Abstract
Natural hypersaline environments are inhabited by an abundance of prokaryotic and eukaryotic microorganisms capable of thriving under extreme saline conditions. Yeasts represent a substantial fraction of halotolerant eukaryotic microbiomes and are frequently isolated as food contaminants and from solar salterns. During the last years, a handful of new species has been discovered in moderate saline environments, including estuarine and deep-sea waters. Although Saccharomyces cerevisiae is considered the primary osmoadaptation model system for studies of hyperosmotic stress conditions, our increasing understanding of the physiology and molecular biology of halotolerant yeasts provides new insights into their distinct metabolic traits and provides novel and innovative opportunities for genome mining of biotechnologically relevant genes. Yeast species such as Debaryomyces hansenii, Zygosaccharomyces rouxii, Hortaea werneckii and Wallemia ichthyophaga show unique properties, which make them attractive for biotechnological applications. Select halotolerant yeasts are used in food processing and contribute to aromas and taste, while certain gene clusters are used in second generation biofuel production. Finally, both pharmaceutical and chemical industries benefit from applications of halotolerant yeasts as biocatalysts. This comprehensive review summarizes the most recent findings related to the biology of industrially-important halotolerant yeasts and provides a detailed and up-to-date description of modern halotolerant yeast-based biotechnological applications.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microorganisms able to colonize hyperosmotic environments are grouped according to the aw, which describes the chemical potential of free water in a given solution. Thus, microbes in growth media can be classified as xerotolerant (no absolute requirement of low aw) and xerophilic (capable of growth, under at least one set of conditions, at a water activity below 0.85) (Pitt and Hocking 2009 and references therein). A more comprehensive microbial classification strategy was proposed recently (Dakal et al. 2014), which considers the non-ionic or ionic osmolyte nature, and catalogues microbes as osmophilic, osmotolerant, and osmosensitive, or halophilic, halotolerant and, halosensitive, respectively. Table 1 shows the grouping of relevant yeast strains into these categories, some of which are considered in this work.
Amongst the 106 orders of fungi known at present, the phenomenon of tolerance for low aw is restricted to only 10. This physiological behavior is in most cases limited to a single genus or just to a few species. In the case of Wallemiales (Basidiomycota), Capnodiales, Dothideales and Eurotiales (Ascomycota), halophily is attributed to several groups within the same order without clear phylogenetical explanation (Gunde-Cimerman et al. 2009). These species grow at salinities exceeding 2.9 M NaCl, but most of them also grow well in salt-free media. In contrast to many typically halophilic prokaryotes isolated from hypersaline environments, most fungal species identified in these media should be considered halotolerant (Gostinčar et al. 2019).
Preferred locations for the isolation of xelotolerant yeasts include brine and soil of solar salterns. Butinar et al. (2005) found 43 unique yeast isolates, which were identified as Pichia guilliermondii, D. hansenii, Yarrowia lipolytica, Metschnikowia bicuspidata, Candida parapsilosis, Rhodosporidium sphaerocarpum, R. babjevae, Rhodotorula laryngis, and Trichosporon mucoides. More recent study by Chung et al. (2019) described fungal isolates from salterns mainly as black yeasts (H. werneckii, Phaeotheca triangularis, Aureobasidium pullulans and Trimmatostroma salinum), and other fungal genera such as Cladosporium, Aspergillus and Penicillium. Seawater remains an excellent source of halotolerant yeasts. Jones et al. (2015) provided an extensive list of identified marine fungi, which included a total number of 1112 species, among them 213 were yeasts. More recently, Mitchison-Field et al. (2019) identified 36 species of fungi in several marine locations and organisms around Woods Hole (MA, USA) including 16 Ascomycetes and 4 Basidiomycetes belonging to Aspergillus, Cladosporium, Candida, Cryptococcus, Metschnikowia, Meyerozyma, Penicillium, Hortaea, Rhodotorula, and Trichoderma genera. Osmotolerant yeasts, such as Candida, Debaryomyces, Pichia, Rhodotorula, and Zygosaccharomyces have been isolated from foods (reviewed in Butinar et al. 2005).
A recent study involving 626 yeast strains implied that yeast species tend to be more resistant to osmotic stress caused by carbohydrates rather than NaCl (Stratford et al. 2019). Osmosensitive yeast species were found to be equally affected by NaCl and glucose, while relative toxicity of salt became more pronounced in more osmoresistant species. The most glucose-resistant species tested were Zygosaccharomyces spp. while salt resistant species included Debaryomyces hansenii var. fabryi and Candida parapsilosis. According to this study, other yeast species such as Candida apicola, Torulaspora microellipsoides, Zygotorulaspora florentina and strains belonging to Candida pseudointermedia, Clavispora lusitania, Wickerhamomyces anomalus and Zygosaccharomyces bailii could be classified as salt-sensitive based on their tolerance to glucose (Stratford et al. 2019).
During the last years, and particularly in the past decade, our knowledge of osmotolerant and halotolerant yeasts has increased considerably due to the identification of an increasing number of isolates and the utilization of novel molecular techniques for their characterization. These discoveries resulted in development of new biotechnological applications, which utilize these microorganisms and/or their components. This review provides a detailed and comprehensive outline of the most recent advantages in this exciting field with a special focus on the main industrially-important halotolerant/halophilic yeasts and their potential applications.
Description of relevant halotolerant yeasts
Few halotolerant yeast species have been extensively studied during the last several years. Species such as H. werneckii, W. ichthyophaga, D. hansenii and Z. rouxii have become model organisms and due to their unique physiological traits provide the biological foundation for numerous biotechnological applications.
Hortaea werneckii, also known as black yeast, grows optimally between 0.8 and 1.7 M NaCl, but can tolerate up to 2 M concentration of MgCl2 or 1.7 M CaCl2, 10% NaCl with 12% glucose, and 50% glucose (reviewed in Zalar et al. 2019). It is the aetiological agent of a skin disorder, tinea nigra, characterized by black spots on hands and feet.
The fungal genus Wallemia comprises eight recognized species; among them W. sebi, W. mellicola, W. muriae and W. ichthyophaga are commonly associated with foods. Wallemia species can easily contaminate and spoil food preserved with high amounts of salt or sugar. An increase in NaCl concentration from 5 to 15% (w/v) in the growth media increases the production of the toxic metabolites wallimidione, walleminol, and walleminone (Jančič et al. 2016). W. ichthyophaga can grow in NaCl-saturated media and is an obligate halophilic microorganism, which requires at least 1.5 M NaCl (or some other osmolyte at equivalent aw) for growth, achieving its maximal proliferation in concentrations ranging between 2.6 and 3.4 M NaCl (Zajc et al. 2014a; Zalar et al. 2005). This species is also capable of growing in saturated KCl and MgSO4 solutions and in 2.1 M MgCl2 (Zajc et al. 2014b).
Debaryomyces hansenii can tolerate as much as 4.0 M NaCl. Its growth rate is not affected in culture media containing 4% (w/v) of marine salts (NaCl content around 0.55 M). However, when the concentration of this salt exceeds 2.0 M both the growth and glucose consumption slow down to 42–46% and 22–24% when compared to control cultures without salts, respectively (Capusoni et al. 2019). This species plays a role in processing and maturation of common foods such as cheeses and sausages. In addition, this yeast limits the presence of undesirable harmful bacteria in those foods. D. hansenii can produce effective killer toxins against other yeasts (reviewed in Prista et al. 2016). It is an oleaginous microorganism that can accumulate massive amounts of lipids (up to 70% of its total biomass). D. hansenii uses a broad spectrum of carbon sources that are generally catabolized by a respiratory metabolism. It is one of the important extremophilic yeasts that can utilize xylose, and it has been shown to be able to produce xylitol, arabitol and riboflavin. D. hansenii genes encoding xylose dehydrogenase and xylose/H+ transporter have been used for second-generation bioethanol produced during the fermentation of pentoses in S. cerevisiae.
The genus Zygosaccharomyces comprises seven species with osmo- and halotolerant Z. rouxii, which is the most phylogenetically related to S. cerevisiae and used in some fermentation processes (Dakal et al. 2014). The most glucose-resistant species identified corresponds to Zygosaccharomyces spp. (Stratford et al. 2019 and references therein).
Other relevant features of these yeasts are compilated in Table 2.
In addition to those described above, many other halotolerant yeast species have been described worldwide (Table S1 in the Supplementary Material). Species belonging to Candida, Meyerozyma, Millerozyma, Pichia, Rhodotorula, Schwannyomyces, Torulaspora, and Yarrowia genera have been broadly employed for biotechnological applications. It is worth mentioning that some of these organisms (Debaryomyces fabryi, Kluyveromyces marxianus, Schwannyomyces etchellsii, and Schwanniomyces polymorphus), have been demonstrated to form biofilms in seawater media in the absence of glucose (Zarnowski et al. 2021).
