Abstract
Tomatoes are an essential part of the human diet worldwide, and their production has been restricted by fungal disease. One of the most common fungal diseases of Lycopersicon esculentum L. is Sclerotinia stem rot, caused by Sclerotinia sclerotiorum. In this work, overexpression of the chimeric chit42 gene from Trichoderma atroviride with Serratia marcescens Chitinase B C-terminal fused chitin-binding domain (ChBD) in tomato was reported. Thirty independent transgenic lines were regenerated and adapted to greenhouse conditions. Five transgenic lines was selected for subsequent molecular and biological analysis. RT-PCR analysis of transgenic plants showed different expression patterns in independent transgenic events. Young leaves of T1 plants challenged with S. sclerotiorum revealed that expression of chimeric chitinase enhanced plant resistance against sclerotinia stem rot. A radial diffusion assay showed that transgenic lines with constitutive expression of the ChBD gene inhibited S. sclerotiorum growth significantly. Lesion sizes of transgenic tomatoes caused by S. sclerotiorum were significantly reduced compared to non-transgenic tomato plants. This is the first study reporting the evaluation of transgenic lines of tomato harboring Trichoderma chitinase (chit42) with chitin binding domain (ChBD) gene for resistance to one of the most important fungal diseases, S. sclerotiorum.
Key message
The chimeric chit42 gene with the Serratia marcescens chitinase binding domain has been expressed in tomato. The resistance of transgenic tomatoes to Sclerotinia stem rot has increased.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tomatoe (Lycopersicon esculentum Mill.) due to its nutrient value such as B-carotene, lycopene, and vitamin C, are a staple ingredient in the human diet (da Silva Santos et al. 2022). Fungal diseases reduce up to 30% of total yield limiting the productivity of tomato in some crucial crops (Chanda et al. 2021). Scleractinia, Alternaria, Aspergillus, Fusarium, and Verticillium are some of the fungal species that cause severe damage to monocotyledonous and dicotyledonous crops, including potato, tomato, cotton, caster, and chickpea (Malik et al. 2022). One of the most destructive diseases of tomato is Sclerotinia stem rot (SSR), caused by Sclerotinia sclerotiorum. The fungus is soil-borne infecting stalks through the midrib/petioles of the soil where the leaves contact (Mazumdar 2021). Transgenic technology has immense potential for resolving biological issues associated with the improvement of crops resistance to biotic and abiotic stress (Nalluri and Karri 2020). However, breeding resistance against S. sclerotiorum has historically been difficult due to a lack of major resistance genes in tomato germplasms (Wang et al. 2019).
Chitin is the second most prevalent polysaccharide in nature which is found in the cell walls of many fungal species. As such, continuous expression of chitin’s degrading enzyme—chitinase, in plants have been suggested as an adequate protection approach against fungi (Ghorbanpour et al. 2018; Chhikara et al. 2012). Chitinase is a biodegradable and safe biological agent compared to the use of traditional fungicides (Berini et al. 2019). Chitinase is a small pathogenesis-related proteins (PRs) with low molecular weight (LMW) secreted from the cell and transported across the plasma membrane via diffusion. Although a small protein, it is resistant to proteolytic degradation by proteases and could be fully soluble in low pH solutions. The cell wall structure of mycelial can be hydrolyzed by Chitinases through catalyzing the b-1, 4 links of d-glucosamine chains, inhibiting fungal growth (El-Sobki and Ali 2020).
Limited growth of different plant fungal pathogens (Rhizoctonia solani, Colletotrichum Gloeosporioides, Alternaria alternata, Fusarium oxysporum) has been reported through in-vitro overexpression of chitinase (Jambhulkar et al. 2018). The expression of chitinase has been promoted resistance to fungal plant pathogens (Zhang et al. 2021). Chitinase-encoding genes from Trichoderma species, in particular, have gotten a lot of attention since they have significantly better antifungal properties than their plant counterparts (Tien et al. 2021). Antifungal activity of chitinases from different Trichoderma species have previously reported (Kowsari et al. 2014a; Solgi et al. 2015; Loc et al. 2020). More than 75 different species, including Trichoderma species, have been utilized in biocontrol and biofertilizer formulations, as active agents (Marra et al. 2021).
Chitinases have a strong chitin-binding domain (ChBD) that functions as a tunnel-like structure that permits tight interaction with the polymeric substrate and N-acetylglucosamine residues in addition to a catalytic domain (Salas et al. 2015). ChBD is usually connected to the catalytic domain which improves chitinase binding to fungal chitins, allowing effective chitin breakdown (Vaghela et al. 2022). To increase the disease suppressive aspect and biocontrol efficiency of Chitinases, genetic engineering has been a gold standard method. This is especially important in the case of Trichoderma as Trichoderma chitinase gene (chit42) does not have a chitin-binding domain (Li et al. 2018).
Chit42 is a Trichoderma atroviride endo-chitinase with the ability to biocontrol against phytopathogenic fungi (Xia et al. 2018). Kowsari et al. (2014b) integrated a ChBD fragment to the chit42 gene, enhancing the ChBd’s binding capacity to insoluble chitin. By fusing Serratia marcescens chitinase binding domain (ChBD) to Trichoderma atroviride Chit42, Matroodi et al. (2013) constructed a chimeric chitinase with a better chitin-binding capability. When compared to the native Chit42, the fusion of ChBD to Chit42 enhanced the affinity of the enzyme to colloidal chitin as well as the enzyme activity of the chimeric chitinase which proved to function more efficiently than the native form of the enzyme against Candida albicans. Chimeric chitinase (Chit42-ChBD) genes were overexpressed in some important crops, including canola (Aghazadeh et al. 2016), tobacco (Badrhadad et al. 2018), Brassica (Zarinpanjeh et al. 2016; Zhu et al. 2021) resulting in an increased tolerance of transgenic plants to fungal diseases. Even though, antifungal genes from other crops have been used for improve tomato resistance to a fungal pathogen (Nuwamanya et al. 2022), in our knowledge, there has been no report on overexpression of the chitin-binding domain fused chitinase gene from Trichoderma atroviride in tomatoes. In this article, we report the expression of a chimeric chitinase gene of the Trichoderma atroviride linked to ChBD from the C-terminal of Serratia marcescens Chitinase B in Tomato for enhanced resistance against S. sclerotiorum.
