Abstract
A halotolerant Planomicrobium species with remarkable lipase activity, isolated from the sediments of a mangrove, was investigated for the lipase purification, yield and production improvement. The enzyme purification was increased by 12.86-folds using a two-step purification process by ammonium sulfate precipitation and gel filtration chromatography. Subsequently, a submerged fermentation was optimized for the maximum lipase production using a response surface method (RSM). Several variables, comprising 8 nutritional variables along with pH, temperature and agitation speed were screened for the effective ones by Placket–Burman design (PBD). The effective variables were further examined for their optimum levels for the enzyme production using a central composite design (CCD). NaCl (1.28 % w/v), glucose (1.20 % w/v), olive oil (1.35 % w/v), peptone (0.69 % w/v) and pH (7.8) were found to be effective in maximizing the enzyme productivity in a lab scale (10.08 U/ml of cell-free supernatant) and a 10 l fermenter (9.89 U/ml). The results suggested that the model is accurate and consistent in predicting the best condition. Therefore, the strain under this specific fermentation condition promises a good prospect for a large-scale lipase production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial lipolytic enzymes have significant biotechnological processes and industrial applications [1]. They have served in many applications, including leather processing, food industry, cosmetics, pharmaceutical products, remediation of wastewater, as an ingredient in dress washing powders and in oleochemical and organic synthesis under organic solvent conditions [2, 3]. The natural function of these enzymes is to hydrolyze steric bonds in various lipids such as triacylglycerol leading to the release of more available compounds of fatty acids and glycerol for assimilation by microbial cells acquiring energy [4]. Versatile microbial species have already been reported to secrete lipases with different enzymological properties and specificities [5–9]. In this regard, extracellular bacterial lipases have attracted considerable interest in biotechnological perspectives, as their bulk production and purification are much easier than intracellular enzymes and enzymes derived from eukaryotic cells [1]. The former needs further purification steps, resulting in low yield of the product, and the latter necessitates a more complex nutritional requirement and sophisticated bioreactors, both of them raise expenses of the production [1, 10].
Once the appreciable activity of a strain is confirmed in the prospect for production of a commercially-relevant compound, the other major issues to be determined is the optimization of the process. Even a little but consistent improvement in yield in industrial scale means a high rise in profit and is highly regarded [10, 11]. Therefore, employment of a method with the prospect of expanding the knowledge on influencing parameters could help to keep the process under control. The more the experimental design could explain the process, the more promising the process is for scale-up success in the maximum yield of a product [12, 13]. In this regard, response surface methods (RSM) with varying experimental designs have shown to provide definitive information as to how to elicit optimum enzyme productivity from a fermentation. These methods have been proven to be more efficient in finding the optimum condition than one factor at a time (OFAT) method [1, 14].
The lipase activity of a halotolerant strain of Planomicrobium has been illustrated along with the efficiency of an enzyme purification process in the current investigation. The main parameters and their interactions in relation to the lipase yield have been examined in a submerged fermentation process using RSM with central composite design (CCD). Based on the empirical results obtained from the experimental runs, the proposed suitable condition for the maximum lipase production was ultimately checked for fidelity and consistency in the enzyme yield in a lab and pilot- scale batch fermentation.
Material and Methods
Materials
All the media components and chemicals were of analytical grade which were purchased from Sigma-Aldrich (St. Louis, Missouri). Dialysis cassettes with 12–14 kDa molecular weight cut off (MWCO) and protein molecular weight markers were also procured from Sigma-Aldrich.
Screening of Lipolytic Bacterial Strain
The Planomicrobium sp. MR23K was isolated from a mangrove rhizosphere in an island Qeshm, south coast of Iran. The microorganism was characterized as a lipase-producing bacterium as a big halo zone formation was observed around their colony on tributyrin agar medium after 48 h incubation at 34 °C [15]. Physicochemical studies followed by 16S rDNA sequence analysis were carried out for identification of the bacterium. The primary culture medium was a synthetic medium containing following nutrients (w/v) as 0.1 % yeast extract, 0.5 % glucose, 0.5 % olive oil, 0.2 % K2HPO4, 2.5 % NaCl, and 0.5 % NH4Cl with initial pH 7.5, whose levels were optimized for the maximum lipase production in the next experiments.
