Abstract
A chitinolytic bacterium Chitinophaga sp. S167 producing extracellular chitinases was isolated from a soil sample in India. The extracellular chitinases produced by S167 were concentrated by ammonium sulphate precipitation (AS70) and seven bands corresponding to chitinases were observed by zymography. Optimum temperature and pH of AS70 were between 40 and 45 °C and pH 6.0 respectively with high stability at 20–40 °C and pH 5–7. AS70 inhibited the growth of Fusarium oxysporum, Alternaria alternata and Cladosporium sp. in vitro. The culture conditions for the high level production of extracellular chitinases were optimized resulting in 48-folds higher chitinase production. As the combination of chitinases could be more potent in biocontrol of plant diseases, it was checked if AS70 could control postharvest fungal infection caused by Fusarium oxysporum on tomatoes. AS70 treated tomatoes showed significant lower incidence of infection (11%) by F. oxysporum as compared with 100% in the control at 5 days post inoculation. Further, AS70 caused significant mortality in second stage juveniles of root knot nematode, Meloidogyne incognita, a major agriculture pest responsible for economic losses in agriculture. This study highlights the antifungal and nematicidal activity of chitinases produced by Chitinophaga sp. S167. To the best of our knowledge, this is the first report of the biocontrol potential of the chitinases produced by Chitinophaga sp.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Chitinophaga, named for the ability to degrade chitin was originally described by Sangkhobol and Skerman (1981). Presently, it comprises of more than 30 species that have been isolated from a variety of habitats such as soil, roots, vermicompost, bark and rocks (Kim et al. 2019). C. pinensis, the type strain of the genus Chitinophaga, secretes biotechnologically relevant carbohydrate active enzymes, some of which are as yet uncharacterized (Del Rio et al. 2010; Larsbrink et al. 2017).
Chitinases (EC 3.2.1.14) are involved in catalysis of β-1 → 4 linkages in chitin and belong to glycoside hydrolase (GH) families 18, 19 or 48 (Henrissat and Bairoch 1993; Seidl 2008). Based on their modes of action, chitinases are classified as endochitinases, that cleave chitin polymers at random sites and exochitinases progressively cleave chitin from non-reducing end of the chitin chain, releasing N-acetyl d-glucosamine (NAG) monomers and diacetyl chitobiose in the process (Horn et al. 2006). Another chitinolytic enzyme, chitobiase, converts NAG dimers to NAG monomers.
Chitin is the second abundant biopolymer on earth, and is present in abundance in marine and terrestrial habitats. It is an important structural component of cell walls in fungi, whereas in nematodes, it provides structural support to eggshell and pharyngeal lining (Hartl et al. 2012; Heustis et al. 2012). Due to the lack of chitin in plants and vertebrates, phytopathogenic fungi and phytophagous nematodes can be targeted by chitinases, without any adverse effect on the environment, vertebrates and plants. Therefore, there is an emphasis on replacing agrochemicals with the microorganisms as biocontrol agents in agriculture. In view of the environmental problems caused by chemicals, there is an enhanced interest in finding new strains of microorganisms to effectively combat the problems of plant pathogenic fungi and nematodes to increase agriculture productivity. The occurrence of chitinases is widespread among bacteria where multiple chitinolytic enzymes produced by an individual strain is a common occurrence (Fuchs et al. 1986; Romaguera et al. 1992; Saito et al. 1999; Tsujibo et al. 2003).
Several studies have highlighted the preponderance of the genus Chitinophaga in natural habitats that abound in phytopathogenic fungi and nematodes. C. pinensis effectively degrades fungal cell walls and utilizes fungal biomass as nutrient source for its growth (McKee et al. 2019). The genus Chitinophaga was found to be enriched in the microbial community associated with the decomposition of fungal mycelia in litter and soil samples (Brabcová et al. 2016; Mehmood et al. 2020). Recently, a study showed that Chitinophaga spp. colonizing the rhizosphere and endosphere of the roots of sugar beet exerted protection against Rhizooctonia solani invasion by secreting chitinases (Carrión et al., 2019). Another study implicated the presence of Chitinophaga in soyabean root associated bacterial community in conferring protection against soyabean cyst nematode (Hu et al. 2019). In spite of the studies which implicate the strongly chitinolytic genus Chitinophaga in suppression of fungal and nematode infections in plants, it has received less attention as a potential biocontrol agent. Chitinases from several yeast, fungi and bacteria have been extensively studied as biocontrol candidates, but to our knowledge, there are no reports on the biocontrol efficacy of chitinases produced by Chitinophaga spp. (Dukare et al. 2019; Subbanna et al. 2018; Carmona-Hernandez et al. 2019).
The annotation of the genome sequence of C. pinensis DSM 2588 revealed that some chitinases encoded in the genome had distinct domain architectures (Del Rio et al. 2010). Recently, GH18 domain of carbohydrate active protein of C. pinensis has been shown to be a potent exo-chitinase with immense potential in bioconversion of chitin waste (Ramakrishna et al. 2018). The present study aimed to determine the potential of the chitinases secreted by the soil isolate, Chitinophaga sp. S167, as a biocontrol agent against fungal pathogens and a nematode pest of the cultivated plants. The potential of the extracellular chitinases produced by S167 as biocontrol agent of postharvest fungal disease caused by F. oxysporum in tomatoes was explored. The effect of S167 chitinases on the second stage juveniles of root node nematode, Meloidogyne incognita, the most destructive pest in agriculture, was also evaluated.
Materials and methods
Sample collection and isolation of bacteria
Soil sample, collected from Guru Nanak Dev University at Amritsar, Punjab, India was air-dried and stored at a cool and dark place. For isolation of the bacteria, soil sample treated with cycloheximide (50 μg/mL) was placed on the paste of E. coli cells laid on the water-clerigel medium (0.1% CaCl2·2H2O, 20 mM HEPES buffer, 0.8% clerigel and 50 μg/mL cycloheximide; pH 7.2) at the temperature ranging from 20 to 32 °C (Reichenbach and Dworkin 1992). The bacteria were picked up from the swarms emerging on the surface of the plate and inoculated on CY medium (0.3% casitone, 0.1% yeast extract, 0.1% CaCl2·2H2O; pH 7.2) containing 1.5% agar powder. The isolated bacteria were purified by repeatedly picking up the cells from the edge of the swarms. The isolates were maintained on CY medium for further experiments.
