Abstract
Background
In recent years, studies have shown that exposure to environmental pollutants (e.g., radiation, heavy metal substances, air pollutants, organic pollutants) is a leading cause of human non-communicable diseases. The key to disease prevention is to clarify the harmful mechanisms and toxic effects of environmental pollutants on the body. Metabolomics is a high-sensitivity, high-throughput omics technology that can obtain detailed metabolite information of an organism. It is a crucial tool for gaining a comprehensive understanding of the pathway network regulation mechanism of the organism. Its application is widespread in many research fields such as environmental exposure assessment, medicine, systems biology, and biomarker discovery.
Aim of review
Recent findings show that metabolomics can be used to obtain molecular snapshots of organisms after environmental exposure, to help understand the interaction between environmental exposure and organisms, and to identify potential biomarkers and biological mechanisms.
Key scientific concepts of review
This review focuses on the application of metabolomics to understand the biological effects of radiation, heavy metals, air pollution, and persistent organic pollutants exposure, and examines some potential biomarkers and toxicity mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Environmental exposure is a complicated and insidious threat, and there is extensive scientific research to prevent, diagnose, and treat its negative effects. As a tool to detect and analyze metabolites produced by cells and discover markers, metabolomics can rapidly promote the development of environmental exposure research. The review combined metabolomics and environmental pollutant exposure keywords to conduct comprehensive searches in CNKI, CBM, NCBI PubMed, and Google Scholar web scientific databases. Based on the research direction of the research group, many high-level reviews, experimental papers, and theoretical articles were accessed. Scientific research published from 2012 to 2022 on radiation, heavy metals, PM2.5, and persistent organic pollutants exposure factors was considered.
1 Metabolomics is viewed as a functional readout of biological samples
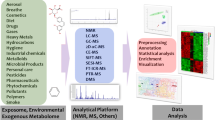
Metabolomics is an emerging experimental tool that has received attention in examining the biological effects of environmental contaminants on animals, plants, and microorganisms (Alseekh et al., 2021). Depending on the specific research objective and/or purposes, the proper metabolomics analysis methods can deal with the associated environmental exposure with less effort than traditional methods. The tools for predicting and evaluating radiation damage are mainly classical cytogenetic experiments, such as micronucleus test and two-point chromosome test (Blumenthal et al., 2014; Dainiak et al., 2019). Metabolomics strategies can be mainly divided into complementary untargeted metabolomics and targeted metabolomics, which are commonly used simultaneously or sequentially (Fig. 1) (González-Peña et al., 2019; Di Minno et al., 2021). Untargeted metabolomics is an exploratory analysis aimed at conducting a comprehensive qualitative and quantitative assessment of the dynamic alterations of the metabolome to get diverse metabolic characteristics of the control and experimental groups, and it is generally used to identify or screen new metabolic biomarkers. In contrast, the goal of targeted metabolomics is to conduct quantitative analysis or verification of specific metabolites (such as potential biomarkers) (Lopes et al., 2017). Generally, targeted metabolomics methods require higher analytical accuracy to accomplish complete quantitative accuracy. The development of metabolomics requires analytical technical support, and thus nuclear magnetic resonance (NMR) and mass spectrometry (MS) have become the main methods for metabolomics research (González-Peña et al., 2019). The former is a rapid, non-destructive separation technique that can measure the chemical and physical properties of molecules and detect metabolites that are difficult to ionize or derivate (Belhaj et al., 2021; Jacob et al., 2019; Nagana et al., 2017). The latter combines diverse chromatographic separation techniques, including gas chromatography-mass spectrometry (GC–MS), liquid chromatography-mass spectrometry (LC–MS), and capillary electrophoresis-mass spectrometry (CE-MS) with higher sensitivity and greater selectivity. GC–MS can specifically distinguish and detect volatile and hydrophilic low-molecular-weight metabolites (Alseekh et al., 2021; Misra, 2021). LC–MS is a practical metabolomic tool. It can detect metabolites with higher molecular weights and more complex chemical structures with suitable repeatability and high specificity (Beale et al., 2018; Pautova et al., 2021). The emerging ultra-high performance liquid chromatography-quadrupole time-of-flight mass spectrometry (UPLC-Q-TOF–MS) has present advantages in capturing metabolite information accurately and rapidly (Zhou et al., 2018). The CE-MS separation technique that relies on the charge-to-size ratio is a powerful tool for analyzing charged metabolites and polar samples (Mever et al., 2019). Although the efficiency and sensitivity of CE-MS are higher than that of GC–MS and LC–MS, this method cannot be applied universally because of its poor reproducibility and technical challenges (Zhang et al., 2021b). Low molecular weight metabolites such as amino acids, nucleic acids, organic acids, and lipids in biological samples can be captured and quantified by this series of technology platforms, which can enhance the utilization of biological samples and obtain the most comprehensive cellular metabolism information (Antcliffe & Gordon, 2016; Au, 2018; Samczuk et al., 2018).
1 Identify the research subjects impacted by environmental exposure; 2 Collect readily accessible samples including plasma, serum, urine, saliva, tissue fluid, etc.; 3 Comprehensively identify metabolites by untargeted metabolomics; 4 Screen meaningful biomarkers; 5 Targeted metabolomics for qualitative and quantitative validation of biomarkers.
2 Metabolomics contributes to an improved understanding of the mechanisms of action of different environmental stimuli
Presently, metabolomics is an advanced and promising omic technology. It can be used to obtain abundant amounts of information on the metabolic state of cells and provide vital data for studying biological effects, which has significant advantages in environmental monitoring, disease detection, diagnosis, treatment, and drug development (Alonso et al., 2015; Klupczyńska et al., 2015; Lee et al., 2019). Similarly, metabolomics has become a sensitive tool for measuring environmental exposures and biological responses, helping to comprehend the mechanism of action of specific environmental factors on the body, as well as diseases (Belhaj et al., 2021; Pernot et al., 2012). Metabolomics research aims to describe the effects of endogenous and exogenous factors on the metabolic characteristics of the body and provides a new methodological framework for environmental exposure research (Pham et al., 2021; Tang et al., 2021). Numerous studies using metabolomics approaches have found that environmental exposures (e.g., physical radiation, air pollutants, heavy metals, and organic pollutants) stimulate metabolic disturbances and induce chronic disease. (Fig. 2). As human exposure data, including data on pollutants, lifestyle factors, and behavior, are often limited or inadequate, the estimation of chronic disease risk is severely hampered. Metabolomics can characterize and quantify small molecules in complex biological samples such as blood, urine, or tissue, describe the dynamic metabolic responses of living systems after environmental exposure stimuli, and trace related biomarkers and metabolic mechanisms (Kyrtopoulos et al., 2013; Rinschen et al., 2019).
3 Metabolomics plays an important role in radiation exposure research
Radiation exposure is a typical environmental exposure. Long-term radiation exposure will produce complex biological effects, causing severe damage to the body (Pan et al., 2021; Singh et al., 2021). Numerous studies have shown that radiation exposure can damage the genetic material of cells and impact the metabolic activities within cells. It is necessary to extract markers of radiation damage from biological samples to help identify exposed individuals and detect both early and delayed systemic and tissue-specific damage (Menon et al., 2016; Reisz et al., 2014). Radiation metabolomics provides meaningful interpretations of changes in radiation stress response metabolites and elucidates the effects of environmental radiation on human physiological states. It determines potential biomarkers and metabolic pathways and helps identify treatments for radiation exposure injury (Fig. 3; Table 1) (Hu et al., 2012; Pannkuk et al., 2017; Satyamitra et al., 2020).
Radiation exposure induces ionization of water molecules in cells, up-regulation of ROS levels in cells, enhanced oxidative stress response, disturbance of metabolic functions, and DNA damage, which ultimately lead to apoptosis.
Receiving different types and doses of radiation can result in various adverse biological effects. Exposure to high doses of radiation will generate heavier unfavorable effects and pose a severe threat to organisms (Chen et al., 2021c). Jelonek and Pernot found that radiation damage is dependent on the dose and time of radiation exposure, and metabolomics can be used to screen the characteristics of metabolic molecules in different periods to obtain time-dose-related biomarkers (Jelonek et al., 2017; Pernot et al., 2012). When plasma lipid metabolism in mice was studied via γ-ray radiation, it was found that radiation can induce manifest changes in lipid metabolism and affect the process of lipid-mediated signal transduction (Ghosh et al., 2013). Epidemiological studies have shown that exposure of the chest or the whole body to ionizing radiation can induce morbidity and mortality of cardiovascular diseases (Azimzadeh et al., 2017; Jones et al., 2019). Potential biomarkers calcium citrate and citrine were identified in the urine of mice exposed to different dose rates of cesium and strontium radiation in vitro and in vivo, suggesting that both the source of exposure and the radiation dose rate had a significant effect on the metabolic activity of the mice (Goudarzi et al., 2016). Radiation can cause significant changes in DNA damage metabolic pathways in the body's tissues, interfere with the body's normal metabolic activities, and cause potential threats to the body (Ghaleb et al., 2022; Kumar et al., 2021). As blood cell metabolites, 2′ -deoxycytidine and choline can be used as effective markers of radiation exposure to evaluate exposure dose during accidental radiation events (Goudarzi et al., 2014). Although numerous meaningful biomarkers have been verified in recent studies, many challenges remain in applying them to medicine. For instance, it is difficult to find a single accurate biomarker with massive metabolite information. Additionally, verification analyses are required under different research conditions and should be executed in the future to follow up on radiation exposure damage research.
