.17.4958
Rayssiguier Y, Noé L, Etienne J, Gueux E, Cardot P, Mazur A (1991) Effect of magnesium deficiency on post-heparin lipase activity and tissue lipoprotein lipase in the rat. Lipids 26(3):182–186. https://doi.org/10.1007/BF02543968
ChengGui L, XuDong G, FeiHong Z, FuXing L (2016) Effect of magnesium ions in water on the activity of lipase. Food Res Dev 37(6):172–175
Boyd CE (2020) Typical chemical characteristics of full-strength seawater, Globalseafood.Org, from https://www.globalseafood.org/advocate/typical-chemical-characteristics-of-full-strength-seawater/. Accessed 3 May 2020
Tsuzuki W, Kitamura Y, Suzuki T, Mase T (1999) Effects of glucose on lipase activity. Biosci Biotechnol Biochem 63(8):1467–1470. https://doi.org/10.1271/bbb.63.1467
Bae J-H, Kwon M-H, Kim I-H, Hou CT, Kim H-R (2014) Purification and characterization of a cold-active lipase from Pichia lynferdii Y-7723: pH-dependant activity deviation. Biotechnol Bioprocess Eng 19(5):851–857. https://doi.org/10.1007/s12257-014-0300-5
Jinwal UK, Roy U, Chowdhury AR, Bhaduri A, Roy P (2003) Purification and characterization of an alkaline lipase from a newly isolated Pseudomonas mendocina PK-12CS and chemoselective hydrolysis of fatty acid ester. Bioorg Med Chem 11(6):1041–1046. https://doi.org/10.1016/S0968-0896(02)00516-3
Acknowledgements
The authors want to thank the faculty of science, at Alexandria University for the support and for using the institute’s resources to conduct this work.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
SAS and HAG designed the study. ASY and BA conducted experiments under the supervision of SAS and HAG. BA wrote the first draft of the manuscript. BA, SAS, and HAG completed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with humans or animals.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: María Martha Martorell
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 469 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abdella, B., Youssif, A.M., Sabry, S.A. et al. Production, purification, and characterization of cold-active lipase from the psychrotroph Pseudomonas sp. A6. Braz J Microbiol 54, 1623–1633 (2023). https://doi.org/10.1007/s42770-023-01079-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01079-y