Abstract
Yeast whole cells have been widely used in modern biotechnology as biocatalysts to generate numerous compounds of industrial, chemical, and pharmaceutical importance. Since many of the biocatalysis-utilizing manufactures have become more concerned about environmental issues, seawater is now considered a sustainable alternative to freshwater for biocatalytic processes. This approach plausibly commenced new research initiatives into exploration of salt-tolerant yeast strains. Recently, there has also been a growing interest in possible applications of microbial biofilms in the field of biocatalysis. In these complex communities, cells demonstrate higher resistance to adverse environmental conditions due to their embedment in an extracellular matrix, in which physical, chemical, and physiological gradients exist. Considering these two topics, seawater and biofilms, in this work, we characterized biofilm formation in seawater-based growth media by several salt-tolerant yeast strains with previously demonstrated biocatalytic capacities. The tested strains formed both air-liquid-like biofilms and biofilms on silicone surfaces, with Debaryomyces fabryi, Schwanniomyces etchellsii, Schwanniomyces polymorphus, and Kluyveromyces marxianus showing the highest biofilm formation. The extracted biofilm extracellular matrices mostly consisted of carbohydrates and proteins. The latter group was primarily represented by enzymes involved in metabolic processes, particularly the biosynthetic ones, and in the response to stimuli. Specific features were also found in the carbohydrate composition of the extracellular matrix, which were dependent both on the yeast isolate and the nature of formed biofilms. Overall, our findings presented herein provide a unique data resource for further development and optimization of biocatalytic processes and applications employing seawater and halotolerant yeast biofilms.
Key points
• Ability for biofilm formation of some yeast-halotolerant strains in seawater medium
• ECM composition dependent on strain and biofilm-forming surface
• Metabolic enzymes in the ECM with potential applications for biocatalysis
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Virtually, all living microorganisms have the capacity to form biofilms that represent communities, in which living cells embedded in an extracellular matrix (ECM) are attached to various biotic and abiotic surfaces (Flemming and Wingender 2010; Strieth et al. 2018). The ECM of biofilms consists of self-produced secreted polymeric substances. Within the biofilm ECM, chemical gradients are generated. These gradients determine the existence of heterogeneous populations with distinct phenotypes and result in manifestation of diverse metabolic pathways, stress responses, and other biological activities (Stewart and Franklin 2008). Biofilm formation is complex and is influenced by multiple biological, chemical, and physical factors. In fact, numerous physiological states can be distinguished during biofilm formation (Halan et al. 2012). Initial interactions of planktonic cells with a surface depend on the availability of nutrients. During proliferation and maturation phases, cells then adhere to it and begin to reproduce and to make the ECM that embeds the cells attached to the surface and to each other. Such a convoluted development leads to the formation of a complex three-dimensional structure with water channels and pores through which nutrients and waste can diffuse. Finally, a pseudo-steady state is established, in which dispersion or release of biofilm biomass particles or individual cells allows for the re-initiation of the cycle.
Cellular organization in biofilms provides enormous advantages, such as colonization of host tissues; expression of virulence characteristics; metabolic cooperation; efficient nutrient capture; cellular communication; and increased tolerance to chemical, physical, and biological stress conditions (Castrillón et al. 2013). These properties make biofilms potentially harmful to human health; in fact, multiple biofilm-associated bacterial and fungal organisms have been shown to be responsible for the majority of microbial infections in humans (Inigo and Del Pozo 2018). While prokaryotic biofilms seem to be more extensively studied (Bales et al. 2013; Cheng et al. 2019; Colvin et al. 2011, 2012), a substantial amount of scientific data on fungal biofilms exists. Research studies have been carried out in the eukaryotic biology model organism Saccharomyces cerevisiae (Andersen et al. 2014), as well as in a few human pathogenic organisms, such as Aspergillus fumigatus, Cryptococcus neoformans, and various Candida species (Dominguez et al. 2018, 2019; Le Mauff et al. 2019; Martinez and Casadevall 2015; Zarnowski et al. 2014). With bacterial biofilms, the secreted high molecular weight exopolysaccharides constitute the scaffold for the formation of this structure, to which other carbohydrates adhere, as well as proteins, lipids, and nucleic acids present in smaller quantities (Hall-Stoodley et al. 2004). The data available for fungal biofilms, obtained in a great extent from Candida spp., revealed the presence of a high content of proteins and polysaccharides with the ECM composition similar to that of the cell wall of the microorganism that forms it, indicating that broad diversity may exist (Mitchell et al. 2016).
The abovementioned features of biofilms can hinder some industrial processes, while also be beneficial and valuable for others, like those related to the biotechnological production of organic compounds and the modification of several foods (Berlanga and Guerrero 2016; Muffler et al. 2014; Strieth et al. 2018). For decades, biofilms have been applied to bioremediation, particularly in the treatment of wastewater and off-gas (Gross et al. 2007).
Biofilms have also been applied to biocatalytic processes, in which natural enzymes or whole cells are used to perform chemical transformations of organic compounds. They have been particularly used for fermentative production of bulk chemicals, such as acetic acid, ethanol, and butanol (Demirci et al. 1997; Ho et al. 1997; Qureshi et al. 2004), as well as in the production of organic acids (citric, fumaric, and succinic acids), enzymes (lignin peroxidase, manganese peroxidase, cellulase), bioactive compounds (bacteriocins such as pediocin or nisin and antibiotics such as cephalosporin C), and fine chemicals (Gross et al. 2007, 2013; Halan et al. 2012, 2014; Muffler et al. 2014; Romero et al. 2018; Willrodt et al. 2017).
Over the last several years, we have demonstrated the usefulness of some Saccharomyces and non-Saccharomyces yeast strains for the stereoselective production of several compounds of chemical and pharmaceutical interest (Andreu and del Olmo 2014, 2018, 2019, 2020; Andreu et al. 2016). A considerable problem faced when whole cells are used as biocatalysts in industrial processes is related to a massive consumption of water due to the low solubility of the hydrophobic substrates. This concern can be addressed by the utilization of seawater as a sustainable alternative to freshwater in chemical reactors (Andreu and del Olmo 2018, 2019, 2020). Finding halotolerant microbial species that could carry out biocatalytic processes in this medium is another logical step in the implementation of this strategy at an industrial scale. In this work, we analyzed the capability of several salt-tolerant yeast strains useful as biocatalysts for biofilm production in seawater medium. The strains considered have been Torulaspora delbrueckii, Kluyveromyces marxianus, Debaryomyces hansenii, Debaryomyces fabryi, Schwanniomyces etchellsii, Schwanniomyces polymorphus, Meyerozyma guilliermondii, Pichia glucozyma, Pichia fermentans, Pichia jadinii, and Saccharomyces cerevisiae Ethanol Red. We also characterized in-depth the composition of biofilm ECMs in those biofilm-forming halotolerant yeast species.
Materials and methods
Yeast strains and growth conditions for biofilm production
The yeast strains used in this work are described in Table 1. To determine the halotolerance of yeast strains, growth experiments were carried out in culture plates of YPD-based medium that contained 2% (w/v) agar supplemented with NaCl to a final concentration between 3.5 and 7.5% (w/v).
The seawater used in the experiments came from the El Perelló beach (Mediterranean Sea), València (Spain). It was sterilized by autoclaving under standard conditions. The water salinity for this area was around 4% (w/v), according to the determination made by weighing the solid residue after lyophilization. The sodium content, analyzed by flame emission spectroscopy (589.0 nm, Thermo Fisher Scientific (Waltham, USA) model iCE 3000 Series), was 12.46 ± 0.17 g/L, and pH ranged from 7.5 to 8.5, depending on the batch.
