Abstract
Giant cell tumor of bone (GCTB) is a rare bone tumor with osteolytic features, composed of stromal cells with a monotonous appearance, macrophages, and osteoclast-like giant cells. GCTB is commonly associated with a pathogenic mutation in the H3-3A gene. While complete surgical resection is the standard cure for GCTB, it often results in local recurrence and, rarely, metastasis. Thus, an effective multidisciplinary treatment approach is necessary. Although patient-derived cell lines is an essential tool for investigating novel treatment strategies, there are only four GCTB cell lines available in public cell banks. Therefore, this study aimed to establish novel GCTB cell lines and successfully created NCC-GCTB6-C1 and NCC-GCTB7-C1 cell lines from two patients' surgically removed tumor tissues. These cell lines exhibited H3-3A gene mutations, consistent proliferation, and invasive properties. After characterizing their behaviors, we performed high-throughput screening of 214 anti-cancer drugs for NCC-GCTB6-C1 and NCC-GCTB7-C1 and integrated their screening data with those of NCC-GCTB1-C1, NCC-GCTB2-C1, NCC-GCTB3-C1, NCC-GCTB4-C1, and NCC-GCTB5-C1 that we previously established. We identified histone deacetylase inhibitor romidepsin as a possible treatment for GCTB. These findings suggest that NCC-GCTB6-C1 and NCC-GCTB7-C1 could be valuable tools for preclinical and basic research on GCTB.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Giant cell tumor of bone (GCTB) is an intermediate-malignant tumor composed of mononuclear stromal cells with a monotonous appearance admixed with macrophages and osteoclast-like giant cells [1]. Genetically, at least 92% of GCTB harbor pathogenic H3-3A gene mutations [1, 2]. Epidemiologically, GCTB is classified as a rare tumor as a study encompassing 10 countries indicated that approximately one person per million per year is affected [3]. GCTB affects the working-age population; approximately 80% of cases occur between 20 and 40 years of age [4, 5].
Clinically, GCTB is locally aggressive, destructive, and grows rapidly, typically around the knee, proximal humerus, distal radius bone, spine, and pelvis [6]. If it is resected surgically, there is a probability of recurrence, which is greater in some locations associated with more difficult treatment, such as the distal radius and proximal femur [7, 8]. Therefore, semi-permanent treatment with denosumab, a receptor activator of nuclear factor kappa-Β ligand (RANKL) inhibitor approved for the treatment of osteoporosis, is currently being discussed as a treatment for GCTB [6, 9, 10]. Although denosumab therapy for GCTB causes dramatic changes in the number of osteoclasts in the tumor tissue, it does not eradicate tumor cells [11, 12]. In addition, jaw necrosis is a principal complication (11%) of denosumab treatment. Thus, there is a need to develop novel therapeutic strategies for treating GCTB.
Using patient-derived cell lines for basic research has been crucial in cancer research and has aided in the development of novel anti-cancer therapies [13,14,15]. However, it has been challenging to use GCTB cell lines for research purposes. According to the Cellosaurus cell line database [16] (https://web.expasy.org/cellosaurus/), although 14 GCTB cell lines (NCI Thesaurus: C4304) have been reported (Supplementary Table 1), only four GCTB cell lines deposited in cell banks are currently available for research. Additionally, a recent study screening 1001 cancer cell lines with 265 compounds to assess the effect of genomic alterations on drug response found that a 50% reduction in the number of cell lines resulted in an 80% loss of significant associations between molecular features and drug response [17, 18]. Given these findings, establishing more cell lines from GCTB patients is crucial.
In this study, we report two novel GCTB cell lines, NCC-GCTB6-C1 and NCC-GCTB7-C1, established from tumor tissues resected from different GCTB patients. We investigated cell characteristics, such as proliferation, spheroid formation, and invasion, to demonstrate the usefulness of these cell lines. We also integrated drug response data from GCTB cell lines in the NCC sarcoma cell line panel (https://web.expasy.org/cgi-bin/cellosaurus/search?input=%22NCC%20sarcoma%20cell%20line%20panel%22), which includes NCC-GCTB1-C1 [19], NCC-GCTB2-C1 [20], NCC-GCTB3-C1 [20], NCC-GCTB4-C1 [21], and NCC-GCTB5-C1 [22], to reveal the potential of these cell lines in drug screening.
