Abstract
Introduction
A large scale population exposure to ionizing radiation during intentional or unintentional nuclear accidents undoubtedly generates a complex scenario with partial-body as well as total-body irradiated victims. A high throughput technique based rapid assessment method is an urgent necessity for stratification of exposed subjects independent of whether exposure is uniform total-body or non-homogenous partial-body.
Objective
Here, we used Nuclear Magnetic Resonance (NMR) based metabolomics approach to compare and identify candidate metabolites differentially expressed in total and partially irradiated mice model.
Methods
C57BL/6 male mice (8–10 weeks) were irradiated total-body or locally to thoracic, hind limb or abdominal regions with 10 Gy of gamma radiation. Urine samples collected at 24 h post irradiation were examined using high resolution NMR spectroscopy and the datasets were analysed using multivariate analysis.
Results
Multivariate and metabolic pathway analysis in urine samples collected at 24 h post-radiation exhibited segregation of all irradiated groups from controls. Metabolites associated with energy metabolism, gut flora metabolism and taurine were common to partial and total-body irradiation, thus making them potential candidates for radiation exposure. Nevertheless, a distinct metabolic pattern was observed in partial-body exposed groups with maximum changes observed in the hind limb region indicating differential tissue associated radiation sensitivity. The organ-specific changes may provide an early warning regarding the physiological system at risk after radiation injury.
Conclusion
The study affirms potentiality of metabolite markers and comparative analysis could be an important piece of information for an integrated solution to a complex research question in terms of radiation biomarkers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
With the increasing threat of terrorist activities, global competition for production of nuclear warheads, and establishment of nuclear reactors to meet growing energy demand, man has risked his life to expose to an invisible weapon namely, ionizing radiation (Hall and Giaccia 2018; Pomper and Tarini 2017; Allison 2018). Radiation accidents would eventually cause a mass casualty with heterogeneous exposure due to partial shielding or non-uniform distance from the radiation exposure source resulting in either total or partial-body radiation exposures (Hasegawa et al. 2016; Ozasa et al. 2019). Heterogeneous exposure further complicates the triage and management procedures as the medical need for total-body radiation exposure would be different from partially exposed victims. Therefore, mass screening of exposed population would be required firstly to triage exposed individuals from non-exposed and thereafter to further segregate the total-body exposed and partially exposed individuals. Thus, there is the utmost requirement of high throughput early biomarkers for triage and appropriate medical management of victims. Since the last decade, extensive research has been carried out in rapid biomarkers identification for radiation exposure. With the advancement of omics technologies, efforts have been placed in all fields of omics, proteomics, transcriptomics, miRNAomics and metabolomics to find biomarkers of interest (Gan et al. 2019; Kultova et al. 2020; Małachowska et al. 2020; Singh et al. 2016). However, most of the research has been focused on identifying markers of radiation injury for total-body radiation exposure scenario and few groups have succeeded in identification of few of biomarkers in preclinical levels (Anderson 2019; Lacombe et al. 2018; Lee et al. 2018; Pannkuk et al. 2016; Valente et al. 2015). None the less, estimation of sensitivity and extent of radiation specificity is still a question of research worldwide.
Studies on partial radiation exposure are equally important not only for the improvement in dose assessment but also for the discovery of organ-specific biomarkers of radiation injury. There are some recent studies that have looked upon the importance of biomarkers identification for partial-body irradiation models (Hérodin et al. 2014; Meadows et al. 2010; Sproull et al. 2017; Valente et al. 2015). Radiation responsive plasma proteins in the cohort have been identified to predict radiation exposure with high accuracy (90–93%) in partial-body irradiation model (Sproull et al. 2017). Another study on baboons has shown a clear distinction of total-body irradiation (TBI) from partial-body situation based on haematological and plasma biochemical parameters (Valente et al. 2015). It is agreeable that the response of an organism to radiation injury differs based on the type of exposure (partial or total-body) and radiation dose; it may be anticipated that partial and total-body radiation exposure may have some common signature.
Metabolomics is one such technique that has been exploited to the maximum nowadays for biomarker research as it has close proximity to the phenotype of an individual with least complexity and maximum information. And being lower in the order of hierarchy, it shall be called a global biomarker in a sense as it is the global representative of all the upstream (genetic, transcriptomic, proteomic) changes experienced by an individual. A metabolomic marker, if found in urine, would be minimally invasive, robust, and also likely to be reproducible. Extensive information on metabolomics based urinary markers for radiation exposure up to primate level have been available in the last few years and has been extensively reviewed (Pannkuk et al. 2016, 2017). However, the non-availability of irradiated human subject data, makes candidate radiation marker still a milestone to achieve. To continue further in this area, efforts have been made across the globe to generate extensive information from different radiation exposure scenario like partial, total-body or organ-specific exposure and also to different animal species with an ultimate aim of finding a robust radiation biomarker (Ghandhi et al. 2018; Golla et al. 2017; Hérodin et al. 2014; Kultova et al. 2020; Pannkuk et al. 2017). Due to ease and reproducibility of technique, NMR based metabolomics holds great potential in biomarker discovery in radiation research and some of the earlier studies have attempted in search of radiation responsive biomarkers (Chen et al. 2011; Coy et al. 2011; Emwas et al. 2019; Khan et al. 2011a, b). Our earlier studies have been able to show radiation-induced changes in urine and serum on exposure to whole-body radiation (Khan et al. 2011b). However, to have complete information on radiation biomarkers, it is necessary to know how much similarity or disparity occurs when the same dose of radiation exposure happens after total or partial-body. Therefore, the present study was conducted to look for comparative changes in mouse urine on exposure to total-body radiation or partial radiation to different regions of the body using Nuclear Magnetic Resonance (NMR) metabolomics approach and to have comparative observations if any between partial and total-body radiation exposure.