Besides H. werneckii, other black yeasts have been identified in hypersaline environments. For example, Phaeotheca triangularis, Aureobasidium pullulans and Trimmatostroma salinum were found in salterns (Chung et al. 2019 and references therein). These microorganisms are polymorphic fungi that undergo yeast-like, filamentous, and meristematic development, and hyphal forms are predominantly observed on solid agar media. While H. werneckii and P. triangularis grow in media containing up to 25% (w/v) NaCl, A. pollutans cannot proliferate at concentrations higher than 10% and T. salinum grows optimally at salinities from 2 to 6% (w/v), but tolerates NaCl concentrations up to 26%. A. penicillioides has been defined as an obligate halophile because its isolates from salterns are unable to grow without the addition of 10% solar salt. Mitchison-Field et al. (2019) have characterized other black yeasts such as Kunifia petricola which produced spherical cells that appeared to expand isotropically and, at the colony level, generated linear chains with branched networks. Aureobasidium pullulans formed up to six buds from a multinuclear mother cell. In Phaeotheca salicorniae, cells have the ability to switch bidirectionally between yeast-like and hyphal-like types; besides, orthogonal divisions not previously described in other fungi, which result in triangular-shaped compartments, have been shown by these authors.
Mechanisms for adaptation of halotolerant/osmotolerant yeasts to extreme environmental conditions
Several research groups have described the main survival strategies of halotolerant and osmotolerant yeasts to counteract changes in turgor pressure and therefore adapt to life at low aw (Kogej et al. 2007; Gunde-Cimerman et al. 2009; Gostinčar et al. 2011; Lenassi et al. 2013; Plemenitaš et al. 2014; Prista et al. 2016; Chung et al. 2019; Sánchez et al. 2020). These strategies include modulation of membrane composition (sterol to phospholipid ratio), synthesis of compatible solutes, maintenance of intracellular Na+ concentrations below toxic levels, activity of transporters of glycerol and alkali-metal-cation, resistance of some enzymes to salt stress, optimized energy-obtaining pathways, and activity of some organelles. In addition, gene duplication events have been described in several yeasts, which one can consider as the strategy of enriching the genomic toolbox that promotes survival under extreme environment conditions (Zalar et al. 2019).
Based on the accumulated information about the ability of H. werneckii to withstand extreme aw, Gostinčar et al. (2011) proposed an integrated model of extreme halotolerance that could be applied to most yeasts showing this property. Figure 1 outlines the sequential steps in response to osmotic stress in halotolerant yeasts that occur according to this model. The High Osmolarity Glycerol (HOG) Mitogen-Activated Protein Kinase (MAPK) pathway was initially described in the baker’s yeast (S. cerevisiae) and is responsible for the transcription response to hyperosmotic stress (Posas et al. 2000). This pathway consists of two upstream branches (SHO1 and SLN1) and common downstream elements Pbs2p MAPKK and Hog1p MAPK. Activation of this pathway causes rapid nuclear accumulation of Hog1p, essentially leading to the expression of target genes. The optimization of energy metabolism that follows the signal transduction pathway is due to an increase in the number of mitochondria and upregulation of expression of genes involved in ATP synthesis, the glycolytic pathway, the tricarboxylic acid cycle, the pentose-phosphate pathway, the modulation of energy storage and, maybe, the harvesting of light by a transmembrane rhodopsin proton pump (Gostinčar et al. 2011). This energy supply is key for other molecular processes that drive the adaptation mechanisms in osmotolerant/halotolerant yeasts, as described below.
Osmosensing proteins
The HOG pathway in Z. rouxii contains two functionally redundant plasma membrane osmosensors Sln1 and Sho1, initially identified in S. cerevisiae (Dakal et al. 2014), whereas in H. werneckii two copies of HwSho1 and HwHhk7 exist. HwHhk7A/B proteins are soluble, cytosolic proteins (Gostinčar et al. 2011), and the transcription of the encoding genes depends on the extracellular salt concentration (Lenassi and Plemenitas 2007). In W. ichthyophaga, WiSho1, a homolog of the ScSho1 protein, has been identified in only one copy but there is no evidence of Sln1-like histidine kinases (Plemenitaš et al. 2014; Konte et al. 2016).
Signaling cascade to Hog1
In Z. rouxii a putative ZrPbs2 MAPKK kinase is present, which is similar to S. cerevisiae Pbs2. In H. werneckii two copies of HwYpd1, HwSsk1 and Ssk2 (SLN branch), Ste20 and Ste11 (SHO1 branch) and Pbs2 have been identified. Heterologously expressed HwPbs2 in S. cerevisiae does not support functionality of SHO1 branch (Plemenitaš et al. 2014). In the case of W. ichthyopaga homologs of these proteins have been found in one copy, but the SHO1 branch components seem not to be involved in HOG signaling, and WiSte11 and WiPbs2 cannot complement S. cerevisiae mutants (Plemenitaš et al. 2014). The signaling cascade in D. hansenii is still poorly understood. The S. cerevisiae PBS2 homolog DPBS2 can partially complement the osmo-sensitivity of pbs2 mutation in S. cerevisiae (Sharma and Mondal 2005 and references therein). These authors have reported that the two MAPKKs exhibit considerable sequence divergence in non-catalytic regions but their kinase domains show strong similarity.
The MAPK Hog1
Depending on the haploidy, Z. rouxii possesses one or two HOG1 (ZrHOG) copies, which are homologs of the S. cerevisiae HOG1 (Iwaki et al. 1999; Kinclová et al. 2001). In H. werneckii HwHog1 shows high homology to S. cerevisiae Hog1 and a typical C-terminal common docking motif (Turk and Plemenitas 2002). These authors found several important differences between the two kinases: HwHog1 is shorter, under nonstress conditions it is located on the plasma membrane instead of being in the cytoplasm, and it is totally activated only at extremely high NaCl concentrations. Kejžar et al. (2015a) reported on the existence of two redundant kinases, HwHog1A and HwHog1B, whose activities are crucial for growth at osmolyte concentrations ≥ 3 M. These two kinases can both rescue the osmosensitive phenotype of the S. cerevisiae hog1∆ strain and the transcription of the encoded genes is salt-dependent. Low basal phosphorylation of HwHog1A/B has been described in non-stressed cells and in those treated with 1.8 M NaCl or KCl, or ≤ 2 M sorbitol (Kejžar et al. 2015b), but this modification is clearly observed under osmolyte concentrations ≥ 3 M, and displays differences depending on the osmolyte.
Among the two Hog1-like kinases, WiHog1A and WiHog1B, identified in W. ichtyophaga, the former cannot fully complement the function of ScHog1, but the latter is a functional Hog1-like kinase that improves halotolerance of the S. cerevisiae hog1∆ strain (Konte and Plemenitas 2013). A particular trait of WiHog1 is the constitutive phosphorylation under optimal osmotic conditions (3.4 M NaCl) and the dephosphorylation when the cells cope with hypo/hyper-osmolar stress (Konte and Plemenitas 2013).
Regarding D. hansenii, DhHOG1 has been cloned and sequenced (Bansal and Mondal 2000), and its expression in the S. cerevisiae hog1∆ mutant strain confers ability to grow under hyperosmotic conditions and to induce glycerol overproduction. The characterization of a null HOG1 mutant in this yeast (Sánchez et al. 2020) has revealed that the HOG pathway is required for survival under high external osmolarity but, similarly to what was described for H. werneckii, is dispensable under low and mid-osmotic conditions. Strong DhHog1 phosphorylation depends on high external osmolarity, and its pattern is essentially consistent with its translocation to the nucleus, in which long accumulation occurs. These authors also demonstrated a role for DhHog1 in the response to oxidative stress, which can be triggered by NaCl or KCl (Ramos-Moreno et al. 2019).
Intracellular levels of compatible osmolytes
Polyols act as compatible solutes to counteract osmotic stress in yeasts, with glycerol being the most abundant among them. The intracellular levels of this molecule in S. cerevisiae depend on glycerol-3-phosphate dehydrogenase (Gpd), dl-glycerol-3-phosphate phosphatase (Gpp) and glycerol dehydrogenase (Gcy) enzymes, and the activity of transporters such as Stl1 (glycerol-proton symporter) and Fps1 (involved in efflux of glycerol and xylitol).
Hortaea werneckii genome contains two salt-inducible GPD1 genes that show similar gene transcription regulation and share 98% amino-acid sequence identity. Leakage of glycerol is reduced by the incorporation of a layer of melanin into the cell wall. Thus, the degree of melanization is altered in response to various exposures to salinities (Plemenitas et al. 2008). In W. ichthyophaga there is only one WiGDP1 gene and its expression is less induced by salt. The gene can rescue the salt tolerance phenotype of the S. cerevisiae gpd1gpd2 double mutant (Lenassi et al. 2011). A second homolog of ScGPD1 has also been found along with homologs of ScGPP1 (one), ScSTL1 (four) and ScFPS1 (three) genes (reviewed in Plemenitaš et al. 2014).
