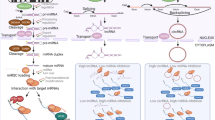
Abstract
Background
Pancreatic ductal adenocarcinoma (PDAC) is a lethal disease with a dismal prognosis for which new therapeutic strategies are desperately needed. Non-coding RNAs (ncRNAs), especially microRNAs (miRNAs) and long non-coding RNAs (lncRNAs), may yield new therapeutic concepts for the treatment of PDAC. A vast number of miRNAs, including the well-studied miR-21, miR-155 and miR-34, has been shown to regulate PDAC growth, invasion and metastasis in vitro and in vivo by targeting members of key signaling pathways. In addition, other miRNAs and lncRNAs, such as HOTTIP and MALAT-1, have been shown to influence the malignant behavior of PDAC cells.
Methods
Here, we discuss several ncRNAs that may be used either as new therapeutic agents or as targets of new therapeutic agents. Furthermore, we discuss the problem of proper delivery of nucleotide-based agents and novel methods that may be used to circumvent this problem.
Conclusions
Although the number of reports addressing the role of ncRNAs in PDAC virtually grows by the day, there are still many steps to be taken before the application of ncRNA-based therapies will become reality in clinical practice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Pancreatic ductal adenocarcinoma (PDAC) is a devastating disease with a poor prognosis. Each year more than 40,000 PDAC-related deaths occur in the European Union, making it the fourth most common cause of cancer death in men and women [1]. The main reasons for its devastating character are the late occurrence of clinical symptoms and the intrinsic malignant nature of the disease [2]. Since curative treatment in the form of radical surgery is only possible in a minority of patients, novel therapeutic options are urgently needed [2]. Common genetic alterations include mutations in the KRAS (>90 %), TP53 (> 50 %), SMAD4 (> 60 %) and CDKN2A (> 80 %) genes [2–4]. Moreover, studies in murine models have shown that TP53 and KRAS mutations, when occurring in a rapid sequence, underlie the development of PDAC in 75 % of the cases [5]. The introduction of the FOLFIRINOX and Gemcitabine plus Nab-Paclitaxel chemotherapy regimens has resulted in a successful prolongation of the survival of PDAC patients, but with frequent and sometimes considerable adverse effects, such as severe rashes [6]. Even though preliminary data show efficacy of small molecule agents such as the tyrosine-kinase inhibitor Erlotinib, successful targeted therapies for the treatment of PDAC have yet to be developed [6].
Non-coding RNAs (ncRNAs) are RNA molecules that are involved in translation, DNA replication, RNA splicing and epigenetic regulation. They are not translated into proteins [7–11]. ncRNAs comprise transfer RNA (tRNA), ribosomal RNA (rRNA), small interfering RNA (siRNA), Piwi-interacting RNA (piRNA), microRNA (miRNA), Y-RNA and small Cajal body-specific RNA (scaRNA). As a group, ncRNAs are believed to be ancient relics of evolution, whereas in modern animals and plants, different mechanisms by which ncRNAs regulate gene expression and DNA replication have evolved. Within the human genome, genes encoding ncRNAs are more than five times as numerous as those encoding proteins. ncRNAs may be transcribed in intergenic regions or regions overlapping with genes encoding proteins [12]. Moreover, mutations in ncRNA genes may underlie disease development, analogous to mutations in protein coding genes [13]. Interestingly, some chemotherapeutic agents, such as Metformin or Vorinostat, are believed to function at least partly through modifying the expression of ncRNAs [14, 15]. Especially miRNAs and lncRNAs have shown therapeutic potential, as they can be introduced into cells via viral-vector transfection or mechanical transfection, conceptually resembling ‘classical’ gene therapy. Delivery of therapeutics to the proper site is important, and methods have been reported that provide the possibility of local application. Of these, endoscopic ultrasound-guided fine-needle injection (EUS-FNI) or computer-tomography (CT)-guided delivery of therapeutic nucleotides have proven to be successful in PDAC-patients [16].
1.1 MicroRNA characteristics
MicroRNAs (miRNAs) represent a subgroup of 19 to 25 nucleotides-long ncRNAs that post-transcriptionally regulate gene expression by a series of steps referred to as RNA interference [17, 18]. miRNAs are evolutionarily conserved and constitute an important part of epigenetic gene regulation, affecting up to 60 % of all mammalian genes, and have been subject to intensive research in recent years [19–25]. From an evolutionary perspective, miRNAs may have facilitated the development of higher complexities in organisms and concomitant biologic functions, making life as we know it now possible [26]. The biogenesis of miRNAs is achieved by the processing of RNAs transcribed by RNA polymerase II (RNAP-II) [27] and the processing of hairpin-shaped pre-miRNAs to miRNAs by the enzymes Drosha and Dicer in the nucleus and the cytoplasm, respectively [28–30]. Subsequently, single stranded miRNA molecules associate with argonaute proteins to form the RNA-induced silencing complex (RISC), which then binds to complementary messenger RNA (mRNA) to either block translation initiation or to initiate mRNA degradation [31]. RISC and its associated miRNAs usually bind to the 3′ untranslated region (3′-UTR) of the complementary mRNA [31]. Perfect complementary binding of a miRNA with its target mRNA leads to mRNA cleavage and degradation [32]. In mammals, miRNA binding to its target mRNA is mostly imperfect. This imperfect binding leads to a blockade of translation of the target mRNA [33]. However, the seed region of a miRNA, which represents a 7 nucleotide long region (nucleotides 2-8) in the 3′-UTR, still needs to bind perfectly to its mRNA target in order to block translation. Some mRNAs may have multiple seed regions complementary to a variety of miRNAs [34]. Hence, a single mammalian miRNA may inhibit the expression of hundreds of genes, while a single gene may undergo translational inhibition by numerous miRNAs [34]. This notion may provide a rationale for the development of future therapeutic strategies, as agents with multiple targets may act synergistically and, as such, may preclude the development of therapy resistance. Related to their function, miRNAs that promote aggressiveness in tumors are termed “onco-miRs”, whereas miRNAs that have the opposite effects are termed “tumor-suppressor-miRs”. Onco-miRs may inhibit the translation of tumor suppressor genes such as PTEN, while tumor-suppressor-miRs may inhibit the translation of oncogenes such as KRAS [35]. Also, the expression of onco-miRs and tumor-suppressor-miRs itself is subject to specific regulatory mechanisms. It has for example been found that KRAS enhances the expression of onco-miR miR-21 while, conversely, the expression of KRAS is reduced by miR-217 [35, 36]. Thus, teleologically, loss of miR-217 function may increase the expression of miR-21 through increased KRAS signaling. This notion adds another layer of complexity to the traditional concept of tumor-suppressor genes and oncogenes. A recent study revealed that at least 500 miRNAs are aberrantly expressed in PDAC [37]. Next to PDAC, the role of some of these miRNAs has also been investigated in the two major precursors of PDAC, i.e., pancreatic intraepithelial neoplasia (PanIN) and intraductal papillary mucinous neoplasm (IPMN) [38–40].
1.2 Long non-coding RNA characteristics
Long non-coding RNAs (lncRNAs) are transcripts defined by a length of over 200 nucleotides that are involved in the regulation of gene transcription, post-transcriptional gene regulation and epigenetic gene regulation [41–43]. lncRNAs are transcribed by RNAP-II, poly-adenylated and spliced similar to mRNAs, but are not translated into protein [44]. lncRNAs represent a heterogeneous group of RNAs that can be divided into different subgroups. From a functional perspective, lncRNAs may be assigned to archetypical categories, i.e., signaling, decoying, scaffolding and guidance, although most lncRNAs are multifunctional [45]. Signaling lncRNAs are RNAs that regulate gene expression intrinsically, i.e., by binding to target genes or regulatory elements. A prime example is the X-chromosomal inactivation-related lncRNA Xist. In female mammals, Xist is expressed from the inactive X-chromosome and covers it to repress gene expression [46]. Subsequent gene silencing and guidance of chromatin modifying enzymes to target locations on the X-chromosome is, however, conducted by distinct RNA sequences, thereby assigning it to both categories of lncRNA function [47]. Decoy lncRNAs affect gene expression in an indirect manner by binding to RNA-binding proteins (RBPs). Decoy lncRNAs consequently titrate away proteins such as transcription factors and chromatin modifying enzymes [45]. Examples of decoy lncRNAs are metastasis-associated MALAT-1 and mTOR pathway-associated Gas5 [45, 48]. Scaffolding lncRNAs serve as platforms for complexes of different effector molecules, analogous to the function of scaffolding proteins [45]. A good example of a scaffolding lncRNA is HOTAIR, which has been implicated in pancreatic carcinogenesis and acts by forming a complex with chromatin modifying enzymes [49]. Regarding their association with genomic regions, lncRNAs may be divided into five classes: promoter-associated lncRNAs (pRNAs), antisense and sense gene body-associated lncRNAs (gsRNAs), enhancer-associated lncRNAs (eRNAs) and intervening lncRNAs (lincRNAs) [50]. In recent years, lncRNAs have been shown to play essential roles in human diseases, such as cancer, endocrine disorders (e.g. diabetes mellitus) and cardiovascular diseases [51–53]. lncRNAs have also been found to be involved in the pathophysiology of PDAC [37] and some of them appear to show potential for diagnostic and/or RNA-based gene therapeutic purposes.
2 ncRNA-based therapy of PDAC
In the search for putative novel therapeutic targets, miRNAs have been studied intensively in recent years. On the one hand as drug targets in the form of miRNA antagonists such as antisense oligonucleotides (ASOs), and on the other hand as miRNA mimics [39, 54] or miRNA-encoding plasmid cDNA (pcDNA) that can restore miRNA or lncRNA function [55]. ASOs are single-stranded RNA molecules that share the same sequence as their target mRNA or ncRNA, but in a reversed order. Binding to their complementary RNA leads to degradation of ASO-mRNA or ASO-ncRNA complexes by RNase H [56]. ASOs that target miRNAs are usually called anti-miRs or antagomiRs. miRNA mimics are synthetic double-stranded RNA molecules that are homologous to endogeneous miRNAs. Similar to miRNAs, lncRNAs have been targeted experimentally, both in vitro and in vivo. It has been found that over-expression of lncRNAs can be achieved by introducing gene-carrying pcDNAs in target cells. lncRNA silencing can, on the other hand, be brought about using siRNAs or shRNAs. Also, methods to target lncRNAs with ASOs have been developed. The latter confers the advantage of nuclear inhibition of lncRNAs rather than cytoplasmic inhibition [57]. A notable exception is lncRNA H19, which is central to a different therapeutic approach and is discussed in more detail under 2.2.8 [58]. Here, emphasis is put on well-investigated ncRNAs in PDAC, some of which have been successfully tested in vitro and in vivo and may serve as candidates for RNA-based gene therapy.
2.1 Potential therapeutic miRNAs
2.1.1 KRAS-related miRNAs
A number of miRNAs has been shown to regulate KRAS expression in PDAC. This is an interesting finding since KRAS is mutated in > 90 % of PDAC precursor lesions and PDACs (see above) and, as such, is thought to be one of the driving oncogenes in pancreatic carcinogenesis [2, 59]. Inhibition of the MAKP/ERK-pathway, of which KRAS is an upstream member, has shown anti-oncogenic effects in PDAC cells [60]. Mutant KRAS, in turn, has shown to regulate miRNAs in PDAC-derived cell lines, as well as in human Nestin-expressing cells (HPNE) derived from non-neoplastic epithelium (NNE) of pancreatic ducts harboring a KRAS mutation [61]. KRAS is a known target of tumor-suppressor-miRs that are deregulated in PDACs. These include miR-217, miR-206, miR-143, miR-145 and Let-7 [35, 55, 62, 63]. It has been shown that transfection with miR-217, which is lowly expressed in PDAC cells, significantly down-regulates KRAS expression and upregulates phosphorylated-AKT1 levels in PDAC-derived cell lines. Additionally, in vitro reduced anchorage-independent colony formation and in vivo reduced tumor growth in xenograft models has been reported [35]. Active KRAS exerts its oncogenic function through up-regulation and repression of miRNAs. miR-31 is a recently discovered KRAS-regulated miRNA that represses RASA1 in a MAPK-dependent manner, leading to increased RhoA-GTPase activity. It has been reported that exogenous expression of miR-31 leads to increased proliferation, migration and invasion of PDAC-derived cells [64]. miR-143 and miR-145, which are down-regulated in the presence of activated KRAS, have been found to lead to inhibition of proliferation and to mediate the induction of apoptosis in PDAC-derived cells in vitro and in vivo [61, 63, 65]. Kent et al. observed down-regulation of miR-143 and miR-145 in PDAC-derived cells and showed that viral-mediated delivery of miR-143 and miR-145 resulted in decreased anchorage-independent growth [61]. KRAS-mediated repression of miR-143 and miR-145 has also been observed in other tumor types, such as colorectal carcinoma (CRC) [66–68]. It has also been found that miR-206 re-expression has inhibitory effects, not only on tumor growth and tumor vascularization in vitro and in vivo, but also on the motility and invasive behavior of PDAC cells. These effects are achieved at least in part through the inhibition of pro-inflammatory and pro-angiogenic genes downstream of the NF-κB pathway, such as those encoding chemokine (C-X-C-motif) ligand 1 (CXCL1), granulocyte colony-stimulating factor-2 (CSF2) and vascular endothelial growth factor A and B (VEGFA, VEGFB), but they may also be achieved through direct inhibition of KRAS mRNA itself [62].
2.1.2 miR-155
Activating KRAS-mutations may also lead to increased expression of onco-miR miR-155, mediated by MAPK and NF-κB signaling, thereby promoting oxidative stress responses by inactivation of the tumor-suppressive transcription factor Foxo3a [69]. miR-155 has been found to be over-expressed in most PDAC cases and to correlate with a poor survival of PDAC patients [70]. During pancreatic carcinogenesis, miR-155 has been found to become over-expressed with a gradually increasing frequency during the progression of PanINs, the most important precursor lesions of PDAC, with highest expression levels in PanIN-3 lesions [39]. miR-155 is a known inhibitor of TP53INP1, which mediates apoptosis upon DNA damage by promoting p53 activation [71]. It is also involved in the creation of a tumor-growth promoting microenvironment by transforming normal pancreatic fibroblasts into cancer-associated fibroblasts (CAFs) that allow PDAC cell invasion, and by promoting tumor-angiogenesis [72]. Anti-miR-155 has been shown to reduce the invasive behavior and migration of PDAC cells by down-regulation of the signal transducer and activator of transcription 3 (STAT3), which can be activated through various signaling pathways, such as the EGF, Il-6 and interferon-related pathways [73, 74]. Hence, miR-155 may serve as a promising target for experimental PDAC therapy, as it is a well-investigated miRNA that has important implications both in early pancreatic carcinogenesis and in advanced stages of PDAC. Research aimed at the development of KRAS-targeting strategies still awaits satisfactory results. The failure so far to develop agents that target KRAS may be due to the complex crystallographic structure of the KRAS protein [6], a problem that may be circumvented by miRNA-based approaches.
2.1.3 miR-21
miR-21 is one of the most extensively investigated miRNAs in malignant tumors, and has been found to be involved in many oncogenic pathways [75–77]. miR-21 has also been investigated in PDAC, and it has been shown that its expression level correlates well with several prognostic parameters and with resistance to Gemcitabine [78–81]. In PDAC it has been found that miR-21 exerts its oncogenic function through up-regulation of Bcl-2 [82], inhibition of PTEN and RECK [83], FasL [80], p85α [84] and PDCD4 [85]. It has also been found to be up-regulated by KRAS signaling, Gemcitabine-induced histone-deacetylation of the MIR21 promoter [36] and by hypoxia [86]. Consequently, miR-21 is a strong enhancer of PI3K-AKT signaling, thus promoting the growth, invasion and migration of PDAC cells [87]. Moreover, miR-21 is involved in creating a tumor-promoting environment by the recruitment of CAFs, which promotes the invasion and metastasis of PDAC cells [88]. Several studies have shown that inhibition of miR-21 via ASOs leads to decreased invasion and metastasis [79], decreased proliferation and increased apoptosis [83] and Gemcitabine sensitivity in PDAC cells [79, 81, 83, 89]. In one study, mice inoculated with tumor cells were treated with anti-miR-21 after palpable tumor formation. Twelve days after intra-tumor injection the control group showed fast progressing tumors, whereas the group of mice treated with anti-miR-21 showed little to no progression [89]. A combined use of anti-miR-21 and the receptor tyrosine-kinase inhibitor Sunitinib led to a synergistic decrease of cell viability, which was more pronounced than with a combination of Sunitinib and anti-miR-221, another highly up-regulated miRNA in PDAC [90, 91]. In view of current data, targeting of miR-21 appears a valid option in the treatment of PDAC, as it is upregulated in most PDACs and interferes with essential malignancy-associated signaling pathways.