Screening for chitinase activity
Swollen chitin (SWC) and glycol chitin were prepared by the method of Monreal and Resse (1969) and Trudel and Asselin (1989) respectively. The screening of bacteria with chitinase activity was carried out on glycol chitin agar (0.01% glycol chitin, 1.5% agar) and SWC agar (0.5% SWC, 1.5% agar) with chitin as the sole carbon source. The cells were spotted on the plates and incubated at 30 °C to check if the halos appeared around the isolated bacteria. Further, the isolated strains were cultivated in CY broth supplemented with 0.5% SWC and the chitinolytic activity of the cell free culture supernatant (CS) was determined on the soluble chitinase substrate (0.01% glycol chitin) and 0.5% SWC incorporated in 1% agarose gel. CS was added in the wells punched in the gel and the zone of clearance was observed after incubating the plates at 30 °C for 72 h after staining with 0.1% Ranipal.
Identification of the isolate and phylogenetic analysis
The isolated bacteria were grown on CY agar medium for 48 h at 30 °C. The cells were scrapped from the surface of the plate, washed with PBS and genomic DNA was extracted using HiYield Genomic DNA Mini Kit (Real Biotech Corporation, Taiwan).
The isolate was identified on the basis of 16S rRNA gene sequence. A set of universal 16S rDNA forward and reverse primers i.e., 8F (5′-AGA GTT TGA TCC TGG CTC AG-3′) and 1492R (5′-GGT TAC CTT GTT ACG ACT T-3′) respectively were used to amplify 16S rRNA gene using genomic DNA as a template. PCR reaction mix (50 µL) consisted of 5 µL 10X Taq reaction buffer, 0.2 µM of each primer, 200 µM dNTPs (New England Biolabs, United States), 20 ng of genomic DNA and 1.5 U Taq DNA polymerase (GeNei, India). The conditions used for PCR amplification were: initial denaturation for 5 min at 95 °C, followed by (i) 5 cycles of denaturation at 95 °C for 45 s, annealing at 45 °C for 45 s, and primer extension at 72 °C for 45 s, then (ii) 5 cycles of denaturation at 95 °C for 45 s, annealing at 50 °C for 45 s, and primer extension at 72 °C for 45 s, followed by (iii) 20 cycles of denaturation at 95 °C for 45 s, annealing at 55 °C for 45 s, and primer extension at 72 °C for 45 s, (iv) final extension step at 72 °C for 10 min. The amplified 16S rRNA gene was resolved on 1.2% agarose gel and purified from the gel using HiYield Gel/PCR DNA Kit (Real Biotech Corporation, Taiwan).
The amplicon of 16S rRNA gene was sequenced at Bioserve Biotechnology Pvt. Ltd., Hyderabad, India. The sequence was submitted to GenBank (www.ncbi.nlm.nih.gov/genbank/) and the accession number was obtained. The bacterial strain was identified on the basis of 16S rRNA gene sequence similarity using the EzTaxon server (https://www.ezbiocloud.net/eztaxon/identify). Phylogenetic analysis based on 16S rRNA sequence of the isolated strain and the sequences of the strain obtained from GenBank database was carried out using Mega 7 software package employing Neighbour-Joining (N-J) method with 1000 bootstrap replicates as statistically significant (Felsenstein 1985; Saitou and Nei 1987; Kumar et al. 2016).
Chitinase assay
CY medium supplemented with 0.5% SWC was used for chitinase production at the shake flask level. After 3 days of growth, CS was obtained by centrifugation of broth at 8000 rpm for 10 min. Chitinase activity was assayed by p-dimethylaminobenzaldehyde (DMAB) method (Reissig et al. 1955). The reaction mix containing 0.5 mL of 1% SWC and 0.5 mL CS was incubated at 40 °C for 1 h followed by centrifugation at 5000 rpm for 10 min. To 500 µL of the supernatant, 100 µL of 0.8 M potassium-tetraborate was added and the reaction mix was placed in the boiling water bath for 3 min. After cooling, 3 mL of diluted DMAB [10 g of DMAB dissolved in 100 mL of acetic acid which contains 12.5% (v/v) 10 N HCl] reagent was added to the reaction mix and incubated at 37 °C for 20 min for color development. Absorbance was measured at 545 nm. Chitinase activity was determined using N-acetylglucosamine as standard. One unit of chitinase activity was defined as the amount of enzyme which liberated 1 µmole of N-acetylglucosamine per second.
Characterization of chitinase
SDS-PAGE and zymogram analysis of chitinase
S167 was grown in CY medium supplemented with 0.5% SWC at 30 °C for 72 h. The enzyme extract was prepared by adding ammonium sulphate to 70% saturation in the CS with stirring. The mixture was incubated for 4 h at 4 °C and centrifuged at 10,000 rpm for 30 min at 4 °C. The resulting precipitate was dissolved in a minimum amount of 25 mM Tris–HCl buffer at pH 8.0 and dialysed against the same buffer for 24 h at 4 °C. The protein concentration of the enzyme extract (AS70) thus obtained was determined by Bradford method with bovine serum albumin (Sigma) as the standard. AS70 was visualized by Blue silver stain after separation by 12% sodium dodecyl sulphate–polyacrylamide gel electrophoresis (SDS-PAGE) (Laemmli 1970; Candiano et al. 2004).
Zymogram analysis was performed by incorporating 0.01% glycol chitin in 12% polyacrylamide gel. After electrophoresis, the gel was incubated in 1% tritonX-100 for 1 h. The gel was washed 3 times in water and incubated in 100 mM Tris–Cl buffer (pH 6.0) for 16 h at 40 °C. The activity bands pertaining to the enzymes were visualized by staining with 0.01% Ranipal for 10 min followed by washing with water to remove the excess stain.
Temperature optimum and stability
To determine the temperature optimum, substrate was mixed with AS70 and assayed after incubation for 1 h at different temperatures (25 °C, 30 °C, 35 °C, 37 °C, 40 °C, 45 °C and 50 °C). The maximum activity of the chitinase was considered as 100%. The stability of the enzyme was determined by pre-incubating the enzymes at different temperatures for 15 min, 30 min, 45 min and 60 min followed by the standard assay. The residual activity of the chitinase was determined and the relative activity was calculated. The activity of AS70 without pre-incubation was considered 100%.
pH optimum and stability
The substrate was prepared in different buffer systems (sodium acetate for pH 4–5, sodium phosphate for pH 6–7, Tris–HCl for pH 8–9). AS70 was incubated with the substrate for 1 h at the optimum temperature with intermittent mixing followed by the standard assay of the chitinase. The stability of the enzyme was determined by pre-incubating AS70 in the buffers of different pH for 16 h followed by enzyme assay. The activity of AS70 without any pre-incubation was considered 100%.
