Abstract
This study aimed to produce a novel, highly thermostable laccase from Bacillus licheniformis O12 bacterial DNA using a recombinant method and investigate the effects of some dyes found in textile wastewaters on decolorization. A putative laccase gene (CotA) from B. licheniformis O12 was cloned and expressed as a fusion protein in Escherichia coli BL21 (DE3) cells. The recombinant laccase was purified using the Ni-NTA affinity chromatography method. Accordingly, 411.76-fold purification was achieved with 0.014 U/mg specific activity, and the molecular mass of the enzyme was calculated as 66 kDa. The optimum temperature and pH values of the laccase were determined as 92 °C and pH 5.0, respectively. It was also found to be quite stable for 75 min under acidic and alkaline conditions. The activity was measured for 12 h at 60 °C and 92 °C. At 92 °C, it was observed that the activity halved after 12 h. The highest value at 60 °C was obtained at 9 h. While an activity decrease of approximately 10 % was observed in the first three hours, a slow increase was detected afterward. These results proved that the obtained laccase was highly resistant to pH and temperature. The recombinant laccase was significantly activated by Al3+, Cd2+, Cr2+, Cu2+, Fe2+, Hg2+, Pb2+, and was inhibited in the presence of organic solvents, surfactants, and laccase inhibitors. Finally, the effects of some textile dyes on decolorization were investigated using the fused recombinant laccase and found to be generally effective. The highest decolorization of the laccase treated with dye for 2 h was 51.2 % in acid black 1, while the lowest decolorization was 1.9 % in the congo red.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Laccases (benzenediol: oxygen oxidoreductases; EC 1.10.3.2) are catalysts that oxidize many phenolic and aromatic compounds belonging to the polyphenol family using free oxygen (Jiang et al. 2021; Zhou et al. 2021). Therefore, there is a wide substrate specificity because they can work with a large number of substrates, be used in the textile industry, fabric bleaching, lignin degradation for paper production, food industry, wine treatment, medicine, synthesis of organic chemicals, and biosensing processes in enzymatic fuel cells (Nakamura 2005; Liu et al. 2011). There are four copper atoms in the structure of laccases, so they are called multi-copper blue proteins. The copper in the type 1 region draws electrons from the substrate and passes to the type 2/type 3 region, releasing H2O. Furthermore, there are three copper atoms in the type 2/type 3 region. Therefore, it is called the trinuclear region (Nakamura 2005).
Laccases are found in plants, insects, bacteria, archaea, and fungi (Claus 2003; Uthandi et al. 2010). Studies on laccases have mostly been conducted with fungi (Baldrian 2006). Most fungal laccases are unstable at alkaline pH values and high temperatures (Guan et al. 2014). The optimum pH values of fungal laccases are generally obtained in slightly acidic environments, and they usually cannot tolerate other pH ranges. Moreover, when the recombinant laccase is produced from fungi, problems occur due to introns in the structure, disulfide bonds, and glycosylation (Rodgers et al. 2010). On the other hand, bacterial laccases can work in a wider pH range and are more resistant to high temperature and alkaline environments (Guan et al. 2018). Genome analysis shows that there are many laccase-encoding genes in bacterial genomes (Ausec et al. 2011). Studies on bacterial laccases have been carried out recently, but they are not sufficient. It is very important for biotechnology to obtain and characterize this important enzyme, which has a wide range of uses, in more bacterial sources (Endo et al. 2003; Guo et al. 2016; Guan et al. 2018).
A place where laccases are frequently used in industries is the decolorization process in textile wastewater. Many textile dyes used cannot be separated by physical and chemical processes, and when separated, they can release toxic substances and harm human health and the ecosystem (Mohorcic et al. 2006; Khlifi et al. 2010; Abdel-Shafy and Mansour 2016). Toxic substances and dyes released into nature can be inhaled or absorbed by living things as contaminants. Laccases are frequently used in the decolorization of textile dyes (Mehandia et al. 2019).
In this study, we cloned the laccase gene from the test strain identified as B. licheniformis O12, with the high laccase activity and expressed in E. coli bacteria, and the recombinant laccase was characterized. We also showed its usability in decolorization applications in textile wastewater.
Materials and methods
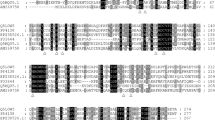
Chemicals used in the study were purchased from Sigma. B. licheniformis O12 (Fig. 1, GenBank accession number KM596797) was obtained by Prof. Dr. Ahmet Adıgüzel from the Molecular Biology and Genetics Department of Atatürk University. The test strain was isolated from Agri-Diyadin hot spring and identified (Baltaci et al. 2017). Primers were purchased from Sentegen Biotech (Turkey). Laccase cloning and expression processes were performed using the Champion ™ pET SUMO protein expression system kit (Invitrogen). The purification of the recombinant laccase was carried out using the ProBond ™ purification system (Novex).