2 Material and methods
2.1 Chemicals
All chemicals, trimethylsilyl-2,2,3,3-tetra deuteropropanoic acid (TSP), deuterium oxide (D2O), Na2HPO4, NaH2PO4 were obtained from Sigma-Aldrich (St Louis, MO, USA).
2.2 Animal handling and radiation exposure
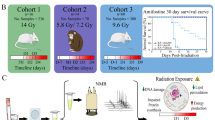
A total of twenty seven ‘C57BL/6’ male mice (8–10 weeks of age) were used in this study. Animals were acclimatized in polypropylene cages for 48 h before group allocation and treatment. During the study, room condition was maintained at 19–23 °C of temperature, 45–65% of humidity and 12 h light/12 h dark cycle. Food and water were provided ad libitum. After acclimatization, animals were randomly allocated to five different groups. Out of five groups, four groups were irradiated using the Tele 60Co irradiation facility (Bhabhatron II). One group was exposed to a single dose of 10 Gy TBI (n = 6) at surface to source distance (SSD) of 80 cm and Field of view (FOV) of 30 × 30 cm2 with a dose rate of 1.096 Gy/min. For partial radiation, three groups of animals were locally irradiated at different regions of the body: abdominal (n = 5), thoracic (n = 5), and hind limb (n = 5), respectively for 10 Gy of single radiation dose at SSD of 80 cm and FOV of 20 × 2 cm2 with a dose rate of 0.66 Gy/min. Animals were partially irradiated in a way that only 20 × 2 cm2 region was exposed during irradiation, rest of the body was lead shielded (Fig. 1). One group was kept as sham control (n = 6). The study was carried out following the guidelines of the institutional animal ethical committee at the Institute of Nuclear Medicine and Allied Sciences (INMAS), DRDO, Delhi, India (8/GO/RBI/S/99/CPCSEA/INM/IAEC/2017/09).
Experimental design for radiation exposure. 10 Gy of 60Co gamma irradiation at dose rate of 1.096 Gy/min with field of view (FOV) of 30 × 30 cm2 in total-body and dose rate of 0.66 Gy/min for 2 × 20 cm2 in partial irradiation groups with source surface distance (SSD) of 80 cm. Blue box represents irradiation area i.e. total-body (TBI), thoracic (TI), abdominal (AI) and hind limb (HI) region
2.3 Urine collection, preparation and NMR experiments
Urine samples were collected in tubes containing 1% sodium azide in cold condition at 24 h from all 4 irradiated and control groups. Following centrifugation, the supernatant was stored at − 80 °C for NMR spectroscopic analysis. Urine samples were centrifuged at 5000 rpm at 4 °C for 10 min. Three hundred fifty microliter of centrifuged urine was diluted with 250 μl of deuterated phosphate buffer (pH 7.4) containing TSP, chemical shift reference. All 1H NMR spectra were acquired using a 600.33 MHz NMR spectrometer (Bruker, Biospin, Switzerland) at 298 K using 1D pulsed sequence along with water saturation (NOESYGPPR1D). A total of 64 scans were collected with a spectral width of 9009 Hz and an acquisition time of 3.63 s and relaxation delay of 4 s was fixed to acquire the spectra.
2.4 Data processing and statistical analysis
Spectral phase, baseline correction and chemical shift referencing to TSP (chemical shift reference) were carried out manually using Topspin3.5.2 (Bruker, Biospin, Switzerland). All processed spectra corresponding to a range of 0.5–9.45 ppm were segmented into consecutive “buckets” (bins) to equal width of 0.02 ppm excluding water (4.61–5.01) and urea (5.61–6.01 ppm) resonance region using AMIX (Bruker, Biospin, Switzerland). Binned data was further analysed using MetaboAnalyst 4.0 (https://www.metaboanalyst.ca). Prior to analysis, data was normalised to total spectral area and pareto-scaling was done to remove inter-sample and sample handling variability. To observe intrinsic metabolic variations, multivariate analysis was commenced with principal component analysis (PCA) that represents a trend and identifies outlier present in the data. Metabolites identified based on chemical shifts information from PCA loading plot using the Human Metabolome Database (HMDB) and the Chenomx NMR Suite software (Chenomx, Edmonton, Canada), were integrated and relative integrals were calculated. Further, to explore clustering behaviour amongst the group and to detect significant metabolites that contribute to variation, Partial Least Squares Discriminant Analysis (PLS-DA) was carried out using MetaboAnalyst 4.0. The model generated was further tested for predictive ability and overfitting using leave-one-out cross validation method and permutation test respectively. Model performance was evaluated from Q2 (predictive ability) and R2 (goodness of fit) whose values if found > 0.5, were considered acceptable.