DhGPD1 and DhGPD2 have been described in D. hansenii. They are upregulated upon hyperosmotic stress and their involvement in NaCl tolerance in this yeast has been demonstrated (Almagro et al. 2000; Gori et al. 2005). It is worth mentioning that DhGPD1 and DhMSN2 are modulated by stress at their transcriptional level (Ramos-Moreno et al. 2019). D. hansenii contains a unique Na+/glycerol symporter that also uses potassium ions and has not been identified in S. cerevisiae. Glycerol accumulation has been reported to be partially dependent on DhHog1, that fully upregulates the transcription of DhSTL1 gene and partially upregulates DhGPD1 (Sánchez et al. 2020).
Zygosaccharomyces rouxii possesses two putative glycerol-3-phosphate dehydrogenases (ZrGpd1p and ZrGpd2p), and two isogenes of the S. cerevisiae GCY1 (ZrGCY1 and ZrGCY2) (Iwaki et al. 2001). ZrGPD1 and ZrGPP2 have a main role in the production of glycerol, but unlike in S. cerevisiae, their expression is not regulated in response to salt stress and hence seems to by-pass the HOG pathway control (Dakal et al. 2014).
Cell turgor is also maintained by the accumulation of large amounts of other compatible solutes, such as erythritol, arabitol, mannitol, and mycosporine-glutaminol-glucoside in H. werneckii; arabitol and traces of mannitol in W. ichthyophaga, and trehalose, arabinitol, glutamic acid, and alanine in D. hansenii (Zajc et al. 2014b; Sánchez et al. 2020). In Z. rouxii, the production of compatible solutes depends upon the osmoticum (Dakal et al. 2014). Under salt stress, d-arabitol is produced in excess and accumulated, and the glycerol concentration remains invariable, while fructose and glucose-containing medium has been associated with mannitol production, which is inhibited by salt.
Levels and activity of membrane transporters
The enrichment in plasma membrane transporters is characteristic of the extremophile yeast H. werneckii (Lenassi et al. 2013; Plemenitaš et al. 2014), with a number of homologs ranging between four and eight like in the case of Trk1/2 channels (for potassium uptake), Tok channels (potassium efflux), Na+, K+/H+ antiporter Nha1, Na+/Pi symporter Pho89, Ena Na+ P-type ATPases and Pma1 P-type H+-ATPase. It is also worth mentioning a salt-dependent transcription enrichment of the HwPMAs genes in this yeast. Homologs of S. cerevisiae proteins involved in cation transport across the vacuole and Golgi apparatus membranes have also been found in H. werneckii and, in some cases, enrichment is found.
Wallemia ichthyophaga cells maintain low intracellular K+ and Na+ at constant salinities, but during hyperosmotic shock the amounts of both cations increase significantly (Zajc et al. 2014b). The ratio of intracellular K+ and Na+ is higher across the whole salinity range than that found in H. werneckii, and decreases with increasing salinities (reviewed in Plemenitaš et al. 2014). The number of cation transporters in W. ichthyophaga is low and mostly independent of salt, except for the P-type ATPases HwEna1 and HwEna2. Considering these findings, it seems that W. ichthyophaga resists higher sodium concentrations and has a poor capability to adjust to changing environments, probably because this microbe prefers relatively constant growth conditions with extremely high salinity levels (Plemenitaš et al. 2014).
In Z. rouxii growing under increasing salinity, intracellular Na+ content rises slightly, intracellular K+ content decreases significantly, and the activity of Na+/K+-ATPase is around four-fold higher (Wang et al. 2020a). To mediate the efflux of excessive cations through the plasma membrane this yeast uses the ZrEna1, ZrNha1 and ZrSod2-22 transporters. Unlike in S. cerevisiae, ZrEna1 has little relevance in salt tolerance and the major pump-out activity relies on ZrSod22 and its variants.
DhTRK1, DhHAK1, DhENA1/DhENA2, DhNHA1, DhKHA1, and DhNHX1, the genes encoding transporters participating in potassium and sodium inward and outward fluxes and organellar sequestration, have been cloned and characterized in D. hansenii (reviewed in Prista et al. 2016). The upregulation of DhENA1 and DhENA2 under hyperosmotic stress has been demonstrated (Almagro et al. 2000; Gori et al. 2007). Recently, Herrera et al. (2017) have unveiled two new halotolerance determinants in D. hansenii: the function of the vacuole in the tight regulation of the maintenance of low Na+ levels in the cytosol and in K+ homeostasis, and the good performance of the cells when cytosolic values of Na+ are significantly higher than those reported in the model yeast. Unlike other fungi, D. hansenii is considered a sodium-ion includer. Cytosolic Na+ levels reach values around three times higher than those reported in S. cerevisiae.
Changes in the membrane properties associated to halotolerance
To preserve the integrity of the cells and the membrane-associated processes In H. werneckii and D. hansenii, membrane lipid composition is restructured in the presence of greater amounts of salt by changes in the expression of genes that are involved in lipid synthesis and modification. Compared to S. cerevisiae, the membranes are more fluid, the sterol-to-phospholipid ratio is lower, and fatty acid unsaturation increases (Gunde-Cimerman et al. 2009). The major fatty acid in the plasma membrane of D. hansenii is oleic acid. Capusoni et al. (2019) have also reported that the exposure to hyper-osmotic conditions elicits reversible membrane depolarization and reduces the membrane permeability, not only in D. hansenii strains but also in other less osmotolerant species like S. cerevisiae. In Z. rouxii, however, ergosterol content increases, the unsaturated/saturated fatty acid ratio reduces, and cell membrane fluidity decreases under NaCl stress (Dakal et al. 2014).
Modifications in morphology and growth
The adaptation of H. werneckii to extremely saline conditions involve changes in cell size and colony appearance, cell-wall ultrastructure, morphology (which includes enhanced melanization), and meristematic growth (Kogej et al. 2007). Striking findings related to long-term adaptation changes in cell wall morphology, signaling pathways and the pentose phosphate cycle were also reported in a recent study of a H. werneckii isolate grown for over 7 years and for at least 800 generations under salinity far above this specie’s preferred optimum (Gostinčar et al. 2021).
Wallemia ichthyophaga grows in multicellular groups or as sarcina-like structures (Zalar et al. 2005). As salinity rises, these clumps increase in size and there is a decrease in the functional cell size due to a threefold thickening of the cell walls. Kralj Kuncic et al. (2010) concluded that the observed about ninefold increase in the expression of the cellulase/exo-1,3-β-glucanase gene when the NaCl concentration elevated from 1.7 M to saturated solution could provide protection to the cells from the hostile environmental conditions. On the other hand, the expansion of the hydrophobins protein family to 26 members from just 15 predicted in the last common ancestor of W. ichthyophaga and W. sebi (Zajc et al. 2013) results in both increased cell wall strength and rigidity and may have a role in the sarcina-like morphology and in the formation of compact cell clusters (Plemenitaš et al. 2014).
Other traits related to osmo(halo)tolerance are described in Table S2 in the Supplementary Material.
Biotechnological applications of halotolerant yeasts
Microorganisms, and especially yeasts, play a significant role in diverse biotechnological applications. Halotolerant yeasts have unique properties for these purposes, and have been broadly used in food production, as biocatalysts in the synthesis of valuable products, and also in biodegradation processes. Figure 2 outlines the most common biotechnological applications. Table S1 in the Supplementary Material contains a description of recently reported or proposed applications of some halotolerant yeast strains, which are used either as whole cells or just as individual enzymes. A more detailed description of several of these applications is provided below.
Food industry
Halophiles have been used in traditional fermentation processes increasing health benefits such as nutrient enrichment. These microbes enter the food chain at different steps, adapt to the environment to survive and grow, and generate useful active substances through metabolic processes, which results in the production of high sugar and/or salt fermented foods. However, the presence of halophiles may have adverse outcomes and result in undesired food spoilage and damage due to the accumulation of potentially toxic compounds such as biogenic amines, nitrates, and nitrosamines. This aspect of halophiles’ biology causes severe economic losses in the production of bread, cereals, spices, dairy products, chocolate, fermented sauces, soft drinks, fruits, jams and syrups (Gostinčar et al. 2011; Dakal et al. 2014; Qi et al. 2014a, b). It has been estimated that food loss caused by yeasts in Australia could exceed $ 10,000,000 annually, and could represent around 5–10% of all food production (Pitt and Hocking 2009).
Zygosaccharomyces yeasts (also called “ZygoFactories”) play a central role in the production of traditional fermented highly sugary and salty food such as soy sauce, soybean paste, miso, kombucha or balsamic vinegar. They also produce secondary metabolites that can serve as relevant flavor components and food additives. In this sense, Z. rouxii has a key role in the production of soy sauce, where it releases flavor compounds, which contribute to its fragrant and smoky aroma. It is also a crucial cell factory in the production of balsamic vinegar, converting concentrated sugars to ethanol as well as in ethanol-free beer production (Solieri 2021). However, due to their tolerance to salt, sugar and acid preservatives, Zygosaccharomyces strains are one of the main spoilage yeasts in the food industry (Dakal et al. 2014). Some wild Z. rouxii are known to impair the quality of soy sauce due to the generation of unpleasant odors caused by the formation of flor velum (Mogi and Watanabe 2020).
