Abstract
Rice sheath blight, caused by Rhizoctonia solani, is a major worldwide rice disease for which little is known about the molecular mechanisms of host immunity to infection. In the present study, a comparative proteomic analysis of two rice cultivars, Teqing (resistant) and Lemont (susceptible), inoculated with R. solani was conducted using an eight-plex iTRAQ (isobaric tags for relative and absolute quantitation) technique, resulting in the identification and quantification of 6560 proteins. A total of 755 proteins showed significant changes in abundance between plants infected with R. solani and control plants, based on an error factor < 2 and a more than 1.5-fold or less than 0.67-fold quantitative difference. The differentially abundant proteins were mainly involved in glyoxylate and dicarboxylate metabolism; glycine, serine and threonine metabolism; unsaturated fatty acid biosynthesis; and glycolysis/gluconeogenesis regulation pathways (p < 0.01). In addition, the expression levels of the genes encoding selected proteins were tested by qRT-PCR, and their functions were tested in Nicotiana benthamiana via agroinfiltration. Based on these proteomic and experimental data, a putative model of the regulation of rice immunity under R. solani infection is proposed. The proteins identified in the present study provide a basis for elucidating the molecular mechanisms underlying rice immunity to infection by R. solani.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice sheath blight is caused by the soil-borne necrotrophic fungus Rhizoctonia solani, which causes disease in a broad spectrum of plants such as rice, wheat, potato, bean, cotton, and sugar beet (Anderson 1982). The fungus invades the roots, stems and other aerial parts of the plant, causing disease symptoms such as root rot, sheath blight, banded leaf, aerial blight and brown patch (Ogoshi 1987; Wrather et al. 1997). R. solani is divided into 14 anastomosis groups (AG1 to AG13 and AGBI) (Ogoshi 1987), in which AG1 IA is one of the most important plant pathogens causing the most serious economic agricultural losses.
Rice sheath blight is considered one of the major rice diseases that leads to significant yield and quality losses (Lee and Rush 1983). The general symptoms of sheath blight include necrotic, dark, reddish-brown, elliptical or oval lesions on the leaf sheath, leaf blade and culm (Kumar et al. 2009; Ghosh et al. 2014). Sheath blight is difficult to manage because of the wide host range, rapid variability, and long survival time in the soil (Taheri 2007). In addition, attempts to control sheath blight by traditional breeding processes have not succeeded due to the lack of strongly resistant cultivars (Bonman et al. 1992). Hence, investigating and utilizing innate rice immune components is an important step toward resistant variety development or breeding.
Proteomic analysis is a valuable tool in dissecting plant-pathogen interactions due to the direct involvement of proteins in molecular processes and biological functions (Kim et al. 2004; Neilson et al. 2011a, b). Recently, proteomic research has made considerable progress in providing functional information regarding abiotic and biotic stress responses (Komatsu and Tanaka 2005). For example, proteomic responses to salinity, drought and cold have been studied in various plants including rice (Neilson et al. 2011a, b; Sarhadi et al. 2012; Wu et al. 2016), canola (Bandehagh et al. 2011), soybean (Ma et al. 2012; Tian et al. 2015), wheat (Guo et al. 2012; Faghani et al. 2015), grape (Delaunois et al. 2013), and barley (Fatehi et al. 2012; Chmielewska et al. 2016). Biotic stresses caused by living organisms (Delaunois et al. 2014), such as fungi (Li et al. 2012; Zhao et al. 2014), bacteria (Gonzalez et al. 2012), viruses (Chen et al. 2014; Berard et al. 2015), and insects (Du et al. 2015) have also been investigated. Zhao et al. reported a total of 27 differentially abundant proteins in response to Pst (Pseudomonas syringae pv. Tomato) inoculation using the two-dimensional electrophoresis (2-DE) technique. Most of these proteins fall into the category “response to stimulus” and are involved in basic resistance processes, such as glycerol-3-phosphate and hydrogen peroxide signaling (Zhao et al. 2014). Li et al. (2012) observed that salicylic acid (SA)-treated rice leaves contained 36 differentially abundant proteins implicated in various functions, including defense, antioxidative enzymes, and signal transduction.
Proteins responsive to R. solani in sheath tissue were investigated in two rice cultivars, Labelle (resistant) and LSBR-5 (susceptible) (Lee et al. 2006). Moreover, an array of antifungal proteins, including chitinases (Datta et al. 2000; Sridevi et al. 2003; Sripriya et al. 2008), thaumatin-like proteins (Datta et al. 1999), nonspecific lipid transfer proteins (Patkar 2006) and plant defense proteins (Jha and Chattoo 2010), have been biotechnologically exploited to generate transgenic plants with resistance to sheath blight. Constitutive overexpression of the OsWRKY4 gene increases resistance to R. solani concomitant with elevated expression of jasmonate (JA)- and ethylene (ET)-responsive pathogenesis-related (PR) genes, implying that the defensive response to rice sheath blight occurs through the JA/ET-dependent signal pathway (Wang et al. 2015). More recently, gas chromatography–mass spectrometry (GC–MS) and RNA sequencing analyses showed that modulating host photosynthesis, respiration, phytohormone signaling, and secondary metabolism is crucial in rice during R. solani infection (Ghosh et al. 2017).
Most rice varieties grown around the world are susceptible to R. solani, although moderate to high levels of resistant have been reported (Pan et al. 1999). Lemont is highly susceptible to sheath blight disease, while Teqing is moderately resistant (Bollich et al. 1985; Tabien et al. 2000; Zheng et al. 2011; Pinson et al. 2012). Although sheath blight disease is one of the most destructive diseases worldwide, limited information is available regarding the mechanisms of rice response to infection. Understanding the genetic mechanisms of plant disease resistance against this pathogen will benefit the development of improved varieties with R. solani resistance enormously. To investigate the rice—R. solani interaction, we conducted a proteomic analysis in resistant and susceptible rice cultivars after R. solani infection The primary objectives of this research were to: (1) identify proteins significantly expressed in response to R. solani infection in rice; (2) analyze the proteins that were differentially expressed between resistant and susceptible rice cultivars; (3) reveal the interaction network and defense mechanism between rice and R. solani. The knowledge gained from this study will be instrumental in understanding the molecular mechanisms involved in the response to R. solani infection in rice.
Materials and methods
Plant materials and inocolalion with R. solani
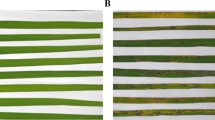
Rice cultivars Teqing (resistant) and Lemont (susceptible) were provided by National Mid-term Genebank for Rice of China National Rice Research (Tabien et al. 2000). After soaking and germination of rice seeds with water, they were planted in soil and placed in the growth room. After then, rice seedlings were transferred into 500 mL pots containing a vermiculite-potting soil mixture. Plants were cultivated in a growth room maintained at 26 °C and 70% relative humidity with a 12/12 h day (200 µE m−2 s−1)/night. R. solani AG1 IA was used in this study. Inoculation was performed using a mycelial plug (5 mm in diameter) containing the actively growing mycelium of R. solani from a culture that had almost covered the surface of the potato dextrose agar (PDA) media in a 90 mm plate. Plugs were inoculated onto the stems of rice seedlings at the four-leaf stage at 20 mm above the surface of the culture liquid. At the same time, the control samples were inoculated with PDA. The samples of each experimental groups were collected 24 h post inoculation (hpi) for proteomic analysis. The stems of 60 plants were collected as one independent biological replicate and two biological replicates were produced in our work, which resulted in eight samples. The eight samples of two biological repeats were analyzed using 8-plex iTRAQ experiment and by running the LC–MS/MS analysis twice. LC–MS/MS analysis for twice resulted in two technical replicates in this study. All samples were immediately frozen in liquid nitrogen and stored at − 80 °C until used. The pictures of phenotype in two rice cultivars infected with R. solani were taken at 24 and 192 hpi.