Univariate analysis was also performed on identified metabolites to evaluate fold change and significant difference between groups using Student’s t-test. Shapiro–Wilk test for normality was performed prior to Student's t-test using IBM SPSS (v 20.2) statistics (http:/www.ibm.com). Further, metabolites with p < 0.05 and q value (FDR) < 0.05 were considered statistically significant. The data of relative intensity of each of the metabolites were expressed as Mean ± Standard deviation (SD). One way analysis of variance (ANOVA) with the Bonferroni post-hoc test was also performed to evaluate significant differences between control and irradiated groups. Statistical significance was considered at p < 0.05. To visualise alterations of identified metabolites in different groups, heatmap was generated using MetaboAnalyst 4.0.
2.5 Metabolic pathway analysis
To evaluate the complex relationships among various metabolites, pathway analysis between irradiated and control groups was performed using MetaboAnalyst 4.0. The significantly perturbed metabolic pathway was screened out on the basis of pathway impact > 0.2 and p-value of < 0.05.
3 Results
This study was performed to determine if total-body and partial-body irradiation (PBI) produce different metabolic profile characteristic of 10 Gy radiation exposure. Also, to determine whether thoracic, abdominal and hind limb irradiation results in unique metabolite change.
3.1 Metabolomic signature and analysis of thoracic region irradiated (TI) group
To determine the distinct signature of radiation exposure for PBI, groups irradiated to different regions were compared to controls independently. PCA analysis showed distinct segregation of TI group from controls without any outlier based on binned data (Supplementary Figs. 1a and 2a). In order to identify distinct metabolic responses to TI, PLS-DA was performed (Supplementary Fig. 2b) that clearly distinguished TI group from control group with an accuracy of 1, R2 of 0.99 and Q2 of 0.95. The metabolites identified through variable importance in projection (VIP) score of > 1, responsible for the classification were citrate, trimethylamine (TMA), trimethylamine N-oxide (TMAO), creatine, taurine and alpha-ketoglutarate (α-KG) (Supplementary Fig. 2c). Univariate analysis of identified metabolites showed up-regulation of two and down-regulation of four out of six identified metabolites in TI group compared to controls (Table 1). Heatmap visualisation also showed altered levels of metabolites in the irradiated group in comparison to controls (Supplementary Fig. 2d).
3.2 Metabolomic signature and analysis of hind limb region irradiated (HI) group
We also identified metabolic profile of HI group (Supplementary Fig. 3). As observed in TI group, HI group was also distinctly clustered away from control group by PCA (Supplementary Fig. 3a) and PLS-DA (Supplementary Fig. 3b) analysis and observed accuracy of 1, R2 of 0.95 and Q2 of 0.89 of PLS-DA model. This group showed maximum changes in metabolic response in PBI groups compared to controls. Univariate analysis showed a total of seventeen significantly altered metabolites out of which, ten metabolites were up-regulated and seven were down-regulated (Table 1). Amongst these significant metabolites, six metabolites were accounted for classification during multivariate analysis with VIP score > 1 (Supplementary Fig. 3c). Taurine, creatine and TMAO represented maximum alteration in their levels in HI group compared to control group. Heatmap also represented an obvious distinction in different metabolites levels in HI and control group (Supplementary Fig. 3d).
3.3 Metabolomic signature and analysis of abdominal region irradiated (AI) group
Similarly, for AI group, multivariate analysis was able to classify irradiated group and control as separate groups with accuracy of 1, Q2 of 0.97 and R2 of 0.87. Ten metabolites were found to be significant by univariate analysis having p-value and FDR < 0.05 (Table 1), out of them, five metabolites had VIP > 1. Besides, TMAO also contributed to classification with VIP > 1 though it was not among significant metabolites (Supplementary Fig. 4).
3.4 Comparative and pooled analysis of PBI group
To identify common and unique metabolic patterns for the localised irradiation, all three PBI groups were together compared with controls. PCA analysis showed clustering of all the three groups whereas, PLS-DA analysis showed separate clusters of all three partially irradiated groups from control. Though some overlapping was observed in TI and HI group, grossly irradiated groups were well segregated. Metabolites held responsible for this classification were mainly citrate, creatine, TMAO, cis-aconitate, α-KG, phosphocholine and allantoin which had a VIP score of > 1 (Supplementary Fig. 5). The classification showed a predictive ability of 0.89 with the goodness of fit of 0.64. Further, all three groups were pooled in a single group as a pooled partial-body irradiated group and analysed with respect to control. Pooled partial irradiated group was also compared with all three different partially irradiated groups using ANOVA and it was observed that dimethylamine (DMA), TMAO and indoxylsulfate were significantly altered in all three partially and pooled partially irradiated groups. Whereas, taurine, TMA, phenylalanine and creatine were observed in two of the three partially irradiated groups along with pooled partially irradiated group (Supplementary Fig. 6).
3.5 Metabolomic signature and analysis of total-body irradiated (TBI) group
There was an obvious distinct segregation of TBI from controls based on PLS-DA. Univariate analysis showed, five metabolites were up-regulated and six metabolites were down-regulated (Table 1) whereas, 8 metabolites contributed to group segregation based on VIP > 1. Changes in each metabolite at individual levels have been well represented in heatmap (Supplementary Fig. 7).
Representative 1H NMR spectrum of urine showing identified metabolites (a) and comparative NMR spectra of control (C) and different irradiated groups (TBI, TI, AI and HI) (b) is shown in Supplementary Fig. 8.