Evaluation of antifungal activity in vitro
Freshly grown fungal cultures (Fusarium oxysporum, Cladosporium sp. and Alternaria alternata) were spread on Potato dextrose agar (PDA) medium and wells were punched in the medium. AS70 (400 μg) was added into the wells and the plates were incubated at 30 °C. To obtain the intracellular fraction of S167 cells (cell lysate), the cell pellet obtained after removing CS was resuspended in 25 mM Tris–Cl pH 8.0 and subjected to sonication. The suspension was centrifuged at 10,000 rpm for 20 min and the supernatant was evaluated for antifungal activity. The plates were observed for the appearance of the zone of growth inhibition of fungi.
Optimization of chitinase production
To maximize chitinase production by Chitinophaga sp. S167, the medium and process were optimized for shake flask cultures by one factor at a time approach.
Media and inducer for chitinase production
S167 was grown in Luria Bertani broth (LB), Nutrient Broth (NB) and CY medium, each supplemented with 0.5% SWC and chitinase activity was determined after every 24 h. The induction of chitinase was checked by incorporating 1% inducer (chitin flakes, swollen chitin or fungal mat) in the selected medium and the enzyme production was determined. To be used as inducer, fungal mat was prepared by growing Cladosporium sp. for 5 days in potato dextrose broth. The cells obtained by centrifugation at 10,000 rpm were washed with water and dried overnight at 50 °C. The dried fungal mass was powdered in mortar and pestle, autoclaved and stored for further use. The concentration of the selected inducer was also optimized.
Effect of incubation time on chitinase production
To determine the incubation period for maximal production of chitinase, S167 was inoculated in the optimized medium and chitinase activity was measured after every 24 h.
Effect of temperature and initial pH on chitinase production
The effect of incubation temperature on the production of chitinase was determined by growing S167 at 30 °C, 35 °C and 40 °C. The effect of the initial hydrogen ion concentration on production of chitinase was determined by growing S167 in the medium of different initial pH values (4–9). Chitinase was assayed every 24 h to determine the production of chitinase.
Effect of carbon and nitrogen sources on chitinase production
To check the effect of carbon source on chitiase production, 0.3% casitone in CY medium was replaced with different carbon sources (sucrose, cellulose, glucose, SWC and casitone) at 1% concentration. The activity of chitinase was determined at a 24 h interval. The carbon source which supported maximum production of chitinase was further tested at different concentrations for the enhancement in the enzyme production.
The effect of different organic and inorganic sources of nitrogen (1%) on chitinase production was studied. Yeast extract, peptone, ammonium sulphate and urea were added in the medium containing the optimized carbon source for chitinase production. Further, the selected nitrogen source was tested for the enhanced production of chitinase at different concentrations ranging from 0.1 to 1%.
Protection assays against fungal infection caused by Fusarium oxysporium in tomatoes
The protective effect of AS70 on the ripe tomato fruit challenged by F. oxysporum was investigated. The tomato fruits were washed with tap water and the surface was disinfected with 0.1% sodium hypochlorite for 10 min followed by washing in tap water and air drying prior to use. Each fruit was wounded (3 mm deep and 3 mm wide) with a sterile razor. 100 µL of conidial suspension (105 conidia/mL) was incubated with 100 μL of AS70 (1 mg/mL) and heat inactivated AS70 (boiled for 15 min) for 12 h at 30 °C. 10 µL of this conidial suspension was then added in each wound. The fruits were kept in plastic boxes lined with moist filter paper to maintain high humidity and incubated at 30 °C. The number of infected wounds and the diameter of infection in each wound were measured each day post inoculation (dpi). Three replicates of each treatment were taken and the experiment was repeated 3 times. The incidence and severity of disease were calculated as:
Evaluation of nematicidal activity in vitro
Nematicidal activity of the AS70 was determined using second stage juveniles (J2s) of Meloidogyne incognita. The suspension of nematode juvenile (250 J2s/50 µL) was added to 1 mL of the extract (1 mg/mL) in 35 mm petri plates. 25 mM Tris–HCl solution of pH 8 served as negative control. The experiment consisted of three replicates. The plates were kept at room temperature (25 ± 2 °C) for 7 days and were daily examined for dead J2s. Living and dead juveniles were counted by using manual counter while viewing under the light microscope. Percentage mortality was calculated by using the equation (Sun et al. 2006):
The malformed, immobile or motionless juveniles when probed with a fine needle were considered to be dead.
Statistical analysis
All experiments were conducted in triplicates and were subjected to statistical analysis. To determine difference in mean, t-test or one-way analysis of variance (ANOVA) was performed.
Results
Isolation and identification of chitinase producing bacteria
During a study carried out to explore the terrestrial myxobacterial diversity, few swarming bacteria designated as S167, S166, S165, S136 and S109 produced clear zones on glycol chitin agar medium after incubation at 30 °C for 24 h indicating the production of extracellular chitinase (Fig. 1a). Among these bacteria, S165 and S167 showed chitinolytic activity on SWC agar medium (Fig. 1b). The cell free culture supernatant (CS) of all the isolates showed hydrolytic activity on glycol chitin (Fig. 1c) whereas CS of S165 and S167 showed hydrolytic activity on SWC (Fig. 1d). Based on the size of zone of clearance on chitin substrate media, the chitinase secreted by S167 in the extracellular medium was selected for further study. S167 is a Gram-negative, yellow coloured, rod-shaped motile bacterium and was identified on the basis of 16S rRNA gene sequence. The 16S rDNA of S167 showed the greatest similarity (98.62%) on EzTaxon to C. ginsengisoli. The phylogenetic relationship of S167 to other strains in chitinophagaceae family showed that S167 clustered with C. ginsengisoli had a bootstrap value of 94 (Fig. 2). The GenBank accession number for the 16S rRNA gene sequence of S167 strain is KP017541.
Characterization of extracellular chitinase from S167
When S167 was cultivated in CY medium without any supplementation with chitinous materials, no chitinase activity could be detected in the CS even after cultivating the cells till 7 days of growth. Therefore, CY medium was supplemented with 0.5% SWC which induced chitinase in S167 as observed in the SWC medium (Fig. 1b). The extracellular medium containing chitinases was concentrated by 70% ammonium sulphate precipitation and analyzed by 12% SDS-PAGE and zymography (Fig. 3a). The zymogram showed seven clear bands of chitinase activity with molecular weights ranging from 20 to 100 kDa (Fig. 3b). The concentrated enzyme extract (AS70) was used further to study its functional characteristics.
