Abstract
α-l-rhamnosidase [EC 3.2.1.40] belongs to glycoside hydrolase (GH) families (GH13, GH78, and GH106 families) in the carbohydrate-active enzymes (CAZy) database, which specifically hydrolyzes the non-reducing end of α-l-rhamnose. Αccording to the sites of catalytic hydrolysis, α-l-rhamnosidase can be divided into α-1, 2-rhamnosidase, α-1, 3-rhamnosidase, α-1, 4-rhamnosidase and α-1, 6-rhamnosidase. α-l-rhamnosidase is an important enzyme for various biotechnological applications, especially in food, beverage, and pharmaceutical industries. α-l-rhamnosidase has a wide range of sources and is commonly found in animals, plants, and microorganisms, and its microbial source includes a variety of bacteria, molds and yeasts (such as Lactobacillus sp., Aspergillus sp., Pichia angusta and Saccharomyces cerevisiae). In recent years, a series of advances have been achieved in various aspects of α-validates the above-described-rhamnosidase research. A number of α-l-rhamnosidases have been successfully recombinant expressed in prokaryotic systems as well as eukaryotic systems which involve Pichia pastoris, Saccharomyces cerevisiae and Aspergillus niger, and the catalytic properties of the recombinant enzymes have been improved by enzyme modification techniques. In this review, the sources and production methods, general and catalytic properties and biotechnological applications of α-l-rhamnosidase in different fields are summarized and discussed, concluding with the directions for further in-depth research on α-l-rhamnosidase.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
α-l-rhamnosidase (EC 3.2.1.40) is a glycoside hydrolase that can hydrolyze terminal l-rhamnose of various natural glycosides, such as naringin, hesperidin, and rutin (Table 1). The enzyme is ubiquitous in nature and is found in animals, plants and microorganisms. Based on amino acid sequence similarity, α-l-rhamnosidases of microbial origin can be classified into GH13, GH78 and GH106 glycoside hydrolase families in the CAZy database. The analysis of the protein structure and reaction mechanism of different α-l-rhamnosidase species revealed that they have some differences: GH78 family α-l-rhamnosidase contains five structural domains and reacts through the substrate binding to the catalytic domain of the barrel structure of (α/α)6; GH106 family α-l-rhamnosidase has a (β/α)8-barrel structure containing five structural domains; GH13 family α-l-rhamnosidase crystal structure has not yet been resolved, and its properties are highly similar to the sequence of amylase but cannot hydrolyze soluble starch (Cui et al. 2007; Terry et al. 2020; Ndeh et al. 2017; Liu et al. 2012a, b). The main production mode of α-l-rhamnosidase is microbial fermentation. However, the current production level of microbial α-l-rhamnosidase needs to be improved, and the α-l-rhamnosidase produced by wild-type strains has problems such as the need for inducers and the difficulty of isolation and purification in the later stage. Although companies have realized the industrial production of this enzyme, the high price of commercial enzymes severely restricts the wide application of α-l-rhamnosidase. Therefore, so many researchers have been involved in the construction and modification of high-yielding α-l-rhamnosidase engineered strains.
α-l-rhamnosidase is one of the important enzymes for various biotechnology applications, especially in the food and pharmaceutical industries. As an enzyme with a natural glycosidic bond hydrolysis function, α-l-Rhamnosidase acts to cleave the α-L-rhamnosyl portion of flavonoids to specifically hydrolyze the α-1,2, α-1,3, α-1,4, α-1, and α-1,6 glycosidic bonds at the ends of glycans or glycosides, releasing L-rhamnose and producing new glycans or glycosides (Slámová et al. 2018). For instance, it is used to remove bitter substances in citrus juice (Li et al. 2018b, 2019; Bodakowska-Boczniewicz and Garncarek 2019); improve the taste and aroma of fresh juice and tea (Fang et al. 2019; Peng et al. 2021a). Prunin, hesperetin 7-O-glucoside, and isoquercitrin with higher bioavailability and bioactivity are prepared using naringin, hesperidin and rutin as substrates (Carceller et al. 2019; Kumar et al. 2019; Wang et al. 2020). Although there have been review articles on α-l-rhamnosidase, many advances have been made in the research related to this enzyme in recent years. The purpose of this review is to summarize and discuss the progress in the sources, production, properties and applications of α-l-rhamnosidase so far, so as to further grasp the key research directions of α-l-rhamnosidase in the future.
Source and production
Animal and plant sources
Naringinase, which has both α-l-rhamnosidase and β-d-glucosidase activities, was first isolated from celery seeds by Hall (1938). Twenty years later, Ting (1958) and Thomas et al. (1958) also discovered naringinase in grape leaves. Since then some scholars have also studied naringinase in buckwheat seeds and the genus Rhamnus (Suzuki 1962; Bourbouze et al. 1975). Liver is the primary location of ɑ-d-rhamnosidase in animals. Qian et al. (2005) first isolated and purified the ɑ-l-rhamnosidase from pig liver, and they found that the enzyme could hydrolyze the terminal rhamnose of dioscin. Later, this dioscin-α-d-rhamnosidase was also identified and purified from bovine liver (Qian et al. 2013).
Microbial sources and production
Molds
The sources of mold include Penicillium, Aspergillus, and Rhizopus genera. By adding corn cob or naringin to the liquid medium, the researchers stimulated the formation of α-l-rhamnosidase. They then sequentially extracted and purified various α-l-rhamnosidase from Penicillium citrinum MTCC-3565, Penicillium citrinum MTCC-8897, and Penicillium corylopholum MTCC-2011 (Yadav et al. 2012a, b, 2013b). They also isolated and identified Penicillium griseoroseum MTCC-9224 from the decaying gooseberry fruit peel, which secretes α-l-rhamnosidase, and determined that gooseberry peel is the best inducer for its production of this enzyme (Yadav et al. 2017).
There are few reports on the production of α-l-rhamnosidase by Rhizopus species. Shanmugam and Yadav (1995) found that Rhizopus nigricans could produce α-l-rhamnosidase extracellularly. However, Rhizopus arrhizus CCF 100 could show obvious α-l-rhamnosidase activity only when it is induced by rutin or naringin (Monti et al. 2004).
The genus Aspergillus contains multiple known producers of α-l-rhamnosidase, including Aspergillus terreus, Aspergillus aculeatus, Aspergillus niger, Aspergillus nidulans, and Aspergillus oryzae. The production methods of α-l-rhamnosidase by Aspergillus mainly include liquid fermentation and solid fermentation. At present, the main international production mode is still liquid deep fermentation. Weignerova et al. (2012) fermented A. terreus CCF 3059 in a 75 L bioreactor to mass-produce alkali- and thermo-stable α-l-rhamnosidase. As with other Aspergillus spp., the α-l-rhamnocidase produced by A. niger tends to be more present as naringinase. Several investigations described the improvement of the fermentation medium and conditions for A. niger DB056 to produce naringinase, and after optimization, the maximum activity of α-l- rhamnosidase was 1069.30 U/mL in a 200 L fermenter (Wu et al. 2010).
Yeasts and other fungi
Pichia angusta X349 is regarded as one of the excellent producers of α-l-rhamnosidase (Yanai and Sato 2000). Furthermore, Cryptococcus albidus and Candida tropicalis can also produce α-l-rhamnosidase (Borzova et al. 2018). Clavispora lusitaniae has the capacity to produce α-l-rhamnosidase, according to Singh et al. (2015b). To improve the enzyme activity, the enzyme production conditions of this strain were further improved (Singh et al. 2015a).