3.6 Common metabolomic signature of radiation exposure
To extend further for any common metabolic signature for radiation exposure (irrespective of partial or total-body irradiation exposure) in urine, data from TBI, pooled partial radiation groups and controls were analysed for all identified metabolites in 1H NMR spectra (Fig. 2). ANOVA based post-hoc comparison represented significant difference in levels of taurine, α-KG, DMA, creatine, phenylalanine, indoxylsulfate, allantoin, TMAO and trans-aconitate in both the irradiated groups compared to controls (Fig. 3). TMA and creatinine are the metabolites of importance for pooled partial radiation as these metabolites represented significant change only in pooled partial radiation group (Fig. 3). However, other important energy metabolites such as citrate and succinate presented a similar trend in both the irradiated groups compared to controls but could not attain significant level in pooled partial radiation group.
Multivariate analysis of pooled partially irradiated and control group a PCA score plot for discriminating control (C), TBI and pooled partially irradiated (PI) groups based on their metabolic profile. Group separation is seen in PC1 with 29%, b PLS-DA score plot after variable selection. Group segregation is seen in component 1 with 37.8% (elliptical boundary shows 95% confidence interval), c VIP plot presenting ranking of metabolites responsible for observed discrimination between the groups. Metabolites having VIP score > 1 are considered for segregation, d Corresponding Heatmap showing comparison of metabolites between the groups. (red = C, control; green = PI, pooled partial-body; blue = TBI, total-body irradiation)
Relative intensity levels of significantly perturbed metabolites using ANOVA followed by Bonferroni post-hoc test with alpha correction in irradiated groups (TBI and Pooled Partial) compared to control group. Data is presented as mean ± standard deviation (*p-value < 0.05, **p-value < 0.01 and ***p-value < 0.001)
3.7 Network visualization of metabolites and pathways
The biological significance of radiation exposure was evaluated by pathway analysis using MetaboAnalyst software. It is essential for understanding the effect of radiation on cellular processes. Pathway analysis revealed alteration in glycine, serine and threonine metabolism, taurine & hypotaurine metabolism, phenylalanine, tyrosine and tryptophan biosynthesis, phenylalanine metabolism and Tri-carboxylic acid (TCA cycle) had good hit with impact > 0.2 and –log(p) < 2 (Fig. 4).
Overview of metabolic pathway analysis discriminating between control and pooled partial body irradiated group, depicting dysregulated metabolic pathways. X axis represents Pathway impact values whereas Y axis represents –log (p) values by pathway enrichment analysis. Glycine, serine and threonine metabolism (creatine, guanidinoacetate, sarcosine, choline, glycine and pyruvate) (p-value = 0.0001), taurine and hypotaurine metabolism (taurine) (p-value = 0.0002), phenylalanine, tyrosine and tryptophan biosynthesis (phenylalanine and hippurate) (p-value = 0.0003), phenylalanine metabolism (phenylalanine) (p-value = 0.0004) and TCA cycle (citrate, α-KG, cis-aconitate, succinate and pyruvate) (p-value = 0.042). Pathways having pathway impact > 0.2 and –log (p) < 2 were considered. Every bubble represents one metabolic pathway and its area is proportional to the impact of pathway and color denotes the significance from highest (red) to lowest (yellow)
4 Discussion
The rise of nuclear or radiological exposure increases the likelihood of partial-body exposure and necessitates the development of appropriate evaluation strategies for nonhomogeneous radiation exposure before treating acute radiation syndrome (Prasanna et al. 2010). Globally, research is going on to identify biomarkers for the identification of nonhomogeneous exposures using in vitro and animal models of partial-body exposure. Still, there is a research gap regarding similarity or dissimilarity coexistence between partial or total-body radiation exposure. Radiation exposure leads to several biological effects, but the mechanism underlying the metabolic effects of radiation are not well known. Pathophysiology involved during partial or total-body radiation further complicates the mechanism. In the present study, we have attempted to elucidate NMR metabolomics based markers common to partial or total-body radiation exposure and further evaluate changes pertaining to specific regions of the body exposed.
NMR based metabolites were evaluated to discriminate between different regional PBI and TBI in mice model for radiation exposure equivalent to 10 Gy. On visual inspection, PCA and PLS-DA plots showed distinct changes in both PBI and TBI groups. Grossly, NMR metabolomics based perturbations were mainly depicted in metabolites associated with energy metabolism, gut flora metabolism and osmolytes in both radiation-exposed groups (Fig. 5). Commonality of metabolite markers in radiation-exposed groups irrespective of partial-body or total-body makes them potential candidates of radiation exposure.
Disturbed energy metabolism is an evident change observed post-radiation exposure. The molecular mechanism of ionizing radiation involves considerable damage to DNA and the indirect reactive oxygen species (ROS) so formed results in cellular stress and death. Mitochondrial DNA is known as sensitive targets to radiation-induced damage that leads to the generation of high levels of ROS (Datta et al. 2012; Kawamura et al. 2018; Yoshida et al. 2012). Therefore, perturbations in TCA intermediates may be a direct indication of mitochondrial dysfunction. Reduction in the energy metabolite (α-KG, citrate, succinate) observed in irradiated groups compared to control suggests mitochondrial dysfunction. The TCA cycle has also been identified as a prominent altered pathway during pathway analysis providing another evidence of mitochondrial dysfunction post-irradiation (Figs. 4 and 5). Our findings of altered TCA intermediates are well in accordance with other recent available literature on identification of metabolomics based biomarkers in different animal models (Chen et al. 2011; Goudarzi et al. 2014; Pannkuk et al. 2017) (Fig. 4).