Materials and methods
Patient history of two cases
Case 1
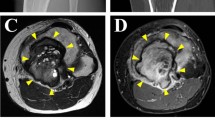
The donor patient was a 38-year-old male with GCTB. The patient had no relevant medical history. He had experienced pain in his right knee without any onset and visited a hospital. X-ray inspection and computed tomography (CT) revealed an osteolytic lesion with subtle cortical destruction in the distal femur (Fig. 1a, b). Magnetic Resonance Imaging (MRI) detected the lesion as homogeneously low-intensity on both T1 and T2 weighted image (Fig. 1c, d). He underwent needle biopsy and GCTB was suspected. The patient was referred to the National Cancer Center Hospital (Tokyo, Japan) for further treatment. No metastasis was detected on the whole-body CT scan. The patient underwent aggressive curettage of the lesion with reinforcement using autologous bone graft implantation and plate fixation. Tumor tissues obtained at the time of surgery were used to establish cell lines. Histologically, the tumor consisted of multinucleated osteoclast-like giant cells and spindle-shaped mononuclear cells. The latter cells were diffusely immunopositive for H3-3A G35W (Fig. 1e, f). The postoperative course was good, and no local recurrence was reported without adjuvant therapy. The use of clinical materials for this study was approved by the ethical committee of the National Cancer Center (Tokyo, Japan) and written informed consent was obtained from the donor patient.
Clinical and pathological data for case 1. X-ray and CT revealed an osteolytic lesion inside the femur delineated by yellow arrows (a: A-P view), with subtle cortical destruction on CT (b: coronal view). MRI detected a mass inside the distal femur and exhibited homogeneously low intensity on T2 (c: coronal view) and T1 weighted image (d: coronal view). H&E staining showed multinucleated osteoclast-like giant cells and spindle-shaped mononuclear cells, where the latter are diffusely immunopositive for H3-3A G35W (e, f)
Case 2
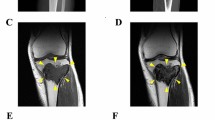
The donor patient was a 31-year-old male with GCTB. The patient had no relevant medical history. He experienced pain in his left wrist and left it for two years. The pain was exacerbated acutely, and he visited the hospital. X-ray inspection and CT revealed an osteolytic lesion at the distal radius entailing the undisposed fracture (Fig. 2a, b). The patient was referred to the National Cancer Center Hospital (Tokyo, Japan) for further treatment. MRI revealed a neoplastic lesion within the radius (Fig. 2c, d). After conservative treatment of the fracture, the patient underwent aggressive curettage of the lesion and was diagnosed with GCTB. Tumor tissues obtained at the time of surgery were used to establish cell lines. Histologically, the tumor consisted of uniform mononuclear neoplastic cells and scattered osteoclast-like giant cells with fresh hemorrhage (Fig. 2e). The neoplastic cells were immunohistochemically diffusely positive for H3-3A G35W (Fig. 2f). Postoperative progression was favorable, and no local recurrence was reported without adjuvant therapy. The use of clinical materials for this study was approved by the ethical committee of the National Cancer Center (Tokyo, Japan) and written informed consent was obtained from the donor patient.
Clinical and pathological data for case 2. Radiography revealed the fracture, indicated by the red arrow (a: lateral view). CT shows an osteolytic lesion inside the radius, delineated by yellow arrows (b: coronal view). MRI revealed a mass inside the distal radius and detected homogeneously high intensity by short tau inversion recovery image (c: coronal view) as well as gadolinium enhanced T1 weighted image (d: axial view) delineated by yellow arrows. H&E staining shows uniform mononuclear neoplastic cells and scattered osteoclast-like giant cells with fresh hemorrhages (e). The neoplastic cells are immunohistochemically positive for H3-3A G35W (f)
Histological analysis of tumor tissue
Histological examination was performed on 4-μm-thick sections of a representative paraffin-embedded tumor sample. The sections were deparaffinized and stained with hematoxylin and eosin (H&E). They were also stained with an H3-3A G35W antibody (1:1000, RM263; RevMAb Biosciences USA, South San Francisco, CA, USA) to investigate H3-3A G35W expression.