Materials and methods
Plant material, fungi, bacterial strains and plasmid
In this study, Agrobacterium-mediated transformation was carried out utilizing the ‘Germeze kamrang urmiya’ tomato line (Lycopersicon esculentum L.). The seeds were provided from the Agricultural and Natural Resources Research and Training Center of West Azerbaijan (Azarbaijan, Iran). The Sclerotinia sclerotiorum strain was obtained from the fungal collection of Tabriz University (Mycology laboratory, Plant protection department, Faculty of Agriculture, Tabriz University, Iran).
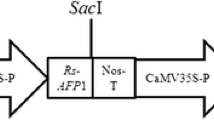
Plasmid pBISM2 (Fig. 1) (Chimeric chit42 from T. atroviride with a S. marcescens ChiB C-terminal fused chitin-binding domain) (Matroodi et al. 2013), was used for plant transformation. Sub-cloning and plant transformation experiments were carried out using Escherichia coli DH5a and Agrobacterium tumefaciens LBA4404, respectively.
The T-DNA region of the pBISM2 construct is shown schematically. RB stands for right boundaries. Nos-T stands for nopaline synthase terminator; NptII stands for neomycin phosphotransferase II; and Nos-P stands for nopaline synthase promoter. Chimeric chitinase (chit42 gene + ChBD (chitin-binding domain)); LB stands for left border
Agrobacterium-mediated transformation of tomato
The tomato seeds were sterilized for 15 min with sodium hypochlorite 2.5% (v/v) and rinsed with sterile distilled water three times in a laminar airflow cabinet. Tomato seeds were sterilized and cultivated on the MS (Murashige and Skoog 1962) medium (0.6% agar, 3% sucrose, and 100 mg/L Myo-inositol). The 7-day-old cotyledons, excreted from germinated seedlings, were used for the transformation of tomatoes.
The pBISM2 constructs were mobilized into LBA4404 strain by the freeze–thaw procedure (Sambrook and Russell 2001). The recombinant A. tumefaciens were grown on LB agar medium complemented with 50 mg/L of Kanamycin and 25 mg/L Rifampicin. Complemented LB liquid medium (10 mL) was then inoculated with a single colony of LBA4404 and kept on shaker at 28 °C for 48 h. Following the inoculation, 500 μL of the bacterial suspension was added to 50 mL of fresh LB media, supplemented with 150 μM acetosyringone (AS) and the bacteria were grown overnight at 28 °C to raise the bacterial OD600 to 0.6. The bacterial culture was centrifuged for 15 min at 4000 rpm and finally resuspended in infection media (IM) for explant inoculation (Zhang et al. 2016).
Cotyledon explants pre-cultured on regeneration media (RM) with 2 mg/L BAP (6-Benzylaminopurine) and 0.2 mg/L IAA (Indole-3-acetic acid) for 2 days in plant growth chamber with 2500 Lux, 16/8 h (dark/day) photoperiod and 25 ± 2 °C. The explants were immersed in prepared bacterial suspension and were placed on shaker for 10 min, the surface was briefly dried on sterilized filter paper and transferred to co-cultivation media (CM) for 2 additional days at 25 °C in the dark. After co-cultivation, the explants were washed with sterile water containing 200 mg/L cefotaxime (Duchefa). The explants were put in regeneration media (RM), supplemented with 200 mg/L cefotaxime and 10 mg/L Kanamycin. After shoot initiation, shoot induced explants were transferred to shoot elongation medium (MS medium containing 0.5 mg/L of BAP, 0.2 mg/L IAA, 10 mg/L of Kanamycin, and 200 mg/L of cefotaxime). The 2–3 cm length shoots were excised and cultured in MS medium containing 0.2 mg/L IAA, 10 mg/L of Kanamycin, and 200 mg/L of cefotaxime to root induction and recover the whole plant.
Extraction of genomic DNA and polymerase chain reaction (PCR) analysis of putative transgenic plants
Genomic DNA was extracted from the leaves of putative transgenic (T0 and T1) plants according to a modified cetyl trimethyl ammonium bromide (CTAB) method (Rogers and Bendich 1985). The presence of transgenes in randomly selected some putative transgenic plants was confirmed by PCR using transgene specific primers (Table 1). The following thermal cycle program was run for DNA amplification: 5-min initial denaturation at 94 °C, followed by 40 cycles of denaturation at 94 °C for 1 min, annealing at 58 °C for minute, extension at 72 °C for 1 min, then a final extension of 10 min at 72 °C. PCR products were separated on a 1% (w/v) agarose gel.
Expression analysis
RT-PCR analysis of the transgene transcription was done for T1 plants using gene specific primers (Table 1). Total RNA was extracted from the powdered leaves of transgenic and control tomato plants using the RNX Plus kit (Cinnagen, Iran). The cDNA synthesis package (YTA) included M-MuLV reverse transcriptase and the oligo (dT) 18 primer, was used to make first strand cDNA.
