Abstract
The fungal system holds morphological plasticity and metabolic versatility which makes it unique. Fungal habitat ranges from the Arctic region to the fertile mainland, including tropical rainforests, and temperate deserts. They possess a wide range of lifestyles behaving as saprophytic, parasitic, opportunistic, and obligate symbionts. These eukaryotic microbes can survive any living condition and adapt to behave as extremophiles, mesophiles, thermophiles, or even psychrophile organisms. This behaviour has been exploited to yield microbial enzymes which can survive in extreme environments. The cost-effective production, stable catalytic behaviour and ease of genetic manipulation make them prominent sources of several industrially important enzymes. Pectinases are a class of pectin-degrading enzymes that show different mechanisms and substrate specificities to release end products. The pectinase family of enzymes is produced by microbial sources such as bacteria, fungi, actinomycetes, plants, and animals. Fungal pectinases having high specificity for natural sources and higher stabilities and catalytic activities make them promising green catalysts for industrial applications. Pectinases from different microbial sources have been investigated for their industrial applications. However, their relevance in the food and textile industries is remarkable and has been extensively studied. The focus of this review is to provide comprehensive information on the current findings on fungal pectinases targeting diverse sources of fungal strains, their production by fermentation techniques, and a summary of purification strategies. Studies on pectinases regarding innovations comprising bioreactor-based production, immobilization of pectinases, in silico and expression studies, directed evolution, and omics-driven approaches specifically by fungal microbiota have been summarized.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The idea of sustainable and innovative bio-economical use of science is the basis for scientific advancements. With more than 4000 different enzymes reported, an average of 200 enzymes has the potential for commercialization, although only 10% can be industrially produced. There is huge potential in the enzyme market, which was reported to be around 6.3 billion dollars in 2017 and has a projection of a compound annual growth rate (CAGR) of 6.8% until 2024. Over the next five years, the food enzyme market is expected to grow by 7.5%, the highest rate of any market projected in the industry (Food enzyme trend gminsight). The thrust to uplift the production of renewable resources is greatly impregnated with the requirement of low-cost yet highly efficient systems (Joshi et al. 2018; Raveendran et al. 2018). White biotechnology is dedicated to harnessing biocatalysts i.e., enzymes and microorganisms at an industrial scale (Meyer et al. 2020, Cairns et al. 2021). The mandate of white biotechnology is to provide pure and replenishable sources as potential alternatives for industrial acceleration resulting in improved, bio-economical, and highly sustainable products (Hyde et al. 2019).
The microbial system is the foundation of biotechnological applications and innovations. The fungal community is a highly exploited eukaryotic system that can be applied directly or by acting as the source to produce industrially important products (Joshi et al. 2018). Filamentous fungi are efficient decomposers that can feed on and break down organic materials and polymeric compounds. Industrial production of commercial citric acid marks the stepping stone of fungal biology as an important commercial industrial product. Enzymes are the backbone of sustainable environments and harnessed industrially for centuries. The fungal system has been a pioneer source of these commercially applicable enzymes. Cellulases, amylases, pectinases, laccases proteases, and lipases are secreted by fungal cells (Hyde et al. 2019; Meyer 2019). These enzymes hydrolyse plant polysaccharides such as cellulose, starch, pectin, proteins, and lipids respectively. Their wide substrate interaction leads to microbial enzymatic intervention in food and feed, pulp and paper, detergent, fuel, pharmaceutical, and chemical sectors (Ahmed et.al. 2021, Kordi et al. 2022).
Pectin alone constitutes approximately 35% of the plant cell wall composition and shows structural complexity and diversity. Pectin polysaccharide gives plant tissues their tensile strength and rigidity. Pectin is frequently employed in the food sector as a gelling agent, thickening, emulsifier, or stabilizer. It can also be used in the pharmaceutical industry as a blood pressure stabilizer, cholesterol controller, and detoxifier (Gawkowska et al. 2018). Natural sources of pectin include fruit waste from pomegranate, banana, lemon, orange, pineapple, wheat bran, malt sprout, and rice bran. Their physiological properties though isolated from different sources remain similar and are beneficial to humankind (de Souza and Kawaguti 2021).
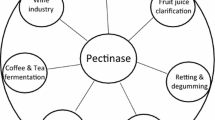
Pectinases are the group of hydrolases, depolymerase, esterases, and lyases enzymes. These act on pectin, protopectin, pectic acid, and galacturonate (Yadav et al. 2009a). Based on the mode of action, specificity of the substrate, and cleavage mechanism there exists a diverse family of pectinolytic microbial enzymes. In this emerging era of biotechnological innovations, fungal pectinase accelerates its way as a promising natural biotechnological innovative agent (Nighojkar et al. 2019; Anand et al. 2020). Microbial pectinases have a broad range of applications and high catalytic effectiveness has significantly raised the global demand. Microbes are natural sources of pectinases that are often employed due to their simplicity of manufacture and distinctive physicochemical features. With a 25% share in the market for food and beverage enzymes, the pectinases family of enzymes is a great part of the biotechnological industry. They are on top of the list of industrial enzymes made for commercial production. Pectinases of acidic nature are preferred for clarification of fruit juices, maceration of vegetables in the manufacturing of pastes and purees, and winemaking. Alkaline pectinases are often used in the retting of natural textile fibres, treatment of pectic-rich wastewater, fermentation of tea, extraction of vegetable oil, and treatment of paper and pulp (Kohli and Gupta 2015; Patidar et al. 2018; Thakur et al. 2021).
The literature available on microbial pectinases has established the importance of the catalyst in industrial sectors. The present review is solely inclined towards fungal pectinolytic interventions in enzyme biotechnology. The status of fungal pectinases and their cost-effective production strategies, the factors affecting production, the large-scale bioreactor-based productions and the purification of the enzyme have been highlighted in this review. Further, emphasis has also been made to include the recent innovations like immobilisation, directed evolution and omics-based approaches targeted in fungal pectinases.
Pectin
Pectin is an abundant natural product predominately observed in dicotyledonous plants. It is secreted by Golgi bodies into the apoplast of cells that are richly methyl-esterified (Sinclair et al. 2018). The committee of the American Society in 1944 accepted definitions of pectic substances, which include pectin acids, pectic acids, and protopectin within the complex class of these macromolecules. The pectinic acids are colloids of galacturonic acids methyl ester and pectic acids without methyl ester (Harholt et al. 2010). Protopectin is considered the parent molecule of a pectic substance and together with pectin and pectic acids was then summarised as “pectin” (Mohen 2008, Anderson 2019).
The solubility behaviour of pectins is categorised as (i) pectins soluble in water or diluted solutions, (ii) pectins soluble in chelators like EDTA, and (iii) protopectin soluble in alkaline or hot solutions based on this observation. The water-soluble and chelator-soluble pectins are derived from the middle lamella of the plant cell wall. These are composed of galacturonic acid residues with a tenth of neutral sugars and barely 2% rhamnose (Voragen et al. 2009; Patidar et al. 2018). The distribution of sugars attached with free carboxyl groups gives the classes their nature of water and chelator solubility. Pectin of alkaline solubility are embedded in part of the cell walls. Alkali-soluble pectins are structured with arabinose and galactose sugars. Typically, softening during ripening or heating is accompanied by a decrease in the proportion of protopectin and an increase in water-soluble pectin (Yapo 2011).
Apart from solubility, pectin has variable percentages of esterification as (i) High-methoxyl pectins, having esterification levels between 40 and 50%, and (ii) Low-methoxyl pectins with esterification levels below 40%. The esterification level can be controlled by acid, alkali, or enzyme treatment of high-methoxyl pectins. This imparts pectin its unique characteristics and helps form gels under specific conditions. The pectin polysaccharide is made up of distinct categories of sugars representing unique structures (Voragen et al. 2009; Wusigale et al. 2020). There are 17 different monosaccharides linked with approximately more than 20 different linkages. These together give pectin a macro-complex structure (Yang and Anderson 2020; Gutierrez-Alvarado et al. 2022). These sugars govern the role of pectin in cell adhesion and separation, expansions and regulation of plant cell walls, and the development of organs and plants.
The complex nature of pectin structure includes components like homogalacturonan (HG), xylogalacturonan (XGA), homogalacturonan, rhamnogalacturonan I (RGI), and rhamnogalacturonan II (RGII) (Yang and Anderson 2020). The key constituent of the spine is α-1,4-linked galacturonic acid (GalA) residues. These residues undergo esterification at six carboxyl carbon and acylation at the third or second oxygen of the chain. This base is alternatively lined by rhamnose sugar and galacturonic acid residues having a structurally similar side chain of arabinose and galactose sugars. Homogalacturonan accounting for 60% of the total pectin structure forms the smooth region of sugar residues. The neutral sugars together are ramified to form a hairy sugar region. Rhamnogalacturonan II (RGII) within HG constitutes twelve different types of sugar resides, including 3-deoxy-lyxo-2-heptulosaric acid (DHA), 3-deoxy-manno-2-octulosonic acid (KDO), apiose and acetic acid. The reproductive tissues, fruits, and seeds store the xylogalacturonan. It forms a single side chain of units of b-D-Xylp-(1 → 3) which is commutated with HG molecules (Zdunek et al. 2021; Shin et al. 2021; Gutierrez-Alvarado et al. 2022).
Pectinases family: an overview
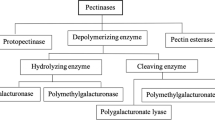
Pectinases act on pectic substances. They possess negative charge, high molecular weight glycosidic bond-linked macromolecules with substrate specificities on pectin (Anderson 2019). Pectinases are classified in respect of the type of modifications of the backbone chain as protopectin, pectic acid, pectin acid, and pectin. Pectinases amalgamate together lyases, hydrolases, and esterases classes of enzymes to act on pectin (Yadav et al. 2009b, Pedrolli et al. 2009). These can work endogenously by cleaving glycosidic bonds to release residues from the inside or in an exogenous manner to cleave residues from the ends. These can be produced through extracellular or intracellular modes. Though intracellular secretion is more costly in comparison to extracellular production. The classification of pectinases or pectinolytic enzymes based on the existence of different pectic substances, reaction mechanisms, and degradation of the hairy and smooth regions has been reported (Kashyap et al. 2001; Jayani et al. 2005; Favela-Torres et al. 2006).
A discrete collection of two hundred and sixty-nine enzymatic families that are similar on grounds of amino acid sequence are called the Carbohydrate-modifying enzymes. This family is broadly distributed under four classes: glycoside hydrolases (GHs), glycosyltransferases (GTs), polysaccharide lyases (PLs), and carbohydrate esterases (CEs). These classes have subgroups of structurally and catalytically related families. This has been listed in the carbohydrate-active enzyme (CAZy) database (www.cazy.org) (Cantarel et al. 2009). Pectinases share a diverse group of enzymes that distinctively occupy their positions in the GH, PL, and CE families (Drula et al. 2022).
Glycoside hydrolases (GH)
Family GH28 is commonly referred to as polygalacturonases, which are glycosidases acting on homogalacturonan and rhamnogalacturonan components of pectin. This includes enzymes with hydrolysis mechanisms. They are capable of hydrolysing glycosidic linkage between carbohydrates -carbohydrates and a non-carbohydrate moiety. GH28 enzymes are also categorized into three distinct categories acting on homogalacturonan, rhamnogalacturonan and xylogalacturonan (Sprockett et al. 2011; Villarreal et al. 2022). It hydrolyses polygalacturonic acid on α-1,4-glycosidic linkages producing d-galacturonate. Fungal polygalacturonase can produce monomeric galacturonic acids on its depolymerization. Mode of action distributes them as Endo-PG (EC 3.2.1.15) which liberates saturated oligogalacturonides and Exo-PG (EC 3.2.1.67) releases saturated galacturonic acid residue. The residue is obtained from the non-reducing end of homogalacturonan by hydrolytic catalysis (Yang et al. 2018; Anand et al. 2020; Christensen 2020). Xylogalacturonans (XG) are enzymes responsible for cleaving glycosidic linkages in the xylose-substituted rhamnogalacturonan chain and the end products are xylose-galacturonate dimers. Rhamnogalacturonan is hydrolytically cleaved by RG galalcturonohydrolase. Its non-reducing end produces monogalacturonate (Villarreal et al. 2022).
Polysaccharide lyases (PL)
These enzymes cleave uronic acid-containing polysaccharide chains. They use the β-elimination mechanism to generate an unsaturated (hexen)uronic acid residue and a new reducing end. PLs, can cleave alginate, heparin, hyaluronan, pectin, xanthan, and several exopolysaccharides (cazy.org/Polysaccharide-Lyases; Yadav et al. 2009c, Chakraborty et al. 2017). PL family 1, 2, and 9 share distributions of lyases degrading pectin. Pectate lyase (PL) results in forming an unsaturated product (α-4,5-d-galacturonate) through a trans-elimination reaction on polygalacturonase acids. Endo-PL (EC 4.2.2.9), acts on a nonreducing end. Pectin lyase (PNL) (EC 4.2.2.10) results in the formation of 4,5- unsaturated oligo-galacturonate. PNL performs a β-elimination mechanism without affecting the ester content of the polymer chain. This ester content is responsible for the specific aroma of fruits. Toxic methanol production is limited by enzymatic degradation. Henceforth, these are preferred in fruit juice clarification industries. Rhamnogalacturonan lyases degrade rhamnogalacturonan I and are distributed in families 4 and 11 (Zheng et al. 2021).