For biofilm production, cells inoculated in YPD medium prepared in seawater (SW-YPD) (1% (w/v) yeast extract, 2% (w/v) bacto peptone, 2% (w/v) glucose) were incubated overnight at 30 °C in an orbital shaker (200 rpm). Then, the cultures were diluted in the same medium without any carbon source (SW-YP) to a final OD600 of 0.3. Fifty milliliters of these suspensions was placed in 250-mL glass beakers and incubated at 30 °C for 14 days without shaking. Alternatively, silicone tubes (length of 25 cm; external diameter of 12 mm; internal diameter of 8 mm) were filled with 10 mL of the same solution, closing both ends by joining them with a polypropylene pipette tip, and kept under the same conditions.
Microscopic visualization of biofilms
The coverslip assay was used for in vitro biofilm imaging (Mitchell et al. 2015). Briefly, in vitro biofilms were grown on sterile coverslips (Thermanox, Thermo Fisher Scientific, Waltham, USA) in 12-well polystyrene plates that were each pre-coated with 10 μL of human NaEDTA plasma and allowed to dry at 30 °C. Fungal cell inocula (106 cells/mL) were prepared out of overnight yeast cultures in artificial SW-YPD (1% (w/v) yeast extract, 2% (w/v) bacto peptone, 2% (w/v) glucose, 2% (w/v) NaCl) at 30 °C followed by cell counting with an automated Countess™ II cell counter (Invitrogen, Carlsbad, USA). Yeast inocula were applied to each coverslip at 30 °C for 60 min. The initial inoculum was then removed, and 1 mL of fresh artificial SW-YPD was added to each well. Lastly, the plates were incubated at 30 °C for 48 h. Samples were imaged on a Leo 1530-1 FESEM/EDS/EBSD scanning electron microscope system (Carl Zeiss Microscopy, White Plains, USA).
Determination of the biofilm formation capacity
For each one of the tested strains growing in glass beakers or silicone tubes under the conditions described above, biofilm and planktonic cells were separated. In the case of experiments carried out with glass beakers, the suspension was passed through a sterile stainless steel colander in which the biofilm was trapped, allowing the separation of the two pools. For silicone tube experiments, planktonic cell solutions were initially obtained by emptying the tubes and, after several washes, the biofilm biomass attached on the silicone was scraped off and collected by adding water. In both cases, the material of each fraction was centrifuged at 2500×g for 3 min, washed with water, and lyophilized. Then, the weights of biofilm and planktonic fractions were determined, and the ratio between them was considered the biofilm formation capacity.
Isolation of the biofilm ECM
Biofilm ECM was obtained as described previously (Zarnowski et al. 2014) for Candida species with minor modifications. Briefly, biofilms were separated from planktonic cells as described above, collected by centrifugation, and washed. Then, the biomass was resuspended in 30 mL of water, and the solution was sonicated for 5 min at 20 kHz in a Vibra Cell VCX-500 sonicator (Sonics & Materials Inc., Newtown, USA). Cells were collected by centrifugation, and the matrix (supernatant) was filtered through 0.22-μm PES filtration units (VWR, Radnor, USA). Sodium azide was added to a final concentration of 0.02% (w/v), and the solution was frozen and lyophilized. After resuspension in water and exhaustive dialysis in 50 mM ammonium bicarbonate buffer, the solution was used for further chemical analyses.
Analyses of the composition in carbohydrates of the biofilm ECM
Carbohydrates in biofilm ECMs were analyzed based on the modified procedures reported elsewhere (Zarnowski et al. 2014). Monosugars were converted to alditol acetate derivatives (Blakeney et al. 1982) and then identified and quantified by gas chromatography on a GC-2010 system (Shimadzu, Kyoto, Japan). A Crossbond® 50% cyanopropylmethyl/50% phenylmethyl polysiloxane column was used (15 m × 0.25 mm with 0.25-μm film thickness, RTX-225, Restek, Bellefonte, USA). The GLC (gas liquid chromatography) conditions were as follows: injector at 220 °C, FID detector at 240 °C, and a temperature program of 215 °C for 2 min, then 4 °C min−1 up to 230 °C before holding for 11.25 min, run at constant linear velocity of 33.4 cm/s and split ratio of 50:1.
Analyses of the ECM proteomes
Enzymatic “in liquid” digestion and mass spectrometric analysis were done at the Mass Spectrometry Facility, Biotechnology Center, University of Wisconsin—Madison. Two hundred micrograms of proteins was extracted by precipitation with 15% TCA (trichloroacetic acid)/60% acetone and then incubated at – 20 °C for 30 min. The ECM preparation was centrifuged at 16,000×g for 10 min, and the resulting pellets were washed twice with ice-cold acetone, followed by an ice-cold MeOH wash. Pelleted proteins were resolubilized and denatured in 10 μL of 8 M urea in 100 mM NH4HCO3 for 10 min, then diluted to 60 μL for tryptic digestion with the following reagents: 3 μL of 25 mM DTT, 4.5 μL of acetonitrile, 36.2 μL of 25 mM NH4HCO3, 0.3 μL of 1M Tris-HCl, and 6 μL of 100 ng/μL Trypsin Gold solution in 25 mM NH4HCO3 (Promega, Madison, USA). Digestion was conducted in two stages. First, the digestion was incubated overnight at 37 °C. Then, an additional 4 μL of trypsin solution was added, and the mixture was incubated at 42 °C for an additional 2 h. The reaction was terminated by acidification with 2.5% TFA (trifluoroacetic acid) to a final concentration of 0.3% and then centrifuged at 16,000×g for 10 min. Trypsin-generated peptides were analyzed by nanoLC-MS/MS using the Agilent 1100 nanoflow system (Agilent, Santa Clara, USA) connected to a hybrid linear ion trap-orbitrap mass spectrometer (LTQ-Orbitrap, Thermo Fisher Scientific, Waltham, USA) equipped with a nanoelectrospray ion source. Capillary HPLC was performed using an in-house fabricated column with an integrated electrospray emitter, as described elsewhere (Martin et al. 2000). Sample loading and desalting were achieved using a trapping column in line with the autosampler (Zorbax 300SB-C18, 5 μm, 5 × 0.3 mm, Agilent, Santa Clara, USA). The LTQ-Orbitrap was set to acquire MS/MS spectra in a data-dependent mode as follows: MS survey scans from 300 to 2000 m/z were collected in profile mode with a resolving power of 100,000. MS/MS spectra were collected on the five most abundant signals in each survey scan. Dynamic exclusion was employed to increase the dynamic range and maximize peptide identifications. Raw MS/MS data were searched against a species-select concatenated amino acid sequence database using an in-house MASCOT search engine (Perkins et al. 1999). The data were searched against concatenated D. fabryi and K. marxianus amino acid sequence databases, whereas the proteomes of S. etchellsii and S. polymorphus were searched against the proteome of D. hansenii. These two species were initially classified within the Debaryomyces genus, but recently reassigned to Schwanniomyces (Kurtzman and Suzuki 2010). Identified proteins were further annotated and filtered to 1.5% peptide and 0.1% protein false-discovery-rate with Scaffold Q+ version 4.10.0 (Proteome Software Inc., Portland, USA) using the protein prophet algorithm (Keller et al. 2002). The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE (Perez-Riverol et al. 2019) partner repository with the dataset identifier PXD022284 and 10.6019/PXD022284.
The ECM proteomes of the tested halotolerant yeast species were analyzed functionally using the UniProt Knowledgebase data, which contains reviewed UniProtKB/Swiss-Prot entries (Uniprot Consortium 2019). Functional Gene Ontology (GO) terms were assigned for each protein identified based on the respective halotolerant yeast genome data. Two sets of hierarchies (“biological process” and “molecular function”) were used to prepare the visualizations of relative quantities of biofilm proteins. On the basis of this hierarchical classification scheme, Voronoi treemaps were constructed (Bernhardt et al. 2009). This approach divides screen space according to hierarchy levels in which the main functional categories determine screen sections on the first level, subsidiary categories on the second level, and so forth. Voronoi treemaps were prepared with Paver version 2.1.9 (DECODON Software UG, Greifswald, Germany).