Certain other fungi, in addition to molds and yeasts, can also produce α-l-rhamnosidase, e.g., the oat pathogenic fungus Stagonospora avenae, the soil fungus Acrostalagmus luteoalbus, and the plant pathogen Fusarium moniliforme (Bleddyn Hughes et al. 2004; Rojas et al. 2011; Kumar et al. 2019). A new α-l-rhamnosidase produced by Talaromyces stollii CLY-6 has been found to more easily hydrolyze the rhamnosidic linkage between the rhamnose and aglycone of epmedin C (Cheng et al. 2022).
Bacteria
Researchers have also focused a lot of emphasis on bacterial sources of α-l-rhamnosidase. The first bacteria found to produce α-l-rhamnosidase was Bacteroides JY-6, a strain of human intestinal bacteria (Jang and Kim 1996). Moreover, it has been demonstrated that the human gut bacteria Fusobacterium K-60 and Enterococcus avium EFEL009 secrete α-l-rhamnosidase (Park et al. 2005; Shin et al. 2016). Subsequent studies have revealed that some Bacillus bacteria, like Bacillus litoralis C44 and Bacillus amyloliquefaciens 11,568, are also able to produce α-l-rhamnosidase (Lyu et al. 2016; Zhu et al. 2017).
The abundance of marine and soil microorganisms has led to a continuous investigation of the productive properties of various bacteria in the ocean and soil by researchers. The bacterium Pseudoalteromonas sp. 005NJ, Brevundimonas sp. Ci19, and Novosphingobium sp. PP1Y screened from sub-Antarctic seawater, Beagle Channel, and contaminated seawater in Italy can produce α-l-rhamnosidase (Gastón Orrillo et al. 2007; Alvarenga et al. 2013; Izzo et al. 2014). As a fermentation strain, Rodrigues et al. (2020) used the Acidobacterium bacterium AB60, which was isolated from Cerrado soils. In the medium that contained xylan as a carbon source, they fermented it. In addition to the primary xylan-degrading enzymes, high enzyme activity α-l-rhamnosidase was present in the fermentation broth.
Molecular biology research of Microbial-derived α-l-rhamnosidase
Cloning of α-l-rhamnosidase genes
At the beginning of the 21st century, researchers have gradually started to study the cloning of α-l-rhamnosidase genes. Initially, PCR amplification and library construction were the most common methods to clone α-l-rhamnosidase genes from microorganisms. Recently, based on the continuous development and advancement of sequencing technology, metagenomic mining has also been applied to the study of α-l-rhamnosidase genes. The cloned α-l-rhamnosidase genes in representative microorganisms are listed in Table 2. Numerous research has revealed that the fungal and bacterial genome may contain multiple genes encoding α-l-rhamnosidase. For instance, the gene sequence of a novel rutin-converting flavonoid glycoside hydrolase was cloned from A. niger DLFCC-90 (accession number EU200666), and the enzyme was classified in the GH 13 family by sequence similarity comparison and hydrolysis characterization (Liu et al. 2012a, b).
Expression of α-l-rhamnosidase genes
So far, the expression of α-l-rhamnosidase of fungal and bacterial origin has been studied by many scholars. The expression of α-l-rhamnosidase from microorganisms in the last decade is demonstrated in Table 3. The prokaryotic (E. coli) expression system has been chosen for protein expression of the bacterial-derived α-l-rhamnosidase genes, and the pET system has been largely utilized. In E. coli BL21, the bacterial-derived α-l-rhamnosidase genes are expressed stably with active protein (Wu et al. 2018; Ferreira-Lazarte et al. 2021).
The expression of α-l-rhamnosidase gene of Aspergillus origin has been more studied. Lyu et al. (2019) expressed the codon-optimized α-l-rhamnosidase gene from n. E. coli and obtained 574.5 U/L of p-nitrophenol-α-rhammnoside (p-NPR) hydrolase activity after fermentation in a 5 L bioreactor. Considering some problems inherent in the prokaryotic expression system, such as unstable disulfide bonds, incorrect protein folding, and inclusion body precipitation, the eukaryotic expression system is preferable for fungal-derived genes to avoid this set of possible problems. Spohner et al. (2015) cloned the codon-optimized gene encoding Aspergillus terreus rhamnosidase, and the hydrolase activity of the enzyme expressed in Pichia pastoris and Kluyveromyces lactis against p-NPR was 17.6 U/mL and 30.6 U/mL, respectively. Moreover, it is not difficult to find that most of the eukaryotic expression systems of Aspergillus-derived α-l-rhamnosidase genes are heterologously expressed using Pichia pastoris, rarely involve homologous expression. Only Ye et al. (2022) homologously expressed the α-l-rhamnosidase gene of A. niger in A. niger 3.350 and the p-NPR hydrolase activity of the target protein obtained after 5 L bioreactor fermentation was 34.43 U/mg.
Properties of α-l-rhamnosidase
General properties and catalytic properties of α-l-rhamnosidase
α-l-rhamnosidase from different sources tends to exhibit different properties. For future research and application of this enzyme, it is advantageous to have adequate knowledge of the properties of α-l-rhamnosidase from different sources. The general properties and catalytic properties of α-l-rhamnosidase from different microbial sources are summarized in Tables 4 and 5, respectively.
General properties of α-l-rhamnosidase
Different plants and animals and microorganisms differ in many aspects of their genetic composition, transcription, translation and post-translational modifications due to their species differences. Various sources of α-l-Rhamnosidase have different levels of glycosylation and different conformations of the enzyme protein due to different post-translational modification mechanisms, resulting in differences in their structure and properties. The enzymes of different sources also have different catalytic properties depending on the properties they carry with their production hosts. Most α-l-rhamnosidases have molecular weights between 50 and 140 kDa, with the majority falling in the around 100 kDa. The optimum pH, optimum temperature and thermal stability of enzymes are crucial for their industrial applications, so it is necessary to focus on these three properties of different α-l-rhamnosidases.
Three bacterial α-l-rhamnosidases have been found to have an acidic pH, despite the fact that the optimum pH range for most bacterial α-l-rhamnosidases is 5.0–8.0. Ram2 from Pediococcus acidilactici and TpeRha form Thermotoga petrophila both have an optimal pH of 4.5 (Michlmayr et al. 2011; Xie et al. 2020). The optimum pH of Ram2 from Lactobacillus plantarum WCFS1 is 3.0, which is the lowest optimum pH reported for acidic α-l-rhamnosidase (Ferreira-Lazarte et al. 2021). The optimum pH of fungal α-l-rhamnosidases is generally in the range of 5.0-6.5. Interestingly, some fungi can produce alkaline α-l-rhamnosidase. Aspergillus clavato-nanicus MTCC-9611, Fusarium moniliforme MTCC-2088, and Aspergillus flavus MTCC-9606 have the high optimum pH of α-l-rhamnosidase, all between 10 and 11 (Yadav et al. 2011, 2012c; Kumar et al. 2019).