Candida versatilis is another of the major microbes in the soy sauce fermentation, which enriches the flavor and improves its quality. Qi et al. (2014a) have described an important effect of the salt stress in the quantity and nature of metabolites produced by this microorganism. Both synthesis and secretion of aldehydes and phenols were enhanced under these conditions. Elevated levels of a variety of biogenic amines were shown during the aging period of soy sauce production, which may be due to amino acid decarboxylation by yeast decarboxylase(s).
Debaryomyces hansenii is the predominant yeast species isolated from multiple common foods. Its metabolic activity is essential to the ripening process of cheeses and sausages such as salami, playing an important role in the final desirable organoleptic properties of these foods, and it can be used as a starter to develop their characteristic flavors and textures (Prista et al. 2016 and references therein). Different strains uniquely contribute to the final aroma and taste of these products, adding a whole range of distinct volatile compounds or altering their composition with distinctive proteolytic enzymes. It has been demonstrated, for instance, that D. hansenii generates ester and sulfur compounds, among other volatiles, responsible for the characteristic meat aroma and taste (Perea-Sanz et al. 2019). Bacterial microbiota on processed foods is influenced by the presence of this yeast due to its striking ability to modulate pH. It has been described as a common contaminant of foods and beverages, being associated with the spoilage of cheese, milk, yogurt, jam, and confectionery. However, the knowledge of its growth behavior under different conditions is useful to the establishment of food preservation strategies (Almagro et al. 2000).
Other described osmotolerant and/or halotolerant food yeasts include Millerozyma farinosa, a yeast species found primarily in alcoholic beverages such as beer and sake, H. werneckii, which is detected on salty foods, and Candida davenportii, C. stellata and C. magnoliae that have been predominantly associated with spoilage of sugary foods (reviewed in Dakal et al. 2014).
It is worth mentioning that halophilic yeasts produce zymocins or yeast killer (K) toxins, which can be developed as potential biocontrol agents in food industries and also in the treatment of fungal infections. Zymocins are heterogeneous antimycotical agents produced by different yeast strains. In fact, several halotolerant species like D. hansenii, Pichia membranifaciens, P. farinosa or C. nodaensis display or even increase K activity in the presence of salts like NaCl, and for this reason they can be interesting tools in several biotechnological applications, especially in the preservation of salt-fermented foods (da Silva et al. 2008 and references therein).
Bioremediation
The progress of industrialization has increased the use of chemicals in products such as fuels and industrial solvents. Pesticides, insecticides, and herbicides in agriculture constitute the main source of water pollution, and their improper disposal results in life-threatening air, water, and soil contamination. Conventional physical and chemical methods for treatment of pollutants are costly, time-consuming and result in generation of other contaminants in the process (Nikolaivits et al. 2020 and references therein). Bioremediation has emerged as a safe, low-cost, and eco-friendly alternative technology that offers sustainable removal of hazardous chemicals. The US Environmental Protection Agency (USEPA) defines bioremediation as a treatment process in which microorganisms are employed to degrade or modify toxic pollutants to less harmful products, thus reducing environmental pollutants generated by various anthropogenic activities.
Polycyclic aromatic hydrocarbons (PAHs) are a group of carcinogenic and highly toxic environmental pollutants with prolonged stability. Bacteria have been used for removal of PAHs, but their effectivity is low when these compounds have high molecular weights, like in the case of benzo[a]pyrene and phenanthrene. Several species of filamentous fungi and lately also yeasts have been used as bioremediating agents because they can use these compounds as growth substrates. As these pollutants are widely distributed in different ecosystems, including saline environments, halotolerant and halophilic yeasts like Candida sp. S1 (Hadibarata et al. 2017), Basidioascopus persicus EBL-C16 (Kamyabi et al. 2018), or R. mucilaginosa EXF-1630 (Martínez-Ávila et al. 2021) are essential bioplayers for the recuperation of these ecological niches.
Phenol and its derivatives are extremely hazardous contaminants, toxic to animals and humans, and are generated from a variety of industrial processes. Phenol is soluble in water, and is frequently found in industrial salty wastewater along with other contaminants like heavy metals. Candida sp. JS3 (Jiang et al. 2015), Debaryomyces sp. JS4 (Jiang et al. 2016) and C. tropicalis SDP-1 (Gong et al. 2021) are halotolerant yeasts that have been shown effective in degradation of these compounds. It has also been described that the immobilisation of Candida sp. in SiO2 nanoparticles had a better phenol removal performance compared to free cells (Jiang et al. 2018). The wastewater from the printing, textile and dyeing industries are representative of hypersaline wastewater which contain azo dyes, a vast group of highly toxic pollutants that are recalcitrant to biodegradation by bacteria. Halotolerant yeasts such as Sterygmatomyces halophilus SSA1575 (Al-Tohamy et al. 2020), Pichia occidentalis A2, Candida tropicalis A1 and C. tropicalis SYF-1 (Tang et al. 2020; Wang et al. 2020b) have been effective in the degradation of these hazardous contaminants.
Biocatalysis
Biocatalysis has been considered an alternative to classic chemical methods that fulfills the green chemistry principles (Anastas and Eghbali 2010). It involves the use of whole cells and enzymes, which are nature’s sustainable, renewable and biodegradable catalysts. They are used under mild temperature and pH conditions; water is used as an environmentally friendly solvent (Ni et al. 2014). In addition, no heavy metals are employed, and good control over chemo-, regio- and stereoselectivity are achieved (Hernáiz et al. 2010). Biocatalysis has a special impact on the pharmaceutical industry as most chiral drugs can be synthesized on a large-scale using enzymes or whole cells in any step of their synthesis (Wells and Meyer 2014).
Water is one of the most preferred solvents by the pharmaceutical industry because its reduced cost and its low impact in the environment (Anderson 2012). However, considerable water usage is another problem in industrial processes, and it is important to investigate alternatives to freshwater. Seawater represents the most important water reservoir with a lot of microorganisms adapted to the high salinity. Marine microorganisms and their enzymes constitute an important source of research into finding new properties as a result of singular physicochemical conditions, such as salt tolerance, hyperthermostability, barophilicity or cold adaptability (Trincone 2010), and in this sense, marine yeasts have been shown promising for biotechnological applications to perform bioprocesses under medium- or high- salinity conditions (Grande et al. 2012; Domínguez de María, 2013; Zaky et al. 2014, 2018; Zambelli et al. 2015; Serra et al. 2016; Andreu and del Olmo 2018, 2019, 2020). In this section, we show some examples of distinct catalytic activities to produce interesting valuable compounds that can be carried out by whole cells or enzymes in salty media.
The ability of halotolerant yeast S. etchellsii to produce the chiral precursor of ephedrine (R)-(-)-phenylacetylcarbinol out of benzaldehyde has been demonstrated in seawater with the conversion rate of the starting material being higher in this medium than in freshwater controls (Andreu and del Olmo 2018). Whole cells of salt-resistant Meyerozyma guilliermondii LM2, have shown nitrile hydrolyzing activity, converting different substrates with high yields and high enantioselectivity toward racemic substrates (Serra et al. 2019). It has also been found that several marine Pichia and Candida yeast isolates can catalyze the conversion reaction of monosodium glutamate to GABA only in the presence of glucose (Masuda et al. 2008).
Debaryomyces and Schwanniomyces halotolerant yeast strains have been used as cell factories for biocatalytic reduction of pro-chiral ketones into chiral alcohols using seawater as a solvent. In this medium, these yeasts show higher thermotolerance, resistance to different organic solvents and enhanced activity. Higher substrate charges and several recycling steps have also been possible in seawater when compared to analogical reactions carried out in freshwater (Andreu and del Olmo 2018, 2019). Other halotolerant yeast strains used for the asymmetric reduction of ketones have been S. cerevisiae FY86, K. marxianus, T delbrueckii, M. guilliermondii (Andreu and del Olmo 2020), and Rhodotorula mucilaginosa (Serra et al. 2016).
The competence of D. hansenii to synthesize biotechnologically relevant products has been well documented. In this sense, it is a valuable alternative to chemical syntheses, not only because of low costs but also for the scope of new products. For instance, as a halotolerant strain, it produces compatible solutes such as d-arabinitol. D. hansenii thiamine auxotrophs under thiamine limitation have also been shown to accumulate pyruvic acid, a product widely used in the chemical, pharmaceutical and agrochemical industries. This yeast has been considered a traditional “cell factory” of alditols, and its capacity to produce xylitol has been exploited for several decades. This product is a metabolic intermediate of d-xylose fermentation and is used in the food industry for its high sweetening properties (Breuer and Harms 2006). Salty substance feedstocks like fermented wastes, soy sauce or marine microalgae are interesting reservoirs for ethanol production due to their abundance. As salt concentration is an important parameter in this process due to its reducing effect on cell growth, the use of halotolerant yeasts for bioethanol production also has gained a lot of attention in recent years. Bioethanol production was achieved from salted algae by Citeromyces matritensis M37 (Okai et al. 2016), in a growth medium supplemented with 10% (w/v) molasses and 4% (w/v) NaCl by two new strains of K. marxianus (Karatay et al. 2019), as well as in seawater by S. cerevisiae AZ65 (Zaky et al. 2020). A recent review by Scapini et al. (2021) provides an updated data for seawater-based biorefineries.