Extraction of total proteins from the rice stem
The stems from three plants (approximately 0.5 g) were immersed in liquid nitrogen, ground to a fine powder, and then suspended in 10% w/v tricarboxylic acid/acetone containing 0.1% dithiothreitol (DTT) in 50 mL centrifuge tube at − 20 °C for 2 h. Thereafter, the tubes were centrifuged at 40,000×g for 20 min and the supernatants discarded. Finally, the protein pellets were washed twice with 30 mL of ice-cold acetone and lyophilized. The vacuum dried pellets were dissolved in 300 µl lysis solution containing 7 M urea, 2 M thiourea, 4% w/v CHAPS, 65 mM DTT and 1 mM phenylmethylsulfonyl fluoride. Insoluble materials were removed by centrifugation, and the protein concentration of the sample was quantified using the Bradford method with bovine serum albumin as standard protein.
Protein digestion, iTRAQ labeling and strong cation exchange
Proteins (100 μg of each sample) were first reduced, alkylated and then labeled using iTRAQ reagents as follows Proteins of each sample were first dissolved in 20 μL of dissolution buffer (7 M urea, 2 M thiourea, 4% w/v CHAPS and 65 mM DTT), reduced with 120 μL of reducing reagent (8 M urea, 0.1 M Tris–HCl and 10 mM DTT), incubated at 36 °C for 1 h, alkylated with 120 μL of cysteine blocking reagent (8 M urea, 0.1 M Tris–HCl and 50 mM IAA), and incubated at room temperature for 10 min (Berger et al. 2007). After then, the samples were digested with trypsin at a 20:1 mass ratio at 37 °C for 14 h, then labeled using the iTRAQ Reagents 8-plex kit according to the manufacturer’s instructions (AB Sciex Inc.). Eight samples of two biological repeats was labeled with iTRAQ tags, as follow: the untreated Teqing replicates were labeled with iTRAQ tags 113 and 114, and the R. solani-treated Teqing replicates were labeled with tags 115 and 116, while the untreated and R. solani-treated Lemont samples were labeled with 117/118 and 119/121, respectively. The labeled samples were then pooled and dried in an Eppendorf vacuum concentrator. Then, the samples were mixed and lyophilized before dissolving in 4 mL of strong cation exchange (SCX) buffer A (25 mM NaH2PO4 in 25% acetonitrile, pH 2.7). The peptides fractionated on Ultremex SCX column (4.6 × 250 mm) using an Agilent 1200 HPLC were fractionated into ten fractions. An Exigent Nano LC-Ultra 2D system (AB Sciex) was used for sample separation. At last, a Triple TOF 5600 mass spectrometer and a Nano Spray III Source (AB Sciex) were used to perform mass spectrometer data acquisition twice, which resulted in two technical replicates (Berger et al. 2007). In brief, peptides were eluted from the C18 column into the mass spectrometer using a linear gradient (5–80%) of acetonitrile (ACN) at a flow rate of 250 μL min−1 for 1 h. The buffers used to create the ACN gradient were: Buffer A [98% H2O, 2% ACN, 0.1% formic acid, and 0.005% trifluoroacetic acid (TFA)] and Buffer B (100% ACN, 0.1% formic acid, and 0.005% TFA). MS/MS data were acquired in a data-dependent manner in which the MS1 data was acquired at a 250 ms at m/z of 400 to 2500 Da and the MS/MS data were acquired from m/z of 50 to 2000 Da. The independent data acquisition parameters were as follows: MS1-TOF acquisition time of 250 ms, followed by 50 MS2 events of 48 ms acquisition time for each event. The ion had the charge state + 2, + 3 and + 4. The ion exclusion time was set to 4 s. The collision energy was set to iTRAQ experiment setting. Finally, the collected data were analyzed using ProteinPilot™ 4.2 (ABSCIEX) for peptide identification.
Database search and iTRAQ quantification
ProteinPilot™ software (Version 4.2) was used for raw data processing against the database of Oryza sativa from UniProt (http://www.uniprot.org). The main database search parameters were as follows: the instrument was TripleTOF 5600, iTRAQ quantification, cysteine modified with iodoacetamide; and biological modifications were selected as ID and trypsin digestion. Peptides with a global false discovery rate (FDR) < 1% were used for further protein annotation. To minimize the incidence of false positives, a strict cutoff of unused ProtScore > 1.3 was applied for protein identification. Furthermore, at least one peptide with the 95% confidence was required for inclusion. The accuracy of each protein ratio is given by a calculated “error factor” in the software. The error factor expresses the 95% uncertainty range (95% confidence error) for a reported ratio, where this 95% confidence error is the weighted standard deviation of the weighted average of log ratios multiplied by the Student’s t factor for n − 1 degrees of freedom, where n is the number of peptides contributing to relative protein quantification. In addition, we performed ANOVA with the biological and technical replicates data individual, and two-way analysis of variance (ANOVA) and LSD analysis (SPSS 18.0) were used to determine if the protein was significantly regulated over time. To control the FDR, we also performed Benjamini–Hochberg correction for multiple testing, and p < 0.05 was accepted. When identifying protein species abundance as significantly different, data were considered reliable when the p value was less than 0.05, the error factor < 2 and a ratio fold change > 1.5 (up-regulated) or < 0.67 (down-regulated) in two biological replicates and two technical replicates to designate significant changes in protein expression.
Gene ontology (GO) and kyoto encyclopedia of genes and genomes (KEGG) analysis
Differentially expressed proteins were classified according to GO (http://www.geneontology.org). GO annotation was used to predict biological processes, cellular component, molecular function for differentially expressed proteins. KEGG (http://www.genome.jp/kegg/ or http://www.kegg.jp/) was used to predict significant pathways of proteins involved in response to R. solani infection. STRING (http://string.embl.de) was used to obtain protein–protein interaction information with the default score setting. Potential paths between differentially expressed proteins and metabolic compounds were used to query the knowledge-driven database with a shortest-path algorithm. Cytoscape (http://cytoscape.org/) (Version 3.1) was used to construct regulatory networks.