The optimum temperature for microbial α-l-rhamnosidase is typically in the range of 40–60 °C, but a few cold-tolerant and thermophilic enzymes do exist. The optimum temperature for α-l-rhamnosidase from the cold-tolerant bacterium, Brevundimonas sp. Ci19, is very low, at 20–37 °C (Alvarenga et al. 2013). Ram2 from Pediococcus acidilactici, RhaL1 from Alternaria sp. L1, and α-l-rhamnosidase from Aspergillus terreus CCF3059 all have an optimum temperature of 70 °C (Birgisson et al. 2004; Michlmayr et al. 2011; (Liu et al. 2012a, b; Weignerova et al. 2012). TpeRha from Thermotoga petrophila and DtRha from Dictyoglomus thermophilum both exhibit much higher optimum temperature than the above-mentioned thermophilic bacteria and fungi, at 90 and 95 °C, respectively (Guillotin et al. 2019; Xie et al. 2020). Moreover, TpeRha is the best heat-stable α-l-rhamnosidases to date, with the residual activity of more than 40% after 1 h incubation at 90 °C (Xie et al. 2020). According to some reports, the thermal stability of enzyme proteins is closely related to their degree of glycosylation modification, and to some extent, the glycosylation level of enzyme proteins correlates with their thermal stability (Manzanares et al. 2001). Besides, many other factors also affect the thermal stability of enzymes, such as salt bridges and hydrophobic interactions (Ge et al. 2018). Metal ions also have a significant effect on α-l-rhamnosidase activity. In the process of microbial metabolism or enzyme reaction in vitro, the participation of metal ions will affect the activity of enzyme. Mn2+ and Mg2+ can promote the conversion of naringin to prunin, while Cu2+ naringin can inhibit the conversion of prunin. The important role of Ca2+ in the catalytic process of α-l-rhamnosidase was revealed by comparing the crystal structures of the five available α-l-rhamnosidases. (Mensitieri et al. 2018) SaRha78A rhamnosidase, Ca2+ binds to an independent domain very close to the catalytic domain. When the substrate binds to the active site of the enzyme, Ca2+ can form coordination bonds with O3 and O4 of the substrate rhamnose-thus promoting the reaction (Fujimoto et al. 2013a, b). Enzyme activity inhibited by the metal chelating agent EDTA can also be fully restored by the addition of Ca2+. Several experiments have demonstrated the positive effect of Ca2+ on glycosidase activity (Miake et al. 2000; Hashimoto et al. 1998). On the contrary, Hg2+ and sulfur-based reagents can affect the sulfur-based groups of enzymes, thus inhibiting the activity of some α-l-rhamnosidases (Yanai et al. 2000). When 1 mmol/L Hg2+ is present, the activity of BtRha and RamA and other glycosidases is significantly reduced or even completely inhibited (Zverlov et al. 2000; Wu et al. 2018).
Catalytic properties of α-l-rhamnosidase
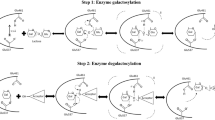
α-l-rhamnosidase can act on a total of two types of glycosidic bonds, one is the bond between glycosyl group and aglycone directly, that is, α-1 glycosidic bond, and the other is the bond between glycosyl group and glycosyl group, including α-1,2, α-1,3, α-1,4, and α-1,6 glycosidic bonds. P-NPR is a synthetic substrate, and all α-l-rhamnosidases can act on its α-1 glycosidic bond except very few α-l-rhamnosidases. Therefore, p-NPR is a universal substrate for the determination of the enzyme activity of α-l-rhamnosidase. However, for quercetin, a natural glycoside containing the α-1 glycosidic bond, only a small fraction of α-l-rhamnosidase can hydrolyze it. For example, the α-l-rhamnosidases from Streptomyces avermitilis, Novosphingobium sp PP1Y, and Alternaria alternata SK37.001, while like most α-l-rhamnosidases, they can hydrolyze α-1,2 and α-1,6 glycosidic bonds (Ichinose et al. 2013; Mensitieri et al. 2018; Zhang et al. 2018). The specific hydrolysis is described by the process of hydrolysis of glycosidic bonds by rhamnosidase SaRha78A as an example. The amino acid Glu636 acts as a proton donor to attack the O1 of the substrate rhamnose to protonate it, while the amino acid Glu895 deprotonates the water molecules that have strong hydrogen bonds with the substrate and the enzyme, and the deprotonated water molecules attack the glycosidic bonds of the substrate to complete the hydrolysis reaction (Fujimoto et al. 2013a, b; David et al. 2000; Zhu et al. 2021).
Interestingly, different α-l-rhamnosidases may have higher specificity for one of the glycosidic bonds. For instance, the α-l-rhamnosidases from Papiliotrema laurentii ZJU-L07, Dictyoglomus thermophilum, and A. nidulans are more specific for the α-1,2 glycosidic bond (Guillotin et al. 2019; Lyu et al. 2019; Lou et al. 2022); whereas the α-l-rhamnosidases from Chloroflexus aurantiacus and Bifidobacterium dentium showed more specific hydrolysis of α-1,6 glycosidic bond (Bang et al. 2015; Shin et al. 2019). Moreover, some α-l-rhamnosidases can hydrolyze only one type of glycosidic bond exclusively, such as A. tubingensis JMU-TS529 whose α-l-rhamnosidase can only hydrolyze α-1,2 glycosidic bond (Li et al. 2019). The number of α-l-rhamnosidases that can hydrolyze α-1,3 and α-1,4 glycosidic bonds is small compared to α-1,2 and α-1,6 glycosidic bonds. Nevertheless, the α-l-rhamnosidase from A. niger S528 has a broad substrate spectrum, and it is the only α-l-rhamnosidase known to hydrolyze the above four glycosidic bonds (Li et al. 2016).
Epimedin C also contains the α-1,2 glycosidic bond, but unlike the glycosidic bonds linking rhamnose and glucose in other compounds, the glycosidic bond in epimedin C links two rhamnose groups. An increasing number of α-l-rhamnosidases have been shown to hydrolyze this α-1,2 glycosidic bond in epimedin C, including DthRha from Dictyoglomus thermophilum DSM 3960, TpeRha from Thermotoga petrophila DSM 13,995, and Rhase-I from Talaromyces stollii CLY-6, and they further hydrolyze the α-1 glycosidic bond between rhamnose and aglycone in the product icariin (Xie et al. 2020; Zhang et al. 2021; Cheng et al. 2022).
Modification of α-l-rhamnosidase
Molecular modification of α-l-rhamnosidase
Thermal stability and catalytic efficiency are two important properties of enzymes, and studies aimed at improving these two properties have gradually increased in recent years. In particular, researchers have achieved many results in improving the thermal stability of r-Rha1 from A. niger JMU-TS528 through directed evolution and semi-rational design strategies (Li et al. 2018b; Liao et al. 2019). The most heat-stable r-Rha1 mutant available today, K406R/K573R, has a 3 h longer half-life at 60 °C than the wild type, which was obtained by replacing the two Lys on the surface of the enzyme with Arg (Li et al. 2018a). The affinity of r-Rha1 has been improved by the substitution of semi-conserved amino acids around the active site, and the affinity is the key to improve the catalytic efficiency (Li et al. 2020).
It should be emphasized that improving the catalytic efficiency needs to increase the flexibility of the structure, and improving the thermal stability needs to increase the rigidity of the structure, both of which have completely opposite structural requirements. Thus, it is difficult to improve the catalytic efficiency and thermal stability of α-l-rhamnosidase simultaneously by adjusting the protein structure. Li et al. (2021) proposed a dual screening strategy, which finally achieved the simultaneous enhancement of the thermal stability and catalytic efficiency of r-Rha1.
Immobilization of α-l-rhamnosidase
Immobilized enzymes can improve the thermal stability and affinity of the enzyme while maintaining the original properties of the enzyme, and at the same time make the enzyme easy to be separated and reusable. Metal-organic frameworks (MOFs), as an emerging porous material, have been successfully applied in the immobilization of α-l-rhamnosidase. For example, the α-l-rhamnosidase from A. niger CCTCC M 2,018,240 and A. niger JMU-TS528 were immobilized on magnetic MOFs and cerium-based metal-organic frameworks nanoparticles, respectively, and the substrate affinity of the enzymes were both significantly enhanced (Peng et al. 2021b; Wang et al. 2021). The co-immobilization of α-l-rhamnosidase Rha1 and β-glucosidase Glu4 from Talaromyces stollii CLY-6 was achieved based on a carrier-free cross-linked enzyme aggregate, and the co-immobilized enzyme exhibited more tolerant to sugars, thus becoming the enzyme that can obtain the highest icaritin yield at the highest epimedin C concentration ever reported (Liu et al. 2022).