4 The potential and usefulness of metabolomics approaches have been widely demonstrated in the heavy metal exposure field
Heavy metals can affect cell activity and lead to cell apoptosis or death. Long-term exposure can lead to acute or chronic tissue, organ damage, and even carcinogenesis (Kosakivska et al., 2021; Wang et al., 2021). Metabolomics can be used to evaluate the toxicity of heavy metals and develop new toxicity biomarkers through models constructed in the laboratory, which creates a new vision and approach for heavy metal exposure research (Table 2) (García-Sevillano et al., 2015; Rodríguez-Moro et al., 2021). For non-occupationally exposed populations, diet is the main source of heavy metal exposure. (Hu et al., 2021; Suomi et al., 2021). In humans, cadmium exposure can result in a variety of adverse effects, such as renal and hepatic dysfunction, damage to the adrenals and hemopoietic system, and it can also cause neurotoxicity (Balali-Mood et al., 2021; Satarug, 2018; Tinkov et al., 2018). For example, using UPLC-Q-TOF–MS to discover the dose-dependent toxicity of CdCl2 to PC-12 cells from a metabolomic perspective, and to identify key metabolic pathways and potential biomarkers of cadmium exposure (Zong et al., 2018). Similarly, mouse urine studies based on UPLC-MS technology have also identified dozens of potential biomarkers of cadmium exposure, such as guanidinosuccinic acid and phenylacetylglycine, which induce renal oxidative stress and interfere with amino acid metabolism, fatty acid metabolism, and energy metabolism (Chen et al., 2018; Zhang et al., 2019). Arsenic is a toxic heavy metal pollutant, and exposure can trigger metabolic disorders, immune dysfunction, oxidative stress, and even cancer (Chen et al., 2019; Ramsey et al., 2013; Xu et al., 2019; Yu et al., 2016). There are shreds of evidence that arsenic exposure has toxic effects on various organ tissues (liver, kidney, and the male reproductive system) (Guvvala et al., 2017; Li et al., 2017; Liu et al., 2020; Silva et al., 2017). LC–MS-based metabolomic studies found that GC-1spg and PC12 cells exposed to arsenic trioxide (ATO) induced metabolic disturbances, enhanced oxidative stress, cell membrane disruption, mitochondrial dysfunction, and autophagy (Chen et al., 2020; Qi et al., 2021). A study on arsenic exposure and male infertility obtained carnitine, estrone, and LysoPC (10:0) a series of potential biomarkers involved in oxidative stress, and lipid metabolism, and may serve as an assessment marker for arsenic-induced male dysfunction (Wu et al., 2021b).
Heavy metal mercury is ubiquitous in the environment. Mercury exposure may increase the expression of heat shock proteins and oxidative phosphorylation genes, promoting detoxification and energy metabolism (Jiang et al., 2021b). In earthworms, low-dose mercury exposure can cause energy metabolism disorders, amino acid metabolism disorders, and osmotic pressure changes, resulting in toxic effects (Tang et al., 2018). Chromium is a metal that is highly toxic to organisms. Accumulation of hexavalent chromium in the human body will inhibit growth, affect components of the antioxidant system, and lead to DNA breakage and irreversible changes in chromosomes (Gutiérrez-Corona et al., 2016; Ventura et al., 2021; Zhao et al., 2019b). The heavy metal manganese is a neurotoxic heavy metal substance. Baker (2017) used metabolomics technology to explore the biomarkers in urine under manganese exposure, helping to better understand the diagnosis and prognosis of manganese exposure (Baker et al., 2017; Zhong et al., 2021). These experimental studies have found that metabolomics methods can provide convincible evidence for understanding the toxic effects of heavy metal pollutants on organisms.
5 Metabolomics has been increasingly applied to the study of PM2.5 exposure damage to the body
The main harmful component of air pollution, particulate matter 2.5 (PM2.5), is considered by the World Health Organization to be a key indicator in evaluating the impact of air pollution on human health (Vo et al., 2020). Human epidemiological studies and controlled animal studies indicated that PM exposure may increase oxidative stress and metabolite changes, leading to various adverse health consequences (e.g., cardiovascular disease, respiratory disease, cancer, central nervous system disease, and adverse pregnancy outcomes) (Malley et al., 2017; Peixoto et al., 2017; Schraufnagel et al., 2019a, 2019b; Turner et al., 2016). Moreover, the severe metabolic disorder caused by PM2.5 exposure can reduce the immune capacity of the body and result in other poisonous substances having toxic effects on the body (Bernatsky et al., 2016; Geng et al., 2021). Human epidemiological studies and untargeted metabolomics assessments have found that the mature mechanism of PM2.5 exposure is the differential response of metabolic pathways associated with oxidative stress and inflammation (Table 3) (Costa et al., 2017; Jin et al., 2021).
Metabolomics is applied to investigate the negative effects of PM2.5 exposure on the metabolic activities of organisms (Liu et al., 2021a). Different human blood metabolomics studies have found that the metabolic pathways of glycerophospholipids, sphingolipids and glutathione are severely disrupted after long-term and short-term exposure to PM2.5, and phospholipid catabolic metabolism is an important pathway of exposure injury. LysoPC (p-20:0) and LysoPC (p-18:1 (9z)) are the potentially effective biomarkers (Chu et al., 2021; Nassan et al., 2021). Further studies in human lung fibroblasts (HEL 299) found that PM2.5 could lead to the reduction of mitochondria-related metabolites, trigger the production of intracellular reactive oxygen species and mitochondrial dysfunction, and induce a high level of apoptosis (Shon et al., 2020). Additionally, GC and LC- MS were used to evaluate the changes in an acute exposure mouse model, and significant changes were found in amino acid metabolism, lipid metabolism, and glucose metabolism in urine. Similar results were obtained using NMR (Du et al., 2020; Zhang et al., 2018). Wang et al. established the time-response relationship between PM2.5 and liver toxicity in mice using LC–MS metabolomics, using different biomarkers to distinguish different stages of liver damage. 4-Pyridine acid and succinate are markers of mild oxidative stress in the liver, proline can indicate severe oxidative stress and inflammation in the liver, and KYNA is a biomarker of significant oxidative damage and inflammation in the liver (Wang et al., 2021c). Exposure to environmental PM2.5 during pregnancy or newborn period can also produce strong neurotoxic effects. For mice in early stages of pregnancy, exposure to PM2.5 can stimulate the dopamine pathway in the brain, inhibit the glutamate pathway, and increase the risk of diseases in the offspring (Church et al., 2018; Cui et al., 2019). Additionally, studies have shown that when mice are sub-chronically exposed to high concentrations of PM2.5, the richness and composition of intestinal and pulmonary microbiota are significantly reduced, leading to glucose metabolism disorder, and five reliable differential metabolites (glutamate, glutamine, formic acid, pyruvate, and lactic acid) are obtained (Ran et al., 2021; Wang et al., 2018). Consequently, metabolomics can be a novel and promising method for assessing PM2.5 damage and elucidating its mechanism of toxicity.