RNA extraction and qRT-PCR analysis of gene expression
To analyze the expression of the target genes, total RNA was extracted from ten rice stems at 0, 24 and 48 hpi following R. solani infection using TRIzol. A NanoDrop-1000 was used to detect the quality and quantity of RNA. Reverse transcription to cDNA was conducted by using the SuperScript first-strand synthesis system (Invitrogen). SYBR Green mix was used in qRT-PCR to determine the expression levels of the target genes. RT-PCR was performed in 10 µL reaction mixture consisting of 5 µL SYBR Green mix (Vazyme), 0.5 µL of 10 mM each primer, and 2 mL of appropriate diluted cDNA. The conditions for RT-qPCR were as follows: 95 °C for 5 min, then 40 cycles at 95 °C for 30 s and 60 °C for 34 s, followed by 72 °C for 35 s for PCR amplification. Transcript levels of each gene were measured used the Applied Biosystems 7500 system according to the manufacturer’s instructions. We used 2∆∆CT method analysis the RT-PCR data. The 18S RNA was used as a housekeeping gene in the qRT-PCR analysis. Primers used in this study are listed in Supplementary Table S8.
Functional analysis of proteins in N. benthamiana
Overexpression constructs of candidate genes were generated by cloning the full-length coding sequences (CDS) into a Gateway destination vector pEG100. The constructs were transformed into Agrobacterium tumefaciens strain GV3101 by electric transfer. The agrobactrium harboring constructs were inoculated in liquid LB medium containing 50 µg mL−1 rifampicin and 50 µg mL−1 kanamycin at 28 °C overnight. Cultured cells were pelleted by centrifugation, washed once with and resuspended in 10 mL infiltration buffer (10 mM MgCl, 100 μM acetosyringone). Transient assays in N. benthamiana were performed by infiltrating whole leaf of 3-week-old N. benthamiana plants with Agrobacterium harboring constructs containing the proteins (OD600 = 1). Two days after infiltration, the leaves were inoculated with R. solani using a mycelial plug (5 mm in diameter) containing the actively growing mycelium from a culture. Lesions were measured and photos taken three days later. We used Photoshop CS6 calculate the lesions areas.
Statistical analysis
All data obtained were subjected to two-way analyses of variance (ANOVA) and mean differences were compared by the least significant difference (L.S.D.) test, and comparisons with p < 0.05 were considered significantly different. Principal component analysis (PCA) was performed with the program SIMCA-P Version 13.0 (Umetrics, Umea, Sweden).
Results
Proteomic changes in rice seedling sheath induced by R. solani infection
Four-leaf stage rice cultivars (resistant: Teqing, susceptible: Lemont) were infected with R. solani. At 24 h post inoculation (hpi), a large numbers of hyphae were produced at the inoculation site the rice stalks gradually became yellow and lodging was observed in Lemont. In contrast, Teqing exhibited fewer disease symptoms. At 192 hpi, the leaves of Lemont showed obvious drooping and the stalk was significantly thinner. In contrast, the leaf drooping of Teqing was not obvious and the stalk was not significantly thinner (Fig. 1). To quantitatively profile the protein abundance pattern in rice seedlings in response to R. solani, we employed the iTRAQ. Before analyzing differentially expressed proteins (DEPs), we made a PCA analysis raw data of iTRAQ from two biological replicates and two technical replicates, and the results showed that there was a good separation of the conditions and no variation (Fig. 2). After then, the total proteins of both Teqing and Lemont rice inoculated with R. solani for 24 h were analyzed by iTRAQ. A total of 6560 proteins were identified with less than 1% false discovery rate in two biological replicates and two technical replicates, and 755 proteins were identified with an error factor (EF) < 2, p value < 0.05 and more than 1.5-fold or less than 0.67-fold quantitative differences in both biological and technical replicates, representing significant changes in protein species abundance (Supplementary Tables S1 and S2). The peptide sequences are listed in Supplementary Table S3. Among these proteins, the abundance of 389 proteins changed significantly in the disease-resistant rice cultivar when infected by R. solani for 24 h, while 448 proteins showed significant abundance changes in the susceptible rice cultivar (Supplementary Table S2). Only 43 proteins showed similar abundance changes in the two rice cultivars (Fig. 2a).
Compared to the untreated plants, in the resistant cultivar, 161 proteins were significantly more abundant, while in the susceptible cultivar, 354 proteins increased in abundance in response to R. solani infection for 24 h (Fig. 3a). Interestingly, 35 identified proteins increased significantly in both Teqing and Lemont at 24 hpi following R. solani infection (Fig. 3a). Among the proteins that were suppressed by R. solani, 185 were identified in only Teqing, whereas 51 proteins were identified in only Lemont (Fig. 3b). Only 8 proteins decreased significantly in both Teqing and Lemont (Fig. 3b). These results show that more proteins were increased by infection in the disease-susceptible than in the resistant cultivar. In contrast, fewer proteins decreased in the disease-susceptible than in the disease-resistant cultivar. Thus, we chose to study the function of these differentially abundant proteins which have differentially relative levels between two cultivars.
Functional annotation and pathway analysis of differentially abundant proteins
The differentially abundant proteins from Teqing and Lemont were analyzed to extract information relevant to the pathways involved in R. solani infection. After KEGG pathway analysis, the top ten significantly enriched terms according to the GO hierarchy (level four) were depicted in Fig. 4. In the biological process analysis, cellular amino acid metabolism was the most significant term (p-value: 2.37e−09), followed by processes such as organonitrogen and alpha-amino acid metabolic (Supplementary Table S4) in Teqing. In contrast, plastid organization was the most representative term (p-value: 2.12e−08) in Lemont and was followed by other biological processes such as chloroplast organization and single-organism metabolic process (Supplementary Table S4). The cell component analysis showed that 21% of annotated proteins were located in the cytoplasm in Teqing (Supplementary Table S5), and 18% annotated proteins were located in the cytoplasm in Lemont (Supplementary Table S5). The GO analysis showed that oxidoreductase activity and GTP binding were the dominant molecular function in Teqing and Lemont, respectively (Supplementary Table S6).
The KEGG pathway enrichment analysis revealed that the differentially abundant proteins at p < 0.01 in the resistant cultivar were mainly involved in glyoxylate and dicarboxylate metabolism; glycine, serine and threonine metabolism; and unsaturated fatty acid biosynthesis pathways (Fig. 5a, Supplementary Table S7). However, the differentially abundant proteins in susceptible plants were mainly involved in the glycolysis/gluconeogenesis regulation (Fig. 5b, Supplementary Table S7). We further focused on 78 significantly differentially abundant proteins based on an overall evaluation of their KEGG analyses and functions. The first 64 proteins are involved in the pathways identified by KEGG analyses, and the last 14 common proteins participate in the signal transduction which have an important role in the course of rice disease resistance (Supplementary Table S9). These candidate proteins were classified into 12 functional categories: pyruvate metabolism; proteasome; photosynthesis; oxidative phosphorylation; metabolic pathways; glyoxylate and dicarboxylate metabolism; glycolysis/gluconeogenesis; glycine, serine and threonine metabolism; carbon fixation in photosynthetic organisms; biosynthesis of unsaturated fatty acids; alpha-Linolenic acid metabolism; and signal transduction. We linked these candidate proteins using a protein interaction network. Functional protein association networks generated with STRING revealed the functional links between different proteins. Three major clusters of interacting proteins in the resistant cultivar are highlighted with circles in Fig. 6a. These proteins are involved in glyoxylate and dicarboxylate metabolism; glycine, serine and threonine metabolism; and unsaturated fatty acid biosynthesis. Ferredoxin-dependent glutamate synthase (Q69RJ0) is the central core protein of this interaction network, due to its interactions with many other proteins involved in glyoxylate/dicarboxylate and glycine/serine/threonine metabolism. A glycolysis/gluconeogenesis protein interaction network was constructed in the susceptible cultivar (Fig. 6b). Enolase 2 (Q10P35), which interacted with other proteins directly or indirectly, is the central core protein of this interaction network.