2.1.4 miR-34
Recently, a number of studies has revealed an important tumor-suppressive activity of miR-34 in PDAC through synergistic interaction with the p53 pathway. This holds true for all three known homologues of miR-34, referred to as miR-34a-c, respectively. Although these homologues all share a seed sequence, their expression levels vary strongly among different tissues, suggesting tissue-specific roles [92]. It has been shown that miR-34a and b can inhibit PDAC growth and viability, although different mechanisms underlying these inhibitions have been reported. Insight into the role of miR-34c is still lacking. It has been reported that miR-34a exerts its function partly by inhibition of target genes encoding Bcl-2, Notch-1 and Notch-2 [93, 94], while miR-34b also inhibits the expression of SMAD3, thereby repressing proliferative signaling through the TGF-β pathway [95]. It has also been found that p53 wild-type PDAC-derived cell lines show considerably increased apoptotic rates when transfected with miR-34 mimics [96]. Interestingly, inhibition of tumor growth has also been seen in p53-deficient PDAC-derived cell lines, suggesting the occurrence of p53-independent interactions of miR-34 [94, 96]. Other investigators found that treatment with the demethylating agent 5-Aza-dC and the HDAC inhibitor Vorinostat resulted in re-expression of miR-34a and a consecutive up-regulation of p53. In addition, it was found that this treatment may lead to an increased expression of the downstream signaling molecules p21, p27 and PUMA, while cell proliferation, cell cycle progression and EMT (epithelial to mesenchymal transition) were inhibited. Additionally, it was found that stem cell characteristics were reduced [15]. Vorinostat is already in clinical use for the treatment of refractory cutaneous T-cell-lymphoma and has an acceptable adverse effects profile, which is probably due to its selective targeting of tumor cells [97]. Also, a successful safety study on PDAC patients has been performed [98]. The estrogenic isoflavone Genistein has been shown to inhibit PDAC growth and to induce apoptosis by enhancing the expression of miR-34a [93]. Genistein has little to no side effects [99], rendering it into an interesting therapeutic option. Clearly, the miR-34 family of miRNAs has therapeutic potential for PDAC, as the p53 pathway is commonly altered in these tumors. Moreover, miR-34 may also exert its tumor-suppressive function via p53-independent pathways, such as the TGF-β pathway. Since at least one of these pathways is altered in most PDACs, miR-34 may be suitable for a broad-scale application [2]. Since the targeted delivery of miRNAs is still challenging (see below), the use of agents such as Vorinostat may support the development of new targeted therapies. As miR-34 is among the most strongly dysregulated miRNAs in different malignancies such as hepatocellular carcinoma (HCC) or leukemia [100, 101], efforts have already been made to utilize miR-34 mimics for tumor therapy. MRX34, which will be discussed in detail later in this article, is a miR-34-mimic-based therapeutic that is currently evaluated for the treatment of unresectable HCCs and has shown promising results in murine models [102]. Apart from miR-34, miR-148 has also been found to be efficacious in PDAC, in part via Bcl-2 inhibition, leading to significant reduction in tumor growth and invasive behavior both in vitro and in vivo [103]. A similar mode of action has been reported for miR-345, although only one study so far has dealt with PDAC [104].
2.1.5 TGF-β-related miRNAs
Other tumor-suppressor-miRs that have been reported to target the TGF-β pathway are miR-483 and miR-367. miR-483-3p has been shown to inhibit PDAC growth through down-regulation of SMAD4, leading to decreased growth-inhibitory signaling of the TGF-β pathway [105]. The SMAD4 gene has been found to be mutated in > 60 % of the PDAC cases (see above). Therefore, it is assumed that tumors that do not exhibit SMAD4 mutations may over-express miR-483-3p in a higher proportion, although this has so far not been documented. Interestingly, the SMAD4 gene status poorly correlates with its mRNA-level, suggesting that post-transcriptional regulation by miRNAs may be at work [106]. It has been found that miR-367 targets the inhibitory SMAD protein SMAD7, leading to an increased in vitro PDAC-derived cell growth [107]. As yet, however, data on the exact role of SMADs, including SMAD7, in the pathobiology of PDAC are still conflicting, although it has been suggested that down-regulation of SMAD4 and SMAD7 is essential [106]. Fact is, nevertheless, that the TGF-β pathway is highly deregulated in PDACs, and that it may represent a pivotal target for targeted therapy.
2.1.6 MUC4-related miRNAs
MUC4 is a membranous high-molecular weight glycoprotein that enhances the malignant behavior of PDAC cells by repressing apoptosis and immune modulation, and by aggravating chemoresistance. It has been shown that MUC4 expression is regulated by miRNAs [108–111]. MUC4 and the epidermal-growth factor receptor-2 (HER-2/neu) are proven targets of miR-150, and they have been found to interact closely to promote HER-2 activation. This interaction leads to an increased stability of the HER-2/neu receptor, thus enhancing downstream signaling and tumor cell proliferation [111, 112]. Restoration of miR-150 expression has led to decreased growth and invasiveness of PDAC-derived cell lines in in vitro studies. The effect could mainly be attributed to repression of MUC4 expression by miR-150, although other targets of miR-150 have also been identified [111]. Another miRNA that targets MUC4 is miR-219-1-3p. It was found that miR-219-1-3p over-expression in PDAC-derived cell lines successfully reduced activation of the AKT and MAPK/ERK pathways through inhibition of MUC4 expression. Furthermore, it was found that injection of miR-219-1-3p mimics reduced tumor growth and MUC4 expression in a murine PDAC xenograft model [113]. The use of miRNAs that target MUC4 seems promising, as MUC4 is a molecule involved in oncogenic pathways essential for PDAC, such as the MAPK/ERK and AKT signaling pathways. Moreover, MUC4 and HER-2 have been found to be over-expressed in most PDAC cases [2], suggesting that targeting these molecules with miRNAs might be considered for the majority of patients.
2.1.7 miR-200
MiR-200, comprising miR-200a-c and miR-141, is known to exhibit tumor suppressive properties and is currently being evaluated as therapeutic agent in a variety of tumors [114–116]. MiR-200 exerts its tumor suppressive action by inhibiting key steps in the EMT cascade, thereby inducing epithelial characteristics in tumor cells [117]. Conversely, miR-200 promotes MET (mesenchymal to epithelial transition), the last step of metastasis in which migratory tumor cells form stable colonies [118]. The expression of miR-200a has been found to be markedly decreased in PDAC cells, especially in its cancer stem-cells (CSCs). Moreover, it has been found that a low expression of miR-200a correlates with EMT-related traits such as a high expression of Vimentin and a low expression of E-cadherin [119]. It has also been found that re-expression of miR-200a using miR-200a-mimics can restore epithelial traits in PDAC CSCs in vitro, while decreasing their invasive and migratory behavior [119]. Ma et al. showed that miR-200c mimics, when transfected into PDAC cells, exhibited a negative effect on their metabolism with a subsequent decrease in survival, invasive behavior and colony forming capacity [120]. Furthermore, subpopulations of CSCs typically expressing the stem-cell markers CD44 and CD24, showed a pronounced dose-dependent decrease in cell viability upon Gemcitabine treatment [120].
2.1.8 miR-141
Another important member of the mir-200 family is miR-141, which exerts its function through inhibition of the Yes-associated protein-1 (YAP1), the transmembrane-4-L-6-family-1 (TM4SF1) protein and the mitogen-activated protein kinase isoform 4 (MAP4K4) [121–123]. A low miR-141 expression is frequently found in PDAC and has been shown to correlate with an impaired overall survival and advanced clinic-pathological features in PDAC patients [123, 124]. In vitro functional studies have shown that re-expression of miR-141 using miR-141 mimics results in a reduced growth and colony forming capacity and an increased apoptosis, effects that are largely attributable to YAP1 inhibition [124]. Lijian et al. showed that miR-141 mimics led to suppressed PDAC cell migration and invasion, although no effect on cell growth or apoptosis was observed [122]. Zhao et al. found that in vitro miR-141 over-expression may lead to G1-phase cell cycle arrest and induction of apoptosis in conjunction with down-regulation of cyclin D1 and Bcl-2 expression. The colony forming and invasive capacities of the respective cells were markedly inhibited. In addition, it was found in murine xenograft models that miR-141 can inhibit in vivo tumor cell growth [123].
2.1.9 miR-375
miR-375 is a miRNA that is expressed especially in pancreatic islet cells under conditions that are necessary for an appropriate development of the endocrine pancreas and the regulation of glucose-dependent insulin secretion [125, 126]. miR-375 has been found to be down-regulated in PDAC cells and to function via the targeting of pyruvate dehydrogenase kinase-1 (PDK-1), Insulin-Like Growth Factor Binding Protein 5 (IGFBP5) and Caveolin-1 [127, 128]. It has also been found that miR-375 signaling leads to reduced activation of the AKT pathway in PDAC cells [128]. Furthermore, over-expression of miR-375 has repeatedly been shown to inhibit PDAC cell growth and to induce apoptosis [127–129]. Zhou et al. found that miR-375 over-expression may lead to a significant increase of cells in the G1/G0-phase of the cell cycle, a reduced cell proliferation and an increased apoptotic rate [128, 129]. The same group showed that miR-375 over-expression may be effective in suppressing tumor growth in xenograft models. It was found that after 1 month mice with solid PDAC tumors that were injected with miR-375 mimics showed significantly smaller tumor volumes than the non-injected controls [130].
2.1.10 miR-221
miR-221 is one of the most strongly deregulated miRNAs in PDACs and PanINs, and has been found to be over-expressed in other tumor entities as well [127, 131, 132]. In PDAC, it effectively enhances tumor viability and resistance to Gemcitabine [83, 133]. It has been found that miR-221 exerts its oncogenic function in PDAC through down-regulation of a number of target genes, including PTEN, p27 (kip1), p57 (kip2), PUMA and tissue inhibitor of metalloproteinases-2 (TIMP-2) [83, 133, 134]. This mode of action may explain its enhancing effects on proliferation, apoptosis and invasive behavior. Xu et al. found that administration of miR-221 mimics inhibited apoptosis and promoted cell proliferation and invasion in vitro, while an anti-miR-221 exhibited opposite effects. These alterations were found to be accompanied by a significant up-regulation of the invasion-associated matrix-metalloproteinases -2 and -9 (MMP-2 and MMP-9) as a result of TIMP-2 inhibition. TIMP-2 is crucial for maintaining a proper ECM-homeostasis that affects the invasive capacities of PDAC cells, and is a target of miR-221 [133]. It has been shown that inhibition of miR-221, using an anti-miR-221, leads to a slow growth, a shortened survival and an up-regulation of tumor-suppressive molecules in PDAC cells in vitro, while miR-221 mimics were found to induce opposite effects [127, 134]. Basu et al. showed in a mouse model that miR-221 expression is triggered by overactive KRAS, i.e., activating KRAS mutations, indicating that KRAS acts as an upstream regulator of miR-221 [127]. Tumor suppressor-miRs and onco-miRs that may be employed for targeted PDAC therapy are listed in Tables 1 and 2, respectively. In Fig. 1 the putative functions of these miRNAs in PDAC are depicted.
Figure 1 illustrates the function of hitherto investigated microRNAs (miRNAs) in PDAC. In the PDAC cell, miRNAs are involved in the regulation of important signaling pathways by inhibiting expression of their target molecules at the mRNA level. miRNAs with oncogenic function are marked in red, whereas miRNAs with tumor suppressive function are marked in green
2.2 Potential therapeutic lncRNAs
In comparison to miRNAs, there is a paucity of literature on lncRNAs in PDAC but, nevertheless, pivotal roles of lncRNAs in this tumor entity have been delineated as well [240]. A recent next generation-based sequencing study revealed that at least 319 lncRNAs may be deregulated during pancreatic carcinogenesis [241]. In addition, it has been shown that lncRNA deregulation constitutes an important aspect of the metastatic process [242]. The affected lncRNAs often appear to be involved in (de)regulation of the MAPK pathway [242], indicating that these ncRNAs may contribute to PDAC metastasis formation by promoting uncontrolled growth. In Table 3 lncRNAs that may potentially be useful for PDAC therapy are listed, whereas in Fig. 2 the functions and effects of these lncRNAs are depicted.
2.2.1 Gas5
Growth arrest-specific 5 (Gas5) is a lncRNA that functions by negatively regulating cell cycle progression. It is expressed at a low level in several tumor entities such as B-cell lymphoma and cervical cancer [243–245]. Gas5 is a downstream member of the mammalian target of rapamycin (mTOR) pathway, which is currently subject of intense research in the field of targeted therapy [48]. It has been found that in prostate cancer mTOR inhibition leads to up-regulation of Gas5 while, in reverse, silencing of Gas5 reduces the effect of mTOR inhibition [246]. In PDAC, Gas5 expression is reduced compared to normal pancreatic ductal epithelium. Lu et al. showed that inoculation of Gas5 pcDNA-containing vectors leads to a significant in vitro reduction of G1/S-phase cells in the PDAC-derived cell lines PANC1 and BxPC3 by negatively regulating cyclin-dependent kinase 6 (CDK6) expression [247]. Based on these observations, reintroduction of Gas5 may therapeutically be beneficial. In addition, Gas5 may have potential as a biomarker for clinical response in mTOR-targeted therapy in PDAC as well as in other tumors due to the close relationship between mTOR signaling and Gas5 expression. As such, Gas5 may facilitate the selection of patients that are eligible for mTOR-targeted therapy and, additionally, for monitoring the efficacy of mTOR-targeted therapy in PDAC patients.
2.2.2 MALAT-1
The > 8000 nucleotides-long intronic ncRNA (lincRNA) metastasis-associated lung adenocarcinoma transcript-1 (MALAT-1) is a highly conserved lncRNA with oncogenic properties that was first detected in non-small cell lung cancer (NSCLC) [248]. MALAT-1 is a lincRNA with a hitherto poorly understood function. It is up-regulated in PDAC and especially in PDAC-CSCs [249] and is located predominantly in the nucleus. It has been shown that MALAT-1 expression correlates with various clinicopathologic features and a poor clinical outcome in PDAC patients [249]. Recent work has revealed a beneficial effect of MALAT-1 inhibition in PDAC through different molecular mechanisms. Silencing of MALAT-1 leads to decreased cellular growth via a p21-mediated cell cycle arrest at the G1/S-phase transition and an increased p53-dependent apoptosis [250]. In addition, MALAT-1 silencing has been found to result in decreased migration and down-regulation of the expression of EMT-associated mesenchymal markers [250]. MALAT-1 silencing has also been found to decrease tumor vascularization and to increase sensitivity to Gemcitabine [251]. These observations provide promising outlooks for an assessment of MALAT-1 as therapeutic target in PDAC. Additionally, it certainly has potential as a prognostic biomarker and as an indicator of metastasis and chemoresistance in PDAC patients.
2.2.3 HULC
Highly up-regulated in liver cancer (HULC) is a cytoplasmic 482 nucleotides-long lncRNA. It was first described in HCC where it regulates the expression of multiple genes associated with the malignant behavior of tumor cells [252]. It has been found to be up-regulated in the presence of Hepatitis B protein x (HBx) and to promote HBV-mediated liver cirrhosis. In PDAC, HULC expression has been found to correlate significantly with a reduced overall survival and time to recurrence. In addition, HULC expression turned out to be highly sensitive and specific in distinguishing cancer tissue from healthy tissue. Silencing of HULC has been demonstrated to result in a decreased viability and proliferation of PDAC cells by blocking the G1/S-phase transition [253]. Thus, HULC may be used to monitor patients after curative surgery and, in addition, may serve as a potential therapeutic target.
2.2.4 HOTAIR
HOX transcript antisense RNA (HOTAIR) is an oncogenic lncRNA and negative prognostic factor in a variety of solid cancers including breast, colon and pancreatic cancer [254]. It has been discovered that increased HOTAIR expression levels correlate with more advanced tumor stages and a short overall-survival in PDAC patients [255]. HOTAIR acts as a molecular scaffold to coordinate the function of the chromatin modifying enzymes Polycomb Repressive Complex 2 (PRC2) and LSD1 to propagate gene suppression of the HOXD locus. In human embryogenesis, it is expressed predominantly in distal anatomic regions, i.e. the head and tail of the embryo [256]. It has been found that HOTAIR promotes the malignant behavior of PDAC cells by different mechanisms [49]. HOTAIR promotes e.g. the invasive behavior of PDAC cells by regulating the expression of genes such as GDF15, although its effects appear to vary drastically between primary tumors and tumor-derived cell lines, which may at least partly be explained by varying PRC2 activities [49, 257]. Recent data indicate that HOTAIR inhibition may reinforce the anti-proliferative and pro-apoptotic effects of radiotherapy, in part by up-regulating Wnt inhibitory factor 1 (WIF-1) [258]. Although HOTAIR may serve as a potential therapeutic target in PDAC, future studies should be aimed at identifying tumors that might benefit most of HOTAIR inhibition. Moreover, the effect of HOTAIR expression on the chemoresistance of PDAC should be evaluated, as recent work has shown that HOTAIR may promote chemoresistance in ovarian cancers [259].
2.2.5 HOTTIP
The lincRNA HOXA transcript at the distal tip (HOTTIP) has been shown to up-regulate HOXA gene expression through interaction with WDR5/MLL and subsequent histone methylation [260, 261]. Recently, HOTTIP expression has been linked to an impaired survival in HCC patients and to overlap with regulation of the HOTAIR gene, although the latter regulates primarily HOXD genes [262]. HOTTIP has been identified as one of the most profoundly up-regulated lncRNAs in PDAC cells compared to non-neoplastic pancreatic duct epithelium [240]. Knock-down of HOTTIP has been found to lead to cell cycle inhibition at the G1/S-phase transition, a decrease in the expression of EMT-associated factors and enhanced sensitivity to Gemcitabine, both in vitro and in vivo. HOTTIP exerts its effects on PDAC cells mainly via up-regulation of HOXA13 expression, but also via up-regulation of Aurora A kinase, which may promote aneuploidy [263, 264]. HOTTIP over-expression has been associated with a low degree of tumor differentiation, lymphatic metastases and early postoperative recurrences [240]. HOTAIR-targeted therapy may hold promise, especially in early stage cancers. Since it is up-regulated especially in tumors prone to recur early in the postoperative phase, it could be used to select patients who might benefit from aggressive adjuvant chemotherapy. These patients would benefit most from simultaneous HOTTIP targeted therapies, as this may enhance the chemosensitivity of PDAC cells.