Energy metabolism associated with metabolic changes observed in our study could be a result of radiation-induced perturbations of metabolic cell signalling pathway. Radiation exposure can initiate a series of complex molecular events involving many signalling pathways including p53 activation that can affect metabolism including mitochondrial biogenesis (Berkers et al. 2013). Amongst the altered energy metabolites, α-KG has been found to be decreased in TI and TBI groups. Decreased α-KG level is suggestive of its increased usage during DNA repair besides an energy substrate intermediate. DNA repair enzymes, Fe2+ binding dioxygenase, are known to be an α-KG dependent enzyme which is involved in removing methyl lesions present in DNA (Bleijlevens et al. 2012).
Another important physiological metabolic system that has been ascertained in our study is gut flora metabolism. The decrease in the urinary level of phenylalanine, indoxylsulfate, TMA, TMAO and DMA in irradiated groups compared to control has been shown to correlate with intestinal micro-biota changes (Fig. 5). Previous studies have established that gut microbial metabolism of choline results in the formation of TMA which is further metabolised by Flavin-monooxygenase (FMO) in host liver to produce TMAO (Hoyles et al. 2018; Wang et al. 2011). A study by Romano et al. 2015 suggests that TMAO level is affected by gut micro-biota composition as some of the bacterial species present in intestinal micro-biota are known to reduce FMO activity. In the present study, reduced level of TMA and TMAO post-irradiation could be due to direct or indirect effect of radiation on microbial diversity. It is evident in literature that high-dose radiation exposure leads to remarkable disturbances in haematopoietic tissue and intestinal mucosa (Ghosh et al. 2013; Shao et al. 2014; Wang et al. 2015). On radiation exposure, intestinal barrier collapses leading to the infiltration of intestinal bacteria into blood circulation. The alteration in the level of TMA and TMAO could be due to transmigration of intestinal bacterial or destruction of probiotic intestinal bacterial flora on radiation exposure. Recent study have shown intestinal cell death along with increased pathogenic proteobacteria on exposure to radiation of more than 10 Gy in rats (Rentea et al. 2016). Also, one of the earlier study have shown that metabolite associated with tryptophan and indole in plasma were found to be associated with radiation-induced gut micro-biota changes in murine model at 24 h after 10 Gy total-body radiation exposure (Kurland et al. 2015). The radiation-induced gut micro-biota changes have been very well supported by altered phenylalanine, tyrosine and tryptophan metabolism by pathway analysis in TBI and PBI group (Fig. 4). The study further observed elevated taurine levels post-irradiation in both the irradiated groups. Elevated level of taurine in urine has consistently been reported in earlier studies on radiation exposure and passive cellular leakage during cell damage or destruction of circulating lymphocytes post-radiation exposure have been proposed as the probable mechanism of release of taurine in urine (Dilley 1972; Fazzino et al. 2006, 2010; Christophersen 2012). The perturbed taurine level has also been substantiated by altered taurine and hypotaurine pathway in the present study (Fig. 4). On a similar note, increased creatine level, a known radiation-induced change in urine, has also been observed in our study on exposure to radiation (Fig. 5).
Concisely, in our study, several metabolite changes were observed in both PBI and TBI groups. Partial-body exposure, similar to total-body exposure, results in decline in blood and immune cell which affect metabolic activities both at cellular and tissue level through signalling. Comparable metabolic changes have been seen in partial or total-body radiation exposure. These familiar changes further affirm the linking of these metabolites associated with radiation injury. However, characteristic behaviour of different irradiated groups could be seen in terms of few metabolites which were disturbed either during TBI and PBI. For instance, citrate and succinate are found to be altered in TBI group only, on the contrary TMA and creatinine were disturbed only in PBI groups. This difference seems obvious due to a distinct level of systemic or localised injury. The impact of partial-body exposure varies with area of body exposed and is expected to be considerably different from a total-body exposure at the identical dose (Prasanna et al. 2010).
Further, distinct degree of injury was also seen amongst PBI groups wherein, most changes were observed in HI irradiated group followed by AI and TI irradiated group respectively. This could be elucidated by differential radiation sensitivity of the organ involved during different localised irradiation. During HI, bone marrow consisting of haematopoietic stem cells (HSC) and mesenchymal stem cells (MSC) is the immediate target of radiation exposure whereas in abdominal region, most affected part is intestinal lining (enterocytes), crypt epithelial stem cells (Green and Rubin 2014; Wang et al. 2015). Differential response at 24 h time point also reflects the variation in the population kinetics of cell renewal system and the difference in the amount of damage that can be tolerated in these different tissue systems associated with different regions during irradiation (Hall and Giaccia 2018). It further substantiates the presence of least changes in thoracic region involving mainly lungs which are considerably late responding critical organ and changes observed at 24 h time point could be due to radiation-induced inflammatory cytokines mediated response (Garofalo et al. 2014; Ghandhi et al. 2018).
Radiation dose assessment and biodosimetry is still a challenge in PBI exposure owing to non-homogeneity of exposure during radiation accident. Radiation exposure of one organ system leads to multi-organ interaction even if they are not directly involved in radiation exposure (Prasanna et al. 2010). Therefore, understanding the cellular, organ injury and system biology may bring out comprehensive information on the pathophysiology of radiation injury that could conclude in identifying the target as an indicator for radiation exposure. Moreover, with robust bioinformatics, integrated approach with network interaction at all levels of information might overcome the gaps inherent in each technique and strengthen the change of getting robust marker of radiation exposure or develop an integrated algorithm for initial triage of victims. Metabolomics based outcome of this study could be an important node or seed point for an integrated solution to a complex research question in terms of radiation biomarkers.