The optimal temperature of AS70 was from 40 to 45 °C. Further, only 63% and 40% relative activity was observed at 50 °C and 37 °C respectively (Fig. 4a). The enzymes retained more than 85% of the initial activity at 20 °C and 40 °C after incubtion for 60 min whereas a drastic reduction in the activity to 24% and 13% was observed within 15 min at 50 °C and 60 °C respectively (Fig. 4b). Further, the enzymes were active at the pH ranging from 5.0 (86% relative activity) to 7.0 (93% realtive activity) with the optimum activity at pH 6.0 (Fig. 4c). The chitinases retained 100% activity at pH 6.0 after 16 h whereas the residual activity of 57% and 71% was observed at pH 5 and 7 respectively (Fig. 4d).
Antifungal activity of Chitinophaga sp. S167 chitinases in vitro
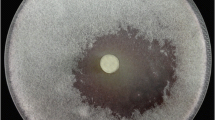
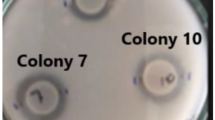
Plate assay was performed to determine if AS70 inhibited the growth of the phytopathogenic fungi, viz. Fusarium oxysporum, Alternaria alternata and Cladosporium sp. AS70 exhibited inhibitory activity against the test fungi whereas S167 cell lysate (intracellular fraction) had no effect on the growth of the test fungi (Fig. 5). Subsequent to the appearance of the zone of clearance, the fungal growth remained inhibited for 10 days on the culture plate.
Antifungal activity against plant pathogenic fungi by plate assay. aAlternaria alternata, bCladosporium sp., cFusarium oxysporum. Cycloheximide was used as positive control and ammonium sulphate (70%) precipitated CS of E. coli was used as negative control. 167 AS: AS 70, 167 lysate: intracellular fraction of S167
Optimization of physiochemical parameters for enhancing chitinase production
For opimization of the enzyme production, various physical and biochemical parameters were determined. Among the growth media tested, LB did not support the growth of S167 and the maximum chitinase activity (2.19 U/mL) was observed in CY medium followed by NB (Fig. 6a). Chitinase activity could be detected after 24 h of growth and reached its maximum after 72 h (Fig. 6b).
Further, chitinase activity reached its highest (4.5 U/mL) when S167 was cultivated at 35 °C (Fig. 6c). Also, chitinase production was affected by the initial pH of the culture medium. It was observed that the culture medium with pH 8.0 supported the maximum production of chitinase after 72 h at 35 °C (8.5 U/mL) (Fig. 6d).
Among the chitinous materials examined for the optimal induction of chitinase, 1% SWC showed the maximal induction (22 U/mL) of the chitinase (Fig. 6e). Using chitin flakes and fungal cell mat, the chitinase activity of 0.8 U/mL and 2.1 U/mL respectively was observed. As 1% SWC showed maximal chitinase production, the concentration of SWC for the enhanced chitinase production was also determined. The supplementation of CY medium with 1.5% and 1% SWC yielded the maximum chitinase production of ~ 24 U/mL (Fig. 6f). Thus, for further optimization, 1% SWC was selected as an inducer of chitinase.
To check the effect of various carbon sources on chitinase production, 0.3% casitone in CY medium was replaced with 1% carbon source (SWC, glucose, sucrose, cellulose, casitine or galactose). All the tested carbon sources except galactose resulted in an increase in the production of chitinase (Fig. 7a). Among all the carbon sources, the maximum production was obtained in SWC (27 U/mL). Further, different concentrations (0.5–2.5%) of SWC were determined to evaluate the production of chitinase by S167. The addition of 1.5% SWC showed the maximal production (44.85 Units/mL) of chitinase (Fig. 7b).
A number of organic and inorganic sources of nitrogen (1% Peptone, yeast extract, ammonium sulphate and urea) were evaluated for the optimum production of chitinase. It was observed that none of the nitrogen sources could elevate the production of chitinase as compared to the control (medium containing 1.5% SWC, 0.1% yeast extract and 0.1% CaCl2) (Fig. 7c). Thus, to deduce the concentration of yeast extract for optimum production of chitinase, different concentrations of yeast extract (0.1–1%) was added in the optimized medium components (1.5% SWC and 0.1% CaCl2). It was observed that 0.5% yeast extract enhanced the chitinase production to 96 U/mL (Fig. 7d).
Protection assays against fungal infection caused by F. oxysporum in tomatoes
The potential of AS70 to protect the tomatoes from the infection caused by F. oxysporum was checked. AS70 exerted significant protection against F. oxysporum in the infected tomatoes (Fig. 8). At 4 dpi, 88% of the wounds treated with heated AS70 showed infection of the fungus whereas none of the AS70 treated wounds was infected. AS70 was efficacious in controlling the infection, as at 6 dpi, the disease incidence of 33% was observed as compared to heated AS70 which had the disease incidence of 100%. Moreover, AS70 displayed a significant decrease in disease severity (10%) in tomatoes as compared to heated AS70 (100%). The mean disease severity caused by F. oxysporum was significantly (p < 0.05) reduced by AS70.
Effect of AS70 on infection of tomatoes caused by Fusarium oxysporum. Representative images of tomatoes treated with AS70 and heated AS70 7dpi (left) and the incidence of infection of the inoculated wounds (right). Asterisks depict significant difference between the infection incidence in AS70 treated and control (heated AS70) treated wounds at each independent day (p < 0.05)
Nematicidal activity of Chitinophaga sp. S167 chitinase in vitro
Further, AS70 caused significant mortality in second stage juveniles of M. incognita. ~ 85% mortality was observed in AS70 (1 mg/mL) treated J2s, whereas Tris–HCl treated J2s showed only 7.2% mortality after 168 h. The effect of AS70 was dose dependent, as with the increasing concentration of AS70, survival of J2s was severely affected. Over the period of 168 h, as the time goes on, the mortality significantly increased (Fig. 9).
Discussion
S167 was isolated from soil in India which exhibited potent chitinolytic activity on SWC and glycol chitin containing medium. On the basis of 16S rRNA gene sequence, the genus of the isolated strain was identified as Chitinophaga sp. S167 and it clustered with C. ginsengisoli in the phylogenetic tree. Though the novel species of Chitinophaga spp. are being isolated and the genome sequence of C. pinensis DSM 2588 has been reported to harbour genes coding for chitin degrading enzymes, only one chitin degrading enzyme has been characterized from C. pinensis till date (Ramakrishna et al. 2018). The potential of chitinases produced by Chitinophaga spp. has not been probed for any biocontrol applications. Chitinases with antifungal and nematicidal activities are viewed as the environmentally friendly alternatives to the use of harmful synthetic chemicals to control plant diseases caused by fungi and nematodes (Berini et al. 2018).