Halotolerant yeasts belonging to the genera Cryptococcus, Schwanniomyces, Yarrowia, Pichia, Schizosaccharomyces and Williopsis have been shown to mediate the synthesis of nanoparticles (Mohite et al. 2017 and references therein). Both gold and silver nanoparticles generated by W. saturnus NCIM 3298 exhibited good catalytic properties in the NaBH4-based reduction of 4-nitrophenol to 4-aminophenol. This observation suggests they could potentially be used as antioxidants and catalysts in the cosmetic and pharmaceutical industries.
Finally, it is worth mentioning that not only whole cells of halotolerant microorganisms have been used as biocatalysts. Several halophilic enzymes, like glycosidases, proteases and lipases have been purified and characterized (Ruginescu et al. 2020). Enzymes derived from halophilic organisms could be useful in low-water or anhydrous solvent applications due to the effect of salt on lowering water activity. Organic solvents enhance reaction rate by increasing the solubility of nonpolar substrates, enhancing thermostability and allowing reactions usually unfavored in aqueous solution (Demirci et al. 2021 and references therein).
Design and use of novel systems for industrial processes
Besides the use of isolates with potential industrial interest, recent research also involves the development of new strategies such as microorganism combinations with enhanced properties or the rational design of efficient halotolerant enzymatic systems. An example of the former is the construction of a novel multipurpose oleaginous consortium designated as OYC-Y.BC.SH developed using three yeast cultures viz. Yarrowia sp. SSA1642, Barnettozyma californica SSA1518 and Sterigmatomyces halophilus SSA1511. This consortium has a high ability to accumulate large amounts of triacylglycerol lipids, grow on xylose, produce lipase and xylanase and it could rapidly decolorize commonly used textile reactive azo dyes. The specific enzyme activities were significantly higher than that of individual strains (Ali et al. 2020). On the other hand, rational design has been used for the development of new halotolerant enzymes for the synthesis of cytidine diphosphate choline, a drug applied in brain surgery patients. An increase in the incidence of surface acidic residues in the enzyme resulted in an improvement of the activity and IC50 of the mutant respect to the wild counterpart (Zheng et al. 2019).
In the future the construction of genetically modified strains useful for industrial processes is expected to become more common. However, this requires the development of vectors, markers and strategies for efficient transformation with heterologous DNA. Unfortunately, these tools have not become available for most of these yeasts yet. In the case of Z. rouxii an improved transformation method by electroporation has been proposed (Watanabe et al. 2010) and, more recently, a set of centromeric and episomal plasmids carrying antibiotic markers and Cre recombinase have been developed (Bizzarri et al. 2019). Metabolic engineering of Y. lipolytica and two-stage continuous fermentation process have been developed for a sustainable eicosapentaenoic acid (C20:5, EPA) production (Xie et al. 2017).
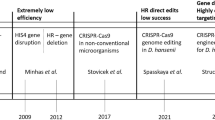
New molecular tools for genetic manipulation have also been developed for D. hansenii. Ricaurte and Govind (1999) introduced the first successful strategy for transformation using dhURA3 gene as a plasmid marker and an ura3 mutant strain. A more effective method of transformation by electroporation based on the DhHIS4 gene as the selectable marker and a histidine auxotrophic recipient strain has been reported more recently; this system is also used for gene disruption by homologous recombination, although with low rate (Minhas et al. 2009; Minhas and Biswas 2019). The ambiguous translation of the CUG codon in D. hansenii also makes genetic manipulation more difficult, while heterologous expression in S. cerevisiae may help with gene function determination. Recently, a novel plasmid-based CRISPRCUG/Cas9 tool for gene editing of prototrophic strains of D. hansenii and a NHEJ-deficient strain have been reported (Strucko et al. 2021). This method allows for a highly efficient introduction of point mutations and single/double gene deletions in D. hansenii.
The knowledge of genetic loci underlying sugar- and salt tolerance in halotolerant yeasts can be utilized to improve the growth and fermentation performance characteristics of laboratory and industrial strains when grown under stressfull conditions. This know-how could potentially alleviate problems caused by extansive soil salinization in plant crops and in agriculture in general (Dakal et al. 2014 and references therein).
Concluding remarks
Halotolerant and osmotolerant yeasts constitute an important group of microorganisms present worldwide. During the last two decades, important advances have been made regarding their identification, characterization, and development of industrial applications. We still need to improve and broaden our knowledge of these yeasts to better understand how they can adapt to the extreme conditions in their natural habitats and to use this information for bioprocesses. As discussed in this review some of these strains have been studied for decades, and now we have some insights into general common strategies that these fungi exploit for halotolerance and osmotolerance. We have also observed differences in several pathways, proteins and enzymes involved when compared among them and with S. cerevisiae, but many intriguing features need to be confirmed and studied in more detail. The omic-based analyses carried out during the last years and many others that will become available and implemented in next forthcoming years, along with molecular tools for which interesting strains remain elusive will increase our knowledge of these halophilic yeasts and their adaptive biology.
References
Ali SS, Al-Tohamy R, Xie R, El-Sheekh MM, Sun J (2020) Construction of a new lipase- and xylanase-producing oleaginous yeast consortium capable of reactive azo dye degradation and detoxification. Bioresour Technol 313:123631. https://doi.org/10.1016/j.biortech.2020.123631
Almagro A, Prista C, Castro S, Quintas C, Madeira-Lopes A, Ramos J, Loureiro-Dias MC (2000) Effects of salts on Debaryomyces hansenii and Saccharomyces cerevisiae under stress conditions. Int J Food Microbiol 56:191–197. https://doi.org/10.1016/s0168-1605(00)00220-8
Al-Tohamy R, Sun J, Fareed MF, Kenawy ER, Ali SS (2020) Ecofriendly biodegradation of Reactive Black 5 by newly isolated Sterigmatomyces halophilus SSA1575, valued for textile azo dye wastewater processing and detoxification. Sci Rep 10:12370. https://doi.org/10.1038/s41598-020-69304-4
Anastas P, Eghbali N (2010) Green chemistry: principles and practice. Chem Soc Rev 39:301–312. https://doi.org/10.1039/b918763b
Anderson NG (2012) Solvent selection. In: Anderson NG (ed) Practical processes & development—a guide for organic chemists, 2nd edn. Academic Press, Oxford, pp 121–168
Andreu C, Del Olmo M (2018) Biotransformation using halotolerant yeast in seawater: a sustainable strategy to produce R-(-)-phenylacetylcarbinol. Appl Microbiol Biotechnol 102:4717–4727. https://doi.org/10.1007/s00253-018-8945-1
Andreu C, del Olmo M (2019) Improved biocatalytic activity of the Debaryomyces species in seawater. ChemCatChem 11:3085. https://doi.org/10.1002/cctc.201900558
Andreu C, Del Olmo M (2020) Whole-cell biocatalysis in seawater: new halotolerant yeast strains for the regio- and stereoselectivity reduction of 1-phenylpropane-1,2-dione in saline-rich media. ChemBioChem 21:1621–1628. https://doi.org/10.1002/cbic.202000023