Applications of α-l-rhamnosidase
Applications in the food industry
Citrus fruits have a bitter taste due to the presence of flavonoid glycosides such as hesperidin, neohesperidin, and naringin. Naringinase is often used for enzymatic debittering (Bodakowska-Boczniewicz and Garncarek 2019; Carceller et al. 2020). In fact, the presence of β-D-glucosidase is not required for debittering, but only the action of α-l-rhamnosidase can reduce the bitter substances. The fungus JMU-TS529 was isolated from rotten pomelo compost and identified as A. tubingensis, whose α-l-rhamnosidase has a strong hydrolytic effect only on naringin, and can remove the bitterness of pomelo juice while still retaining the aroma of pomelo (Li et al. 2019).
α-l-rhamnosidase alone or in combination with β-D-glucosidase can increase the floral flavor of orange juice, and the combined treatment has a more obvious effect on the taste and aroma quality of orange juice (Peng et al. 2021a). Ultrasonic action assisted α-l-rhamnosidase and β-d-glucosidase in the fermentation broth of A. niger to debitter Ouguan juice, and the rate of debittering increased while increasing the content of sweet and fruity aroma compounds in the juice, which improved the flavor of Ouguan juice (Gao et al. 2021). In addition to improving the aroma components of the juice, these two enzymes could also increase the content of aroma components in tea broth (Fang et al. 2019).
The hydrolysis products of α-l-rhamnosidase, flavonoid monoglycosides, have higher bioavailability and enhanced efficacy, so the beverage after the effect of α-l-rhamnosidase can be used as functional beverage, such as ginkgo tea drink (Fang et al. 2019). The addition of sorbitol promoted the hydrolysis of hesperidin by A. niger α-l-rhamnosidase, which helped to accelerate the production of hesperetin 7-O-glucoside, a sweetener precursor (Sun et al. 2022). The α-l-rhamnosidase immobilized on magnetic Fe3O4/MIL-101(Cr) nanoparticles could hydrolyze hesperidin dihydrochalcone to hesperidin dihydrochalcone glucoside, which is a sweetener (Wang et al. 2022).
Applications in the pharmaceutical industry
Rutin is converted to isoquercitrin by the action of α-l-rhamnosidase with the removal of one molecule of rhamnose. Isoquercitrin has anti-inflammatory, antioxidant, anti-allergic, and antihypertensive effects, and has important applications in the pharmaceutical industry. Yet, the industrial production of isoquercitrin has problems such as high cost, slow reaction, and low yield. Ionic liquids as co-solvents, high hydrostatic pressure treatment, addition of sorbitol, and addition of organic solvents have all been used to address these issues (Wang et al. 2013; Kim et al. 2016; Ge et al. 2017a; Shin et al. 2019), but the conversion rate of rutin or the yield of isoquercitrin still could not reach the standard of industrial application. Wang et al. (2020) first hydrolyzed rutin to isoquercitrin using α-l-rhamnosidase in deep eutectic solvents. 130 g/L rutin could be completely converted to isoquercitrin, with the yield of isoquercetin reaching 208.68 mm/h.
Herba Epimedii is a famous Chinese herbal medicine. The total flavonoids of Epimedii (TFE) are the main active ingredients in Epimedii, including icariin, epimedin A, epimedin B, epimedin C and icariside II (Jiang et al. 2016). Among them, icariin has the most significant pharmacological activities, such as anti-osteoporosis, antidepressant, and treatment of cardiovascular diseases (Zhang et al. 2017). However, the content of icariin is limited, while epimedin C, which structurally has only one more rhamnose than icariin, has a higher content, so epimedin C can be used as a substrate to convert it into icariin using α-l-rhamnosidase, which can hydrolyze the α-1,2 glycosidic bond between two rhamnoses. E.coli BL21 cells expressing A. nidulans synAnRhaE can completely convert 1 g/L of epimedin C to icariin within 90 min (Lyu et al. 2019). There are also many drug precursors or substances with multiple biological activities, such as tilianin and cyanidin-3-O-rutinoside, which can be prepared by α-l-rhamnosidase (Cui et al. 2016; Li et al. 2023).
L-rhamnose is used in the synthesis of rare bioactive rhamnosylated compounds in medicine and chemistry, and also functions as a chiral intermediate in plant protection agents (Yadav et al. 2010). All of the above mentioned substrates are subjected to α-l-rhamnosidase action to take off the terminal L-rhamnose, so α-l-rhamnosidase can be used to produce L-rhamnose (Wang et al. 2020). Moreover, some α-l-rhamnosidases can also synthesize glycosidic bonds through a reverse hydrolysis (Ge et al. 2017b). RhaL1 from Alternaria sp. L1 has been shown to perform rhamnosylation of anticancer drugs such as 2′-deoxy-5-fluorouridine, cytosine arabinoside, hydroxyurea, etc., and the rhamnosylated drugs are potentially valuable in enzyme-activated prodrug systems (Xu et al. 2019).
Conclusion
In recent years, many scholars have studied α-l-rhamnosidase from different levels. Firstly, the screening of α-l-rhamnosidase production strains and the optimization of fermentation conditions are becoming more and more mature, which provides a certain reference value for large-scale production of α-l-rhamnosidase preparations. Secondly, the cloning and expression of α-l-rhamnosidase from different microorganisms were studied at the molecular level. The constructed engineered strains with high enzyme activity can be used for producing α-l-rhamnosidase, which is of great significance for increasing the yield of this enzyme. Meanwhile, the properties of α-l-rhamnosidase from different microbial sources were studied, and the enzyme was modified by molecular modification or immobilization techniques, which is helpful to promote the practical application of α-l-rhamnosidase in industrial conditions. Finally, the catalytic ability of α-l-rhamnosidase to various natural substrates was studied, which provides ideas for the application of α-l-rhamnosidase in different production processes.
But so far, there are few studies on the induction mechanism and synthesis pathway of α-l-rhamnosidase from different sources. More in-depth studies on α-l-rhamnosidase should be carried out using structural biology and molecular biology techniques. Meanwhile, in order to better meet the demanding industrial production process, the enzyme modification work must continue to advance, and the search for more strategies that can simultaneously improve the catalytic efficiency and thermal stability of α-l-rhamnosidase should be taken as a research priority in the molecular modification of this enzyme. Furthermore, since the main applications of α-l-rhamnosidase include food and drug fields, α-l-rhamnosidase produced by A. niger, A. oryzae, and other filamentous fungi recognized as safe strains by the U.S. Food and Drug Administration, deserve the attention of researchers. In particular, the presence of β-D-glucosidase in the preparation of flavonoid monoglycosides catalyzed by α-l-rhamnosidase will cause the production of flavonoid aglycones as a by-product. It can be considered to use CRISPR/Cas9 gene editing technology to knockout the β-D-glucosidase gene, so that the complete transformation of flavonoid diglycosides to flavonoid monoglycosides can be achieved without the isolation and purification of α-l-rhamnosidase. The above research will continue to promote the deep understanding and further development of α-l-rhamnosidase in the future.