Carbohydrate esterases (CE)
These enzymes catalyse acylation at the oxygen or nitrogen end. The members of this family remove esterified modifications from mono-, oligo- and polysaccharides. The acylation provides easy access to glycoside hydrolase (cazy.org/Carbohydrate-Esterases; Wardman et al. 2022). Pectin methyl esterases are grouped in CE 8 family and act preferentially on a methyl ester group of galacturonate to produce methanol and pectic acid. The action of PMEs forms pectate gel from homogalacturonan. The action of esterases can hinder the action of polygalacturonases (Benen et al. 2002). Rhamnogalacturonan acetyl esterase is responsible for cleaving acetyl groups of the rhamnogalacturonan chain that constitutes the major part of the hairy portion of pectin and belongs to the family CE12. Pectin acetyl esterase belongs to CE13 and hydrolyses the acetyl ester of pectin. They help the formation of pectic acid and acetate and acylation affects the age and differentiation of plant tissues. It even acts as protection from different enzymatic interactions. The esterases assist actively in biomass saccharification and have diverse biological and biotechnological applications (Benen et al. 2002; Bonnin and Pelloux 2020). The diversity of pectinases and their potential for industrial application is depicted in Fig. 1.
Fungal pectinases
Microorganisms have been in the environment from the beginning of time on this planet. In the scientific world, the study of the structural, functional, and ecological attributes of microorganisms is significant (Prasad et al. 2021). Microbial enzymes particularly from fungi are preferred over other sources because: (i) Their content is more predictable. (ii) They have a wide range of enzymes; (iii) Bulk production generally resin with low costs and reliable raw materials. (iv) Their productivity rate is high and they contain a greater amount of active ingredients. (v) Fungi can be easily managed to take the desired enzymes, and they can be made in large quantities rapidly and inexpensively through existing fermentation techniques and sophisticated instrumentation. (vi) Enzyme production may be programmable in various environments. and (vii) more potentially hazardous components like phenolic compounds, endogenous enzyme inhibitors, and proteases are found in plant and animal tissues than in microorganisms (Sharma et al. 2013; Singh et al. 2019).
Fungal microbiota cover about 50% of microbial enzyme production. Around 35% of the production shared is held by bacteria and only 15% is produced from higher organisms (Wösten 2019). The fungal system is a popular source of enzymes because they provide a cost-effective technology with reduced resource consumption and minimal emissions, as opposed to animal and plant sources. Fungi and yeast alone are a producer of half of the globally used enzymes (Lübeck and Lübeck 2022). The secretion of pectinases by fungi assists in the breakdown of the middle lamella in plants (Prasad et al. 2021). Soil is a diverse and dynamic environment that is home to a diverse range of microorganisms, especially fungi. The traditional laboratory culture techniques have made the greatest contribution to gaining access to microbial diversity. Despite its familiarity and use, it is one of the world’s least explored environments. Soil microorganisms play a crucial role in plant development and carbon and nutrient cycling. The bulk of soil microorganisms, on the other hand, have yet to be isolated, and their roles are mostly unknown. These microbial communities are exploited as a sustainable source for microbial enzymatic production systems (Baldrian 2019; Selvasekaran and Chidambaram 2020).
Fungi generate a plethora of extracellular enzymes capable of degrading organic materials, one of which is pectinolytic enzymes. Commercial enzymes have been produced using filamentous fungi for more than 50 years (Haile and Ayele 2022). Pectinolytic enzymes are one of the extracellular enzymes that fungi produce that can break down organic molecules. One of the most potent sources of pectinases is filamentous fungi, which can be extensively exploited in the production of SSF at a low cost. Many different fungal species have been reported to produce pectinases. Aspergillus niger is the most typical fungus used in the production of pectinolytic enzymes for industrial use (Gutiérrez-Correa et al. 2012). A. oryzae, A. fumigatus, A. terreus, A. sojoe, A. awamori, and other Aspergillus species are also known to produce pectinase. A. giganteus was the first species whose production of endo-PGL was noted. Additionally, species of Penicillium, Fusarium, Mucor, Neurospora crass, Sclerotinia sclerotium, and others play a part in the manufacture of pectinase (Sharma et al. 2013; Haile and Ayele 2022). Fungal pectinases play a part in the phytopathological process. They interact in plant–microbe symbiosis, and the decomposition of dead plant material, thereby, contributing to the natural carbon cycle. In the context of mining fungal pectinolytic sources, soil samples have been indefinitely explored for the isolation of novel fungal strains as listed in Table 1.
Production strategies
Fermentation-based production of microbial pectinases is facilitated by solid-state fermentation and submerged fermentation industrially and at a small scale. The advantages of fermentation-based production of enzymes include low costs, low energy consumption, and low waste-water generation, and it can be exploited to repurpose organic wastes into value-added products. Fermentation-based microbial enzyme mass production uses either solid-state fermentation (SSF) or submerged fermentation (SmF). SmF technology is often used to produce microbial enzymes, especially from bacterial sources and the major advantage is easy to control the process as compared to SSF (Sharma et al. 2013).
Solid State Fermentation uses a solid substrate that acts as a natural habitat for fungi to attach. The fermentation requires lower to no moisture content occurring in the absence or near absence of free water. The sturdy foundation offers support, or occasionally both support and sustenance. The main benefits of SSF include low capital expenditure, reduced levels of catabolite repression and end-product inhibition, low wastewater output, improved productivity, higher enzyme yields, and better product recovery. SSF has been used predominantly as it triggers the production of various enzymes directly from raw materials rich in lignocellulose (Kumar and Verma 2020). SSF is highly favourable for fungal microflora as it is like their natural habitat. Some of the limitations of the SSF include the need for proper aeration and humidity control and a time-consuming scale-up process. Pectinases of fungal origin have been extensively reported by using the solid-state fermentation method (Soccol et al. 2017; Lizardi-Jiménez and Hernández-Martínez 2017). The production of fungal pectinases by SSF requires optimization of several parameters which can directly affect the enzyme production.
Factors influencing the production of pectinases by SSF
The process of fermentation is dependent on biological and physio-chemical parameters that greatly affect the kinetics of the microbial enzymes. To improve the efficiency of the enzymes, these parameters need to be optimised and microbes, the size of the inoculum, and substrates are some of the important biological parameters. Further, incubation temperatures, pH specificities, moisture content, aerations, rotations, and heat transfer affect the performance of the enzymes (Soccol et al. 2017).
Fungal spores can directly be added as inocula and have a very fast production rate. These can grow over a range of temperature conditions between 24 and 30 °C. Thermophilic fungi can also grow optimally in this range. The pH range may change according to the substrate used, however, for the best growth, fungi strains prefer an acidic to a neutral range (Prado Barragán et al. 2016). Optimizations of moisture content have resulted in more stress-resistant pectinase production. The rotation and agitation affect microbial growth and contamination. Production of fungal pectinase is severely constrained by bacterial contamination (Prado Barragán et al. 2016; Chen and Wang 2017).
The mesophilic and thermophilic fungal pectinase production during SSF is also affected by heat transfer during the process. The gases produced by the fungal inoculum and moisture vaporization regulate the heat of the system (Kumar et al. 2021; Chilakamarry et al. 2022).
Substrates used for the production of fungal pectinase
Higher fungi have well-tuned enzymes, spores, and metabolites for development on solid, moist substrates. For instance, fungus spores produced by SSF display greater stability, are more resistant to drying, and have higher germination rates for longer periods (Arun et al. 2020). The substrate acts as structural ed support rich in nitrogen and carbon for the growth of microorganisms. The nutritional composition and quality problems may affect the fermentation batches. This variation could lead to decreased production. The choice of substrate will determine how much heterogeneity is introduced during the process. The most often utilised substrates for SSF include agricultural and food processing wastes such as wheat bran, sawdust, apple pomace, cassava, sugar beetroot pulp, citrus waste, maize cob and banana waste. Innovations in the production of pectinases using different agro-wastes like peels and pulps of citrus, orange, coffee, grapefruit, and banana using both SSF and SmF have been reported recently (Bharathiraja et al. 2017; Chilakamarry et al. 2022).
Fruits and vegetable peels are rapidly utilized nowadays as they are environment-friendly and immensely nutritious for microbes. Peels of citrus fruits, bananas, sweet potatoes, and mango are being vigorously studied. The pomace of apple, kiwi, peach, and grapes are pectin-rich biomass for valorisation via fermentation. Other agro-industrial residues such as oil cakes of pumpkin, sesame, groundnut, and sunflower oil have also been used as substrates (Lopes and Ligabue-Braun 2021). Additionally, pectinolytic enzyme production has been reported by the use of sugarcane bagasse, corn cobs, soybean hulls, sugar beetroot pulp, barley husks and straws as sources of carbon. Tea extract serves as an important source of nitrogen. In addition to these, brewery waste, sewage wastewater, drainage effluents, tobacco stalks, molasses, and vegetable and fruit juices work excellently as liquid substrates for the fermentation of fungal pectinases (Sadh et al. 2018; Cano et al. 2020; Chukwuma et al. 2020).
Bioreactors for the production of fungal pectinases
For large-scale bulk production, bioreactors have been used. These bioreactors or fermenters are designed for processing biological products under a specifically controlled environment. Bioreactors for fungal pectinases have used lignocellulosic wastes, and agricultural wastes as substrates for industry efficient scaled production of enzymes (Cerda et al. 2019). Bioreactors prefer solid state-based fermentation methods for the production of fungal pectinases. In the light of fungal pectinases, Aspergillus niger has been extensively utilized for pectinases production by solid-state fermentation using the packed bed, and bench scale rotating-drum reactors (Finkler et al. 2017; Poletto et al. 2017; Reginatto et al. 2022). A 40 cm high packed bed bioreactor yielded productivity of 1840 U/g pectinases using Aspergillus niger (Pitol et al. 2016). Raimbault columns, packed-bed bioreactors, Erlenmeyer flasks, perforated trays, and other static bioreactors have been used to produce pectinases (Yang and Sha 2019). These bioreactors are chosen because of their usability and simplicity. A. niger on sugarcane bagasse and orange pomace has been utilized as solid-state substrates for production using a tray and rotating drum bioreactors (Mahmoodi et al. 2019). Agitated bioreactors utilise intermittent or continuous mixing to homogenise substrate using solid-state fermentation. It is possible to construct agitated bioreactors with or without a water jacket to regulate temperature (Mitchell and Krieger 2019). This type of reactor may be continuously or intermittently agitated. Shear problems and damage to the fungal mycelium’s structural integrity may occur, depending on the degree of agitation (Shanmugam et al. 2022). Large fermenters are commonly built of stainless steel in the food and beverage industries because of their ability to resist corrosion. A bioreactor’s design incorporates numerous essential engineering elements that are regularly updated and modernised to increase the final product’s productivity and quality (Kaur and Kaur 2019). Basal stirred tank fermenters utilised A. foetidus strain for optimization and evaluation of pH effect on microbial enzymes including pectinases (Li et al. 2018). Innovative forms of bioreactor-based fermentation largely depend on aeration techniques. These reactors have been modernised with the inclusion of steam traps, valves, mechanical foam breakers, pH temperature and pO2 monitors, micro-spargers for self-cleaning, and other sampling ports. Connecting to computers is a crucial advancement for novel bioreactors since it speeds up data processing and calculation and facilitates operational optimisation (Mitchell et al. 2019; John et al. 2020; Leite et al. 2021).
Response surface methodology (RSM) utilisation for bioreactor-based production using shake flasks has been utilised recently to produce pectinases at a concentration of 380 U/ml by A. sojoe (Fratebianchi et al. 2017). Similarly, an indigenous Aspergillus sp. isolated from coffee waste was used in response surface methodology designed on an SSF-based tan ray bioreactor to yield 29.9 IU/g of pectinases (Núñez Pérez et al. 2022).
Purification of fungal pectinases
Enzyme purification can be achieved by using a variety of conventional and modern techniques. The choice of the best treatment stage is a prerequisite for the enzyme purification process to be successful. Depending on the intended usage of the enzyme, the degree of purification may vary. Purification of microbial pectinases has been attained by simple centrifugation, sedimentation, or precipitation (Holm et al. 2018). The removal of inorganic and organic impurities is highly feasible by salting out using ammonium sulphate salts. This method of purification or partial purification has yielded a stable protein with better activity. Solvent precipitation using acetone, ethanol, and methanol, based on the solubility of protein is a cost-effective method for the removal of organic and inorganic impurities. The salt-based precipitation has been preferred as other solvent methods for pectinase. This is generally followed by dialysis to yield salt unbound proteins which are dissolved in buffers for optimal activities. Purification using counter solvents like butanol or octanol or by ultrafiltration facilitates the generation of aqueous pectinase. This eliminates the need for precipitation with dialysis of salt-based methods (Patel et al. 2017; Raina et al. 2022).
Purification of pectin lyases produced from Penicillium oxalicum, P. citrinum, Aspergillus flavus, A. ficuum, A. terricola, Fusarium decemcellulare, and F. lateritum has been performed simply by using ammonium sulphate precipitation and column chromatography method (Yadav and Shastri 2007; Yadav et al. 2008, 2009a, c, 2013, 2014, 2017b). Exo-polygalacturonase from Aspergillus flavus has been purified using solvent-based acetone purification, followed by cellulose column and gel filtration chromatography (Anand et al. 2017a).