Bansal PK, Mondal AK (2000) Isolation and sequence of the HOG1 homologue from Debaryomyces hansenii by complementation of the hog1Delta strain of Saccharomyces cerevisiae. Yeast 16(1):81–88. https://doi.org/10.1002/(SICI)1097-0061(20000115)16:1%3c81::AID-YEA510%3e3.0.CO;2-I
Bizzarri M, Cassanelli S, Pryszcz LP, Gawor J, Gromadka R, Solieri L (2018) Draft genome sequences of the highly halotolerant strain Zygosaccharomyces rouxii ATCC 42981 and the novel allodiploid strain Zygosaccharomyces sapae ATB301T obtained using the MinION platform. Microbiol Resour Announc 7:e00874-e918. https://doi.org/10.1128/MRA.00874-18
Bizzarri M, Cassanelli S, Dušková M, Sychrová H, Solieri L (2019) A set of plasmids carrying antibiotic resistance markers and Cre recombinase for genetic engineering of nonconventional yeast Zygosaccharomyces rouxii. Yeast 36:711–722. https://doi.org/10.1002/yea.3438
Breuer U, Harms H (2006) Debaryomyces hansenii—an extremophilic yeast with biotechnological potential. Yeast 23:415–437. https://doi.org/10.1002/yea.1374
Butinar L, Santos S, Spencer-Martins I, Oren A, Gunde-Cimerman N (2005) Yeast diversity in hypersaline habitats. FEMS Microbiol Lett 244:229–234. https://doi.org/10.1016/j.femsle.2005.01.043
Capusoni C, Arioli S, Donzella S, Guidi B, Serra I, Compagno C (2019) Hyper-osmotic stress elicits membrane depolarization and decreased permeability in halotolerant marine Debaryomyces hansenii strains and in Saccharomyces cerevisiae. Front Microbiol 10:64. https://doi.org/10.3389/fmicb.2019.00064
Chung D, Kim H, Choi HS (2019) Fungi in salterns. J Microbiol 57:717–724. https://doi.org/10.1007/s12275-019-9195-3
da Silva S, Calado S, Lucas C, Aguiar C (2008) Unusual properties of the halotolerant yeast Candida nodaensis Killer toxin, CnKT. Microbiol Res 163:243–251. https://doi.org/10.1016/j.micres.2007.04.002
Dakal TC, Solieri L, Giudici P (2014) Adaptive response and tolerance to sugar and salt stress in the food yeast Zygosaccharomyces rouxii. Int J Food Microbiol 185:140–157. https://doi.org/10.1016/j.ijfoodmicro.2014.05.015
Demirci H, Kurt-Gur G, Ordu E (2021) Microbiota profiling and screening of the lipase active halotolerant yeasts of the olive brine. World J Microbiol Biotechnol 37:23. https://doi.org/10.1007/s11274-020-02976-2
Domínguez de María P (2013) On the use of seawater as reaction media for large-scale applications in biorefineries. ChemCatChem 5:1643–1648. https://doi.org/10.1002/cctc.201200877
Dujon B, Sherman D, Fischer G et al (2004) Genome evolution in yeasts. Nature 430:35–44. https://doi.org/10.1038/nature02579
Génolevures Consortium, Souciet JL, Dujon B, Gaillardin C et al (2009) Comparative genomics of protoploid Saccharomycetaceae. Genome Res 19:1696–1709. https://doi.org/10.1101/gr.091546.109
Gong Y, Ding P, Xu MJ, Zhang CM, Xing K, Qin S (2021) Biodegradation of phenol by a halotolerant versatile yeast Candida tropicalis SDP-1 in wastewater and soil under high salinity conditions. J Environ Manag 289:112525. https://doi.org/10.1016/j.jenvman.2021.112525
Gordon JL, Wolfe KH (2008) Recent allopolyploid origin of Zygosaccharomyces rouxii strain ATCC 42981. Yeast 25:449–456. https://doi.org/10.1002/yea.1598
Gori K, Mortensen HD, Arneborg N, Jespersen L (2005) Expression of the GPD1 and GPP2 orthologues and glycerol retention during growth of Debaryomyces hansenii at high NaCl concentrations. Yeast 22:1213–1222. https://doi.org/10.1002/yea.1306
Gori K, Hébraud M, Chambon C, Mortensen HD, Arneborg N, Jespersen L (2007) Proteomic changes in Debaryomyces hansenii upon exposure to NaCl stress. FEMS Yeast Res 7:293–303. https://doi.org/10.1111/j.1567-1364.2006.00155.x
Gostinčar C, Lenassi M, Gunde-Cimerman N, Plemenitaš A (2011) Fungal adaptation to extremely high salt concentrations. In: Laskin AI, Sariaslani S, Gadd GM (eds) Advances in applied microbiology, vol 77. Elsevier, Amsterdam, pp 71–96
Gostinčar C, Sun X, Zajc J, Fang C, Hou Y, Luo Y, Gunde-Cimerman N, Song Z (2019) Population genomics of an obligately halophilic Basidiomycete Wallemia ichthyophaga. Front Microbiol 10:2019. https://doi.org/10.3389/fmicb.2019.02019
Gostinčar C, Stajich JE, Kejžar A, Sinha S, Nislow C, Lenassi M, Gunde-Cimerman N (2021) Seven years at high salinity—experimental evolution of the extremely halotolerant black yeast Hortaea werneckii. J Fungi 7:723. https://doi.org/10.3390/jof7090723
Grande PM, Bergs C, Domínguez de María P (2012) Chemo-enzymatic conversion of glucose into 5-hydroxymethylfurfural in seawater. Chemsuschem 5:1203–1206. https://doi.org/10.1002/cssc.201200065
Gunde-Cimerman N, Ramos J, Plemenitas A (2009) Halotolerant and halophilic fungi. Mycol Res 113:1231–1241. https://doi.org/10.1016/j.mycres.2009.09.002
Hadibarata T, Khudhair AB, Kristanti RA, Kamyab H (2017) Biodegradation of pyrene by Candida sp. S1 under high salinity conditions. Bioprocess Biosyst Eng 40:1411–1418. https://doi.org/10.1007/s00449-017-1798-7
Hernáiz MJ, Alcántara AR, García JI, Sinisterra JV (2010) Applied biotransformations in green solvents. Chem Eur J 16:9422–9437. https://doi.org/10.1002/chem.201000798
Herrera R, Salazar A, Ramos-Moreno L, Ruiz-Roldan C, Ramos J (2017) Vacuolar control of subcellular cation distribution is a key parameter in the adaptation of Debaryomyces hansenii to high salt concentrations. Fungal Genet Biol 100:52–60. https://doi.org/10.1016/j.fgb.2017.02.002
Iwaki T, Tamai Y, Watanabe Y (1999) Two putative MAP kinase genes, ZrHOG1 and ZrHOG2, cloned from the salt-tolerant yeast Zygosaccharomyces rouxii are functionally homologous to the Saccharomyces cerevisiae HOG1 gene. Microbiology (Reading) 145:241–248. https://doi.org/10.1099/13500872-145-1-241
Iwaki T, Kurono S, Yokose Y, Kubota K, Tamai Y, Watanabe Y (2001) Cloning of glycerol-3-phosphate dehydrogenase genes (ZrGPD1 and ZrGPD2) and glycerol dehydrogenase genes (ZrGCY1 and ZrGCY2) from the salt-tolerant yeast Zygosaccharomyces rouxii. Yeast 18:737–744. https://doi.org/10.1002/yea.722
Jančič S, Frisvad JC, Kocev D, Gostinčar C, Džeroski S, Gunde-Cimerman N (2016) Production of secondary metabolites in extreme environments: food- and airborne Wallemia spp. produce toxic metabolites at hypersaline conditions. PLoS ONE 11:e0169116. https://doi.org/10.1371/journal.pone.0169116
Jiang Y, Yang K, Wang H, Shang Y, Yang X (2015) Characteristics of phenol degradation in saline conditions of a halophilic strain JS3 isolated from industrial activated sludge. Mar Pollut Bull 99:230–234. https://doi.org/10.1016/j.marpolbul.2015.07.021
Jiang Y, Shang Y, Yang K, Wang H (2016) Phenol degradation by halophilic fungal isolate JS4 and evaluation of its tolerance of heavy metals. Appl Microbiol Biotechnol 100:1883–1890. https://doi.org/10.1007/s00253-015-7180-2
Jiang Y, Yang K, Deng T, Ji B, Shang Y, Wang H (2018) Immobilization of halophilic yeast for effective removal of phenol in hypersaline conditions. Water Sci Technol 77:706–713. https://doi.org/10.2166/wst.2017.576
Jones EBG, Suetrong S, Sakayaroj J et al (2015) Classification of marine Ascomycota, Basidiomycota, Blastocladiomycota and Chytridiomycota. Fungal Divers 73:1–72. https://doi.org/10.1007/s13225-015-0339-4
Kamyabi A, Nouri H, Moghimi H (2018) Characterization of pyrene degradation and metabolite identification by Basidioascus persicus and mineralization enhancement with bacterial-yeast co-culture. Ecotoxicol Environ Saf 15:471–477. https://doi.org/10.1016/j.ecoenv.2018.07.098
Karatay SE, Demiray E, Dönmez G (2019) Bioethanol production by newly isolated halotolerant Kluyveromyces marxianus strains. Environ Prog Sustain Energy 38:542–547. https://doi.org/10.1002/ep.12935
Kejžar A, Grötli M, Tamás MJ, Plemenitaš A, Lenassi M (2015a) HwHog1 kinase activity is crucial for survival of Hortaea werneckii in extremely hyperosmolar environments. Fungal Genet Biol 74:45–58. https://doi.org/10.1016/j.fgb.2014.11.004