References
Alvarenga AE, Romero CM, Castro GR (2013) A novel α-L-rhamnosidase with potential applications in citrus juice industry and in winemaking. Eur Food Res Technol 237:977–985. https://doi.org/10.1007/s00217-013-2074-y
Avila M, Jaquet M, Moine D et al (2009) Physiological and biochemical characterization of the two α-L-rhamnosidases of Lactobacillus plantarum NCC245. Microbiology 155:2739–2749. https://doi.org/10.1099/mic.0.027789-0
Bang SH, Hyun YJ, Shim J et al (2015) Metabolism of rutin and poncirin by human intestinal microbiota and cloning of their metabolizing α-L-rhamnosidase from Bifidobacterium dentium. J Microbiol Biotechnol 25:18–25. https://doi.org/10.4014/jmb.1404.04060
Baudrexl M, Schwarz WH, Zverlov VV et al (2019) Biochemical characterisation of four rhamnosidases from thermophilic bacteria of the genera Thermotoga. Caldicellulosiruptor and Thermoclostridium Sci Rep 9:15924. https://doi.org/10.1038/s41598-019-52251-0
Beekwilder J, Marcozzi D, Vecchi S et al (2009) Characterization of rhamnosidases from Lactobacillus plantarum and Lactobacillus acidophilus. Appl Environ Microbiol 75:3447–3454. https://doi.org/10.1128/AEM.02675-08
Birgisson H, Hreggvidsson GO, Fridjónsson OH et al (2004) Two new thermostable α-L-rhamnosidases from a novel thermophilic bacterium. Enzyme Microb Technol 34:561–571. https://doi.org/10.1016/j.enzmictec.2003.12.012
Bleddyn Hughes H, Morrissey JP, Osbourn AE (2004) Characterisation of the saponin hydrolysing enzyme avenacoside-α-L-rhamnosidase from the fungal pathogen of cereals, Stagonospora avenae. Eur J Plant Pathol 110:421–427. https://doi.org/10.1023/b:ejpp.0000021083.94707
Bodakowska-Boczniewicz J, Garncarek Z (2019) Immobilization of naringinase from Penicillium decumbens on chitosan microspheres for debittering grapefruit juice. Molecules 24:4234. https://doi.org/10.3390/molecules24234234
Borzova N, Gudzenko O, Varbanets L (2018) Purification and characterization of a naringinase from Cryptococcus albidus. Appl Biochem Biotechnol 184:953–969. https://doi.org/10.1007/s12010-017-2593-2
Bourbouze R, Pratviel-Sosa F, Percheron F (1975) Rhamnodiastase et α-L-rhamnosidase de Fagopyrum esculentum. Phytochemistry 14:1279–1282. https://doi.org/10.1016/S0031-9422(00)98610-2
Carceller JM, Martínez Galán JP, Monti R et al (2019) Selective synthesis of citrus flavonoids prunin and naringenin using heterogeneized biocatalyst on graphene oxide. Green Chem 21:839–849. https://doi.org/10.1039/c8gc03661f
Carceller JM, Martínez Galán JP, Monti R et al (2020) Covalent immobilization of naringinase over two-dimensional 2D zeolites and its applications in a continuous process to produce citrus flavonoids and for debittering of juices. ChemCatChem 12:4502–4511. https://doi.org/10.1002/cctc.202000320
Cheng L, Zhang H, Cui H et al (2021) Efficient enzyme-catalyzed production of diosgenin: inspired by the biotransformation mechanisms of steroid saponins in Talaromyces stollii CLY-6. Green Chem 23:5896–5910. https://doi.org/10.1039/d0gc04152a
Cheng L, Zhang H, Cui H et al (2022) A novel α-L-rhamnosidase renders efficient and clean production of icaritin. J Clean Prod 341:130903. https://doi.org/10.1016/j.jclepro.2022.130903
Cui Z, Maruyama Y, Mikami B et al (2007) Crystal structure of glycoside hydrolase family 78 alpha-L-Rhamnosidase from Bacillus sp. GL1. J Mol Biol 374:384–398. https://doi.org/10.1016/j.jmb.2007.09.003
Cui P, Dou TY, Li SY et al (2016) Highly selective and efficient biotransformation of linarin to produce tilianin by naringinase. Biotechnol Lett 38:1367–1373. https://doi.org/10.1007/s10529-016-2116-1
David LZ, Stephen GW (2000) Glycosidase mechanisms: anatomy of a finely tuned catalyst. Acc Chem Res 33:11–18. https://doi.org/10.1021/ar970172
De Lise F, Mensitieri F, Tarallo V et al (2016) RHA-P: isolation, expression and characterization of a bacterial α-L-rhamnosidase from Novosphingobium sp. PP1Y. J Mol Catal B: Enzym 134:136–147. https://doi.org/10.1016/j.molcatb.2016.10.002
Fang X, Dong Y, Xie Y et al (2019) Effects of β-glucosidase and α-rhamnosidase on the contents of flavonoids, ginkgolides, and aroma components in ginkgo tea drink. Molecules 24:2009. https://doi.org/10.3390/molecules24102009
Ferreira-Lazarte A, Plaza-Vinuesa L, de Las Rivas B et al (2021) Production of α-rhamnosidases from Lactobacillus plantarum WCFS1 and their role in deglycosylation of dietary flavonoids naringin and rutin. Int J Biol Macromol 193:1093–1102. https://doi.org/10.1016/j.ijbiomac.2021.11.053
Fujimoto Z, Jackson A, Michikawa M et al (2013a) The structure of a Streptomyces avermitilis α-L-rhamnosidase reveals a novel carbohydrate- binding module CBM67 within the six-domain arrangement. J Biol Chem 288:12376–12385. https://doi.org/10.1074/jbc.M113.460097
Gao X, Feng T, Liu E et al (2021) Ougan juice debittering using ultrasound-aided enzymatic hydrolysis: impacts on aroma and taste. Food Chem 345:128767. https://doi.org/10.1016/j.foodchem.2020.128767
Gastón Orrillo A, Ledesma P, Delgado OD et al (2007) Cold-active α-L-rhamnosidase from psychrotolerant bacteria isolated from a sub-antarctic ecosystem. Enzyme Microb Technol 40:236–241. https://doi.org/10.1016/j.enzmictec.2006.04.002
Ge L, Chen A, Pei J et al (2017a) Enhancing the thermostability of α-L-rhamnosidase from aspergillus terreus and the enzymatic conversion of rutin to isoquercitrin by adding sorbitol. BMC Biotechnol 17:21. https://doi.org/10.1186/s12896-017-0342-9
Ge L, Xie J, Wu T et al (2017b) Purification and characterisation of a novel α-L-rhamnosidase exhibiting transglycosylating activity from aspergillus oryzae. Int J Food Sci Tech 52:2596–2603. https://doi.org/10.1111/ijfs.13546
Ge L, Li D, Wu T et al (2018) B-factor-saturation mutagenesis as a strategy to increase the thermostability of α-L-rhamnosidase from aspergillus terreus. J Biotechnol 275:17–23. https://doi.org/10.1016/j.jbiotec.2018.03.013
Gerstorferová D, Fliedrová B, Halada P et al (2012) Recombinant α-L-rhamnosidase from aspergillus terreus in selective trimming of rutin. Process Biochem 47:828–835. https://doi.org/10.1016/j.procbio.2012.02.014
Guillotin L, Kim H, Traore Y et al (2019) Biochemical characterization of the α-L-rhamnosidase DtRha from Dictyoglomus thermophilum: application to the selective derhamnosylation of natural flavonoids. ACS Omega 4:1916–1922. https://doi.org/10.1021/acsomega.8b03186
Hall DH (1938) A new enzyme of the glycosidase type. Nature 142:150–150. https://doi.org/10.1038/142150a0