To examine the relative transcript levels of transgenic tomato with defensive genes Real time PCR analyses of the gene was performed in an ABI PRISM 7500 Fast Real-Time PCR System (Applied Biosystems, CA, USA) with the program as follow: 50 °C for 2 min, 95 °C for 10 min, 45 cycles of 95 °C for 2 min, 62 °C for 30 s, and 72 °C for 30 s. To normalize of expression data, the ubiquitin gene of Solanum lycopersicum (ubi3) (NM_001346406.1) was employed as an internal reference gene. The quantification of transgene expression levels was determined by 2−ΔΔCt method (Litvak and Schmittgen 2001). All reactions were performed in three replicates, with P-values less than 0.05 considered significant.
Chitinase activity assay
Young leaves of putative transgenic lines and control tomato plants (i.e., not transformed) were frozen in liquid N2 and milled into a fine powder. The soluble proteins were extracted in 50 mM sodium acetate buffer (pH 7.0). Chitinase activity was measured in the reaction mixture (total 500 mL) comprising colloidal chitin as the substrate (3.8 mg) and the crude of transgenic plant enzymes (200 µg/mL). The reaction was performed for 1 h at 37 °C and then centrifuged at 6000 rpm for 5 min. The supernatant was boiled in 100 µL of potassium tetraborate buffer for 3 min. Three mL sample of DMAB reagent (10 g di-methyl aminobenzaldehyde in 100 mL glacial acetic acid and 10 M hydrochloric acid) was added to the reaction. The supernatant was incubated at 37 °C for 20 min, and the quantity of N-acetylglucosamine (GLcNAc) generated was determined using the technique published by Zeilinger et al. (1999), using GLcNAc as a standard. The amount of enzyme required to catalyze the release of 1 µmol GLcNAc in 60 min at 37 °C was defined as one unit of enzyme activity. Each assay was replicated three times.
Fungal growth inhibition assay
The fungus was grown at 22 °C for 24 h using a medium of potato dextrose agar (PDA) (potatoes, infusion 200 g/L, dextrose 20 g/L, and agar 15 g/L) (Ayers et al. 1976). The concentration of total soluble protein of T1 plants was quantified using the Bradford method with bovine serum albumin (BSA) as a reference (Bradford 1976). A modified radial diffusion approach was used to conduct an in vitro antifungal activity assay (Broekaert et al. 1992). The pathogenic fungi's agar discs (5 mm diam) including, S. sclerotiorum, which was produced from fungus in active growth that had previously been grown on PDA, were mounted in the center of Petri plates (100 × 15 mm) with 25 mL of PDA. Following the forming of the mycelial colony, 5 mm holes were drilled 1 cm away from the colony around the rim. To extract crude protein from transgenic plants, 50 mM Tris–HCl (pH 8.8) was employed, and the wells were filled with equal aliquots (55–60 µg). As a control, non-transgenic tomato plant crude protein was used. The plates were incubated at 28 °C for 48 h, which during this period the hyphae began to extend outwards from the center. After treatment, hyphae inhibition was monitored regularly. The following formula was used to measure the inhibition rate (%): [(the distance between the borders of the control hypha’s and the middle of the hyphal—the distance between the borders of the treated hypha’s and the center)/the distance between the borders of the control hypha's and the center] × 100%. Each sample was subjected to the assay three times.
Transgenic plant resistance bioassays
The transgenic line’s resistance bioassays were carried out using detached leaves, as stated previously (Yang et al. 2019). Briefly, S. sclerotiorum was grown on potato dextrose agar (PDA) at room temperature, and the inoculum was made up of 5-mm mycelial plugs from the active developing tip. Inoculation was carried out using young completely expanded leaves from a 4 weeks old wild-type (WT) and T1 transgenic plants. A standard expanded leaf of normal color and shape was chosen for inoculation from each plant. Each leaf’s inoculation site was located in the center of the leaf and incubated in the dark at 25 °C for detached leaf assays. 72 h after inoculation, necrotic lesions (cm2) were monitored and photographed according to Cunha et al. (2010). Every transgenic line was inoculated with three detached leaves.
Experiments and data investigation
The whole experiments were carried out with three replications in Completely Randomized Design (CRD) and One-Way ANOVA method was used to assess the significance of the variations. The analysis was performed by the SPSS (v. 17, USA) program using Duncan’s Multiple Range Test and SNK test. Significant differences were described as P values less than 0.05.
Results
Transgenic plant transformation, selection, and regeneration
The recombinant construct, assigned as pBISM2 (Fig. 1) was mobilized into Agrobacterium tumefaciens and used for transformation of 500 cotyledonary explants of tomato. Results indicate 56% of explants with shoot regeneration on a selective medium containing 10 mg/L Kanamycin, and up to 36% remained green and developed stems in selective medium (Fig. 2).
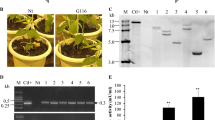
Transformation and regeneration of transgenic tomato plants. a Seven days old seedlings that have been grown at half strength-MS media. b Regenerated shoots in MS medium containing 2 mg/L BAP and 0.2 mg/L IAA without selection media. c Development of putative transgenic shoots in a selective medium. d Plantlet with the elongated shoot and induced roots. e Covered regenerated plantlets in pots. f, g Acclimated plantlet in the greenhouse. h Tomato flowers and green fruit. i Tomato fruit. (Color figure online)
Stable integration of transgene was confirmed in randomly selected putative transgenic plants using gene specific primers (Table 1). Amplification of a 900-bp fragment of chimeric chit42 was shown in 22 putative transgenic lines (Fig. 3a). A set of virG primers (Table 1) were employed to detect Agrobacterium contamination of the putative plants. Vir gene fragment was amplified only in Agrobacterium genomic DNA supplemented reaction (data not shown). Existence of the Chitin binding domain (ChBD) (225 bp) was confirmed using transgenic specific primers (Fig. 3b).