Laccase cloning and plasmid construction
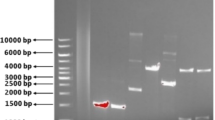
The genomic DNA of B. licheniformis O12 bacterium, a test strain, was obtained using a commercial kit (Pure Link Hi-Pure plasmid DNA purification kit Invitrogen). Primers used in the polymerase chain reaction were designed on the NCBI site by comparing the gene sequence in the laccase studies previously performed with bacteria of the Bacillus genus (Forward primer 5’-ATG AAA CTT GAA AAA TTC GTT GAC-3’ and reverse primer 5’-TTA TTG ATG ACG AAC ATC TGT CAC- 3’). PCR was performed using B. licheniformis O12 bacteria genomic DNA and the designed primers. After pre-denaturation at 94 °C for DNA denaturation, 35 cycles were performed at 94 °C for 1 min, at 52 °C for 1 min, and at 72 °C for 1.5 min. The PCR product was controlled by 1 % agarose gel electrophoresis. The PCR product, containing 3′ deoxyadenosine (A) residues, was ligated by T4 DNA ligase into the linearized pET SUMO vector, containing deoxythymidine (T) overhangs.
The laccase genes amplified by PCR were inserted into the pET SUMO vector by ligation. For the ligation to be successful, the concentration of the laccase gene and a molar vector-to-insert ratio should be 1:5 (Demir and Beydemir 2015). The A (adenine) extensions in the laccase gene were attached to the (thymine) extensions in the plasmid (A-T cloning). The recombinant vector obtained as a result of the ligation process was transformed into E. coli One Shot® Mach1™-T1R competent cells to produce large numbers. The transformation was performed as described in the Champion pET SUMO expression system user manual (Demir and Beydemir 2015).
At the end of the transformation process, PCR was performed with colonies grown on a solid medium containing kanamycin antibiotics (50 µg/mL). The aim was to check whether the cells formed after transformation contained the recombinant protein. After the PCR result was controlled by 1 % agarose gel electrophoresis, plasmid isolation was conducted following the procedure, and sequence analysis was performed using a Thermo Scientific GeneJET Plasmid Miniprep kit. The analysis result was verified by blasting on the NCBI site. It was observed that the laccase gene was 100 % compatible with the pET SUMO expression system (Invitrogen).
Expression and purification of recombinant His-SUMO-laccase
Expression was initiated with the laccase genes proved to be compatible with the literature. The transformation of the recombinant plasmid into E. coli BL21DE3 cells was performed as described in the Champion pET SUMO expression system. Then, the pilot expression process was carried out for 8 h with the kanamycin antibiotic with LB broth. Cells after lysis were lysed with lysis buffer, heat shock, and sonicator. Then, the presence of laccase was confirmed by the SDS-PAGE process. The water solubility of the obtained enzyme was proved by the SDS-PAGE method (Demir 2019).
Purification of laccase with the Ni-NTA affinity column
One of the features of the pET SUMO One Shot cloning system is the histidine tails in the plasmid structure. During cloning, histidine tails are added to laccase. This tail structure is very small, increases the water solubility of the enzyme, and also allows it to attach to the Ni-NTA column because the nickel ligands in the structure of the affinity column show high affinity for histidines in the tail structure. The elution solution contains imidazole, and the structure of imidazole also contains histidine. Due to the affinity of the column to the high concentration of imidazole, laccases detach from the column, and imidazoles are bound to nickel ligands in the column. Thus, we can easily purify the laccase in one step. After the purification step, laccase was checked by the SDS-PAGE method. The molecular mass of laccase was calculated.
Cutting the fusion recombinant laccase with SUMO protease
The laccase we obtained using the pET SUMO expression system is a fusion protein, and the recombinant laccase has SUMO. From this fusion protein structure, the SUMO protein was cut with the SUMO protease enzyme. To this end, it was kept at 30 °C for 5 h. At the end of the period, the result was checked by the SDS-PAGE method (Demir 2020).
Enzyme activity assay and protein determination
The laccase activity was measured by the oxidation of 2,2-azino-bis(3-ethylbenzothiazoline-6-sulfonate) (ABTS) substrate. The activity mixture contains 2 mM Na-acetate buffer (pH 5.0), 1 mM ABTS, and enzyme. This mixture was incubated at 55 °C for 15 min, and then absorbance was measured at 420 nm (ε420 = 36,000 M-1 cm-1) (Heinfling et al. 1998). The protein concentration of laccase was measured by the Bradford method using bovine serum albumin as the standard (Bradford 1976).
Effect of pH on the laccase activity and stability
Activity measurements were made in the range of pH 2.0 to 10.5 to determine the optimum pH value of the laccase we obtained as a recombinant. In the optimal pH study, glycine-HCl buffer at pH 2.0–3.0, acetate buffer at pH 4.0–5.0, phosphate buffer at pH 6.0–8.0, and glycine-NaOH buffer at pH 9.0-10.5 were used. The stability of laccase at pH 5.0 and pH 8.0 was determined by measuring the activity periodically, and then the residual activity was calculated.
Effects of temperature on laccase and stability
B. licheniformis O12 was chosen because it is a thermophilic bacterium and can work at high temperatures. Therefore, it was studied in a wide temperature range. The activity measurements were made in the range of 30 to 92 °C. 92 °C is the boiling point under study conditions in Erzurum. Then, the stability of the enzyme activity versus time was investigated at 60 and 92 °C. The activity measurement was carried out at 1-hour intervals for 12 h, and the residual activity was calculated.