As mentioned above, S167 was cultivated in CY medium supplemented with 0.5% SWC to induce chitinolytic genes resulting in the detection of chitinase activity in the CS. Chitinase encoding genes are induced by different forms of chitin in different organisms (Keyhani and Roseman 1996; Miyashita et al. 2000; Li and Roseman 2004). The extracellular chitinases produced by S167 were concentrated by ammonium sulphate precipitation for preliminary biochemical and functional characterization. Zymogram using glycol chitin as substrate revealed that S167 secreted seven chitinases into the medium. It is known that bacteria generally produce more than one chitinase, presumably, to act on the diverse forms of chitin (Berini et al. 2018). Bacterial chitinases active over a wide range of pH and temperature have been isolated. Chitinases in AS70 had optimal temperatures of 40–45 °C and retained ˃ 85% relative activity at 20–40 °C. Chitinases from Serratia marcescens B4A, B. subtilis B298 and Enterobacter spp. showed optimal temperatures of 40–45 °C (Zarei et al. 2011; Lestari et al. 2017; Park et al. 1997). The enzymes that are active at low temperature can be economically beneficial as they can save energy in the biotechnological and ecological processes by obviating the need to operate at elevated temperatures. AS70 showed maximum activity between pH 5 and 7 with the highest activity at pH 6.0. The enzymes were highly stable (100% activity) up to 16 h at the optimal pH, whereas at pH 7.0, the enzymes were highly active (> 70% residual activity). Various chitinases from Bacillus spp. have pH optimums similar to S167 chitinases (Wang et al. 2006, 2018; Chang et al. 2010).
Owing to the lack of chitin in vertebrates and plants, chitinases are deemed as potent alternatives to the toxic fungicides to control phytopathogens, the causative agents of many plant diseases that pose threat to production and quality of crop. The extracellular chitinases produced by S167 inhibited the growth of phytopathogenic F. oxysporum, Cladosporium and A. alternata. To the best of our knowledge, this is the first report pertaining to the antifungal activity of the chitinases produced by Chitinophaga sp. The only chitinase studied from C. pinensis (GenBank accession number ACU59676.1) has been suggested to be efficient in the bioconversion of chitin waste (Ramakrishna et al. 2018). The chitinases secreted by S167 might have varied spectrum of antifungal activities or could be acting synergistically on chitin in fungal cell walls. The culture filtrate of Serratia marcescens bearing chitinolytic activity was shown to be a potent biocontrol agent against Sclerotium rolfsii (Ordentlich et al. 1988). The chitinases produced by S. marcescens 2170 have been reported to synergistically act in degradation of chitin (Suzuki et al. 2002). Several chitinases from Gram-negative bacteria i.e., Serratia spp., Enterobacter sp., Alcaligenes spp. and Streptomyces spp. are known to exhibit antifungal activity against phytopathogenic fungi (Gherbawy et al. 2012; Berini et al. 2018).
The production of different enzymes in microbial cultures can be increased by optimizing the media components and growth conditions. The increased yield of the enzymes would be required to test the utility of the enzymes in various processes. As extracellular chitinases secreted by S167 showed potential to inhibit phytopathogenic fungi, the conditions for the production of these enzymes were optimization. In CY medium supplemented with 0.5% SWC, S167 produced too few enzyme (2 U/mL) in the extracellular medium. The pH of the growth medium influences the permeability of the cell membranes in bacteria which affects the enzyme secretion into the medium. The enzyme production was increased more than 4 -fold after 72 h of growth of S167 in the medium of initial pH 8.0 at 35 °C. Bacillus lichenformis and B. pabuli K1 have been shown to produce maximum chitinase in the medium of pH 8 (Gomaa 2012; Frändberg and Schnürer 1994). The decrease in the production of the enzyme thereafter could be due to the accumulation of the hydrolysis products of SWC in the medium.
It is well-documented that the expression of chitinase in bacteria is induced by chitin (Uchiyama et al. 2003). S167 showed a further threefold increase in chitinase expression with the supplementation of 1% SWC in the medium. It is known that carbon source not only supports growth of the organism but also influences the product formation. Chitnase production may be enhanced with a particular carbon source supporting its growth whereas others may decrease its production (Beier and Bertilsson 2013). Among the different carbon sources used, 1.5% SWC as the sole source of carbon in the medium further enhanced chitinase production by further twofold. Thus, the presence of casitone in the (CY) medium might be inhibiting the induction of one or more secretory chitinase(s) in S167. The production of chitinase in B. thuringiensis and B. licheniformis was optimized using colloidal chitin medium supplemented with 1.5% chitin (Gomaa 2012).
Among the different organic and inorganic nitrogen sources added to the medium, yeast extract was the most effective and resulted in the highest chitinase production. The organic source of nitrogen is expected to enhance the yield of the chitinase because it contains amino acids or some growth factors that can be metabolised directly by the cells (Bhattacharya et al. 2016). The addition of 0.5% yeast extract in the medium resulted in a final yield of 96 U/mL chitinase. In Aeromonas punctata HS6, yeast extract was found to be the most favourable nitrogen source for chitinase production (Saima and Roohi 2013). Therefore, after the process and medium optimization, a substantial overall enhancement (48-fold) in chitinase production was observed.
The post-harvest fungal diseases in tomatoes caused by various soil borne fungi such as Fusarium sp. and Alternata sp. cause considerable loss of produce (Agrios 1997). To control these pathogens, chemical fungicides are used which are hazardous to human health and environment. Our results suggest that S167 chitinases have the great potential to control postharvest diseases of tomatoes caused by F. oxysporum. In the wounds treated with heat inactivated AS70 (control), the fungal infection started appearing after 24 h of inoculation whereas the infection in the enzyme treated wounds was evident after 120 h of inoculation. For the control of post harvest fungal diseases in fruits, the action of chitinases as antagonists produced predominantly by Bacillus spp. is reported (Swain et al. 2008; Wang et al. 2013). This is the first report of application of chitinases to control the postharvest decay of tomatoes by Fusarium sp. Kong et al. (2016) reported the antifungal activity of combination of thymol and salicylic acid to control Fusarium sp. infection in tomatoes. Another study reported the suppression of growth of Fusarium spp. on tomato slices by the volatile and non-volatile metabolites of Trichoderma spp. (El-Katatny et al. 2011). Thus, the potential of Chitinophaga sp. S167 should be explored as a biocontrol agent because of its ability to secrete multiple chitinases with potent antifungal activity.
S167 chitinases also exhibited nematicidal activity against the J2 stage of M. incognita. In the literature, there is no report of nematicidal activity of the chitinases produced by Chitinophaga spp. There are reports of J2 mortality by the CS of Bacillus spp. which contained chitinases and proteases (Lee and Kim 2016; Wei et al. 2014). We did not detect any protease activity in AS70 indicating that chitinases are causing J2 mortality. Thus, the potent nematicidal effect of the S167 chitinases against M. incognita is a promising alternatve for synthetic toxic nematicides. Moreover, chitinases are superior biocontrol agents over proteases because plants and vertebrates do not contain chitin (Neeraja et al. 2010). Recently, Chitinophaga has been identified as one of the key taxa contributing to the suppression of soyabean cyst nematode (Hu et al. 2019; Hamid et al. 2017).