Ion exchange, gel filtration, and affinity-based chromatographic methods are used to produce samples with a comparatively greater level of purity. The form, size, charge, hydrophobicity, or binding ability of the stationary phase are criteria used in chromatographic procedures to purify microbial pectinases. The molecular properties and interactions that underlie ion exchange, surface adsorption, partition, and size exclusion are also important considerations (Coskun 2016). Pectinolytic purification has been predominately accomplished by column chromatography (Smith 2005; Ullah 2012; Bassim Atta and Ruiz-Larrea 2022). Ion exchange or gel filtration, which gives rise to purer fractions of pectinases, along with a significant increase in its specific activity has also been reported. Anion exchange column-based purification for polygalacturonase from Calonectria pteridis utilized eucalyptus leaves in submerged fermentation (Ladeira Ázar et al. 2020). An indigenously isolated soil-borne Aspergillus japonicus yielded 2.9-fold purified polygalacturonase using two chromatographic techniques simultaneously (Cavalieri de Alencar Guimarães et al. 2022). A repertoire of purification strategies has been adopted for the purification of fungal pectinases from different fungal strains as shown in Table 2.
Innovations: diverse approaches
Immobilisation
The pectinolytic industrial intervention is disrupted due to their recovery rates, and low stability. Immobilization of enzymes enhances storage, reduces product contamination, and simplifies the separation of products, which in their free form is challenging. It improvises the catalytic properties of enzymes and enhances their functioning in adverse conditions (Bashir et al. 2020). Thereby, facilitating the recovery and reuse of enzymes in the medium and enhancing the economic feasibility of the enzymes. Suitable immobilization protocols and supportive environments are required for enzyme biocatalysts with high enzymatic activity (Patel et al. 2022). Pectinases have been immobilized using diverse supports by membrane adsorption, covalent binding, and cross-linking mechanisms. A variety of supports, including beads, microspheres, pulp fibre, matrix, resins, capsules, nanoparticles pumice, and magnetic beads have been deployed (Martín et al. 2019; Karataş et al. 2021). The magnetic core of magnetic particles as beads makes it simple, rapid, and effective to separate the enzyme from the reaction mixture using an external magnetic field, making them suitable support for enzyme immobilization. Additionally, the size of the particle can be adjusted to give a large surface area and high enzyme activity (Soozanipour et al. 2019; Trindade Ximenes et al. 2021). Direct crosslinking of different enzyme preparations is the most typical technique for producing cross-linked enzyme aggregates (CLEAs). The advantages of this approach are highly concentrated enzyme activity, greater stability, and the absence of an extra carrier’s associated production costs (Nouri and Khodaiyan 2020).
Adsorption, covalent binding, and entrapment are just a few of the methods utilised to keep enzymes inside the membrane. Enzymes are frequently attached to membranes by chemical bonds and adsorption. Pectinase is frequently bound to membranes using adsorption techniques. Chemical enzyme binders including glutaraldehyde, glycidyl methacrylate, and carbonyl diimidazole are used to adsorb membranes. It has been observed that membrane-bound enzyme exhibits enhanced thermal stability and temperature optima. Among the different methods of immobilising enzymes, covalent immobilisation is frequently preferred. This is so that it won’t allow the enzyme to desorb from the support during the process (Nadar and Rathod 2019).
A scale bioreactor used in stainless steel bases matrix was immobilized to get a titre of 307.5 and 242.6 U/ml of exo and endo PG respectively from Rhizopus oryzae (Zheng et al. 2017). Beads of alginate-montmorillonite were used to immobilize pectinase from A. aculeatus recovering 53% of its initial activity (Mohammadi et al. 2019). Gel-based beads of alginate and agar facilitate the immobilization of pectinase from A. awamori. This retained initial activity even after 8 cycles of reaction (Abdel Wahab et al. 2018). An indigenously isolated pectinolytic yeast strain, Geotrichum candidum was immobilized retaining 70% of its initial activity using corn cob matrix (Ejaz et al. 2018). Similarly, beads of sodium alginate were used in different strains of Geotrichum candidum to immobilize pectinase enhancing its activity from 0.046 to 0.115 IU mL−1 (Ejaz et al. 2020). Pectinases have also been immobilized using magnetic chitosan particles by direct extraction from fruit juices without the intervention of microbes (Dal Magro et al. 2018, 2019; Soozanipour et al. 2019). Efforts on the immobilization of pectinases from fungal strains have been summarized in Table 3.
Directed evolution
The state-of-the-art technology of directed evolution for the desired manipulation of enzymes for industrial application has been attempted for pectinases. Mutation using a UV range of 254 nm has been used for the enhancement of polygalacturonases production of Aspergillus and Penicillium species (Heerd et al. 2014; Kamalambigeswari et al. 2018; Nawaz et al. 2019). Mutated strains have also been used to study evolutionary relationships between PEL and PL subclasses of pectinases (Yang et al. 2020). Mutation of gaaX and gaaR allowed A. niger to express pectinases without an inducer (Alazi et al. 2019). The approach of directed evolution combined with computational technologies has been used to access different metabolic pathways of fungal pectinases (Wang et al. 2021). For fungal pectinases, artificial environments can be simulated through strain mutation, recombination, and gene overexpression. With this modification, the pectinolytic mechanism can be accelerated to catalyse chemical reactions in an entirely new environment employing a newer substrate, resulting in increased catalytic activity. Chromosomal mapping was used to analyse S. bayanus var. uvarum strains, and the results revealed three divergent genes, PGU1b, PGU2b, and PGU3b, which are situated on chromosomes X, I, and XIV, respectively. As a result, it was demonstrated that these yeasts’ strong pectinolytic activity might be caused by the existence of many PGU polymeric genes in their genomes (Naumova et al. 2019). Heterologous expression of fungal pectinase targeting expression using microbes with a high capacity for protein production and enzyme secretion has been performed. It is a good alternative to the fermentation technique for the desired production of enzymes by targeting the relevant genes. The expression of pectinolytic genes has been summarized in Table 4.
Omics interventions
The omics-driven approach is the current trend in enzyme research which aims to analyse the potential of fungal species in terms of enzyme production by targeting the whole genome or proteome. Over 50% of the currently available eukaryotic genome sequences are from the kingdom of Fungi. Several fungal genome sequences have been targeted to decipher the diversity of pectinases. Recently using a shotgun proteomics approach two pectin lyase and one pectate lyase from Saccharomyces cerevisiae produced using passion fruit flour by solid-state fermentation has been reported (Takeyama et al. 2022). Two-dimensional electrophoresis-based proteomic analysis of Aspergillus niger EIMU2 has been attempted. It revealed that the mutant EIMU2’s multiple enzyme systems used for the degradation of pectin included the main-chain cleaving enzymes polygalacturonase, pectate lyase, and pectin esterase, as well as some accessory enzymes rhamnogalacturonan lyase (Lin et al. 2021). Studying the interaction of wood rotting fungi, pectinases proteomics profiling helped analysed other proteins secreted which might have a significant role in degrading wood (Presley et al. 2020). CRISPR/Cas9 system generated three chimeric GaaR-XlnR induces by D-galacturonic acid from Aspergillus niger. Their proteomics investigation verified that the gaaR mutants carrying the chimeric transcription factor produced several pectinolytic enzymes (Kun et al. 2021). The PL7 and PL8 enzymes required for the breakdown of laminarin, cellulose, lipids, and peptides, were found to be abundantly secreted by Paradendryphiella salina cultured on brown algae using proteomic analysis (Pilgaard et al. 2019). However, a significant issue with the existing fungal pectinases proteomics is to fully understand the expression, operation, and regulation of the entire set of fungus-genome-encoded proteins. Moreover, the sequencing of several fungal proteomes is in progress (Sudhakar et al. 2018).
Meta-omics approach collects total environmental DNA which is targeted for metagenomic studies. A metagenomic system can be any arbitrary environmental sample defining the collection of microbes. Soil, water, air, cow rumen, and composts are such systems, thus, opening doors for unculturable and novel sources for catalytic enzymes. metagenomic approach for pectinase enzyme mining from soil resulted in the isolation of thermostable pectinase (Singh et al. 2012a, b). This approach has been used for identifying novel fungal sources for pectinases (Tanveer et al. 2016; Pilgaard et al. 2019; Ahmad et al. 2021). The metagenomic studies exclusively for fungal pectinases are summarized in Table 5.
Industrial applications
Pectin in plant cells is degraded by pectinases. They were first used commercially in the 1930s, and since then, they govern 25% of industrial applications. Wide-ranging industrial uses for pectin-degrading enzymes include degumming and retting of plant fibres, oil extraction, fruit juice clarification, wine production, fermentation of tea and coffee, bioconversion of wastes, and protoplast fusion technology (Singhania et al. 2015). Since 40% of the dry weight of plant cambium cells is made up of pectin, pectinases are essential for digesting natural fibres. With the aid of pectinases, the bast fibres of jute, flax, hemp, ramie, banana, pineapple leaf, and bamboo can be successfully degummed, macerated, and retted because they break down the pectin in the middle lamella and primary cell walls. Their wide applicability in the textile industry makes their study essential. Microbial pectinases-based natural fibre retting and extraction is biodegradable, recyclable, cuts production costs and is energy sustainable (Kumari et al. 2021). The fibres produced are reported with higher strength, shinier, easy to obtain and light weighted. The increasing demands on enzyme applications are growing as replacements for traditional harsh chemical processes. Fungal pectinases are also used for degumming natural fibres, bio scouring, bio bleaching and in wastewater treatment of textile power plants (Sharma et al. 2017).
They are also used to produce effective viral preparation from plant tissues, in the treatment of wastewater and for the isolation of protoplasts. Protoplasts are isolated from the mycelia of Pleurotuseous and Pleurotus flabellatus using enzymes comprising commercial cellulases, crude pectinases, and crude chitinases (Eyini et al. 2006; Ruiz et al. 2017). Pectinases are also applied in animal feeds as it helps in the efficient absorption of nutrients by animals by degrading the fibres that entrap them. These groups of enzymes have been used for biofuel production like bioethanol. The rate of ethanol generation rises when pectinaceous structures in the feedstock are destroyed and hydrolyzed by pectinases. Biomass enzymatic hydrolysis is a cost-effective and efficient treatment method that produces no hazardous waste (Samanta 2019). Sugar becomes more accessible and sensitive to hydrolytic enzymes after being treated with liquid hot water. Alkaline pectinases both from fungal and bacterial sources are also applied in the fermentation of coffee and tea. Degrading pectin, pectinase increases the pace of tea fermentation and reduces the foaming ability of instant tea granules (Tatta et al. 2022).
The fruit and food processing industries have wide applicability of pectinases. Fruits have a complicated pectin structure, making it challenging to extract juice from this very viscous, jellified pulp (Pagnonceli et al. 2019). The pectinase enzyme acts on the pectin of fruit peels and dissolves the glycosidic linkages between the galacturonic acid monomers, reducing the amount of water that may be held by pectin enzymatic treatment is the most frequently used method for juice extraction and clarity (Anand et al. 2017b). The enzymatic hydrolysis of cell walls enhances the extraction yield, soluble dry matter content, galacturonic acid content, and titratable acidity of the products. The amount of waste pomace decreased and the resulting pulp had a lower viscosity. The biomaterial is enzymatically degraded depending on the type of enzyme, incubation period, temperature, concentration, agitation, pH, and the use of various enzyme combinations. The wine industry chooses pectinases as they increase wine quality, and facilitate extraction, filtering, and taste and colour intensification (Gunjal et al. 2020). Pectinases were also used in extracting essential oils from a variety of sources like olives, flaxseed oil, dates, and other fruits and vegetables (Nagpal et al. 2021). These enzymes help to enhance the fatty acids, peroxide value, and colour intensity as compared to chemical treatment. In the paper industry, pectinases along with xylanases are preferred as a bio-bleaching agent. Enzymatic intervention is eco-friendly, less abrasive, and effective in improving paper quality (Nagpal et al. 2020). Biological bleaching with pectinases and xylanases brightens the paper and improves its physical characteristics, as well as lowers the kappa number and permanganate number of the pulp. In comparison to those chemical alternative solutions, the substitution of pectinases contributes to a reduction in chlorine discharge into the environment (Nagpal et al. 2020; Tatta et al. 2022). The diverse industrial application of pectinases has been summarized in Table 6.
The bottom line and future prospects
Pectinases represent an important group of enzymes with immense potential for diverse industrial applications. Substantial efforts have been made to explore the possibility of diverse approaches for enhancing pectinases production, manipulation and elucidating industrial applications, exclusively from fungal sources. The cost-effective production of fungal pectinases using agro-wastes is an eco-friendly approach that has immense potential for converting waste biomass. It also results in the production of different value-added products. This is also added to the saccharification potential of pectinases. Efforts have been made to optimize growth conditions as a precursor to enhanced fungal bioproduct production. Utilising waste valorisation techniques, it is possible to take advantage of the diversity of fungi by using contaminated items as a source of fungi. The fungus system offers many advantages and benefits, but it also poses a hazard due to its pathogenicity and ability to mitigate spoilage and damage. Recombinant and mutagenic approaches can be used to change the pathogenicity of native fungus hosts. According to industrial needs, the fusion of traditional and modern state-of-the-art technology has enormous potential.