6 Metabolomics analysis can provide scientific tools for exploring toxicity and risk assessment of POPs
Persistent organic pollutants (POPs) are the most common endogenous and exogenous exposures. Because they are highly resistant to metabolic degradation and readily accumulate in adipose tissue, long-term exposure can lead to neurodegenerative diseases, inflammation, hepatotoxicity, nephrotoxicity, insulin resistance, allergy, metabolic diseases, and carcinogenesis (Ranjbar et al., 2020; Żwierełło et al., 2020). The compounds 2,3,7,8-tetrachlorodibenzo-p-dioxins (TCDD), 2,3,7,8-tetrachlorodibenzo-furan (TCDF), and polychlorinated biphenyls (PCBs) are typical environmental pollutants as well as aryl hydrocarbon receptor (AHR) agonists (AHR activation regulates intestinal microbial community structure and function) (Chen et al., 2021b; Stedtfeld et al., 2017; Zhang et al., 2015a). Presently, many laboratories use metabolomics studies to reveal and explain exposure-related metabolic disturbances and their risks to chronic diseases, providing a scientific basis for elucidating the toxicity mechanisms of environmental pollutant exposure (Table 4) (You et al., 2022). The environmental pollutant TCDD can cause cancer, reproductive and developmental issues, damage to the immune system, and interfere with the endocrine system (Gaspari et al., 2021; Maqbool et al., 2016; Patrizi B et al., 2018). Untargeted metabolomics was used to analyze inflammatory lipid metabolites in mouse liver, serum, and urine to elucidate the association between TCDD exposure and hepatotoxicity. The results suggest that TCDD can activate the AHR and induce the release of pro-inflammatory factors, which may lead to the development of steatohepatitis (Doskey et al., 2020). A study on acute exposure to TCDD in mice showed that acute exposure to TCDD caused dysregulation of extracellular signal-regulated protein kinases, metabolic disturbances of biologically active lipids, amino acids, etc., resulting in various organ damage and dysfunction (Dopkins et al., 2021). TCDF is the most typical environmental pollutant of the dioxins. When mice were exposed to TCDF, 1H-NMR analysis of their metabolism revealed that gluconeogenesis and glycogenolysis were inhibited, which stimulated the synthesis of liver fat and induced inflammatory reaction (Yuan et al., 2020). Exposure to TCDF caused intestinal microflora disorder, with increased production of lipopolysaccharide and glutamate resulting in intestinal inflammation (Nichols et al., 2019; Zhang et al., 2015b). Of PCBs, 3,3′,4,4′,5-pentachlorobiphenyl (PCB126) is the most toxic. Long-term exposure to low-dose PCB126 can cause an accumulation of fatty acids (e.g., palmitic acid, palmitoleic acid, and linoleic acid) in heart tissue, leading to cardiac hypertrophy. The increase in collagen synthase and extracellular matrix protein induced by glycine and threonine up-regulation indicates cardiac fibrosis (Wang et al., 2021a). Polycyclic aromatic hydrocarbons (PAHs) are recognized as carcinogens. Recent epidemiological studies have demonstrated links between cardiovascular disease, diabetes, and neurodegenerative diseases (Lu et al., 2021; Wu et al., 2021a). In human and animal experiments, PAHs have been found to cause oxidative stress and injury, protein biosynthesis disorders, organ dysfunction, and metabolic disorders (Gao et al., 2018). LC–MS-based untargeted metabolomics was used to analyze urine samples from children and adolescents from industrially polluted areas to explore the metabolic pathways of pollutants associated with liver dysfunction and chronic kidney disease. The study obtained a series of biomarkers of lysophosphatidylcholine (LPCs), phosphatidylcholine (PCs), and sphingomyelin to clarify their biological mechanism (Chen et al., 2021a). The organic pollutant 2,2,4,4′ -tetrabromodiphenyl ether (BDE-47) can cause obesity disease. When mice are exposed to BDE-47 through a high-fat diet, lipid metabolism disorders occur, and the accumulation of saturated fatty acids and triglycerides in fat induces inflammation and obesity (Yang et al., 2021a). In a separate study, a mouse model of BDE-47 exposure showed that BDE-47 mainly affected glycerophospholipid metabolism in the brain, and potential biomarkers were obtained, including phosphatidylcholine, lysophosphatidylcholine, sphingomyelin, biogenic amines, and various amino acids (Li et al., 2020a). Additionally, POPs can produce adverse biological effects. Triphenyl phosphate (TPP) can affect glycolysis, citric acid cycling, oxidative phosphorylation, and lipid and protein metabolism pathways; induce apoptosis of normal liver cells (L02); damage cell ultrastructure; and increase reactive oxygen species (Wang et al., 2020). Triclocarban (TCC) is an endocrine disruptor widely present in nursing products, which can inhibit the activity of human soluble cyclooxyhydrolase involved in the regulation of blood pressure and inflammation and is a potential cause of colon diseases (Zhang et al., 2020). Zhang et al. used untargeted metabolomics to study the effects of Triclocarban exposure on the liver and gut microbiota in mice. This study found that Triclocarban exposure caused metabolic disturbances in the liver microenvironment and enhanced oxidative stress in mice. At the same time, Triclocarban derivatives interfered with the composition of gut microbes and induced chronic diseases such as colitis. (Li et al., 2018; Zhang et al., 2021a). These studies revealed the mechanism of POPs toxicity to the body and screened a series of meaningful biomarkers. Existing research and data prove that metabolomics is an effective means to find environmental exposure disease-related biomarkers to elucidate the mechanism of action.
7 The potential for efficient and sensitive analysis of metabolomics is a boon for scientific research
Metabolomics aims to comprehensively monitor the level of metabolite changes induced by endogenous and exogenous factors in living systems, screen significantly differential metabolites, and obtain potential biomarkers (Wang et al., 2021b). It has been widely used in disease diagnosis, nutrition, new drug research and development, drug toxicity assessment, environmental toxicology, systems biology, and other research fields (Fu et al., 2022; Steuer et al., 2019; Tran et al., 2020). Metabolomics-based toxicity studies of environmental pollutants can provide new insights into the impact of environmental exposures on human health and enhance the study of pathogenic mechanisms of environmental pollutant exposures. We reviewed biomarkers and pathogenesis associated with exposure to various environmental pollutants from a metabolomic perspective. The findings enhanced our understanding of the biological effects of exposure to environmental pollutants. However, there are still many gaps in the research on metabolic disorders and signaling disorders caused by environmental pollutants exposure, which cannot fully meet our needs for disease prevention and treatment. Most metabolomics studies are still in the stage of biomarker discovery using a large amount of sample data, lacking targeted validation of potential biomarkers and exploration of metabolic pathways. Metabolomics can be combined with advanced sequencing technologies to screen and validate biomarkers associated with specific environmental exposures and diseases, providing new ideas and insights for exploring pathogenic mechanisms.
Data availability
The data and materials in this article are online and publicly available without request.
References
Alonso, A., Marsal, S., & Julià, A. (2015). Analytical methods in untargeted metabolomics: State of the art in 2015. Frontiers Bioengineering Biotechnology, 3, 23.
Alseekh, S., Aharoni, A., Brotman, Y., Contrepois, K., D’Auria, J., Ewald, J., Fraser, P. D., Giavalisco, P., Hall, R. D., Heinemann, M., Link, H., Luo, J., Neumann, S., Nielsen, J., Perez de Souza, L., Saito, K., Sauer, U., Schroeder, F. C., & Fernie, A. R. (2021). Mass spectrometry-based metabolomics: a guide for annotation, quantification and best reporting practices. Nature Methods., 18(7), 747–756.
Antcliffe, D., & Gordon, A. C. (2016). Metabonomics and intensive care. Critical Care, 20, 68.
Au, A. (2018). Metabolomics and lipidomics of ischemic stroke. Advances in Clinical Chemistry, 85, 31–69.
Azimzadeh, O., & Tapio, S. (2017). Proteomics landscape of radiation-induced cardiovascular disease: Somewhere over the paradigm. Expert Review of Proteomics, 14(11), 987–996.
Baker, M. G., Simpson, C. D., Lin, Y. S., Shireman, L. M., & Seixas, N. (2017). The use of metabolomics to identify biological signatures of manganese exposure. Annals of Work Exposures and Health, 61(4), 406–415.
Balali-Mood, M., Naseri, K., Tahergorabi, Z., Khazdair, M. R., & Sadeghi, M. (2021). Toxic mechanisms of five heavy metals: Mercury, lead, chromium, cadmium, and arsenic. Frontiers in Pharmacology, 12, 643972.
Beale, D. J., Pinu, F. R., Kouremenos, K. A., Poojary, M. M., Narayana, V. K., Boughton, B. A., Kanojia, K., Dayalan, S., Jones, O. A. H., & Dias, D. A. (2018). Review of recent developments in GC-MS approaches to metabolomics-based research. Metabolomics, 14(11), 152.
Belhaj, M. R., Lawler, N. G., & Hoffman, N. J. (2021). Metabolomics and lipidomics: Expanding the molecular landscape of exercise biology. Metabolites, 11(3), 151.
Bernatsky, S., Smargiassi, A., Barnabe, C., Svenson, L. W., Brand, A., Martin, R. V., Hudson, M., Clarke, A. E., Fortin, P. R., van Donkelaar, A., Edworthy, S., Bélisle, P., & Joseph, L. (2016). Fine particulate air pollution and systemic autoimmune rheumatic disease in two Canadian provinces. Environmental Research, 146, 85–91.