Primary cell isolation and culture
Primary tumor cells of GCTB were collected from surgically resected tumor tissues that were dissected into small pieces with scissors. Using the gentleMACS™ Octo Dissociator with Heaters (Miltenyi Biotec, Bergisch Gladbach, Germany), the dissected tissues in the gentleMACS™ C Tube (Miltenyi Biotec) were digested and homogenized using the Tumor Cell Dissociation Kit, human (Miltenyi Biotec) for 30 min at 37 °C. The obtained cells were seeded on 60 mm collagen type I-coated culture plates (Sumitomo Bakelite Co. Ltd., Tokyo, Japan). To maintain the cells, we used DMEM/F-12 medium (Thermo Fisher Scientific Inc., Waltham, MA, USA) supplemented with GlutaMAX (Thermo Fisher Scientific Inc.), 5% heat-inactivated fetal bovine serum (FBS) (Thermo Fisher Scientific Inc.), 10 µM Y-27632 (ROCK inhibitor; Selleck Chemicals, Houston, TX, USA), 10 ng/mL basic fibroblast growth factor (bFGF; Sigma-Aldrich Co. LLC, St. Louis, MO, USA), 5 ng/mL epidermal growth factor (EGF; Sigma-Aldrich Co. LLC), 5 µg/mL insulin (Sigma-Aldrich Co. LLC), 0.4 µg/mL hydrocortisone (Sigma-Aldrich Co. LLC), 100 μg/mL penicillin, and 100 µg/mL streptomycin (Nacalai Tesque Inc., Kyoto, Japan). The culture medium was changed every 2–3 days. Cell confluence was observed using a microscope (Carl Zeiss AG, Land Baden-Württemberg, Germany). When the cultured cells reached sub-confluency, they were washed with PBS (–) (Nacalai Tesque Inc.), followed by dissociation with Trypsin–EDTA solution (Nacalai Tesque Inc.). The cells were then transferred to tissue culture plates and maintained at 37 °C in a humidified atmosphere containing 5% CO2.
Authentication and quality control
For authentication and quality control of the established cell lines, DNA was extracted from the tumor tissues and established cell lines. The Wizard® Genomic DNA Purification Kit (Promega Co., Madison, WI, USA) and Qiagen DNeasy Blood and Tissue Kit (QIAGEN N.V., Hilden, Germany) were used to extract DNA, and the DNA concentration was measured using the NanoDrop 8000 (Thermo Fisher Scientific Inc.). Authentication of the established cell lines was conducted by short tandem repeat (STR) analysis for 10 loci using the GenePrint 10 system (Promega Co.) and a 3500xL Genetic Analyzer (Thermo Fisher Scientific Inc.).
Sequence of mutation
Using the QIAzol Lysis Reagent (QIAGEN) and miRNeasy Mini Kit (QIAGEN), total RNA was extracted from GCTB cells to detect gene mutations. The extracted RNA was reverse-transcribed to complementary DNA using SuperScript III reverse transcriptase (Invitrogen) according to the manufacturer’s instructions. The H3-3A gene was amplified by PCR using the H3-3A forward primer H3-3A_F (5′-TAAAGCACCCAGGAAGCAAC-3′), H3-3A reverse primer H3-3A_R (5′-CAAGAGAGACTTTGTCCCATTTTT-3′), and Platinum Taq DNA Polymerase High Fidelity (Life Technologies Co., Carlsbad, CA, USA). PCR products were purified using the Wizard SV Gel and PCR Clean-Up System (Promega Co.), and direct sequencing was performed using the BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Waltham, MA, USA) and Applied Biosystems 3130xL Genetic Analyzer (Thermo Fisher Scientific Inc.) by GENEWIZ. Plasmid Editor v2.0.61 was used to analyze the sequence results.
Spheroid formation assay
The spheroid formation ability was assessed as previously described [21]. The established cells were seeded at a density of 1 × 105 cells/well in a 96-well Clear Round Bottom Ultra-Low Attachment Microplate (Corning Inc., NY, USA), and spheroid formation was confirmed by microscopic observation (KEYENCE Co., Osaka, Japan). After 3 days of culture, the spheroids picked from the plate were covered in gel using iPGell (GenoStaff Co. Ltd., Bunkyo-ku, Tokyo, Japan) and fixed with 10% formalin neutral buffer solution. To prepare paraffin sections of the fabricated spheroids, gel-covered spheroids were embedded in paraffin and sliced into 4 μm-thick paraffin sections. The sectioned spheroids were subjected H&E staining, followed by microscopic observation.
Tumor cell proliferation assay
The proliferative ability of the cells was measured using the Cell Counting Kit-8 (Agilent Technologies Inc., CA, USA). The cells (2.5 × 104) were placed in DMEM/F12 supplemented with GlutaMAX, 5% FBS, 10 µM Y-27632, 10 ng/mL bFGF, 5 ng/mL EGF, 5 µg/mL insulin, 0.4 µg/mL hydrocortisone, 100 µg/mL penicillin, and 100 µg/mL streptomycin.