Several chitinolytic microorganisms abound in diverse environments and they present a sustainable approach in control of phytopathogenic fungi, insects and nematodes. The use of chitinase producing organisms viz., actinomycetes, bacteria and fungi as biocontrol agents have been extensively exploited (Veliz et al. 2017; Subbanna et al. 2018). Due to the limited success obtained in the use of whole microorganisms as biocontrol agents, there is an enhanced focus on the bioproducts (enzymes, secondary metabolites etc.) as plant pathogen antagonists and chitinases figure prominently in this list (Raymaekers et al. 2020; Swiontek-Brzezinska et al. 2014; Alves et al. 2020; Subbanna et al. 2018; Kumar et al. 2018). Moreover, chitinases are outstanding candidates to prevent contamination of fruits in the postharvest stage, because they are selective against fungi without causing any damage to the fruit (daSilva 2019). Another advantage of using enzymes is that they can be produced in bioreactors for large scale applications.
The present study underlines the importance of chitinases produced by Chitinophaga sp. S167 in controlling the phytopathogenic fungi and M. incognita, a parasitic nematode. Our results showed that chitinases produced by S167 inhibited the infection of F. oxysporum through postharvest wounds on tomatoes. Further work will focus on the evaluation of potential of S167 chitinases in inhibiting diverse fungal pathogens that infect various fruits at the postharvest stage. The stability and activity of the enzyme preparation on fruits stored at different temperatures would be probed in the future course of investigation. It will be tested if S167 chitinases can act synergistically with the cocktail of other cell degrading enzymes for more efficacious control of postharvest decay of fruits. Moreover, the in vitro nematicidal activity of S167 chitinases will be further validated by carrying out in vivo studies in the nematode infested plants.
Conclusion
The present study is the first one to investigate the biocontrol potential of the multiple chitinases secreted in the extracellular medium by newly isolated Chitinophaga sp. S167. The extracellular concentrated crude extract of chitinases from S167 (AS70) was optimally active at 40 °C and pH 6. AS70 inhibited the growth of phytopathogenic fungi in vitro, highlighting its potential use as a biocontrol agent. To overcome the bottleneck of low production of enzymes in native host to study their applications, various physical and biochemical parameters were optimized to increase the production of chitinases by S167. Chitinase production could be enhanced by 48 folds by growing S167 in the medium supplemented with (1%) SWC as carbon source and (0.5%) yeast extract as nitrogen source at 35 °C and pH 8.0. The application of AS70 conferred complete protection to tomatoes against the F. oxysporum infection till 4 dpi. AS70 also caused significant mortality in second stage juveniles of M. incognita showing its nematicidal potential. Thus, this study has pointed out the importance of this underexplored genus for its application as biocontrol agent in plant diseases.
References
Agrios G (1997) Plant pathology, 4th edn. Academic Press, San Diego, p 184
Alves EA, Schmaltz S, Tres VM, Zabot LG, Kuhn CR, Muzutti AM (2020) Process development to obtain a cocktail containing cell-wall degrading enzymes with insecticidal activity from Beauveria bassiana. Biochem Eng J 156:107484. https://doi.org/10.1016/j.bej.2019.107484
Beier S, Bertilsson S (2013) Bacterial chitin degradation—mechanisms and ecophysiological strategies. Front Microbiol 4:149. https://doi.org/10.3389/fmicb.2013.00149
Berini F, Katz C, Gruzdev N, Casartelli M, Tettamanti G, Marinelli F (2018) Microbial and viral chitinases: attractive biopesticides for integrated pest management. Biotechnol Adv 36:818–838. https://doi.org/10.1016/j.biotechadv.2018.01.002
Bhattacharya S, Das A, Samadder S, Rajan SS (2016) Biosynthesis and characterization of a thermostable, alkali-tolerant chitinase from Bacillus pumilus JUBCH08 displaying antagonism against phytopathogenic Fusarium oxysporum. 3 Biotech 6:87. https://doi.org/10.1007/s13205-016-0406-x
Brabcová V, Nováková M, Davidová A, Baldrian P (2016) Dead fungal mycelium in forest soil represents a decomposition hotspot and a habitat for a specific microbial community. New Phytol 210:1369–1381. https://doi.org/10.1111/nph.13849
Candiano G, Bruschi M, Musante L, Santucci L, Ghiggeri GM, Carnemolla B, Orecchia P, Zardi L, Righetti PG (2004) Blue silver: a very sensitive colloidal Coomassie G-250 staining for proteome analysis. Electrophoresis 25:1327–1333. https://doi.org/10.1002/elps.200305844
Carmona-Hernandez S, Reyes-Pérez JJ, Chiquito-Contreras RG, Rincon-Enriquez G, Cerdan-Cabrera RC, Hernandez-Montiel GL (2019) Biocontrol of postharvest fruit fungal diseases by bacterial antagonists: a review. Agronomy 9:121. https://doi.org/10.3390/agronomy9030121
Carrión VJ, Perez-Jaramillo J, Cordovez V, Tracanna V, Hollander DM et al (2019) Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366:606–612. https://doi.org/10.1126/science.aaw9285
Chang WT, Chen ML, Wang SL (2010) An antifungal chitinase produced by Bacillus subtilis using chitin waste as a carbon source. World J Microbiol Biotechnol 26:945–950. https://doi.org/10.1007/s11274-009-0244-7
da Silva RR (2019) Enzyme technology in food preservation: a promising and sustainable strategy for biocontrol of post-harvest fungal pathogens. Food Chem 277:531–532. https://doi.org/10.1016/j.foodchem.2018.11.022
Del Rio TG, Abt B, Spring S, Lapidus A, Nolan M, Tice H, Copeland A, Cheng JF, Chen F, Bruce D, Goodwin L (2010) Complete genome sequence of Chitinophaga pinensis type strain (UQM 2034 T). Stand Genomic Sci 2:87–95. https://doi.org/10.4056/sigs.661199