Results
The ability of halotolerant yeast strains for biofilm and ECM production in seawater
The requirement for large quantities of water at the industrial scale makes the use of seawater as a sustainable, freely available solvent, more conveniently. Recently, we have reported the versatility of several halotolerant strains for biocatalytic processes in the presence of seawater (Andreu and del Olmo 2018, 2019, 2020). Some of the tested yeasts such as T. delbrueckii and K. marxianus were capable of growing in YPD supplemented with NaCl up to 5% (w/v), while D. hansenii, D. fabryi, S. etchellsii, S. polymorphus, and M. guilliermondii tolerated even 7.5% NaCl (Andreu and del Olmo 2018, 2020). In this work, we have expanded this growth characteristic analysis to four additional strains of biotechnological interest: P. glucozyma, P. fermentans, P. jadinii, and S. cerevisiae Ethanol Red) (Table 1). Only the latter two tolerated salt concentrations up to 5% (w/v) and were thus also considered for additional analyses in this work (Fig. 1a).
Salt resistance and biofilm-forming capacity in seawater-based medium for the strains considered in this work. Panel a: Growth of several yeast strains on the YPD-derived plates supplemented with NaCl concentrations of 3.5, 5, or 7.5% (w/v), or without salt (YPD). 106 cells from the overnight cultures in YPD were spread on plates, which were then incubated for 48 h at 28 °C. This figure shows the result of a representative experiment. Panel b: Comparison of the 14-day growth appearance of one strain capable of biofilm formation in glass beakers (Schwanniomyces polymorphus) and another in which only planktonic cells are observed (Pichia jadinii). Panel c: Biofilm formation capacity was determined as the ratio between the weight of the dry biomass corresponding to biofilm respect to the weight of the dry planktonic cells after 14-day incubation. The values shown correspond to the average of three experiments
We further characterized the ability of the tested halotolerant strains to form biofilms in seawater-growth media on two types of solid surfaces: in glass beakers and in silicone tubes. These complex microbial community structures appeared mainly as a white veil of cell aggregates forming air-liquid biofilms on the liquid surface as well as to some extent on the glass walls (Fig. 1b). In silicone tubes, the biofilms were observed attached to the silicone surface. Except for P. jadinii and S. cerevisiae Ethanol Red, all the tested yeast strains formed visible biofilms both on silicone and glass, but their ability to grow as biofilms varied (Fig. 1c). The highest biofilm-forming capacities were determined for both Schwanniomyces species, D. fabryi, and for K. marxianus. Microscopic studies of biofilm architecture revealed the presence of visible ECM deposits (Fig. 2). These four strong biofilm producers were subjected to biochemical analyses of the ECM composition. It is noteworthy that no biofilm-like structures were observed in yeast cultures grown in freshwater-based media.
Scanning electron microscopy of extracellular matrix deposits present in the halotolerant yeast biofilms of Schwannyomyces etchellsii, Schwanniomyces polymorphus, Debaryomyces fabryi, and Kluyveromyces marxianus. Forty-eight-hour-old biofilms were grown in seawater-YPD medium (SW-YPD). Scale bars represent 10 μm
Glycosyl composition of the biofilm ECM of halotolerant yeast strains
Fungal biofilm functional biology has been linked to the ECM quantity and its composition, in which carbohydrates constitute a considerable fraction (Reichhardt et al. 2015; Zarnowski et al. 2014). The ECM carbohydrate profiles of the tested halotolerant yeast strains consisted of three pentoses (ribose, arabinose, and xylose) and two hexoses (mannose, glucose), but their ratios varied between species and, even to a higher degree, growth conditions (Fig. 3). Mannose represented the most abundant sugar in the biofilm ECMs formed in silicone tubes for K. marxianus and S. polymorphus, with percentages reaching 50%, whereas comparable amounts of mannose and arabinose were found in D. fabryi and S. etchellsii. In the case of biofilms formed in glass beakers, mannose was present at lower concentrations, while arabinose was usually the most abundant carbohydrate found. In fact, this monosaccharide was especially predominant in air-medium biofilms formed by D. fabryi, S. polymorphus, and K. marxianus, where its content reached up to 80%. The amount of arabinose in the S. etchellsii ECMs was low, whereas higher ribose concentration was detected in comparison with the other strains. Interestingly, significant variability in the content of this monosaccharide was observed, with percentages usually higher in the exopolysaccharides of the biofilms formed on silicone. The highest glucose content was found in the S. etchellsii ECMs (around 25%), while in the other cases, percentages between 2 and 15 were detected. Two additional monocarbohydrates, xylose and rhamnose, were identified in the tested ECMs, but in low or even trace amounts with percentages up to 11 in silicone biofilms and 4 in glass beaker biofilms, respectively.
Proteomic analysis of the biofilm ECM of halotolerant yeast strains
Proteins are the major component of the ECM in yeast biofilms (Mitchell et al. 2016). In this work, proteins that represented > 0.05% of the respective proteome were considered (Table 2). Bearing in mind this cutoff, the analyzed proteomes of biofilm ECMs of the halotolerant yeasts contained between 220 and 390 proteins depending on the strain and growth condition. Unlike the observed differences in the ECM carbohydrate profiles, the analysis of the ECM proteomes revealed striking similarities with reference to the most abundant functional categories and subcategories. Gene Ontology–based functional predictions grouped most of these proteins into the “metabolic process” category with a larger number classified as “biosynthetic process” when compared to “catabolic process,” which reflects a relevant molecular “building” activity in these structures (Table 2 and Fig. 4). Three other major functional protein groups, including “establishment of localization,” “response to stress,” and “cellular component organization,” were also abundant (Figs. 4, 5, 6, 7, and 8 and Fig. S1 in the Supplementary Material). Tables S1 to S8 in the Supplementary Material compilate the information about the proteins found in diverse categories and subcategories ordered by their weight percentage.