Tumor cell invasion assay
The invasive ability of the GCTB cell lines NCC-GCTB1-C1 [19], NCC-GCTB2-C1 [20], NCC-GCTB3-C1 [20], NCC-GCTB4-C1 [21], NCC-GCTB5-C1 [22], NCC-GCTB6-C1, and NCC-GCTB7-C1 was measured using a real-time cell analyzer, xCELLigence (Agilent Technologies Inc.). DMEM/F12 supplemented with GlutaMAX, 5% FBS, 10 µM Y-27632, 10 ng/mL bFGF, 5 ng/mL EGF, 5 µg/mL insulin, 0.4 µg/mL hydrocortisone, 100 µg/mL penicillin, and 100 µg/mL streptomycin was added to the lower chamber. The membrane in the upper chamber was coated with Matrigel basement membrane matrix (Corning Inc.) and the chamber was filled with 4 × 104 cells suspended in DMEM/F12 without any supplements. The cells invaded the bottom chamber from the upper chamber through a membrane and were attached to the electronic sensors on the underside of the membrane. The real-time cell analyzer monitored the electrical impedance of the electronic sensors that were influenced by the invading cells every 15 min for 72 h. The MG-63 osteosarcoma cell line (Japanese Collection of Research Bioresources Cell Bank, Osaka, Japan) [23] was used as a control.
Tumorigenicity assay in nude mice
Tumorigenesis of established cell lines in nude mice was assessed as previously described [21]. Cells (1 × 106) suspended in 50 µL of PBS (–) were mixed with an equal volume of Matrigel (21.2 mg/mL). The cell suspension was subcutaneously injected into BALB/c nude mice (CLEA Japan Inc., Tokyo, Japan) using a 5 mL syringe (Terumo Corp., Tokyo, Japan) and a 26 G needle (Terumo Corp.). Tumor size was measured weekly. Animal experiments were conducted in compliance with the guidelines of the Institute for Laboratory Animal Research, National Cancer Center Research Institute.
Screening for anti-cancer drugs
Drug screening tests were performed as previously described [21]. In this experiment, the anti-cancer effects of 214 drugs, including Food and Drug Administration-approved anti-cancer agents (Selleck Chemicals, Houston, TX, USA; Supplementary Table 2), were assessed. Cells (5 × 103) were seeded in a 384-well plate and incubated for 1 day (n = 2). On the second day, 10 µM of each drug was added to the wells. On the fifth day, cell proliferation was measured based on the absorbance at 450 nm after treatment with Cell Counting Kit-8. Cell viability was calculated by comparison with the DMSO control. The obtained data were analyzed using the results for NCC-GCTB1-C1 [19], NCC-GCTB2-C1 [20], NCC-GCTB3-C1 [20], NCC-GCTB4-C1 [21], and NCC-GCTB5-C1 [22], which we previously reported. Quantile normalization was performed using R (version 4.0.3, limma package version 3.46.0, Bioconductor) and unsupervised hierarchical clustering was performed using the ‘gplots’ package (version 3.1.0, CRAN, https://cran.r-project.org).
The half-maximal inhibitory concentration (IC50) value was calculated for drugs that were identified in the preceding screening study and standard treatment drugs, as previously reported. The drugs were added to similarly seeded cells at 10 different concentrations from 0.1 nM to 100 µM. Based on the cell viability calculated similarly, IC50 values were calculated using GraphPad Prism 9.1.1 (GraphPad Software, San Diego, CA, USA).
Results
Establishment and authentication of GCTB cell lines
GCTB cell lines were established from the patients in cases 1 and 2 and named NCC-GCTB6-C1 and NCC-GCTB7-C1, respectively. These two cell lines were maintained for over 25 passages for 5 months. To authenticate the established cell lines, we examined 10 microsatellite sites (STRs) and found that these STRs were identical in the original tumor tissues and the established cell lines (Table 1, Supplementary Fig. 1). A database search revealed that the STR patterns of NCC-GCTB6-C1 and NCC-GCTB7-C1 cells did not match those of the existing cell lines deposited in public cell banks. These results indicate that NCC-GCTB6-C1 and NCC-GCTB7-C1 are novel cell lines. NCC-GCTB6-C1 and NCC-GCTB7-C1 cells were negative for mycoplasma contamination as no mycoplasma-specific DNA was found in the cell-conditioned medium (data not shown).