Dukare AS, Paul S, Nambi VE, Gupta KR, Rajbir S, Sharma K, Vishwakarma KR (2019) Exploitation of microbial antagonists for the control of postharvest diseases of fruits: a review. Crit Rev Food Sci Nutr 59:1498–1513. https://doi.org/10.1080/10408398.2017.1417235
El-Katatny MH, El-Katatny MS, Fadl-Allah EM, Emam AS (2011) Antagonistic effect of two isolates of Trichoderma harzianum against postharvest pathogens of tomato (Lycopersicon esculentum). Arch Phytopathol Plant Prot 44:637–654. https://doi.org/10.1080/03235400903266438
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Frändberg E, Schnürer J (1994) Chitinolytic properties of Bacillus pabuli K1. J Appl Bacteriol 76:361–367
Fuchs RL, McPherson SA, Drahos DJ (1986) Cloning of a Serratia marcescens gene encoding chitinase. Appl Environ Microbiol 51:504–509
Gherbawy Y, Elhariry H, Altalhi A, El-Deeb B, Khiralla G (2012) Molecular screening of Streptomyces isolates for antifungal activity and family 19 chitinase enzymes. J Microbiol 50:459–468. https://doi.org/10.1007/s12275-012-2095-4
Gomaa EZ (2012) Chitinase production by Bacillus thuringiensis and Bacillus licheniformis: their potential in antifungal biocontrol. J Microbiol 50:103–111. https://doi.org/10.1007/s12275-012-1343-y
Hamid MI, Hussain M, Wu Y, Zhang X, Xiang M, Liu X (2017) Successive soybean-monoculture cropping assembles rhizosphere microbial communities for the soil suppression of soybean cyst nematode. FEMS Microbiol Ecol 93:fiw122. https://doi.org/10.1093/femsec/fiw222
Hartl L, Zach S, Seidl-Seiboth V (2012) Fungal chitinases: diversity, mechanistic properties and biotechnological potential. Appl Microbiol Biotechnol 93:533–543. https://doi.org/10.1007/s00253-011-3723-3
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293:781–788
Heustis RJ, Ng HK, Brand KJ, Rogers MC, Le LT, Specht CA, Fuhrman JA (2012) Pharyngeal polysaccharide deacetylases affect development in the nematode C. elegans and deacetylate chitin in vitro. PLoS ONE 7:e40426. https://doi.org/10.1371/journal.pone.0040426
Horn SJ, Sørbotten A, Synstad B, Sikorski P, Sørlie M, Vårum KM, Eijsink VG (2006) Endo/exo mechanism and processivity of family 18 chitinases produced by Serratia marcescens. FEBS J 273:491–503. https://doi.org/10.1111/j.1742-4658.2005.05079.x
Hu W, Strom NB, Haarith D, Chen S, Bushley KE (2019) Seasonal variation and crop sequences shape the structure of bacterial communities in cysts of soybean cyst nematode. Front Microbiol 10:2671. https://doi.org/10.3389/fmicb.2019.02671
Keyhani NO, Roseman S (1996) The chitin catabolic cascade in the marine bacterium Vibrio furnissii molecular cloning, isolation, and characterization of a periplasmic chitodextrinase. J Biol Chem 271:33414–33424
Kim SK, Kook M, Yan ZF, Park SY, Kim SS, Kim HB, Trinh H, Won KH, Yang JE, Yi TH (2019) Chitinophaga aurantiaca sp. nov., isolated from a soil sample from a tangerine field. Antonie Van Leeuwenhoek 112:1189–1197. https://doi.org/10.1007/s10482-019-01251-1
Kong J, Xie YF, Guo YH, Cheng YL, Qian H, Yao WR (2016) Biocontrol of postharvest fungal decay of tomatoes with a combination of thymol and salicylic acid screening from 11 natural agents. LWT Food Sci Technol 72:215–222. https://doi.org/10.1016/j.lwt.2016.04.020
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kumar M, Brar A, Yadav M, Chawade A, Vivekanand V, Pareek N (2018) Chitinases—potential candidates for enhanced plant resistance towards fungal pathogens. Agric 8:1–12. https://doi.org/10.3390/agriculture8070088
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Larsbrink J, Tuveng TR, Pope PB, Bulone V, Eijsink VG, Brumer H, McKee LS (2017) Proteomic insights into mannan degradation and protein secretion by the forest floor bacterium Chitinophaga pinensis. J Proteomics 156:63–74. https://doi.org/10.1016/j.jprot.2017.01.003
Lee YS, Kim KY (2016) Antagonistic potential of Bacillus pumilus L1 against root-knot nematode, Meloidogyne arenaria. J Phytopathol 164:29–39. https://doi.org/10.1111/jph.12421
Lestari P, Prihatiningsih P, Djatmiko AH (2017) Partial biochemical characterization of crude extract extracellular chitinase enzyme from Bacillus subtilis B 298. IOP Conf Ser: Mater Sci Eng 172:012041. https://doi.org/10.1088/1757-899X/172/1/012041
Li X, Roseman S (2004) The chitinolytic cascade in Vibrios is regulated by chitin oligosaccharides and a two-component chitin catabolic sensor/kinase. Proc Natl Acad Sci USA 101:627–631. https://doi.org/10.1073/pnas.0307645100
Mckee LS, Ruthes AC, Vilaplana F, Brumer H (2019) Focused metabolism of β–glucans by the soil Bacteroidetes species Chitinophaga pinensis. Appl Environ Microbiol. https://doi.org/10.1128/AEM.02231-18
Mehmood MA, Zhao H, Cheng J, Xie J, Jiang D, Fu Y (2020) Sclerotia of a phytopathogenic fungus restrict microbial diversity and improve soil health by suppressing other pathogens and enriching beneficial microorganisms. J Environ Manag 259:109857. https://doi.org/10.1016/j.jenvman.2019.109857
Miyashita K, Fujii T, Saito A (2000) Induction and repression of a Streptomyces lividans chitinase gene promoter in response to various carbon sources. Biosci Biotechnol Biochem 64:39–43. https://doi.org/10.1271/bbb.64.39
Monreal J, Reese ET (1969) The Chitinases of Serratia marcescens. Can J Microbiol 15:689–696. https://doi.org/10.1139/m69-122
Neeraja C, Anil K, Purushotham P, Suma K, Sarma P, Moerschbacher BM, Podile AR (2010) Biotechnological approaches to develop bacterial chitinases as a bioshield against fungal diseases of plants. Crit Rev Biotechnol 30:231–241. https://doi.org/10.3109/07388551.2010.487258
Ordentlich A, Elad Y, Chet I (1988) The role of chitinase of Serratia marcescens in biological control of Sclerotium rolfsii. Phytopathol 78:84–88