Lipase Extraction and Purification
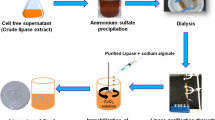
All purification steps were conducted at 4 °C. The cell-free supernatant was served for the lipase enzyme purification. Thus, an appropriate volume of the supernatant was mixed with 70 % w/v glacial ammonium sulfate solution (1:1) and kept in an ice-bath for the enzyme precipitation for 20 min. The precipitant was then collected by centrifugation at 3000g for 15 min at 4 °C, dissolved in phosphate buffer (0.1 M, pH 7) and dialyzed overnight against an extensive volume of phosphate buffer (0.01 M). The enzyme purification was conducted using gel filtration chromatography with a Sephadex G-100 column (1.25 × 20 cm). Ten milliliters of the dialysate were loaded onto the column pre-equilibrated with Tris-HCl buffer (0.1 M, pH 8) and eluted with the same buffer until 20 fractions of 2 ml were taken. The fractions were collected with every 10 min for determination of the lipase activity and protein assay.
Protein Quantification
Total protein content was determined according to Bradford method, using Bradford reagent and bovine serum albumin (BSA) as the reagent for standard curve [16]. The protein content was measured in the cell-free substrate, the dialysate and for all fractions withdrawn from the Sephadex column.
Lipase Activity Assay
Lipase activity was assayed according to Gupta et al. [17], using 4-nitrophenyl palmitate (4-NPP) as the substrate and indicator of hydrolysis activity. A 50 mM stock solution of 4-NPP was prepared using 1:1 ratio of acetonitrile and n-butanol. The reaction mixture was prepared by adding 20 µl of the 4-NPP stock solution into 200 µl of the solutions which were to be tested for the activity in Tris-HCl buffer (1.8 ml). The reaction mixture was incubated at 37 °C for 15 min and then the reaction was halted by adding 100 µl trichloroacetic acid (0.5 M). The respective absorbance value was recorded at 410 nm by spectrophotometry. The unknown concentration of 4-nitrophenol released was determined from the standard curve of 4-nitrophenol. One unit of lipase was defined as an amount of enzyme releasing 1 µmol of 4-NP under standard assay conditions.
Determination of Molecular Weight
The molecular weight of the purified protein sample was determined by SDS–PAGE [18]. Electrophoresis was carried out with 4 % stacking gel and 12 % separating gel at 120 V for 150 min. The protein band was visualized using a silver-staining kit. Relative molecular mass (Mr) was estimated as compared to molecular mass markers. The purified protein sample was also run on the PAGE with 12 % separating gel without SDS and the enzymatic feature of the loaded protein was determined with a chromogenic zymography as a yellow color band was developed on the gel after incubation at 37 °C for 15 min [19].
Optimization of the Enzyme Fermentation
Plackett–Burman Design Experiment
A Plackett–Burman design (PBD) was employed for identification of main variables affecting the crude lipase production. A total of 11 independent variables with the possible low and high levels were assessed for their significance in the crude enzyme yield using outcomes of 12 experimental runs. These variables were as NaCl (0.5–3), glucose (1–3), yeast extract (0.5–1), olive oil (0.5–1), NH4Cl (0.25–0.5), peptone (0–0.1), MgSO4 (0–0.1), KH2PO4 (0.03–0.06) in w/v percentage, pH (4–8), agitation speed (100–120 rpm) and temperature (25–35 °C). The statistical software package of Design Expert® 7.0.0 (Stat-Ease, Inc., Minneapolis, MN, USA) was applied to generate the design and analyze the outputs of the experiments. The lipase activity of the media was assessed in the cell-free supernatants precipitated by the ammonium sulfate solution after 72 h of incubation.