Characterization of GCTB cell lines
Sanger sequencing revealed the presence of the H3-3A G35W mutation, the most typical mutation in GCTB, in NCC-GCTB6-C1 (Fig. 3a) and NCC-GCTB7-C1 cells (Fig. 3b). NCC-GCTB6-C1 and NCC-GCTB7-C1 cells exhibited adherent characteristics with a spindle and polygonal appearance (Fig. 4a, b). The cells were able to form spheroids when seeded on low-attachment plates. H&E-stained spheroid sections showed spindle and round cell morphologies. There were no giant cells in the spheroid (Fig. 4c, d). Based on the growth curve, the population doubling times of NCC-GCTB6-C1 and NCC-GCTB7-C1 were 21 and 28 h, respectively (Fig. 4e). NCC-GCTB6-C1 and NCC-GCTB7-C1 cells demonstrated a stronger invasion than the MG63 osteosarcoma cell line (Fig. 4f). After subcutaneous injection of NCC-GCTB6-C1 (passage 20) and NCC-GCTB7-C1 (passage 20), tumorigenesis in nude mice was not observed for 2 months (data not shown).
Characterization of GCTB cell lines. a, b Microscopic appearance of a NCC-GCTB6-C1 (passage 21) and b NCC-GCTB7-C1 (passage 20) cells under 2D culture conditions. c, d The H&E stained spheroid sections of c NCC-GCTB6-C1 (passage 22) and d NCC-GCTB7-C1 (passage 21) fabricated in 96-well low-attachment round bottom plates. e Growth curve of NCC-GCTB6-C1 (passage 23) and NCC-GCTB7-C1 (passage 21) cell lines. Each point represents the mean ± standard deviation (n = 3). f Invasion capability of NCC-GCTB1-C1 (passage 32), NCC-GCTB2-C1 (passage 14), NCC-GCTB3-C1 (passage 29), NCC-GCTB4-C1 (passage 16), NCC-GCTB5-C1 (passage 28), NCC-GCTB6-C1 (passage 22), and NCC-GCTB7-C1 (passage 21) cells was observed using a Real-Time Cell Analyzer. MG63 osteosarcoma cell line (passage 25) was used as control
Sensitivity to anti-cancer drugs
The anti-cancer effects of 214 drugs on NCC-GCTB6-C1 and NCC-GCTB7-C1 cells were evaluated. The cell viability of NCC-GCTB6-C1 and NCC-GCTB7-C1 after drug treatment is summarized in Supplementary Table 3 along with the five previously established GCTB cell lines in our laboratory. The 214 drugs were classified into three categories depending on their anti-cancer effects: cluster A was the effective group, cluster B was the intermediate effect group, and cluster C was the poor effect group (Fig. 5a). The proportion of tyrosine kinase inhibitors and molecular-targeted agents was higher in clusters A and B than in cluster C (Fig. 5b). Cluster A included a higher proportion of topoisomerase inhibitors and anti-microtubule agents than the other clusters (Fig. 5c). As for tyrosine kinase inhibitors, ALK inhibitors occupied a high proportion of the effective group (Fig. 5d). Among molecular-targeted drugs, a high proportion of HDAC, MDM2 and PI3K inhibitors belonged to the effective group (Fig. 5e).
Drug screening tests on GCTB cell lines. a The drugs were categorized into three groups according to their anti-cancer effects: cluster A, effective group; cluster B, intermediate effect group; cluster C, poor effect group. b–e The proportion of each drug type belonging to each cluster. The graphs are depicted after the normalization of the number of drugs. Data concerning NCC-GCTB1-C1, NCC-GCTB2-C1, NCC-GCTB3-C1, NCC-GCTB4-C1, and NCC-GCTB5-C1 were previously reported. Passage number of NCC-GCTB6-C1 and NCC-GCTB7-C1 were 20 and 16, respectively
The IC50 values were calculated, and the growth curves of the HDAC inhibitor romidepsin, with the lowest IC50 values, are shown in Fig. 6 and Table 2, along with those of NCC-GCTB1-C1 [18], NCC-GCTB2-C1 [19], NCC-GCTB3-C1 [19], NCC-GCTB4-C1 [17], NCC-GCTB5-C1 [22], NCC-GCTB6-C1, and NCC-GCTB7-C1.