Molecular detection of putative transgenic tomato. a PCR analysis of transgenic plants to determine the chitinase gene. b PCR analysis of transgenic plants to determine chimeric chitinase (ChBD) gene. c RT-PCR of transgenic plants to determine the expression of the chitinase gene. M: 1 kb DNA ladder, line 1: positive control (pBISM2), line 2: wild type plant, line 3: negative control (Deionized Water), lines 4–9: transgenic plants
The transformation efficiency was estimated to be 20%. All transgenic plants had a typical Lycopersicon esculentum L. phenotype in terms of morphology and growth characteristics compared to non-transformed plants. A total of 40 elongated shoots were successfully rooted in kanamycin-containing root induction medium, followed by a transplantation to soil after a brief hardening period and allowed for blossoming and seedling in the greenhouse (Fig. 2d–i).
Expression analysis of the chimeric chit42
RT-PCR was used to validate the chimeric chit42 transcripts in randomly selected five PCR-positive transgenic plants. RT-PCR analysis confirmed the presence of the specific transcription of chit42 in T1 transgenic tomatoes plants (Fig. 3C). A single 900 bp sharp fragment was detected in all evaluated samples, similar to the positive control (pBISM2). There were no amplified products in RT-PCR reactions in the negative control assays when wild-type DNA was used as template (data not shown). Different levels of expression of the chimeric chit42 gene were detected using RT-PCR in distinct transgenic strains. (Fig. 4). All transgenic lines showed a significant (P < 0.05) increase in expression of chimeric chit42 chitinase gene when compared to control with T10 showing the highest increase expression.
Chitinase activity assay
The chimeric chitinase activity was assayed using colloidal chitin as a substrate with crude protein extract from leaf tissues of PCR positive transgenic plants. The specific enzyme activity of different transgenic plants varied from 4.79 ± 0.22 to 6.02 ± 0.38 U/μg (Fig. 5). All five transgenic lines that showed different expression patterns of the transgene were evaluated for enzyme activity and result showed that the T10 (6.02 ± 0.38 U/μg) had the highest activity flowed by T14 (5.82 ± 0.36 U/μg), T22 (5.42 ± 0.36 U/μg), T11 (4.98 ± 0.74 U/μg) and T15 (4.79 ± 0.22 U/μg).
Chitinase activity in leaf tissues of transgenic tomato lines (T) and untransformed control plant (WT). One unit of activity is defined as the enzyme activity catalyzing the formation of one µmol of N-acetylglucosamine h−1 (µg protein)−1. Difference was significance at P < 0.05 using Duncan’s multiple range tests. Results represent the average and standard deviation of three biological replicates
Radial diffusion assay for antifungal activity
A radial diffusion assay was used to investigate the antifungal activity against the growth of Sclerotiorum sclerotiorum in the chosen five transgenic plants. Crude protein extracts from all transgenic lines significantly inhibited fungus hypha development compared to the wild type non-transgenic plants (Fig. 6). Similar to the expression analysis where T10 showed the highest expression level, radial diffusion assay also showed less redial extension.
a Radial diffusion assay of transgenic tomato crude protein extracts against S. sclerotiorum. T10, T14, T22, T11 and T15, the crude proteins from transgenic lines with high chitinase expression. (WT), the crude of proteins from wild-type negative control. b Growth inhibition zone (mm2) of S. sclerotiorum by crude extracts of transgenic and non-transgenic plants. Values represent the mean ± SD according to Duncan’s multiple range test (P < 0.05) of three replications
Resistance assay to S. sclerotiorum in greenhouse
To determine the severity of disease caused by S. sclerotiorum infection leaf inoculation assay was used. The lesion region of the bacteria on the leaves was measured. The results revealed a significant difference in fungus lesion diameter between transgenic lines and untransformed plants. The lesion on the non-transgenic leaves progressed rapidly 72 h after inoculation with S. sclerotiorum but lesion extended slowly and lesion sizes was significantly smaller in transgenic lines. Consistent with the other results, T10 showed the smallest lesion (Fig. 7). In detached leaf experiments, consistent results showed that (ChBD) gene expression increased transgenic tomato resistance to S. sclerotiorum (Table 2).
Discussion
Tomatoes are an important global commercial crop used both in the fresh fruit market and in the processed food industry. The phytopathogenic fungus can damage the vegetative and reproductive parts of the plants as well as affect tomato growth and subsequently it’s yield. Plant genetic manipulation is an effective strategy in plant protection. Chemical control is replaced by biological control, which is the most environmentally friendly option (Abbas et al. 2018). Several species of Trichoderma have been investigated extensively against a variety of soil and foliar diseases, including S. sclerotiorum, R. cinerea solani, Pythium spp., and B. fusarium spp. (Olowe et al. 2022). Previously, we have shown that overexpression of chimeric chitinase gene in transgenic plants induce systemic resistance against pathogens by affecting salicylic acid and jasmonic acid signaling pathways (Eslahi et al. 2021). Here, we have introduced chimeric chitinase of Trichoderma atroviride with chitin-binding domain and have shown its resistance to Lycopersicon esculentum L.