Response Surface Method
The five significant variables, suggested by the PBD analysis (Fig. 1), were examined for their optimum levels suitable for the maximum lipase production. Hence, RSM with a 25 full factorial CCD was employed with a total of 32 runs using Design-Expert software. The CCD that consisted of factorial trails and star points was employed to get a second-order model, explaining the quadratic effects of the variables and the combined effects of their interactions on the process variability in terms of lipase production measured as the dependent response. The remaining non-significant variables as KH2PO4, NH4Cl, MgSO4, temperature, and agitation speed were maintained at their respective minimum values. The basic quadratic polynomial model explaining linear, quadratic and interactive variables in relation to the system response was subjected to model modification, whose insignificant terms were omitted to increase the model fitness for the empirical responses. This model is as follows.
where Y represents predicted response, Xi and Xj are the input variables, and β0, βi, βii and βij are the regression coefficients for intercept, linear, quadratic and interaction terms, respectively. The following equation was employed for coding the experimental values of the factors in the range of −1 to +1:
where xi is the dimensionless value of an independent variable, Xi is the actual value of the independent variable, X0 is the value of Xi at the center point, and ∆Xi is the step change. The relation among the factors and the response values was determined by analyzing the data in a statistical approach, using multiple regression analysis. Analysis of variance (ANOVA) was performed by Design-expert software to determine the significance of each term in the final equation and to estimate the goodness of fit for the equation and every term. Moreover, the fitted polynomial equation was utilized to generate three-dimensional response plots to display the main and their combined effects of the independent factors on the dependent response. The concentration of different variables suggested by the model, the best fermentation condition for enzyme productivity, was double-checked in lab scale and pilot-scale fermentations to verify whether the model was reliable and consistent in achieving the optimum lipase productivity.
Molecular weight estimation of the bacterial lipase by comparing the location of the associated band with those of a molecular marker derived from bovine serum (Lane M). A single band of a nearly 50 kDa on silver staining indicates the protein’s purity (Lane 1) and the corresponding zymogram obtained from the native pure protein on the PAGE supports the enzymatic feature of the protein (Lane 2)
Submerge Fermentation for Lipase Production
The fermentation was performed according to the favorable condition, concluded from PBD and RSM analyses in a 10 l propeller-stirred batch fermenter (Biolab, B. Braun, Germany). The crude enzyme yield was determined in the ammonium sulfate precipitants. These included the desirable nutritional condition consisted of significant variables (% w/v) as NaCl (1.28), glucose (1.20), olive oil (1.35) and peptone (0.69) at pH 7.8, along with other nutrients affecting lipolytic productivity insignificantly at their respective minimum levels. Accordingly, a broth with the bacterial growth at log phase was inoculated into a 6 l fresh medium at the ratio of 1:9 (v/v) and the submerged culture was incubated at 25 °C for 3 days at 100 rpm. During the incubation time, samples were taken from the fermented culture at 8 h intervals to monitor the lipase production and microbial growth rate. The results were utilized to determine the period as when the productivity reached its maximum level. Furthermore, the maximum production was compared with the yield predicted by the equation and ones obtained in the lab scale test to verify the validity and ensure the consistency of the model in the prediction.
Results and Discussion
Purification of Lipase Enzyme from Planomicrobium sp. MR23K
It was rational to first investigate how much purifying the lipase would be successful before optimizing fermentation condition for the crude lipase production. The lipolytic activity in the cell-free supernatant was 22.31 U/mg of total protein after 3 days of incubation (Table 1). After the ammonium sulfate precipitation process, the recovery of total protein was accompanied by increased lipase harvesting yield (17.7-folds) with a specific activity of 29.39 U/mg that infers a 32 % increase in purification. Further a considerable amount of purification (nearly 13-folds) was accomplished in the second step of the purification, after gel filtration chromatography. In this step, the enzyme yield was 21 % and the activity of the enzyme reached up to 286.78 U/mg. According to Ogino et al. [19], the purification of LST-03 organic solvent-tolerant lipase was increased by 34.7-folds with an overall yield of 12.6 %. The lipase from Bacillus sphaericus was purified 17-folds, with a specific activity of 377.69 U/mg, using ammonium sulfate precipitation and DEAE-Sepharose anion exchange chromatography [20]. In the present study, the purification of a lipase from a halotolerant Planomicrobium species increased by 12.86-folds, associated with a specific activity of 286.78 U/mg, using ammonium sulfate precipitation and gel filtration column. The 16S rDNA sequence of the bacterium was deposited in NCBI Genbank with accession number of KT260165 and the bacterium was termed as Planomicrobium sp. MR23K.