Discussion
In this study, two novel cell lines from patients with GCTB, NCC-GCTB6-C1 and NCC-GCTB7-C1, were established and characterized. The current treatment strategy for GCTB carries the risk of recurrence after resection as well as the side effects of denosumab. Although denosumab therapy for GCTB causes dramatic changes in the number of osteoclasts in the tumor tissue, it does not eradicate tumor cells [11, 12]. Patient-derived cell lines are a promising tool for developing a replaceable treatment strategy. The basic study of GCTB is stagnant owing to the lack of well-characterized cell line models. We believe that the establishment of GCTB cell lines will accelerate in vitro preclinical research for the development of novel therapeutic strategies.
NCC-GCTB6-C1 and NCC-GCTB7-C1 cell lines represent typical characteristics analyzed from several aspects. Errani et al. reviewed 349 GCTB cases and reported that the most common location was the knee, followed by the distal radius and proximal femur [7]. The prevalence of GCTB is higher in females than males [24, 25]. NCC-GCTB6-C1 and NCC-GCTB7-C1 cell lines were derived from the right knee and left wrist (distal radius), respectively, of males in their thirties. Thus, NCC-GCTB6-C1 and NCC-GCTB7-C1 were established from a typical GCTB site with minor sex distribution. Original tumor tissues of NCC-GCTB6-C1 and NCC-GCTB7-C1 cells were diffusely immunopositive for H3-3A G35W. Sanger sequencing revealed a H3-3A G35W mutation in both NCC-GCTB6-C1 and NCC-GCTB7-C1 cells consisted with these original tissues. Considering that at least 92% of GCTB harbor pathogenic H3-3A gene mutations [1, 2], NCC-GCTB6-C1 and NCC-GCTB7-C1 are genetically representative GCTB cell lines.
These novel GCTB cell lines showed several characteristics suitable for in vitro functional research. The constant proliferation and invasive capabilities of NCC-GCTB6-C1 and NCC-GCTB7-C1 are able to utilize as indicators of the sensitivity to anti-cancer agents. In addition, we confirmed the spheroid formation abilities of NCC-GCTB6-C1 and NCC-GCTB7-C1 cells. Currently, tumor cell spheroids have attracted attention as models for evaluating chemotherapy [26, 27]. NCC-GCTB6-C1 and NCC-GCTB7-C1 cells have the potential to conduct research using their spheroids.
In the screening of 214 anti-cancer drugs, we classified them based on their anti-cancer effects. An effective group of cytotoxic drugs included topoisomerase inhibitors and alkylating drugs, that of tyrosine kinase inhibitors included ALK inhibitors, and that of molecular-targeted drugs included HDAC, MDM2, and PI3K inhibitors. According to ClinicalTrials.gov (https://www.clinicaltrials.gov/), none of the alkylating drugs and topoisomerase, ALK, HDAC, MDM2, and PI3K inhibitors have been clinically investigated for use in GCTB treatment and are thus worth exploring further. Of these drugs, romidepsin, an HDAC inhibitor, approved by the US Food and Drug Administration (FDA) for the clinical use of peripheral T-cell lymphoma, showed the lowest IC50 value for GCTB cell lines [28, 29]. HDAC acts as a transcription repressor, owing to its histone deacetylation effect, and consequently promotes chromatin condensation [30]. Aberrant HDAC expression is associated with advanced disease and poor clinical outcomes in several cancer types, such as prostate, colorectal, breast, lung, liver, and gastric cancers [31, 32]. The results of our drug screening with GCTB cell lines are concordant with those reported previously. Venneker et al. established four patient-derived cell lines of GCTB and found them to be sensitive to five HDAC inhibitors including romidepsin, in both 2D and 3D models [33]. In a patient-derived xenograft model of GCTB, the anti-cancer effects of the novel HDAC inhibitor ITF-2357 were revealed via drug screening [34]. Thus, using GCTB cell lines, additional investigation of romidepsin is warranted for developing treatment strategies for GCTB.
This study had several limitations. Although NCC-GCTB6-C1 and NCC-GCTB7-C1 cells are suitable for in vitro studies, tumorigenesis was not observed in BALB/c nude mice injected with these cells. Therefore, NCC-GCTB6-C1 and NCC-GCTB7-C1 cells may not be suitable for xenograft experiments. Severely immunosuppressed mice may be utilized to observe tumorigenesis of NCC-GCTB6-C1 and NCC-GCTB7-C1 cells. In addition, considering that the 13,117 drugs for human were approved as reported in the fact sheet of the US FDA, the number of drugs screened in this study may not be sufficient. We report novel cell lines of GCTB, and the application of these cell lines should be challenged in future studies.