Kejžar A, Cibic M, Grøtli M, Plemenitaš A, Lenassi M (2015b) The unique characteristics of HOG pathway MAPKs in the extremely halotolerant Hortaea werneckii. FEMS Microbiol Lett 362:fnv046. https://doi.org/10.1093/femsle/fnv046
Kinclová O, Potier S, Sychrová H (2001) The Zygosaccharomyces rouxii strain CBS732 contains only one copy of the HOG1 and the SOD2 genes. J Biotechnol 88:151–158. https://doi.org/10.1016/s0168-1656(01)00274-7
Kogej T, Stein M, Volkmann M, Gorbushina AA, Galinski EA, Gunde-Cimerman N (2007) Osmotic adaptation of the halophilic fungus Hortaea werneckii: role of osmolytes and melanization. Microbiology (Reading) 153:4261–4273. https://doi.org/10.1099/mic.0.2007/010751-0
Konte T, Plemenitas A (2013) The HOG signal transduction pathway in the halophilic fungus Wallemia ichthyophaga: identification and characterisation of MAP kinases WiHog1A and WiHog1B. Extremophiles 17:623–636. https://doi.org/10.1007/s00792-013-0546-4
Konte T, Terpitz U, Plemenitaš A (2016) Reconstruction of the high-osmolarity glycerol (HOG) signaling pathway from the halophilic fungus Wallemia ichthyophaga in Saccharomyces cerevisiae. Front Microbio 7:901. https://doi.org/10.3389/fmicb.2016.00901
Kralj Kuncic M, Kogej T, Drobne D, Gunde-Cimerman N (2010) Morphological response of the halophilic fungal genus Wallemia to high salinity. Appl Environ Microbiol 76:329–337. https://doi.org/10.1128/AEM.02318-09
Lenassi M, Plemenitas A (2007) Novel group VII histidine kinase HwHhk7B from the halophilic fungi Hortaea werneckii has a putative role in osmosensing. Curr Genet 51:393–405. https://doi.org/10.1007/s00294-007-0131-4
Lenassi M, Zajc J, Gostinčar C, Gorjan A, Gunde-Cimerman N, Plemenitaš A (2011) Adaptation of the glycerol-3-phosphate dehydrogenase Gpd1 to high salinities in the extremely halotolerant Hortaea werneckii and halophilic Wallemia ichthyophaga. Fungal Biol 115:959–970. https://doi.org/10.1016/j.funbio.2011.04.001
Lenassi M, Gostinčar C, Jackman S, Turk M, Sadowski I, Nislow C, Jones S, Birol I, Cimerman NG, Plemenitaš A (2013) Whole genome duplication and enrichment of metal cation transporters revealed by de novo genome sequencing of extremely halotolerant black yeast Hortaea werneckii. PLoS ONE 8:e71328. https://doi.org/10.1371/journal.pone.0071328
Martínez-Ávila L, Peidro-Guzmán H, Pérez-Llano Y et al (2021) Tracking gene expression, metabolic profiles, and biochemical analysis in the halotolerant basidiomycetous yeast Rhodotorula mucilaginosa EXF-1630 during benzo[a]pyrene and phenanthrene biodegradation under hypersaline conditions. Environ Pollut 271:116358. https://doi.org/10.1016/j.envpol.2020.116358
Masuda K, Guo X-F, Uryu N, Hagiwara T, Watabe S (2008) Isolation of marine yeasts collected from the Pacific Ocean showing a high production of γ-aminobutyric acid. Biosci Biotechnol Biochem 72:3265–3272. https://doi.org/10.1271/bbb.80544
Minhas AP, Biswas D (2019) Development of an efficient transformation system for halotolerant yeast Debaryomyces hansenii CBS767. Bio Protoc 9:e3352. https://doi.org/10.21769/BioProtoc.3352
Minhas A, Biswas D, Mondal AK (2009) Development of host and vector for high-efficiency transformation and gene disruption in Debaryomyces hansenii. FEMS Yeast Res 9:95–102. https://doi.org/10.1111/j.1567-1364.2008.00457.x
Mitchison-Field LMY, Vargas-Muñiz JM, Stormo BM, Vogt EJD, Van Dierdonck S, Pelletier JF, Ehrlich C, Lew DJ, Field CM, Gladfelter AS (2019) Unconventional cell division cycles from marine-derived yeasts. Curr Biol 29:3439-3456.e5. https://doi.org/10.1016/j.cub.2019.08.050
Mogi R, Watanabe J (2020) Identification of SFL1 as a positive regulator for flor formation in Zygosaccharomyces rouxii. Biosci Biotechnol Biochem 84:1291–1298. https://doi.org/10.1080/09168451.2020.1732187
Mohite P, Kumar AR, Zinjarde S (2017) Relationship between salt tolerance and nanoparticle synthesis by Williopsis saturnus NCIM 3298. World J Microbiol Biotechnol 33:163. https://doi.org/10.1007/s11274-017-2329-z
Ni Y, Holtmann D, Hollmann F (2014) How green is biocatalysis? To calculate is to know. ChemCatChem 6:930–943. https://doi.org/10.1002/cctc.201300976
Nikolaivits E, Agrafiotis A, Baira E, Le Goff G, Tsafantakis N, Chavanich SA, Benayahu Y, Ouazzani J, Fokialakis N, Topakas E (2020) Degradation mechanism of 2,4-dichlorophenol by fungi isolated from marine invertebrates. Int J Mol Sci 21:3317. https://doi.org/10.3390/ijms21093317
Okai M, Betsuno A, Shirao A, Obara N, Suzuki K, Takei T, Takashio M, Ishida UN (2016) Citeromyces matritensis M37 is a salt-tolerant yeast that produces ethanol from salted algae. Can J Microbiol 63:20–26. https://doi.org/10.1139/cjm-2016-0259
Perea-Sanz L, Peris D, Belloch C, Flores D (2019) Debaryomyces hansenii metabolism of sulfur amino acids as precursors of volatile sulfur compounds of interest in meat products. Agric Food Chem 67(33):9335–9343. https://doi.org/10.1021/acs.jafc.9b03361
Petersen KM, Jespersen L (2004) Genetic diversity of the species Debaryomyces hansenii and the use of chromosome polymorphism for typing of strains isolated from surface-ripened cheeses. J Appl Microbiol 97:205–213. https://doi.org/10.1111/j.1365-2672.2004.02293.x
Pitt JI, Hocking AD (2009) Fungi and food spoilage, 3rd edn. Springer, Boston
Plemenitas A, Vaupotic T, Lenassi M, Kogej T, Gunde-Cimerman N (2008) Adaptation of extremely halotolerant black yeast Hortaea werneckii to increased osmolarity: a molecular perspective at a glance. Stud Mycol 61:67–75. https://doi.org/10.3114/sim.2008.61.06
Plemenitaš A, Lenassi M, Konte T, Kejžar A, Zajc J, Gostinčar C, Gunde-Cimerman N (2014) Adaptation to high salt concentrations in halotolerant/halophilic fungi: a molecular perspective. Front Microbiol 5:199. https://doi.org/10.3389/fmicb.2014.00199
Posas F, Chambers JR, Heyman JA, Hoeffler JP, de Nadal E, Ariño J (2000) The transcriptional response of yeast to saline stress. J Biol Chem 275:17249–17255. https://doi.org/10.1074/jbc.M910016199
Prista C, Michán C, Miranda IM, Ramos J (2016) The halotolerant Debaryomyces hansenii, the Cinderella of non-conventional yeasts. Yeast 33:523–533. https://doi.org/10.1002/yea.3177
Qi W, Fan ZC, Wang CL, Hou LH, Liu JF, Cao XH (2014a) Non-targeted metabolomic reveals the effect of salt stress on global metabolite of halotolerant yeast Candida versatilis and principal component analysis. J Ind Microbiol Biotechnol 41:1553–1562. https://doi.org/10.1007/s10295-014-1475-7
Qi W, Hou LH, Guo HL, Wang CL, Fan ZC, Liu JF, Cao XH (2014b) Effect of salt-tolerant yeast of Candida versatilis and Zygosaccharomyces rouxii on the production of biogenic amines during soy sauce fermentation. J Sci Food Agric 94:1537–1542. https://doi.org/10.1002/jsfa.6454
Ramos-Moreno L, Ramos J, Michán C (2019) Overlapping responses between salt and oxidative stress in Debaryomyces hansenii. World J Microbiol Biotechnol 35:170. https://doi.org/10.1007/s11274-019-2753-3
Ricaurte ML, Govind NS (1999) Construction of plasmid vectors and transformation of the marine yeast Debaryomyces hansenii. Mar Biotechnol (NY) 1:15–19. https://doi.org/10.1007/pl00011745
Romeo O, Marchetta A, Giosa D, Giuffrè L, Urzì C, De Leo F (2020) Whole genome sequencing and comparative genome analysis of the halotolerant deep sea black yeast Hortaea werneckii. Life (Basel) 10:229. https://doi.org/10.3390/life10100229
Ruginescu R, Gomoiu I, Popescu O, Cojoc R, Neagu S, Lucaci I, Batrinescu-Moteau C, Enache M (2020) Bioprospecting for novel halophilic and halotolerant sources of hydrolytic enzymes in brackish, saline and hypersaline lakes of Romania. Microorganisms 8:1903. https://doi.org/10.3390/microorganisms8121903
Sánchez NS, Calahorra M, González J, Defosse T, Papon N, Peña A, Coria R (2020) Contribution of the mitogen-activated protein kinase Hog1 to the halotolerance of the marine yeast Debaryomyces hansenii. Curr Genet 66:1135–1153. https://doi.org/10.1007/s00294-020-01099-3