2.2.6 ENST00000480739
The recently discovered lncRNA ENST00000480739 has been shown to exhibit tumor-suppressive properties in PDACs, and to be lowly expressed in > 90 % of primary tumor samples as well as in a number of PDAC-derived cell lines. Sun et al. found that a high ENST00000480739 expression negatively correlates with the occurrence of lymph node metastases and positively with an improved clinical outcome, thus considering it as an interesting candidate PDAC biomarker. In addition, low ENST00000480739 expression was found to serve as an independent risk factor for impaired postoperative survival. Experimental work has shown that ENST00000480739 over-expression results in suppression of tumor cell invasion, both in vitro and in vivo. Its effects were found to be mediated by up-regulation of osteosarcoma amplified-9 (OS-9), which leads to a decrease in hypoxia inducible factor-1-alpha (HIF-1α) expression levels and a subsequent down-regulation of matrix-metalloproteinase (MMP) expression levels [265]. Because of its anti-invasive effect on PDAC cells, ENST00000480739 may have potential as a therapeutic agent, i.e., its exogenous over-expression may reduce the likelihood of recurrence. Moreover, a low expression after surgery may be considered as an indicator of metastasis and, thus, as a need for re-evaluation of the patient’s TNM status.
2.2.7 LOC389641
The newly discovered lncRNA LOC389641 was found to play an important role in PDAC. Statistical analyses revealed that LOC389641 expression closely correlates with the overall survival and the occurrence of lymph node metastasis in PDAC patients [266]. LOC389641 has been found to be over-expressed in primary PDAC tissues and in PDAC-derived cell lines and, in addition, to contribute to PDAC cell proliferation through cell cycle-independent mechanisms. Additional in vitro and in vivo studies have shown that LOC389641 can increase the invasion and migration of PDAC cells. The latter is caused by inhibition of E-cadherin expression and a subsequent induction of EMT in the tumor cells, indicated by increased expression of the mesenchymal marker Vimentin and the EMT-associated transcription factor Snail. In addition, it has been found that LOC389641 can up-regulate the expression of TNFRSF10A, a gene encoding a TRAIL-receptor, which is a mediator of metastasis-promoting signals in TRAIL-resistant cancer cells [266]. Targeting LOC389641 may be considered as an actionable approach, since the up-regulation of TRAIL receptors is highly specific for tumor cells [267] and the damage of normal cells, therefore, appears unlikely. Moreover, E-cadherin down-regulation is commonly seen in PDACs, suggesting a broad efficacy of the use of LOC389641 inhibition.
2.2.8 H19
lncRNA H19 is highly expressed in many human tissues during embryogenesis until the time of birth, after which its expression levels drop to practically zero [268, 269]. Expression in adult individuals appears to be confined to cancer cells [269–271]. H19 has been found to be highly expressed in PDAC cells, and seems to be involved in the modulation of their migration, invasion and metastasis. This effect is at least partially due to the antagonistic effects of H19 on let-7 activity, which results in up-regulation of HGMA2 expression levels and, subsequently, tumorigenesis [272]. Recently, H19 has been subjected to targeted tumor therapy research. The plasmid-based drug BC-819 that utilizes polyethylenimine-coated plasmid DNA (also known as DTA-H19), induces expression of diphtheria toxin (DT) and cell death in the presence of H19-specific transcription factors, which are restricted to malignant tumors in adults [273]. This drug has been shown to be effective in the treatment of human bladder and ovarian cancer in case studies [274, 275]. In a phase I/IIa trial the effects of ultrasound-guided endoscopic or CT-guided injection of BC-819 (see below) were investigated in patients suffering from PDAC. The results were promising with respect to reducing tumor mass, even leading to resectability in one case by tumor shrinkage. Although the trial was not powered to provide statistically significant results, the safety of treatment was confirmed [58]. Currently, a multicenter randomized phase-IIb trial with the aim to assess the effect of BC-819 on progression-free survival is under way [276].
3 Delivery systems for ncRNA-based therapeutics
Whereas many questions about the pharmacokinetic properties of ncRNA-based therapies remain, the major challenges of ncRNA-based therapies lie in the development of reliable delivery systems. Several systemic therapies with miRNA-carrying viral vectors and non-viral agents have been studied in recent years. Current therapeutic delivery methods that utilize ncRNA can be divided into several different categories: chemically modified ncRNAs, viral and non-viral vectors, ncRNA “sponges”, and endoscopic ultrasound and CT-guided injections [230, 278, 279].
3.1 Chemically modified ncRNAs
It has been reported that by adding chemical backbone modifications to a miRNA mimic or anti-miR, its degradation can be inhibited and its in vitro and in vivo activity can be improved [280]. A recent study has shown that miR-155 could be targeted more effectively when a pH-low insertion peptide (pHLIP) was chemically conjugated with anti-miR-155 oligonucleotides, enabling their accumulation specifically in acidotic tumor tissues. Using this approach, the authors of this study found that intravenous application of this combined agent led to a significant reduction in tumor volume in mice with solid lymphomas [279]. This seems to be an attractive approach, as it overcomes the problems posed by systemic introduction of unmodified oligonucleotides, which mainly undergo hepatic accumulation and subsequent clearance [56]. Moreover, hypoxia and acidosis are typical characteristics of PDAC that could be used in this concept, whereas they represent barriers for conventional therapeutics [281]. Even though data showing that pHLIP-conjugated ncRNAs indeed do accumulate in PDAC tissue are lacking, PDAC-imaging data from mouse models with multispectral optoacoustic tomography do suggest this [282].
3.2 Viral and non-viral vectors
The use of viral vectors may have potential for future therapeutic application, as tropism against cells that express certain molecules can be established. Lentiviral, adeno-associated-viral and adenoviral vectors have been used to express ncRNA in target cells. Viruses possess a complex endogenous machinery for gene processing, which makes them highly effective in transferring nucleotides [283]. Disadvantages of viral vectors are their complicated manufacturing process and possible immunologic, hematologic or oncologic side effects. Non-viral vectors have the advantage of permitting a larger size nucleotide-cargo and an uncomplicated large scale production [283]. The non-viral miR-34-mimic-based therapeutic MRX34 is currently under evaluation in a phase-I clinical trial for the treatment of HCC and, as such, represents the first clinical application of miRNA-mimics [102]. MRX34 utilizes the liposomal carrier-system Smarticles™ for delivery into tumor cells, which has shown promising results in both in vitro and in vivo studies [284]. Whether an analogous approach would lead to satisfactory accumulation in PDACs is unlikely, because of the aforementioned hepatic clearance. This obstacle may be bypassed by the use of poly-ethylene-glycol (PEG)-coated liposomes, also called “stealth”-liposomes. Stealth-liposomes can evade degradation in the reticuloendothelial system and remain in circulation for up to 48 h. The PEG-coats of stealth-liposomes can be chemically equipped with functional groups to enable coupling with tumor-specific antibodies and proteins to target tumor cells or other cell types [285, 286]. Another interesting option that has been known for almost a decade are Carbon nanotobes (CNTs). CNTs are small versatile molecules of condensed rings of carbon atoms. It has been shown that CNTs permit the transport of nucleic acids in vitro and in vivo. In addition, they have the advantage of a high transport capacity. They also allow diverse surface modifications, turning them into interesting candidate non-viral vectors. The specific targeting of solid tumors after systemic delivery still remains a challenge [287]. However, modifications that enable a pH-dependent nucleotide-cargo release have been found to result in an adequate accumulation in solid tumors [278].
3.3 ncRNA sponges
Yet another interesting system is the use of a transfectable “miRNA sponge”, as was recently demonstrated by Jung et al. [230]. In this study, the authors simultaneously inhibited five miRNAs, i.e., miR-21, miR-155, miR-10, miR-211 and miR-222, by sequestration into a circular sponge carrying binding sites for the respective miRNAs, which resulted in a significant reduction in tumor growth and viability [230]. The antiviral drug Miravirsen, which has been successfully tested in humans for the treatment of hepatitis C-virus (HCV), has an analogous function. It binds intracellular miR-122 to inhibit the propagation of viral RNA without toxic side-effects [288]. An equivalent approach is, however, unlikely to work for the treatment of PDAC without further modification, as oligonucleotides are subject to hepatic clearance as outlined above. Therefore, more work is needed to improve the applicability of nucleotide-based therapies, which remains one of the biggest challenges of this research field.
3.4 Endoscopic ultrasound and CT-guided injections
Endoscopic ultrasound-guided fine needle injections (EUS-FNI) for different diagnostic and therapeutic applications in PDAC have been reported [289]. This method involves the use of fine needles attached to an endoscope by which different fluids can be injected into the pancreas during retrograde-cholangiopancreatography (ERCP). The use of EUS-FNI for celiac plexus neurolysis is an upcoming method to treat PDAC-associated pain, whereas also the effect of local application of chemotherapeutics is currently being investigated [289]. CT-guided delivery is performed via a posterior percutaneous access to reach retroperitoneal organs. CT-guided injections have also been used to treat PDAC-associated pain with good results after unsuccessful celiac plexus neurolysis, turning it into a back-up system or alternative method for EUS-FNI [290]. Endoscopic ultrasound-guided or CT-guided injection of BC-819 in PDAC patients has been shown to improve clinical outcomes in case studies, but as yet without statistical benefit [58]. Mouse model data have shown that BC-819 is able to significantly reduce tumor growth and to counteract metastasis in treated mice compared to untreated controls [291]. As yet, however, limitations of this procedure, such as its accuracy or chemoresistance to BC-819, are not well defined. Intraperitoneal injection of miR-26-carrying viral vectors has been proven to successfully induce gene expression in pancreatic beta-cells in murine models [292]. In humans, this approach may provide a simpler, cheaper and less invasive access to the pancreas than EUS-FNI or CT-guided injections, although questions regarding efficacy and side effects can be addressed only by speculation at this point.
4 Perspectives of ncRNA-based therapies in PDAC
Recent research has revealed many ncRNAs that are involved in the pathobiology of various human cancers [293]. Especially miRNAs and lncRNAs have been shown to regulate multiple crucial genes involved in cell cycle progression, apoptosis, invasion, migration and vascularization, often in concert with ncRNAs regulating multiple genes [34]. In PDAC, several miRNAs and lncRNAs have been found to substantially affect tumor behavior by regulating multiple target genes and pathways. This was exploited by using ncRNA mimics or ASOs, delivered by viral or non-viral vectors, for the design of in vitro and in vivo PDAC therapies. In addition, more subtle methods have been employed to create ncRNA-based therapeutics, such as the experimental agent BC-819 and the chemical conjugation of pHLIP to ncRNAs. It has also been found that drugs such as Metformin and Vorinostat inhibit in vitro PDAC cell growth at least in part via down-regulation of certain key miRNAs [14, 15]. Additional more complicated systems, such as the plasmid-based agent BC-819 that utilizes the abnormally expressed lncRNA H19 in PDAC cells resulting in DT production and cell death, may be promising.
Whether combinations of different ncRNA mimics and ASOs exhibit synergistic effects to positively interfere with tumor biology, or whether simultaneous application of ncRNAs may abrogate or enhance the anti-tumor effects of each other is currently unknown, but it has been shown that a combined approach may be superior [230]. This holds also true for the combined use of therapeutic ncRNAs and conventional chemotherapeutic agents such as Gemcitabine or 5-FU, as also other agents such as the tyrosine-kinase inhibitor Sunitinib [90], although clinical trials are still needed. Ye et al. showed that interactions of miRNAs, mRNAs, lncRNAs and transcription factors encompass an intriguing network of regulation, in which several miRNAs and lncRNAs inhibit each other and/or lead to degradation of mRNA [37], which clearly indicates that the administration of several ncRNAs may result in adverse effects. Further knowledge on the regulation of ncRNAs in PDAC may lead to the identification of pivotal ncRNAs that may serve as the most promising candidates for therapeutic approaches and help to understand the subtleties of PDAC biology in further detail.
An important aspect of personalized cancer therapy is knowledge about the genetic and molecular differences between individual patients that result in different responses to therapy. This holds true especially for PDAC, as only 25 % of the patients respond well to Gemcitabine treatment, and the overall survival is only marginally improved in responders [294]. It has amply been shown that tumor-related miRNAs may be easily identified in blood, stool or saliva [25, 295, 296]. This circumstance may enable us to create patient-specific ncRNA profiles to guide tumor therapy. A recent clinical study has shown that miR-21, miR-100 and miR-99 expression levels in tumor specimens after curative surgery in patients with UICC stage II PDACs may predict the response to adjuvant Gemcitabine therapy [297]. So, the measurement of miRNA levels may become a useful tool to guide adjuvant chemotherapy, since alternatives to Gemcitabine treatment have proven more efficacious, but also since they carry the risk of more severe side-effects that may not be acceptable.
5 Conclusions
To date, many ncRNAs have been identified that may serve as targets for tumor therapy, but additional information clearly needs to be gathered before clinical application can be considered. The use of currently available viral delivery systems may pose problems, such as adverse effects or treatment failure due to unknown toxicity, host immunity and the risk of genomic insertion and a consequent risk of (secondary) cancer. Although, non-viral delivery systems have shown promising results, it still needs to be established whether systemic application will result in adequate concentrations at the a priori target site. The intravenously applied miR-34-based non-viral therapeutic agent MRX34 is currently undergoing evaluation in a phase I trial, which may be refined so as to target PDAC cells. In case of insufficient efficacy, local application via EUS-FNI or CT-guided injection may be considered. However, systemic therapy should be the primary aim, as most PDACs are incurable by operation due to the frequent presence of metastases at the time of discovery [2].
As yet, little is known about the possibility of resistance to ncRNA-based therapy, and how to chose the right ncRNA as target out of a plethora of candidate ncRNAs. This connects the search for potential ncRNA therapeutic targets with that for ncRNA-based diagnostics. Studies aimed at early-stage recognition of PDAC using miRNAs alone or in combination with CA19-9 have yielded a higher sensitivity and specificity than using CA19-9 alone [298]. In addition, it has been found that a postoperative prognosis can be established precisely through miRNA measurement [299]. The promising miRNA-mimic-based therapeutic agent MRX34 and several lncRNA-based therapeutics are currently being evaluated in clinical studies, but it will require intensive additional research to finally reach systemic therapy. Therefore, we anticipate that the clinical role of ncRNAs in the near future may primarily lie in its diagnostic use, such as the monitoring of response to conventional chemotherapy, and their use as prognostic markers, as they represent stable differentially expressed molecules that are amenable to sensitive detection. Interestingly, recent high throughput studies have revealed that the lncRNAs that are most widely deregulated in PDAC cells have not been thoroughly investigated yet [241]. Therefore, there may still be many of them around of which the biological role and clinical applicability await discovery.