In conclusion, we established two novel GCTB cell lines—NCC-GCTB6-C1 and NCC-GCTB7-C1—that exhibit several characteristics suitable for in vitro functional research. Using these cell lines, we revealed the anti-cancer effects of romidepsin, an HDAC inhibitor, on GCTB. According to the ClinicalTrials.gov, romidepsin has not been used in clinical trials of patients with GCTB. Further analysis of the efficacy of romidepsin in GCTB is hence required. Together, these results indicate that patient-derived GCTB cell lines have potential use in preclinical and basic research on GCTB.
References
Organisation mondiale de la sant Cidrslc (2020) Soft tissue and bone tumours
Behjati S, Tarpey PS, Presneau N, et al. Distinct H3F3A and H3F3B driver mutations define chondroblastoma and giant cell tumor of bone. Nat Genet. 2013;45:1479–82.
Liede A, Bach BA, Stryker S, et al. Regional variation and challenges in estimating the incidence of giant cell tumor of bone. JBJS. 2014;96:1999.
Jamshidi K, Karimi A, Mirzaei A. Epidemiologic characteristics, clinical behavior, and outcome of the giant cell tumor of the bone: a retrospective single-center study. Arch Bone Jt Surg. 2019;7:538–44.
Sobti A, Agrawal P, Agarwala S, Agarwal M. Giant cell tumor of bone—an overview. Arch Bone Jt Surg. 2016;4:2–9.
Borkowska AM, Szumera-Ciećkiewicz A, Szostakowski B, Pieńkowski A, Rutkowski PL. Denosumab in giant cell tumor of bone: multidisciplinary medical management based on pathophysiological mechanisms and real-world evidence. Cancers (Basel). 2022;14:2290.
Errani C, Ruggieri P, Asenzio MA, et al. Giant cell tumor of the extremity: a review of 349 cases from a single institution. Cancer Treat Rev. 2010;36:1–7.
van der Heijden L, Dijkstra PDS, van de Sande MAJ, et al. The clinical approach toward giant cell tumor of bone. Oncologist. 2014;19:550–61.
Zaheer S, LeBoff M, Lewiecki EM. Denosumab for the treatment of osteoporosis. Expert Opin Drug Metab Toxicol. 2015;11:461–70.
Palmerini E, Chawla NS, Ferrari S, et al. Denosumab in advanced/unresectable giant-cell tumour of bone (GCTB): for how long? Eur J Cancer. 2017;76:118–24.
Kato I, Furuya M, Matsuo K, Kawabata Y, Tanaka R, Ohashi K. Giant cell tumours of bone treated with denosumab: histological, immunohistochemical and H3F3A mutation analyses. Histopathology. 2018;72:914–22.
Forsyth RG, Krenács T, Athanasou N, Hogendoorn PCW. cell biology of giant cell tumour of bone: crosstalk between m/wt Nucleosome H3.3, telomeres and osteoclastogenesis. Cancers (Basel). 2021;13:5119.
Barretina J, Caponigro G, Stransky N, et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Garnett MJ, Edelman EJ, Heidorn SJ, et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature. 2012;483:570–5.
Folkesson E, Niederdorfer B, Nakstad VT, et al. High-throughput screening reveals higher synergistic effect of MEK inhibitor combinations in colon cancer spheroids. Sci Rep. 2020;10:11574.
Bairoch A. The cellosaurus, a cell-line knowledge resource. J Biomol Tech JBT. 2018;29:25–38.
Williams SP, McDermott U. The pursuit of therapeutic biomarkers with high-throughput cancer cell drug screens. Cell Chem Biol. 2017;24:1066–74.
Iorio F, Knijnenburg TA, Vis DJ, et al. A landscape of pharmacogenomic interactions in cancer. Cell. 2016;166:740–54.
Noguchi R, Yoshimatsu Y, Ono T, et al. Establishment and characterization of NCC-GCTB1-C1: a novel patient-derived cancer cell line of giant cell tumor of bone. Hum Cell. 2020;33:1321–8.
Yoshimatsu Y, Noguchi R, Tsuchiya R, et al. Establishment and characterization of novel patient-derived cell lines from giant cell tumor of bone. Hum Cell. 2021;34:1899–910.
Ono T, Noguchi R, Yoshimatsu Y, et al. Establishment and characterization of the NCC-GCTB4-C1 cell line: a novel patient-derived cell line from giant cell tumor of bone. Hum Cell. 2022;35:392–9.