5 Conclusion
In the present study, metabolomics based changes were observed in some metabolites irrespective of total or partial-body radiation exposure. Additionally, differential regional response was also reflected for some of the metabolites; exhibiting radiation sensitivity of the organs involved in different regions exposed during irradiation. In future, the authors intend to extend the study further with the inclusion of more parameters and to look for integrated network analysis with gene expression or micro-biome studies.
References
Allison, G. (2018). Nuclear terrorism: Did we beat the odds or change them? PRISM, NDU press, 7(3). https://ndupress.ndu.edu/Journals/PRISM/.
Anderson, R. M. (2019). Cytogenetic biomarkers of radiation exposure. Clinical Oncology, 31(5), 311–318.
Berkers, C. R., Maddocks, O. D. K., Cheung, E. C., Mor, I., & Vousden, K. H. (2013). Metabolic regulation by p53 family members. Cell Metabolism, 18(5), 617–633.
Bleijlevens, B., Shivarattan, T., van den Boom, K. S., de Haan, A., van der Zwan, G., Simpson, P. J., et al. (2012). Changes in protein dynamics of the DNA repair dioxygenase AlkB upon binding of Fe2+ and 2-oxoglutarate. Biochemistry, 51(16), 3334–3341.
Chen, C., Brenner, D. J., & Brown, T. R. (2011). Identification of urinary biomarkers from X-irradiated mice using NMR spectroscopy. Radiation Research, 175(5), 622–630.
Christophersen, O. A. (2012). Radiation protection following nuclear power accidents: A survey of putative mechanisms involved in the radioprotective actions of taurine during and after radiation exposure. Microbial Ecology in Health and Disease, 23(1), 14787.
Coy, S. L., Cheema, A. K., Tyburski, J. B., Laiakis, E. C., Collins, S. P., & Fornace, A. (2011). Radiation metabolomics and its potential in biodosimetry. International Journal of Radiation Biology, 87(8), 802–823.
Datta, K., Suman, S., Kallakury, B. V., & Fornace, A. J., Jr. (2012). Exposure to heavy ion radiation induces persistent oxidative stress in mouse intestine. PLoS ONE, 7(8), e42224.
Dilley, J. V. (1972). The origin of urinary taurine excretion during chronic radiation injury. Radiation Research, 50(1), 191–196.
Emwas, A. H., Roy, R., McKay, R. T., Tenori, L., Saccenti, E., Gowda, G. A. N., et al. (2019). NMR spectroscopy for metabolomics research. Metabolites, 9(7), 123.
Fazzino, F., Obregón, F., & Lima, L. (2010). Taurine and proliferation of lymphocytes in physically restrained rats. Journal of Biomedical Science, 17(Suppl 1), S24.
Fazzino, F., Urbina, M., Mata, S., & Lima, L. (2006). Taurine transport and transporter localization in peripheral blood lymphocytes of controls and major depression patients. Advances in Experimental Medicine and Biology, 583, 423–426.
Gan, W. Z., Ramachandran, V., Lim, C. S. Y., & Koh, R. Y. (2019). Omics-based biomarkers in the diagnosis of diabetes. Journal of Basic and Clinical Physiology and Pharmacology, 31(2), 1–21.
Garofalo, M., Bennett, A., Farese, A. M., Harper, J., Ward, A., Taylor-Howell, C., et al. (2014). The delayed pulmonary syndrome following acute high-dose irradiation: A rhesus macaque model. Health Physics, 106(1), 56–72.
Ghandhi, S. A., Turner, H. C., Shuryak, I., Dugan, G. O., Bourland, J. D., Olson, J. D., et al. (2018). Whole thorax irradiation of non-human primates induces persistent nuclear damage and gene expression changes in peripheral blood cells. PLoS ONE, 13(1), e0191402.
Ghosh, S. P., Singh, R., Chakraborty, K., Kulkarni, S., Uppal, A., Luo, Y., et al. (2013). Metabolomic changes in gastrointestinal tissues after whole body radiation in a murine model. Molecular BioSystems, 9(4), 723–731.
Golla, S., Golla, J. P., Krausz, K. W., Manna, S. K., Simillion, C., Beyoğlu, D., et al. (2017). Metabolomic analysis of mice exposed to gamma radiation reveals a systemic understanding of total-body exposure. Radiation Research, 187(5), 612–629.
Goudarzi, M., Mak, T. D., Chen, C., Smilenov, L. B., Brenner, D. J., & Fornace, A. J. (2014). The effect of low dose rate on metabolomic response to radiation in mice. Radiation and Environmental Biophysics, 53(4), 645–657.
Green, D. E., & Rubin, C. T. (2014). Consequences of irradiation on bone and marrow phenotypes, and its relation to disruption of hematopoietic precursors. Bone, 63, 87–94.
Hall, E. J., & Giaccia, A. J. (2018). Radiobiology for the radiologist (8th ed.). Philadelphia: Wolters Kluwer Health.
Hasegawa, A., Ohira, T., Maeda, M., Yasumura, S., & Tanigawa, K. (2016). Emergency responses and health consequences after the Fukushima accident evacuation and relocation. Clinical Oncology, 28(4), 237–244.