Distribution of main functional categories of proteins identified in the biofilm ECMs of the halotolerant yeasts, Schwannyomyces etchellsii, Schwanniomyces polymorphus, Debaryomyces fabryi, and Kluyveromyces marxianus. Functional protein assignments were done based on the Gene Ontology data in the UniProt Knowledgebase
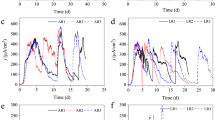
Comparative ECM proteomics of Debaryomyces fabryi biofilms grown in glass beakers and silicone tubes. a A Venn diagram illustrating the number of exclusive (green and red) and common (yellow) biofilm ECM proteins. b, c, d Smallest clusters represent individual identified proteins and are arranged inside higher level regions according to their GO biological process and molecular function assignments. Z-scores were calculated based on log2 silicone tube/glass beaker ratios of biofilm proteins and mapped to a color ramp starting with red (more protein in glass beaker biofilms), passing yellow (similar protein proportions under both growth conditions), and reaching green (more protein in silicone tube biofilms)
Comparative ECM proteomics of Schwannyomyces etchellsii biofilms grown in glass beakers and silicone tubes. a A Venn diagram illustrating the number of exclusive (green and red) and common (yellow) biofilm ECM proteins. b, c, d Smallest clusters represent individual identified proteins and are arranged inside higher level regions according to their GO biological process and molecular function assignments. Z-scores were calculated based on log2 silicone tube/glass beaker ratios of biofilm proteins and mapped to a color ramp starting with red (more protein in glass beaker biofilms), passing yellow (similar protein proportions under both growth conditions), and reaching green (more protein in silicone tube biofilms)
Comparative ECM proteomics of Schwanniomyces polymorphus biofilms grown in glass beakers and silicone tubes. a A Venn diagram illustrating the number of exclusive (green and red) and common (yellow) biofilm ECM proteins. b, c, d Smallest clusters represent individual identified proteins and are arranged inside higher level regions according to their GO biological process and molecular function assignments. Z-scores were calculated based on log2 silicone tube/glass beaker ratios of biofilm proteins and mapped to a color ramp starting with red (more protein in glass beaker biofilms), passing yellow (similar protein proportions under both growth conditions), and reaching green (more protein in silicone tube biofilms)
Comparative ECM proteomics of Kluyveromyces marxianus biofilms grown in glass beakers and silicone tubes. a A Venn diagram illustrating the number of exclusive (green and red) and common (yellow) biofilm ECM proteins. b, c, d Smallest clusters represent individual identified proteins and are arranged inside higher level regions according to their GO biological process and molecular function assignments. Z-scores were calculated based on log2 silicone tube/glass beaker ratios of biofilm proteins and mapped to a color ramp starting with red (more protein in glass beaker biofilms), passing yellow (similar protein proportions under both growth conditions), and reaching green (more protein in silicone tube biofilms)
Our detailed analysis revealed the presence of multiple enzymes participating in glycolysis/gluconeogenesis and in other processes strictly related to carbohydrate metabolism. For example, about 9% and 12% of the ECM proteins of D. fabryi grown on silicone and glass, respectively, were functionally classified as such (Tables S1 and S2 in the Supplementary Material), while these numbers were higher for K. marxianus and amounted 23% and 18%, respectively (Tables S3 and S4 in the Supplementary Material). Proteins involved in the biosynthesis of nitrogen compounds such as amino acids, proteins, and nucleotides were also found in abundance. Among the proteins involved in the response to stress, heat shock proteins and enzymes participating in the decomposition of reactive oxygen species were prevalent. In the functional categories of establishment of localization (localization) and cellular component organization (and biogenesis) proteins involved in transport processes, folding and ribosomal structure are mainly found (Figs. 4, 5, 6, 7, and 8 and Fig. S1 and Tables S1 to S8 in the Supplementary Material).
A compelling correlation between the ECM of the biofilms formed on silicone and glass beakers was observed with reference to the most abundant proteins with percentages higher than 0.7–1% (proteins in bold in Figs. 4, 5, 6, 7, and 8). The vast majority of those proteins were represented by heat shock proteins (such as SSA1, SSA2, and sphingolipid long chain base-responsive protein PIL1) or enzymes involved in glycolysis/gluconeogenesis (glyceraldehyde-3-phosphate dehydrogenase, pyruvate kinase, and enolase), pentose phosphate pathway (6-phosphogluconate dehydrogenase), Krebs cycle (malate dehydrogenase), and ATP synthesis (several subunits of the ATP synthase). In addition, some ribosomal proteins were also identified.
Despite these common aspects described above, there were also considerable differences found in the ECM protein composition of the biofilms formed on silicone and in glass beakers. There were pools of proteins detected exclusively under one type of the tested biofilm growth condition as well as groups of proteins that were abundant under both growth conditions, but their expression levels differed (see Figs. 5, 6, 7, and 8, Fig. S1 and Table S9 in the Supplementary Material). Detailed analysis of all results presented in this work revealed the number of biofilm proteins identified solely in silicone tubes was lower than that observed in the case of glass beakers (being the only exception S. etchellsii). About 3-fold more proteins were identified in glass beakers than in silicone tubes ECMs in the case of D. fabryi and K. marxianus. Interestingly, some of these proteins were exclusive of each strain, but others were found in two, three, or even in the four studied yeast species. For instance, 5-(hydroxymethyl) glutathione dehydrogenase was identified in all four yeast strains while grown in glass beaker biofilms. Other enzymes involved in carbohydrate metabolism, such as ATP-dependent 6-phosphofructokinase, glycerol-3-phosphate dehydrogenase, and pyruvate carboxylase, were detected only in these biofilms in three out of the four herein tested strains. Some aminoacyl-tRNA-synthetases (especially those involved in alanine, arginine, leucine, and lysine metabolism) were only observed in D. fabryi and in S. polymorphus. Similarly, glyoxalase II and nuclear transport 2 protein were detected in silicone biofilms in three of the four strains, while some components of the proteasome endopeptidase complex appeared uniquely in those formed by S. etchellsii and S. polymorphus.
Our in-depth analysis of the ECM proteomes laid a solid base for potential protein-based discriminatory studies among the halotolerant yeast species considered herein. For instance, D. fabryi had the lowest percentage of proteins involved in metabolism (Table 2). This value was 58–59%, whereas this number was around 70% in other species. In addition, proteins involved in lipid metabolism were found in higher amounts in the K. marxianus biofilms grown on silicone and glass (see Supplemental Tables S3 and S4 in comparison with S1, S2, and S5 to S8). Enzymes involved in amino acid catabolism were also more abundant in the ECM of these biofilms. As mentioned above, some proteins were abundantly formed under both biofilm growth conditions (Figs. 5, 6, 7, and 8 and Table S9 in the Supplementary Material). For instance, β-glucosidases, previously reported in the C. albicans biofilm ECM (Taff et al. 2012; Zarnowski et al. 2014), were detected in the biofilms formed only in silicone tubes by the tested Schwanniomyces species (Supplemental Tables S1–S9). Regulatory subunits of 26S protease as well as some ribosomal proteins, d-3-phosphoglycerate dehydrogenase, GTP-binding nuclear protein, and homocitrate synthase were found only in these K. marxianus biofilm ECMs. In the case of the other strains, catalase (D. fabryi), cystathionine beta-synthase and glutamate-5-semialdehyde dehydrogenase (S. etchellsii), and 3-isopropylmalate dehydrogenase and calmodulin (S. polymorphus) were unique in the biofilm ECMs formed in this material. Likewise, several proteins were only identified in the biofilms formed by individual strains in glass beakers. Catalase, some components of the cytochrome b-c1, and the proteasome endopeptidase complexes were formed in K. marxianus, whereas several nucleolar proteins, the ubiquitin-activating enzyme E1 and the pyruvate dehydrogenase E1 component subunit beta were also unique of these structures in D. fabryi.
Discussion
Biofilm formation is an elegant, powerful, and inexpensive solution for cell immobilization. One of the preferred strategies involves physical entrapment of the cells in a polymeric matrix (Halan et al. 2012; Muffler et al. 2014; Rosche et al. 2009). In the biocatalysis field, cell immobilization by means of biofilms provides some benefits, such as the ease of their formation; the low cost of the process compared to other methods of immobilizing enzymes or whole cells, especially for large-scale use; and the increased stability and robustness of these structures to high reactants/product concentrations (Winn et al. 2012). In addition, this approach offers use of high cell densities, in which enzymes can remain active at higher levels over long periods of operation and formed products can be easily recovered from the cells (Muffler et al. 2014). Unfortunately, certain disadvantages can also be experienced in these communities when used for biocatalysis, including undesired side reactions, potential contamination due to cell leakage, or limited mass transfer of substrates and products through the cell membrane and immobilizing matrix (Muffler et al. 2014). Although the use of biofilm communities for the production of fine chemicals has not yet been widely implemented at an industrial scale, several research studies have been reported within the last few years, indicating its great future potential. For example, native and recombinant strains of Pseudomonas sp. have been used to catalyze the styrene asymmetric epoxidation, the selective octane hydroxylation to octanol, and the production of (R)-(+) perillic acid from (R)-(+)-limonene (Gross et al. 2007, 2013; Halan et al. 2014; Willrodt et al. 2017). Besides, Bacillus sp. Mcn4 biofilms have been demonstrated as an excellent support for enzymes of its extracellular matrix (Romero et al. 2018). Based on these studies, it is imperative to gain more detailed information on the ability of biocatalysts to form biofilms under potentially advantageous growth conditions leading to production of desired fine chemicals.