Over the years, several fungal genera have been targeted for the production of pectinases and efforts have been made to enhance the catalytic activity, specificity, and applicability for industrial applications. Dual culture inoculums for fermentation-based manufacturing have been employed to increase enzyme productivity. These involve using more than one fungal species for the production of the same biocatalyst. But they strictly demand more comprehension of how various hosts interact with one another. The metagenomics approach has resulted in the deciphering of novel microbes with enhanced pectinase activity, thereby giving the world new industrially potent species. Despite metagenomics inclination in microbial studies, fungal metagenomic library construction and diversity studies are minimal. Though purity of metagenomic DNA from humic acid contamination and the easy extraction of prokaryotic diversity in metagenomics DNA limits the studies of pectinases of fungal metagenomic origin from s potential. The directed evolution approach for altered pectinases activity and specificity has resulted in diverse industrial applications predominately in the textile and food industries. Omics-driven approaches including genomics, proteomics, and metabolomics have been used for understanding the production and expression of pectinase genes. Sequencing of fungal strains, genome-wide mining of pectinases using a bioinformatics approach, and expression of the identified pectinases are intensely investigated areas of research in fungal pectinases. Immobilisation of fungal pectinases using novel approaches for enhancing stability and reuse for industrial application has also been attempted.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article. Data sharing does not apply to this article as no new data were created or analysed in this study.
References
Abd El-Rahim WM, Moawad H, Hashem MM et al (2020) Highly efficient fungal pectinase and laccase producers among isolates from flax retting liquor. Bicoastal Agric Biotechnol 25:101570. https://doi.org/10.1016/j.bcab.2020.101570
Abdel Wahab WA, Karam EA, Hassan ME et al (2018) Optimization of pectinase immobilization on grafted alginate-agar gel beads by 24 full factorial CCD and thermodynamic profiling for evaluating of operational covalent immobilization. Int J Biol Macromol 113:159–170. https://doi.org/10.1016/j.ijbiomac.2018.02.086
Abdullah R, Farooq I, Kaleem A et al (2018a) Pectinase production from Aspergillus niger IBT-7 using solid-state fermentation. Bangladesh J Bot 47:473–478. https://doi.org/10.3329/bjb.v47i3.38714
Abdullah R, Jafer A, Nisar K et al (2018b) Process optimization for pectinase production by locally isolated fungal strain using submerged fermentation. Biosci J. https://doi.org/10.14393/BJ-v34n1a2018-39947
Abdulrachman D, Thongkred P, Kocharin K et al (2017) Heterologous expression of Aspergillus aculeatus endo-polygalacturonase in Pichia pastoris by high cell density fermentation and its application in textile scouring. BMC Biotechnol 17:15. https://doi.org/10.1186/s12896-017-0334-9
Adedeji OE, Ezekiel OO (2019) Pretreatment of selected peels for polygalacturonase production by Aspergillus awamori CICC 2040: purification and application in mango juice extraction. Bioresour Technol Rep 7:100306. https://doi.org/10.1016/j.biteb.2019.100306
Adeleke AJ, Odunfa SA, Olanbiwonninu A, Owoseni MC (2012) Production of cellulase and pectinase from orange peels by fungi. Nat Sci 10:107–112
Aggarwal R, Dutta T, Sheikh J (2020) Extraction of pectinase from Candida isolated from textile mill effluent and its application in bio-scouring of cotton. Sustain Chem Pharm 17:100291. https://doi.org/10.1016/j.scp.2020.100291
Ahmad T, Gupta G, Sharma A et al (2021) Metagenomic analysis exploring taxonomic and functional diversity of bacterial communities of a Himalayan urban fresh water lake. PLoS ONE 16:e0248116. https://doi.org/10.1371/journal.pone.0248116
Ahmed A, Sohail M (2020) Characterization of pectinase from Geotrichum candidum AA15 and its potential application in orange juice clarification. J King Saud Univ Sci 32:955–961. https://doi.org/10.1016/j.jksus.2019.07.002
Ahmed I, Zia MA, Hussain MA et al (2016) Bioprocessing of citrus waste peel for induced pectinase production by Aspergillus niger; its purification and characterization. J Radiat Res Appl Sci 9:148–154. https://doi.org/10.1016/j.jrras.2015.11.003
Ahmed J, Thakur A, Goyal A (2021) Emerging trends on the role of recombinant pectinolytic enzymes in industries—an overview. Biocatal Agric Biotechnol 38:102200. https://doi.org/10.1016/j.bcab.2021.102200
Ajayi AA, Lawal B, Salubi AE et al (2021) Pectinase production by Aspergillus niger using pineapple peel pectin and its application in coconut oil extraction. IOP Conf Ser Earth Environ Sci 655:012014. https://doi.org/10.1088/1755-1315/655/1/012014
Alazi E, Niu J, Otto SB et al (2019) W361R mutation in GaaR, the regulator of D-galacturonic acid-responsive genes, leads to constitutive production of pectinases in Aspergillus niger. Microbiologyopen 8:e00732. https://doi.org/10.1002/mbo3.732
Almowallad SA, Alshammari GM, Alsayadi MM et al (2022) Partial purification and characterization of exo-polygalacturonase produced by Penicillium oxalicum AUMC 4153. Life 12:284. https://doi.org/10.3390/life12020284
Amin F, Bhatti HN, Bilal M, Asgher M (2017) Purification, kinetic, and thermodynamic characteristics of an exo-polygalacturonase from Penicillium notatum with industrial perspective. Appl Biochem Biotechnol 183:426–443. https://doi.org/10.1007/s12010-017-2455-y
Amin F, Mohsin A, Bhatti HN, Bilal M (2020) Production, thermodynamic characterization, and fruit juice quality improvement characteristics of an exo-polygalacturonase from Penicillium janczewskii. Biochimica Biophysica Acta 1868:140379. https://doi.org/10.1016/j.bbapap.2020.140379
Anand G, Yadav S, Yadav D (2017a) Purification and biochemical characterization of an exo-polygalacturonase from Aspergillus flavus MTCC 7589. Biocatal Agric Biotechnol 10:264–269. https://doi.org/10.1016/j.bcab.2017.03.018
Anand G, Yadav S, Yadav D (2017b) Production, purification and biochemical characterization of an exo-polygalacturonase from Aspergillus niger MTCC 478 suitable for clarification of orange juice. 3 Biotech 7:122. https://doi.org/10.1007/s13205-017-0760-3
Anand G, Yadav S, Gupta R, Yadav D (2020) Pectinases: from microbes to industries. In: Chowdhary P, Raj A, Verma D, Akhter Y (eds) Microorganisms for sustainable environment and health. Elsevier, Amsterdam, pp 287–313
Anderson CT (2019) Pectic polysaccharides in plants: structure, biosynthesis, functions, and applications. Springer, New York, pp 487–514
Anuradha K, Padma PN, Venkateshwar S, Reddy G (2016) Mango juice clarification with polygalacturonase produced by Aspergillus awamori MTCC 9166-Optimization of conditions. Int Food Res J 23:147
Arun KB, Madhavan A, Sindhu R et al (2020) Remodelling agro-industrial and food wastes into value-added bioactive and biopolymers. Ind Crops Prod 154:112621. https://doi.org/10.1016/j.indcrop.2020.112621
Atanasova L, Dubey M, Grujić M et al (2018) Evolution and functional characterization of pectate lyase PEL12, a member of a highly expanded Clonostachys rosea polysaccharide lyase 1 family. BMC Microbiol 18:178. https://doi.org/10.1186/s12866-018-1310-9
Azzaz HH, Murad HA, Hassaan NA, Fahmy M (2019) Pectinase production optimization for improving dairy animal’s diets degradation. Int J Dairy Sci 15:54–61. https://doi.org/10.3923/ijds.2020.54.61
Bader AN, Sanchez Rizza L, Consolo VF, Curatti L (2020) Efficient saccharification of microalgal biomass by Trichoderma harzianum enzymes for the production of ethanol. Algal Res 48:101926. https://doi.org/10.1016/j.algal.2020.101926
Baldrian P (2019) The known and the unknown in soil microbial ecology. FEMS Microbiol Ecol 95:fiz005. https://doi.org/10.1093/femsec/fiz005
Banakar SP, Thippeswamy B (2014) Isolation, production and partial purification of fungal extracellular pectinolytic enzymes from the forest soils of Bhadra Wildlife Sanctuary, Western Ghats of Southern India. J Biochem Technol 3:138–143
Banu AR, Devi MK, Gnanaprabhal GR et al (2010) Production and characterization of pectinase enzyme from Penicillium chrysogenum. Indian J Sci Technol 4:377–381
Bashir N, Sood M, Bandral JD (2020) Enzyme immobilization and its applications in food processing: a review. Int J Chem Stud 8:254–261. https://doi.org/10.22271/chemi.2020.v8.i2d.8779
Bassim Atta M, Ruiz-Larrea F (2022) Fungal pectinases in food technology. In: Pectins—the new-old polysaccharides. Intech Open
Belda I, Conchillo LB, Ruiz J et al (2016) Selection and use of pectinolytic yeasts for improving clarification and phenolic extraction in winemaking. Int J Food Microbiol 223:1–8. https://doi.org/10.1016/j.ijfoodmicro.2016.02.003
Benen JA, van Alebeek GJW, Voragen AG, Visser J (2002) Pectic esterases. In: Whitaker JR, Voragen AG, Wong DW (eds) Handbook of food enzymology, 1st edn. CRC Press, Boca Raton, pp 864–871
Bezawada P, Raju KJ (2018) Screening of pectinolytic fungi and optimization of process parameters using guava peel powder as substrate under solid state fermentation. Int J Eng Sci Invention 7:43–47
Bharathiraja S, Suriya J, Krishnan M, et al (2017) Production of enzymes from agricultural wastes and their potential industrial applications, pp 125–148
Bhattacharyya R, Mukhopadhyay D, Nagarakshita VK et al (2021) Thermostable and organic solvent-tolerant acid pectinase from Aspergillus terreus FP6: purification, characterization and evaluation of its phytopigment extraction potential. 3 Biotech 11:487. https://doi.org/10.1007/s13205-021-03033-x
Bonnin E, Pelloux J (2020) Pectin degrading enzymes. In: Kontogiorgos V (ed) Pectin: technological and physiological properties. Springer, Cham, pp 37–60
Cairns TC, Barthel L, Meyer V (2021) Something old, something new: challenges and developments in Aspergillus niger biotechnology. Essays Biochem 65:213–224. https://doi.org/10.1042/EBC20200139
Cano ME, García-Martin A, Comendador Morales P et al (2020) Production of oligosaccharides from agrofood wastes. Fermentation 6:31. https://doi.org/10.3390/fermentation6010031
Cantarel BL, Coutinho PM, Rancurel C et al (2009) The Carbohydrate-Active Enzymes database (CAZy): an expert resource for Glycogen omics. Nucleic Acids Res 37:D233–D238. https://doi.org/10.1093/nar/gkn663
Cavello I, Albanesi A, Fratebianchi D et al (2017) Pectinolytic yeasts from cold environments: novel findings of Guehomyces pullulans, Cystofilobasidium infirmominiatum and Cryptococcus adeliensis producing pectinases. Extremophiles 21:319–329. https://doi.org/10.1007/s00792-016-0904-0
cazy.org/Carbohydrate-Esterases Carbohydrate Esterase family classification. http://www.cazy.org/Carbohydrate-Esterases.html
cazy.org/Glycoside-Hydrolases. Glycoside Hydrolase family classification. http://www.cazy.org/Glycoside-Hydrolases.html
cazy.org/Polysaccharide-Lyases Polysaccharide Lyase family classification. http://www.cazy.org/Polysaccharide-Lyases.html
Cerda A, Artola A, Barrena R et al (2019) Innovative production of bioproducts from organic waste through solid-state fermentation. Front Sustain Food Syst 3:63. https://doi.org/10.3389/fsufs.2019.00063
Chakraborty S, Rani A, Dhillon A, Goyal A (2017) Polysaccharide Lyases. In: Ashok PA, Sangeeta Negi S, Soccol CR (eds) Current developments in biotechnology and bioengineering. Elsevier, Amsterdam, pp 527–539
Chang H-X, Yendrek CR, Caetano-Anolles G, Hartman GL (2016) Genomic characterization of plant cell wall degrading enzymes and in silico analysis of xylanases and polygalacturonases of Fusarium virguliforme. BMC Microbiol 16:147. https://doi.org/10.1186/s12866-016-0761-0
Chen H, Wang L (2017) Microbial fermentation strategies for biomass conversion. In: Technologies for biochemical conversion of biomass. Elsevier, Amsterdam, pp 165–196
Chen X, Li L, He Z et al (2018) Molecular cloning and functional analysis of two novel polygalacturonase genes in Rhizoctonia solani. Can J Plant Path 40:39–47. https://doi.org/10.1080/07060661.2017.1417915