Determination of Molecular Weight
A single band appeared on a 12 % gel after silver staining, with a molecular weight of roughly 50 kDa (Fig. 1). Panigrahi et al. [21] reported a thermostable lipase from a Brevibacillus species with a molecular weight of 70 kDa. In contrast, Planococcus halocryophilus Or1 produced a lipase with a low molecular weight of 33 kDa [22]. The molecular weight of the most lipases from other gram-positive bacilli has already been reported to be around 43–45 kDa [23]. Regarding the molecular weight of the lipase in the present investigation, the study corroborates with those of other lipases exemplified here [21–23].
Optimization of Fermentation Condition
PBD suggests a fractional factorial design with a minimum collection of experimental runs to screen many factors for significant ones. In this statistical approach, no interactions between different parameters in a specific range were observed. Furthermore, a linear correlation between the response and a variable is sufficient for screening the main variables. Among 11 variables considered in the PBD approach, 5 variables were found to exert a significant change in the enzyme yield (Fig. 2). These were NaCl, glucose, olive oil, peptone and pH, indicating that their changes could influence the yield of the lipase enzyme. To discover the optimum levels of these significant variables, a set of different experimental runs was employed.
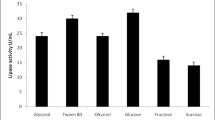
Pareto chart showing the effect of different fermentation variables on the lipase activity. Given the t value limit, it has been revealed that pH, glucose, olive oil, NaCl and peptone variation could influence the enzyme productivity as they exceed the t value limit. On the other hand, as none of the parameters exceeds the Bonferroni limit, it implies that all significant parameters share similar involvement in the enzyme productivity
Response surface methodology (RSM) is a collection of empirical techniques for evaluation of relations between controlled variables and measured response. To understand the combined effects of interacting variables as well as their main impacts on the lipase productivity, RSM with CCD was applied for the selected variables obtained by the PBD. By employing multiple regression analysis on the experimental data, a second-order polynomial equation was achieved describing the levels of lipase productivity as a function of the significant variables. The quadratic polynomial equation for the lipase productivity was explained in terms of the coded values as following.
where Y was the predicted crude lipase production, and \(x_{1}\), \(x_{2}\), \(x_{3}\), \(x_{4}\) and \(x_{5}\) were the coded factors for NaCl, glucose, pH, peptone and olive oil, respectively.
The analysis of variance (ANOVA) for the regression equation is listed in Table 2. The pure error was very low, indicating a perfect fit between the predicted values suggested by the model and the real data. F-value is a ratio of the mean square from the regression to the mean square residual and P value is the estimated probability of rejecting the null hypothesis (H0) when the null hypothesis is true. The high F-value (47.06) and low P value (<0.0001) implied that the model was very predictive and significant. In other words, the sum of squares of “Lack of Fit” with the low F value of 1.62 and P value >0.05 indicated that the Lack of Fit was not significant relative to pure error. Taken together, the results indicated that there would be a little chance to get the model due to errors.
The value of the predicted determination coefficient (R 2Pred = 0.81) implied that 81 % of the total variation could be explained by the regression equation. The value of the adjusted determination coefficient (R 2Adj = 0.97) was also very high, indicating that the equation was highly significant. In comparison, the predicted R square value was very close to that of adjusted R square, which was an accurate measure of precision [24].