Hérodin, F., Valente, M., & Abend, M. (2014). Useful radiation dose biomarkers for early identification of partial-body exposures. Health Physics, 106(6), 750–754.
Hoyles, L., Jiménez-Pranteda, M. L., Chilloux, J., Brial, F., Myridakis, A., Aranias, T., et al. (2018). Metabolic retroconversion of trimethylamine N-oxide and the gut microbiota. Microbiome, 6(1), 73.
Kawamura, K., Qi, F., & Kobayashi, J. (2018). Potential relationship between the biological effects of low-dose irradiation and mitochondrial ROS production. Journal of Radiation Research, 59(Suppl 2), ii91–ii97.
Khan, A. R., Rana, P., Devi, M. M., Chaturvedi, S., Javed, S., Tripathi, R. P., et al. (2011a). Nuclear magnetic resonance spectroscopy-based metabonomic investigation of biochemical effects in serum of γ-irradiated mice. International Journal of Radiation Biology, 87(1), 91–97.
Khan, A. R., Rana, P., Tyagi, R., Kumar, I. P., Devi, M. M., Javed, S., et al. (2011b). NMR spectroscopy based metabolic profiling of urine and serum for investigation of physiological perturbations during radiation sickness. Metabolomics, 7(4), 583–592.
Kultova, G., Tichy, A., Rehulkova, H., & Myslivcova-Fucikova, A. (2020). The hunt for radiation biomarkers: Current situation. International Journal of Radiation Biology, 96(3), 370–382.
Kurland, I. J., Broin, P. Ó., Golden, A., Su, G., Meng, F., Liu, L., et al. (2015). Integrative metabolic signatures for hepatic radiation injury. PLoS ONE, 10(6), e0124795.
Lacombe, J., Sima, C., Amundson, S. A., & Zenhausern, F. (2018). Candidate gene biodosimetry markers of exposure to external ionizing radiation in human blood: A systematic review. PLoS ONE, 13(6), e0198851.
Lee, Y., Pujol Canadell, M., Shuryak, I., Perrier, J. R., Taveras, M., Patel, P., et al. (2018). Candidate protein markers for radiation biodosimetry in the hematopoietically humanized mouse model. Scientific Reports, 8(1), 13557.
Małachowska, B., Tomasik, B., Stawiski, K., Kulkarni, S., Guha, C., Chowdhury, D., et al. (2020). Circulating microRNAs as biomarkers of radiation exposure: A systematic review and meta-analysis. International Journal of Radiation Oncology Biology Physics, 106(2), 390–402.
Meadows, S. K., Dressman, H. K., Daher, P., Himburg, H., Russell, J. L., Doan, P., et al. (2010). Diagnosis of partial body radiation exposure in mice using peripheral blood gene expression profiles. PLoS ONE, 5(7), e11535.
Ozasa, K., Cullings, H. M., Ohishi, W., Hida, A., & Grant, E. J. (2019). Epidemiological studies of atomic bomb radiation at the Radiation Effects Research Foundation. International Journal of Radiation Biology, 95(7), 879–891.
Pannkuk, E. L., Laiakis, E. C., Authier, S., Wong, K., & Fornace, A. J. (2017). Gas chromatography/mass spectrometry metabolomics of urine and serum from nonhuman primates exposed to ionizing radiation: Impacts on the tricarboxylic acid cycle and protein metabolism. Journal of Proteome Research, 16(5), 2091–2100.
Pannkuk, E. L., Laiakis, E. C., Mak, T. D., Astarita, G., Authier, S., Wong, K., et al. (2016). A lipidomic and metabolomic serum signature from nonhuman primates exposed to ionizing radiation. Metabolomics: Official Journal of the Metabolomic Society, 12(5), 80.
Pomper, M. A., & Tarini, G. (2017). Nuclear terrorism—Threat or not? AIP Conference Proceedings, 1898, 050001. https://doi.org/10.1063/1.5009230
Prasanna, P. G. S., Moroni, M., & Pellmar, T. C. (2010). Triage dose assessment for partial-body exposure: Dicentric analysis. Health Physics, 98(2), 244–251.
Rentea, R. M., Lam, V., Biesterveld, B., Fredrich, K. M., Callison, J., Fish, B. L., et al. (2016). Radiation-induced changes in intestinal and tissue-nonspecific alkaline phosphatase: Implications for recovery after radiation therapy. American Journal of Surgery, 212(4), 602–608.
Romano, K. A., Vivas, E. I., Amador-Noguez, D., & Rey, F. E. (2015). Intestinal microbiota composition modulates choline bioavailability from diet and accumulation of the proatherogenic metabolite trimethylamine-N-oxide. American Society for Microbiology, 6(2), e02481.
Shao, L., Luo, Y., & Zhou, D. (2014). Hematopoietic stem cell injury induced by ionizing radiation. Antioxidants & Redox Signaling, 20(9), 1447–1462.
Singh, V. K., Newman, V. L., Romaine, P. L., Hauer-Jensen, M., & Pollard, H. B. (2016). Use of biomarkers for assessing radiation injury and efficacy of countermeasures. Expert Review of Molecular Diagnostics, 16(1), 65–81.
Sproull, M., Kramp, T., Tandle, A., Shankavaram, U., & Camphausen, K. (2017). Multivariate analysis of radiation responsive proteins to predict radiation exposure in total-body irradiation and partial-body irradiation models. Radiation Research, 187(2), 251–258.