Hashimoto W, Murata K (1998a) alpha-L-rhamnosidase of Sphingomonas sp. R1 producing an unusual exopolysaccharide of sphingan. Biosci Biotechnol Biochem 62(6):1068–1074. https://doi.org/10.1271/bbb.62.1068
Hashimoto W, Miyake O, Nankai H et al (2003) Molecular identification of an α-L-rhamnosidase from Bacillus sp. strain GL1 as an enzyme involved in complete metabolism of gellan. Arch Biochem Biophys 415:235–244. https://doi.org/10.1016/s0003-9861(03)00231-5
Ichinose H, Fujimoto Z, Kaneko S (2013) Characterization of an α-L-rhamnosidase from Streptomyces avermitilis. Biosci Biotechnol Biochem 77:213–216. https://doi.org/10.1271/bbb.120735
Ishikawa M, Shiono Y, Koseki T (2017) Biochemical characterization of aspergillus oryzae recombinant α-L-rhamnosidase expressed in Pichia pastoris. J Biosci Bioeng 124:630–634. https://doi.org/10.1016/j.jbiosc.2017.07.007
Izzo V, Tedesco P, Notomista E et al (2014) α-Rhamnosidase activity in the marine isolate Novosphingobium sp. PP1Y and its use in the bioconversion of flavonoids. J Mol Catal B: Enzym 105:95–103. https://doi.org/10.1016/j.molcatb.2014.04.002
Jang IS, Kim DH (1996) Purification and characterization of α-L-rhamnosidase from Bacteroides JY-6, a human intestinal bacterium. Biol Pharm Bull 19:1546–1549. https://doi.org/10.1248/bpb.19.1546
Jiang J, Zhao B-j, Song J et al (2016) Pharmacology and clinical application of plants in Epimedium L. Chin Herb Med 8:12–23. https://doi.org/10.1016/s1674-6384(16)60003-5
Kim DY, Yeom SJ, Park CS et al (2016) Effect of high hydrostatic pressure treatment on isoquercetin production from rutin by commercial α-L-rhamnosidase. Biotechnol Lett 38:1775–1780. https://doi.org/10.1007/s10529-016-2157-5
Kumar D, Yadav S, Yadava S et al (2019) An alkali tolerant α-L-rhamnosidase from Fusarium moniliforme MTCC-2088 used in de-rhamnosylation of natural glycosides. Bioorg Chem 84:24–31. https://doi.org/10.1016/j.bioorg.2018.11.027
Li L, Yu Y, Zhang X et al (2016) Expression and biochemical characterization of recombinant α-L-rhamnosidase r-Rha1 from Aspergillus niger JMU-TS528. Int J Biol Macromol 85:391–399. https://doi.org/10.1016/j.ijbiomac.2015.12.093
Li L, Liao H, Yang Y et al (2018a) Improving the thermostability by introduction of arginines on the surface of α-L-rhamnosidase (r-Rha1) from Aspergillus niger. Int J Biol Macromol 112:14–21. https://doi.org/10.1016/j.ijbiomac.2018.01.078
Li LJ, Wu ZY, Yu Y et al (2018b) Development and characterization of an α-L-rhamnosidase mutant with improved thermostability and a higher efficiency for debittering orange juice. Food Chem 245:1070–1078. https://doi.org/10.1016/j.foodchem.2017.11.064
Li L, Gong J, Wang S et al (2019) Heterologous expression and characterization of a new clade of aspergillus α-L-rhamnosidase suitable for citrus juice processing. J Agric Food Chem 67:2926–2935. https://doi.org/10.1021/acs.jafc.8b06932
Li L, Gong J, Li W et al (2020) Enhancement in affinity of Aspergillus niger JMU-TS528 α-L-rhamnosidase (r-Rha1) by semiconservative site-directed mutagenesis of (α/α)6 catalytic domain. Int J Biol Macromol 151:845–854. https://doi.org/10.1016/j.ijbiomac.2020.02.157
Li L, Li W, Gong J et al (2021) An effective computational-screening strategy for simultaneously improving both catalytic activity and thermostability of a-L-rhamnosidase. Biotechnol Bioeng 118:3409–3419. https://doi.org/10.1002/bit.27758
Li Q, Ge L, Zheng D et al (2022) Screening and characterization of a GH78 α-L-rhamnosidase from aspergillus terreus and its application in the bioconversion of icariin to icaritin with recombinant β-glucosidase. Enzyme Microb Technol 153:109940. https://doi.org/10.1016/j.enzmictec.2021.109940
Li YT, Huang T, Wang JZ et al (2023) Efficient acquisition of high-purity cyanidin-3-O-glucoside from mulberry fruits: an integrated process of ATPS whole-cell transformation and semi-preparative HPLC purification. Food Chem 404:134651. https://doi.org/10.1016/j.foodchem.2022.134651
Liao H, Gong JY, Yang Y et al (2019) Enhancement of the thermostability of Aspergillus niger α-L-rhamnosidase based on PoPMuSiC algorithm. J Food Biochem 43:e12945. https://doi.org/10.1111/jfbc.12945
Liu Q, Lu L, Xiao M (2012a) Cell surface engineering of α-L-rhamnosidase for naringin hydrolysis. Bioresour Technol 123:144–149. https://doi.org/10.1016/j.biortech.2012a.05.083
Liu TQ, Hong SY, Chun ZZ et al (2012b) Aspergillus niger DLFCC-90 Rhamnoside Hydrolase, a New type of Flavonoid Glycoside Hydrolase. Appl Environ Microbiol 78:4752–4754. https://doi.org/10.1128/AEM.00054-12
Liu F, Wei B, Cheng L et al (2022) Co-immobilizing two glycosidases based on cross-linked enzyme aggregates to enhance enzymatic properties for achieving high titer icaritin biosynthesis. J Agric Food Chem 70:11631–11642. https://doi.org/10.1021/acs.jafc.2c04253
Lou H, Liu X, Liu S et al (2022) Purification and characterization of a novel α-L-Rhamnosidase from Papiliotrema laurentii ZJU-L07 and its application in production of icariin from epimedin C. J Fungi 8:644. https://doi.org/10.3390/jof8060644
Lu L, Liu Q, Jin L et al (2015) Enzymatic synthesis of rhamnose containing chemicals by reverse hydrolysis. PLoS ONE 10:e0140531. https://doi.org/10.1371/journal.pone.0140531
Lyu W, Wu Y, Liu Y et al (2016) Draft genome sequence of Bacillus litoralis C44, isolated from chinese scholar tree (Sophora japonica) forest soil. Genome Announc 4:e01059–e01016. https://doi.org/10.1128/genomeA.01059-16
Lyu Y, Zeng W, Du G et al (2019) Efficient bioconversion of epimedin C to icariin by a glycosidase from aspergillus nidulans. Bioresour Technol 289:121612. https://doi.org/10.1016/j.biortech.2019.121612
Manzanares P, van den Broeck HC, de Graaff LH et al (2001) Purification and characterization of two different α-L-rhamnosidases, RhaA and RhaB, from Aspergillus aculeatus. Appl Environ Microbiol 67:2230–2234. https://doi.org/10.1128/aem.67.5.2230-2234.2001
Markosova K, Weignerova L, Rosenberg M et al (2015) Upscale of recombinant α-L-rhamnosidase production by Pichia pastoris MutS strain. Front Microbiol 6:1140. https://doi.org/10.3389/fmicb.2015.01140
Mensitieri F, De Lise F, Strazzulli A et al (2018) Structural and functional insights into RHA-P, a bacterial GH106 α-L-rhamnosidase from Novosphingobium sp PP1Y. Arch Biochem Biophys 648:1–11. https://doi.org/10.1016/j.abb.2018.04.013