The bacterial growth decreased as NaCl concentration in the medium increased, which was accompanied by a decrease in the enzyme activity (Fig. 3a). Considering the strong coefficient of determination (R2 = 0.73), the reduced lipase activity could be attributed to the reduced bacterial density in the media containing the elevated NaCl concentration (Fig. 3b). It was observed that the main effects were significant for all coded factors, with the exception of peptone (P < 0.001). Additionally, the interacting effects of glucose with peptone, glucose with olive oil and olive oil with pH were found to be significant on the lipase production in the fermentation (P < 0.05). The three-dimensional response surface graphs displayed the combined effect of two interacting variables on the lipase production while the other variables were maintained at their average levels (Fig. 4). According to Fig. 4a, increase in NaCl concentration in the medium resulted in a decrease in the lipase production probably due to the bacterial growth reduction, although the strain was tolerant to elevated concentrations of salts. On the other hand, the increase in peptone concentration enhanced the lipase productivity to about 9.87 U/ml and after that it was decreased with further increase in peptone concentration. However, variation in the lipase productivity caused by the peptone concentration was observed insignificant. Increase in olive oil concentration enhanced the lipase productivity up to 9.84 U/ml at pH 8 (Fig. 4b), while it did not affect the lipase activity at pH 6, indicating that the lipase probably was active at only pH 8. The major activity of the lipase at alkaline condition is of great significance as alkaline lipases are more suitable for application in industrial sectors, like enzymatic bioscouring process in textile industries as well as in laundry detergents [25, 26]. Increase in olive oil concentration increased the productivity steadily and the elevation of the glucose concentration enhanced firstly the lipase activity up to 9.65 U/ml, but thereafter the lipolytic activity was decreased with further increase in glucose (Fig. 4c). The same trend was also observed for the lipase activity in relation to the glucose concentration in Fig. 4d. The peptone concentration variation did not influence the lipase activity, given the P value >0.05 for the peptone parameter. The reduction in lipase productivity could be attributed to osmotic pressure caused by elevated levels of glucose influencing both bacterial growth and lipase productivity due to probably catabolic carbon repression of the enzyme production [27].
Enzyme activity reduced along with the bacterial growth (optical density or OD) as the NaCl concentration increases in the media (a). Given the strong coefficient of determination (R2 = 0.73), the enzyme activity variation can be attributed to the bacterial growth in the media (b). Asterisk depicts significant differences as compared with the respective values of the enzyme activity and OD at zero NaCl concentration, using one-way ANOVA and Tukey’s multiple comparison post-test at the significance level of 0.05 from GraphPad prism version 5.0 (n = 3)
According to the model, the maximum lipase activity was estimated to be achieved at 1.28 % w/v NaCl, 1.20 % w/v glucose, 1.35 % w/v olive oil, 0.69 % w/v peptone and at pH 7.8, which was predicted at 10.22 U/ml. Under this condition, where the remaining insignificant variables were maintained at their minimum levels, the lipase productivity was improved and increased up to 10.08 U/ml in the lab-scale and 9.89 U/ml in the pilot-scale experiments after 48 h of incubation (Fig. 5). Considering the optical density of the media, the increase in the enzyme activity could be correlated to the enhanced bacterial biomass. Overall, the model was confirmed as to be accurate and consistent in predicting the enzyme productivity under the suggested condition. Although some factors like agitation speed were found to be unimportant in influencing the lipase production, it may affect the productivity in a broader range than the range (100–120 rpm) studied in the current investigation. Furthermore, it may affect the productivity in large-scale fermentation system. On the other hand, parameters like agitation and temperature are considered to be important in large-scale production as they involve the costs of the production. How much variation can be applied with regard to these parameters without a significant alteration in the productivity is a highly interesting question from the industrial perspectives, especially in large-scale fermentation systems, as the information helps to reduce the costs of the production.