Blumenthal, D. J., Sugarman, S. L., Christensen, D. M., Wiley, A. L., Livingston, G. K., Glassman, E. S., Koerner, J. F., Sullivan, J. M., & Hinds, S. (2014). Role of dicentric analysis in an overarching biodosimetry strategy for use following a nuclear detonation in an urban environment. Health Physics, 106(4), 516–522.
Chen, C. S., Kuo, T. C., Kuo, H. C., Tseng, Y. J., Kuo, C. H., Yuan, T. H., & Chan, C. C. (2021a). Lipidomics of children and adolescents exposed to multiple industrial pollutants. Environmental Research, 201, 111448.
Chen, H., Liu, G., Qiao, N., Kang, Z., Hu, L., Liao, J., Yang, F., Pang, C., Liu, B., Zeng, Q., Li, Y., & Li, Y. (2020). Toxic effects of arsenic trioxide on spermatogonia are associated with oxidative stress, mitochondrial dysfunction, autophagy and metabolomic alterations. Ecotoxicology and Environmental Safety, 190, 110063.
Chen, J., Yang, Y., Russu, W. A., & Chan, W. K. (2021b). The aryl hydrocarbon receptor undergoes chaperone-mediated autophagy in triple-negative breast cancer cells. International Journal of Molecular Sciences, 22(4), 1654.
Chen, L., Song, D., Zhang, W., Zhang, C., & Zhang, L. (2019). The dynamic changes of arsenic bioaccumulation and antioxidant responses in the marine medaka Oryzias melastigma during chronic exposure. Aquat Toxicology., 212, 110–119.
Chen, S., Zhang, M., Bo, L., Li, S., Hu, L., Zhao, X., & Sun, C. (2018). Metabolomic analysis of the toxic effect of chronic exposure of cadmium on rat urine. Environmental Science and Pollution Research International, 25(4), 3765–3774.
Chen, Y., Cui, J., Gong, Y., Wei, S., Wei, Y., & Yi, L. (2021c). MicroRNA: A novel implication for damage and protection against ionizing radiation. Environmental Science and Pollution Research International, 28(13), 15584–15596.
Chi, L., Tu, P., Liu, C. W., Lai, Y., Xue, J., Ru, H., & Lu, K. (2019). Chronic arsenic exposure induces oxidative stress and perturbs serum lysolipids and fecal unsaturated fatty acid metabolism. Chemical Research in Toxicology, 32(6), 1204–1211.
Chu, H., Huang, F. Q., Yuan, Q., Fan, Y., Xin, J., Du, M., Wang, M., Zhang, Z., & Ma, G. (2021). Metabolomics identifying biomarkers of PM2.5 exposure for vulnerable population: Based on a prospective cohort study. Environmental Science Pollution Research International, 28(12), 14586–14596.
Church, J. S., Tijerina, P. B., Emerson, F. J., Coburn, M. A., Blum, J. L., Zelikoff, J. T., & Schwartzer, J. J. (2018). Perinatal exposure to concentrated ambient particulates results in autism-like behavioral deficits in adult mice. Neurotoxicology, 65, 231–240.
Costa, L. G., Cole, T. B., Coburn, J., Chang, Y. C., Dao, K., & Roqué, P. J. (2017). Neurotoxicity of traffic-related air pollution. Neurotoxicology, 59, 133–139.
Cui, J., Fu, Y., Lu, R., Bi, Y., Zhang, L., Zhang, C., Aschner, M., Li, X., & Chen, R. (2019). Metabolomics analysis explores the rescue to neurobehavioral disorder induced by maternal PM2.5 exposure in mice. Ecotoxicology and Environmental Safety, 169, 687–695.
Dainiak, N., Albanese, J., Kaushik, M., Balajee, A. S., Romanyukha, A., Sharp, T. J., & Blakely, W. F. (2019). Concepts of operations for a US dosimetry and biodosimetry network. Radiation Protection Dosimetry, 186(1), 130–138.
Di Minno, A., Gelzo, M., Stornaiuolo, M., Ruoppolo, M., & Castaldo, G. (2021). The evolving landscape of untargeted metabolomics. Nutrition, Metabolism, and Cardiovascular Diseases, 31(6), 1645–1652.
Dopkins, N., Neameh, W. H., Hall, A., Lai, Y., Rutkovsky, A., Gandy, A. O., Lu, K., Nagarkatti, P. S., & Nagarkatti, M. (2021). Effects of acute 2,3,7,8-tetrachlorodibenzo-p-dioxin exposure on the circulating and cecal metabolome profile. International Journal of Molecular Sciences, 22(21), 11801.
Doskey, C. M., Fader, K. A., Nault, R., Lydic, T., Matthews, J., Potter, D., Sharratt, B., Williams, K., & Zacharewski, T. (2020). 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) alters hepatic polyunsaturated fatty acid metabolism and eicosanoid biosynthesis in female Sprague-Dawley rats. Toxicology and Applied Pharmacology, 398, 115034.
Du, X., Zeng, X., Pan, K., Zhang, J., Song, L., Zhou, J., Chen, R., Xie, Y., Sun, Q., Zhao, J., & Kan, H. (2020). Science of the Total Environment, 714, 136790.
Fu, J., Zhang, L. L., Li, W., Zhang, Y., Zhang, Y., Liu, F., & Zou, L. (2022). Application of metabolomics for revealing the interventional effects of functional foods on metabolic diseases. Food Chemistry, 367, 130697.
Gao, P., da Silva, E., Hou, L., Denslow, N. D., Xiang, P., & Ma, L. Q. (2018). Human exposure to polycyclic aromatic hydrocarbons: Metabolomics perspective. Environment International, 119, 466–477.
García-Sevillano, M. Á., García-Barrera, T., & Gómez-Ariza, J. L. (2015). Environmental metabolomics: Biological markers for metal toxicity. Electrophoresis, 36(18), 2348–2365.
Gaskins, A. J., Tang, Z., Hood, R. B., Ford, J., Schwartz, J. D., Jones, D. P., Laden, F., Liang, D., EARTH Study Team. (2021). Periconception air pollution, metabolomic biomarkers, and fertility among women undergoing assisted reproduction. Environ Int., 155, 106666.
Gaspari, L., Paris, F., Kalfa, N., Soyer-Gobillard, M. O., Sultan, C., & Hamamah, S. (2021). Experimental Evidence of 2,3,7,8-tetrachlordibenzo-p-dioxin (TCDD) transgenerational Effects on reproductive health. International Journal of Molecular Sciences, 22(16), 9091.
Geng, N., Ren, X., Gong, Y., Zhang, H., Wang, F., Xing, L., Cao, R., Xu, J., Gao, Y., Giesy, J. P., & Chen, J. (2019). Integration of metabolomics and transcriptomics reveals short-chain chlorinated paraffin-induced hepatotoxicity in male Sprague-Dawley rat. Environment International, 133(Pt B), 105231.
Geng, N., Song, X., Cao, R., Luo, Y., & A M, Cai Z, Yu K, Gao Y, Ni Y, Zhang H, Chen J. (2021). The effect of toxic components on metabolomic response of male SD rats exposed to fine particulate matter. Environmental Pollution, 272, 115922.
Ghaleb, A., Roa, L., & Marchenko, N. (2022). Low-dose but not high-dose γ-irradiation elicits the dominant-negative effect of mutant p53 in vivo. Cancer Letters, 530, 128–141.
Ghosh, S. P., Singh, R., Chakraborty, K., Kulkarni, S., Uppal, A., Luo, Y., Kaur, P., Pathak, R., Kumar, K. S., Hauer-Jensen, M., & Cheema, A. K. (2013). Metabolomic changes in gastrointestinal tissues after whole body radiation in a murine model. Molecular BioSystems, 9(4), 723–731.
González-Peña, D., & Brennan, L. (2019). Recent advances in the application of metabolomics for nutrition and health. Annual Review of Food Science and Technology, 10, 479–519.
Goudarzi, M., Chauthe, S., Strawn, S. J., Weber, W. M., Brenner, D. J., & Fornace, A. J. (2016). Quantitative metabolomic analysis of urinary citrulline and calcitroic acid in mice after exposure to various types of ionizing radiation. International Journal of Molecular Sciences, 17(5), 782.
Goudarzi, M., Mak, T. D., Chen, C., Smilenov, L. B., Brenner, D. J., & Fornace, A. J. (2014). The effect of low dose rate on metabolomic response to radiation in mice. Radiation and Environmental Biophysics, 53(4), 645–657.
Gutiérrez-Corona, J. F., Romo-Rodríguez, P., Santos-Escobar, F., Espino-Saldaña, A. E., & Hernández-Escoto, H. (2016). Microbial interactions with chromium: Basic biological processes and applications in environmental biotechnology. World Journal of Microbiology & Biotechnology, 32(12), 191.