Effects of metal ions, various chemical reagents, and organic solvents on laccase
Laccases are used for many purposes in the industry, so they are exposed to a wide variety of chemicals. Therefore, the activity measurement was performed in the presence of 10 metal ions (Al3+, Cd2+, Co2+, Cu2+, Fe2+, Hg2+, Cr2+ Mn2+, Pb2+, and Zn2+), organic solvents (methanol, ethanol, t-butanol, 1-propanol, DMSO, and acetone), surfactants (SDS, Triton X-100, Tween-20, and Tween-80), chemicals known as laccase inhibitors (sodium azide, sodium fluoride, and EDTA), and H2O2. During the activity measurement, 100 µL of the solutions of these substances were added, and the same amount was subtracted from the buffer solution. The residual activity was determined. All of the characterization studies were performed with the His-SUMO-laccase form.
Kinetic assays
The kinetic parameters of SUMO-laccase and laccase were investigated. The activity measurements were made in five different concentrations of ABTS, 2,6-dimethoxyphenol, and guaiacol substrates. The Lineweaver–Burk graphs were drawn, and KM, Vmax, kcat, and kcat/KM values were calculated. Galai’s procedure was applied to measure activity with the 2,6 DMP substrate. The spectrophotometric measurement was performed at 468 nm after 15 min of incubation in the presence of pH 5.0, 0.1 M phosphate buffer, and 0.2 mM CuSO4 (Galai et al. 2011). A 2 mM solution of the guaiacol substrate in 95 % ethanol was prepared. The spectrophotometric measurement was carried out at 465 nm after 5 min of incubation with 10 mM CuSO4 and pH 7.5 50 mM phosphate buffer (Liu et al. 2015).
Decolorization of dyes by the purified laccase
Laccases are frequently used in the decolorization processes of dyes in textile wastewater. In our study, we investigated the usability of laccase in the removal of dyes, such as acid black 1 (620 nm), congo red (350 nm), methylene blue (660 nm), orange (622 nm), acid red 27 (520 nm), and reactive black 5 (597 nm). Laccase was added to the dye solutions, spectrophotometric measurements were made at periodic intervals (15–120 min), and dye decolorization (%) was calculated (Lorenzo et al. 2006). No mediators were added to the reaction mixture. Dye solution without enzyme was used as a control.
Results and discussion
Laccase gene production and reproduction
We designed primers to obtain the laccase gene using gene information from Bacillus spp. in the literature. We applied the PCR protocol using primers and validated this process with 1 % agarose gel electrophoresis. After the ligation process, we transformed the recombinant vector we obtained into the E. coli One Shot® Mach1 ™-T1R competent cell. The sequence analysis was performed on plasmids after the transformation was proven by colony PCR. The sequence analysis of two colonies was carried out. One of them gave a 100 % concordant result with the Bacillus sp. SL-1 (CotA) gene, and the other one yielded a 98.12 % concordant result (Table 1). After seeing that we obtained the correct gene, the recombinant plasmids were transformed into E.coli BL21(DE3) cells, and pilot expression was performed in a kanamycin antibiotic medium.
Purification of the recombinant laccase with the Ni-NTA affinity column
The Ni-NTA affinity column is designed for recombinant enzymes and has nickel ligands. During cloning, histidine tails are added to the enzyme, and these tails are attached to nickel ligands and easily separated from other proteins. The elution process is carried out with imidazole solution, and imidazole also has an affinity for nickel. We purified the laccases obtained after the expression process using the mentioned method and performed quantitative protein determination by employing the Bradford method (Bradford 1976). Using the Ni-NTA affinity column, 411.76-fold purification was achieved with 0.014 U/mg specific activity (Table 2). The expression system used increases the water solubility of the resulting recombinant enzymes (Invitrogen). Subsequently, SUMO was cleavaged from laccase and verified by the SDS-PAGE method (Fig. 2). The molecular weight of the recombinant laccase was calculated to be 66 kDa. Previous studies have reported that the molecular weight is between 50 and 100 kDa for laccases originating from bacteria and yeasts (Zhu et al. 2016; Callejon et al. 2017) (Table 3).
SDS-PAGE analysis of SUMO-laccase fusion protein cleaved by SUMO protease and recombinant laccase. (a) SDS-PAGE analysis of SUMO-laccase fusion protein cleaved by SUMO protease, lane 1: Mixture of SUMO – laccase fusion protein by SUMO protease cleavage. (b) SDS – PAGE analysis of recombinant laccase, lane 1: Purified recombinant laccase by Ni-NTA. Lane M: Standard molecular weight markers (PageRegulerTM Prestained Protein Ladder)
Effects of pH on the recombinant laccase
The optimal pH study was conducted between pH 2.0 and pH 10.5, and the optimum value was found as pH 5.0 (Fig. 3). The optimum pH value of laccase can vary according to the source from which it is obtained. For the laccase obtained from Ganoderma sp., the optimum pH value was pH 4.0 (Kumar et al. 2017); for the laccase obtained from Myceliophthora thermophila and Trametes trogii, the optimum pH value was 4.5 (Herkommerova et al. 2018); for the laccase obtained from Pestalotiopsis sp. with different substrates, the optimum pH values were 2.0 and 5.0 (Wikee et al. 2019); for the laccase obtained from Bacillus subtitlis st. OH67, the optimum pH value was 6.5 (Hajipour et al. 2020). The optimum pH for ABTS and phenolic substances is similar for laccases and reflects the different oxidation mechanisms that depend on the substrate used (Xu 1997).