Akiyama T, Yoshimatsu Y, Noguchi R, et al. Establishment and characterization of NCC-GCTB5-C1: a novel cell line of giant cell tumor of bone. Hum Cell. 2022;35:1621–9.
Billiau A, Edy VG, Heremans H, et al. Human interferon: mass production in a newly established cell line, MG-63. Antimicrob Agents Chemother. 1977;12:11–5.
Beebe-Dimmer JL, Cetin K, Fryzek JP, Schuetze SM, Schwartz K. The epidemiology of malignant giant cell tumors of bone: an analysis of data from the surveillance, epidemiology and end results program (1975–2004). Rare Tumors. 2009;1: e52.
Chakarun CJ, Forrester DM, Gottsegen CJ, Patel DB, White EA, George R, Matcuk J. Giant cell tumor of bone: review, mimics, and new developments in treatment. Radiographics. 2013;33:197–211.
Perche F, Torchilin VP. Cancer cell spheroids as a model to evaluate chemotherapy protocols. Cancer Biol Ther. 2012;13:1205–13.
Sant S, Johnston PA. The production of 3D tumor spheroids for cancer drug discovery. Drug Discov Today Technol. 2017;23:27–36.
Grant C, Rahman F, Piekarz R, et al. Romidepsin: a new therapy for cutaneous T-cell lymphoma and a potential therapy for solid tumors. Expert Rev Anticancer Ther. 2010;10:997–1008.
Lee HZ, Kwitkowski VE, Del Valle PL, et al. FDA approval: belinostat for the treatment of patients with relapsed or refractory peripheral T-cell lymphoma. Clin Cancer Res. 2015;21:2666–70.
Xu WS, Parmigiani RB, Marks PA. Histone deacetylase inhibitors: molecular mechanisms of action. Oncogene. 2007;26:5541–52.
Seto E, Yoshida M. Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harb Perspect Biol. 2014;6: a018713.
West AC, Johnstone RW. New and emerging HDAC inhibitors for cancer treatment. J Clin Investig. 2014;124:30–9.
Venneker S, van Eenige R, Kruisselbrink AB, et al. Histone deacetylase inhibitors as a therapeutic strategy to eliminate neoplastic “stromal” cells from giant cell tumors of bone. Cancers (Basel). 2022;14:4708.
Yafei J, Haoran M, Wenyan J, et al. Personalized medicine modality based on patient-derived xenografts from a malignant transformed GCTB harboring H3F3A G34W mutation. J Orthop Transl. 2021;29:106–12.
Acknowledgements
We thank Drs. E. Kobayashi, S. Iwata, K. Ogura, K. Sato (Department of Musculoskeletal Oncology), Dr. Y. Takahashi (Department of Diagnostic Pathology), and the National Cancer Center Hospital for sampling tumor tissue specimens from surgically resected materials. We appreciate the technical support provided by Mrs. Y. Shiotani, Mr. N. Uchiya, and Dr. T. Imai (Central Animal Division, National Cancer Center Research Institute). We would like to thank Editage (www.editage.jp) for providing English language editing services and for their constructive comments on this manuscript. This study was technically assisted by the Fundamental Innovative Oncology Core of the National Cancer Center.
Funding
This research was supported by the Japan Agency for Medical Research and Development (Grant number: 20ck0106537h0003).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethics approval
The ethical committee of the National Cancer Center (Tokyo, Japan) approved the use of clinical materials for this study (approval number 2004-050). Animal experiments were conducted in compliance with the guidelines of the Institute for Laboratory Animal Research, National Cancer Center Research Institute.
Informed consent
Written informed consent was provided by the patients.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13577_2023_928_MOESM1_ESM.tif
Supplementary file1 Supplementary Fig. 1 Short tandem repeat patterns of GCTB cell lines and original tumor tissues. (a) Short tandem repeat patterns of NCC-GCTB6-C1 (passage 20), (b) Short tandem repeat patterns of original tumor tissue of NCC-GCTB6-C1, (c) Short tandem repeat patterns of NCC-GCTB7-C1 (passage 20), (d) Short tandem repeat patterns of original tumor tissue of NCC-GCTB7-C1 (TIF 517 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ono, T., Noguchi, R., Yoshimatsu, Y. et al. Establishment and characterization of two novel patient-derived cell lines from giant cell tumor of bone. Human Cell 36, 1804–1812 (2023). https://doi.org/10.1007/s13577-023-00928-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-023-00928-0