Valente, M., Denis, J., Grenier, N., Arvers, P., Foucher, B., Desangles, F., et al. (2015). Revisiting biomarkers of total-body and partial-body exposure in a baboon model of irradiation. PLoS ONE, 10(7), e0132194.
Wang, J., Shao, L., Hendrickson, H. P., Liu, L., Chang, J., Luo, Y., et al. (2015). Total body irradiation in the “hematopoietic” dose range induces substantial intestinal injury in non-human primates. Radiation Research, 184(5), 545–553.
Wang, Z., Klipfell, E., Bennett, B. J., Koeth, R., Levison, B. S., DuGar, B., et al. (2011). Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature, 472(7341), 57–63.
Yoshida, T., Goto, S., Kawakatsu, M., Urata, Y., & Li, T. S. (2012). Mitochondrial dysfunction, a probable cause of persistent oxidative stress after exposure to ionizing radiation. Free Radical Research, 46(2), 147–153.
Funding
This work was supported by Defence R & D Organsiation (DRDO), Ministry of Defence, India (INM 313). KM was supported by University Grant Commission (UGC), India.
Author information
Authors and Affiliations
Contributions
PR and RB conceived the project and designed the study, PR supervised the NMR experiments and data analysis. PR, AD and KM involved in experimentation. KM and RT involved in analysis and interpretation of data and writing of manuscript. PR and RB evaluated the manuscript critically and all the authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Research involving human and/or animal rights
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Electronic supplementary material 1 (TIF 5841 kb)
Fig. 1: PCA Loading plots of (PC1 and PC2): (a) Control and TI groups (b) Control and HI groups, (c) Control and AI groups (d) Control, TI, AI and HI groups, (e) Control and TBI groups and (f) Control, TBI and PI groups.
Electronic supplementary material 2 (TIF 3234 kb)
Fig. 2: Multivariate analysis in TI (Thoracic Irradiated) group (a) PCA score plot showing variation between control and TI groups, (b) PLS-DA score plot showing complete separation of metabolic profile of TI from control group. Group segregation is seen in component 1 with 55.7% (elliptical boundary shows 95% confidence interval), (c) VIP plot presenting top 15 metabolites responsible for observed discrimination between the groups, (d) Corresponding Heatmap showing comparison of metabolites between the groups. (C-Red, TI-Green).
Electronic supplementary material 3 (TIF 3595 kb)
Fig. 3: Multivariate analysis in HI (Hind Limb Irradiated) group: (a) PCA score plot showing variation between control and HI groups. (b) PLS-DA score plot showing complete separation of metabolic profile of HI from control group. Group segregation is seen in component 1 with 49.4% (elliptical boundary shows 95% confidence interval). (c) VIP plot presenting top 15 metabolites responsible for observed discrimination between the groups (d) Corresponding Heatmap showing comparison of metabolites between the groups. (C-Red, HI-Green).
Electronic supplementary material 4 (TIF 3282 kb)
Fig. 4: Multivariate analysis in AI (Abdominal region Irradiated) group (a) PCA score plot showing variation between control and AI groups, (b) PLS-DA score plot showing complete separation of metabolic profile of AI from control group. Group segregation is seen in component 1 with 46.8% (elliptical boundary shows 95% confidence interval), (c) VIP plot presenting top 15 metabolites responsible for observed discrimination between the groups, (d) Corresponding Heatmap showing comparison of metabolites between the groups (C-Green, AI-Red).
Electronic supplementary material 5 (TIF 3189 kb)
Fig. 5: Pattern recognition analysis of urine samples (a) PCA score plot showing variation between control, AI, HI and TI groups, (b) PLS-DA score plot showing separation of all irradiated groups from control group (elliptical boundary shows 95% confidence interval) (c) VIP plot presenting top 15 metabolites responsible for observed discrimination between the groups, (d) Corresponding Heatmap showing comparison of metabolites between the groups (C-Green, AI- Red, HI- Purple and TI-Blue).
Electronic supplementary material 6 (TIF 8113 kb)
Fig. 6: Relative levels of peak intensity of significantly perturbed metabolites in all irradiated groups (TI, AI, HI and Pooled Partial) compared to control group. Data is presented as mean ± standard deviation. (*p-value < 0.05, **p-value < 0.01 and ***p-value < 0.001).
Electronic supplementary material 7 (TIF 3422 kb)
Fig. 7: Pattern recognition analysis of urine samples (a) PCA score plot showing variation between control and TBI groups, (b) PLS-DA score plot showing complete separation of metabolic profile of TBI from control group. Group segregation is seen in component 1 with 54% (elliptical boundary shows 95% confidence interval), (c) VIP plot presenting top 15 metabolites are responsible for observed discrimination between the groups, (d) Corresponding Heatmap showing comparison of metabolites between the groups (C-Red, TBI-Green).
Electronic supplementary material 8 (TIF 5557 kb)
Fig. 8: Representative 1H NMR spectrum of urine showing identified metabolites (a) and comparative NMR spectra of control (C) and different irradiated groups (TBI, TI, AI and HI) (b).
Rights and permissions
About this article
Cite this article
Maan, K., Tyagi, R., Dutta, A. et al. Comparative metabolic profiles of total and partial body radiation exposure in mice using an untargeted metabolomics approach. Metabolomics 16, 124 (2020). https://doi.org/10.1007/s11306-020-01742-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-01742-7