To measure the long-term stability of laccase in acidic and alkali environments, activity was measured in pH 5.0 and pH 8.0 buffers for 75 min (Fig. 2). It was observed that laccase maintained its activity at both pH 5.0 and pH 8.0 and even increased its activity. The laccase obtained from Paenibacillus glucanolyticus SLM1 decreased its activity after 4 h at the optimum pH value of 7.0 (Mathews et al. 2016). In the laccase obtained from Cohnella sp. A01, the optimum pH value was reported as 8.0, and the highest stability was measured at pH 6.0 and the lowest at pH 10.0 (Shafiei et al. 2019). In pH stability studies of two different laccase genes obtained from Pestalotiopsis sp., it was observed that both enzymes could not maintain stability at pH 2.0, and the PsLac1 gene was reported to be more stable between pH 3.0 and pH 6.0 (Wikee et al. 2019). Therefore, it can be said that the recombinant laccase produced from B. licheniformis O12 preserves its activity very well under acidic and alkaline conditions. In contrast to fungal laccases, which are restrictively stable in the acid-neutral pH range, bacterial laccases have been shown to active in a broader pH range and be more stable (Guan et al. 2014).
Effect of temperature on the recombinant laccase
The activity measurements were made between 30 and 92 °C, and the optimum temperature value was found to be 92 °C (Fig. 4). 92 °C is the boiling point of water under the study conditions. The optimum temperature value varies according to the source from which laccase is obtained. The optimum temperature value for the laccase obtained from Cohnella sp. A01 was found to be 90 °C (Shafiei et al. 2019). Likewise, the optimum temperature value for the laccase obtained from Thermus thermophilus SG0.5JP17-16 was reported as 90 °C (Liu et al. 2015). The optimum temperature value for the laccase obtained from Kurthia huakuii LAM0618T was determined as 85 °C (Guo et al. 2016). The optimum temperature for the laccase obtained from Paenibacillus glucanolyticus SLM1 was determined as 40 °C; for the laccase obtained from Bacillus subtitlis St. OH67, it was determined as 50 °C (Mathews et al. 2016; Hajipour et al. 2020). The optimum temperature for the laccase obtained from Pedioccocus acidilactici CECT5930 was reported as 28 °C, while it was 60 °C for the laccase obtained from Staphylococcus haemolyticus (Callejon et al. 2017; Li et al. 2020). It is advantageous for the industry to produce laccase that maintains its activity at high temperatures.
We measured activity per hour at both 60 and 92 °C for 12 h and found that the recombinant laccase obtained from B. licheniformis O12 continued to work for a very long time (Fig. 3). Therefore, 12-hour enzyme stability is a very efficient result. The laccase obtained from B. licheniformis O12 thermophilic bacteria maintained at least half of its initial activity for a very long time. It was revealed to be much more stable at 60 °C. These values demonstrate that the laccase produced is highly resistant to high temperatures. At the end of the period, it was observed that the laccase continued to work at both temperatures. The activity of the laccase obtained from Thermus thermophilus SG0.5JP17-16 was measured for 4 h at different temperatures (70 °C, 80 °C, and 90 °C), and it was reported that the enzyme did not lose any activity at 70 °C. At 80 °C and 90 °C, the laccase was been found to retain most of its activity (Liu et al. 2015). It was reported that the activity of the laccase obtained from Myceliophthora thermophila was up to 60 °C and that of the laccase obtained from Tramates tragii was stable up to 70 °C (Herkommerova et al. 2018). For the laccase obtained from Cohnella sp. A01, measurements were made at 60 °C, 90 °C, and 100 °C for 2 h, and the highest activity was obtained at 60 °C, while the lowest value was obtained at 100 °C (Shafiei et al. 2019). The laccase purified from Bacillus subtilis St. OH67 was reported to have decreased activity within 1 h (Hajipour et al. 2020). As can be seen from the results of the temperature stability experiments, the recombinant laccase is extremely robust, and this high thermostability appears to be the most striking feature of the recombinant laccase (Fig. 4).