References
M. Malvezzi, P. Bertuccio, F. Levi, C. La Vecchia, E. Negri, European cancer mortality predictions for the year 2014. Ann. Oncol. 25, 1650–1656 (2014)
D.P. Ryan, T.S. Hong, N. Bardeesy, Pancreatic adenocarcinoma. N. Engl. J. Med. 371, 1039–1049 (2014)
T. Furukawa, R. Fujisaki, Y. Yoshida, N. Kanai, M. Sunamura, T. Abe, K. Takeda, S. Matsuno, A. Horii, Distinct progression pathways involving the dysfunction of DUSP6/MKP-3 in pancreatic intraepithelial neoplasia and intraductal papillary-mucinous neoplasms of the pancreas. Mod. Pathol. 18, 1034–1042 (2005)
S.A. Hahn, A.T. Hoque, C.A. Moskaluk, L.T. da Costa, M. Schutte, E. Rozenblum, A.B. Seymour, C.L. Weinstein, C.J. Yeo, R.H. Hruban, S.E. Kern, Homozygous deletion map at 18q21.1 in pancreatic cancer. Cancer Res. 56, 490–494 (1996)
J.M. Bailey, A.M. Hendley, K.J. Lafaro, M.A. Pruski, N.C. Jones, J. Alsina, M. Younes, A. Maitra, F. McAllister, C.A. Iacobuzio-Donahue, S.D. Leach, p53 mutations cooperate with oncogenic Kras to promote adenocarcinoma from pancreatic ductal cells. Oncogene (2015). doi:10.1038/onc.2015.441
E.G. Chiorean, A.L. Coveler, Pancreatic cancer: optimizing treatment options, new, and emerging targeted therapies. Drug Des. Devel. Ther. 9, 3529–3545 (2015)
C.P. Christov, T.J. Gardiner, D. Szuts, T. Krude, Functional requirement of noncoding Y RNAs for human chromosomal DNA replication. Mol. Cell. Biol. 26, 6993–7004 (2006)
S. Kishore, S. Stamm, Regulation of alternative splicing by snoRNAs. Cold Spring Harb. Symp. Quant. Biol. 71, 329–334 (2006)
A.T. Zhang, A.R. Langley, C.P. Christov, E. Kheir, T. Shafee, T.J. Gardiner, T. Krude, Dynamic interaction of Y RNAs with chromatin and initiation proteins during human DNA replication. J. Cell Sci. 124, 2058–2069 (2011)
Y. Zhu, V. Stribinskis, K.S. Ramos, Y. Li, Sequence analysis of RNase MRP RNA reveals its origination from eukaryotic RNase P RNA. RNA 12, 699–706 (2006)
A. Huttenhofer, P. Schattner, N. Polacek, Non-coding RNAs: hope or hype? Trends Genet. 21, 289–297 (2005)
P. Kapranov, J. Cheng, S. Dike, D.A. Nix, R. Duttagupta, A.T. Willingham, P.F. Stadler, J. Hertel, J. Hackermuller, I.L. Hofacker, I. Bell, E. Cheung, J. Drenkow, E. Dumais, S. Patel, G. Helt, M. Ganesh, S. Ghosh, A. Piccolboni, V. Sementchenko, H. Tammana, T.R. Gingeras, RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007)
R.J. Taft, K.C. Pang, T.R. Mercer, M. Dinger, J.S. Mattick, Non-coding RNAs: regulators of disease. J. Pathol. 220, 126–139 (2010)
R. Tanaka, M. Tomosugi, M. Horinaka, Y. Sowa, T. Sakai, Metformin causes G1-phase arrest via down-regulation of MiR-221 and enhances TRAIL sensitivity through DR5 up-regulation in pancreatic cancer cells. PLoS One 10, e0125779 (2015)
D. Nalls, S.N. Tang, M. Rodova, R.K. Srivastava, S. Shankar, Targeting epigenetic regulation of miR-34a for treatment of pancreatic cancer by inhibition of pancreatic cancer stem cells. PLoS One 6, e24099 (2011)
L. Buscail, B. Bournet, F. Vernejoul, G. Cambois, H. Lulka, N. Hanoun, M. Dufresne, A. Meulle, A. Vignolle-Vidoni, L. Ligat, N. Saint-Laurent, F. Pont, S. Dejean, M. Gayral, F. Martins, J. Torrisani, O. Barbey, F. Gross, R. Guimbaud, P. Otal, F. Lopez, G. Tiraby, P. Cordelier, First-in-man phase I clinical trial of gene therapy for advanced pancreatic cancer: safety, biodistribution and preliminary clinical findings. Mol. Ther. 23, 202–214 (2015)
V. Ambros, The functions of animal microRNAs. Nature 431, 350–355 (2004)
D.P. Bartel, MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004)
K. Chen, N. Rajewsky, The evolution of gene regulation by transcription factors and microRNAs. Nat. Rev. Genet. 8, 93–103 (2007)
A. Tanzer, P.F. Stadler, Molecular evolution of a microRNA cluster. J. Mol. Biol. 339, 327–335 (2004)
M.J. Axtell, D.P. Bartel, Antiquity of microRNAs and their targets in land plants. Plant Cell 17, 1658–1673 (2005)
R.C. Friedman, K.K. Farh, C.B. Burge, D.P. Bartel, Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009)
I. Fkih M’hamed, M. Privat, F. Ponelle, F. Penault-Llorca, A. Kenani, Y.J. Bignon, Identification of miR-10b, miR-26a, miR-146a and miR-153 as potential triple-negative breast cancer biomarkers. Cell. Oncol. 38, 433–442 (2015)
E. Yiannakopoulou, Targeting epigenetic mechanisms and microRNAs by aspirin and other non steroidal anti-inflammatory agents--implications for cancer treatment and chemoprevention. Cell. Oncol. 37, 167–178 (2014)
C. Salazar, R. Nagadia, P. Pandit, J. Cooper-White, N. Banerjee, N. Dimitrova, W.B. Coman, C. Punyadeera, A novel saliva-based microRNA biomarker panel to detect head and neck cancers. Cell. Oncol. 37, 331–338 (2014)
K.J. Peterson, M.R. Dietrich, M.A. McPeek, MicroRNAs and metazoan macroevolution: insights into canalization, complexity, and the Cambrian explosion. Bioessays 31, 736–747 (2009)
Y. Lee, M. Kim, J. Han, K.H. Yeom, S. Lee, S.H. Baek, V.N. Kim, MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060 (2004)
R.I. Gregory, T.P. Chendrimada, R. Shiekhattar, MicroRNA biogenesis: isolation and characterization of the microprocessor complex. Methods Mol. Biol. 342, 33–47 (2006)
E. Lund, J.E. Dahlberg, Substrate selectivity of exportin 5 and Dicer in the biogenesis of microRNAs. Cold Spring Harb. Symp. Quant. Biol. 71, 59–66 (2006)
E. Prodromaki, A. Korpetinou, E. Giannopoulou, E. Vlotinou, M. Chatziathanasiadou, N.I. Papachristou, C.D. Scopa, H. Papadaki, H.P. Kalofonos, D.J. Papachristou, Expression of the microRNA regulators Drosha, Dicer and Ago2 in non-small cell lung carcinomas. Cell. Oncol. 38, 307–317 (2015)
X.J. Wang, J.L. Reyes, N.H. Chua, T. Gaasterland, Prediction and identification of Arabidopsis thaliana microRNAs and their mRNA targets. Genome Biol. 5, R65 (2004)
A. Eulalio, E. Huntzinger, T. Nishihara, J. Rehwinkel, M. Fauser, E. Izaurralde, Deadenylation is a widespread effect of miRNA regulation. RNA 15, 21–32 (2009)
A.A. Bazzini, M.T. Lee, A.J. Giraldez, Ribosome profiling shows that miR-430 reduces translation before causing mRNA decay in zebrafish. Science 336, 233–237 (2012)
B.P. Lewis, I.H. Shih, M.W. Jones-Rhoades, D.P. Bartel, C.B. Burge, Prediction of mammalian microRNA targets. Cell 115, 787–798 (2003)
W.G. Zhao, S.N. Yu, Z.H. Lu, Y.H. Ma, Y.M. Gu, J. Chen, The miR-217 microRNA functions as a potential tumor suppressor in pancreatic ductal adenocarcinoma by targeting KRAS. Carcinogenesis 31, 1726–1733 (2010)
M.C. du Rieu, J. Torrisani, J. Selves, T. Al Saati, A. Souque, M. Dufresne, G.J. Tsongalis, A.A. Suriawinata, N. Carrere, L. Buscail, P. Cordelier, MicroRNA-21 is induced early in pancreatic ductal adenocarcinoma precursor lesions. Clin. Chem. 56, 603–612 (2010)
S. Ye, L. Yang, X. Zhao, W. Song, W. Wang, S. Zheng, Bioinformatics method to predict two regulation mechanism: TF-miRNA-mRNA and lncRNA-miRNA-mRNA in pancreatic cancer. Cell Biochem. Biophys. 70, 1849–1858 (2014)
N. Habbe, J.B. Koorstra, J.T. Mendell, G.J. Offerhaus, J.K. Ryu, G. Feldmann, M.E. Mullendore, M.G. Goggins, S.M. Hong, A. Maitra, MicroRNA miR-155 is a biomarker of early pancreatic neoplasia. Cancer Biol. Ther. 8, 340–346 (2009)
J.K. Ryu, S.M. Hong, C.A. Karikari, R.H. Hruban, M.G. Goggins, A. Maitra, Aberrant MicroRNA-155 expression is an early event in the multistep progression of pancreatic adenocarcinoma. Pancreatology 10, 66–73 (2010)
C. Yu, M. Wang, Z. Li, J. Xiao, F. Peng, X. Guo, Y. Deng, J. Jiang, C. Sun, MicroRNA-138-5p regulates pancreatic cancer cell growth through targeting FOXC1. Cell. Oncol. 38, 173–181 (2015)
J.A. Goodrich, J.F. Kugel, Non-coding-RNA regulators of RNA polymerase II transcription. Nat. Rev. Mol. Cell Biol. 7, 612–616 (2006)
M. Beltran, I. Puig, C. Pena, J.M. Garcia, A.B. Alvarez, R. Pena, F. Bonilla, A.G. de Herreros, A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev. 22, 756–769 (2008)
A. Mazo, J.W. Hodgson, S. Petruk, Y. Sedkov, H.W. Brock, Transcriptional interference: an unexpected layer of complexity in gene regulation. J. Cell Sci. 120, 2755–2761 (2007)
J.H. Yoon, J. Kim, M. Gorospe, Long noncoding RNA turnover. Biochimie 1859, 209–221 (2015)
K.C. Wang, H.Y. Chang, Molecular mechanisms of long noncoding RNAs. Mol. Cell 43, 904–914 (2011)
D.B. Pontier, J. Gribnau, Xist regulation and function explored. Hum. Genet. 130, 223–236 (2011)
A. Wutz, T.P. Rasmussen, R. Jaenisch, Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nat. Genet. 30, 167–174 (2002)
M. Mourtada-Maarabouni, A.M. Hasan, F. Farzaneh, G.T. Williams, Inhibition of human T-cell proliferation by mammalian target of rapamycin (mTOR) antagonists requires noncoding RNA growth-arrest-specific transcript 5 (GAS5). Mol. Pharmacol. 78, 19–28 (2010)
K. Kim, I. Jutooru, G. Chadalapaka, G. Johnson, J. Frank, R. Burghardt, S. Kim, S. Safe, HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene 32, 1616–1625 (2013)
R. Bonasio, R. Shiekhattar, Regulation of transcription by long noncoding RNAs. Annu. Rev. Genet. 48, 433–455 (2014)
S. Uchida, S. Dimmeler, Long noncoding RNAs in cardiovascular diseases. Circ. Res. 116, 737–750 (2015)
M. Knoll, H.F. Lodish, L. Sun, Long non-coding RNAs as regulators of the endocrine system. Nat. Rev. Endocrinol. 11, 151–160 (2015)
K. Tano, N. Akimitsu, Long non-coding RNAs in cancer progression. Front. Genet. 3, 219 (2012)
A.G. Bader, D. Brown, J. Stoudemire, P. Lammers, Developing therapeutic microRNAs for cancer. Gene Ther. 18, 1121–1126 (2011)
J. Torrisani, B. Bournet, M.C. du Rieu, M. Bouisson, A. Souque, J. Escourrou, L. Buscail, P. Cordelier, let-7 MicroRNA transfer in pancreatic cancer-derived cells inhibits in vitro cell proliferation but fails to alter tumor progression. Hum.Gene Ther. 20, 831–844 (2009)
P.J. White, F. Anastasopoulos, C.W. Pouton, B.J. Boyd, Overcoming biological barriers to in vivo efficacy of antisense oligonucleotides. Expert Rev. Mol. Med. 11, e10 (2009)
X. Zong, L. Huang, V. Tripathi, R. Peralta, S.M. Freier, S. Guo, K.V. Prasanth, Knockdown of nuclear-retained long noncoding RNAs using modified DNA antisense oligonucleotides. Methods Mol. Biol. 1262, 321–331 (2015)
N. Hanna, P. Ohana, F.M. Konikoff, G. Leichtmann, A. Hubert, L. Appelbaum, Y. Kopelman, A. Czerniak, A. Hochberg, Phase 1/2a, dose-escalation, safety, pharmacokinetic and preliminary efficacy study of intratumoral administration of BC-819 in patients with unresectable pancreatic cancer. Cancer Gene Ther. 19, 374–381 (2012)
M. Lohr, P. Maisonneuve, A.B. Lowenfels, K-Ras mutations and benign pancreatic disease. Int. J. Pancreatol. 27, 93–103 (2000)
M. Wang, X. Lu, X. Dong, F. Hao, Z. Liu, G. Ni, D. Chen, pERK1/2 silencing sensitizes pancreatic cancer BXPC-3 cell to gemcitabine-induced apoptosis via regulating Bax and Bcl-2 expression. World J.Surg.Oncol. 13, 66 (2015). doi:10.1186/s12957-015-0451-7
O.A. Kent, R.R. Chivukula, M. Mullendore, E.A. Wentzel, G. Feldmann, K.H. Lee, S. Liu, S.D. Leach, A. Maitra, J.T. Mendell, Repression of the miR-143/145 cluster by oncogenic Ras initiates a tumor-promoting feed-forward pathway. Genes Dev. 24, 2754–2759 (2010)
I. Keklikoglou, K. Hosaka, C. Bender, A. Bott, C. Koerner, D. Mitra, R. Will, A. Woerner, E. Muenstermann, H. Wilhelm, Y. Cao, S. Wiemann, MicroRNA-206 functions as a pleiotropic modulator of cell proliferation, invasion and lymphangiogenesis in pancreatic adenocarcinoma by targeting ANXA2 and KRAS genes. Oncogene 34, 4867–4878 (2014)
X. Yan, X. Chen, H. Liang, T. Deng, W. Chen, S. Zhang, M. Liu, X. Gao, Y. Liu, C. Zhao, X. Wang, N. Wang, J. Li, R. Liu, K. Zen, C.Y. Zhang, B. Liu, Y. Ba, miR-143 and miR-145 synergistically regulate ERBB3 to suppress cell proliferation and invasion in breast cancer. Mol. Cancer. 13, 220 (2014). doi:10.1186/1476-4598-13-220
O.A. Kent, J.T. Mendell, R. Rottapel, Transcriptional regulation of miR-31 by oncogenic KRAS mediates metastatic phenotypes by repressing RASA1. Mol. Cancer. Res. (2016). doi:10.1158/1541-7786.MCR-15-0456
J. Su, H. Liang, W. Yao, N. Wang, S. Zhang, X. Yan, H. Feng, W. Pang, Y. Wang, X. Wang, Z. Fu, Y. Liu, C. Zhao, J. Zhang, C.Y. Zhang, K. Zen, X. Chen, Y. Wang, MiR-143 and MiR-145 regulate IGF1R to suppress cell proliferation in colorectal cancer. PLoS One 9, e114420 (2014)
X. Chen, X. Guo, H. Zhang, Y. Xiang, J. Chen, Y. Yin, X. Cai, K. Wang, G. Wang, Y. Ba, L. Zhu, J. Wang, R. Yang, Y. Zhang, Z. Ren, K. Zen, J. Zhang, C.Y. Zhang, Role of miR-143 targeting KRAS in colorectal tumorigenesis. Oncogene 28, 1385–1392 (2009)
C. Clape, V. Fritz, C. Henriquet, F. Apparailly, P.L. Fernandez, F. Iborra, C. Avances, M. Villalba, S. Culine, L. Fajas, miR-143 interferes with ERK5 signaling, and abrogates prostate cancer progression in mice. PLoS One 4, e7542 (2009)
M. Sachdeva, S. Zhu, F. Wu, H. Wu, V. Walia, S. Kumar, R. Elble, K. Watabe, Y.Y. Mo, p53 represses c-Myc through induction of the tumor suppressor miR-145. Proc. Natl. Acad. Sci. U. S. A. 106, 3207–3212 (2009)
P. Wang, C.F. Zhu, M.Z. Ma, G. Chen, M. Song, Z.L. Zeng, W.H. Lu, J. Yang, S. Wen, P.J. Chiao, Y. Hu, P. Huang, Micro-RNA-155 is induced by K-Ras oncogenic signal and promotes ROS stress in pancreatic cancer. Oncotarget 6, 21148–21158 (2015)
T. Greither, L.F. Grochola, A. Udelnow, C. Lautenschlager, P. Wurl, H. Taubert, Elevated expression of microRNAs 155, 203, 210 and 222 in pancreatic tumors is associated with poorer survival. Int. J. Cancer 126, 73–80 (2010)