Cheng Z, Chen D, Lu B et al (2016) A novel acid-stable endo-polygalacturonase from Penicillium oxalicum CZ1028: purification, characterization, and application in the beverage industry. J Microbiol Biotechnol 26:989–998. https://doi.org/10.4014/jmb.1511.11045
Cheng Z, Chen D, Wang Q et al (2017) Identification of an acidic endo-polygalacturonase from Penicillium oxalicum CZ1028 and its broad use in major tropical and subtropical fruit juices production. J Biosci Bioeng 123:665–672. https://doi.org/10.1016/j.jbiosc.2017.01.013
Cheng Z, Xian L, Chen D et al (2020) Development of an innovative process for high-temperature fruit juice extraction using a novel thermophilic endo-polygalacturonase from Penicillium oxalicum. Front Microbiol 11:1200. https://doi.org/10.3389/fmicb.2020.01200
Chilakamarry CR, Mimi Sakinah AM, Zularisam AW et al (2022) Advances in solid-state fermentation for bioconversion of agricultural wastes to value-added products: opportunities and challenges. Bioresour Technol 343:126065. https://doi.org/10.1016/j.biortech.2021.126065
Christensen SH (2020) Pectins. In: Glicksman M (ed) Food hydrocolloids. CRC Press, Boca Raton, pp 205–230
Chukwuma OB, Rafatullah M, Tajarudin HA, Ismail N (2020) Lignocellulolytic enzymes in biotechnological and industrial processes: a review. Sustainability 12:7282. https://doi.org/10.3390/su12187282
Coskun O (2016) Separation techniques: chromatography. North Clin Istanb. https://doi.org/10.14744/nci.2016.32757
Dal Magro L, Silveira VCC, de Menezes EW et al (2018) Magnetic biocatalysts of pectinase and cellulase: synthesis and characterization of two preparations for application in grape juice clarification. Int J Biol Macromol 115:35–44. https://doi.org/10.1016/j.ijbiomac.2018.04.028
Dal Magro L, de Moura KS, Backes BE et al (2019) Immobilization of pectinase on chitosan-magnetic particles: Influence of particle preparation protocol on enzyme properties for fruit juice clarification. Biotechnol Rep 24:e00373. https://doi.org/10.1016/j.btre.2019.e00373
Davanso M, Atsakou AE, Gattás EA, de Paula AV (2019) Assessment of pectinase-producing fungi isolated from soil and the use of orange waste as a substrate for pectinase production. Revista De Ciências Farmacêuticas Básica e Aplicada 40:1–5
de Alencar Cavalieri, Guimarães N, Glienke NN, Silva Galeano RM et al (2022) Polygalacturonase from Aspergillus japonicus (PGAj): Enzyme production using low-cost carbon source, biochemical properties and application in clarification of fruit juices. Biocatal Agric Biotechnol 39:102233. https://doi.org/10.1016/j.bcab.2021.102233
de Oliveira RL, Dias JL, da Silva OS, Porto TS (2018) Immobilization of pectinase from Aspergillus aculeatus in alginate beads and clarification of apple and umbu juices in a packed bed reactor. Food Bioprod Process 109:9–18. https://doi.org/10.1016/j.fbp.2018.02.005
de Souza TSP, Kawaguti HY (2021) Cellulases, hemicellulases, and pectinases: applications in the food and beverage industry. Food Bioproc Tech 14:1446–1477. https://doi.org/10.1007/s11947-021-02678-z
Deng Z, Wang F, Zhou B et al (2019) Immobilization of pectinases into calcium alginate microspheres for fruit juice application. Food Hydrocoll 89:691–699. https://doi.org/10.1016/j.foodhyd.2018.11.031
Dhital R, Panta OP, Karki TB (2014) Optimization of cultural conditions for the production of pectinase from selected fungal strain. J Food Sci Technol Nepal 8:65–70. https://doi.org/10.3126/jfstn.v8i0.11752
Dong Z, Wang Z (2011) Isolation and characterization of an exo polygalacturonase from Fusarium oxysporum f.sp. cubense race 1 and race 4. BMC Biochem 12:51. https://doi.org/10.1186/1471-2091-12-51
Dong Z, Wang Z (2015) Isolation and heterologous expression of a polygalacturonase produced by Fusarium oxysporum f. sp. cubense Race 1 and 4. Int J Mol Sci 16:7595–7607. https://doi.org/10.3390/ijms16047595
Dong Z, Luo M, Wang Z (2020) An exo-polygalacturonase Pgc4 regulates aerial hyphal growth and virulence in Fusarium oxysporum f. sp. cubense race 4. Int J Mol Sci 21:5886. https://doi.org/10.3390/ijms21165886
Drula E, Garron M-L, Dogan S et al (2022) The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res 50:D571–D577. https://doi.org/10.1093/nar/gkab1045
Ejaz U, Ahmed A, Sohail M (2018) Statistical optimization of immobilization of yeast cells on corncob for pectinase production. Biocatal Agric Biotechnol 14:450–456. https://doi.org/10.1016/j.bcab.2018.04.011
Ejaz U, Hanif H, Sohail M (2020) Two layered strategy for cost effective production of pectinase: immobilization of yeast and utilization of crude substrate. Heliyon 6:e05456. https://doi.org/10.1016/j.heliyon.2020.e05456
El-Ghomary AE, Shoukry AA, El-Kotkat MB (2021) Productivity of pectinase enzymes by Aspergillus sp. isolated from Egyptian soil. Al-Azhar J Agric Res 46:79–87. https://doi.org/10.21608/ajar.2021.245617
Esawy MA, Gamal AA, Kamel Z et al (2013) Evaluation of free and immobilized Aspergillus niger NRC1ami pectinase applicable in industrial processes. Carbohydr Polym 92:1463–1469. https://doi.org/10.1016/j.carbpol.2012.10.061
Esawy MA, Gamal AA, Kamel Z (2022) Optimization of Aspergillus niger NRC1ami pectinase using citrus peel pectin, purification, and thermodynamic characterization of the free and modified enzyme. Waste Biomass Valoriz. https://doi.org/10.1007/s12649-022-01838-2
Eyini M, Rajkumar K, Balaji P (2006) Isolation, regeneration and PEG-induced fusion of protoplasts of Pleurotus pulmonarius and Pleurotus florida. Mycobiology 34:73. https://doi.org/10.4489/MYCO.2006.34.2.073
Favela-Torres E, Aguilar C, Contreras-Esquivel JC, Viniegra-González G (2006) Pectinases. In: Pandey A, Webb C, Soccol CR, Larroche C (eds) Enzyme technology. Springer, New York, pp 273–296
Finkler ATJ, Biz A, Pitol LO et al (2017) Intermittent agitation contributes to uniformity across the bed during pectinase production by Aspergillus niger grown in solid-state fermentation in a pilot-scale packed-bed bioreactor. Biochem Eng J 121:1–12. https://doi.org/10.1016/j.bej.2017.01.011
Food enzyme trend gminsight food enzymes market size by product (proteases, lipases, carbohydrases [amylases, xylanases, cellulases, pectinases, lactases], polymerases & nucleases, phytases, catalases), by application (food & beverage, processed food, diary, bakery, confectionary), industry analysis report, regional outlook, growth potential, Covid-19 impact analysis, price trend, competitive market share & forecast, 2021–2027. https://www.gminsights.com/industry-analysis/food-enzymes-market.
Fratebianchi D, Crespo JM, Tari C, Cavalitto S (2017) Control of agitation rate and aeration for enhanced polygalacturonase production in submerged fermentation by Aspergillus sojae using agro-industrial wastes. J Chem Technol Biotechnol 92:305–310. https://doi.org/10.1002/jctb.5006
Gacura MD, Sprockett DD, Heidenreich B, Blackwood CB (2016) Comparison of pectin-degrading fungal communities in temperate forests using glycosyl hydrolase family 28 pectinase primers targeting Ascomycete fungi. J Microbiol Methods 123:108–113. https://doi.org/10.1016/j.mimet.2016.02.013
Garlapati VK (2015) Isolation and screening of fungal isolates for multienzyme production through submerged and solid-state fermentations. J Bioprocess Biotech 5:1. https://doi.org/10.4172/2155-9821.1000249
Gawkowska D, Cybulska J, Zdunek A (2018) Structure-related gelling of pectins and linking with other natural compounds: a review. Polymers 10:762. https://doi.org/10.3390/polym10070762
Ghosh R, Kar R, Bhattacharyya S, Majumdar S (2015) Efficient retting of bast fibre yielding stems by extracellular enzyme conglomerate and oxalic acid produced by a newly isolated Penicillium sp. Int J Adv Res 3:291–300
Gunjal AB, Patil NN, Shinde SS (2020) Pectinase in degradation of lignocellulosic wastes. In: Enzymes in degradation of the lignocellulosic wastes. Springer, Cham, pp 71–103
Gutierrez-Alvarado K, Chacón-Cerdas R, Starbird-Perez R (2022) Pectin microspheres: synthesis methods, properties, and their multidisciplinary applications. Chemistry 4:121–136. https://doi.org/10.3390/chemistry4010011
Gutiérrez-Correa M, Ludeña Y, Ramage G, Villena GK (2012) Recent advances on filamentous fungal biofilms for industrial uses. Appl Biochem Biotechnol 167:1235–1253. https://doi.org/10.1007/s12010-012-9555-5
Haile S, Ayele A (2022) Pectinase from microorganisms and its industrial applications. Sci World J 2022:1–15. https://doi.org/10.1155/2022/1881305
Harholt J, Suttangkakul A, Vibe Scheller H (2010) Biosynthesis of pectin. Plant Physiol 153:384–395. https://doi.org/10.1104/pp.110.156588
Hassan SS, Tiwari BK, Williams GA, Jaiswal AK (2020a) Bioprocessing of brewers’ spent grain for production of xylan pectinolytic enzymes by Mucor sp. Bioresour Technol Rep 9:100371. https://doi.org/10.1016/j.biteb.2019.100371
Hassan SS, Williams GA, Jaiswal AK (2020b) Computational modelling approach for the optimization of apple juice clarification using immobilized pectinase and xylanase enzymes. Curr Res Food Sci 3:243–255. https://doi.org/10.1016/j.crfs.2020.09.003
He Y, Pan L, Wang B (2018) Efficient over-expression and application of high-performance pectin lyase by screening Aspergillus niger pectin lyase gene family. Biotechnol Bioprocess Eng 23:662–669. https://doi.org/10.1007/s12257-018-0387-1
Heerd D, Tari C, Fernández-Lahore M (2014) Microbial strain improvement for enhanced polygalacturonase production by Aspergillus sojae. Appl Microbiol Biotechnol 98:7471–7481. https://doi.org/10.1007/s00253-014-5657-z
Holm HC, Nielsen PM, Longin F (2018) Upscaling of enzymatic processes. In: Lipid modification by enzymes and engineered microbes. Elsevier, pp 343–373
Huang D, Song Y, Liu Y, Qin Y (2019) A new strain of Aspergillus tubingensis for high-activity pectinase production. Braz J Microbiol 50:53–65. https://doi.org/10.1007/s42770-018-0032-3
Hyde KD, Xu J, Rapior S et al (2019) The amazing potential of fungi: 50 ways we can exploit fungi industrially. Fungal Divers 97:1–136. https://doi.org/10.1007/s13225-019-00430-9
Ibrahim E, Jones KD, Taylor KE et al (2017) Molecular and biochemical characterization of recombinant cel12B, cel8C, and peh28 overexpressed in Escherichia coli and their potential in biofuel production. Biotechnol Biofuels 10:52. https://doi.org/10.1186/s13068-017-0732-1
Ibrahim NA, Eid BM, Abdel Aziz MS et al (2019) Environmentally benign scouring of cotton knits using locally produced acid pectinase enzyme. Fibers Polym 20:787–793. https://doi.org/10.1007/s12221-019-1207-8
Ire F, Vinking E (2016) Production, purification and characterization of polygalacturonase from Aspergillus niger in solid state and submerged fermentation using banana peels. J Adv Biol Biotechnol 10:1–15. https://doi.org/10.9734/JABB/2016/29593
Islam S, Feroza B, Alam A, Begum S (2013) Pectinase production by Aspergillus niger isolated from decomposed apple skin. Bangladesh J Sci Ind Res 48:25–32. https://doi.org/10.3329/bjsir.v48i1.15410
Ismail A-MS, Abo-Elmagd HI, Housseiny MM (2016) A safe potential juice clarifying pectinase from Trichoderma viride EF-8 utilizing Egyptian onion skins. J Genet Eng Biotechnol 14:153–159. https://doi.org/10.1016/j.jgeb.2016.05.001
Jagajanantha P, Morey M, Satankar V, Mageshwaran V (2022) Bio-scouring of non-spinnable cotton by a crude enzyme of a new fungal strain Aspergillus sp. VM-1, isolated from banana pseudo stem waste. Waste Biomass Valoriz 13:1849–1858. https://doi.org/10.1007/s12649-021-01621-9
Jaramillo PMD, Andreaus J, de Neto GPS et al (2015) The characterization of a pectin-degrading enzyme from Aspergillus oryzae grown on passion fruit peel as the carbon source and the evaluation of its potential for industrial applications. Biocatal Biotransform 33:310–322. https://doi.org/10.3109/10242422.2016.1168817
Jayani RS, Saxena S, Gupta R (2005) Microbial pectinolytic enzymes: a review. Process Biochem 40:2931–2944. https://doi.org/10.1016/j.procbio.2005.03.026
John J, Kamal KKS, Smith ML et al (2020) Advances in upstream and downstream strategies of pectinase bioprocessing: a review. Int J Biol Macromol 162:1086–1099. https://doi.org/10.1016/j.ijbiomac.2020.06.224
Joshi S, Mohapatra B, Mishra JPN (2018) Microbial soil enzymes: implications in the maintenance of rhizosphere ecosystem and soil health, pp 179–192