Validation of protein profiles by qRT-PCR
To investigate whether changes in the expression of proteins were regulated at the transcriptional level, the expression pattern of 12 selected genes which have roles in disease resistance was further validated experimentally by qRT-PCR. As shown in Fig. 7, PR1 (Os07g0129300), PBZ1 (Os12g36880), LOX (rci-1), BAP (OJ1509C06.18) and PAL (PAL) were up-regulated in both disease-resistant and disease-susceptible cultivars after 24 h and 48 h of infection. These genes may represent immune regulators acting in basal responses, since their expression was increased in both cultivars upon infection. However, their expression levels were different in the two cultivars. For example, PBZ1 showed a higher expression level in Teqing than in Lemont after pathogen stress. PR2 (Gns6), MDAR (Os08g0557600) and POD (APX1) were induced in only the disease-resistant cultivar. In contrast, MDH (OsI29345) was induced in the susceptible cultivar but down-regulated in the resistant cultivar after R. solani infection. Therefore, it may negatively regulate rice immunity. However, PR1 expression was induced approximately fourfold and sevenfold in Teqing and Lemont, respectively, at 48 hpi following R. solani infection. The expression level of CS (OsI06215) was similar in the two cultivars against R. solani infection (data not shown). In the proteomic analysis, this protein was unchanged in Lemont but increased in abundance in Teqing.
Assessment of protein function
To further understand the role of the differently abundant proteins in plant immunity, a total of four proteins were chosen for function assessment based on their protein expression during infection and the putative functions. PR1 which involved in SA signal pathway, was induced in both cultivars (Fig. 7). PBZ1, a JA signal pathway inducible protein, was also induced in both cultivars (Fig. 7). Monodehydroascorbate reductase (MDAR) is crucial for Ascorbate (AsA) regeneration and essential for maintaining a reduced pool of AsA. Overexpression of MDAR confers enhanced resistance against ozone, salt and polyethylene glycol (PEG) stress (Eltayeb et al. 2007). Blight-associated proteins (BAPs) are found throughout the plant, most abundantly in the xylem and vascular cambial zone of roots and stems. One BAP that was analyzed and selected for further characterization was found to resemble β-1,3-glucanase (Derrick et al. 1993). In this study, both MDAR (Q6ZJ08) and BAP proteins (Q6K4C4) were increased in the resistant cultivar. We introduced the open reading frames of candidate protein under the control of the CaMV 35S promoter into Nicotiana benthamiana and inoculated the leaves with R. solani at 48 h after agroinfiltration. Compared with the empty vector (EV), overexpression of these proteins in N. benthamiana increased resistance to R. solani (Fig. 8a). In addition, small lesions were observed on leaves expressing each individual protein at 48 hpi (Fig. 8b).
Functional tests of proteins in promoting resistance to R. solani infection. a Photos of agroinfiltrated N. benthamiana leaves expressing recombinant proteins and subsequently inoculated with R. solani. b Lesion sizes on R. solani-inoculated N. benthamiana leaves expressing recombinant proteins. Different letters indicate significant differences based on one-way ANOVA (p < 0.05). Similar results were obtained from three biological replicates
Discussion
R. solani is an important soil-borne pathogen that causes significant damage to rice globally (Zheng et al. 2013). To date, little is known about the mechanisms of host defense in response to infection. In this study, an iTRAQ-based comparative proteomic analysis between the resistant (Teqing) and susceptible (Lemont) rice cultivars was employed to investigate the potential functional proteins related to challenge with the fungal pathogen R. solani. A total of 755 differentially expressed proteins were identified in the resistant and susceptible rice cultivars after inoculation with R. solani (Table S2).
Influence of R. solani infection on glyoxylate and dicarboxylate metabolism
In plants, the glyoxylate cycle has a key role in conversion of acetyl coenzyme A (CoA) into oxaloacetate via fatty acid β-oxidation and in the subsequent conversion into sugar (Beevers 1961). In this proteomic study, five proteins, including ferredoxin-dependent glutamate synthase (Q69RJ0), glycine decarboxylase complex H-protein (Q655T6), formate dehydrogenase (Q0DC43), ribulose bisphosphate carboxylase small chain (Q67IU5) and pyridoxal phosphate-dependent transferase (Q6ZFI6) were identified (Supplementary Table S9). Formate dehydrogenase (FDH) is used to oxidize formate to CO2 (Garrett and Grisham 2000), which is involved in abiotic and biotic stress responses in a variety of higher plants (Hourton-Cabassa et al. 1998; Suzuki et al. 1998; David et al. 2010). Our results showed that FDH was up-regulated in the resistant cultivar (Supplementary Table S9), suggesting putative role in immunity to R. solani. Ribulose bisphosphate carboxylase small chain (RBCS) is known to be one of the most important enzymes involved in glyoxylate and dicarboxylate metabolism (Yang et al. 2015). RBCS increased in both the susceptible and resistant rice cultivars after pathogen challenge (Supplementary Table S9). This result may indicate that increased glyoxylate and dicarboxylate metabolism is needed to support basic defense during rice immunity to R. solani. Ferredoxin-dependent glutamate synthase (Fd-GOGAT) is a chloroplastic enzyme responsible for the reassimilation of photorespiratory ammonia as well as for primary nitrogen assimilation, with concomitant consumption of both ATP and reducing power (Leegood et al. 1995; Douce et al. 2001). In our study, Fd-GOGAT decreased in the resistant cultivar and increased in the susceptible cultivar. It is likely that decreasing of Fd-GOGAT reduced energy consumption in resistant cultivar than in susceptible cultivar response to R. solani infection, which may be an important reason why Teqing is more resistance than Lemont.
Proteins involved in glycolysis/gluconeogenesis
Glycolysis/gluconeogenesis is a catabolic anaerobic pathway that oxidizes hexoses to generate ATP, reducing agents, and pyruvate and produces building blocks for anabolism (Plaxton 1996). In this study, dihydrolipoyl dehydrogenase (Q94CN9), triosephosphate isomerase (Q69K00), 2,3-bisphosphoglycerate-independent phosphoglycerate mutase (Q5KQH5), enolase 2 (Q10P35), phosphoenolpyruvate carboxykinase (Q10NX2) and alcohol dehydrogenase 2 (Q10A46) were identified as significantly differentially expressed proteins (Supplementary Table S9), providing evidence that the glycolysis/gluconeogenesis pathway (Houde and Diallo 2008) is involved in response to infection. Alteration in glycolysis/gluconeogenesis metabolism has also been reported previously during R. solani infection (Mutuku and Nose 2012; Ghosh et al. 2017). The abundance of all identified proteins related to glycolysis (except alcohol dehydrogenase 2) increased in the susceptible cultivar, but no variation was observed in the resistant cultivar, indicating that the susceptible cultivar needs more ATP by strengthening the glycolysis/gluconeogenesis pathway under R. solani infection.