M. Gironella, M. Seux, M.J. Xie, C. Cano, R. Tomasini, J. Gommeaux, S. Garcia, J. Nowak, M.L. Yeung, K.T. Jeang, A. Chaix, L. Fazli, Y. Motoo, Q. Wang, P. Rocchi, A. Russo, M. Gleave, J.C. Dagorn, J.L. Iovanna, A. Carrier, M.J. Pebusque, N.J. Dusetti, Tumor protein 53-induced nuclear protein 1 expression is repressed by miR-155, and its restoration inhibits pancreatic tumor development. Proc. Natl. Acad. Sci. U. S. A. 104, 16170–16175 (2007)
W. Pang, J. Su, Y. Wang, H. Feng, X. Dai, Y. Yuan, X. Chen, W. Yao, Pancreatic cancer-secreted miR-155 implicates in the Conversion from Normal Fibroblasts to Cancer-Associated Fibroblasts. Cancer. Sci. 106, 1362–1369 (2015)
C. Huang, H. Li, W. Wu, T. Jiang, Z. Qiu, Regulation of miR-155 affects pancreatic cancer cell invasiveness and migration by modulating the STAT3 signaling pathway through SOCS1. Oncol. Rep. 30, 1223–1230 (2013)
C. Huang, G. Yang, T. Jiang, G. Zhu, H. Li, Z. Qiu, The effects and mechanisms of blockage of STAT3 signaling pathway on IL-6 inducing EMT in human pancreatic cancer cells in vitro. Neoplasma 58, 396–405 (2011)
I.A. Asangani, S.A. Rasheed, D.A. Nikolova, J.H. Leupold, N.H. Colburn, S. Post, H. Allgayer, MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 27, 2128–2136 (2008)
J.A. Chan, A.M. Krichevsky, K.S. Kosik, MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res. 65, 6029–6033 (2005)
D.L. Vaux, S. Cory, J.M. Adams, Bcl-2 gene promotes haemopoietic cell survival and cooperates with c-myc to immortalize pre-B cells. Nature 335, 440–442 (1988)
M. Dillhoff, J. Liu, W. Frankel, C. Croce, M. Bloomston, MicroRNA-21 is overexpressed in pancreatic cancer and a potential predictor of survival. J. Gastrointest. Surg. 12, 2171–2176 (2008)
W.F. Song, L. Wang, W.Y. Huang, X. Cai, J.J. Cui, L.W. Wang, MiR-21 upregulation induced by promoter zone histone acetylation is associated with chemoresistance to gemcitabine and enhanced malignancy of pancreatic cancer cells. Asian Pac. J. Cancer Prev. 14, 7529–7536 (2013)
P. Wang, L. Zhuang, J. Zhang, J. Fan, J. Luo, H. Chen, K. Wang, L. Liu, Z. Chen, Z. Meng, The serum miR-21 level serves as a predictor for the chemosensitivity of advanced pancreatic cancer, and miR-21 expression confers chemoresistance by targeting FasL. Mol. Oncol. 7, 334–345 (2013)
J.H. Hwang, J. Voortman, E. Giovannetti, S.M. Steinberg, L.G. Leon, Y.T. Kim, N. Funel, J.K. Park, M.A. Kim, G.H. Kang, S.W. Kim, M. Del Chiaro, G.J. Peters, G. Giaccone, Identification of microRNA-21 as a biomarker for chemoresistance and clinical outcome following adjuvant therapy in resectable pancreatic cancer. PLoS One 5, e10630 (2010)
P. Liu, H. Liang, Q. Xia, P. Li, H. Kong, P. Lei, S. Wang, Z. Tu, Resveratrol induces apoptosis of pancreatic cancers cells by inhibiting miR-21 regulation of BCL-2 expression. Clin. Transl. Oncol. 15, 741–746 (2013)
J.K. Park, E.J. Lee, C. Esau, T.D. Schmittgen, Antisense inhibition of microRNA-21 or -221 arrests cell cycle, induces apoptosis, and sensitizes the effects of gemcitabine in pancreatic adenocarcinoma. Pancreas 38, e190–e199 (2009)
P.A. Toste, L. Li, B.E. Kadera, A.H. Nguyen, L.M. Tran, N. Wu, D.L. Madnick, S.G. Patel, D.W. Dawson, T.R. Donahue, p85alpha is a microRNA target and affects chemosensitivity in pancreatic cancer. J. Surg. Res. 196, 285–293 (2015)
W.H. Paik, H.R. Kim, J.K. Park, B.J. Song, S.H. Lee, J.H. Hwang, Chemosensitivity induced by down-regulation of microRNA-21 in gemcitabine-resistant pancreatic cancer cells by indole-3-carbinol. Anticancer Res. 33, 1473–1481 (2013)
T.A. Mace, A.L. Collins, S.E. Wojcik, C.M. Croce, G.B. Lesinski, M. Bloomston, Hypoxia induces the overexpression of microRNA-21 in pancreatic cancer cells. J. Surg. Res. 184, 855–860 (2013)
H. Ying, K.G. Elpek, A. Vinjamoori, S.M. Zimmerman, G.C. Chu, H. Yan, E. Fletcher-Sananikone, H. Zhang, Y. Liu, W. Wang, X. Ren, H. Zheng, A.C. Kimmelman, J.H. Paik, C. Lim, S.R. Perry, S. Jiang, B. Malinn, A. Protopopov, S. Colla, Y. Xiao, A.F. Hezel, N. Bardeesy, S.J. Turley, Y.A. Wang, L. Chin, S.P. Thayer, R.A. DePinho, PTEN is a major tumor suppressor in pancreatic ductal adenocarcinoma and regulates an NF-kappaB-cytokine network. Cancer Discov. 1, 158–169 (2011)
B.E. Kadera, L. Li, P.A. Toste, N. Wu, C. Adams, D.W. Dawson, T.R. Donahue, MicroRNA-21 in pancreatic ductal adenocarcinoma tumor-associated fibroblasts promotes metastasis. PLoS One 8, e71978 (2013)
F. Sicard, M. Gayral, H. Lulka, L. Buscail, P. Cordelier, Targeting miR-21 for the therapy of pancreatic cancer. Mol. Ther. 21, 986–994 (2013)
M. Passadouro, M.C. Pedroso de Lima, H. Faneca, MicroRNA modulation combined with sunitinib as a novel therapeutic strategy for pancreatic cancer. Int. J. Nanomedicine 9, 3203–3217 (2014)
Y. Zhang, M. Li, H. Wang, W.E. Fisher, P.H. Lin, Q. Yao, C. Chen, Profiling of 95 microRNAs in pancreatic cancer cell lines and surgical specimens by real-time PCR analysis. World J. Surg. 33, 698–709 (2009)
H. Hermeking, The miR-34 family in cancer and apoptosis. Cell Death Differ. 17, 193–199 (2010)
J. Xia, Q. Duan, A. Ahmad, B. Bao, S. Banerjee, Y. Shi, J. Ma, J. Geng, Z. Chen, K. Rahman, L. Miele, F. Sarkar, Z. Wang, Genistein inhibits cell growth and induces apoptosis through up-regulation of miR-34a in pancreatic cancer cells. 13, 1750–1756 (2012)
Q. Ji, X. Hao, M. Zhang, W. Tang, M. Yang, L. Li, D. Xiang, J.T. Desano, G.T. Bommer, D. Fan, E.R. Fearon, T.S. Lawrence, L. Xu, MicroRNA miR-34 inhibits human pancreatic cancer tumor-initiating cells. PLoS One 4, e6816 (2009)
C. Liu, H. Cheng, S. Shi, X. Cui, J. Yang, L. Chen, P. Cen, X. Cai, Y. Lu, C. Wu, W. Yao, Y. Qin, L. Liu, J. Long, J. Xu, M. Li, X. Yu, MicroRNA-34b inhibits pancreatic cancer metastasis through repressing Smad3. Curr. Mol. Med. 13, 467–478 (2013)
T.C. Chang, E.A. Wentzel, O.A. Kent, K. Ramachandran, M. Mullendore, K.H. Lee, G. Feldmann, M. Yamakuchi, M. Ferlito, C.J. Lowenstein, D.E. Arking, M.A. Beer, A. Maitra, J.T. Mendell, Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol. Cell 26, 745–752 (2007)
A.K. Bubna, Vorinostat-An overview. Indian J. Dermatol. 60, 419 (2015). doi:10.4103/0019-5154.160511
D.A. Deming, J. Ninan, H.H. Bailey, J.M. Kolesar, J. Eickhoff, J.M. Reid, M.M. Ames, R.M. McGovern, D. Alberti, R. Marnocha, I. Espinoza-Delgado, J. Wright, G. Wilding, W.R. Schelman, A Phase I study of intermittently dosed vorinostat in combination with bortezomib in patients with advanced solid tumors. Invest. New Drugs 32, 323–329 (2014)
J.H. Mitchell, E. Cawood, D. Kinniburgh, A. Provan, A.R. Collins, D.S. Irvine, Effect of a phytoestrogen food supplement on reproductive health in normal males. Clin. Sci. (Lond.) 100, 613–618 (2001)
S. Babashah, M. Sadeghizadeh, M.R. Tavirani, S. Farivar, M. Soleimani, Aberrant microRNA expression and its implications in the pathogenesis of leukemias. Cell. Oncol. 35, 317–334 (2012)
J. Haybaeck, N. Zeller, M. Heikenwalder, The parallel universe: microRNAs and their role in chronic hepatitis, liver tissue damage and hepatocarcinogenesis. Swiss Med. Wkly. 141, w13287 (2011)
ClinicalTrials.gov [Internet] Identifier: NCT01829971, A multicenter phase I study of MRX34, MicroRNA miR-RX34 liposomal injection, 2016 (2015)
R. Zhang, M. Li, W. Zang, X. Chen, Y. Wang, P. Li, Y. Du, G. Zhao, L. Li, MiR-148a regulates the growth and apoptosis in pancreatic cancer by targeting CCKBR and Bcl-2. Tumour Biol. 35, 837–844 (2014)
S.K. Srivastava, A. Bhardwaj, S. Arora, N. Tyagi, S. Singh, J. Andrews, S. McClellan, B. Wang, A.P. Singh, MicroRNA-345 induces apoptosis in pancreatic cancer cells through potentiation of caspase-dependent and -independent pathways. Br. J. Cancer 113, 660–668 (2015)
J. Hao, S. Zhang, Y. Zhou, X. Hu, C. Shao, MicroRNA 483-3p suppresses the expression of DPC4/Smad4 in pancreatic cancer. FEBS Lett. 585, 207–213 (2011)
P. Singh, J.D. Wig, R. Srinivasan, B.D. Radotra, A comprehensive examination of Smad4, Smad6 and Smad7 mRNA expression in pancreatic ductal adenocarcinoma. Indian J. Cancer 48, 170–174 (2011)
Z. Zhu, Y. Xu, J. Zhao, Q. Liu, W. Feng, J. Fan, P. Wang, miR-367 promotes epithelial-to-mesenchymal transition and invasion of pancreatic ductal adenocarcinoma cells by targeting the Smad7-TGF-beta signalling pathway. Br. J. Cancer 112, 1367–1375 (2015)
X. Zhi, J. Tao, K. Xie, Y. Zhu, Z. Li, J. Tang, W. Wang, H. Xu, J. Zhang, Z. Xu, MUC4-induced nuclear translocation of beta-catenin: a novel mechanism for growth, metastasis and angiogenesis in pancreatic cancer. Cancer Lett. 346, 104–113 (2014)
Y. Zhu, J.J. Zhang, W.B. Liang, R. Zhu, B. Wang, Y. Miao, Z.K. Xu, Pancreatic cancer counterattack: MUC4 mediates Fas-independent apoptosis of antigen-specific cytotoxic T lymphocyte. Oncol. Rep. 31, 1768–1776 (2014)
D. Ansari, C. Urey, K.S. Hilmersson, M.P. Bauden, F. Ek, R. Olsson, R. Andersson, Apicidin sensitizes pancreatic cancer cells to gemcitabine by epigenetically regulating MUC4 expression. Anticancer Res. 34, 5269–5276 (2014)
S.K. Srivastava, A. Bhardwaj, S. Singh, S. Arora, B. Wang, W.E. Grizzle, A.P. Singh, MicroRNA-150 directly targets MUC4 and suppresses growth and malignant behavior of pancreatic cancer cells. Carcinogenesis 32, 1832–1839 (2011)
P. Chaturvedi, A.P. Singh, S. Chakraborty, S.C. Chauhan, S. Bafna, J.L. Meza, P.K. Singh, M.A. Hollingsworth, P.P. Mehta, S.K. Batra, MUC4 mucin interacts with and stabilizes the HER2 oncoprotein in human pancreatic cancer cells. Cancer Res. 68, 2065–2070 (2008)
F. Lahdaoui, Y. Delpu, A. Vincent, F. Renaud, M. Messager, B. Duchene, E. Leteurtre, C. Mariette, J. Torrisani, N. Jonckheere, I. Van, Seuningen, miR-219-1-3p is a negative regulator of the mucin MUC4 expression and is a tumor suppressor in pancreatic cancer. Oncogene 34, 780–788 (2015)
D. Chen, Y. Zhang, J. Wang, J. Chen, C. Yang, K. Cai, X. Wang, F. Shi, J. Dou, MicroRNA-200c overexpression inhibits tumorigenicity and metastasis of CD117+CD44+ ovarian cancer stem cells by regulating epithelial-mesenchymal transition. J. Ovarian Res. 6, 50 (2013). doi:10.1186/1757-2215-6-50
J. Dou, X.F. He, W.H. Cao, F.S. Zhao, X.Y. Wang, Y.R. Liu, J. Wang, Overexpression of microRna-200c in CD44+CD133+ CSCS inhibits the cellular migratory and invasion as well as tumorigenicity in mice. Cell. Mol. Biol. Suppl 59, OL1861-8 (2013)
F.F. Ibrahim, R. Jamal, S.E. Syafruddin, N.S. Ab Mutalib, S. Saidin, R.R. MdZin, M.M. Hossain Mollah, N.M. Mokhtar, MicroRNA-200c and microRNA-31 regulate proliferation, colony formation, migration and invasion in serous ovarian cancer. J. Ovarian Res. 8, 56 (2015). doi:10.1186/s13048-015-0186-7
J. Lu, G. Getz, E.A. Miska, E. Alvarez-Saavedra, J. Lamb, D. Peck, A. Sweet-Cordero, B.L. Ebert, R.H. Mak, A.A. Ferrando, J.R. Downing, T. Jacks, H.R. Horvitz, T.R. Golub, MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005)
D.M. Dykxhoorn, MicroRNAs and metastasis: little RNAs go a long way. Cancer Res. 70, 6401–6406 (2010)
Y. Lu, J. Lu, X. Li, H. Zhu, X. Fan, S. Zhu, Y. Wang, Q. Guo, L. Wang, Y. Huang, M. Zhu, Z. Wang, MiR-200a inhibits epithelial-mesenchymal transition of pancreatic cancer stem cell. BMC Cancer 14, 85 (2014). doi:10.1186/1471-2407-14-85
C. Ma, T. Huang, Y.C. Ding, W. Yu, Q. Wang, B. Meng, S.X. Luo, microRNA-200c overexpression inhibits chemoresistance, invasion and colony formation of human pancreatic cancer stem cells. Int. J. Clin. Exp. Pathol. 8, 6533–6539 (2015)
Y. Imanaka, S. Tsuchiya, F. Sato, Y. Shimada, K. Shimizu, G. Tsujimoto, MicroRNA-141 confers resistance to cisplatin-induced apoptosis by targeting YAP1 in human esophageal squamous cell carcinoma. J. Hum. Genet. 56, 270–276 (2011)
L. Xu, Q. Li, D. Xu, Q. Wang, Y. An, Q. Du, J. Zhang, Y. Zhu, Y. Miao, hsa-miR-141 downregulates TM4SF1 to inhibit pancreatic cancer cell invasion and migration. Int. J. Oncol. 44, 459–466 (2014)
G. Zhao, B. Wang, Y. Liu, J.G. Zhang, S.C. Deng, Q. Qin, K. Tian, X. Li, S. Zhu, Y. Niu, Q. Gong, C.Y. Wang, miRNA-141, downregulated in pancreatic cancer, inhibits cell proliferation and invasion by directly targeting MAP4K4. Mol. Cancer. Ther. 12, 2569–2580 (2013)
Z.M. Zhu, Y.F. Xu, Q.J. Su, J.D. Du, X.L. Tan, Y.L. Tu, J.W. Tan, H.B. Jiao, Prognostic significance of microRNA-141 expression and its tumor suppressor function in human pancreatic ductal adenocarcinoma. Mol. Cell. Biochem. 388, 39–49 (2014)
T. Avnit-Sagi, L. Kantorovich, S. Kredo-Russo, E. Hornstein, M.D. Walker, The promoter of the pri-miR-375 gene directs expression selectively to the endocrine pancreas. PLoS One 4, e5033 (2009)
M.N. Poy, L. Eliasson, J. Krutzfeldt, S. Kuwajima, X. Ma, P.E. Macdonald, S. Pfeffer, T. Tuschl, N. Rajewsky, P. Rorsman, M. Stoffel, A pancreatic islet-specific microRNA regulates insulin secretion. Nature 432, 226–230 (2004)
A. Basu, H. Alder, A. Khiyami, P. Leahy, C.M. Croce, S. Haldar, MicroRNA-375 and MicroRNA-221: potential noncoding RNAs associated with antiproliferative activity of benzyl isothiocyanate in pancreatic cancer. Genes Cancer 2, 108–119 (2011)
J. Zhou, S. Song, S. He, X. Zhu, Y. Zhang, B. Yi, B. Zhang, G. Qin, D. Li, MicroRNA-375 targets PDK1 in pancreatic carcinoma and suppresses cell growth through the Akt signaling pathway. Int. J. Mol. Med. 33, 950–956 (2014)
J. Zhou, S. Song, J. Cen, D. Zhu, D. Li, Z. Zhang, MicroRNA-375 is downregulated in pancreatic cancer and inhibits cell proliferation in vitro. Oncol. Res. 20, 197–203 (2012)