Guvvala, P. R., Ravindra, J. P., Rajani, C. V., Sivaram, M., & Selvaraju, S. (2017). Protective role of epigallocatechin-3-gallate on arsenic induced testicular toxicity in Swiss albino mice. Biomedicine & Pharmacotherapy, 96, 685–694.
Hu, Y., Wang, C., Song, Z., Chen, M., Ding, L., Liang, X., Bi, X., Li, Z., Li, P., & Zheng, W. (2021). Heavy metal in rice and vegetable and human exposure near a large Pb/Zn smelter in central China. International Journal of Environmental Research and Public Health, 18(23), 12631.
Hu, Z. P., Kim, Y. M., Sowa, M. B., Robinson, R. J., Gao, X., Metz, T. O., Morgan, W. F., & Zhang, Q. (2012). Metabolomic response of human skin tissue to low dose ionizing radiation. Molecular BioSystems, 8(7), 1979–1986.
Jacob, M., Lopata, A. L., Dasouki, M., & Abdel Rahman, A. M. (2019). Metabolomics toward personalized medicine. Mass Spectrometry Reviews, 38(3), 221–238.
Jelonek, K., Pietrowska, M., & Widlak, P. (2017). Systemic effects of ionizing radiation at the proteome and metabolome levels in the blood of cancer patients treated with radiotherapy: The influence of inflammation and radiation toxicity. International Journal of Radiation Biology, 93(7), 683–696.
Jia, C., Wei, Y., Lan, Y., Hou, X., Zuo, J., Wang, T., Li, J., Guan, X., Yang, H., & Mao, G. (2019). Comprehensive analysis of the metabolomic characteristics on the health lesions induced by chronic arsenic exposure: A metabolomics study. International Journal of Hygiene and Environmental Health, 222(3), 434–445.
Jiang, L., Hong, Y., Xie, G., Zhang, J., Zhang, H., & Cai, Z. (2021a). Comprehensive multi-omics approaches reveal the hepatotoxic mechanism of perfluorohexanoic acid (PFHxA) in mice. Science of the Total Environment, 790, 148160.
Jiang, W., Fang, J., Du, M., Gao, Y., Fang, J., & Jiang, Z. (2021b). Integrated transcriptomics and metabolomics analyses reveal benzo[a]pyrene enhances the toxicity of mercury to the Manila clam ruditapes philippinarum. Ecotoxicology Environment Safety, 213, 112038.
Jin, L., Godri Pollitt, K. J., Liew, Z., Rosen Vollmar, A. K., Vasiliou, V., Johnson, C. H., & Zhang, Y. (2021). Use of untargeted metabolomics to explore the air pollution-related disease continuum. Current Environment Health Reports, 8(1), 7–22.
Jones, J. W., Alloush, J., Sellamuthu, R., Chua, H. L., MacVittie, T. J., Orschell, C. M., & Kane, M. A. (2019). Effect of sex on biomarker response in a mouse model of the hematopoietic acute radiation syndrome. Health Physics, 16(4), 484–502.
Klupczyńska, A., Dereziński, P., & Kokot, Z. J. (2015). Metabolomics in medical sciences–trends, challenges and perspectives. Acta Poloniae Pharmaceutica, 72(4), 629–641.
Kosakivska, I. V., Babenko, L. M., Romanenko, K. O., Korotka, I. Y., & Potters, G. (2021). Molecular mechanisms of plant adaptive responses to heavy metals stress. Cell Biology International, 45(2), 258–272.
Kumar, P., Wang, P., Farese, A. M., MacVittie, T. J., & Kane, M. A. (2021). Metabolomics of multiorgan radiation injury in non-human primate model reveals system-wide metabolic perturbations. Health Physics, 121(4), 395–405.
Kyrtopoulos, S. A. (2013). Making sense of OMICS data in population-based environmental health studies. Environmental and Molecular Mutagenesis, 54(7), 468–479.
Laiakis, E. C., Mak, T. D., Strawn, S. J., Wang, Y. W., Moon, B. H., Ake, P., & Fornace, A. J., Jr. (2018). Global metabolomic responses in urine from atm deficient mice in response to LD50/30 gamma irradiation doses. Environmental and Molecular Mutagenesis, 59(7), 576–585.
Laiakis, E. C., Pannkuk, E. L., Chauthe, S. K., Wang, Y. W., Lian, M., Mak, T. D., Barker, C. A., Astarita, G., & Fornace, A. J., Jr. (2017). A serum small molecule biosignature of radiation exposure from total body irradiated patients. Journal of Proteome Research, 16(10), 3805–3815.
Lee, M. Y., & Hu, T. (2019). Computational methods for the discovery of metabolic markers of complex traits. Metabolites, 9(4), 66.
Li, J., Hu, Y., Liu, L., Wang, Q., Zeng, J., & Chen, C. (2020a). PM25 exposure perturbs lung microbiome and its metabolic profile in mice. Science Total Environment, 721, 137432.
Li, S. W., Sun, X., He, Y., Guo, Y., Zhao, H. J., Hou, Z. J., & Xing, M. W. (2017). Assessment of arsenic trioxide in the heart of Gallus gallus: Alterations of oxidative damage parameters, inflammatory cytokines, and cardiac enzymes. Environmental Science and Pollution Research International, 24(6), 5781–5790.
Li, W., Zhang, W., Chang, M., Ren, J., Xie, W., Chen, H., Zhang, Z., Zhuang, X., Shen, G., & Li, H. (2018). Metabolomics reveals that triclocarban affects liver metabolism by affecting glucose metabolism, β-oxidation of fatty acids, and the TCA cycle in male mice. Toxicology Letters, 299, 76–85.
Li, X., Brejnrod, A. D., Ernst, M., Rykær, M., Herschend, J., Olsen, N. M. C., Dorrestein, P. C., Rensing, C., & Sørensen, S. J. (2019). Heavy metal exposure causes changes in the metabolic health-associated gut microbiome and metabolites. Environment International, 126, 454–467.
Li, Y., Yu, N., Li, M., Li, K., Shi, W., Yu, H., & Wei, S. (2020b). Metabolomic insights into the lasting impacts of early-life exposure to BDE-47 in mice. Environmental Pollution, 263(Pt B), 114524.
Liang, Y., Tang, Z., Jiang, Y., Ai, C., Peng, J., Liu, Y., Chen, J., Zhang, J., & Cai, Z. (2020). Serum metabolic changes associated with dioxin exposure in a Chinese male cohort. Environment International, 143, 105984.
Liu, F., Chen, X., Liu, Y., Niu, Z., Tang, H., Mao, S., Li, N., Chen, G., & Xiang, H. (2021a). Serum cardiovascular-related metabolites disturbance exposed to different heavy metal exposure scenarios. Journal of Hazardous Materials, 415, 125590.
Liu, H., Li, H., Zhang, X., Gong, X., Han, D., Zhang, H., Tian, X., & Xu, Y. (2021b). Metabolomics comparison of metabolites and functional pathways in the gills of Chlamys farreri under cadmium exposure. Environmental Toxicology and Pharmacology, 86, 103683.
Liu, P., Xue, Y., Zheng, B., Liang, Y., Zhang, J., Shi, J., Chu, X., Han, X., & Chu, L. (2020). Crocetin attenuates the oxidative stress, inflammation and apoptosisin arsenic trioxide-induced nephrotoxic rats: Implication of PI3K/AKT pathway. International Immunopharmacology, 88, 106959.
Locci, E., Lecca, L. I., Piras, R., Noto, A., Pilia, I., d’Aloja, E., & Campagna, M. (2019). Urinary 1H NMR metabolomics profile of Italian citizens exposed to background levels of arsenic: A (pre)cautionary tale. Biomarkers, 24(8), 727–734.
Long, C., Hu, G., Zheng, P., Chen, T., Su, Z., Zhang, Y., Ding, C., Peng, F., Yu, S., Wang, T., & Jia, G. (2021). Analysis of serum metabolome of workers occupationally exposed to hexavalent chromium: A preliminary study. Toxicology Letters, 349, 92–100.
Lopes, A. S., Cruz, E. C., Sussulini, A., & Klassen, A. (2017). Metabolomic strategies involving mass spectrometry combined with liquid and gas chromatography. Advances in Experimental Medicine and Biology, 965, 77–98.
Lu, X., Lin, Y., Qiu, X., Liu, J., Zhu, T., Araujo, J. A., Zhang, J., & Zhu, Y. (2021). Metabolomic changes after subacute exposure to polycyclic aromatic hydrocarbons: A natural experiment among healthy travelers from los angeles to Beijing. Environmental Science and Technology, 55(8), 5097–5105.