Effects of metal ions, inhibitors, organic solvents, and surfactants effects on the laccase activity
The activity measurement was performed in the presence of some metal ions (Al3+, Cd2+, Co2+, Cu2, Fe2+, Hg2+, Cr2+, Mn2+, Pb2+, and Zn2+), organic solvents (methanol, ethanol, t-butanol, 1-propanol, DMSO, and acetone), surfactants (SDS, Triton X-100, Tween-20, and Tween-80), chemicals known as laccase inhibitors (sodium azide, sodium fluoride, and EDTA), and H2O2. Metal ions showed an activator effect on laccase, except for Zn2+ (Table 3). In general, laccases appear to be resistant to metal ions (Litwinska et al. 2019; Zhang et al. 2019). Ethanol, methanol, 1-propanol, t-butanol, and DMSO inhibited the recombinant laccase by approximately 90 % (Table 4). It is known that organic solvents inhibit laccase activity by promoting protein unfolding (D’Acunzo et al. 2004; Li et al. 2019). NaF, NaN3, and EDTA also inhibited the laccase (Table 5). H2O2 inhibited approximately 60 %. These substances are already known as laccase inhibitors (Sadhasivam et al. 2008; Liu et al. 2010; Afreen et al. 2017). When the inhibition of surfactants is examined, it is observed that SDS has the highest inhibition effect. If surfactants are added at high rates, they can have an inhibitory effect on laccases (Shafiei et al. 2019). In the study conducted with Morchella importuna, EDTA and SDS were found to inhibit laccase (Zhang et al. 2019). However, in the study carried out with Pediococcus acidilactici CECT 5930, it was observed that EDTA and SDS almost did not affect the laccase activity (Callejon et al. 2017).
Kinetic parameters
Substrates ABTS, 2,6-DMP, and guaiacol were used to measure the kinetic parameters of the laccase obtained from B. licheniformis O12. The measurements were made for the recombinant laccase in the fused state (laccase + SUMO) and the laccase obtained by cutting the SUMO (Table 6). The aim was to investigate whether SUMO had a negative or positive effect on the enzyme activity. Although the presence of SUMO affects the affinity of the enzyme to the substrate, it was found that it did not have an adverse effect on the kinetic parameters. The recombinant fusion laccase showed the highest affinity to the 2,6-DMP substrate (0.0102 mM). The highest catalytic efficiency (kcat/KM) was also measured with the same substrate (1.52 min− 1mM− 1). Among these substrates, it was observed that the guaiacol substrate showed the least affinity, and the catalytic efficiency was also lower in the guaiacol substrate than the others (0.0000017 min− 1mM− 1). The recombinant laccase obtained after cutting the SUMO showed the highest affinity to the guaiacol substrate (0.0136 mM), and the catalytic efficiency was also very high (156.4 min− 1 mM− 1) (Table 3). When the results are examined, it is revealed that the affinity of laccase for 2,6-DMP is always higher than that of guaiacol. The second methoxyl group in 2,6-DMP has a high redox ability. Guaiacol contains one methoxy group in its structure (Huang et al. 2011).
Dye decolorization
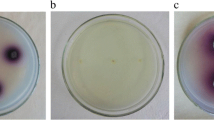
To investigate the effect of the laccase obtained from B. licheniformis O12 on the decolorization of dyes in textile wastewater, measurements were made with six dyes (acid black 1, orange, congo red, acid red 27, reactive black 5, and methylene blue). The spectrophotometric measurement was performed by adding the enzyme to the dye solutions, and no redox mediators were used. As a result of the measurements made at periodic intervals of 120 min, the highest decolorization was obtained in acid black 1 (51.2 %). There was decolorization of 36.2 % in methylene blue and 32.05 % in reactive black 5. The lowest decolorization was achieved in congo red with 1.9 %. It was observed that the decolorization rate increased over time; if the time was extended, the amount of decolorization increased (Table 7). The laccase originating from Staphylococcus haemolyticus was been used for the decolorization of congo red, brilliant green, bromophenol blue, and crystal violet dyes. The best decolorization was detected in brilliant green with 80 %, while the lowest decolorization was observed in crystal violet with approximately 40 %. The dye decolorization has been shown to stop after 3 h (Li et al. 2020). It was reported that the decolorizing effect of laccases was stronger when added to the laccase mediator medium (Herkommerova et al. 2018). Galai et al. carried out a decolorization study in the presence of four different mediators, including ABTS. A slight increase was reported when compared with the media-free results, and there was 80 % decolorization only in the presence of ABTS (Galai et al. 2011). Guo et al. stated that decolorization increased up to 5 times in the presence of two different mediators (ABTS, acetosyringone) (Guo et al. 2016). Considering that most textile effluents are characterized by high temperature and alkaline pH (Rodrigues et al. 2009), it appears that the recombinant laccase may have a promising application in this area. However, it is thought that the dye decolorization efficiency can be increased by adding a mediator.