Chitinase is an essential enzyme present widely in plants, and it plays a role in physiological processes as plant growth, development, and immunity (Zhang et al. 2018). Transformation of some important crops with chitinase improved resistance against fungal pathogen infection (Girhepuje and Shinde 2011; Khan et al. 2014; Karmakar et al. 2016; Dowd et al. 2018; Tue et al. 2022). Overexpression of a Chitinase gene from Indica rice has enhanced rice resistance to Rhizoctonia solani (Richa et al. 2017) and introducing the chimeric chit42 from Trichoderma harzianum to carrots reduced the severity of carrot rot caused by three isolates of Sclerotiorum, according to Ojaghian et al. (2018). To date, there has been no study on the effect of introducing chimeric chitinase genes into tomatoes on S. sclerotiorum. Given the necessity of developing new Trichoderma strains with improved antifungal activity for agricultural use, Matroodi et al. (2013) used ChBD to create Chit42 with a higher chitinase activity. The ChBD binds to insoluble chitin-like fungal cell walls and boosts chitinase activity (Eslahi et al. 2021). The findings of Wang et al. (2021) showed that overexpression of MdCHI1 (Malus × domestica), which includes the chitin binding domain and glycoside hydrolase domain, increase resistant of apples to Colletotrichum gloeosporioides and Alternaria alternata.
Following the creation of chimeric chitinase of Trichoderma atroviride, we detected the relative transcript levels of PCR-positive plants in five transgenic lines, and various levels of expression of the chimeric chit42 gene were measured in distinct transgenic strains. Moreover, the antifungal efficacy of chimeric chit42 extracted protein from transgenic lines was evaluated using a radial diffusion assay. The generation of five T1 events was selected for subsequent analysis. The homozygous plants were challenged with S. sclerotiorum in the greenhouse assay. Five transgenic lines of tomato were evaluated for resistance against Sclerotinia sclerotiorum, which causes tomato white mold, one of the deadliest diseases affecting tomato yield (Mazumdar 2021). Sclerotinia sclerotiorum infection behavior showed that none of the chimeric chitinase expressing transgenic lines nor the wilt type inhibited fungal penetration but, the fungal infection symptoms (necrosis) on the leaves was limited in the transgenic plants (T10 and T14) but not in non-transgenic plant leaf. The chimeric chitinase used in this work had a chitin-binding domain with higher chitin-binding capacity, which improved chitinase activity and inhibited fungus growth more effectively. Hou et al. (2019) investigated the influence of ChBD on chitin binding and showed that removing ChBD from chitinase reduces the efficiency of chitin degradation significantly. According to Zarinpanjeh et al. (2016), the chimeric chitinase’s antifungal activity was more efficient against chitin than the wild-type enzyme, particularly in its crystalline form. The conclude that Chitin binding characteristics of the chimeric chitinase were significantly higher (approximately twofold) than Chit42, owing to the inclusion of the ChBD, this appears to be due to the subsite structure in this enzyme's binding cleft. In-vitro and in-vivo studies of T. harzianum recombinant strains (containing chimeric chitinase) and wild-type strains against R. solani revealed that recombinant strains had higher biocontrol activity and contributed to plant health (Eslahi et al. 2021).
Compared to non-transformed plants, the results indicated that constitutive chitinase gene expression might be sufficient to prevent S. sclerotiorum growth. The earlier reports of chitinase gene overexpressed in transgenic plants showed enhanced resistance against soil borne and foliar fungal pathogens. Several in-vitro inhibition towards Sclerotinia sclerotiorum, Sclerotium rolfsii, Fusarium oxysporum, revealed that chitinases could inhibit a wide range of fungal infections (Zarinpanjeh et al. 2016; Loc et al. 2020; Sharma et al. 2020; Santoso et al. 2022). Overexpression of chit42 gene in transgenic tomatoes increased resistance of transgenic plants to the fungal disease compared to wild-type plants. Consistent with these reports, the preliminary study with this gene reveals the overexpression of chimeric chitinase gene in B. napus improved resistance against Sclerotinia sclerotiorum and disease symptoms was significant in B. napus control plants than transgenic lines. There have been numerous instances of transgenic plants producing chitinases alone or combined with other proteins, expressing chitinases enhancing resistance to fungal pathogens in greenhouse studies (Huang et al. 2013; Zarinpanjeh et al. 2016; Yang et al. 2020). Enzyme activity of the chitinase in transgenic extract were different in evaluated transgenic plants. Chititinas activity of the extract from T10 and T14 lines were significantly higher than others accordingly to the expression pattern of the gene (Fig. 4).
Chitinases have a double function: they limit fungal development by digesting cell walls and they release pathogen-borne elicitors, which cause the host to respond with more defense responses (Dana et al. 2006). Overexpressing a chitinase gene in plants improved pathogen resistance, likely accompanied by the activation of other defense-related systems due to their increased activity to degrade chitin.