Kamalambigeswari R, Alagar S, Sivvaswamy N (2018) Strain improvement through mutation to enhance pectinase yield from Aspergillus niger and molecular characterization of polygalacturonase gene. J Pharm Sci Res 10:989–994
Karataş E, Tülek A, Çakar MM et al (2021) From secretion in Pichia pastoris to application in apple juice processing: exo-Polygalacturonase from Sporothrix schenckii 1099–18. Protein Pept Lett 28:817–830. https://doi.org/10.2174/1871530321666210106110400
Kashyap DR, Vohra PK, Chopra S, Tewari R (2001) Applications of pectinases in the commercial sector: a review. Bioresour Technol 77:215–227. https://doi.org/10.1016/S0960-8524(00)00118-8
Kaur A, Kaur J (2019) Cultivation strategies with special reference to bioreactor design and operation for industrial production in biotechnology. In: Biotechnology of microorganisms. Apple Academic Press, Series statement: Innovations in biotechnology, vol 2, pp 23–44
Kc S, Upadhyaya J, Joshi DR et al (2020) Production, characterization, and industrial application of pectinase enzyme isolated from fungal strains. Fermentation 6:59. https://doi.org/10.3390/fermentation6020059
Ketipally R, Kumar GK, Ram MR (2019) Polygalacturonase production by Aspergillus nomius MR103 in solid state fermentation using Agro-industrial wastes. J Appl Nat Sci 11:305–310. https://doi.org/10.31018/jans.v11i2.2039
Khan S, Saleem S, Azhar A (2014) Isolation and selection of Aspergillus species for hyper-production of polygalacturonases. Pak J Biochem Mol Biol 47–125
Khatri BP, Bhattarai T, Shrestha S, Maharjan J (2015) Alkaline thermostable pectinase enzyme from Aspergillus niger strain MCAS2 isolated from Manaslu Conservation Area, Gorkha. Nepal Springerplus 4:488. https://doi.org/10.1186/s40064-015-1286-y
Kohli P, Gupta R (2015) Alkaline pectinases: a review. Biocatal Agric Biotechnol 4:279–285. https://doi.org/10.1016/j.bcab.2015.07.001
Kordi M, Salami R, Bolouri P et al (2022) White biotechnology and the production of bio-products. Syst Microbiol Biomanuf 2:413–429. https://doi.org/10.1007/s43393-022-00078-8
Kumar B, Verma P (2020) Enzyme mediated multi-product process: a concept of bio-based refinery. Ind Crops Prod 154:112607. https://doi.org/10.1016/j.indcrop.2020.112607
Kumar YS, Kumar PV, Reddy OVS (2012) Pectinase production from mango peel using Aspergillus foetidus and its application in processing of mango juice. Food Biotechnol 26:107–123. https://doi.org/10.1080/08905436.2012.670830
Kumar V, Ahluwalia V, Saran S et al (2021) Recent developments on solid-state fermentation for production of microbial secondary metabolites: challenges and solutions. Bioresour Technol 323:124566. https://doi.org/10.1016/j.biortech.2020.124566
Kumari M, Padhi S, Sharma S et al (2021) Biotechnological potential of psychrophilic microorganisms as the source of cold-active enzymes in food processing applications. 3 Biotech 11:479. https://doi.org/10.1007/s13205-021-03008-y
Kun RS, Garrigues S, di Falco M et al (2021) The chimeric GaaR-XlnR transcription factor induces pectinolytic activities in the presence of D-xylose in Aspergillus niger. Appl Microbiol Biotechnol 105:5553–5564. https://doi.org/10.1007/s00253-021-11428-2
Ladeira Ázar RIS, da Luz MM, Piccolo Maitan-Alfenas G et al (2020) Apple juice clarification by a purified polygalacturonase from Calonectria pteridis. Food Bioprod Process 119:238–245. https://doi.org/10.1016/j.fbp.2019.11.013
Laha S, Sarkar D, Chaki S (2014) Research article optimization of production and molecular characterization of pectinase enzyme produced from Penicillium chrysogenum. 326–335
Leite P, Sousa D, Fernandes H et al (2021) Recent advances in production of lignocellulolytic enzymes by solid-state fermentation of agro-industrial wastes. Curr Opin Green Sustain Chem 27:100407. https://doi.org/10.1016/j.cogsc.2020.100407
Li Q, Al Loman A, Callow NV et al (2018) Leveraging pH profiles to direct enzyme production (cellulase, xylanase, polygalacturonase, pectinase, α-galactosidase, and invertase) by Aspergillus foetidus. Biochem Eng J 137:247–254. https://doi.org/10.1016/j.bej.2018.06.008
Li Q, Ray CS, Callow NV et al (2020) Aspergillus niger production of pectinase and α-galactosidase for enzymatic soy processing. Enzyme Microb Technol 134:109476. https://doi.org/10.1016/j.enzmictec.2019.109476
Lin W, Xu X, Lv R et al (2021) Differential proteomics reveals main determinants for the improved pectinase activity in UV-mutagenized Aspergillus niger strain. Biotechnol Lett 43:909–918. https://doi.org/10.1007/s10529-020-03075-w
Liu C-Q, Hu K-D, Li T-T et al (2017a) Polygalacturonase gene pgxB in Aspergillus niger is a virulence factor in apple fruit. PLoS ONE 12:e0173277. https://doi.org/10.1371/journal.pone.0173277
Liu M, Ale MT, Kołaczkowski B et al (2017b) Comparison of traditional field retting and Phlebia radiata Cel 26 retting of hemp fibres for fibre-reinforced composites. AMB Express 7:58. https://doi.org/10.1186/s13568-017-0355-8
Liu C, Zhang L, Tan L et al (2021) Immobilized crosslinked pectinase preparation on porous ZSM-5 zeolites as reusable biocatalysts for ultra-efficient hydrolysis of β-glycosidic bonds. Front Chem. https://doi.org/10.3389/fchem.2021.677868
Lizardi-Jiménez MA, Hernández-Martínez R (2017) Solid state fermentation (SSF): diversity of applications to valorize waste and biomass. 3 Biotech 7:44. https://doi.org/10.1007/s13205-017-0692-y
Lodhi MS, Shaheen A, Khan MT et al (2022) A novel method of affinity purification and characterization of polygalacturonase of Aspergillus flavus by galacturonic acid engineered magnetic nanoparticle. Food Chem 372:131317. https://doi.org/10.1016/j.foodchem.2021.131317
Lopes FC, Ligabue-Braun R (2021) Agro-industrial residues: eco-friendly and inexpensive substrates for microbial pigments production. Front Sustain Food Syst. https://doi.org/10.3389/fsufs.2021.589414
Lu X, Lin J, Wang C et al (2016) Purification and characterization of exo-polygalacturonase from Zygoascus hellenicus V25 and its potential application in fruit juice clarification. Food Sci Biotechnol 25:1379–1385. https://doi.org/10.1007/s10068-016-0215-3
Lu B, Xian L, Zhu J et al (2022) A novel endo-polygalacturonase from Penicillium oxalicum: gene cloning, heterologous expression and its use in acidic fruit juice extraction. J Microbiol Biotechnol 32:464–472. https://doi.org/10.4014/jmb.2112.12023
Lübeck M, Lübeck PS (2022) Fungal cell factories for efficient and sustainable production of proteins and peptides. Microorganisms 10:753. https://doi.org/10.3390/microorganisms10040753
Ma Y, Sun S, Hao H, Xu C (2016) Production, purification and characterization of an exo-polygalacturonase from Penicillium janthinellum sw09. An Acad Bras Cienc 88:479–487. https://doi.org/10.1590/0001-3765201620150051
Mahmoodi M, Najafpour GD, Mohammadi M (2017) Production of pectinases for quality apple juice through fermentation of orange pomace. J Food Sci Technol 54:4123–4128. https://doi.org/10.1007/s13197-017-2829-8
Mahmoodi M, Najafpour GD, Mohammadi M (2019) Bioconversion of agro-industrial wastes to pectinases enzyme via solid state fermentation in trays and rotating drum bioreactors. Biocatal Agric Biotechnol 21:101280. https://doi.org/10.1016/j.bcab.2019.101280
Makky EA, Yusoff M (2014) Bioeconomy: fermented waste management and pectinases purification from Thermomyceslanuginosus. J Mech Eng Sci 7:1196–1207. https://doi.org/10.15282/jmes.7.2014.19.0117
Martin N, Mau G, Sette LD et al (2010) Pectinase production by a Brazilian thermophilic fungus Thermomucor indicae-seudaticae N31 in solid-state and submerged fermentation. Microbiology 79:306–313. https://doi.org/10.1134/S0026261710030057
Martín MC, López OV, Ciolino AE et al (2019) Immobilization of ecological pectinase in calcium alginate hydrogels: a potential biocatalyst for winemaking. Biocatal Agric Biotechnol 18:101091. https://doi.org/10.1016/j.bcab.2019.101091
Mat Jalil MT, Ibrahim D (2021) Partial purification and characterisation of pectinase produced by Aspergillus niger LFP-1 grown on pomelo peels as a substrate. Trop Life Sci Res 32:1–22. https://doi.org/10.21315/tlsr2021.32.1.1
Mehmood T, Saman T, Irfan M et al (2019) Pectinase production from Schizophyllum commune through central composite design using citrus waste and its immobilization for industrial exploitation. Waste Biomass Valoriz 10:2527–2536. https://doi.org/10.1007/s12649-018-0279-9
Meyer V (2019) Merging science and art through fungi. Fungal Biol Biotechnol 6:5. https://doi.org/10.1186/s40694-019-0068-7
Meyer V, Basenko EY, Benz JP et al (2020) Growing a circular economy with fungal biotechnology: a white paper. Fungal Biol Biotechnol 7:5. https://doi.org/10.1186/s40694-020-00095-z
Mitchell DA, Krieger N (2019) Solid-state cultivation bioreactors, pp 105–133
Mitchell DA, Pitol LO, Biz A, et al (2019) Design and operation of a pilot-scale packed-bed bioreactor for the production of enzymes by solid-state fermentation, pp 27–50
Mohammadi M, Khakbaz Heshmati M, Sarabandi K et al (2019) Activated alginate-montmorillonite beads as an efficient carrier for pectinase immobilization. Int J Biol Macromol 137:253–260. https://doi.org/10.1016/j.ijbiomac.2019.06.236
Mohammadi M, Rezaei Mokarram R, Shahvalizadeh R et al (2020) Immobilization and stabilization of pectinase on an activated montmorillonite support and its application in pineapple juice clarification. Food Biosci 36:100625. https://doi.org/10.1016/j.fbio.2020.100625
Mohd AS, Veena P, Mohammad A (2013) Polygalacturonase production from Rhizomucor pusillus isolated from fruit markets of Uttar Pradesh. Afr J Microbiol Res 7:252–259
Mohen D (2008) Pectin structure and biosynthesis. Curr Opin Plant Biol 11:266–277. https://doi.org/10.1016/j.pbi.2008.03.006
Mondal S, Soren JP, Mondal J et al (2020) Contemporaneous synthesis of multiple carbohydrate debranching enzymes from newly isolated Aspergillus fumigatus SKF-2 under solid state fermentation: a unique enzyme mixture for proficient saccharification of plant bioresources. Ind Crops Prod 150:112409. https://doi.org/10.1016/j.indcrop.2020.112409
Mondal S, Santra S, Rakshit S et al (2022) Saccharification of lignocellulosic biomass using an enzymatic cocktail of fungal origin and successive production of butanol by Clostridium acetobutylicum. Bioresour Technol 343:126093. https://doi.org/10.1016/j.biortech.2021.126093
Monjed MK, Robson GD, Pittman JK (2020) Isolation of fungal strains for biodegradation and saccharification of microalgal biomass. Biomass Bioenergy 137:105547. https://doi.org/10.1016/j.biombioe.2020.105547
Monjed MK, Achour B, Robson GD, Pittman JK (2021) Improved saccharification of Chlorella vulgaris biomass by fungal secreted enzymes for bioethanol production. Algal Res 58:102402. https://doi.org/10.1016/j.algal.2021.102402
Mukunda S, Onkarappa R, Prashith K (2013) Isolation and Screening of industrially important fungi from the soils of Western Ghats of Agumbe and Koppa, Karnataka, India. Sci Technol Arts Res J 1:27. https://doi.org/10.4314/star.v1i4.98816
Munir MD, Abdullah R, Haq I et al (2020) Purification, characterization, kinetics and thermodynamic analysis of polygalacturonase from Aspergillus tamarii for industrial applications. Rev Mex Ing Quim 19:293–304. https://doi.org/10.24275/rmiq/Bio1753
Nadar SS, Rathod VK (2019) A co-immobilization of pectinase and cellulase onto magnetic nanoparticles for antioxidant extraction from waste fruit peels. Biocatal Agric Biotechnol 17:470–479. https://doi.org/10.1016/j.bcab.2018.12.015
Naga Padma P, Anuradha K, Reddy G (2011) Pectinolytic yeast isolates for cold-active polygalacturonase production. Innov Food Sci Emerg Technol 12:178–181. https://doi.org/10.1016/j.ifset.2011.02.001
Nagpal R, Bhardwaj NK, Mahajan R (2020) Synergistic approach using ultrafiltered xylano-pectinolytic enzymes for reducing bleaching chemical dose in manufacturing rice straw paper. Environ Sci Pollut Res 27:44637–44646. https://doi.org/10.1007/s11356-020-11104-4
Nagpal R, Bhardwaj NK, Mahajan R (2021) Potential of crude xylano-pectinolytic enzymes in bleaching of rice straw pulp for improving paper quality and reducing toxic effluent load generation. Environ Sci Pollut Res 28:18284–18293. https://doi.org/10.1007/s11356-021-13204-1