Maan, K., Tyagi, R., Dutta, A., Bakhshi, R., & Rana, P. (2020). Comparative metabolic profiles of total and partial body radiation exposure in mice using an untargeted metabolomics approach. Metabolomics, 16(12), 124.
Malley, C. S., Kuylenstierna, J. C., Vallack, H. W., Henze, D. K., Blencowe, H., & Ashmore, M. R. (2017). Preterm birth associated with maternal fine particulate matter exposure: A global, regional and national assessment. Environment International, 101, 173–182.
Maqbool, F., Mostafalou, S., Bahadar, H., & Abdollahi, M. (2016). Review of endocrine disorders associated with environmental toxicants and possible involved mechanisms. Life Sciences, 145, 265–273.
Menon, S. S., Uppal, M., Randhawa, S., Cheema, M. S., Aghdam, N., Usala, R. L., Ghosh, S. P., Cheema, A. K., & Dritschilo, A. (2016). Radiation metabolomics: Current status and future directions. Frontiers in Oncology, 6, 20.
Mever, M., Hankemeier, T., & Ramautar, R. (2019). CE-MS for anionic metabolic profiling: An overview of methodological developments. Electrophoresis, 40(18–19), 2349–2359.
Misra, B. B. (2021). Advances in high resolution GC-MS technology: A focus on the application of GC-Orbitrap-MS in metabolomics and exposomics for FAIR practices. Analytical Methods, 13(20), 2265–2282.
Nagana Gowda, G. A., & Raftery, D. (2017). Recent advances in NMR-based metabolomics. Analytical Chemistry, 89(1), 490–510.
Nassan, F. L., Wang, C., Kelly, R. S., Lasky-Su, J. A., Vokonas, P. S., Koutrakis, P., & Schwartz, J. D. (2021). Ambient PM2.5 species and ultrafine particle exposure and their differential metabolomic signatures. Environ Int., 151, 106447.
Nichols, R. G., Zhang, J., Cai, J., Murray, I. A., Koo, I., Smith, P. B., Perdew, G. H., & Patterson, A. D. (2019). Metatranscriptomic analysis of the mouse gut microbiome response to the persistent organic pollutant 2,3,7,8-tetrachlorodibenzofuran. Metabolites, 10(1), 1.
Pan, Y., Lei, X., & Zhang, Y. (2021). Association predictions of genomics, proteinomics, transcriptomics, microbiome, metabolomics, pathomics, radiomics, drug, symptoms, environment factor, and disease networks: A comprehensive approach. Medicinal Research Reviews. https://doi.org/10.1002/med.21847
Pannkuk, E. L., Fornace, A. J., Jr., & Laiakis, E. C. (2017). Metabolomic applications in radiation biodosimetry: Exploring radiation effects through small molecules. International Journal of Radiation Biology, 93(10), 1151–1176.
Pannkuk, E. L., Laiakis, E. C., Mak, T. D., Astarita, G., Authier, S., Wong, K., & Fornace, A. J., Jr. (2016). A lipidomic and metabolomic serum signature from nonhuman primates exposed to ionizing radiation. Metabolomics, 12(5), 80.
Patrizi, B., & Siciliani de Cumis, M. (2018). TCDD toxicity mediated by epigenetic mechanisms. International Journal of Molecular Sciences, 19(12), 4101.
Pautova, A., Burnakova, N., & Revelsky, A. (2021). Metabolic profiling and quantitative analysis of cerebrospinal fluid using gas chromatography-mass spectrometry: Current methods and future perspectives. Molecules, 26(12), 3597.
Peixoto, M. S., de Oliveira Galvão, M. F., & Batistuzzo de Medeiros, S. R. (2017). Cell death pathways of particulate matter toxicity [published correction appears in Chemosphere. 2017 Nov 21]. Chemosphere, 188, 32–48.
Pernot, E., Hall, J., Baatout, S., Benotmane, M. A., Blanchardon, E., Bouffler, S., El Saghire, H., Gomolka, M., Guertler, A., Harms-Ringdahl, M., Jeggo, P., Kreuzer, M., Laurier, D., Lindholm, C., Mkacher, R., Quintens, R., Rothkamm, K., Sabatier, L., Tapio, S., … Cardis, E. (2012). Ionizing radiation biomarkers for potential use in epidemiological studies. Mutation Research, 751(2), 258–286.
Pham, Y. L., & Beauchamp, J. (2021). Breath biomarkers in diagnostic applications. Molecules, 26(18), 5514.
Qi, Z., Wang, Q., Wang, H., & Tan, M. (2021). Metallothionein attenuated arsenic-induced cytotoxicity: The underlying mechanism reflected by metabolomics and lipidomics. Journal of Agriculture and Food Chemistry, 69(18), 5372–5380.
Ramsey, K. A., Bosco, A., McKenna, K. L., Carter, K. W., Elliot, J. G., Berry, L. J., Sly, P. D., Larcombe, A. N., & Zosky, G. R. (2013). In utero exposure to arsenic alters lung development and genes related to immune and mucociliary function in mice. Environmental Health Perspectives, 121(2), 244–250.
Ran, Z., An, Y., Zhou, J., Yang, J., Zhang, Y., Yang, J., Wang, L., Li, X., Lu, D., Zhong, J., Song, H., Qin, X., & Li, R. (2021). Subchronic exposure to concentrated ambient PM2.5 perturbs gut and lung microbiota as well as metabolic profiles in mice. Environment Pollution, 272, 115987.
Ranjbar Jafarabadi, A., Mashjoor, S., Riyahi Bakhtiari, A., & Jadot, C. (2020). Dietary intake of polycyclic aromatic hydrocarbons (PAHs) from coral reef fish in the Persian gulf - human health risk assessment. Food Chemistry, 329, 127035.
Reisz, J. A., Bansal, N., Qian, J., Zhao, W., & Furdui, C. M. (2014). Effects of ionizing radiation on biological molecules–mechanisms of damage and emerging methods of detection. Antioxidants & Redox Signaling, 21(2), 260–292.
Rinschen, M. M., Ivanisevic, J., Giera, M., & Siuzdak, G. (2019). Identification of bioactive metabolites using activity metabolomics. Nature Reviews Molecular Cell Biology, 20(6), 353–367.
Rodríguez-Moro, G., Ramírez-Acosta, S., Callejón-Leblic, B., Arias-Borrego, A., García-Barrera, T., & Gómez-Ariza, J. L. (2021). Environmental metal toxicity assessment by the combined application of metallomics and metabolomics. Environmental Science and Pollution Research International. https://doi.org/10.1007/s11356-021-13507-3
Samczuk, P., Ciborowski, M., & Kretowski, A. (2018). Application of metabolomics to study effects of bariatric surgery. Journal of Diabetes Research, 2018, 6270875.
Satarug, S. (2018). Dietary cadmium intake and its effects on kidneys. Toxics., 6(1), 15.
Satyamitra, M. M., Cassatt, D. R., Hollingsworth, B. A., Price, P. W., Rios, C. I., Taliaferro, L. P., Winters, T. A., & DiCarlo, A. L. (2020). Metabolomics in radiation biodosimetry: Current approaches and advances. Metabolites, 10(8), 328.
Schraufnagel, D. E., Balmes, J. R., Cowl, C. T., De Matteis, S., Jung, S. H., Mortimer, K., Perez-Padilla, R., Rice, M. B., Riojas-Rodriguez, H., Sood, A., Thurston, G. D., To, T., Vanker, A., & Wuebbles, D. J. (2019a). Air pollution and noncommunicable diseases: A review by the forum of International respiratory societies’ environmental committee, Part 1: The damaging effects of air pollution. Chest, 155(2), 409–416.
Schraufnagel, D. E., Balmes, J. R., Cowl, C. T., De Matteis, S., Jung, S. H., Mortimer, K., Perez-Padilla, R., Rice, M. B., Riojas-Rodriguez, H., Sood, A., Thurston, G. D., To, T., Vanker, A., & Wuebbles, D. J. (2019b). Air pollution and noncommunicable diseases: A review by the forum of International respiratory societies’ environmental committee, part 2: Air pollution and organ systems. Chest., 155(2), 417–426.
Shon, J. C., Lee, S. M., Jung, J. H., Wu, Z., Kwon, Y. S., Sim, H. J., & Seo, J. S. (2020). Integrated metabolomics and lipidomics reveals high accumulation of polyunsaturated lysoglycerophospholipids in human lung fibroblasts exposed to fine particulate matter. Ecotoxicology and Environmental Safety, 202, 110896.