Conclusions
In this study, the laccase-encoding gene from B. licheniformis O12 was expressed in the E.coli BL21 (DE3) expression system. Then, the recombinant laccase was purified and biochemically characterized. The enzyme showed the highest activity at a very high temperature (92 °C). At 92 °C, its activity halved after 12 h. At 60 °C, it retained more than half of its activity for 12 h. This is a very important feature because textile wastewater has high temperatures and it takes time and is expensive to cool it. The recombinant laccase is stable under both acidic and alkaline conditions, and its activity increased after 75 min at pH 5.0 and pH 8.0. It is known that the color of textile dyes increases as pH increases, so pH stability is important. It was observed that metal ions did not have a significant inhibitory effect on the recombinant laccase. The activating effects of Al3+, Cd2+, Cr2+, Cu2+, Fe2+, Hg2+, and Pb2+ ions were also found. Organic solvents (methanol, ethanol, 1-propanol, t-butanol, acetone, and DMSO) and surfactants (Tween-20, Tween-80, and Triton X-100) reduced the activity of the recombinant laccase by approximately 90 % after 2 h. SDS showed a strong inhibitory property, just like NaF and EDTA. H2O2 caused 60 % activity loss after 2 h. NaN3, known as a laccase inhibitor, caused 90 % activity loss after 3 h. The decolorization effect of recombinant laccase was 51.2 % in acid black 1 and 32.5 % in reactive black 5. Decolorization was performed without adding mediators to the medium. As a result, the laccase obtained from a novel strain by the recombinant method is an enzyme that exhibits very high thermostability, is resistant to pH changes and metal ions. It was concluded that the laccase obtained from B. licheniformis O12 has prominent properties in many aspects. Its use in the industry will be beneficial in many ways and economically.
In this study, laccase was obtained from B. licheniformis O12 by recombinant method and many properties of the enzyme were investigated. In line with the results obtained, the ability of the enzyme in different applications such as delignification or clarification can be examined. With the mutagenesis method, the properties of the enzyme can be improved and a stronger and more durable laccase can be obtained.
Data Availability
All authors approved that all data and materials comply with field standards.
Abbreviations
- EDTA:
-
Ethylenediamine tetraacetic acid
- DMSO:
-
Dimethyl sulfoxide
- IPTG:
-
Izopropil-β-D-tiyogalaktopiranozit
- kDa:
-
Kilo dalton
- kcat :
-
A unit showing catalytic ability of enzyme (turnover number)
- KM :
-
Unit showing the enzyme’s affinity for the substrate
- mg:
-
Miligram
- min:
-
Minute
- mM:
-
Milimole
- Ni-NTA:
-
Nickel-nitrilotriacetic acid
- nm:
-
Nanometer
- SDS-PAGE:
-
Sodium dodecyl sulfate polyacrylamide gel electrophoresis
- SUMO:
-
Small ubiquitin-like modifier
- µM:
-
Micromolar
- Vmax :
-
Maximum speed the enzyme reaches
- Ɛ:
-
Molecular extinction coefficient
References
Abdel-Shafy HI, Mansour MSM (2016) A review on polycyclic aromatic hydrocarbons: Source, environmental impact, effect on human health and remediation. Egypt J Pet 25:107–123. https://doi.org/10.1016/j.ejpe.2015.03.011
Afreen S, Shamsi TN, Baig MA, Ahmad N, Fatima S, Qureshi MI (2017) A novel multicopper oxidase (laccase) from cyanobacteria: Purification, characterization with potential in the decolorization of anthraquinonic dye. PLoS One 12(4):e0175144. https://doi.org/10.1371/journal.pone.0175144
Ausec L, Zakrzewski M, Goesmann A, Schlüter A, Mandic MI (2011) Bioinformatic Analysis Reveals High Diversity of Bacterial Genes for Laccase-Like Enzymes. PLoS One:6e25724. https://doi.org/10.1371/journal.pone.0025724
Baldrian P (2006) Fungal laccases-occurrence and properties. FEMS Microbiol Rev 30:215–242. https://doi.org/10.1111/j.1574-4976.2005.00010.x
Baltaci MO, Genc B, Arslan S, Adıguzel G, Adıguzel A (2017) Isolation and characterization of thermophilic bacteria from geothermal areas in Turkey and preliminary research on biotechnologically important enzyme production. Geomicrobiol J 34(1):53–62. https://doi.org/10.1080/01490451.2015.1137662
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248. https://doi.org/10.1006/abio.1976.9999
CallejoÂn S, Sendra R, Ferreri S, Pardol I (2017) Recombinant laccase from Pediococcus acidilactici CECT 5930 with ability to degrade Tyramine. PLoS One. https://doi.org/10.1371/journal.pone.0186019
Claus H (2003) Laccases and their occurrence in prokaryotes. Arch Microbiol 179:145–150. https://doi.org/10.1007/s00203-002-0510-7
D’Acunzo F, Barreca AM, Galli C (2004) Determination of the activity of laccase, and mediated oxidation of a lignin model compound, in aqueous-organic mixed solvents. Mol Catal B: Enzym 31:25–30. https://doi.org/10.1016/j.molcatb.2004.07.001