Naumova ES, Shalamitskiy MYu, Naumov GI (2019) Molecular polymorphism of pectinase genes PGU of Saccharomyces bayanus var. uvarum Yeast. Appl Biochem Microbiol 55:882–887. https://doi.org/10.1134/S0003683819090059
Nawaz A, Hussain M, Munir M et al (2019) Strain improvement and assessment of cultural conditions for improved biosynthesis of pectinase using Penicillium notatum. Int J Biol Biotech 16:1–8
Nighojkar A, Patidar MK, Nighojkar S (2019) Pectinases: production and applications for fruit juice beverages. In: Processing and sustainability of beverages. Elsevier, pp 235–273
Nouri M, Khodaiyan F (2020) Magnetic biocatalysts of pectinase: synthesis by macromolecular cross-linker for application in apple juice clarification. Food Technol Biotechnol 58:391–401. https://doi.org/10.17113/ftb.58.04.20.6737
Núñez Pérez J, Chávez Arias BS, de la Vega Quintero JC et al (2022) Multi-objective statistical optimization of pectinolytic enzymes production by an Aspergillus sp. on dehydrated coffee residues in solid-state fermentation. Fermentation 8:170. https://doi.org/10.3390/fermentation8040170
Okonji RE, Ovumedia JO, Adedeji OS (2019) Purification and biochemical characterization of pectinase produced by Aspergillus fumigatus isolated from soil of decomposing plant materials. J Appl Biol Biotechnol 7:1–8. https://doi.org/10.7324/JABB.2019.70301
Ortiz GE, Ponce-Mora MC, Noseda DG et al (2017) Pectinase production by Aspergillus giganteus in solid-state fermentation: optimization, scale-up, biochemical characterization and its application in olive-oil extraction. J Ind Microbiol Biotechnol 44:197–211. https://doi.org/10.1007/s10295-016-1873-0
Padma PN, Anuradha K, Nagaraju B (2012) Use of pectin rich fruit wastes for polygalacturonase production by Aspergillus awamori MTCC 9166 in solid state fermentation. J Bioprocess Biotech. https://doi.org/10.4172/2155-9821.1000116
Pagnonceli J, Rasbold LM, Rocha GB et al (2019) Biotechnological potential of an exo-polygalacturonase of the new strain Penicillium janthinellum VI2R3M: biochemical characterization and clarification of fruit juices. J Appl Microbiol 127:1706–1715. https://doi.org/10.1111/jam.14426
Panda S, Sahoo K, Das R, Dhal KN (2012) Pectinolytic and cellulolytic activity of soil fungal isolates from similipal bioreserve forest. World Environ 2:1–3. https://doi.org/10.5923/j.env.20120202.01
Patel AK, Singhania RR, Pandey A (2017) Production, purification, and application of microbial enzymes. In: Biotechnology of microbial enzymes. Elsevier, pp 13–41
Patel VB, Chatterjee S, Dhoble AS (2022) A review on pectinase properties, application in juice clarification, and membranes as immobilization support. J Food Sci 87:3338–3354. https://doi.org/10.1111/1750-3841.16233
Patidar MK, Nighojkar A, Nighojkar S, Kumar A (2017) Purification and Characterization of Polygalacturonase Produced by Aspergillus niger AN07 in Solid State Fermentation. Canadian Journal of Biotechnology 1:11–18. https://doi.org/10.24870/cjb.2017-000102
Patidar MK, Nighojkar S, Kumar A, Nighojkar A (2018) Pectinolytic enzymes-solid state fermentation, assay methods and applications in fruit juice industries: a review. 3 Biotech 8:199. https://doi.org/10.1007/s13205-018-1220-4
Patidar MK, Nighojkar S, Kumar A, Nighojkar A (2020) Production of polygalacturonase using Carica papaya peel biowaste and its application for pomegranate juice clarification. Environ Sustain 3:509–520. https://doi.org/10.1007/s42398-020-00138-6
Patil NP, Chaudhari BL (2010) Production and purification of pectinase by soil isolate Penicillium sp. and search for better agro-residue for its SSF. Recent Res Sci Technol 7:36–42
Patil NP, Patil KP, Chaudhari BL, Chincholkar SB (2012) Production, purification of exo-polygalacturonase from soil isolate Paecilomyces variotii NFCCI 1769 and its application. Indian J Microbiol 52:240–246. https://doi.org/10.1007/s12088-011-0162-x
Pedrolli DB, Monteiro AC, Gomes E, Carmona EC (2009) Pectin and pectinases: production, characterization and industrial application of microbial pectinolytic enzymes. Open Biotechnol J 3:9–18. https://doi.org/10.2174/1874070700903010009
Pilgaard B, Wilkens C, Herbst F-A et al (2019) Proteomic enzyme analysis of the marine fungus Paradendryphiella salina reveals alginate lyase as a minimal adaptation strategy for brown algae degradation. Sci Rep 9:12338. https://doi.org/10.1038/s41598-019-48823-9
Pitol LO, Biz A, Mallmann E et al (2016) Production of pectinases by solid-state fermentation in a pilot-scale packed-bed bioreactor. Chem Eng J 283:1009–1018. https://doi.org/10.1016/j.cej.2015.08.046
Poletto P, Polidoro TA, Zeni M, da Silveira MM (2017) Evaluation of the operating conditions for the solid-state production of pectinases by Aspergillus niger in a bench-scale, intermittently agitated rotating drum bioreactor. LWT Food Sci Technol 79:92–101. https://doi.org/10.1016/j.lwt.2017.01.018
Poondla V, Bandikari R, Subramanyam R, Reddy Obulam VS (2015) Low temperature active pectinases production by Saccharomyces cerevisiae isolate and their characterization. Biocatal Agric Biotechnol 4:70–76. https://doi.org/10.1016/j.bcab.2014.09.008
Poturcu K, Ozmen I, Biyik HH (2017) Characterization of an alkaline thermostable pectin lyase from newly isolated Aspergillus niger _WHAK1 and its application on fruit juice clarification. Arab J Sci Eng 42:19–29. https://doi.org/10.1007/s13369-016-2041-6
Prado Barragán LA, Figueroa JJB, Rodríguez Durán LV, et al (2016) Fermentative production methods. In: Biotransformation of agricultural waste and by-products. Elsevier, Amsterdam, pp 189–217
Prasad S, Malav LC, Choudhary J et al (2021) Soil microbiomes for healthy nutrient recycling, pp 1–21
Presley GN, Zhang J, Purvine SO, Schilling JS (2020) Functional genomics, transcriptomics, and proteomics reveal distinct combat strategies between lineages of wood-degrading fungi with redundant wood decay mechanisms. Front Microbiol 11:1646. https://doi.org/10.3389/fmicb.2020.01646
Priya G, Lau N-S, Furusawa G et al (2018) Metagenomic insights into the phylogenetic and functional profiles of soil microbiome from a managed mangrove in Malaysia. Agric Gene 9:5–15. https://doi.org/10.1016/j.aggene.2018.07.001
Raina D, Kumar V, Saran S (2022) A critical review on exploitation of agro-industrial biomass as substrates for the therapeutic microbial enzymes production and implemented protein purification techniques. Chemosphere 294:133712. https://doi.org/10.1016/j.chemosphere.2022.133712
Raveendran S, Parameswaran B, Ummalyma SB et al (2018) Applications of microbial enzymes in food industry. Food Technol Biotechnol 56:16. https://doi.org/10.17113/ftb.56.01.18.5491
Ketipally R, Ram MR (2018) Optimization of pectinase production by Aspergillus oryzae RR 103. Current agriculture research journal, 6(1):37. https://doi.org/10.12944/CARJ.6.1.05
Reddy PL, Sreeramulu A (2012) Isolation, identification and screening of pectinolytic fungi from different soil samples of Chittoor district. Int J Life Sci Biotechnol Pharma Res 1:1–10
Reginatto C, dos SantosPosso G, Costa Ramos K et al (2022) Inoculation conditions improved the pectinase productivity in Aspergillus niger LB-02-SF solid-state cultivation. Biocatal Agric Biotechnol 42:102354. https://doi.org/10.1016/j.bcab.2022.102354
Rollero S, Zietsman AJJ, Buffetto F et al (2018) Kluyveromyces marxianus secretes a pectinase in shiraz grape must that impacts technological properties and aroma profile of wine. J Agric Food Chem 66:11739–11747. https://doi.org/10.1021/acs.jafc.8b03977
Ruiz HA, Rodríguez-Jasso RM, Hernandez-Almanza A et al (2017) Pectinolytic enzymes. In: Current Developments in Biotechnology and Bioengineering. Elsevier, Amsterdam, pp 47–71
Sadh PK, Duhan S, Duhan JS (2018) Agro-industrial wastes and their utilization using solid state fermentation: a review. Bioresour Bioprocess 5:1. https://doi.org/10.1186/s40643-017-0187-z
Safran J, Habrylo O, Cherkaoui M et al (2021) New insights into the specificity and processivity of two novel pectinases from Verticillium dahliae. Int J Biol Macromol 176:165–176. https://doi.org/10.1016/j.ijbiomac.2021.02.035
Samanta S (2019) Microbial pectinases: a review on molecular and biotechnological perspectives. J Microbiol Biotechnol Food Sci 9:248–266. https://doi.org/10.15414/jmbfs.2019.9.2.248-266
Sarkar D (2014) A study on optimization of Penicillium chrysogenum culture media in solid state fermentation process for pectinase enzyme production. Int J Pharm Life Sci 5:3966–3971
Satapathy S, Soren JP, Mondal KC et al (2021) Industrially relevant pectinase production from Aspergillus parvisclerotigenus KX928754 using apple pomace as the promising substrate. J Taibah Univ Sci 15:347–356. https://doi.org/10.1080/16583655.2021.1978833
Sathya TA, Jacob AM, Khan M (2014) Cloning and molecular modelling of pectin degrading glycosyl hydrolase of family 28 from soil metagenomic library. Mol Biol Rep 41:2645–2656. https://doi.org/10.1007/s11033-014-3123-8
Selvasekaran P, Chidambaram R (2020) Agriculturally important fungi for crop protection, pp 1–53
Sethi BK, Nanda PK, Sahoo S (2016) Enhanced production of pectinase by Aspergillus terreus NCFT 4269.10 using banana peels as substrate. 3 Biotech 6:36. https://doi.org/10.1007/s13205-015-0353-y
Shanmugam MK, Mandari V, Devarai SK, Gummadi SN (2022) Types of bioreactors and important design considerations. In: Current developments in biotechnology and bioengineering. Elsevier, pp 3–30
Shanmugavel M, Vasantharaj S, Yazhmozhi A et al (2018) A study on pectinases from Aspergillus tamarii: toward greener approach for cotton bio scouring and phytopigments processing. Biocatal Agric Biotechnol 15:295–303. https://doi.org/10.1016/j.bcab.2018.06.013
Sharma S, Mandhan RP, Sharma J (2011) Pseudozyma sp. SPJ: an economic and eco-friendly approach for degumming of flax fibres. World J Microbiol Biotechnol 27:2697–2701. https://doi.org/10.1007/s11274-011-0743-1
Sharma N, Rathore M, Sharma M (2013) Microbial pectinase: sources, characterization and applications. Rev Environ Sci Biotechnol 12:45–60. https://doi.org/10.1007/s11157-012-9276-9
Sharma HP, Patel H, Sugandha (2017) Enzymatic added extraction and clarification of fruit juices–A review. Crit Rev Food Sci Nutr 57:1215–1227. https://doi.org/10.1080/10408398.2014.977434
Shin Y, Chane A, Jung M, Lee Y (2021) Recent advances in understanding the roles of pectin as an active participant in plant signaling networks. Plants 10:1712. https://doi.org/10.3390/plants10081712
Siddiqa A, Noreen S, Khalid AM et al (2018) Statistical optimization of pectin lyase from Penicillium digitatum in solid state fermentation. Int J Appl Biol Forensics 2:157–170
Sinclair R, Rosquete MR, Drakakaki G (2018) Post-golgi trafficking and transport of cell wall components. Front Plant Sci 9:1784. https://doi.org/10.3389/fpls.2018.01784
Singh P, Hamid B, Lone MA, et al (2012a) Evaluation of pectinase activity from the psychrophilic fungal strain Truncatella angustata-BPF5 for use in wine industry. J Endocytobiosis Cell Res 57–61
Singh R, Dhawan S, Singh K, Kaur J (2012b) Cloning, expression and characterization of a metagenome derived thermoactive/thermostable pectinase. Mol Biol Rep 39:8353–8361. https://doi.org/10.1007/s11033-012-1685-x
Singh J, Kundu D, Das M, Banerjee R (2019) Enzymatic processing of juice from fruits/vegetables: an emerging trend and cutting-edge research in food biotechnology. In: Enzymes in food biotechnology. Elsevier, pp 419–432
Singhania RR, Patel AK, Thomas L, et al (2015) Industrial enzymes. In: Industrial biorefineries & white biotechnology. Elsevier, pp 473–497
Smith C (2005) Striving for purity: advances in protein purification. Nat Methods 2:71–77. https://doi.org/10.1038/nmeth0105-71
Soccol CR, da Costa ESF, Letti LAJ et al (2017) Recent developments and innovations in solid state fermentation. Biotechnol Res Innov 1:52–71. https://doi.org/10.1016/j.biori.2017.01.002
Soozanipour A, Taheri-Kafrani A, Barkhori M, Nasrollahzadeh M (2019) Preparation of a stable and robust nanobiocatalyst by efficiently immobilizing of pectinase onto cyanuric chloride-functionalized chitosan grafted magnetic nanoparticles. J Colloid Interface Sci 536:261–270. https://doi.org/10.1016/j.jcis.2018.10.053