Time-course illustration of bacterial growth and enzyme production using optical density (OD) of media and the enzyme activity of the cell free supernatant. There is found no difference between pilot scale and lab scale results in the enzyme activity and OD, using two-way ANOVA and Tukey’s multiple comparison post test at the significance level of 0.05 from GraphPad prism version 5.0 (n = 3)
Conclusion
In the present work, a halotolerant Planomicrobium with appreciable lipase activity was selected from the sediments of a mangrove. The major accomplishment of a highly purified lipase enzyme with a simple two-step purification process is an encouraging success in industrial point of view, regarding usually high expenses of down-stream purification steps and low product yield. This encouraged the authors to optimize the fermentation condition to obtain maximum lipase production. In this regard, it was found that adopting RSM approach could contribute to the comprehension of the effective parameters and discovery of the optimum condition for the highest lipase productivity in fermentation. Moreover, RSM could minimize the experimental runs needed for an objective, which was the maximum lipase productivity in the present study. The proposed model derived from the empirical experiments was also proven to be reliable, since employing the predicted optimum condition resulted in consistently similar outputs and maximum lipase productivity, in both the lab scale and pilot-scale fermentations. Knowing the information that lipase purification steps are amenable for a large-scale lipase harvesting and that the estimated optimum fermentation condition is reliable, this strain has a promising prospect for application in the industrial production of the lipase enzyme.
References
Gupta R, Gupta N, Rathi P (2004) Bacterial lipases: an overview of production, purification and biochemical properties. Appl Microbiol Biotechnol 64(6):763–781. doi:10.1007/s00253-004-1568-8
Bora L, Gohain D, Das R (2013) Recent advances in production and biotechnological applications of thermostable and alkaline bacterial lipases. J Chem Technol Biotechnol 88(11):1959–1970. doi:10.1002/jctb.4170
Jaeger KE, Reetz MT (1998) Microbial lipases form versatile tools for biotechnology. Trends Biotechnol 16(9):396–403. doi:10.1016/S0167-7799(98)01195-0
Bussamara R, Fuentefria AM, Oliveira ESd, Broetto L, Simcikova M, Valente P, Schrank A, Vainstein MH (2010) Isolation of a lipase-secreting yeast for enzyme production in a pilot-plant scale batch fermentation. Bioresour Technol 101(1):268–275. doi:10.1016/j.biortech.2008.10.063
Abramić M, Leščić I, Korica T, Vitale L, Saenger W, Pigac J (1999) Purification and properties of extracellular lipase from Streptomyces rimosus. Enzyme Microb Technol 25(6):522–529. doi:10.1016/S0141-0229(99)00077-0
Adan Gökbulut A, Arslanoǧlu A (2013) Purification and biochemical characterization of an extracellular lipase from psychrotolerant Pseudomonas fluorescens KE38. Turk J Biol 37(5):538–546. doi:10.3906/biy-1211-10
Ahmed EH, Raghavendra T, Madamwar D (2010) An alkaline lipase from organic solvent tolerant Acinetobacter sp. EH28: application for ethyl caprylate synthesis. Bioresour Technol 101(10):3628–3634. doi:10.1016/j.biortech.2009.12.107
Amoozegar MA, Salehghamari E, Khajeh K, Kabiri M, Naddaf S (2008) Production of an extracellular thermohalophilic lipase from a moderately halophilic bacterium, Salinivibrio sp. strain SA-2. J Basic Microbiol 48(3):160–167. doi:10.1002/jobm.200700361
Anbu P, Hur BK (2014) Isolation of an organic solvent-tolerant bacterium Bacillus licheniformis PAL05 that is able to secrete solvent-stable lipase. Biotechnol Appl Biochem. doi:10.1002/bab.1202
Gunasekaran V, Das D (2005) Lipase fermentation: progress and prospects. Indian J Biotechnol 4(4):437–445