Both formation and composition of fungal biofilms have been extensively studied for more than three decades, with special emphasis on C. albicans biofilms due to their role in colonization and infections in humans. Thus, over the last several years, many research groups have provided important information about the C. albicans biofilm life cycle and its clinical consequences (Wall et al. 2019). Furthermore, one of the first studies on S. cerevisiae biofilms revealed its capability to adhere to polystyrene and polypropylene surfaces at low-glucose concentrations and demonstrated the requirement of Flo11 in this attachment process (Reynolds and Fink 2001). The flor or velum formation by the S. cerevisiae flor strains is also a form of air-liquid interfacial biofilm that is essential for aerobic growth during the production of sherry or sherry-like wines (Zara et al. 2011). The abovementioned protein Flo11 was also found to be essential in this biological system (Zara et al. 2005). The ability of some commercial wine yeasts to form biofilms has been described with the presence of cells with different lifestyles and growth modes, such as invasive growth, bud elongation, and sporulation (Tek et al. 2018). These authors have proposed that such cellular organization allows for yeasts to colonize the winery environment. In this work, we report on the biofilm formation and composition of the halotolerant yeast strains, K. marxianus, D. fabryi, S. etchellsii, and S. polymorphus, that are of great importance in several essential industrial and biotechnological processes. We demonstrate that they can form two kinds of such communities in the presence of seawater: air-liquid biofilms on the liquid surface in glass beakers and biofilms attached to silicone tubes.
The biofilm ECM is not a passive structural material in biofilms; rather, it is dynamic and involved in nutrient sequestration, water adsorption, protection of free-living cells from environmental stress, signaling, migration, and genetic exchange (Dragos and Kovacs 2017). Both its structure and composition depend on the organism; the type of media; if any, supporting biofilm growth; and environmental or process characteristics (Edel et al. 2019). Overall, fungal biofilm ECMs appear to be complex blends of all types of biologics: proteins, carbohydrates, lipids, and nucleic acids (Mitchell et al. 2016). In all the cases described up to now, both protein and carbohydrates were the main components (40% and 43% in A. fumigatus, and 55% and 25% in C. albicans, respectively) (Reichhardt et al. 2015; Zarnowski et al. 2014).
Polysaccharides represent a key component of ECMs in all the biofilms described so far. In the yeast strains considered herein, mannose represented a very abundant monosugar in the biofilms formed in silicone tubes, and its weight percentages were around 50% for K. marxianus and S. polymorphus (Fig. 3). Arabinose was usually the most abundant carbohydrate in biofilms formed on glass beakers, whereas xylose and rhamnose were present in all tested biofilm ECMs, but at very low concentrations. The content of two other identified carbohydrates, ribose and glucose, varied significantly in the analyzed ECMs. The monosaccharide profiles of the biofilm ECMs formed in silicone tubes were similar for the four analyzed strains and completely different to those observed in glass beakers. In this sense, the main distinctive feature among the tested yeast strains was found for S. etchellsii biofilms grown in glass beakers, which had noticeably lower levels of arabinose. In addition, higher ribose and lower arabinose concentrations were determined in these biofilms when compared to their silicone counterparts. Overall, we concluded that the measured monocarbohydrate profiles were relatively simple yet offering some potential for discriminatory studies.
Proteins represent a large portion of the biomass in most of the reported ECM analyses (Branda et al. 2005; Branda et al. 2006; Flemming and Wingender 2010). In all of the proteomes analyzed in this work (Figs. 5, 6, 7, and 8, Tables 2 and Fig. S1 and Tables S1–S9 in the Supplementary Material), the overrepresented functional protein categories corresponded to metabolism, particularly carbohydrate and amino acid biosynthesis, and response to stimuli. Glyceraldehyde-3-phosphate dehydrogenases (GAPDH), along with other glycolytic and gluconeogenic enzymes, several heat shock proteins, and antioxidant enzymes such as superoxide dismutase and catalase, were particularly abundant. These results are in agreement with previously published data on protein composition in yeast biofilm ECMs. For instance, Lattif et al. (2008) showed GAPDH to be the only upregulated enzyme of the glycolysis/gluconeogenesis pathways during the early phase of biofilm formation by C. albicans, and its expression level was 9.1 fold higher when compared to cell walls. Thus, GAPDH was identified as a key player during biofilm formation. Two other enzymes belonging to these pathways, phosphoglycerate kinase and fructose bisphosphate aldolase, were also upregulated in maturing biofilms (Lattif et al. 2008). In addition, glutamate and nitrogen metabolism pathways were represented only in early phase biofilms, while proteins involved in purine metabolism, glycine/serine/threonine metabolism, carbon fixation, and inositol metabolism were primarily upregulated in mature phase biofilms. In the ECM of S. cerevisiae biofilm-like mats, GAPDH was also found (Faria-Oliveira et al. 2015) along with other proteins that were grouped into distinct functional classes, mostly including metabolism, protein fate/remodeling, and cell rescue and defense mechanisms. Interestingly, heat shock chaperones, metalloproteinases, superoxide dismutase, broad signaling cross-talkers, and other putative signaling proteins were present in abundance. In the case of A. fumigatus biofilm-like structures, catalase B was detected as one of the major proteins in the ECM; the presence of this enzyme was associated with its protective role against reactive oxygen species, which microbes encounter in excessive amounts while infecting a host (Reichhardt et al. 2015). In fact, catalases have been reported to play a critical role in other microbial biofilms, such as those formed by Pseudomonas aeruginosa (Elkins et al. 1999). It is worth mentioning that elevated expression levels of heat shock proteins have been reported during C. albicans biofilm formation (Becherelli et al. 2013). The similarities found between the protein composition of the biofilm ECMs of non-pathogenic yeasts tested in this work and those causing infection suggest that the presence of antioxidant enzymes and other stress response proteins is not only relevant to yeast protection during infection but also from the environmental conditions in which biofilms exist.
Many of the proteins identified herein did not contain secretion signals, and their extracellular location could not be easily explained. This striking observation is in agreement with previously published studies characterizing biofilm ECM composition. The presence of these proteins in the biofilm extracellular milieu could be due to either the use of a non-secretory pathway, which does not require leading secretion sequences, or leakage of proteins from cells undergoing cellular death (Nickel and Rabouille 2009; Nombela et al. 2006; Nosanchuk et al. 2008; Zarnowski et al. 2014). However, the existence of several hundred moonlighting proteins, which in many cases perform multiple Gene Ontology–based unpredictable functions, has been documented recently (Jeffery 2018). Often, these classically intracellular proteins are secreted extracellularly by unknown mechanisms and become involved in completely different, functionally distinct processes. For example, cellular ribosomal proteins have been identified in the Staphylococcus aureus biofilm ECM, where they are responsible for physical stabilization of the microbial community (Graf et al. 2019). According to these authors, the oxygen limitation in biofilms determined the release of fermentation products like formate, lactate, and acetate, which resulted in an acidic environment, while the strong positive charge located on those specific ECM proteins probably mediated electrostatic interactions with anionic cell surface components and anionic metabolites leading to strong aggregation and biofilm stabilization. The glycolytic enzyme GAPDH, overexpressed in many yeast biofilms as commented above, has also been shown to play an important role in the organization of these communities by C. albicans; the ability of this microorganism to form early and mature phase biofilms was significantly reduced in the presence of sodium iodoacetate, which specifically and irreversibly inhibits this enzyme (Lattif et al. 2008). It is hence difficult to assign a proper function that an individual protein could have once located in the biofilm ECM; actually, it could not be related to its cellular role.