Miake F, Satho T, Takesue H et al (2000) Purification and characterization of intracellular α-L-rhamnosidase from Pseudomonas paucimobilis FP2001. Arch Microbiol 173:65–70. https://doi.org/10.1007/s002030050009
Michlmayr H, Brandes W, Eder R et al (2011) Characterization of two distinct glycosyl hydrolase family 78 α-L-rhamnosidases from Pediococcus acidilactici. Appl Environ Microbiol 77:6524–6530. https://doi.org/10.1128/AEM.05317-11
Miyata T, Kashige N, Satho T et al (2005) Cloning, sequence analysis, and expression of the gene encoding Sphingomonas paucimobilis FP2001 α-L-rhamnosidase. Curr Microbiol 51:105–109. https://doi.org/10.1007/s00284-005-4487-8
Monti D, Pisvejcová A, Kren V et al (2004) Generation of an α-L-rhamnosidase library and its application for the selective derhamnosylation of natural products. Biotechnol Bioeng 87:763–771. https://doi.org/10.1002/bit.20187
Ndeh D, Rogowski A, Cartmell A et al (2017) Complex pectin metabolism by gut bacteria reveals novel catalytic functions. Nature 544:65–70. https://doi.org/10.1038/nature21725
Nghi do H, Bittner B, Kellner H et al (2012) The wood rot ascomycete Xylaria polymorpha produces a novel GH78 glycoside hydrolase that exhibits α-L-rhamnosidase and feruloyl esterase activities and releases hydroxycinnamic acids from lignocelluloses. Appl Environ Microbiol 78:4893–4901. https://doi.org/10.1128/AEM.07588-11
O’Neill EC, Stevenson CE, Paterson MJ et al (2015) Crystal structure of a novel two domain GH78 family α-rhamnosidase from Klebsiella oxytoca with rhamnose bound. Proteins 83:1742–1749. https://doi.org/10.1002/prot.24807
Park SY, Kim JH, Kim DH (2005) Purification and characterization of quercitrin-hydrolyzing α-L-rhamnosidase from Fusobacterium K-60, a human intestinal bacterium. J Microbiol Biotechnol 15:519–524
Peng C, Li R, Ni H et al (2021a) The effects of α-L-rhamnosidase, β-D-glucosidase, and their combination on the quality of orange juice. J Food Process Pres 45:e15604. https://doi.org/10.1111/jfpp.15604
Peng L, Tan W, Lu Y et al (2021) Convenient immobilization of α-L-rhamnosidase on cerium-based metal-organic frameworks nanoparticles for enhanced enzymatic activity and recyclability. ChemCatChem. https://doi.org/10.1002/cctc.202101489
Qian S, Yu H, Zhang C et al (2005) Purification and characterization of dioscin-α-L-rhamnosidase from pig liver. Chem Pharm Bull 53:911–914. https://doi.org/10.1248/cpb.53.911
Qian S, Wang H, Zhang C et al (2013) Isolation and characterization of dioscin-α-L-rhamnosidase from bovine liver. J Mol Catal B: Enzym 97:31–35. https://doi.org/10.1016/j.molcatb.2013.07.007
Rodrigues GR, Pinto OHB, Schroeder LF et al (2020) Unraveling the xylanolytic potential of Acidobacteria bacterium AB60 from Cerrado soils. FEMS Microbiol Lett. https://doi.org/10.1093/femsle/fnaa149
Rojas NL, Voget CE, Hours RA et al (2011) Purification and characterization of a novel alkaline α-L-rhamnosidase produced by Acrostalagmus luteoalbus. J Ind Microbiol Biotechnol 38:1515–1522. https://doi.org/10.1007/s10295-010-0938-8
Shanmugam V, Yadav KD (1995) Extracellular production of α-rhamnosidase by Rhizopus nigricans. Indian J Exp Biol 33:705–707
Shin NR, Moon JS, Shin SY et al (2016) Isolation and characterization of human intestinal Enterococcus avium EFEL009 converting rutin to quercetin. Lett Appl Microbiol 62:68–74. https://doi.org/10.1111/lam.12512
Shin KC, Seo MJ, Oh DK et al (2019) Cloning and characterization of α-L-rhamnosidase from Chloroflexus aurantiacus and its application in the production of isoquercitrin from rutin. Biotechnol Lett 41:419–426. https://doi.org/10.1007/s10529-019-02648-8
Singh P, Sahota P, Bhadra F et al (2015a) Optimization, production and scale up of debittered kinnow beverage by α-L-rhamnosidase producing yeast. Emir J Food Agr 27:548–555. https://doi.org/10.9755/ejfa.184383
Singh P, Sahota PP, Singh RK (2015b) Evaluation and characterization of new α-L-rhamnosidase-producing yeast strains. J Gen Appl Microbiol 61:149–156. https://doi.org/10.2323/jgam.61.149
Slámová K, Kapešová J, Valentová K (2018) Sweet Flavonoids”: glycosidase-catalyzed modifications. Int J Mol Sci 19:2126. https://doi.org/10.3390/ijms19072126
Spohner SC, Zahn D, Schaum V et al (2015) Recombinant α-L-rhamnosidase from aspergillus terreus in selective trimming of α-L-rhamnose from steviol glycosides. J Mol Catal B: Enzym 122:248–254. https://doi.org/10.1016/j.molcatb.2015.09.009
Sun J, Li W, Liao H et al (2022) Adding sorbitol improves the thermostability of α-L-rhamnosidase from Aspergillus niger and increases the conversion of hesperidin. J Food Biochem 46:e14055. https://doi.org/10.1111/jfbc.14055
Suzuki H (1962) Hydrolysis of flavonoid glycosides by enzymes (rhamnodiastase) from Rhamnus and other sources. Arch Biochem Biophys 99:476–483. https://doi.org/10.1016/0003-9861(62)90296-5
Terry B, Ha J, De Lise F et al (2020) The crystal structure and insight into the substrate specificity of the α-L rhamnosidase RHA-P from Novosphingobium sp PP1Y. Arch Biochem Biophys 679:108–189. https://doi.org/10.1016/j.abb.2019.108189
Thomas DW, Smythe CV, Labbee MD (1958) Enzymatic hydrolysis of naringin, the bitter principle of grapefruit. J Food Sci 23:591–598. https://doi.org/10.1111/j.1365-2621.1958.tb17609.x
Ting SV (1958) Fruit glycoside hydrolysis, enzymic hydrolysis of naringin in grapefruit. J Agr Food Chem 6:546–549. https://doi.org/10.1021/jf60089a010
Wang J, Sun GX, Yu L et al (2013) Enhancement of the selective enzymatic biotransformation of rutin to isoquercitrin using an ionic liquid as a co-solvent. Bioresour Technol 128:156–163. https://doi.org/10.1016/j.biortech.2012.10.098
Wang D, Zheng P, Chen P (2019) Production of a recombinant α-L-rhamnosidase from Aspergillus niger CCTCC M 2018240 in Pichia pastoris. Appl Biochem Biotechnol 189:1020–1037. https://doi.org/10.1007/s12010-019-03020-2
Wang D, Zheng P, Chen P et al (2020) Highly efficient enzymatic conversion of rutin to isoquercitrin and L-rhamnose using deep eutectic solvents. ACS Sustain Chem Eng 8:14905–14913. https://doi.org/10.1021/acssuschemeng.0c04797
Wang D, Zheng P, Chen P et al (2021) Immobilization of α-L-rhamnosidase on a magnetic metal-organic framework to effectively improve its reusability in the hydrolysis of rutin. Bioresour Technol 323:124611. https://doi.org/10.1016/j.biortech.2020.124611
Wang C, Xia N, Zhu S et al (2022) Green synthesis of hesperitin dihydrochalcone glucoside by immobilized α-L-rhamnosidase biocatalysis based on Fe3O4/MIL-101(cr) metal-organic framework. Polyhedron 220:115856. https://doi.org/10.1016/j.poly.2022.115856