Souza RC, Cantão ME, Nogueira MA et al (2018) Outstanding impact of soil tillage on the abundance of soil hydrolases revealed by a metagenomic approach. Braz J Microbiol 49:723–730. https://doi.org/10.1016/j.bjm.2018.03.001
Sprockett DD, Piontkivska H, Blackwood CB (2011) Evolutionary analysis of glycosyl hydrolase family 28 (GH28) suggests lineage-specific expansions in necrotrophic fungal pathogens. Gene 479:29–36. https://doi.org/10.1016/j.gene.2011.02.009
Sudhakar N, Shanmugam H, Kumaran S et al (2018) Omics approaches in fungal biotechnology. In: Omics technologies and bio-engineering. Elsevier, pp 53–70
Szabo OE, Csiszar E, Toth K et al (2015) Ultrasound-assisted extraction and characterization of hydrolytic and oxidative enzymes produced by solid state fermentation. Ultrason Sonochem 22:249–256. https://doi.org/10.1016/j.ultsonch.2014.07.001
Takeyama MM, de Carvalho MC, Carvalho HS et al (2022) Pectinases secretion by Saccharomyces cerevisiae: optimization in solid-state fermentation and identification by a shotgun proteomics approach. Molecules 27:4981. https://doi.org/10.3390/molecules27154981
Tan H, Yang G, Chen W et al (2020) Identification and characterization of thermostable endo-polygalacturonase II B from Aspergillus luchuensis. J Food Biochem 44:e13133. https://doi.org/10.1111/jfbc.13133
Tanaka Y, Suzuki T, Nakamura L et al (2019) A GH family 28 endo-polygalacturonase from the brown-rot fungus Fomitopsis palustris: purification, gene cloning, enzymatic characterization and effects of oxalate. Int J Biol Macromol 123:108–116. https://doi.org/10.1016/j.ijbiomac.2018.11.004
Tanveer A, Yadav S, Yadav D (2016) Comparative assessment of methods for metagenomic DNA isolation from soils of different crop growing fields. 3 Biotech 6:220. https://doi.org/10.1007/s13205-016-0543-2
Tatta ER, Imchen M, Moopantakath J, Kumavath R (2022) Bioprospecting of microbial enzymes: current trends in industry and healthcare. Appl Microbiol Biotechnol 106:1813–1835. https://doi.org/10.1007/s00253-022-11859-5
Teixeira WFA, Batista RD, do Amaral Santos CCA et al (2021) Minimal enzymes cocktail development by filamentous fungi consortia in solid-state cultivation and valorization of pineapple crown waste by enzymatic saccharification. Waste Biomass Valoriz 12:2521–2539. https://doi.org/10.1007/s12649-020-01199-8
Thakur P, Singh AK, Singh M, Mukherjee G (2021) Extracellular alkaline pectinases production: a review. J Microbiol Biotechnol Food Sci 11:e3745. https://doi.org/10.55251/jmbfs.3745
Trindade LV, Desagiacomo C, Polizeli MD (2016) Biochemical characterization, thermal stability, and partial sequence of a novel exo-polygalacturonase from the thermophilic fungus Rhizomucor pusillus A13.36 obtained by submerged cultivation. Biomed Res Int 2016:1–10. https://doi.org/10.1155/2016/8653583
Trindade Ximenes IA, de Oliveira PCO, Wegermann CA, de Moraes MC (2021) Magnetic particles for enzyme immobilization: a versatile support for ligand screening. J Pharm Biomed Anal 204:114286. https://doi.org/10.1016/j.jpba.2021.114286
Ullah H (2012) The role of ion exchange chromatography in purification and characterization of molecules. In: Ion Exchange Technologies. InTech
Usha DK, Kanimozhi G, Panneerselvam A (2014) Isolation and screening of pectin lyase producing fungi from soil sample of dead organic matters. World J Pharm Res 3:563–569
Villarreal F, Stocchi N, ten Have A (2022) Functional classification and characterization of the fungal glycoside hydrolase 28 protein family. Journal of Fungi 8:217. https://doi.org/10.3390/jof8030217
Voragen AGJ, Coenen G-J, Verhoef RP, Schols HA (2009) Pectin, a versatile polysaccharide presents in plant cell walls. Struct Chem 20:263–275. https://doi.org/10.1007/s11224-009-9442-z
Wagschal K, Rose Stoller J, Chan VJ et al (2016) Expression and characterization of hyperthermostable exo-polygalacturonase TtGH28 from Thermotoga thermophilus. Mol Biotechnol 58:509–519. https://doi.org/10.1007/s12033-016-9948-8
Wang H, Li X, Ma Y, Song J (2014) Characterization and high-level expression of a metagenome-derived alkaline pectate lyase in recombinant Escherichia coli. Process Biochem 49:69–76. https://doi.org/10.1016/j.procbio.2013.10.001
Wang C, Dong D, Wang H et al (2016) Metagenomic analysis of microbial consortia enriched from compost: new insights into the role of Actinobacteria in lignocellulose decomposition. Biotechnol Biofuels 9:22. https://doi.org/10.1186/s13068-016-0440-2
Wang J, Zhang Y, Qin X et al (2017) Efficient expression of an acidic endo-polygalacturonase from Aspergillus niger and Its application in juice production. J Agric Food Chem 65:2730–2736. https://doi.org/10.1021/acs.jafc.6b05109
Wang J, Chio C, Chen X et al (2019) Efficient saccharification of agave biomass using Aspergillus niger produced low-cost enzyme cocktail with hyperactive pectinase activity. Bioresour Technol 272:26–33. https://doi.org/10.1016/j.biortech.2018.09.069
Wang Y, Xue P, Cao M et al (2021) Directed evolution: methodologies and applications. Chem Rev 121:12384–12444. https://doi.org/10.1021/acs.chemrev.1c00260
Wardman JF, Bains RK, Rahfeld P, Withers SG (2022) Carbohydrate-active enzymes (CAZymes) in the gut microbiome. Nat Rev Microbiol 20:542–556. https://doi.org/10.1038/s41579-022-00712-1
Wong LY, Saad WZ, Mohamad R, Tahir PMd (2017) Optimization of cultural conditions for polygalacturonase production by a newly isolated Aspergillus fumigatus R6 capable of retting kenaf. Ind Crops Prod 97:175–183. https://doi.org/10.1016/j.indcrop.2016.12.019
Wösten HAB (2019) Filamentous fungi for the production of enzymes, chemicals and materials. Curr Opin Biotechnol 59:65–70. https://doi.org/10.1016/j.copbio.2019.02.010
Wulandari AP, Purba JR, Irawan B et al (2021) Effect of harvesting age of plant and pectinolytic selected-fungi in bio degumming ramie performance. Heliyon 7:e08392. https://doi.org/10.1016/j.heliyon.2021.e08392
Wusigale LL, Luo Y (2020) Casein and pectin: structures, interactions, and applications. Trends Food Sci Technol 97:391–403. https://doi.org/10.1016/j.tifs.2020.01.027
Xu H, Zhang P, Zhang Y et al (2020) Overexpression and biochemical characterization of an Endo-α-1,4-polygalacturonase from Aspergillus nidulans in Pichia pastoris. Int J Mol Sci 21:2100. https://doi.org/10.3390/ijms21062100
Yadav S, Shastri NV (2007) Purification and properties of an extracellular pectin lyase produced by the strain of Penicillium oxalicum in solid-state fermentation. Indian J Biochem Biophys 44:247–251
Yadav S, Yadav PK, Yadav D, Yadav KDS (2008) Purification and characterisation of an acidic pectin lyase produced by Aspergillus ficuum strain MTCC 7591 suitable for clarification of fruit juices. Ann Microbiol 58:61–65. https://doi.org/10.1007/BF03179446
Yadav S, Yadav PK, Yadav D, Yadav KDS (2009a) Purification and characterization of pectin lyase secreted by Penicillium citrinum. Biochem Mosc 74:800–806. https://doi.org/10.1134/S0006297909070141
Yadav S, Yadav PK, Yadav D, Yadav KDS (2009b) Pectin lyase: a review. Process Biochem 44:1–10. https://doi.org/10.1016/j.procbio.2008.09.012
Yadav S, Yadav PK, Yadav D, Yadav KDS (2009c) Purification and characterization of pectin lyase produced by Aspergillus terricola and its application in retting of natural fibers. Appl Biochem Biotechnol 159:270–283. https://doi.org/10.1007/s12010-008-8471-1
Yadav S, Dubey AK, Anand G, Yadav D (2013) Purification and characterization of pectin lyase secreted by Aspergillus flavus MTCC 10938. Appl Biochem Microbiol 49:396–401. https://doi.org/10.7868/S0555109913040156
Yadav S, Dubey AK, Anand G et al (2014) Purification and biochemical characterization of an alkaline pectin lyase from Fusarium decemcellulare MTCC 2079 suitable for Crotolaria juncea fibre retting. J Basic Microbiol 54:S161–S169. https://doi.org/10.1002/jobm.201300281
Yadav S, Maurya SK, Anand G et al (2017a) Purification, characterization and retting of Crotolaria juncea fibres by an alkaline pectin lyase from Fusarium oxysporum MTCC 1755. 3 Biotech 7:136. https://doi.org/10.1007/s13205-017-0750-5
Yadav S, Maurya SK, Anand G et al (2017b) Purification and characterization of a highly alkaline pectin lyase from Fusarium lateritum MTCC 8794. Biologia 72:245–251. https://doi.org/10.1515/biolog-2017-0038
Yamada H, Kubo S, Kunishige Y et al (2021) Homogalacturonan and Xylogalacturonan region specificity of self-cloning vector-expressed pectin methylesterases (AoPME1–3) in Aspergillus oryzae. Enzyme Microb Technol 150:109894. https://doi.org/10.1016/j.enzmictec.2021.109894
Yang Y, Anderson CT (2020) Biosynthesis, localisation, and function of pectins in plants. In: Pectin: technological and physiological properties. Springer, Cham, pp 1–15
Yang Y, Sha M (2019) A beginner’s guide to bioprocess modes–batch, fed-batch, and continuous fermentation. Eppendorf Inc, Enfield
Yang Y, Yu Y, Liang Y et al (2018) A profusion of molecular scissors for pectins: classification, expression, and functions of plant polygalacturonases. Front Plant Sci 9:1208. https://doi.org/10.3389/fpls.2018.01208
Yang G, Chen W, Tan H et al (2020) Biochemical characterization and evolutionary analysis of a novel pectate lyase from Aspergillus parasiticus. Int J Biol Macromol 152:180–188. https://doi.org/10.1016/j.ijbiomac.2020.02.279
Yapo BM (2011) Pectic substances: from simple pectic polysaccharides to complex pectins—a new hypothetical model. Carbohydr Polym 86:373–385. https://doi.org/10.1016/j.carbpol.2011.05.065
Yoshino-Yasuda S, Kato M, Kitamoto N (2011) Sequence analysis and heterologous expression of polygalacturonase gene (AspecA) from a Shoyu Koji Mold, Aspergillus sojae KBN1340. Food Sci Technol Res 17:579–584. https://doi.org/10.3136/fstr.17.579
Yuan Y, Zhang X-Y, Zhao Y et al (2019) A novel PL9 pectate lyase from Paenibacillus polymyxa KF-1: cloning, expression, and its application in pectin degradation. Int J Mol Sci 20:3060. https://doi.org/10.3390/ijms20123060
Zdunek A, Pieczywek PM, Cybulska J (2021) The primary, secondary, and structures of higher levels of pectin polysaccharides. Compr Rev Food Sci Food Saf 20:1101–1117. https://doi.org/10.1111/1541-4337.12689
Zheng Y, Wang Y, Pan J et al (2017) Semi-continuous production of high-activity pectinases by immobilized Rhizopus oryzae using tobacco wastewater as substrate and their utilization in the hydrolysis of pectin-containing lignocellulosic biomass at high solid content. Bioresour Technol 241:1138–1144. https://doi.org/10.1016/j.biortech.2017.06.066
Zheng L, Xu Y, Li Q, Zhu B (2021) Pectinolytic lyases: a comprehensive review of sources, category, property, structure, and catalytic mechanism of pectate lyases and pectin lyases. Bioresour Bioprocess 8:79. https://doi.org/10.1186/s40643-021-00432-z
Zhou M, Guo P, Wang T et al (2017) Metagenomic mining pectinolytic microbes and enzymes from an apple pomace-adapted compost microbial community. Biotechnol Biofuels 10:198. https://doi.org/10.1186/s13068-017-0885-y
Acknowledgements
The financial support provided by the Department of Science and Technology, New Delhi in the form of the DST Women Scientist Scheme-B (Grant. No. DST/WOS-B/2018/1707) and DST-INSPIRE Fellowship (DST/INSPIRE Fellowship/2019/IF190080) to Dr Aiman Tanveer and Kanchan Yadav are sincerely acknowledged.
Funding
Funding was provided by DST-INSPIRE Fellowship (grant no. DST/INSPIRE Fellowship/2019/IF190080), DST, Women Scientist-B, New Delhi INDIA (Grant no. DST/WOS-B/2018/1707).
Author information
Authors and Affiliations
Contributions
SD developed the concept and collected the relevant information, analysed and framed the first draft of the manuscript. KY and SG made a substantial contribution to the concept and design of the article. Dr AT and Dr SY reviewed and revised it critically for important intellectual content, and Prof. DY approved the concept and the version to be published. The authors agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dwivedi, S., Yadav, K., Gupta, S. et al. Fungal pectinases: an insight into production, innovations and applications. World J Microbiol Biotechnol 39, 305 (2023). https://doi.org/10.1007/s11274-023-03741-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03741-x