Park JK, Morita K, Fukumoto I, Yamasaki Y, Nakagawa T, Kawamukai M, Matsuda H (1997) Purification and characterization of the chitinase (ChiA) from Enterobacter sp. Gl. Biosci Biotechnol Biochem 61:684–689. https://doi.org/10.1271/bbb.61.684
Ramakrishna B, Vaikuntapu P, Mallakuntla MK, Bhuvanachandra B, Sivaramakrishna D, Uikey S, Podile AR (2018) Carboxy-terminal glycosyl hydrolase 18 domain of a carbohydrate active protein of Chitinophaga pinensis is a non-processive exochitinase. Int J Biol Macromol 115:1225–1232. https://doi.org/10.1016/j.ijbiomac.2018.04.159
Raymaekers K, Ponet L, Holtappels D, Berckmans B, Cammue PAB (2020) Screening for novel biocontrol agents applicable in plant disease management—a review. Biol Control 144:104240. https://doi.org/10.1016/j.biocontrol.2020.104240
Reichenbach H, Dworkin M (1992) The prokaryotes. In: Balows A, Truper HG, Dworkin M, Harder W, Schleifer KH (eds) The myxobacteria. Springer, New York, pp 3416–3487
Reissig JL, Strominger JL, Leloir LF (1955) A modified colorimetric method for the estimation of N-acetylamino sugars. J Biol Chem 217:959–966
Romaguera A, Menge U, Breves R, Diekmann H (1992) Chitinases of Streptomyces olivaceoviridis and significance of processing for multiplicity. J Bacteriol 174:3450–3454
Saima KM, Roohi AIZ (2013) Isolation of novel chitinolytic bacteria and production optimization of extracellular chitinase. J Genet Eng Biotechnol 11:39–46. https://doi.org/10.1016/j.jgeb.2013.03.001
Saito A, Fujii T, Yoneyama T, Redenbach M, Ohno T, Watanabe T, Miyashita K (1999) High-multiplicity of chitinase genes in Streptomyces coelicolor A3 (2). Biosci Biotechnol Biochem 63:710–718. https://doi.org/10.1271/bbb.63.710
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sangkhobol V, Skerman VBD (1981) Chitinophaga, a new genus of chitinolytic myxobacteria. Int J Syst Bacteriol 31:285–293
Seidl V (2008) Chitinases of filamentous fungi: a large group of diverse proteins with multiple physiological functions. Fungal Biol Rev 22:36–42. https://doi.org/10.1016/j.fbr.2008.03.002
Subbanna ARNS, Rajasekhara H, Stanley J, Mishra KK, Pattanayak A (2018) Pesticidal prospectives of chitinolytic bacteria in agricultural pest management. Soil Biol Biochem 116:52–66. https://doi.org/10.1016/j.soilbio.2017.09.019
Sun MH, Gao L, Shi YX, Li BJ, Liu XZ (2006) Fungi and actinomycetes associated with Meloidogyne spp. eggs and females in China and their biocontrol potential. J Invertebr 93:22–28. https://doi.org/10.1016/j.jip.2006.03.006
Suzuki K, Sugawara N, Suzuki M, Uchiyama T, Katouno F, Nikaidou N, Watanabe T (2002) Chitinases A, B, and C1 of Serratia marcescens 2170 produced by recombinant Escherichia coli: enzymatic properties and synergism on chitin degradation. Biosci Biotechnol Biochem 66:1075–1083. https://doi.org/10.1271/bbb.66.1075
Swiontek Brzezinska M, Jankiewicz U, Burkowska A, Walczak M (2014) Chitinolytic microorganisms and their possible application in environmental protection. Curr Microbiol 68:71–81. https://doi.org/10.1007/s00284-013-0440-4
Swain MR, Ray RC, Nautiyal CS (2008) Biocontrol efficacy of Bacillus subtilis strains isolated from cow dung against postharvest yam (Dioscorea rotundata L.) pathogens. Curr Microbiol 57:407–411. https://doi.org/10.1007/s00284-008-9213-x
Trudel J, Asselin A (1989) Detection of chitinase activity after polyacrylamide gel electrophoresis. Anal Biochem 178:362–366
Tsujibo H, Kubota T, Yamamoto M, Miyamoto K, Inamori Y (2003) Characterization of chitinase genes from an alkaliphilic actinomycete, Nocardiopsis prasina OPC-131. Appl Environ Microbiol 69:894–900. https://doi.org/10.1128/AEM.69.2.894-900.2003
Uchiyama T, Kaneko R, Yamaguchi J, Inoue A, Yanagida T, Nikaidou N, Regue M, Watanabe T (2003) Uptake of N, N′-diacetylchitobiose [(GlcNAc) 2] via the phosphotransferase system is essential for chitinase production by Serratia marcescens 2170. J Bacteriol 185:1776–1782. https://doi.org/10.1128/JB.185.6.1776-1782.2003
Veliz AE, Martinez-Hidalgo P, Hirsch MA (2017) Chitinase-producing bacteria and their role in biocontrol. AIMS Microbiol 3:689–705. https://doi.org/10.3934/microbiol.2017.3.689
Wang SL, Lin TY, Yen YH, Liao HF, Chen YJ (2006) Bioconversion of shellfish chitin wastes for the production of Bacillus subtilis W-118 chitinase. Carbohydr Res 341:2507–2515. https://doi.org/10.1016/j.carres.2006.06.027
Wang X, Xu F, Wang J, Jin P, Zheng Y (2013) Bacillus cereus AR156 induces resistance against Rhizopus rot through priming of defense responses in peach fruit. Food Chem 136:400–406. https://doi.org/10.1016/j.foodchem.2012.09.032
Wang D, Li A, Han H, Liu T, Yang Q (2018) A potent chitinase from Bacillus subtilis for the efficient bioconversion of chitin-containing wastes. Int J Biol Macro 116:863–868. https://doi.org/10.1016/j.ijbiomac.2018.05.122
Wei L, Shao Y, Wan J, Feng H, Zhu H, Huang H, Zhou Y (2014) Isolation and characterization of a rhizobacterial antagonist of root-knot nematodes. PLoS ONE 9:e85988. https://doi.org/10.1371/journal.pone.0085988
Zarei M, Aminzadeh S, Zolgharnein H, Safahieh A, Daliri M, Noghabi KA, Motallebi A (2011) Characterization of a chitinase with antifungal activity from a native Serratia marcescens B4A. Braz J Microbio 42:1017–1029. https://doi.org/10.1590/S1517-838220110003000022
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: RK, PO. Performed the experiments: Chitinase experiments-SS, Nematicidal Activity-AK, Strain isolation-SK. Analyzed the data: RK, RK, PO. Wrote the manuscript: SS, AK (initial draft), RK (final draft).
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sharma, S., Kumar, S., Khajuria, A. et al. Biocontrol potential of chitinases produced by newly isolated Chitinophaga sp. S167. World J Microbiol Biotechnol 36, 90 (2020). https://doi.org/10.1007/s11274-020-02864-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02864-9