Conclusion
The present work has demonstrated that expression of chimeric chitinase successfully increased resistance of transgenic plants to S. sclerotiorum. These transgenic lines show promise for developing a variety of tomato lines with improved resistance to S. sclerotiorum, as well as serving as a source of germplasm for enhanced resistance to other critical fungal infections. This transgenic line can also be manipulated with other resistance gene candidates to accumulate the resistance to fungal disease.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Abbas T, Zahir ZA, Naveed M, Kremer RJ (2018) Limitations of existing weed control practices necessitate development of alternative techniques based on biological approaches. Adv Agron 147:239–280. https://doi.org/10.1016/bs.agron.2017.10.005
Aghazadeh R, Zamani M, Motallebi M, Moradyar M (2016) Co-transformation of canola by chimeric chitinase and tlp genes towards improving resistance to Sclerotinia sclerotiorum. World J Microbiol Biotechnol 32(9):144. https://doi.org/10.1007/s11274-016-2104-6
Ayers AR, Ebel J, Valent B, Albersheim P (1976) Host–pathogen interactions: X. Fractionation and biological activity of an elicitor isolated from the mycelial walls of Phytophthora megasperma var. sojae. Plant Physiol 57:760–765. https://doi.org/10.1104/pp.57.5.760
Badrhadad A, Nazarian F, Ismaili A (2018) Fusion of a chitin-binding domain to an antibacterial peptide to enhance resistance to Fusarium solani in tobacco (Nicotiana tabacum). 3 Biotech 8:391. https://doi.org/10.1007/s13205-018-1416-7
Berini F, Casartelli M, Montali A, Reguzzoni M, Tettamanti G, Marinelli F (2019) Metagenome-sourced microbial chitinases as potential insecticide proteins. Front Microbiol 18(10):1358. https://doi.org/10.3389/fmicb.2019.01358
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Broekaert WF, Marien W, Terras F, Bolle M, Proost P, Damme J, Dillen L, Claeys M, Rees S (1992) Antimicrobial peptides from Amaranthus caudatus seeds with sequence homology to the cysteine/glycinerich domain of chitin-binding proteins. Biochemistry 31:4308–4314. https://doi.org/10.1021/bi00132a023
Chanda B, Shamimuzzaman M, Gilliard A, Ling KS (2021) Effectiveness of disinfectants against the spread of tobamoviruses: Tomato brown rugose fruit virus and Cucumber green mottle mosaic virus. Virol J 18(1):1–2. https://doi.org/10.1186/s12985-020-01479-8
Chhikara S, Chaudhury D, Dhankher OP, Jaiwal PK (2012) Combined expression of a barley class II chitinase and type I ribosome inactivating protein in transgenic Brassica juncea provides protection against Alternaria brassicae. Plant Cell Tissue Organ Cult 108:83–89. https://doi.org/10.1007/s11240-011-0015-7
Cunha WG, Tinoco MLP, Pancoti HL, Ribeiro RE, Aragão FJL (2010) High resistance to Sclerotinia sclerotiorum in transgenic soybean plants transformed to express an oxalate decarboxylase gene. Plant Pathol 59:654–660. https://doi.org/10.1111/j.1365-3059.2010.02279.x
da Silva Santos SD, da Silva AA, Polonio JC, Polli AD, Orlandelli RC, dos Santos Oliveira JA, Brandão Filho JU, Azevedo JL, Pamphile JA (2022) Influence of plant growth-promoting endophytes Colletotrichum siamense and Diaporthe masirevici on tomato plants (Lycopersicon esculentum Mill.). Mycology 18:1–4. https://doi.org/10.1080/21501203.2022.2050825
Dana M, Pintor-Toro T, Cubero B (2006) Transgenic tobacco plants overexpressing chitinases of fungal origin show enhanced resistance to biotic and abiotic stress agents. Plant Physiol 142:722–730. https://doi.org/10.1104/pp.106.086140
Dowd PF, Naumann TA, Price NP, Johnson ET (2018) Identification of a maize (Zea mays) chitinase allele sequence suitable for a role in ear rot fungal resistance. Agri Gene 7:15–22. https://doi.org/10.1016/j.aggene.2017.10.001
El-Sobki AE, Ali AA (2020) Biochemical effects of some chitin synthesis inhibitors against red palm weevil, Rhynchophorus ferrugineus insect. Egypt Acad J Biol Sci F Toxicol Pest Control 12(1):127–39. https://doi.org/10.21608/eajbsf.2020.83542
Eslahi N, Kowsari M, Zamania MR, Motallebia M (2021) The profile change of defense pathways in Phaseouls vulgaris L. by biochemical and molecular interactions of Trichoderma harzianum transformants overexpressing a chimeric chitinase. Biol Control 152:104304. https://doi.org/10.1016/j.biocontrol.2020.104304
Ghorbanpour M, Omidvari M, Abbaszadeh-Dahaji P, Omidvar R, Kariman K (2018) Mechanisms underlying the protective effects of beneficial fungi against plant diseases. Biol Control 117:147–157. https://doi.org/10.1016/j.biocontrol.2017.11.006
Girhepuje PV, Shinde GB (2011) Transgenic tomato plants expressing a wheat endochitinase gene demonstrate enhanced resistance to Fusarium oxysporum f. sp. lycopersici. Plant Cell Tissue Organ Cult 105:243–251. https://doi.org/10.1007/s11240-010-9859-5
Hou J, Li X, Kaczmarek MB, Chen P, Li K, Jin P, Liang Y, Daroch M (2019) Accelerated CO2 hydration with thermostable sulfurihydrogenibium azorense carbonic anhydrase-chitin binding domain fusion protein immobilised on chitin support. Int J Mol Sci 20(6):1494. https://doi.org/10.3390/ijms20061494