Glycine, serine and threonine metabolism-related proteins
A general symptom of photosynthetic plants under stress is energy deficiency (Baena-Gonzalez et al. 2007), glycine, serine and threonine metabolism is vital to the regulation of plant energy metabolism(Ma et al. 2017; Igamberdiev and Kleczkowski 2018; Matityahu et al. 2019). In response to high soil temperature, glycine, serine, and threonine accumulate in the roots of foxtail millet, which accumulates more protective metabolites and is more resistant to high soil temperature than rice (Aidoo et al. 2016). By identification of genes underlying drought resistance, alanine-glyoxylate aminotransferase 2 (AGT) was found to have significant role in energy and carbon metabolism (Sehgal et al. 2012). In our study, tryptophan synthase alpha chain protein (Q6ZL61) abundance was increased in the resistant cultivar, and malate dehydrogenase (MDH) protein (Q6YYW3) was down-regulated in the susceptible cultivar. Moreover, AGT (Q10R45) displayed higher induction in the resistant cultivar than that in the susceptible cultivar (Supplementary Table S9). These results also suggest that the resistant cultivar perceived environmental stress and reduced energy consumption under infection.
Biosynthesis of unsaturated fatty acids
In this study, two 3-ketoacyl-CoA thiolase (KAT) proteins (Q94LR9 and Q84P96) showed increased abundance in the resistant cultivar (Supplementary Table S9). KAT catalysis is a key step in fatty acid β-oxidation, which has an important role in seed development, via thiolase activity (Germain et al. 2001). KAT catalysis is a key step in fatty acid β-oxidation, which is a multistep process to produce energy (Lopaschuk et al. 2010). However, the exact role of these proteins in rice immunity remains unknown.
Potential model of rice immunity under R. solani infection
Based on the functions of the differentially expressed proteins and the expression patterns found using comparative proteomics, qRT-PCR and protein function, a possible mechanism is proposed for rice resistance to R. solani (Fig. 9). This network consists of several functional components, such as the SA and JA signaling pathways, reactive oxygen species (ROS) production and the TCA cycle.
SA plays a role in rice resistance to pathogen infection (Li et al. 2012). Phenylalanine ammonia-lyase (PAL) is a crucial enzyme in phenylpropanoid metabolism, catalyzing the formation of trans-cinnamic acid via the l-deamination of phenylalanine, which is a key regulator of SA accumulation (Sendon et al. 2011). The abundance of PAL proteins was up-regulated in both the resistant (A0A0E0K2L5 and Q7X720) and susceptible rice cultivars (Q75HQ7 and I1P2D6) (Supplementary Table S9), but the expression levels were higher in Teqing than in Lemont (Fig. 8). The marker genes PR1 and PR2 in the SA signal pathway were induced strongly in the resistant cultivar, but not in the susceptible cultivar (Supplementary Table S9). However, in the transcriptional analysis, the expression of PR1 was more highly up-regulated in Lemont than in Teqing (Fig. 8). The difference between the qRT-PCR and proteomic results may be due to mRNA stability, splicing, and translational regulation (Fabian et al. 2010; Brown et al. 2015). So, we assume that resistance to R. solani mediated by SA is due to higher levels of PR1 and PR2, which in turn give higher resistance.
JA signaling also plays an important role in rice basal defense against pathogen infection (Mei et al. 2006; Yamada et al. 2012). LOX catalyzes the oxygenation of linoleic polyunsaturated fatty acids, resulting in the formation of hydroperoxides important for JA biosynthesis (Christensen et al. 2014). AOS2 encodes an allene oxide synthase, a key enzyme in JA biosynthesis, which plays an important role in resistance to rice blast (Mei et al. 2006). Incereased expression of LOX and the defense gene, PBZ1, can enhance resistance to R. solani and M. oryzae (Peng et al. 2012). In the present study, LOX protein abundance in the susceptible cultivar (Q9FSE5, A0A0E0QL12, and I1PFA2) was higher than in the resistant cultivar (Supplementary Table S9). However, LOX expression level was higher in the resistant than that in the susceptible cultivar at 48 hpi according to qRT-PCR (Fig. 8). The AOS protein (Q10EK5) was up-regulated in only the resistant cultivar (Supplementary Table S9). This result implies that the AOS protein is likely a key factor in JA biogenesis when rice is challenged with R. solani. Indeed, PBZ1, which is a JA-inducible gene (Mahmood et al. 2006), was more up-regulated in the resistant cultivar than in the susceptible cultivar according to qRT-PCR (Fig. 8).
The generation of ROS is one of the first universal reactions to abiotic or biotic challenges in plants (Torres 2010; Suzuki et al. 2012). ROS can be either beneficial or harmful to cells and tissues. Plants have evolved complex regulatory mechanisms to maintain steady-state levels of ROS and to respond to different environmental and developmental signals (Mittler et al. 2004; Circu and Aw 2010). The peroxidase family plays a major role in regulating the levels of ROS, and these detoxifying enzymes oxidize a wide variety of compounds in the presence of H2O2 (Teixeira et al. 2004). In this study, ten (APX1_ORYSI, Q5U1S3, Q5U1T0, Q9ST80, O22438, Q5U1Q2, Q9FYP0, Q5U1F5, Q6AVZ3 and Q6AVZ3) and seven peroxidase homologous proteins (Q7XSU7, A0A0P0VZ16, Q6K4J4, Q9SMG8, Q9FYP0, Q9LDL0 and A0A0E0H918) (Supplementary Table S9) with putative functions in ROS level regulation were detected in the resistant and susceptible cultivars, respectively. In the resistant cultivar, five proteins were up-regulated, and five were down-regulated. In the susceptible cultivar, six proteins were up-regulated, and one protein was down-regulated. Interestingly, one peroxidase protein (Q9FYP0) had different expression patterns in the susceptible and resistant cultivars, suggesting that plants have different strategies in response to pathogen infection depending on the plant variety. ROS have been reported to play a role in the interaction between R. solani and wheat (Foley et al. 2016). The accumulation of ROS may be essential for cell death in the infected tissues.
The TCA cycle is one of the iconic pathways in metabolism and is associated with energy metabolism or the oxidation of respiratory substrates (Sweetlove et al. 2010). In the present study, two identified proteins were involved in the TCA cycle. Citrate synthase (Q9FUJ7) was up-regulated in only the resistant cultivar (Supplementary Table S9). However, MD was down-regulated in the resistant cultivar (A0A0E0KJK7) and up-regulated in the susceptible cultivar (A2YVI5), which was consistent with the transcriptional data (Supplementary Table S9 and Fig. 8). The increase in the levels of metabolic proteins related to the TCA cycle indicates enhanced respiration in rice after infection. Increased respiration has previously been observed during the interaction between R. solani and rice (Suharti et al. 2016; Ghosh et al. 2017). Plants usually increase respiration to support the induction of defense programs upon pathogen infection (Berger et al. 2007). Our recent study found that SA and JA signal pathways are also involved in defense response to the blast disease (Liu et al. 2014). In contrast, ROS production and TCA cycle are only found in rice immunity against R. solani infection. So we inferred that SA and JA are the general defense machinery to broad-spectrum pathogen.