Silva, R. F., Borges, C. D. S., de Almeida, L. C., Cagnon, V. H. A., & de Grava, K. W. (2017). Arsenic trioxide exposure impairs testicular morphology in adult male mice and consequent fetus viability. Journal of Toxicology and Environmental Health. Part A, 80(19–21), 1166–1179.
Singh, V. K., Seed, T. M., & Cheema, A. K. (2021). Metabolomics-based predictive biomarkers of radiation injury and countermeasure efficacy: Current status and future perspectives. Expert Review of Molecular Diagnostics, 21(7), 641–654.
Stedtfeld, R. D., Stedtfeld, T. M., Fader, K. A., Williams, M. R., Bhaduri, P., Quensen, J., Zacharewski, T. R., Tiedje, J. M., & Hashsham, S. A. (2017). TCDD influences reservoir of antibiotic resistance genes in murine gut microbiome. FEMS Microbiology Ecology, 93(5), fix058.
Steuer, A. E., Brockbals, L., & Kraemer, T. (2019). Metabolomic strategies in biomarker research-new approach for indirect identification of drug consumption and sample manipulation in clinical and forensic toxicology? Frontiers in Chemistry, 7, 319.
Suomi, J., Valsta, L., & Tuominen, P. (2021). Dietary heavy metal exposure among finnish adults in 2007 and in 2012. International Journal of Environmental Research and Public Health, 18(20), 10581.
Tang, R., Ding, C., Dang, F., Ma, Y., Wang, J., Zhang, T., & Wang, X. (2018). NMR-based metabolic toxicity of low-level Hg exposure to earthworms. Environmental Pollution, 239, 428–437.
Tang, S., Li, T., Fang, J., Chen, R., Cha, Y., Wang, Y., Zhu, M., Zhang, Y., Chen, Y., Du, Y., Yu, T., Thompson, D. C., Godri Pollitt, K. J., Vasiliou, V., Ji, J. S., Kan, H., Zhang, J. J., & Shi, X. (2021). The exposome in practice: An exploratory panel study of biomarkers of air pollutant exposure in Chinese people aged 60–69 years (China BAPE Study). Environment International, 157, 106866.
Tinkov, A. A., Filippini, T., Ajsuvakova, O. P., Skalnaya, M. G., Aaseth, J., Bjørklund, G., Gatiatulina, E. R., Popova, E. V., Nemereshina, O. N., Huang, P. T., Vinceti, M., & Skalny, A. V. (2018). Cadmium and atherosclerosis: A review of toxicological mechanisms and a meta-analysis of epidemiologic studies. Environmental Research, 162, 240–260.
Tran, H., McConville, M., & Loukopoulos, P. (2020). Metabolomics in the study of spontaneous animal diseases. Journal of Veterinary Diagnostic Investigation, 32(5), 635–647.
Turner, M. C., Jerrett, M., Pope, C. A., 3rd., Krewski, D., Gapstur, S. M., Diver, W. R., Beckerman, B. S., Marshall, J. D., Su, J., Crouse, D. L., & Burnett, R. T. (2016). Long-term ozone exposure and mortality in a large prospective study. American Journal of Respiratory and Critical Care Medicine, 193(10), 1134–1142.
Tyagi, R., Maan, K., Khushu, S., & Rana, P. (2020). Urine metabolomics based prediction model approach for radiation exposure. Science and Reports, 10(1), 16063.
Upadhyay, M., Rajagopal, M., Gill, K., Li, Y., Bansal, S., Sridharan, V., Tyburski, J. B., Boerma, M., & Cheema, A. K. (2020). Identification of plasma lipidome changes associated with low dose space-type radiation exposure in a murine model. Metabolites, 10(6), 252.
Valcke, M., Ouellet, N., Dubé, M., Laouan Sidi, E. A., LeBlanc, A., Normandin, L., Balion, C., & Ayotte, P. (2019). Biomarkers of cadmium, lead and mercury exposure in relation with early biomarkers of renal dysfunction and diabetes: Results from a pilot study among aging Canadians. Toxicology Letters, 312, 148–156.
Ventura, C., Gomes, B. C., Oberemm, A., Louro, H., Huuskonen, P., Mustieles, V., Fernández, M. F., Ndaw, S., Mengelers, M., Luijten, M., Gundacker, C., & Silva, M. J. (2021). Biomarkers of effect as determined in human biomonitoring studies on hexavalent chromium and cadmium in the period 2008–2020. Environmental Research, 197, 110998.
Vo, T. T. T., Wu, C. Z., & Lee, I. T. (2020). Potential effects of noxious chemical-containing fine particulate matter on oral health through reactive oxygen species-mediated oxidative stress: Promising clues. Biochemical Pharmacology, 182, 114286.
Wang, C., Cui, R., Niu, C., Zhong, X., Zhu, Q., Ji, D., Li, X., Zhang, H., Liu, C., Zhou, L., Li, Y., Xu, G., & Wei, Y. (2021a). Low-dose PCB126 exposure disrupts cardiac metabolism and causes hypertrophy and fibrosis in mice. Environ Pollut., 290, 118079.
Wang, M., Xu, J., Zhang, Y., Yang, N., Ge, W., & Song, R. (2021b). Integrated multiplatform-based metabolomics and network analysis to explore the mechanism of Polygonum cuspidatum on hyperlipidemia. Journal of Chromatography. B, Analytical Technologies in the Biomedical and Life Sciences, 1176, 122769.
Wang, R., Han, X., Pang, H., Hu, Z., & Shi, C. (2021c). Illuminating a time-response mechanism in mice liver after PM2.5 exposure using metabolomics analysis. Science Total Environment, 767, 144485.
Wang, W., Zhou, J., Chen, M., Huang, X., Xie, X., Li, W., Cao, Q., Kan, H., Xu, Y., & Ying, Z. (2018). Exposure to concentrated ambient PM2.5 alters the composition of gut microbiota in a murine model. Part Fibre Toxicology, 15(1), 17.
Wang, X., Li, F., Liu, J., Ji, C., & Wu, H. (2020). Transcriptomic, proteomic and metabolomic profiling unravel the mechanisms of hepatotoxicity pathway induced by triphenyl phosphate (TPP). Ecotoxicology and Environmental Safety, 205, 111126.
Wu, F., Chi, L., Ru, H., Parvez, F., Slavkovich, V., Eunus, M., Ahmed, A., Islam, T., Rakibuz-Zaman, M., Hasan, R., Sarwar, G., Graziano, J. H., Ahsan, H., Lu, K., & Chen, Y. (2018). Arsenic exposure from drinking water and urinary metabolomics: Associations and long-term reproducibility in Bangladesh adults. Environmental Health Perspectives, 126(1), 017005.
Wu, S., Chen, Z., Yang, L., Zhang, Y., Luo, X., Guo, J., & Shao, Y. (2021a). Particle-bound PAHs induced glucose metabolism disorders through HIF-1 pathway. Science of the Total Environment, 797, 149132.
Wu, Y., Ding, R., Zhang, X., Zhang, J., Huang, Q., Liu, L., & Shen, H. (2021b). Meet-in-metabolite analysis: A novel strategy to identify connections between arsenic exposure and male infertility. Environment International, 147, 106360.
Xu, L., Lu, Z., Ji, C., Cong, M., Li, F., Shan, X., & Wu, H. (2019). Toxicological effects of As (V) in juvenile rockfish Sebastes schlegelii by a combined metabolomic and proteomic approach. Environmental Pollution, 255(Pt 2), 113333.
Xu, Y., Liu, H., Han, D., Ren, L., Gong, X., Jiang, F., Cui, Y., Liu, X., Ren, C., Xue, J., & Tian, X. (2021). Metabolomic alterations in the digestive system of the mantis shrimp oratosquilla oratoria following short-term exposure to cadmium. Frontiers in Physiology, 12, 706579.
Yan, Q., Liew, Z., Uppal, K., Cui, X., Ling, C., Heck, J. E., von Ehrenstein, O. S., Wu, J., Walker, D. I., Jones, D. P., & Ritz, B. (2019). Maternal serum metabolome and traffic-related air pollution exposure in pregnancy. Environment International, 130, 104872.
Yang, C., Wei, J., Cao, G., & Cai, Z. (2021a). Lipid metabolism dysfunction and toxicity of BDE-47 exposure in white adipose tissue revealed by the integration of lipidomics and metabolomics. Science Total Environment, 806(Pt 1), 150350.
Yang, J., Chen, W., Sun, Y., Liu, J., & Zhang, W. (2021b). Effects of cadmium on organ function, gut microbiota and its metabolomics profile in adolescent rats. Ecotoxicology and Environmental Safety, 222, 112501.
Yang, L., Liu, Y., Cui, Z., Zhang, Y., Zhang, J., & Lian, K. (2021c). Metabolomic mechanisms of short chain chlorinated paraffins toxicity in rats. Environmental Research, 197, 111060.