Demir Y (2019) The behaviour of some antihypertension drugs on human serum paraoxonase-1: an important protector enzyme against atherosclerosis. J Pharmacol 71(10):1976–1583. https://doi.org/10.1111/jphp.13144
Demir Y (2020) Naphthoquinones, benzoquinones, and anthraquinones: Molecular docking, ADME and inhibition studies on human serum paraoxonase-1 associated with cardiovascular diseases. Drug Dev Res 81(5):628–636. https://doi.org/10.1002/ddr.21667
Demir Y, Beydemir Ş (2015) Puri_cation, refolding, and characterization of recombinant human paraoxonase-1. Turk J Chem 39:764–776. https://doi.org/10.3906/kim-1501-51
Endo K, Hayashi Y, Hibi T, Hosono K, Beppu T, Ueda K (2003) Enzymological characterization of EpoA, a laccase-like phenol oxidase produced by Streptomyces 401 griseus. J Biochem 133:671–677. https://doi.org/10.1093/jb/mvg086
Galai S, Lucas EP, Marzouki MN, Sanchez AA (2011) Molecular cloning of a copper-dependent laccase from the dye-decolorizing strain Stenotrophomonas maltophilia AAP56. J Appl Microbiol :1364–5072. https://doi.org/10.1111/j.1365-2672.2011.05164.x
Guan ZB, Zhang N, Song CM, Zhou W, Zhou LX, Zhao H, Xu CW, Cai YJ, Liao XR (2014) Molecular cloning, characterization, and dye-decolorizing ability of a temperature and pH-stable laccase from Bacillus subtilis X. Appl Biochem Biotechnol 172:1147–1157. https://doi.org/10.1007/s12010-013-0614-3
Guan ZB, Luo Q, Wang HR, Chen Y, Liao XR (2018) Bacterial laccases: promising biological green tools for industrial applications. Cell Mol Life Sci 75:3569–3592. https://doi.org/10.1007/s00018-018-2883-z
Guo X, Zhou S, Wang Y, Song J, Wang H, Kong D, Zhu J, Dong W, He M, Hu G, Ruan Z (2016) Characterization of a highly thermostable and organic solvent-tolerant copper containing polyphenol oxidase with dye-decolorizing ability from Kurthia huakuii LAM0618T. PLoS One 11:e0164810. https://doi.org/10.1371/journal.pone.0164810
Hajipour O, Mercan Dogan N, Dincer S, Norizadehazehkand M (2020) Cloning, expression, and characterization of novel laccase enzyme from native Bacillus subtilis strain OH67. Mol Biol 54:611–617. https://doi.org/10.1134/S0026893320040068
Heinfling A, Martinez AT, Martinez MJ, Bergbauer M, Szewzyk U (1998) Purification and characterization of peroxidases from the dye-decolorizing fungus Bjerkandera adusta. FEMS Microbiol Lett 165:43e50. https://doi.org/10.1111/j.1574-6968.1998.tb13125.x
Herkommerová K, Dostál J, Pichová I (2018) Decolorization and detoxification of textile wastewaters by recombinant Myceliophthora thermophila and Trametes trogii laccases. 3 Biotech 8:505. https://doi.org/10.1007/s13205-018-1525-3
Huang WT, Tai R, Hseu RS, Huang CT (2011) Overexpression and characterization of a thermostable, pH-stable and organic solvent-tolerant Ganoderma fornicatum laccase in Pichia pastoris. Process Biochem 46:1469–1474. https://doi.org/10.1016/j.procbio.2011.03.020
Jiang Y, Cai J, Pei J, Li Q, Zhao L (2021) Cloning, overexpression, and characterization of a thermostable, organic solvent-tolerant laccase from Bacillus pumilus ARA and its application to dye decolorization. ACS Omega 6:9741–9749. https://doi.org/10.1021/acsomega.1c00370
Khlifi R, Belbahri L, Woodward S, Ellouz M, Dhouib A, Sayadi S, Mechichi T (2010) Decolourization and detoxification of textile industry wastewater by the laccase-mediator system. J Hazard Mater 175:802–808. https://doi.org/10.1016/j.jhazmat.2009.10.079
Kumar A, Singh D, Sharma KK, Arora S, Singh AK, Gill SS, Singhal B (2017) Gel-based purification and biochemical study of laccase isozymes from Ganoderma sp. and its role in enhanced cotton callogenesis. Front Microbiol 8:674. https://doi.org/10.3389/fmicb.2017.00674
Li X, Liu D, Wu Z, Li D, Cai Y, Lu Y, Zhao X, Xue H (2020) Multiple Tolerances and Dye Decolorization Ability of a Novel Laccase Identified from Staphylococcus Haemolyticus. J Microbiol Biotechnol 30(4):615–621. https://doi.org/10.4014/jmb.1910.10061
Litwińska K, Bischoff F, Matthes F, Bode R, Rutten T, Kunze G (2019) Characterization of recombinant laccase from Trametes versicolor synthesized by Arxula adeninivorans and its application in the degradation of pharmaceuticals. AMB Expr 9:102. https://doi.org/10.1186/s13568-019-0832-3
Liu Z, Zhang D, Hua Z, Li J, Du G, Chen J (2010) Improvement of laccase production and its properties by lowenergy ion implantation. Bioprocess Biocyst Eng 33:639–646. https://doi.org/10.1007/s00449-009-0389-7
Liu X, Gillespie M, Ozel AD, Dikici E, Daunert S, Bachas LG (2011) Electrochemical properties and temperature dependence of a recombinant laccase from Thermus thermophilus. Anal Bioanal Chem 399:361–366. https://doi.org/10.1007/s00216-010-4345-9