Conclusions
In conclusion, our findings provide a possible model by which multiple signal pathways are activated to regulate rice immunity following R. solani infection. These pathways do not function separately; instead, they work together as an integral network via multilateral cross talk. However, further research is needed to explore the importance of these proteins and their potential roles during the establishment of rice sheath blight disease.
Abbreviations
- 2-DE:
-
Two-dimensional electrophoresis
- DEPs:
-
Differentially expressed proteins
- EF:
-
Error factor
- EV:
-
Empty vector
- FDR:
-
False discovery Rate
- GC–MS:
-
Gas chromatography–mass spectrometry
- GO:
-
Gene ontology
- iTRAQ:
-
Isobaric tags for relative and absolute quantification
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- LC–MS:
-
Liquid chromatography–tandem mass spectrometry
- MDH:
-
Malate dehydrogenase
- TFA:
-
Trifluoroacetic acid
- PDA:
-
Potato dextrose agar
- PR:
-
Pathogenesis-related
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- ROS:
-
Reactive oxygen species
- SA:
-
Salicylic acid
- TCA:
-
Tricarboxylic acid cycle
References
Aidoo MK, Bdolach E, Fait A, Lazarovitch N, Rachmilevitch S (2016) Tolerance to high soil temperature in foxtail millet (Setaria italica L.) is related to shoot and root growth and metabolism. Plant Physiol Biochem 106:73–81
Anderson NA (1982) The genetics and pathology of Rhizoctonia solani. Annu Rev Phytopathol 20:329–347
Baena-Gonzalez E, Rolland F, Thevelein JM, Sheen J (2007) A central integrator of transcription networks in plant stress and energy signalling. Nature 448:938U910
Bandehagh A, Salekdeh GH, Toorchi M, Mohammadi A, Komatsu S (2011) Comparative proteomic analysis of canola leaves under salinity stress. Proteomics 11(10):1965–1975
Beevers H (1961) Metabolic production of sucrose from fat. Nature 191:433–436
Berard AR, Coombs KM, Severini A (2015) Quantification of the host response proteome after herpes simplex virus type 1 infection. J Proteome Res 14(5):2121–2142
Berger S, Sinha AK, Roitsch T (2007) Plant physiology meets phytopathology: plant primary metabolism and plant-pathogen interactions. J Exp Bot 58:4019–4026
Bollich CN, Webb BD, Marchetti MA, Scott JE (1985) Registration of lemont rice. Crop Sci 25(5):883–885
Bonman JM, Khush GS, Nelson RJ (1992) Breeding rice for resistance to pests. Annu Rev Phytopathol 30:507–528
Brown JWS, Simpson CG, Marquez Y, Gadd GM, Barta A, Kalyna M (2015) Lost in translation: pitfalls in deciphering plant alternative splicing transcripts. Plant Cell 27(8):2083–2087
Chen Z, Guo Q, Chen BH, Li XY, Wang ZC, He P, Yan F, Hu DY, Yang S (2014) Development of proteomic technology of shotgun and label free combined with multiple reaction monitoring to simultaneously detect southern rice black-streaked dwarf virus and rice ragged stunt virus. Virusdisease 25(3):322–330
Chmielewska K, Rodziewicz P, Swarcewicz B, Sawikowska A, Krajewski P, Marczak L, Ciesiolka D, Kuczynska A, Mikolajczak K, Ogrodowicz P, Krystkowiak K, Surma M, Adamski T, Bednarek P, Stobiecki M (2016) Analysis of drought-induced proteomic and metabolomic changes in barley (Hordeum vulgare L.) leaves and roots unravels some aspects of biochemical mechanisms involved in drought tolerance. Front Plant Sci 7:1108
Christensen SA, Nemchenko A, Park YS, Borrego E, Huang PC, Schmelz EA, Kunze S, Feussner I, Yalpani N, Meeley R, Kolomiets MV (2014) The novel monocot-specific 9-lipoxygenase ZmLOX12 is required to mount an effective jasmonate-mediated defense against Fusarium verticillioides in maize. Mol Plant Microbe Interact 27(11):1263–1276
Circu ML, Aw TY (2010) Reactive oxygen species, cellular redox systems, and apoptosis. Free Radic Biol Med 48(6):749–762
Datta K, Velazhahan R, Oliva N, Ona I, Mew T, Khush GS, Muthukrishnan S, Datta SK (1999) Over-expression of the cloned rice thaumatin-like protein (PR-5) gene in transgenic rice plants enhances environmental friendly resistance to Rhizoctonia solani causing sheath blight disease. Theor Appl Gent 98:1138–1145
Datta K, Koukolikova-Nicola Z, Baisakh N, Oliva N, Datta SK (2000) Agrobacterium-mediated engineering for sheath blight resistance of indica rice cultivars from different ecosystems. Theor Appl Genet 100(6):832–839
David P, des Francs-Small CC, Sevignac M, Thareau V, Macadre C, Langin T, Geffroy V (2010) Three highly similar formate dehydrogenase genes located in the vicinity of the B4 resistance gene cluster are differentially expressed under biotic and abiotic stresses in Phaseolus vulgaris. Theor Appl Gent 121(1):87–103
Delaunois B, Colby T, Belloy N, Conreux A, Harzen A, Baillieu F, Clement C, Schmidt J, Jeandet P, Cordelier S (2013) Large -scale proteomic annalysis of the grapevine apoplastic fluid revels mainly stress-related proteins and cell wall modifying enzymes. BMC Plant Biol 13:24
Delaunois B, Jeandet P, Clement C, Baillieu F, Dorey S, Cordelier S (2014) Uncovering plant-pathogen crosstalk through apoplastic proteomic studies. Front Plant Sci 5:249
Derrick KS BG, Hewitt BG, Lee RF (1993) Serological tests for citrus blight. In: Moreno P, da Graca JV, Timmer LW (eds) Proceedings 12th Conference International Organization of Citrus Virologists, New Delhi, India, pp 121–126
Douce R, Bourguignon J, Neuburger M, Rebeille F (2001) The glycine decarboxylase system: a fascinating complex. Trends Plant Sci 6(4):167–176
Du B, Wei Z, Wang ZQ, Wang XX, Peng XX, Du B, Chen RZ, Zhu LL, He GC (2015) Phloem-exudate proteome analysis of response to insect brown plant-hopper in rice. J Plant Physiol 183:13–22
Eltayeb AE, Kawano N, Badawi GH, Kaminaka H, Sanekata T, Shibahara T, Inanaga S, Tanaka K (2007) Overexpression of monodehydroascorbate reductase in transgenic tobacco confers enhanced tolerance to ozone, salt and polyethylene glycol stresses. Planta 225(5):1255–1264
Fabian MR, Sonenberg N, Filipowicz W (2010) Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem 79:351–379
Faghani E, Gharechahi J, Komatsu S, Mirzaei M, Khavarinejad RA, Najafi F, Farsad LK, Salekdeh GH (2015) Comparative physiology and proteomic analysis of two wheat genotypes contrasting in drought tolerance. J Proteom 114:1–15