Sato A, Matsushima K, Oshima K, Hattori M, Koyama Y (2017) Draft genome sequencing of the highly halotolerant and allopolyploid yeast Zygosaccharomyces rouxii NBRC 1876. Genome Announc 5:e01610-e1616. https://doi.org/10.1128/genomeA.01610-16
Scapini T, Dalastra C, Camargo AF, Kubeneck S, Modkovski TA, Júnior SLA, Treichel H (2021) Seawater-based biorefineries: a strategy to reduce the water footprint in the conversion of lignocellulosic biomass. Bioresour Technol 14:126325. https://doi.org/10.1016/j.biortech.2021.126325
Serra I, Guidi B, Burgaud G, Contente ML, Ferraboschi P, Pinto A, Compagno C, Molinari F, Romano D (2016) Seawater-based biocatalytic strategy: stereoselective reductions of ketones with marine yeasts. ChemCatChem 8:3254. https://doi.org/10.1002/cctc.201600947
Serra I, Capusoni C, Molinari F, Musso L, Pellegrino L, Compagno C (2019) Marine microorganisms for biocatalysis: selective hydrolysis of nitriles with a salt-resistant strain of Meyerozyma guilliermondii. Mar Biotechnol (NY) 21:229–239. https://doi.org/10.1007/s10126-019-09875-0
Sharma P, Mondal AK (2005) Evidence that C-terminal non-kinase domain of Pbs2p has a role in high osmolarity-induced nuclear localization of Hog1p. Biochem Biophys Res Commun 328:906–913. https://doi.org/10.1016/j.bbrc.2005.01.039
Sinha S, Flibotte S, Neira M, Formby S, Plemenitaš A, Cimerman NG, Lenassi M, Gostinčar C, Stajich JE, Nislow C (2017) Insight into the recent genome duplication of the halophilic yeast Hortaea werneckii: combining an improved genome with gene expression and chromatin structure. G3 (Bethesda) 7:2015–2022. https://doi.org/10.1534/g3.117.040691
Solieri L (2021) The revenge of Zygosaccharomyces yeasts in food biotechnology and applied microbiology. World J Microbiol Biotechnol 37:96. https://doi.org/10.1007/s11274-021-03066-7
Solieri L, Cassanelli S, Croce MA, Giudici P (2008) Genome size and ploidy level: new insights for elucidating relationships in Zygosaccharomyces species. Fungal Genet Biol 45:1582–1590. https://doi.org/10.1016/j.fgb.2008.10.001
Stratford M, Steels H, Novodvorska M, Archer DB, Avery SV (2019) Extreme osmotolerance and halotolerance in food-relevant yeasts and the role of glycerol-dependent cell individuality. Front Microbiol 9:3238. https://doi.org/10.3389/fmicb.2018.03238
Strucko T, Andersen NL, Mahler MR, Martínez JL, Mortensen UH (2021) A CRISPR/Cas9 method facilitates efficient oligo-mediated gene editing in Debaryomyces hansenii. Synth Biol 6:ysab031. https://doi.org/10.1093/synbio/ysab031
Tang W, Zhou B, Xing K, Tan L (2020) Co-enhanced activated sludge system by static magnetic field and two halotolerant yeasts for azo dye treatment. Water Environ Res 92:2095–2104. https://doi.org/10.1002/wer.1375
Trincone A (2010) Potential biocatalysts originating from sea environments. J Mol Catal B 66:241–256. https://doi.org/10.1016/j.molcatb.2010.06.004
Turk M, Plemenitas A (2002) The HOG pathway in the halophilic black yeast Hortaea werneckii: isolation of the HOG1 homolog gene and activation of HwHog1p. FEMS Microbiol Lett 216:193–199. https://doi.org/10.1111/j.1574-6968.2002.tb11435.x
Wang D, Zhang M, Huang J, Zhou R, Jin Y, Wu C (2020a) Zygosaccharomyces rouxii combats salt stress by maintaining cell membrane structure and functionality. J Microbiol Biotechnol 30:62–70. https://doi.org/10.4014/jmb.1904.04006
Wang X, Wang Y, Ning S, Shi S, Tan L (2020b) Improving azo dye decolorization performance and halotolerance of Pichia occidentalis A2 by static magnetic field and possible mechanisms through comparative transcriptome analysis. Front Microbiol 11:712. https://doi.org/10.3389/fmicb.2020.00712
Watanabe J, Uehara K, Mogi Y, Suzuki K, Watanabe T, Yamazaki T (2010) Improved transformation of the halo-tolerant yeast Zygosaccharomyces rouxii by electroporation. Biosci Biotechnol Biochem 74:1092–1094. https://doi.org/10.1271/bbb.90865
Wells A, Meyer H-P (2014) Biocatalysis as a strategic green technology for the chemical industry. ChemCatChem 6:918–920. https://doi.org/10.1002/cctc.201402065
Xie D, Miller E, Sharpe P, Jackson E, Zhu Q (2017) Omega-3 production by fermentation of Yarrowia lipolytica: from fed-batch to continuous. Biotechnol Bioeng 114:798–812. https://doi.org/10.1002/bit.26216
Zajc J, Liu Y, Dai W, Yang Z, Hu J, Gostinčar C, Gunde-Cimerman N (2013) Genome and transcriptome sequencing of the halophilic fungus Wallemia ichthyophaga: haloadaptations present and absent. BMC Genomics 14:617. https://doi.org/10.1186/1471-2164-14-617
Zajc J, Džeroski S, Kocev D, Oren A, Sonjak S, Tkavc R, Gunde-Cimerman N (2014a) Chaophilic or chaotolerant fungi: a new category of extremophiles? Front Microbiol 5:708. https://doi.org/10.3389/fmicb.2014.00708
Zajc J, Kogej T, Galinski EA, Ramos J, Gunde-Cimerman N (2014b) Osmoadaptation strategy of the most halophilic fungus, Wallemia ichthyophaga, growing optimally at salinities above 15% NaCl. Appl Environ Microbiol 80:247–256. https://doi.org/10.1128/AEM.02702-13
Zaky AS, Tucker GA, Daw ZY, Du C (2014) Marine yeast isolation and industrial application. FEMS Yeast Res 14:813–825. https://doi.org/10.1111/1567-1364.12158
Zaky AS, Greetham D, Tucker GA, Du C (2018) The establishment of a marine focused biorefinery for bioethanol production using seawater and a novel marine yeast strain. Sci Rep 8:12127. https://doi.org/10.1038/s41598-018-30660-x
Zaky AS, French CE, Tucker GA, Du C (2020) Improving the productivity of bioethanol production using marine yeast and seawater-based media. Biomass Bioenergy 139:105615. https://doi.org/10.1016/j.biombioe.2020.105615
Zalar P, Sybren de Hoog G, Schroers HJ, Frank JM, Gunde-Cimerman N (2005) Taxonomy and phylogeny of the xerophilic genus Wallemia (Wallemiomycetes and Wallemiales, cl. et ord. nov.). Antonie Van Leeuwenhoek 87:311–328. https://doi.org/10.1007/s10482-004-6783-x
Zalar P, Zupančič J, Gostinčar C, Zajc J, de Hoog GS, De Leo F, Azua-Bustos A, Gunde-Cimerman N (2019) The extremely halotolerant black yeast Hortaea werneckii—a model for intraspecific hybridization in clonal fungi. IMA Fungus 10:10. https://doi.org/10.1186/s43008-019-0007-5
Zambelli P, Serra I, Fernandez-Arrojo L, Plou FJ, Tamborini L, Conti P, Contente ML, Molinari F, Romano D (2015) Sweet-and-salty biocatalysis: fructooligosaccharides production using Cladosporium cladosporioides in seawater. Process Biochem 50:1086–1090. https://doi.org/10.1016/j.procbio.2015.04.006
Zarnowski R, Sanchez H, Andreu C, Andes D, Del Olmo ML (2021) Formation and characterization of biofilms formed by salt-tolerant yeast strains in seawater-based growth medium. Appl Microbiol Biotechnol 105:2411–2426. https://doi.org/10.1007/s00253-021-11132-1
Zheng C, Li Z, Yang H, Zhang T, Niu H, Liu D, Wang J, Ying H (2019) Computation-aided rational design of a halophilic choline kinase for cytidine diphosphate choline production in high-salt condition. J Biotechnol 290:59–66. https://doi.org/10.1016/j.jbiotec.2018.11.008
Acknowledgements
This work has been supported by Grants UV-INV-AE19-1199043 and UV-INV-AE-1560151 from the Universitat de València. We thank Ms. Hanna Anhalt for linguisitic comments on this manuscript.
Author information
Authors and Affiliations
Contributions
All the authors contributed to the writing, reviewing and editing of the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest, financial or otherwise.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Andreu, C., Zarnowski, R. & del Olmo, M. Recent developments in the biology and biotechnological applications of halotolerant yeasts. World J Microbiol Biotechnol 38, 27 (2022). https://doi.org/10.1007/s11274-021-03213-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-021-03213-0