Liu H, Cheng Y, Du B, Tong C, Liang S, Han S (2015) Overexpression of a novel thermostable and chloride-tolerant laccase from Thermus thermophilus SG0.5JP17-16 in Pichia pastoris and its application in synthetic dye decolorization. PLoS One 10(3):e0119833. https://doi.org/10.1371/journal.pone.0119833
Lorenzo M, Moldes D, Sanromán MA (2006) Effect of heavy metals on the production of several laccase isoenzymes by Trametes versicolor and on their ability to decolourise dyes. Chemosphere 63:912–917. https://doi.org/10.1016/j.chemosphere.2005.09.046
Mathews SL, Smithson CE, Grunden AM (2016) Purification and characterization of a recombinant laccase-like multi-copper oxidase from Paenibacillus glucanolyticus SLM1. J Appl Microbiol :1364–5072. https://doi.org/10.1111/jam.13241
Mehandia S, Sharma SC, Kumar Arya S (2019) Isolation and characterization of an alkali and thermostable laccase from a novel Alcaligenes faecalis and its application in decolorization of synthetic dyes. Biotechnol Rep 25:e00413. https://doi.org/10.1016/j.btre.2019.e00413
Mohorcic M, Teodorovic S, Golob V, Friedrich J (2006) Fungal and enzymatic decolourisation of artificial textile dye baths. Chemosphere 63:1709–1717. https://doi.org/10.1016/j.chemosphere.2005.09.063
Nakamura K, Go N (2005) Function and molecular evolution of multicopper blue proteins. Cell Mol Life Sci 62:2050–2066. https://doi.org/10.1007/s00018-004-5076-x
Rodgers CJ, Blanford CF, Giddens SR, Skamnioti P, Armstrong FA, Gurr SJ (2010) Designer laccases: a vogue for high-potential fungal enzymes. Trends Biotechnol 28: 63–72. https://doi.org/10.1016/j.tibtech.2009.11.001
Rodrigues CSD, Madeira LM, Boaventura RAR (2009) Treatment of textile effluent by chemical (Fenton’s Reagent) and biological (sequencing batch reactor) oxidation. J Hazard Mater 172:1551–1559. https://doi.org/10.1016/j.jhazmat.2009.08.027
Sadhasivam S, Savitha S, Swaminathan K, Lin FH (2008) Production purification and characterization of mid redox potential laccase from a newly isolated Trichoderma harzianum WL. Process Biochem 43:736–774. https://doi.org/10.1016/j.procbio.2008.02.017
Shafiei M, Afzali F, Karkhane AA, Ebrahimi SM, Haghbeen K, Aminzadeh S (2019) Cohnella sp. A01 laccase: thermostable, detergent resistant, anti-environmental and industrial pollutants enzyme. Heliyon 5:e02543. https://doi.org/10.1016/j.heliyon.2019.e02543
Uthandi S, Saad B, Humbard MA, Maupin-Furlow JA (2010) LccA, an archaeal laccase secreted as a highly stable glycoprotein into the extracellular medium by Haloferax volcanii. Appl Environ Microbiol 76:733–743. https://doi.org/10.1128/AEM.01757-09
Wikee S, Hatton J, Turbé-Doan A, Mathieu Y, Daou M, Lomascolo A, Kumar A, Lumyong S, Sciara G, Faulds CB, Record E (2019) Characterization and dye decolorization potential of two laccases from the marine-derived fungus Pestalotiopsis sp. Int J Mol Sci 20:1864. https://doi.org/10.3390/ijms20081864
Xu F (1997) Effects of redox potential and hydroxide inhibition on the pH activity profile of fungal laccases. J Biol Chem 72(2):924–928. https://doi.org/10.1074/jbc.272.2.924
Zhang Q, Miao R, Liu T, Huang Z, Peng W. Gan B, Zhang X, Tan H (2019) Biochemical characterization of a key laccase–like multicopper oxidase of artificially cultivable Morchella importuna provides insights into plant–litter decomposition. 3 Biotech 9:171. https://doi.org/10.1007/s13205-019-1688-6
Zhou W, Zhang W, Cai Y (2021) Laccase immobilization for water purification: a comprehensive review. Chem Eng J 403:126272. https://doi.org/10.1016/j.cej.2020.126272
Zhu M, Zhang G, Meng L, Wang H, Gao K, Ng T (2016) Purification and characterization of a white laccase with pronounced dye decolorizing ability and HIV-1 reverse transcriptase inhibitory activity from Lepista nuda. Molecules 21:415. https://doi.org/10.3390/molecules21040415
Acknowledgements
This study was funded by TUBITAK project number 218Z032. Thank you for the institutional contributions of TUBITAK. We would also like to thank Assistant Mustafa Ozkan BALTACI for their support during primer design.
Funding
This study was funded by TUBITAK project number 218Z032.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation and analysis were performed by Arzu Öztürk Kesebir. The first draft of the manuscript was written by Arzu Öztürk Kesebir, Prof. Dr. Ömer İrfan Küfrevioğlu, and Prof. Dr. Melda Sisecioglu and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
Not applicable
Consent for publication
Not applicable
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kesebir, A.Ö., Kılıç, D., Şişecioğlu, M. et al. Recombinant laccase production from Bacillus licheniformis O12: Characterization and its application for dye decolorization. Biologia 76, 3429–3438 (2021). https://doi.org/10.1007/s11756-021-00847-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-021-00847-1