S.D. Song, J. Zhou, J. Zhou, H. Zhao, J.N. Cen, D.C. Li, MicroRNA-375 targets the 3-phosphoinositide-dependent protein kinase-1 gene in pancreatic carcinoma. Oncol. Lett. 6, 953–959 (2013)
J. Li, Y. Wang, W. Yu, J. Chen, J. Luo, Expression of serum miR-221 in human hepatocellular carcinoma and its prognostic significance. Biochem. Biophys. Res. Commun. 406, 70–73 (2011)
M. Bloomston, W.L. Frankel, F. Petrocca, S. Volinia, H. Alder, J.P. Hagan, C.G. Liu, D. Bhatt, C. Taccioli, C.M. Croce, MicroRNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA 297, 1901–1908 (2007)
Q. Xu, P. Li, X. Chen, L. Zong, Z. Jiang, L. Nan, J. Lei, W. Duan, D. Zhang, X. Li, H. Sha, Z. Wu, Q. Ma, Z. Wang, miR-221/222 induces pancreatic cancer progression through the regulation of matrix metalloproteinases. Oncotarget 6, 14153–14164 (2015)
S. Sarkar, H. Dubaybo, S. Ali, P. Goncalves, S.L. Kollepara, S. Sethi, P.A. Philip, Y. Li, Down-regulation of miR-221 inhibits proliferation of pancreatic cancer cells through up-regulation of PTEN, p27(kip1), p57(kip2), and PUMA. Am. J. Cancer. Res. 3, 465–477 (2013)
Y. Li, T.G. VandenBoom 2nd, D. Kong, Z. Wang, S. Ali, P.A. Philip, F.H. Sarkar, Up-regulation of miR-200 and let-7 by natural agents leads to the reversal of epithelial-to-mesenchymal transition in gemcitabine-resistant pancreatic cancer cells. Cancer Res. 69, 6704–6712 (2009)
K. Patel, A. Kollory, A. Takashima, S. Sarkar, D.V. Faller, S.K. Ghosh, MicroRNA let-7 downregulates STAT3 phosphorylation in pancreatic cancer cells by increasing SOCS3 expression. Cancer Lett. 347, 54–64 (2014)
Y.D. Bhutia, S.W. Hung, M. Krentz, D. Patel, D. Lovin, R. Manoharan, J.M. Thomson, R. Govindarajan, Differential processing of let-7a precursors influences RRM2 expression and chemosensitivity in pancreatic cancer: role of LIN-28 and SET oncoprotein. PLoS One 8, e53436 (2013)
S. Watanabe, Y. Ueda, S. Akaboshi, Y. Hino, Y. Sekita, M. Nakao, HMGA2 maintains oncogenic RAS-induced epithelial-mesenchymal transition in human pancreatic cancer cells. Am. J. Pathol. 174, 854–868 (2009)
A. Druz, Y.C. Chen, R. Guha, M. Betenbaugh, S.E. Martin, J. Shiloach, Large-scale screening identifies a novel microRNA, miR-15a-3p, which induces apoptosis in human cancer cell lines. RNA Biol. 10, 287–300 (2013)
S. Guo, X. Xu, Y. Tang, C. Zhang, J. Li, Y. Ouyang, J. Ju, P. Bie, H. Wang, miR-15a inhibits cell proliferation and epithelial to mesenchymal transition in pancreatic ductal adenocarcinoma by down-regulating Bmi-1 expression. Cancer Lett. 344, 40–46 (2014)
F. Wang, X. Xue, J. Wei, Y. An, J. Yao, H. Cai, J. Wu, C. Dai, Z. Qian, Z. Xu, Y. Miao, hsa-miR-520h downregulates ABCG2 in pancreatic cancer cells to inhibit migration, invasion, and side populations. Br. J. Cancer 103, 567–574 (2010)
J. Jiang, C. Yu, M. Chen, H. Zhang, S. Tian, C. Sun, Reduction of miR-29c enhances pancreatic cancer cell migration and stem cell-like phenotype. Oncotarget 6, 2767–2778 (2015)
M.K. Muniyappa, P. Dowling, M. Henry, P. Meleady, P. Doolan, P. Gammell, M. Clynes, N. Barron, MiRNA-29a regulates the expression of numerous proteins and reduces the invasiveness and proliferation of human carcinoma cell lines. Eur. J. Cancer 45, 3104–3118 (2009)
S. Yu, Z. Lu, C. Liu, Y. Meng, Y. Ma, W. Zhao, J. Liu, J. Yu, J. Chen, miRNA-96 suppresses KRAS and functions as a tumor suppressor gene in pancreatic cancer. Cancer Res. 70, 6015–6025 (2010)
J. Feng, J. Yu, X. Pan, Z. Li, Z. Chen, W. Zhang, B. Wang, L. Yang, H. Xu, G. Zhang, Z. Xu, HERG1 functions as an oncogene in pancreatic cancer and is downregulated by miR-96. Oncotarget 5, 5832–5844 (2014)
C. Li, X. Du, S. Tai, X. Zhong, Z. Wang, Z. Hu, L. Zhang, P. Kang, D. Ji, X. Jiang, Q. Zhou, M. Wan, G. Jiang, Y. Cui, GPC1 regulated by miR-96-5p, rather than miR-182-5p, in inhibition of pancreatic carcinoma cell proliferation. Int. J. Mol. Sci. 15, 6314–6327 (2014)
X. Huang, W. Lv, J.H. Zhang, D.L. Lu, miR96 functions as a tumor suppressor gene by targeting NUAK1 in pancreatic cancer. Int. J. Mol. Med. 34, 1599–1605 (2014)
D. Li, X. Li, W. Cao, Y. Qi, X. Yang, Antagonism of microRNA-99a promotes cell invasion and down-regulates E-cadherin expression in pancreatic cancer cells by regulating mammalian target of rapamycin. Acta Histochem. 116, 723–729 (2014)
Z. Li, X. Li, C. Yu, M. Wang, F. Peng, J. Xiao, R. Tian, J. Jiang, C. Sun, MicroRNA-100 regulates pancreatic cancer cells growth and sensitivity to chemotherapy through targeting FGFR3. Tumour Biol. 35, 11751–11759 (2014)
W. Jiang, W. Gu, R. Qiu, C. Shen, E.Y. YaohaoWu, J. Zhang, J. Zhou, Y. Guo, Z. Li, J. Deng, L. Zeng, J. Tang, Q. Zhi, X. Deng, miRNA-101 suppresses epithelial-to-mesenchymal transition by targeting HMGA2 in pancreatic cancer cells. Anticancer Agents Med. Chem. (2015). doi:10.2174/1871520615666150507122142
A.M. Qazi, O. Gruzdyn, A. Semaan, S. Seward, S. Chamala, V. Dhulipala, S. Sethi, R. Ali-Fehmi, P.A. Philip, D.L. Bouwman, D.W. Weaver, S.A. Gruber, R.B. Batchu, Restoration of E-cadherin expression in pancreatic ductal adenocarcinoma treated with microRNA-101. Surgery 152, 704–711 (2012)
K.H. Lee, C. Lotterman, C. Karikari, N. Omura, G. Feldmann, N. Habbe, M.G. Goggins, J.T. Mendell, A. Maitra, Epigenetic silencing of MicroRNA miR-107 regulates cyclin-dependent kinase 6 expression in pancreatic cancer. Pancreatology 9, 293–301 (2009)
P. Wang, L. Chen, J. Zhang, H. Chen, J. Fan, K. Wang, J. Luo, Z. Chen, Z. Meng, L. Liu, Methylation-mediated silencing of the miR-124 genes facilitates pancreatic cancer progression and metastasis by targeting Rac1. Oncogene 33, 514–524 (2014)
S. Hamada, K. Satoh, W. Fujibuchi, M. Hirota, A. Kanno, J. Unno, A. Masamune, K. Kikuta, K. Kume, T. Shimosegawa, MiR-126 acts as a tumor suppressor in pancreatic cancer cells via the regulation of ADAM9. Mol. Cancer Res. 10, 3–10 (2012)
G. Zhao, J.G. Zhang, Y. Shi, Q. Qin, Y. Liu, B. Wang, K. Tian, S.C. Deng, X. Li, S. Zhu, Q. Gong, Y. Niu, C.Y. Wang, MiR-130b is a prognostic marker and inhibits cell proliferation and invasion in pancreatic cancer through targeting STAT3. PLoS One 8, e73803 (2013)
S. Zhang, J. Hao, F. Xie, X. Hu, C. Liu, J. Tong, J. Zhou, J. Wu, C. Shao, Downregulation of miR-132 by promoter methylation contributes to pancreatic cancer development. Carcinogenesis 32, 1183–1189 (2011)
Z. Dang, W.H. Xu, P. Lu, N. Wu, J. Liu, B. Ruan, L. Zhou, W.J. Song, K.F. Dou, MicroRNA-135a inhibits cell proliferation by targeting Bmi1 in pancreatic ductal adenocarcinoma. Int. J. Biol. Sci. 10, 733–745 (2014)
C. Yu, M. Wang, M. Chen, Y. Huang, J. Jiang, Upregulation of microRNA1385p inhibits pancreatic cancer cell migration and increases chemotherapy sensitivity. Mol. Med. Rep. 12, 5135–5140 (2015)
S. Liang, X. Gong, G. Zhang, G. Huang, Y. Lu, Y. Li, MicroRNA-140 regulates cell growth and invasion in pancreatic duct adenocarcinoma by targeting iASPP. Acta Biochim. Biophys. Sin. (Shanghai) 48, 174–181 (2016)
T.N. MacKenzie, N. Mujumdar, S. Banerjee, V. Sangwan, A. Sarver, S. Vickers, S. Subramanian, A.K. Saluja, Triptolide induces the expression of miR-142-3p: a negative regulator of heat shock protein 70 and pancreatic cancer cell proliferation. Mol. Cancer Ther. 12, 1266–1275 (2013)
Y. Hu, Y. Ou, K. Wu, Y. Chen, W. Sun, miR-143 inhibits the metastasis of pancreatic cancer and an associated signaling pathway. Tumour Biol. 33, 1863–1870 (2012)
H. Pham, C.E. Rodriguez, G.W. Donald, K.M. Hertzer, X.S. Jung, H.H. Chang, A. Moro, H.A. Reber, O.J. Hines, G. Eibl, miR-143 decreases COX-2 mRNA stability and expression in pancreatic cancer cells. Biochem. Biophys. Res. Commun. 439, 6–11 (2013)
T. Han, X.P. Yi, B. Liu, M.J. Ke, Y.X. Li, MicroRNA-145 suppresses cell proliferation, invasion and migration in pancreatic cancer cells by targeting NEDD9. Mol. Med. Rep. 11, 4115–4120 (2015)
S. Khan, M.C. Ebeling, M.S. Zaman, M. Sikander, M.M. Yallapu, N. Chauhan, A.M. Yacoubian, S.W. Behrman, N. Zafar, D. Kumar, P.A. Thompson, M. Jaggi, S.C. Chauhan, MicroRNA-145 targets MUC13 and suppresses growth and invasion of pancreatic cancer. Oncotarget 5, 7599–7609 (2014)
S. Ali, A. Ahmad, A. Aboukameel, A. Ahmed, B. Bao, S. Banerjee, P.A. Philip, F.H. Sarkar, Deregulation of miR-146a expression in a mouse model of pancreatic cancer affecting EGFR signaling. Cancer Lett. 351, 134–142 (2014)
Y. Li, T.G. VandenBoom 2nd, Z. Wang, D. Kong, S. Ali, P.A. Philip, F.H. Sarkar, Up-regulation of miR-146a contributes to the inhibition of invasion of pancreatic cancer cells. Cancer Res. 70, 5703 (2010)
F. Lin, X. Wang, Z. Jie, X. Hong, X. Li, M. Wang, Y. Yu, Inhibitory effects of miR-146b-5p on cell migration and invasion of pancreatic cancer by targeting MMP16. J. Huazhong Univ. Sci. Technolog. Med. Sci. 31, 509–514 (2011)
M. Azizi, L. Teimoori-Toolabi, M.K. Arzanani, K. Azadmanesh, P. Fard-Esfahani, S. Zeinali, MicroRNA-148b and microRNA-152 reactivate tumor suppressor genes through suppression of DNA methyltransferase-1 gene in pancreatic cancer cell lines. Cancer Biol. Ther. 15, 419–427 (2014)
X. Bofill-De Ros, M. Gironella, C. Fillat, miR-148a- and miR-216a-regulated oncolytic adenoviruses targeting pancreatic tumors attenuate tissue damage without perturbation of miRNA activity. Mol. Ther. 22, 1665–1677 (2014)
S.T. Liffers, J.B. Munding, M. Vogt, J.D. Kuhlmann, B. Verdoodt, S. Nambiar, A. Maghnouj, A. Mirmohammadsadegh, S.A. Hahn, A. Tannapfel, MicroRNA-148a is down-regulated in human pancreatic ductal adenocarcinomas and regulates cell survival by targeting CDC25B. Lab. Invest. 91, 1472–1479 (2011)
G. Zhao, J.G. Zhang, Y. Liu, Q. Qin, B. Wang, K. Tian, L. Liu, X. Li, Y. Niu, S.C. Deng, C.Y. Wang, miR-148b functions as a tumor suppressor in pancreatic cancer by targeting AMPKalpha1. Mol. Cancer. Ther. 12, 83–93 (2013)
L. Farhana, M.I. Dawson, F. Murshed, J.K. Das, A.K. Rishi, J.A. Fontana, Upregulation of miR-150* and miR-630 induces apoptosis in pancreatic cancer cells by targeting IGF-1R. PLoS One 8, e61015 (2013)
Y. Sun, X.L. Jin, T.T. Zhang, C.W. Jia, J. Chen, MiR-150-5p inhibits the proliferation and promoted apoptosis of pancreatic cancer cells. Zhonghua Bing Li Xue Za Zhi 42, 460–464 (2013)
L. Zhou, W.G. Zhang, D.S. Wang, K.S. Tao, W.J. Song, K.F. Dou, MicroRNA-183 is involved in cell proliferation, survival and poor prognosis in pancreatic ductal adenocarcinoma by regulating Bmi-1. Oncol. Rep. 32, 1734–1740 (2014)
H. Liu, X.F. Xu, Y. Zhao, M.C. Tang, Y.Q. Zhou, J. Lu, F.H. Gao, MicroRNA-191 promotes pancreatic cancer progression by targeting USP10. Tumour Biol. 35, 12157–12163 (2014)
J. Li, F. Kong, K. Wu, K. Song, J. He, W. Sun, miR-193b directly targets STMN1 and uPA genes and suppresses tumor growth and metastasis in pancreatic cancer. Mol. Med. Rep. 10, 2613–2620 (2014)
C. Marin-Muller, D. Li, U. Bharadwaj, M. Li, C. Chen, S.E. Hodges, W.E. Fisher, Q. Mo, M.C. Hung, Q. Yao, A tumorigenic factor interactome connected through tumor suppressor microRNA-198 in human pancreatic cancer. Clin. Cancer Res. 19, 5901–5913 (2013)
P. Radhakrishnan, A.M. Mohr, P.M. Grandgenett, M.M. Steele, S.K. Batra, M.A. Hollingsworth, MicroRNA-200c modulates the expression of MUC4 and MUC16 by directly targeting their coding sequences in human pancreatic cancer. PLoS One 8, e73356 (2013)
J. Yu, K. Ohuchida, K. Mizumoto, N. Sato, T. Kayashima, H. Fujita, K. Nakata, M. Tanaka, MicroRNA, hsa-miR-200c, is an independent prognostic factor in pancreatic cancer and its upregulation inhibits pancreatic cancer invasion but increases cell proliferation. Mol. Cancer. 9, 169 (2010). doi:10.1186/1476-4598-9-169
O. Soubani, A.S. Ali, F. Logna, S. Ali, P.A. Philip, F.H. Sarkar, Re-expression of miR-200 by novel approaches regulates the expression of PTEN and MT1-MMP in pancreatic cancer. Carcinogenesis 33, 1563–1571 (2012)
L. Miao, X. Xiong, Y. Lin, Y. Cheng, J. Lu, J. Zhang, N. Cheng, miR-203 inhibits tumor cell migration and invasion via caveolin-1 in pancreatic cancer cells. Oncol. Lett. 7, 658–662 (2014)
D. Xu, Q. Wang, Y. An, L. Xu, MiR203 regulates the proliferation, apoptosis and cell cycle progression of pancreatic cancer cells by targeting survivin. Mol. Med. Rep. 8, 379–384 (2013)
N. Ikenaga, K. Ohuchida, K. Mizumoto, J. Yu, T. Kayashima, H. Sakai, H. Fujita, K. Nakata, M. Tanaka, MicroRNA-203 expression as a new prognostic marker of pancreatic adenocarcinoma. Ann. Surg. Oncol. 17, 3120–3128 (2010)
A. Mittal, D. Chitkara, S.W. Behrman, R.I. Mahato, Efficacy of gemcitabine conjugated and miRNA-205 complexed micelles for treatment of advanced pancreatic cancer. Biomaterials 35, 7077–7087 (2014)
C. Stahlhut, Y. Suarez, J. Lu, Y. Mishima, A.J. Giraldez, miR-1 and miR-206 regulate angiogenesis by modulating VegfA expression in zebrafish. Development 139, 4356–4364 (2012)
I. Keklikoglou, K. Hosaka, C. Bender, A. Bott, C. Koerner, D. Mitra, R. Will, A. Woerner, E. Muenstermann, H. Wilhelm, Y. Cao, S. Wiemann, MicroRNA-206 functions as a pleiotropic modulator of cell proliferation, invasion and lymphangiogenesis in pancreatic adenocarcinoma by targeting ANXA2 and KRAS genes. Oncogene 34, 4867–4878 (2015)
M. Maftouh, A. Avan, N. Funel, A.E. Frampton, H. Fiuji, S. Pelliccioni, L. Castellano, V. Galla, G.J. Peters, E. Giovannetti, miR-211 modulates gemcitabine activity through downregulation of ribonucleotide reductase and inhibits the invasive behavior of pancreatic cancer cells. Nucleosides Nucleotides Nucleic Acids 33, 384–393 (2014)