You, L., Zheng, F., Su, C., Wang, L., Li, X., Chen, Q., Kou, J., Wang, X., Wang, Y., Wang, Y., Mei, S., Zhang, B., Liu, X., & Xu, G. (2022). Metabolome-wide association study of serum exogenous chemical residues in a cohort with 5 major chronic diseases. Environment International, 158, 106919.
Yu, D., Ji, C., Zhao, J., & Wu, H. (2016). Proteomic and metabolomic analysis on the toxicological effects of As (III) and As (V) in juvenile mussel Mytilus galloprovincialis. Chemosphere, 150, 194–201.
Yuan, P., Dong, M., Lei, H., Xu, G., Chen, G., Song, Y., Ma, J., Cheng, L., & Zhang, L. (2020). Targeted metabolomics reveals that 2,3,7,8-tetrachlorodibenzofuran exposure induces hepatic steatosis in male mice. Environmental Pollution, 259, 113820.
Zhan, J., Wang, S., Li, F., Ji, C., & Wu, H. (2021). Dose-dependent responses of metabolism and tissue injuries in clam Ruditapes philippinarum after subchronic exposure to cadmium. Science of the Total Environment, 779, 146479.
Zhang, H., Liang, Y., Wu, P., Shi, X., Zhang, G., & Cai, Z. (2021a). Continuous dermal exposure to triclocarban perturbs the homeostasis of liver-gut axis in mice: Insights from metabolic interactions and microbiome shifts. Environmental Science and Technology, 55(8), 5117–5127.
Zhang, H., Lu, Y., Liang, Y., Jiang, L., & Cai, Z. (2020). Triclocarban-induced responses of endogenous and xenobiotic metabolism in human hepatic cells: Toxicity assessment based on nontargeted metabolomics approach. Journal of Hazardous Materials, 392, 122475.
Zhang, J., Wen, X., Li, Y., Zhang, J., Li, X., Qian, C., Tian, Y., Ling, R., & Duan, Y. (2021b). Diagnostic approach to thyroid cancer based on amino acid metabolomics in saliva by ultra-performance liquid chromatography with high resolution mass spectrometry. Talanta, 235, 122729.
Zhang, L., Hatzakis, E., Nichols, R. G., Hao, R., Correll, J., Smith, P. B., Chiaro, C. R., Perdew, G. H., & Patterson, A. D. (2015a). Metabolomics reveals that aryl hydrocarbon receptor activation by environmental chemicals induces systemic metabolic dysfunction in mice. Environmental Science and Technology, 49(13), 8067–8077.
Zhang, L., Nichols, R. G., Correll, J., Murray, I. A., Tanaka, N., Smith, P. B., Hubbard, T. D., Sebastian, A., Albert, I., Hatzakis, E., Gonzalez, F. J., Perdew, G. H., & Patterson, A. D. (2015b). Persistent organic pollutants modify gut microbiota-host metabolic homeostasis in mice through aryl hydrocarbon receptor activation. Environmental Health Perspectives, 123(7), 679–688.
Zhang, M., Jia, S., Liu, Y., Liu, Y., Li, S., Bo, L., Zhao, X., & Sun, C. (2019). Metabonomics analysis of kidneys in rats administered with chronic low-dose cadmium by ultra-performance liquid chromatography-mass spectrometry. Journal of Applied Toxicology, 39(3), 441–450.
Zhang, P., Zhu, W., Wang, D., Yan, J., Wang, Y., Zhou, Z., & He, L. (2017). A combined NMR- and HPLC-MS/MS-based metabolomics to evaluate the metabolic perturbations and subacute toxic effects of endosulfan on mice. Environmental Science and Pollution Research International, 24(23), 18870–18880.
Zhang, X., Zhang, J., Wu, Y., Nan, B., Huang, Q., Du, X., Tian, M., Liu, L., Xin, Y., Li, Y., Duan, J., Chen, R., Sun, Z., & Shen, H. (2021c). Dynamic recovery after acute single fine particulate matter exposure in male mice: Effect on lipid deregulation and cardiovascular alterations. Journal of Hazardous Materials, 414, 125504.
Zhang, Y., Li, Y., Shi, Z., Wu, J., Yang, X., Feng, L., Ren, L., Duan, J., & Sun, Z. (2018). Metabolic impact induced by total, water soluble and insoluble components of PM2.5acute exposure in mice. Chemosphere, 207, 337–346.
Zhao, C., Niu, M., Song, S., Li, J., Su, Z., Wang, Y., Gao, Q., & Wang, H. (2019a). Serum metabolomics analysis of mice that received repeated airway exposure to a water-soluble PM2.5 extract. Ecotoxicology and Environmental Safety, 168, 102–109.
Zhao, H., Xi, C., Tian, M., Lu, X., Cai, T. J., Li, S., Tian, X. L., Gao, L., Liu, H. X., Liu, K. H., & Liu, Q. J. (2020). Identification of potential radiation responsive metabolic biomarkers in plasma of rats exposed to different doses of cobalt-60 gamma rays. Dose Response, 18(4), 1559325820979570.
Zhao, L., Fang, J., Tang, S., Deng, F., Liu, X., Shen, Y., Liu, Y., Kong, F., Du, Y., Cui, L., Shi, W., Wang, Y., Wang, J., Zhang, Y., Dong, X., Gao, Y., Dong, L., Zhou, H., Sun, Q., … Shi, X. (2022). PM2.5 and serum metabolome and insulin resistance, potential mediation by the gut microbiome: A population-based panel study of older adults in China. Environment Health Perspective, 130(2), 27007.
Zhao, M., Lau, K. K., Zhou, X., Wu, J., Yang, J., & Wang, C. (2017). Urinary metabolic signatures and early triage of acute radiation exposure in rat model. Molecular BioSystems, 13(4), 756–766.
Zhao, Y., Zhang, H., Wu, X., Zhang, T., Shen, K., Li, L., Peng, Y., Mehmood, K., & Zhou, D. (2019b). Metabonomic analysis of the hepatic injury suffer from hexavalent chromium poisoning in broilers. Environmental Science and Pollution Research International, 26(18), 18181–18190.
Zhong, G., Wan, F., Wu, S., Jiang, X., Tang, Z., Zhang, X., Huang, R., & Hu, L. (2021). Arsenic or/and antimony induced mitophagy and apoptosis associated with metabolic abnormalities and oxidative stress in the liver of mice. Science of the Total Environment, 777, 146082.
Zhou, J., Ma, H., Wu, Y., Lv, X., Wang, J., Liu, S., Li, D., Wang, H., Yan, Y., Luo, N., Li, Q., Xu, H., Zhang, Q., Yu, L., Guo, H., Kuzmanov, U., Di, L., Wu, Q., & Duan, J. (2019). Lipidomic profiling of subchronic As4S4 exposure identifies inflammatory mediators as sensitive biomarkers in rats. Metallomics, 11(3), 576–585.
Zhou, Y., Men, L., Pi, Z., Wei, M., Song, F., Zhao, C., & Liu, Z. (2018). Fecal metabolomics of type 2 diabetic rats and treatment with Gardenia jasminoides ellis based on mass spectrometry technique. Journal of Agriculture and Food Chemistry, 66(6), 1591–1599.
Zong, L., Xing, J., Liu, S., Liu, Z., & Song, F. (2018). Cell metabolomics reveals the neurotoxicity mechanism of cadmium in PC12 cells. Ecotoxicology and Environmental Safety, 147, 26–33.
Żwierełło, W., Maruszewska, A., Skórka-Majewicz, M., Goschorska, M., Baranowska-Bosiacka, I., Dec, K., Styburski, D., Nowakowska, A., & Gutowska, I. (2020). The influence of polyphenols on metabolic disorders caused by compounds released from plastics - review. Chemosphere, 240, 124901.
Funding
This work was supported by Key Scientific Research Project of Hunan Health Commission (No.202102051816), Hunan Natural Science Foundation (2019JJ40238) and (2019JJ50509), Hunan Health Commission (C2019096), Hunan Provincial Department of Education (10C0479), the project of Hengyang Science and Technology Bureau (2020jh042).
Author information
Authors and Affiliations
Contributions
SW and YW contributed equally to the writing of this article and were major contributors in draft manuscripts. YG, YC, JC, LL, HY, YY, XL, and GL were involved in literature search. LY was involved in conception and instruction of the study and critically revised this manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no other competing interests or conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wei, S., Wei, Y., Gong, Y. et al. Metabolomics as a valid analytical technique in environmental exposure research: application and progress. Metabolomics 18, 35 (2022). https://doi.org/10.1007/s11306-022-01895-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-022-01895-7