Weignerova L, Marhol P, Gerstorferova D et al (2012) Preparatory production of quercetin-3-β-D-glucopyranoside using alkali-tolerant thermostable α-L-rhamnosidase from aspergillus terreus. Bioresour Technol 115:222–227. https://doi.org/10.1016/j.biortech.2011.08.029
Wu S, Ni H, Xiao A et al (2010) Preliminary optimization of fermentation conditions for production of naringinase by Aspergillus niger DB056. Microbiol China 37:1305–1311. https://doi.org/10.13344/j.microbiol.china.2010.09.008
Wu T, Pei J, Ge L et al (2018) Characterization of a α-L-rhamnosidase from Bacteroides thetaiotaomicron with high catalytic efficiency of epimedin C. Bioorg Chem 81:461–467. https://doi.org/10.1016/j.bioorg.2018.08.004
Xie J, Zhang S, Tong X et al (2020) Biochemical characterization of a novel hyperthermophilic α-L-rhamnosidase from Thermotoga petrophila and its application in production of icaritin from epimedin C with a thermostable β-glucosidase. Process Biochem 93:115–124. https://doi.org/10.1016/j.procbio.2020.03.019
Xie J, Zhao J, Zhang N et al (2022) Efficient production of isoquercitin, icariin and icariside II by a novel thermostable α-L-rhamnosidase PodoRha from Paenibacillus odorifer with high α-1,6-/α-1,2-glycoside specificity. Enzyme Microb Technol 158:110039. https://doi.org/10.1016/j.enzmictec.2022.110039
Xu L, Liu X, Yin Z et al (2016) Site-directed mutagenesis of α-L-rhamnosidase from Alternaria sp L1 to enhance synthesis yield of reverse hydrolysis based on rational design. Appl Microbiol Biotechnol 100:10385–10394. https://doi.org/10.1007/s00253-016-7676-4
Xu L, Liu X, Li Y et al (2019) Enzymatic rhamnosylation of anticancer drugs by an α-L-rhamnosidase from Alternaria sp L1 for cancer-targeting and enzyme-activated prodrug therapy. Appl Microbiol Biotechnol 103:7997–8008. https://doi.org/10.1007/s00253-019-10011-0
Yadav V, Yadav PK, Yadav S et al (2010) α-l-Rhamnosidase: a review. Process Biochem 45:1226–1235. https://doi.org/10.1016/j.procbio.2010.05.025
Yadav V, Yadav S, Yadava S et al (2011) α-L-Rhamnosidase from Aspergillus flavus MTCC-9606 isolated from lemon fruit peel. Int J Food Sci Tech 46:350–357. https://doi.org/10.1111/j.1365-2621.2010.02498.x
Yadav S, Yadav V, Yadav S et al (2012a) Purification, characterisation and application of α-L-rhamnosidase from Penicillium citrinum MTCC-8897. Int J Food Sci Tech 47:290–298. https://doi.org/10.1111/j.1365-2621.2011.02838.x
Yadav S, Yadav V, Yadava S et al (2012b) Purification and functional characterisation of an α-L-rhamnosidase from Penicillium citrinum MTCC-3565. Int J Food Sci Tech 47:1404–1410. https://doi.org/10.1111/j.1365-2621.2012.02987.x
Yadav V, Yadav S, Yadav S et al (2012c) α-L-rhamnosidase from Aspergillus clavato-nanicus MTCC-9611 active at alkaline pH. Appl Biochem Micro 48:295–301. https://doi.org/10.1134/s0003683812030155
Yadav S, Yadav RSS, Yadav KDS (2013a) An α-L-rhamnosidase from Aspergillus awamori MTCC-2879 and its role in debittering of orange juice. Int J Food Sci Tech 48:927–933. https://doi.org/10.1111/ijfs.12043
Yadav S, Yadava S, Yadav KDS (2013b) Purification and characterization of α-L-rhamnosidase from Penicillium corylopholum MTCC-2011. Process Biochem 48:1348–1354. https://doi.org/10.1016/j.procbio.2013.05.001
Yadav S, Yadava S, Yadav KD (2017) α-L-rhamnosidase selective for rutin to isoquercitrin transformation from Penicillium griseoroseum MTCC-9224. Bioorg Chem 70:222–228. https://doi.org/10.1016/j.bioorg.2017.01.002
Yanai T, Sato M (2000) Purification and characterization of an α-L-rhamnosidase from Pichia angusta X349. Biosci Biotechnol Biochem 64:2179–2185. https://doi.org/10.1271/bbb.64.2179
Ye H, Li X, Li L et al (2022) Homologous expression and characterization of α-L-rhamnosidase from Aspergillus niger for the transformation of flavonoids. Appl Biochem Biotechnol 194:3453–3467. https://doi.org/10.1007/s12010-022-03894-9
Yu S, Shan X, Lyv Y et al (2022) Bioproduction of quercetin using recombinant thermostable glycosidases from Dictyoglomus thermophilum. Bioresour Bioprocess 9:48. https://doi.org/10.1186/s40643-022-00538-y
Zhang R, Zhang BL, Xie T et al (2015) Biotransformation of rutin to isoquercitrin using recombinant α-L-rhamnosidase from Bifidobacterium breve. Biotechnol Lett 37:1257–1264. https://doi.org/10.1007/s10529-015-1792-6
Zhang X, Sun H, Su Q et al (2017) Antidepressant-like activity of icariin mediated by group I mGluRs in prenatally stressed offspring. Brain Dev 39:593–600. https://doi.org/10.1016/j.braindev.2017.03.021
Zhang T, Yuan W, Li M et al (2018) Purification and characterization of an intracellular α-L-rhamnosidase from a newly isolated strain, Alternaria alternata SK37.001. Food Chem 269:63–69. https://doi.org/10.1016/j.foodchem.2018.06.134
Zhang S, Luo J, Dong Y et al (2021) Biotransformation of the total flavonoid extract of epimedium into icaritin by two thermostable glycosidases from Dictyoglomus thermophilum DSM3960. Process Biochem 105:8–18. https://doi.org/10.1016/j.procbio.2021.03.002
Zhu XC, Tang SY (2021) Enzymatic properties of α-L-rhamnosidase and the factors affecting its activity: a review. Chin J Biotechnol 37:2623–2632. https://doi.org/10.13345/j.cjb.200565
Zhu Y, Jia H, Xi M et al (2017) Purification and characterization of a naringinase from a newly isolated strain of Bacillus amyloliquefaciens 11568 suitable for the transformation of flavonoids. Food Chem 214:39–46. https://doi.org/10.1016/j.foodchem.2016.06.108
Zverlov VV, Hertel C, Bronnenmeier K et al (2000) The thermostable α-L-rhamnosidase RamA of Clostridium stercorarium: biochemical characterization and primary structure of a bacterial α-L-rhamnoside hydrolase, a new type of inverting glycoside hydrolase. Mol Microbiol 35:173–179. https://doi.org/10.1046/j.1365-2958.2000.01691.x
Funding
This research was financially supported by the Technology Innovation and Application Development Special Key Project of Chongqing (cstc2021jscx-jbgsX0002).
Author information
Authors and Affiliations
Contributions
LP, YZ and FZ: writing-original draft. ZW and JZ: supervision and funding acquisition. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pan, L., Zhang, Y., Zhang, F. et al. α-l-rhamnosidase: production, properties, and applications. World J Microbiol Biotechnol 39, 191 (2023). https://doi.org/10.1007/s11274-023-03638-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03638-9