Esteban-Torres M, Mancheño JM, de las Rivas B, Munoz R (2015) Characterization of a halotolerant lipase from the lactic acid bacteria Lactobacillus plantarum useful in food fermentations. LWT Food Sci Technol 60(1):246–252
Ruchi G, Anshu G, Khare S (2008) Lipase from solvent tolerant Pseudomonas aeruginosa strain: production optimization by response surface methodology and application. Bioresour Technol 99(11):4796–4802
Hill WJ, Hunter WG (1966) A review of response surface methodology: a literature survey. Technometrics 8(4):571–590
Winkler UK, Stuckmann M (1979) Glycogen, hyaluronate, and some other polysaccharides greatly enhance the formation of exolipase by Serratia marcescens. J Bacteriol 138(3):663–670
Esteban-Torres M, Mancheño JM, de las Rivas B, Muñoz R (2014) Production and characterization of a tributyrin esterase from Lactobacillus plantarum suitable for cheese lipolysis. J Dairy Sci. doi:10.3168/jds.2014-8234
Kruger NJ (2002) The Bradford method for protein quantitation. In: Walker JW (ed) The protein protocols handbook, 2nd edn. Humana Press Inc., Totowa, pp 15–21
Gupta N, Rathi P, Gupta R (2002) Simplified para-nitrophenyl palmitate assay for lipases and esterases. Anal Biochem 311(1):98–99
Singh R, Gupta N, Goswami VK, Gupta R (2006) A simple activity staining protocol for lipases and esterases. Appl Microbiol Biotechnol 70(6):679–682
Ogino H, Nakagawa S, Shinya K, Muto T, Fujimura N, Yasuda M, Ishikawa H (2000) Purification and characterization of organic solvent-stable lipase from organic solvent-tolerant Pseudomonas aeruginosa LST-03. J Biosci Bioeng 89(5):451–457
Tamilarasan K, Kumar MD (2012) Purification and characterization of solvent tolerant lipase from Bacillus sphaericus MTCC 7542. Biocatal Agric Biotechnol 1(4):309–313
Panigrahi AK, Panda AK, Bisht SPS (2014) Production and purification of a thermostable lipase from a hyperactive bacterial isolate Brevibacillus sp. from Taptapani hot spring, Odisha. Int J Pharma Bio Sci 5(4):B320–B327
Mykytczuk NC, Foote SJ, Omelon CR, Southam G, Greer CW, Whyte LG (2013) Bacterial growth at −15 °C; molecular insights from the permafrost bacterium Planococcus halocryophilus Or1. ISME J 7(6):1211–1226
Lee D-W, Kim H-W, Lee K-W, Kim B-C, Choe E-A, Lee H-S, Kim D-S, Pyun Y-R (2001) Purification and characterization of two distinct thermostable lipases from the gram-positive thermophilic bacterium Bacillus thermoleovorans ID-1. Enzyme Microb Technol 29(6):363–371
Chang S-W, Shaw J-F, Yang K-H, Chang S-F, Shieh C-J (2008) Studies of optimum conditions for covalent immobilization of Candida rugosa lipase on poly (γ-glutamic acid) by RSM. Bioresour Technol 99(8):2800–2805
Hasan F, Shah AA, Hameed A (2006) Industrial applications of microbial lipases. Enzyme Microb Technol 39(2):235–251
Nerurkar M, Joshi M, Adivarekar R (2015) Bioscouring of cotton using lipase from marine bacteria Bacillus sonorensis. Appl Biochem Biotechnol 175(1):253–265
Lan D, Hou S, Yang N, Whiteley C, Yang B, Wang Y (2011) Optimal production and biochemical properties of a lipase from Candida albicans. Int J Mol Sci 12(10):7216–7237
Acknowledgments
The authors are very thankful of Dr. Kavyani for helpful advices and also appreciate of all postgraduate students in Shahid Beheshti University for scientific suggestions about statistical analysis and fermentation method.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors would like to confirm that there is no conflict of interest associated with this publication and there has been no significant facilities that cover financial matters for this work.
Rights and permissions
About this article
Cite this article
Teymouri, M., Karkhane, M., Gilavand, F. et al. Extracellular Lipase Purification from a Marine Planomicrobium sp. MR23K and Productivity Optimization in a Pilot-Scale Submerged Bioreactor. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 88, 739–746 (2018). https://doi.org/10.1007/s40011-016-0812-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-016-0812-1