Fatehi F, Hosseinzadeh A, Alizadeh H, Brimavandi T, Struik PC (2012) The proteome response of salt-resistant and salt-sensitive barley genotypes to long-term salinity stress. Mol Biol Rep 39(5):6387–6397
Foley RC, Kidd BN, Hane JK, Anderson JP, Singh KB (2016) Reactive oxygen species play a role in the infection of the necrotrophic fungi, Rhizoctonia solani in wheat. PLoS One 11(3):e0152548
Garrett RH, Grisham CM (2000) Biochimie. Ann Biol Clin (Paris) 58(6):767–768
Germain V, Rylott EL, Larson TR, Sherson SM, Bechtold N, Carde JP, Bryce JH, Graham IA, Smith SM (2001) Requirement for 3-ketoacyl-CoA thiolase-2 in peroxisome development, fatty acid beta-oxidation and breakdown of triacylglycerol in lipid bodies of Arabidopsis seedlings. Plant J 28(1):1–12
Ghosh S, Gupta SK, Jha G (2014) Identification and functional analysis of AG1-IA specific genes of Rhizoctonia solani. Curr Genet 60:327–341
Ghosh S, Kanwar P, Jha G (2017) Alterations in rice chloroplast integrity, photosynthesis and metabolome associated with pathogenesis of Rhizoctonia solani. Sci Rep 7:41610
Gonzalez JF, Degrassi G, Devescovi G, De Vleesschauwer D, Hofte M, Myers MP, Venturi V (2012) A proteomic study of Xanthomonas oryzae pv. oryzae in rice xylem sap. J Proteom 75(18):5911–5919
Guo GF, Ge P, Ma CY, Li XH, Lv DW, Wang SL, Ma WJ, Yan YM (2012) Comparative proteomic analysis of salt response proteins in seedling roots of two wheat varieties. J Proteom 75(6):1867–1885
Houde M, Diallo AO (2008) Identification of genes and pathways associated with aluminum stress and tolerance using transcriptome profiling of wheat near-isogenic lines. Bmc Genom 9:400
Hourton-Cabassa C, Ambard-Bretteville F, Moreau F, de Virville JD, Remy R, des Francs-Small CC (1998) Stress induction of mitochondrial formate dehydrogenase in potato leaves. Plant Physiol 116(2):627–635
Igamberdiev AU, Kleczkowski LA (2018) The glycerate and phosphorylated pathways of serine synthesis in plants: the branches of plant glycolysis linking carbon and nitrogen metabolism. Front Plant Sci 9:318
Jha S, Chattoo BB (2010) Expression of a plant defensin in rice confers resistance to fungal phytopathogens. Transgenic Res 19(3):373–384
Kim ST, Kim SG, Hwang DH, Kang SY, Kim HJ, Lee BH, Lee JJ, Kang KY (2004) Proteomic analysis of pathogen-responsive proteins from rice leaves induced by rice blast fungus, Magnaporthe grisea. Proteomics 4(11):3569–3578
Komatsu S, Tanaka N (2005) Rice proteome analysis: a step toward functional analysis of the rice genome. Proteomics 5(4):938–949
Kumar KVK, Raju SK, Reddy MS, Kloepper JW, Lawrence KS, Groth DE, Miller ME, Sudini H, Du BH (2009) Evaluation of commercially available PGPR for control of rice sheath blight caused by Rhizoctonia solani. J Pure Appl Microbiol 3(2):485–488
Lee FN, Rush MC (1983) Rice sheath blight: a major rice disease. Plant Dis 67:829–832
Lee J, Bricker TM, Lefevre M, Pinson SR, Oard JH (2006) Proteomic and genetic approaches to identifying defence-related proteins in rice challenged with the fungal pathogen Rhizoctonia solani. Mol Plant Pathol 7(5):405–416
Leegood RC, Lea PJ, Adcock MD, Hausler RE (1995) The regulation and control of photorespiration. J Exp Bot 46:1397–1414
Li Y, Zhang Z, Nie Y, Zhang L, Wang Z (2012) Proteomic analysis of salicylic acid-induced resistance to Magnaporthe oryzae in susceptible and resistant rice. Proteomics 12(14):2340–2354
Liu GT, Ma L, Duan W, Wang BC, Li JH, Xu HJ, Yan XQ, Yan BF, Li SH, Wang LJ (2014) Differential proteomic analysis of grapevine leaves by iTRAQ reveals responses to heat stress and subsequent recovery. BMC Plant Biol 14:110–127
Lopaschuk GD, Ussher JR, Folmes CD, Jaswal JS, Stanley WC (2010) Myocardial fatty acid metabolism in health and disease. Physiol Rev 90:207–258
Ma HY, Song LR, Shu YJ, Wang S, Niu J, Wang ZK, Yu T, Gu WH, Ma H (2012) Comparative proteomic analysis of seedling leaves of different salt tolerant soybean genotypes. J Proteom 75(5):1529–1546
Ma Q, Cao X, Xie Y, Xiao H, Tan X, Wu L (2017) Effects of glucose on the uptake and metabolism of glycine in pakchoi (Brassica chinensis L.) exposed to various nitrogen sources. BMC Plant Biol 17(1):58
Mahmood T, Jan A, Kakishima M, Komstsu S (2006) Proteomic analysis of bacterial-blight defense-responsive proteins in rice leaf blades. Proteomics 6:6053–6065
Matityahu I, Godo I, Hacham Y, Amir R (2019) The level of threonine in tobacco seeds is limited by substrate availability, while the level of methionine is limited also by the activity of cystathionine gamma-synthase. Plant Sci 283:195–201
Mei C, Qi M, Sheng G, Yang Y (2006) Inducible overexpression of a rice allene oxide synthase gene increases the endogenous jasmonic acid level, PR gene expression, and host resistance to fungal infection. Mol Plant Microbe Interact 19(10):1127–1137
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9(10):490–498
Mutuku JM, Nose A (2012) Changes in the contents of metabolites and enzyme activities in Rice plants responding to Rhizoctonia solani Kuhn infection: activation of glycolysis and connection to phenylpropanoid pathway. Plant Cell Physiol 53(6):1017–1032
Neilson KA, Mariani M, Haynes PA (2011a) Quantitative proteomic analysis of cold-responsive proteins in rice. Proteomics 11(9):1696–1706
Neilson KA, Mariani M, Haynes PA (2011b) Quantitative proteomic analysis of cold-responsive proteins in rice. Proteomics 11(9):1696–1706
Ogoshi A (1987) Ecology and pathogenicity of anastomosis and intraspecific groups of Rhizoctonia-Solani Kuhn. Annu Rev Phytopathol 25:125–143
Pan XB, Rush MC, Sha XY, Xie QJ, Linscombe SD, Stetina SR, Oard JH (1999) Major gene, nonallelic sheath blight resistance from the rice cultivars Jasmine 85 and Teqing. Crop Sci 39(2):338–346
Patkar RCB (2006) Transgenic indica rice expressing ns-LTP-like protein shows enhanced resistance to both fungal and bacterial pathogens. Mol Breed 17:159–171
Peng XX, Hu YJ, Tang XK, Zhou PL, Deng XB, Wang HH, Guo ZJ (2012) Constitutive expression of rice WRKY30 gene increases the endogenous jasmonic acid accumulation, PR gene expression and resistance to fungal pathogens in rice. Planta 236(5):1485–1498