Huang JK, Ma PL, Ji SY, Zhao XL, Tan JX, Sun XJ, Huang FD (2013) Age-dependent alterations in the presynaptic active zone in a Drosophila model of Alzheimer’s disease. Neurobiol Dis 51:161–167. https://doi.org/10.1016/j.nbd.2012.11.006
Jambhulkar PP, Sharma P, Manokaran R, Lakshman DK, Rokadia P, Jambhulkar N (2018) Assessing synergism of combined applications of Trichoderma harzianum and Pseudomonas fluorescens to control blast and bacterial leaf blight of rice. Eur J Plant Pathol 152(3):747–757. https://doi.org/10.1007/s10658-018-1519-3
Karmakar S, Molla KA, Chanda PK, Sarkar SN, Datta SK, Datta K (2016) Green tissue-specific co-expression of chitinase and oxalate oxidase 4 genes in rice for enhanced resistance against sheath blight. Planta 243(1):115–130. https://doi.org/10.1007/s00425-015-2398-x
Khan RS, Darwish NA, Khattak B, Ntui VO, Kong K, Shimomae K, Nakamura I, Mii M (2014) Retransformation of marker-free potato for enhanced resistance against fungal pathogens by pyramiding chitinase and wasabi defensin genes. Mol Biotechnol 56(9):814–823. https://doi.org/10.1007/s12033-014-9760-2
Kowsari M, Zamani MR, Motallebi M (2014a) Enhancement of Trichoderma harzianum activity against Sclerotinia sclerotiorum by overexpression of Chit42. Iran J Biotechnol 12(2):e13869
Kowsari M, Motallebi M, Zamani MR (2014b) Protein engineering of chit42 towards improvement of chitinase and antifungal activities. Curr Microbiol 68:495–502. https://doi.org/10.1007/s00284-013-0494-3
Li YH, Zhang XY, Zhang F, Peng LC, Zhang DB, Kondo A, Zhao XQ (2018) Optimization of cellulolytic enzyme components through engineering Trichoderma reesei and on-site fermentation using the soluble inducer for cellulosic ethanol production from corn stover. Biotechnol Biofuels 11:49. https://doi.org/10.1186/s13068-018-1048-5
Litvak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Loc NH, Huy ND, Quang HT, Lan TT, Ha TT (2020) Characterisation and antifungal activity of extracellular chitinase from a biocontrol fungus, Trichoderma asperellum PQ34. Mycology 11(1):38–48. https://doi.org/10.1080/21501203.2019.1703839
Malik MS, Haider S, Rehman A, Rehman SU, Jamil M, Naz I, Anees M (2022) Biological control of fungal pathogens of tomato (Lycopersicon esculentum) by chitinolytic bacterial strains. J Basic Microbiol 62(1):48–62. https://doi.org/10.1002/jobm.202100512
Marra R, Lombardi N, Piccolo A, Bazghaleh N, Prashar P, Vandenberg A, Woo S (2021) Mineral biofortification and growth stimulation of lentil plants inoculated with Trichoderma strains and metabolites. Microorganisms 10(1):87. https://doi.org/10.3390/microorganisms10010087
Matroodi S, Motallebi M, Zamani MR, Moradyar M (2013) Designing a new chitinase with more chitin binding and antifungal activity. World J Microbiol Biotechnol 29:1517–1523. https://doi.org/10.1007/s11274-013-1318-0
Mazumdar P (2021) Sclerotinia stem rot in tomato: a review on biology, pathogenicity, disease management and future research priorities. J Plant Dis Prot 128(6):1403–1431. https://doi.org/10.1007/s41348-021-00509-z
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Nalluri N, Karri VR (2020) Recent advances in genetic manipulation of crops: a promising approach to address the global food and industrial applications. Plant Sci Today 7(1):70–92. https://doi.org/10.14719/pst.2020.7.1.659
Nuwamanya AM, Runo S, Mwangi M (2022) In-vitro sensitivity of Alternaria solani isolates to azoxystrobin and difenoconazole fungicides in Kenya and detection of Cyt b mutations associated with azoxystrobin resistance. Crop Prot 30:106010. https://doi.org/10.1016/j.cropro.2022.106010
Ojaghian S, Wang L, Xie GL, Zhang JZ (2018) Increased resistance against storage rot in transgenic carrots expressing chitinase chit42 from Trichoderma harzianum. Sci Hortic 234:81–86. https://doi.org/10.1016/j.scienta.2018.02.025
Olowe OM, Nicola L, Asemoloye MD, Akanmu AO, Babalola OO (2022) Trichoderma: Potential bio-resource for the management of tomato root rot diseases in Africa. Microbiol Res 2:126978. https://doi.org/10.1016/j.micres.2022.126978
Richa K, Tiwari IM, Devanna BN, Botella JR, Sharma V, Sharma TR (2017) Novel CHITINASE Gene LOC_Os11g47510 from indica rice tetep provides enhanced resistance against sheath blight pathogen Rhizoctonia solani in rice. Front Plant Sci 8:596. https://doi.org/10.3389/fpls.2017.00596
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76. https://doi.org/10.1007/BF00020088
Salas CE, Badillo-Corona JA, Ramírez-Sotelo G, Oliver-Salvador C (2015) Biologically active and antimicrobial peptides from plants. Biomed Res Int 2015:1–11. https://doi.org/10.1155/2015/102129
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Santoso P, Minamihata K, Ishimine Y, Taniguchi H, Komada T, Sato R, Goto M, Takashima T, Taira T, Kamiya N (2022) Enhancement of the antifungal activity of chitinase by palmitoylation and the synergy of palmitoylated chitinase with amphotericin B. ACS Infect Dis. https://doi.org/10.1021/acsinfecdis.2c00052
Sharma S, Kumar S, Khajuria A, Ohri P, Kaur R, Kaur R (2020) Biocontrol potential of chitinases produced by newly isolated Chitinophaga sp. S167. World J Microbiol Biotechnol 36(6):1–5. https://doi.org/10.1007/s11274-020-02864-9