The results described in this work reveal important differences in the content of the biofilm ECM considered both in monosugars and proteins. It is worth mentioning that the atomic composition in both materials is the same, silicon and oxygen, but their synthesis, structure, and other features are very different. In glass, atoms are linked in a crisscross network; in silicone, long lineal chains are formed and joined by cross links. Physical properties (hardness, elasticity, porosity, rugosity …) are also very different. In fact, in glass beakers, biofilms are not attached to the glass surface (they are mainly formed in the interface between the liquid medium and air), while in silicone tubes, biofilms are made on the solid material.
Many of the enzymes found in the biofilm ECM function as an external digestive system that breaks down extracellular biopolymers later used as energy sources (Zarnowski et al. 2014). In fact, other studies of bacterial biofilms have demonstrated their ECMs to be enzymatically active (Absalon et al. 2012; Chen et al. 2013; Jiao et al. 2011; Romani et al. 2008; Sutherland 2001). Relatively high percentages of metabolic enzymes in the biofilm ECMs of K. marxianus and both Schwanniomyces strains used in this work suggest that these yeast biofilms are more suitable as potential biocatalysts, although more thorough experimental verification would be required. In addition, differences were found with reference to the enzymes that are unique or present in higher percentages in the ECM of biofilms formed in silicone tubes and in glass beakers (see Figs. 5, 6, 7, and 8, and Table S9 and Fig. S1 in the Supplementary Material) for each one of the tested strains. This information could be helpful for the selection of the most appropriate surface for biofilm formation depending on the biocatalytic purposes.
The halotolerant yeast strains tested in this work have been extensively used in our laboratory for the stereoselective synthesis of compounds of interest for the pharmaceutical and chemical industries, using seawater in some cases as a sustainable solvent (Andreu and del Olmo 2018, 2019, 2020). Their ability to form biofilms in seawater-based growth medium as well as both the abundance and the complexity of enzymes located in these extracellular structures offer unlimited possibilities for their use as a convenient strategy for cell immobilization for biocatalytic purposes. Compared to planktonic cells, the ability of microorganisms to live in complex biofilms results in higher stability, physical robustness, and increased resistance to toxic compounds (Halan et al. 2012). All these factors strongly support the idea of using immobilized biofilms in biocatalytic applications. Although this strategy has not been widely implemented at an industrial scale, these biofilm systems have already been used for the production of valuable chemicals or manufacturing of value-added products (Edel et al. 2019). With reference to novel industrial applications, it is still necessary to gain more in-depth knowledge about and understanding of long-term operability and techniques to steer the desired processes and large-scale reactor systems. In fact, existing membrane biofilm reactors and bioelectrochemical systems appear to be promising tools for highly productive biofilms (Edel et al. 2019; Ontiveros-Valencia et al. 2018). Future studies will determine factors related to the applicability of individual biofilm structure types in existing and developing biocatalytic processes and provide better understanding of some aspects related to biotechnological capabilities of these halotolerant yeast-based biofilms.
Data availability
Exhaustive information about the proteins identified in the extracellular matrix of the biofilms formed by the yeast strains considered in this work is included in the Supplementary Material.
References
Absalon C, Ymele-Leki P, Watnick PI (2012) The bacterial biofilm matrix as a platform for protein delivery. mBio 3:e00127-12. https://doi.org/10.1128/mBio.00127-12
Andersen KS, Bojsen R, Sorensen LGR, Nielsen MW, Lisby M, Folkesson A, Regenberg B (2014) Genetic basis for Saccharomyces cerevisiae biofilm in liquid medium. G3-Genes Genom Genet 4:1671–1680. https://doi.org/10.1534/g3.114.010892
Andreu C, del Olmo M (2014) Potential of some yeast strains in the stereoselective synthesis of (R)-(-)-phenylacetylcarbinol and (S)-(+)-phenylacetylcarbinol and their reduced 1,2-dialcohol derivatives. Appl Microbiol Biotechnol 98:5901–5913. https://doi.org/10.1007/s00253-014-5635-5
Andreu C, Peña M, Del Olmo M (2016) Biocatalytic reduction of racemic 2-arenoxycycloalkanones by yeasts P. glucozyma and C. glabrata: one way of achieving chiral 2-arenoxycycloalcohols. Appl Microbiol Biotechnol 100:4865–4873. https://doi.org/10.1007/s00253-015-7261-2
Andreu C, del Olmo M (2018) Biotransformation using halotolerant yeast in seawater: a sustainable strategy to produce R-(-)-phenylacetylcarbinol. Appl Microbiol Biotechnol 102:4717–4727. https://doi.org/10.1007/s00253-018-8945-1
Andreu C, del Olmo M (2019) Improved biocatalytic activity of the Debaryomyces species in seawater. Chemcatchem 11:3085–3092. https://doi.org/10.1002/cctc.201900558
Andreu C, del Olmo M (2020) Whole-cell biocatalysis in seawater: new halotolerant yeast strains for the regio- and stereoselectivity reduction of 1-phenylpropane-1,2-dione in saline-rich media. Chembiochem 21:1621–1628. https://doi.org/10.1002/cbic.202000023
Bales PM, Renke EM, May SL, Shen Y, Nelson DC (2013) Purification and characterization of biofilm-associated EPS exopolysaccharides from ESKAPE organisms and other pathogens. PLoS One 8:e67950. https://doi.org/10.1371/journal.pone.0067950
Becherelli M, Tao J, Ryder NS (2013) Involvement of heat shock proteins in Candida albicans biofilm formation. J Mol Microbiol Biotechnol 23:396–400. https://doi.org/10.1159/000351619
Berlanga M, Guerrero R (2016) Living together in biofilms: the microbial cell factory and its biotechnological implications. Microb Cell Fact 15:165. https://doi.org/10.1186/s12934-016-0569-5
Bernhardt J, Funke S, Hecker M, Siebourg J (2009) Visualizing gene expression data via Voronoi treemaps. In: 2010 International Symposium on Voronoi Diagrams in Science and Engineering. Copenhagen, Denmark, pp 233–241. https://doi.org/10.1109/ISVD.2009.33
Blakeney AB, Harris PJ, Henry RJ, Stone BA, Norris T (1982) Gas-chromatography of alditol acetates on a high-polarity bonded-phase vitreous-silica column. J Chromatogr 249:180–182. https://doi.org/10.1016/S0021-9673(00)80246-0
Branda SS, Vik S, Friedman L, Kolter R (2005) Biofilms: the matrix revisited. Trends Microbiol 13:20–26. https://doi.org/10.1016/j.tim.2004.11.006