Pinson SRM, Liu GJ, Jia MH, Jia YL, Fjellstrom RG, Sharma A, Wang YG, Tabien RE, Li ZK (2012) Registration of a rice gene-mapping population consisting of ‘TeQing’-into-’Lemont’ backcross introgression lines. J Plant Regist 6(1):128–135
Plaxton WC (1996) The organization and regulation of plant glycolysis. Annu Rev Plant Physiol Plant Mol Biol 47:185–214
Sarhadi E, Bazargani MM, Sajise AG, Abdolahi S, Vispo NA, Arceta M, Nejad GM, Singh RK, Salekdeh GH (2012) Proteomic analysis of rice anthers under salt stress. Plant Physiol Biochem 58:280–287
Sehgal D, Rajaram V, Armstead IP, Vadez V, Yadav YP, Hash CT, Yadav RS (2012) Integration of gene-based markers in a pearl millet genetic map for identification of candidate genes underlying drought tolerance quantitative trait loci. BMC Plant Biol 12:9
Sendon PM, Seo HS, Song JT (2011) Salicylic Acid signaling: biosynthesis, metabolism, and crosstalk with jasmonic acid. J Korean Soc Appl Biol Chem 54(4):501–506
Sridevi G, Sabapathi N, Meena P, Nandakumar R, Samiyappan R, Muthukrishnan S, Veluthambi K (2003) Transgenic indica rice variety Pusa Basmati 1 constitutively expressing a rice chitinase gene exhibits enhanced resistance to Rhizoctonia solani. J Plant Biochem Biotechnol 12(2):93–101
Sripriya R, Raghupathy V, Veluthambi K (2008) Generation of selectable marker-free sheath blight resistant transgenic rice plants by efficient co-transformation of a cointegrate vector T-DNA and a binary vector T-DNA in one Agrobacterium tumefaciens strain. Plant Cell Rep 27(10):1635–1644
Suharti WS, Nose A, Zheng SH (2016) Metabolomic study of two rice lines infected by Rhizoctonia solani in negative ion mode by CE/TOF-MS. J Plant Physiol 206:13–24
Suzuki K, Itai R, Suzuki K, Nakanishi H, Nishizawa NK, Yoshimura E, Mori S (1998) Formate dehydrogenase, an enzyme of anaerobic metabolism, is induced by iron deficiency in barley roots. Plant Physiol 116(2):725–732
Suzuki N, Koussevitzky S, Mittler R, Miller G (2012) ROS and redox signalling in the response of plants to abiotic stress. Plant Cell Environ 35(2):259–270
Sweetlove LJ, Beard KF, Nunes-Nesi A, Fernie AR, Ratcliffe RG (2010) Not just a circle: flux modes in the plant TCA cycle. Trends Plant Sci 15(8):462–470
Tabien RE, Li Z, Paterson AH, Marchetti MA, Stansel JW, Pinson SRM (2000) Mapping of four major rice blast resistance genes from ‘Lemont’ and ‘Teqing’ and evaluation of their combinatorial effect for field resistance. Theor Appl Gent 101(8):1215–1225
Taheri PHM (2007) Riboflavin-induced resistance against rice sheath blight functions through the potentiation of lignin formation and jasmonic acid signalling pathway. Commun Agric Appl Biol Sci 72:309–313
Teixeira FK, Menezes-Benavente L, Margis R, Margis-Pinheiro M (2004) Analysis of the molecular evolutionary history of the ascorbate peroxidase gene family: inferences from the rice genome. J Mol Evol 59(6):761–770
Tian X, Liu Y, Huang ZG, Duan HP, Tong JH, He XL, Gu WH, Ma H, Xiao LT (2015) Comparative proteomic analysis of seedling leaves of cold-tolerant and -sensitive spring soybean cultivars. Mol Biol Rep 42(3):581–601
Torres MA (2010) ROS in biotic interactions. Physiol Plant 138(4):414–429
Wang HH, Meng J, Peng XX, Tang XK, Zhou PL, Xiang JH, Deng XB (2015) Rice WRKY4 acts as a transcriptional activator mediating defense responses toward Rhizoctonia solani, the causing agent of rice sheath blight. Plant Mol Biol 89:157–171
Wrather JA, Anderson TR, Arsyad DM, Gai J, Ploper LD, PortaPuglia A, Ram HA, Yorinori JT (1997) Soybean disease loss estimates for the top 10 soybean producing countries in 1994. Plant Dis 81(1):107–110
Wu Y, Mirzaei M, Pascovici D, Chick JM, Atwell BJ, Haynes PA (2016) Quantitative proteomic analysis of two different rice varieties reveals that drought tolerance is correlated with reduced abundance of photosynthetic machinery and increased abundance of ClpD1 protease. J Proteomics 143:73–82
Yamada S, Kano A, Tamaoki D, Miyamoto A, Shishido H, Miyoshi S, Taniguchi S, Akimitsu K, Gomi K (2012) Involvement of OsJAZ8 in jasmonate-induced resistance to bacterial blight in rice. Plant Cell Physiol 53(12):2060–2072
Yang SY, Hao DL, Song ZZ, Yang GZ, Wang L, Su YH (2015) RNA-Seq analysis of differentially expressed genes in rice under varied nitrogen supplies. Gene 555(2):305–317
Zhao J, Yang Y, Kang Z (2014) Proteomic analysis of rice nonhost resistance to Puccinia striiformis f. sp. tritici using two-dimensional electrophoresis. Int J Mol Sci 15(12):21644–21659
Zheng TQ, Wang Y, Ali AJ, Zhu LH, Sun Y, Zhai HQ, Mei HW, Xu ZJ, Xu JL, Li ZK (2011) Genetic effects of background-independent loci for grain weight and shape identified using advanced reciprocal introgression lines from lemont × teqing in rice. Crop Sci 51(6):2525–2534
Zheng A, Lin R, Zhang D, Qin P, Xu L, Ai P, Ding L, Wang Y, Chen Y, Liu Y, Sun Z, Feng H, Liang X, Fu R, Tang C, Li Q, Zhang J, Xie Z, Deng Q, Li S, Wang S, Zhu J, Wang L, Liu H, Li P (2013) The evolution and pathogenic mechanisms of the rice sheath blight pathogen. Nat Commun 4:1424
Acknowledgements
The authors thank the National Mid-term Genebank for Rice of China National Rice Research Institute for providing the rice cultures of Teqing and Lemont. This work was supported by the Natural Science Foundation of Jiangsu Province of China (BK20171382), the Fundamental Research Funds for the Central Universities (Y0201700152, KJQN201916) and the National Natural Science Foundation of China (31501621, 31801778).
Author information
Authors and Affiliations
Contributions
DN conceived the idea and designed the project; HM, CS, and LQ performed the experiments; DN, HM, CS, and HZ analysed and interpreted the data; DN, HM and CS wrote the manuscript; all authors read and approved the final manuscript for publication.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ma, H., Sheng, C., Qiao, L. et al. A comparative proteomic approach to identify defence-related proteins between resistant and susceptible rice cultivars challenged with the fungal pathogen Rhizoctonia solani. Plant Growth Regul 90, 73–88 (2020). https://doi.org/10.1007/s10725-019-00551-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-019-00551-w