S. Wang, X. Chen, M. Tang, MicroRNA-216a inhibits pancreatic cancer by directly targeting Janus kinase 2. Oncol. Rep. 32, 2824–2830 (2014)
X. Zhang, H. Shi, S. Lin, M. Ba, S. Cui, MicroRNA-216a enhances the radiosensitivity of pancreatic cancer cells by inhibiting beclin-1-mediated autophagy. Oncol. Rep. 34, 1557–1564 (2015)
H. He, S.J. Hao, L. Yao, F. Yang, Y. Di, J. Li, Y.J. Jiang, C. Jin, D.L. Fu, MicroRNA-218 inhibits cell invasion and migration of pancreatic cancer via regulating ROBO1. Cancer Biol. Ther. 15, 1333–1339 (2014)
C.H. Li, K.F. To, J.H. Tong, Z. Xiao, T. Xia, P.B. Lai, S.C. Chow, Y.X. Zhu, S.L. Chan, V.E. Marquez, Y. Chen, Enhancer of zeste homolog 2 silences microRNA-218 in human pancreatic ductal adenocarcinoma cells by inducing formation of heterochromatin. Gastroenterology 144, 1086–1097.e9 (2013)
Z.L. Zhang, Z.H. Bai, X.B. Wang, L. Bai, F. Miao, H.H. Pei, miR-186 and 326 predict the prognosis of pancreatic ductal adenocarcinoma and affect the proliferation and migration of cancer cells. PLoS One 10, e0118814 (2015)
L. Gao, Y. Yang, H. Xu, R. Liu, D. Li, H. Hong, M. Qin, Y. Wang, MiR-335 functions as a tumor suppressor in pancreatic cancer by targeting OCT4. Tumour Biol. 35, 8309–8318 (2014)
R. Guo, J. Gu, Z. Zhang, Y. Wang, C. Gu, MicroRNA-410 functions as a tumor suppressor by targeting angiotensin II type 1 receptor in pancreatic cancer. IUBMB Life 67, 42–53 (2015)
R. Guo, Y. Wang, W.Y. Shi, B. Liu, S.Q. Hou, L. Liu, MicroRNA miR-491-5p targeting both TP53 and Bcl-XL induces cell apoptosis in SW1990 pancreatic cancer cells through mitochondria mediated pathway. Molecules 17, 14733–14747 (2012)
Y. Liu, X. Li, S. Zhu, J.G. Zhang, M. Yang, Q. Qin, S.C. Deng, B. Wang, K. Tian, L. Liu, Y. Niu, C.Y. Wang, G. Zhao, Ectopic expression of miR-494 inhibited the proliferation, invasion and chemoresistance of pancreatic cancer by regulating SIRT1 and c-Myc. Gene Ther. 22, 729–738 (2015)
L. Li, Z. Li, X. Kong, D. Xie, Z. Jia, W. Jiang, J. Cui, Y. Du, D. Wei, S. Huang, K. Xie, Down-regulation of MicroRNA 494 via loss of SMAD4 increases FOXM1 and beta-catenin signaling in pancreatic ductal adenocarcinoma cells. Gastroenterology 147, 485–497 (2014)
J.W. Xu, T.X. Wang, L. You, L.F. Zheng, H. Shu, T.P. Zhang, Y.P. Zhao, Insulin-like growth factor 1 receptor (IGF-1R) as a target of MiR-497 and plasma IGF-1R levels associated with TNM stage of pancreatic cancer. PLoS One 9, e92847 (2014)
J. Xu, T. Wang, Z. Cao, H. Huang, J. Li, W. Liu, S. Liu, L. You, L. Zhou, T. Zhang, Y. Zhao, MiR-497 downregulation contributes to the malignancy of pancreatic cancer and associates with a poor prognosis. Oncotarget 5, 6983–6993 (2014)
J. Du, X. Zheng, S. Cai, Z. Zhu, J. Tan, B. Hu, Z. Huang, H. Jiao, MicroRNA506 participates in pancreatic cancer pathogenesis by targeting PIM3. Mol. Med. Rep. 12, 5121–5126 (2015)
B. Song, W. Ji, S. Guo, A. Liu, W. Jing, C. Shao, G. Li, G. Jin, miR-545 inhibited pancreatic ductal adenocarcinoma growth by targeting RIG-I. FEBS Lett. 588, 4375–4381 (2014)
H. Heyn, S. Schreek, R. Buurman, T. Focken, B. Schlegelberger, C. Beger, MicroRNA miR-548d is a superior regulator in pancreatic cancer. Pancreas 41, 218–221 (2012)
Y. Sun, T. Zhang, C. Wang, X. Jin, C. Jia, S. Yu, J. Chen, MiRNA-615-5p Functions as a Tumor Suppressor in Pancreatic Ductal Adenocarcinoma by Targeting AKT2. PLoS One 10, e0119783 (2015)
Y. Harazono, T. Muramatsu, H. Endo, N. Uzawa, T. Kawano, K. Harada, J. Inazawa, K. Kozaki, miR-655 Is an EMT-suppressive microRNA targeting ZEB1 and TGFBR2. PLoS One 8, e62757 (2013)
S. Muller, S. Raulefs, P. Bruns, F. Afonso-Grunz, A. Plotner, R. Thermann, C. Jager, A.M. Schlitter, B. Kong, I. Regel, W.K. Roth, B. Rotter, K. Hoffmeier, G. Kahl, I. Koch, F.J. Theis, J. Kleeff, P. Winter, C.W. Michalski, Next-generation sequencing reveals novel differentially regulated mRNAs, lncRNAs, miRNAs, sdRNAs and a piRNA in pancreatic cancer. Mol. Cancer. 14, 94 (2015). doi:10.1186/s12943-015-0358-5
J. Jiang, Z. Li, C. Yu, M. Chen, S. Tian, C. Sun, MiR-1181 inhibits stem cell-like phenotypes and suppresses SOX2 and STAT3 in human pancreatic cancer. Cancer Lett. 356, 962–970 (2015)
S. Shi, Y. Lu, Y. Qin, W. Li, H. Cheng, Y. Xu, J. Xu, J. Long, L. Liu, C. Liu, X. Yu, miR-1247 is correlated with prognosis of pancreatic cancer and inhibits cell proliferation by targeting neuropilins. Curr. Mol. Med. 14, 316–327 (2014)
K. Ohuchida, K. Mizumoto, C. Lin, H. Yamaguchi, T. Ohtsuka, N. Sato, H. Toma, M. Nakamura, E. Nagai, M. Hashizume, M. Tanaka, MicroRNA-10a is overexpressed in human pancreatic cancer and involved in its invasiveness partially via suppression of the HOXA1 gene. Ann. Surg. Oncol. 19, 2394–2402 (2012)
F.U. Weiss, I.J. Marques, J.M. Woltering, D.H. Vlecken, A. Aghdassi, L.I. Partecke, C.D. Heidecke, M.M. Lerch, C.P. Bagowski, Retinoic acid receptor antagonists inhibit miR-10a expression and block metastatic behavior of pancreatic cancer. Gastroenterology 137, 2136–45.e1-7 (2009)
H. Ouyang, J. Gore, S. Deitz, M. Korc, microRNA-10b enhances pancreatic cancer cell invasion by suppressing TIP30 expression and promoting EGF and TGF-beta actions. Oncogene 33, 4664–4674 (2014)
K. Nakata, K. Ohuchida, K. Mizumoto, T. Kayashima, N. Ikenaga, H. Sakai, C. Lin, H. Fujita, T. Otsuka, S. Aishima, E. Nagai, Y. Oda, M. Tanaka, MicroRNA-10b is overexpressed in pancreatic cancer, promotes its invasiveness, and correlates with a poor prognosis. Surgery 150, 916–922 (2011)
J. Yu, K. Ohuchida, K. Mizumoto, H. Fujita, K. Nakata, M. Tanaka, MicroRNA miR-17-5p is overexpressed in pancreatic cancer, associated with a poor prognosis, and involved in cancer cell proliferation and invasion. Cancer Biol. Ther. 10, 748–757 (2010)
H.J. Yan, W.S. Liu, W.H. Sun, J. Wu, M. Ji, Q. Wang, X. Zheng, J.T. Jiang, C.P. Wu, miR-17-5p inhibitor enhances chemosensitivity to gemcitabine via upregulating Bim expression in pancreatic cancer cells. Dig. Dis. Sci. 57, 3160–3167 (2012)
Y. Nagao, M. Hisaoka, A. Matsuyama, S. Kanemitsu, T. Hamada, T. Fukuyama, R. Nakano, A. Uchiyama, M. Kawamoto, K. Yamaguchi, H. Hashimoto, Association of microRNA-21 expression with its targets, PDCD4 and TIMP3, in pancreatic ductal adenocarcinoma. Mod. Pathol. 25, 112–121 (2012)
J. Dong, Y.P. Zhao, L. Zhou, T.P. Zhang, G. Chen, Bcl-2 upregulation induced by miR-21 via a direct interaction is associated with apoptosis and chemoresistance in MIA PaCa-2 pancreatic cancer cells. Arch. Med. Res. 42, 8–14 (2011)
T. Moriyama, K. Ohuchida, K. Mizumoto, J. Yu, N. Sato, T. Nabae, S. Takahata, H. Toma, E. Nagai, M. Tanaka, MicroRNA-21 modulates biological functions of pancreatic cancer cells including their proliferation, invasion, and chemoresistance. Mol. Cancer Ther. 8, 1067–1074 (2009)
Y. Ma, S. Yu, W. Zhao, Z. Lu, J. Chen, miR-27a regulates the growth, colony formation and migration of pancreatic cancer cells by targeting Sprouty2. Cancer Lett. 298, 150–158 (2010)
G. He, L. Zhang, Q. Li, L. Yang, miR-92a/DUSP10/JNK signalling axis promotes human pancreatic cancer cells proliferation. Biomed. Pharmacother. 68, 25–30 (2014)
W.G. Li, Y.Z. Yuan, M.M. Qiao, Y.P. Zhang, High dose glargine alters the expression profiles of microRNAs in pancreatic cancer cells. World J. Gastroenterol. 18, 2630–2639 (2012)
P. Li, Q. Xu, D. Zhang, X. Li, L. Han, J. Lei, W. Duan, Q. Ma, Z. Wu, Z. Wang, Upregulated miR-106a plays an oncogenic role in pancreatic cancer. FEBS Lett. 588, 705–712 (2014)
Z. Bai, J. Sun, X. Wang, H. Wang, H. Pei, Z. Zhang, MicroRNA-153 is a prognostic marker and inhibits cell migration and invasion by targeting SNAI1 in human pancreatic ductal adenocarcinoma. Oncol. Rep. 34, 595–602 (2015)
D. Takiuchi, H. Eguchi, H. Nagano, Y. Iwagami, Y. Tomimaru, H. Wada, K. Kawamoto, S. Kobayashi, S. Marubashi, M. Tanemura, M. Mori, Y. Doki, Involvement of microRNA-181b in the gemcitabine resistance of pancreatic cancer cells. Pancreatology 13, 517–523 (2013)
Z. Song, H. Ren, S. Gao, X. Zhao, H. Zhang, J. Hao, The clinical significance and regulation mechanism of hypoxia-inducible factor-1 and miR-191 expression in pancreatic cancer. Tumour Biol. 35, 11319–11328 (2014)
C. Zhao, J. Zhang, S. Zhang, D. Yu, Y. Chen, Q. Liu, M. Shi, C. Ni, M. Zhu, Diagnostic and biological significance of microRNA-192 in pancreatic ductal adenocarcinoma. Oncol. Rep. 30, 276–284 (2013)
J. Zhang, C.Y. Zhao, S.H. Zhang, D.H. Yu, Y. Chen, Q.H. Liu, M. Shi, C.R. Ni, M.H. Zhu, Upregulation of miR-194 contributes to tumor growth and progression in pancreatic ductal adenocarcinoma. Oncol. Rep. 31, 1157–1164 (2014)
M. Liu, Y. Du, J. Gao, J. Liu, X. Kong, Y. Gong, Z. Li, H. Wu, H. Chen, Aberrant expression miR-196a is associated with abnormal apoptosis, invasion, and proliferation of pancreatic cancer cells. Pancreas 42, 1169–1181 (2013)
F. Huang, J. Tang, X. Zhuang, Y. Zhuang, W. Cheng, W. Chen, H. Yao, S. Zhang, MiR-196a promotes pancreatic cancer progression by targeting nuclear factor kappa-B-inhibitor alpha. PLoS One 9, e87897 (2014)
S. Hamada, K. Satoh, S. Miura, M. Hirota, A. Kanno, A. Masamune, K. Kikuta, K. Kume, J. Unno, S. Egawa, F. Motoi, M. Unno, T. Shimosegawa, miR-197 induces epithelial-mesenchymal transition in pancreatic cancer cells by targeting p120 catenin. J. Cell. Physiol. 228, 1255–1263 (2013)
A. Liu, C. Shao, G. Jin, R. Liu, J. Hao, B. Song, L. Ouyang, X. Hu, miR-208-induced epithelial to mesenchymal transition of pancreatic cancer cells promotes cell metastasis and invasion. Cell Biochem. Biophys. 69, 341–346 (2014)
J. Jung, C. Yeom, Y.S. Choi, S. Kim, E. Lee, M.J. Park, S.W. Kang, S.B. Kim, S. Chang, Simultaneous inhibition of multiple oncogenic miRNAs by a multi-potent microRNA sponge. Oncotarget 6, 20370–20387 (2015)
J. Ma, B. Fang, F. Zeng, C. Ma, H. Pang, L. Cheng, Y. Shi, H. Wang, B. Yin, J. Xia, Z. Wang, Down-regulation of miR-223 reverses epithelial-mesenchymal transition in gemcitabine-resistant pancreatic cancer cells. Oncotarget 6, 1740–1749 (2015)
J. Ma, L. Cheng, H. Liu, J. Zhang, Y. Shi, F. Zeng, L. Miele, F.H. Sarkar, J. Xia, Z. Wang, Genistein down-regulates miR-223 expression in pancreatic cancer cells. Curr. Drug Targets 14, 1150–1156 (2013)
Z. Chen, L.Y. Chen, H.Y. Dai, P. Wang, S. Gao, K. Wang, miR-301a promotes pancreatic cancer cell proliferation by directly inhibiting Bim expression. J. Cell. Biochem. 113, 3229–3235 (2012)
Z. Lu, Y. Li, A. Takwi, B. Li, J. Zhang, D.J. Conklin, K.H. Young, R. Martin, Y. Li, miR-301a as an NF-kappaB activator in pancreatic cancer cells. EMBO J. 30, 57–67 (2011)
N. Funamizu, C.R. Lacy, S.T. Parpart, A. Takai, Y. Hiyoshi, K. Yanaga, MicroRNA-301b promotes cell invasiveness through targeting TP63 in pancreatic carcinoma cells. Int. J. Oncol. 44, 725–734 (2014)
S. Hamada, A. Masamune, S. Miura, K. Satoh, T. Shimosegawa, MiR-365 induces gemcitabine resistance in pancreatic cancer cells by targeting the adaptor protein SHC1 and pro-apoptotic regulator BAX. Cell. Signal. 26, 179–185 (2014)
D. He, H. Miao, Y. Xu, L. Xiong, Y. Wang, H. Xiang, H. Zhang, Z. Zhang, MiR-371-5p facilitates pancreatic cancer cell proliferation and decreases patient survival. PLoS One 9, e112930 (2014)
K. Wu, G. Hu, X. He, P. Zhou, J. Li, B. He, W. Sun, MicroRNA-424-5p suppresses the expression of SOCS6 in pancreatic cancer. Pathol. Oncol. Res. 19, 739–748 (2013)
S. Hasegawa, H. Eguchi, H. Nagano, M. Konno, Y. Tomimaru, H. Wada, N. Hama, K. Kawamoto, S. Kobayashi, N. Nishida, J. Koseki, T. Nishimura, N. Gotoh, S. Ohno, N. Yabuta, H. Nojima, M. Mori, Y. Doki, H. Ishii, MicroRNA-1246 expression associated with CCNG2-mediated chemoresistance and stemness in pancreatic cancer. Br. J. Cancer 111, 1572–1580 (2014)
Y. Wang, Z. Li, S. Zheng, Y. Zhou, L. Zhao, H. Ye, X. Zhao, W. Gao, Z. Fu, Q. Zhou, Y. Liu, R. Chen, Expression profile of long non-coding RNAs in pancreatic cancer and their clinical significance as biomarkers. Oncotarget 6, 35684–35698 (2015)
Q. Wang, H. Jiang, C. Ping, R. Shen, T. Liu, J. Li, Y. Qian, Y. Tang, S. Cheng, W. Yao, L. Wang, Exploring the Wnt pathway-associated LncRNAs and genes involved in pancreatic carcinogenesis driven by Tp53 mutation. Pharm. Res. 32, 793–805 (2015)
A.C. Tahira, M.S. Kubrusly, M.F. Faria, B. Dazzani, R.S. Fonseca, V. Maracaja-Coutinho, S. Verjovski-Almeida, M.C. Machado, E.M. Reis, Long noncoding intronic RNAs are differentially expressed in primary and metastatic pancreatic cancer. Mol. Cancer. 10, 141 (2011). doi:10.1186/1476-4598-10-141
S. Cao, W. Liu, F. Li, W. Zhao, C. Qin, Decreased expression of lncRNA GAS5 predicts a poor prognosis in cervical cancer. Int. J. Clin. Exp. Pathol. 7, 6776–6783 (2014)
M. Mourtada-Maarabouni, M.R. Pickard, V.L. Hedge, F. Farzaneh, G.T. Williams, GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene 28, 195–208 (2009)
Y. Nakamura, N. Takahashi, E. Kakegawa, K. Yoshida, Y. Ito, H. Kayano, N. Niitsu, I. Jinnai, M. Bessho, The GAS5 (growth arrest-specific transcript 5) gene fuses to BCL6 as a result of t(1;3)(q25;q27) in a patient with B-cell lymphoma. Cancer Genet. Cytogenet. 182, 144–149 (2008)
K. Yacqub-Usman, M.R. Pickard, G.T. Williams, Reciprocal regulation of GAS5 lncRNA levels and mTOR inhibitor action in prostate cancer cells. Prostate 75, 693–705 (2015)