Branda SS, Chu F, Kearns DB, Losick R, Kolter R (2006) A major protein component of the Bacillus subtilis biofilm matrix. Mol Microbiol 59:1229–1238. https://doi.org/10.1111/j.1365-2958.2005.05020.x
Castrillón RLE, Palma RA, Padilla DMC (2013) Biopelículas fúngicas. Dermatología Rev Mex 57:350–361
Chen C, Krishnan V, Macon K, Manne K, Narayana SVL, Schneewind O (2013) Secreted proteases control autolysin-mediated biofilm growth of Staphylococcus aureus. J Biol Chem 288:29440–29452. https://doi.org/10.1074/jbc.M113.502039
Cheng YF, Feng GP, Moraru CI (2019) Micro- and nanotopography sensitive bacterial attachment mechanisms: a review. Front Microbiol 10:191. https://doi.org/10.3389/fmicb.2019.00191
Colvin KM, Gordon VD, Murakami K, Borlee BR, Wozniak DJ, Wong GCL, Parsek MR (2011) The Pel polysaccharide can serve a structural and protective role in the biofilm matrix of Pseudomonas aeruginosa. PLoS Pathog 7:e1001264. https://doi.org/10.1371/journal.ppat.1001264
Colvin KM, Irie Y, Tart CS, Urbano R, Whitney JC, Ryder C, Howell PL, Wozniak DJ, Parsek MR (2012) The Pel and Psl polysaccharides provide Pseudomonas aeruginosa structural redundancy within the biofilm matrix. Environ Microbiol 14:1913–1928. https://doi.org/10.1111/j.1462-2920.2011.02657.x
Demirci A, Pometto AL, Ho KLG (1997) Ethanol production by Saccharomyces cerevisiae in biofilm reactors. J Ind Microbiol Biot 19:299–304. https://doi.org/10.1038/sj.jim.2900464
Dominguez E, Zarnowski R, Sanchez H, Covelli AS, Westler WM, Azadi P, Nett J, Mitchell AP, Andes DR (2018) Conservation and divergence in the Candida species biofilm matrix mannan-glucan complex structure, function, and genetic control. Mbio 9:e00451–e00418. https://doi.org/10.1128/mBio.00451-18
Dominguez EG, Zarnowski R, Choy HL, Zhao M, Sanchez H, Nett JE, Andes DR (2019) Conserved role for biofilm matrix polysaccharides in Candida auris drug resistance. Msphere 4:e00680–e00618. https://doi.org/10.1128/mSphereDirect.00680-18
Dragos A, Kovacs AT (2017) The peculiar functions of the bacterial extracellular matrix. Trends Microbiol 25:257–266. https://doi.org/10.1016/j.tim.2016.12.010
Edel M, Horn H, Gescher J (2019) Biofilm systems as tools in biotechnological production. Appl Microbiol Biotechnol 103:5095–5103. https://doi.org/10.1007/s00253-019-09869-x
Elkins JG, Hassett DJ, Stewart PS, Schweizer HP, McDermott TR (1999) Protective role of catalase in Pseudomonas aeruginosa biofilm resistance to hydrogen peroxide. Appl Environ Microbiol 65:4594–4600. https://doi.org/10.1128/AEM.65.10.4594-4600.1999
Faria-Oliveira F, Carvalho J, Ferreira C, Hernaez ML, Gil C, Lucas C (2015) Quantitative differential proteomics of yeast extracellular matrix: there is more to it than meets the eye. BMC Microbiol 15:271. https://doi.org/10.1186/s12866-015-0550-1
Flemming HC, Wingender J (2010) The biofilm matrix. Nat Rev Microbiol 8:623–633. https://doi.org/10.1038/nrmicro2415
Graf AC, Leonard A, Schauble M, Rieckmann LM, Hoyer J, Maass S, Lalk M, Becher D, Pane-Farre J, Riedel K (2019) Virulence factors produced by Staphylococcus aureus biofilms have a moonlighting function contributing to biofilm integrity. Mol Cell Proteomics 18:1036–1053. https://doi.org/10.1074/mcp.RA118.001120
Gross R, Hauer B, Otto K, Schmid A (2007) Microbial biofilms: new catalysts for maximizing productivity of long-term biotransformations. Biotechnol Bioeng 98:1123–1134. https://doi.org/10.1002/bit.21547
Gross R, Buehler K, Schmid A (2013) Engineered catalytic biofilms for continuous large scale production of n-octanol and (S)-styrene oxide. Biotechnol Bioeng 110:424–436. https://doi.org/10.1002/bit.24629
Halan B, Buehler K, Schmid A (2012) Biofilms as living catalysts in continuous chemical syntheses. Trends Biotechnol 30:453–465. https://doi.org/10.1016/j.tibtech.2012.05.003
Halan B, Letzel T, Schmid A, Buehler K (2014) Solid support membrane-aerated catalytic biofilm reactor for the continuous synthesis of (S)-styrene oxide at gram scale. Biotechnol J 9:1339–1349. https://doi.org/10.1002/biot.201400341
Hall-Stoodley L, Costerton JW, Stoodley P (2004) Bacterial biofilms: from the natural environment to infectious diseases. Nat Rev Microbiol 2:95–108. https://doi.org/10.1038/nrmicro821
Ho KLG, Pometto AL, Hinz PN (1997) Optimization of L-(+)-lactic acid production by ring and disc plastic composite supports through repeated-batch biofilm fermentation. Appl Environ Microbiol 63:2533–2542. https://doi.org/10.1128/Aem.63.7.2533-2542.1997
Inigo M, Del Pozo JL (2018) Fungal biofilms: from bench to bedside. Rev Esp Quimioter 31(Suppl 1):35–38
Jeffery CJ (2018) Protein moonlighting: what is it, and why is it important? Philos Trans R Soc Lond B Biol Sci 373:20160523. https://doi.org/10.1098/rstb.2016.0523
Jiao YQ, D’haeseleer P, Dill BD, Shah M, VerBerkmoes NC, Hettich RL, Banfield JF, Thelen MP (2011) Identification of biofilm matrix-associated proteins from an acid mine drainage microbial community. Appl Environ Microbiol 77:5230–5237. https://doi.org/10.1128/Aem.03005-10
Keller A, Purvine S, Nesvizhskii AI, Stolyar S, Goodlett DR, Kolker E (2002) Experimental protein mixture for validating tandem mass spectral analysis. OMICS 6:207–212. https://doi.org/10.1089/153623102760092805
Kurtzman CP, Suzuki M (2010) Phylogenetic analysis of ascomycete yeasts that form coenzyme Q-9 and the proposal of the new genera Babjeviella, Meyerozyma, Millerozyma, Priceomyces, and Scheffersomyces. Mycoscience 51:2–14. https://doi.org/10.1007/s10267-009-0011-5
Lattif AA, Chandra J, Chang J, Liu S, Zhou G, Chance MR, Ghannoum MA, Mukherjee PK (2008) Proteomics and pathway mapping analyses reveal phase-dependent over-expression of proteins associated with carbohydrate metabolic pathways in Candida albicans biofilms. The Open Proteomics J 1:5–26. https://doi.org/10.2174/1875039700801010005
Le Mauff F, Bamford NC, Alnabelseya N, Zhang YZ, Baker P, Robinson H, Codee JDC, Howell PL, Sheppard DC (2019) Molecular mechanism of Aspergillus fumigatus biofilm disruption by fungal and bacterial glycoside hydrolases. J Biol Chem 294:10760–10772. https://doi.org/10.1074/jbc.RA119.008511
Martin SE, Shabanowitz J, Hunt DF, Marto JA (2000) Subfemtomole MS and MS/MS peptide sequence analysis using nano-HPLC micro-ESI Fourier transform ion cyclotron resonance mass spectrometry. Anal Chem 72:4266–4274. https://doi.org/10.1021/ac000497v
Martinez LR, Casadevall A (2015) Biofilm formation by Cryptococcus neoformans. Microbiol Spectr 3:MB-0006-2014. https://doi.org/10.1128/microbiolspec.MB-0006-2014
Mitchell KF, Zarnowski R, Andes DR (2016) The extracellular matrix of fungal biofilms. Adv Exp Med Biol 931:21–35. https://doi.org/10.1007/5584_2016_6
Mitchell KF, Zarnowski R, Sanchez H, Edward JA, Reinicke EL, Nett JE, Mitchell AP, Andes DR (2015) Community participation in biofilm matrix assembly and function. Proc Natl Acad Sci U S A 112:4092–4097. https://doi.org/10.1073/pnas.1421437112