Solgi T, Moradyar M, Zamani MR, Motallebi M (2015) Transformation of canola by chit33 gene towards improving resistance to Sclerotinia sclerotiorum. Plant Prot Sci 51(1):6–12. https://doi.org/10.17221/83/2013-PPS
Tien NQ, Hoa PT, Tue NH, Thanh DV, Thi HA, Luong NN, Huy NX, Loc NH (2021) Transient expression of chi42 genes from Trichoderma asperellum in Nicotiana benthamiana by agroinfiltration. Int J Agric Biol 26:177–184. https://doi.org/10.17957/IJAB/15.1822
Tue NH, Tuong TG, Trang PT, Chung ND, Hoa PT, Tien NQ, Loc NH (2022) Cloning the root-specific Asy promoter and genes encoding chitinase 42 kDa of Trichoderma asperellum into the plant expression vector. J Appl Biol Biotechnol. https://doi.org/10.7324/JABB.2022.100302
Vaghela B, Vashi R, Rajput K, Joshi R (2022) Plant chitinases and their role in plant defense—a comprehensive review. Enzym Microb Technol 30:110055. https://doi.org/10.1016/j.enzmictec.2022.110055
Wang Z, Ma LM, Cao J, Li YL, Ding LN, Zhu KM, Yang YH, Tan ZL (2019) Recent advances in mechanisms of plant defense to Sclerotinia sclerotiorum. Front Plant Sci 10:1314. https://doi.org/10.3389/fpls.2019.01314
Wang F, Yang S, Wang Y, Zhang B, Zhang F, Xue H, Jiang Q, Ma Y (2021) Overexpression of chitinase gene enhances resistance to Colletotrichum gloeosporioides and Alternaria alternata in apple (Malus × domestica). Sci Hortic 277:109779. https://doi.org/10.1016/j.scienta.2020.109779
Xia H, Li YY, Liu ZC, Li YQ, Chen J (2018) Transgenic expression of chit42 gene from Metarhizium anisopliae in Trichoderma harzianum enhances antagonistic activity against Botrytis cinerea. Mol Biol 52(5):668–675. https://doi.org/10.1134/S002689331805014X
Yang X, Yang J, Wang Y, He H, Niu L, Guo D, Xing G, Zhao Q, Zhong X, Sui L, Li Q, Dong Y (2019) Enhanced resistance to sclerotinia stem rot in transgenic soybean that overexpresses a wheat oxalate oxidase. Transgenic Res 28(1):103–114. https://doi.org/10.1007/s11248-018-0106-x
Yang X, Yang J, Li H, LuNiu L, Xing G, Zhang Y, Xu W, Zhao Q, Li Q, Dong Y (2020) Overexpression of the chitinase gene CmCH1 from Coniothyrium minitans renders enhanced resistance to Sclerotinia sclerotiorum in soybean. Transgenic Res 29:187–198. https://doi.org/10.1007/s11248-020-00190-2
Zarinpanjeh N, Motallebi M, Zamani MR, Ziaie M (2016) Enhanced resistance to Sclerotinia sclerotiorum in Brassica napus by co-expression of defensin and chimeric chitinase genes. J Appl Genet 57:417–425. https://doi.org/10.1007/s13353-016-0340-y
Zeilinger S, Galhaup C, Payer K, Woo SL, Mach RL, Fekete C, Lorito M, Kubicek CP (1999) Chitinase gene expression during mycoparasitic interaction of Trichoderma harzianum with its host. Fungal Genet Biol 26:131–140. https://doi.org/10.1006/fgbi.1998.1111
Zhang F, Ge H, Zhang F, Guo N, Wang Y, Chen L, Li C (2016) Biocontrol potential of Trichoderma harzianum isolate T-aloe against Sclerotinia sclerotiorum in soybean. Plant Physiol Biochem 100:64–74. https://doi.org/10.1016/j.plaphy.2015.12.017
Zhang WY, Zhang SY, Chen XH, Qiao H, Jiang SF, Xiong YW, Jin SB, Gong YS, Fu HT (2018) Molecular cloning and expression analysis of MnCht1C gene from oriental river pawn (Macrobrachium nipponense). Genomics Appl Biol 37(2):723–732
Zhang Y, Chen W, Sang X, Wang T, Gong H, Zhao Y, Zhao P, Wang H (2021) Genome-wide identification of the thaumatin-like protein family genes in Gossypium barbadense and analysis of their responses to Verticillium dahliae infection. Plants 10(12):2647. https://doi.org/10.3390/plants10122647
Zhu M, Lu S, Zhuang M, Zhang Y, Lv H, Ji J, Hou X, Fang Z, Wang Y, Yang L (2021) Genome-wide identification and expression analysis of the Brassica oleracea L. chitin-binding genes and response to pathogens infections. Planta 253:80. https://doi.org/10.1007/s00425-021-03596-2
Acknowledgements
The authors thank to Dr. Mostafa Motallebi from National Institute of Genetic Engineering and Biotechnology (NIGEB) for provide the chimeric chitinase construct and the Mycology laboratory, Plant protection department of Tabriz University to provide the fungi and support us for the bioassay experiments.
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
Lab work have been done by VG by adviser of ED. ED conceived and designed the experiment. MZ help us in all bioassay analysis test and give the good advises in writing the manuscript. VG wrote the manuscript with support of ED. MV give the good and benefit advice in lab techniques and revised the manuscript finally. All author critical feedback and helped shape the research, analysis and manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that there is no conflict of interest.
Additional information
Communicated by Joyce Van Eck.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gougerdchi, V., Dorani, E., Valizadeh, M. et al. Overexpression of the chimeric chitinase (ChBD) gene in Lycopersicon esculentum L. enhanced resistance to Sclerotinia sclerotiorum. Plant Cell Tiss Organ Cult 151, 165–175 (2022). https://doi.org/10.1007/s11240-022-02340-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-022-02340-2