X. Lu, Y. Fang, Z. Wang, J. Xie, Q. Zhan, X. Deng, H. Chen, J. Jin, C. Peng, H. Li, B. Shen, Downregulation of gas5 increases pancreatic cancer cell proliferation by regulating CDK6. Cell Tissue Res. 354, 891–896 (2013)
P. Ji, S. Diederichs, W. Wang, S. Boing, R. Metzger, P.M. Schneider, N. Tidow, B. Brandt, H. Buerger, E. Bulk, M. Thomas, W.E. Berdel, H. Serve, C. Muller-Tidow, MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22, 8031–8041 (2003)
E.J. Pang, R. Yang, X.B. Fu, Y.F. Liu, Overexpression of long non-coding RNA MALAT1 is correlated with clinical progression and unfavorable prognosis in pancreatic cancer. Tumour Biol. 36, 2403–2407 (2015)
F. Jiao, H. Hu, C. Yuan, L. Wang, W. Jiang, Z. Jin, Z. Guo, L. Wang, Elevated expression level of long noncoding RNA MALAT-1 facilitates cell growth, migration and invasion in pancreatic cancer. Oncol. Rep. 32, 2485–2492 (2014)
F. Jiao, H. Hu, T. Han, C. Yuan, L. Wang, Z. Jin, Z. Guo, L. Wang, Long noncoding RNA MALAT-1 enhances stem cell-like phenotypes in pancreatic cancer cells. Int. J. Mol. Sci. 16, 6677–6693 (2015)
K. Panzitt, M.M. Tschernatsch, C. Guelly, T. Moustafa, M. Stradner, H.M. Strohmaier, C.R. Buck, H. Denk, R. Schroeder, M. Trauner, K. Zatloukal, Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology 132, 330–342 (2007)
W. Peng, W. Gao, J. Feng, Long noncoding RNA HULC is a novel biomarker of poor prognosis in patients with pancreatic cancer. Med. Oncol. 31, 346 (2014). doi:10.1007/s12032-014-0346-4
M. Hajjari, A. Salavaty, HOTAIR: an oncogenic long non-coding RNA in different cancers. Cancer. Biol. Med. 12, 1–9 (2015)
J.K. Stratford, D.J. Bentrem, J.M. Anderson, C. Fan, K.A. Volmar, J.S. Marron, E.D. Routh, L.S. Caskey, J.C. Samuel, C.J. Der, L.B. Thorne, B.F. Calvo, H.J. Kim, M.S. Talamonti, C.A. Iacobuzio-Donahue, M.A. Hollingsworth, C.M. Perou, J.J. Yeh, A six-gene signature predicts survival of patients with localized pancreatic ductal adenocarcinoma. PLoS Med. 7, e1000307 (2010)
M.C. Tsai, O. Manor, Y. Wan, N. Mosammaparast, J.K. Wang, F. Lan, Y. Shi, E. Segal, H.Y. Chang, Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010)
R.A. Gupta, N. Shah, K.C. Wang, J. Kim, H.M. Horlings, D.J. Wong, M.C. Tsai, T. Hung, P. Argani, J.L. Rinn, Y. Wang, P. Brzoska, B. Kong, R. Li, R.B. West, M.J. van de Vijver, S. Sukumar, H.Y. Chang, Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010)
Y. Jiang, Z. Li, S. Zheng, H. Chen, X. Zhao, W. Gao, Z. Bi, K. You, Y. Wang, W. Li, L. Li, Y. Liu, R. Chen, The long non-coding RNA HOTAIR affects the radiosensitivity of pancreatic ductal adenocarcinoma by regulating the expression of Wnt inhibitory factor 1. Tumour Biol. (2015). doi:10.1007/s13277-015-4234-0
A.E. Teschendorff, S.H. Lee, A. Jones, H. Fiegl, M. Kalwa, W. Wagner, K. Chindera, I. Evans, L. Dubeau, A. Orjalo, H.M. Horlings, L. Niederreiter, A. Kaser, W. Yang, E.L. Goode, B.L. Fridley, R.G. Jenner, E.M. Berns, E. Wik, H.B. Salvesen, G.B. Wisman, A.G. van der Zee, B. Davidson, C.G. Trope, S. Lambrechts, I. Vergote, H. Calvert, I.J. Jacobs, M. Widschwendter, HOTAIR and its surrogate DNA methylation signature indicate carboplatin resistance in ovarian cancer. Genome Med. 7, 108 (2015). doi:10.1186/s13073-015-0233-4
K.C. Wang, Y.W. Yang, B. Liu, A. Sanyal, R. Corces-Zimmerman, Y. Chen, B.R. Lajoie, A. Protacio, R.A. Flynn, R.A. Gupta, J. Wysocka, M. Lei, J. Dekker, J.A. Helms, H.Y. Chang, A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011)
J.J. Song, R.E. Kingston, WDR5 interacts with mixed lineage leukemia (MLL) protein via the histone H3-binding pocket. J. Biol. Chem. 283, 35258–35264 (2008)
L. Quagliata, M.S. Matter, S. Piscuoglio, L. Arabi, C. Ruiz, A. Procino, M. Kovac, F. Moretti, Z. Makowska, T. Boldanova, J.B. Andersen, M. Hammerle, L. Tornillo, M.H. Heim, S. Diederichs, C. Cillo, L.M. Terracciano, Long noncoding RNA HOTTIP/HOXA13 expression is associated with disease progression and predicts outcome in hepatocellular carcinoma patients. Hepatology 59, 911–923 (2014)
Z. Li, X. Zhao, Y. Zhou, Y. Liu, Q. Zhou, H. Ye, Y. Wang, J. Zeng, Y. Song, W. Gao, S. Zheng, B. Zhuang, H. Chen, W. Li, H. Li, H. Li, Z. Fu, R. Chen, The long non-coding RNA HOTTIP promotes progression and gemcitabine resistance by regulating HOXA13 in pancreatic cancer. J. Transl. Med. 13, 84 (2015). doi:10.1186/s12967-015-0442-z
Y. Cheng, I. Jutooru, G. Chadalapaka, J.C. Corton, S. Safe, The long non-coding RNA HOTTIP enhances pancreatic cancer cell proliferation, survival and migration. Oncotarget 6, 10840–10852 (2015)
Y.W. Sun, Y.F. Chen, J. Li, Y.M. Huo, D.J. Liu, R. Hua, J.F. Zhang, W. Liu, J.Y. Yang, X.L. Fu, T. Yan, J. Hong, H. Cao, A novel long non-coding RNA ENST00000480739 suppresses tumour cell invasion by regulating OS-9 and HIF-1alpha in pancreatic ductal adenocarcinoma. Br. J. Cancer 111, 2131–2141 (2014)
S. Zheng, H. Chen, Y. Wang, W. Gao, Z. Fu, Q. Zhou, Y. Jiang, Q. Lin, L. Tan, H. Ye, X. Zhao, Y. Luo, G. Li, L. Ye, Y. Liu, W. Li, Z. Li, R. Chen, Long non-coding RNA LOC389641 promotes progression of pancreatic ductal adenocarcinoma and increases cell invasion by regulating E-cadherin in a TNFRSF10A-related manner. Cancer Lett. 37, 354–365 (2016)
A. Ashkenazi, R.C. Pai, S. Fong, S. Leung, D.A. Lawrence, S.A. Marsters, C. Blackie, L. Chang, A.E. McMurtrey, A. Hebert, L. DeForge, I.L. Koumenis, D. Lewis, L. Harris, J. Bussiere, H. Koeppen, Z. Shahrokh, R.H. Schwall, Safety and antitumor activity of recombinant soluble Apo2 ligand. J. Clin. Invest. 104, 155–162 (1999)
I. Ariel, S. Ayesh, E.J. Perlman, G. Pizov, V. Tanos, T. Schneider, V.A. Erdmann, D. Podeh, D. Komitowski, A.S. Quasem, N. de Groot, A. Hochberg, The product of the imprinted H19 gene is an oncofetal RNA. Mol. Pathol. 50, 34–44 (1997)
T. Arima, T. Matsuda, N. Takagi, N. Wake, Association of IGF2 and H19 imprinting with choriocarcinoma development. Cancer Genet. Cytogenet. 93, 39–47 (1997)
G. Banet, O. Bibi, I. Matouk, S. Ayesh, M. Laster, K.M. Kimber, M. Tykocinski, N. de Groot, A. Hochberg, P. Ohana, Characterization of human and mouse H19 regulatory sequences. Mol. Biol. Rep. 27, 157–165 (2000)
I.J. Matouk, N. DeGroot, S. Mezan, S. Ayesh, R. Abu-lail, A. Hochberg, E. Galun, The H19 non-coding RNA is essential for human tumor growth. PLoS One 2, e845 (2007)
C. Ma, K. Nong, H. Zhu, W. Wang, X. Huang, Z. Yuan, K. Ai, H19 promotes pancreatic cancer metastasis by derepressing let-7’s suppression on its target HMGA2-mediated EMT. Tumour Biol. 35, 9163–9169 (2014)
D. Amit, A. Hochberg, Development of targeted therapy for a broad spectrum of cancers (pancreatic cancer, ovarian cancer, glioblastoma and HCC) mediated by a double promoter plasmid expressing diphtheria toxin under the control of H19 and IGF2-P4 regulatory sequences. Int. J. Clin. Exp. Med. 5, 296–305 (2012)
A.A. Sidi, P. Ohana, S. Benjamin, M. Shalev, J.H. Ransom, D. Lamm, A. Hochberg, I. Leibovitch, Phase I/II marker lesion study of intravesical BC-819 DNA plasmid in H19 over expressing superficial bladder cancer refractory to bacillus Calmette-Guerin. J. Urol. 180, 2379–2383 (2008)
A. Mizrahi, A. Czerniak, T. Levy, S. Amiur, J. Gallula, I. Matouk, R. Abu-lail, V. Sorin, T. Birman, N. de Groot, A. Hochberg, P. Ohana, Development of targeted therapy for ovarian cancer mediated by a plasmid expressing diphtheria toxin under the control of H19 regulatory sequences. J.Transl.Med. 7, 69 (2009). doi:10.1186/1479-5876-7-69
ClinicalTrials.gov [Internet] Identifier: NCT01413087, Efficacy and safety of BC-819 and gemcitabine in patients with locally advanced pancreatic adenocarcinoma (LAPC-BC-819), 2015 (2012)
J.H. Liu, G. Chen, Y.W. Dang, C.J. Li, D.Z. Luo, Expression and prognostic significance of lncRNA MALAT1 in pancreatic cancer tissues. Asian Pac. J. Cancer Prev. 15, 2971–2977 (2014)
E. Heister, V. Neves, C. Lamprecht, S.R.P. Silva, H.M. Coley, J. McFadden, Drug loading, dispersion stability, and therapeutic efficacy in targeted drug delivery with carbon nanotubes. 50, 622–632 (2012)
C.J. Cheng, R. Bahal, I.A. Babar, Z. Pincus, F. Barrera, C. Liu, A. Svoronos, D.T. Braddock, P.M. Glazer, D.M. Engelman, W.M. Saltzman, F.J. Slack, MicroRNA silencing for cancer therapy targeted to the tumour microenvironment. Nature 518, 107–110 (2015)
C.F. Bennett, E.E. Swayze, RNA targeting therapeutics: molecular mechanisms of antisense oligonucleotides as a therapeutic platform. Annu. Rev. Pharmacol. Toxicol. 50, 259–293 (2010)
C.M. Sousa, A.C. Kimmelman, The complex landscape of pancreatic cancer metabolism. Carcinogenesis 35, 1441–1450 (2014)
C.W. Kimbrough, A. Khanal, M. Zeiderman, B.R. Khanal, N.C. Burton, K.M. McMasters, S.M. Vickers, W.E. Grizzle, L.R. McNally, Targeting acidity in pancreatic adenocarcinoma: multispectral optoacoustic tomography detects pH-low insertion peptide probes in vivo. Clin. Cancer Res. 21, 4576–4585 (2015)
H. Atkinson, R. Chalmers, Delivering the goods: viral and non-viral gene therapy systems and the inherent limits on cargo DNA and internal sequences. Genetica 138, 485–498 (2010)
B.D. Adams, C. Parsons, F.J. Slack, The tumor-suppressive and potential therapeutic functions of miR-34a in epithelial carcinomas. Expert Opin. Ther. Targets (2015). doi:10.1517/14728222.2016.1114102
G. Almer, K.L. Summers, B. Scheicher, J. Kellner, I. Stelzer, G. Leitinger, A. Gries, R. Prassl, A. Zimmer, H. Mangge, Interleukin 10-coated nanoparticle systems compared for molecular imaging of atherosclerotic lesions. Int. J. Nanomedicine 9, 4211–4222 (2014)
S. Zalba, A.M. Contreras, A. Haeri, T.L. Ten Hagen, I. Navarro, G. Koning, M.J. Garrido, Cetuximab-oxaliplatin-liposomes for epidermal growth factor receptor targeted chemotherapy of colorectal cancer. J. Control. Release 210, 26–38 (2015)
K. Bates, K. Kostarelos, Carbon nanotubes as vectors for gene therapy: past achievements, present challenges and future goals. Adv. Drug Deliv. Rev. 65, 2023–2033 (2013)
H.L. Janssen, H.W. Reesink, E.J. Lawitz, S. Zeuzem, M. Rodriguez-Torres, K. Patel, A.J. van der Meer, A.K. Patick, A. Chen, Y. Zhou, R. Persson, B.D. King, S. Kauppinen, A.A. Levin, M.R. Hodges, Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 368, 1685–1694 (2013)
E.C. Verna, V. Dhar, Endoscopic ultrasound-guided fine needle injection for cancer therapy: the evolving role of therapeutic endoscopic ultrasound. Ther. Adv. Gastroenterol. 1, 103–109 (2008)
A. Kambadakone, A. Thabet, D.A. Gervais, P.R. Mueller, R.S. Arellano, CT-guided celiac plexus neurolysis: a review of anatomy, indications, technique, and tips for successful treatment. Radiographics 31, 1599–1621 (2011)
V. Scaiewicz, V. Sorin, Y. Fellig, T. Birman, A. Mizrahi, J. Galula, R. Abu-Lail, T. Shneider, P. Ohana, L. Buscail, A. Hochberg, A. Czerniak, Use of H19 gene regulatory sequences in DNA-based therapy for pancreatic cancer. J. Oncol. 2010, 178174 (2010)
J. Kota, R.R. Chivukula, K.A. O’Donnell, E.A. Wentzel, C.L. Montgomery, H.W. Hwang, T.C. Chang, P. Vivekanandan, M. Torbenson, K.R. Clark, J.R. Mendell, J.T. Mendell, Therapeutic delivery of miR-26a inhibits cancer cell proliferation and induces tumor-specific apoptosis. Cell 137, 1005–1017 (2009)
G.A. Calin, C.D. Dumitru, M. Shimizu, R. Bichi, S. Zupo, E. Noch, H. Aldler, S. Rattan, M. Keating, K. Rai, L. Rassenti, T. Kipps, M. Negrini, F. Bullrich, C.M. Croce, Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. U. S. A. 99, 15524–15529 (2002)
H.A. Burris 3rd, M.J. Moore, J. Andersen, M.R. Green, M.L. Rothenberg, M.R. Modiano, M.C. Cripps, R.K. Portenoy, A.M. Storniolo, P. Tarassoff, R. Nelson, F.A. Dorr, C.D. Stephens, D.D. Von Hoff, Improvements in survival and clinical benefit with gemcitabine as first-line therapy for patients with advanced pancreas cancer: a randomized trial. J. Clin. Oncol. 15, 2403–2413 (1997)
Y. Ren, J. Gao, J.Q. Liu, X.W. Wang, J.J. Gu, H.J. Huang, Y.F. Gong, Z.S. Li, Differential signature of fecal microRNAs in patients with pancreatic cancer. Mol. Med. Rep. 6, 201–209 (2012)
M. Humeau, A. Vignolle-Vidoni, F. Sicard, F. Martins, B. Bournet, L. Buscail, J. Torrisani, P. Cordelier, Salivary MicroRNA in pancreatic cancer patients. PLoS One 10, e0130996 (2015)
S.A. Dhayat, B. Abdeen, G. Kohler, N. Senninger, J. Haier, W.A. Mardin, MicroRNA-100 and microRNA-21 as markers of survival and chemotherapy response in pancreatic ductal adenocarcinoma UICC stage II. Clin. Epigenetics 7, 132 (2015). doi:10.1186/s13148-015-0166-1. eCollection 2015
J. Liu, J. Gao, Y. Du, Z. Li, Y. Ren, J. Gu, X. Wang, Y. Gong, W. Wang, X. Kong, Combination of plasma microRNAs with serum CA19-9 for early detection of pancreatic cancer. Int. J. Cancer 131, 683–691 (2012)
K.H. Lee, J.K. Lee, D.W. Choi, I.G. Do, I. Sohn, K.T. Jang, S.H. Jung, J.S. Heo, S.H. Choi, K.T. Lee, Postoperative prognosis prediction of pancreatic cancer with seven MicroRNAs. Pancreas 44, 764–768 (2015)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors of this article, V.Taucher, J. Haybaeck and H. Mangge declare no conflicts of interest.
Rights and permissions
About this article
Cite this article
Taucher, V., Mangge, H. & Haybaeck, J. Non-coding RNAs in pancreatic cancer: challenges and opportunities for clinical application. Cell Oncol. 39, 295–318 (2016). https://doi.org/10.1007/s13402-016-0275-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-016-0275-7