Muffler K, Lakatos M, Schlegel C, Strieth D, Kuhne S, Ulber R (2014) Application of biofilm bioreactors in white biotechnology. Adv Biochem Eng Biot 146:123–161. https://doi.org/10.1007/10_2013_267
Nickel W, Rabouille C (2009) Mechanisms of regulated unconventional protein secretion. Nat Rev Mol Cell Biol 10:148–155. https://doi.org/10.1038/nrm2617
Nombela C, Gil C, Chaffin WL (2006) Non-conventional protein secretion in yeast. Trends Microbiol 14:15–21. https://doi.org/10.1016/j.tim.2005.11.009
Nosanchuk JD, Nimrichter L, Casadevall A, Rodrigues ML (2008) A role for vesicular transport of macromolecules across cell walls in fungal pathogenesis. Commun Integr Biol 1:37–39. https://doi.org/10.4161/cib.1.1.6639
Ontiveros-Valencia A, Zhou C, Zhao HP, Krajmalnik-Brown R, Tang Y, Rittmann BE (2018) Managing microbial communities in membrane biofilm reactors. Appl Microbiol Biotechnol 102:9003–9014. https://doi.org/10.1007/s00253-018-9293-x
Perez-Riverol Y, Csordas A, Bai J, Bernal-Llinares M, Hewapathirana S, Kundu DJ, Inuganti A, Griss J, Mayer G, Eisenacher M, Pérez E, Uszkoreit J, Pfeuffer J, Sachsenberg T, Yilmaz S, Tiwary S, Cox J, Audain E, Walzer M, Jarnuczak AF, Ternent T, Brazma A, Vizcaíno JA (2019) The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res 47(D1):D442–D450. https://doi.org/10.1093/nar/gky1106
Perkins DN, Pappin DJC, Creasy DM, Cottrell JS (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20:3551–3567. https://doi.org/10.1002/(Sici)1522-2683(19991201)20:18<3551::Aid-Elps3551>3.0.Co;2-2
Qureshi N, Karcher P, Cotta M, Blaschek HP (2004) High-productivity continuous biofilm reactor for butanol production: effect of acetate, butyrate, and corn steep liquor on bioreactor performance. Appl Biochem Biotechnol 113-116:713–721. https://doi.org/10.1385/abab:114:1-3:713
Reichhardt C, Ferreira JA, Joubert LM, Clemons KV, Stevens DA, Cegelski L (2015) Analysis of the Aspergillus fumigatus biofilm extracellular matrix by solid-state nuclear magnetic resonance spectroscopy. Eukaryot Cell 14:1064–1072. https://doi.org/10.1128/EC.00050-15
Reynolds TB, Fink GR (2001) Bakers’ yeast, a model for fungal biofilm formation. Science 291:878–881. https://doi.org/10.1126/science.291.5505.878
Romani AM, Fund K, Artigas J, Schwartz T, Sabater S, Obst U (2008) Relevance of polymeric matrix enzymes during biofilm formation. Microb Ecol 56:427–436. https://doi.org/10.1007/s00248-007-9361-8
Romero CM, Martorella PV, Lopez AG, Penalver CGN, Chaves S, Mechetti M (2018) Architecture and physicochemical characterization of Bacillus biofilm as a potential enzyme immobilization factory. Colloid Surface B 162:246–255. https://doi.org/10.1016/j.colsurfb.2017.11.057
Rosche B, Li XZ, Hauer B, Schmid A, Buehler K (2009) Microbial biofilms: a concept for industrial catalysis? Trends Biotechnol 27:636–643. https://doi.org/10.1016/j.tibtech.2009.08.001
Stewart PS, Franklin MJ (2008) Physiological heterogeneity in biofilms. Nat Rev Microbiol 6:199–210. https://doi.org/10.1038/nrmicro1838
Strieth D, Ulber R, Muffler K (2018) Application of phototrophic biofilms: from fundamentals to processes. Bioprocess Biosyst Eng 41:295–312. https://doi.org/10.1007/s00449-017-1870-3
Sutherland IW (2001) The biofilm matrix - an immobilized but dynamic microbial environment. Trends Microbiol 9:222–227. https://doi.org/10.1016/S0966-842x(01)02012-1
Taff HT, Nett JE, Zarnowski R, Ross KM, Sanchez H, Cain MT, Hamaker J, Mitchell AP, Andes DR (2012) A Candida biofilm-induced pathway for matrix glucan delivery: implications for drug resistance. PLoS Pathog 8:e1002848. https://doi.org/10.1371/journal.ppat.1002848
Tek EL, Sundstrom JF, Gardner JM, Oliver SG, Jiranek V (2018) Evaluation of the ability of commercial wine yeasts to form biofilms (mats) and adhere to plastic: implications for the microbiota of the winery environment. FEMS Microbiol Ecol 94(2). https://doi.org/10.1093/femsec/fix188
UniProt Consortium (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47(D1):D506–D515. https://doi.org/10.1093/nar/gky1049
Wall G, Montelongo-Jauregui D, Vidal Bonifacio B, Lopez-Ribot JL, Uppuluri P (2019) Candida albicans biofilm growth and dispersal: contributions to pathogenesis. Curr Opin Microbiol 52:1–6. https://doi.org/10.1016/j.mib.2019.04.001
Willrodt C, Halan B, Karthaus L, Rehdorf J, Julsing MK, Buehler K, Schmid A (2017) Continuous multistep synthesis of perillic acid from limonene by catalytic biofilms under segmented flow. Biotechnol Bioeng 114:281–290. https://doi.org/10.1002/bit.26071
Winn M, Foulkes JM, Perni S, Simmons MJH, Overton TW, Goss RJM (2012) Biofilms and their engineered counterparts: a new generation of immobilised biocatalysts. Catal Sci Technol 2:1544–1547. https://doi.org/10.1039/c2cy20085f
Zara G, Budroni M, Mannazzu I, Zara S (2011) Air-liquid biofilm formation is dependent on ammonium depletion in a Saccharomyces cerevisiae flor strain. Yeast 28:809–814. https://doi.org/10.1002/yea.1907
Zara S, Bakalinsky AT, Zara G, Pirino G, Demontis MA, Budroni M (2005) FLO11-based model for air-liquid interfacial biofilm formation by Saccharomyces cerevisiae. Appl Environ Microbiol 71:2934–2939. https://doi.org/10.1128/AEM.71.6.2934-2939.2005
Zarnowski R, Westler WM, Lacmbouh GA, Marita JM, Bothe JR, Bernhardt J, Lounes-Hadj Sahraoui A, Fontaine J, Sanchez H, Hatfield RD, Ntambi JM, Nett JE, Mitchell AP, Andes DR (2014) Novel entries in a fungal biofilm matrix encyclopedia. Mbio 5:e01333–e01314. https://doi.org/10.1128/mBio.01333-14
Acknowledgements
We gratefully acknowledge Dr. J. Ramos for providing us with the Debaryomyces hansenii strain CBS767 and the SCSIE (Universitat de València) for access to its instrumental facilities. We thank Ms. Marienela Y. Heredia for comments on this manuscript.
Funding
This work was supported by grants from the Universitat de València (UV-INV-AE15-323062) and the National Institutes of Health (R01 AI073289).
Author information
Authors and Affiliations
Contributions
Conceptualization and supervision by C.A. and M.O. R.Z; C.A. and M.O. conceived, designed, performed research and analysis of data and wrote the paper. H.S. contributed with some experiments. D.A. was responsible for the discussion of results and supervision of the paper.
Corresponding author
Ethics declarations
Ethics approval
The article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 1105 kb)
Rights and permissions
About this article
Cite this article
Zarnowski, R., Sanchez, H., Andreu, C. et al. Formation and characterization of biofilms formed by salt-tolerant yeast strains in seawater-based growth medium. Appl Microbiol Biotechnol 105, 2411–2426 (2021). https://doi.org/10.1007/s00253-021-11132-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11132-1