Abstract
The WHO 2021 classification defines IDH wild type (IDHw) histologically lower-grade glioma (hLGG) as molecular glioblastoma (mGBM) if TERT promoter mutation (pTERTm), EGFR amplification or chromosome seven gain and ten loss aberrations are indicated. We systematically reviewed articles of IDHw hLGGs studies (49 studies, N = 3748) and meta-analyzed mGBM prevalence and overall survival (OS) according to the PRISMA statement. mGBM rates in IDHw hLGG were significantly lower in Asian regions (43.7%, 95% confidence interval [CI: 35.8–52.0]) when compared to non-Asian regions (65.0%, [CI: 52.9–75.4]) (P = 0.005) and were significantly lower in fresh-frozen specimen when compared to formalin-fixed paraffin-embedded samples (P = 0.015). IDHw hLGGs without pTERTm rarely expressed other molecular markers in Asian studies when compared to non-Asian studies. Patients with mGBM had significantly longer OS times when compared to histological GBM (hGBM) (pooled hazard ratio (pHR) 0.824, [CI: 0.694–0.98], P = 0.03)). In patients with mGBM, histological grade was a significant prognostic factor (pHR 1.633, [CI: 1.09–2.447], P = 0.018), as was age (P = 0.001) and surgical extent (P = 0.018). Although bias risk across studies was moderate, mGBM with grade II histology showed better OS rates when compared to hGBM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diffuse gliomas with histologically lower-grade features (hLGG) are heterogeneous tumors with different clinical outcomes. Some patients may experience long stable disease, while others rapidly deteriorate with fatal outcomes. Classifications based on histological findings have been used to discriminate subgroups with distinct prognosis outcomes. Molecular studies have reported hLGG sub-classification into different molecular subgroups [13, 14, 17, 55]. The first group is IDH mutant-type (IDHm) and chromosome 1p/19q co-deleted tumors which correspond to oligodendrogliomas. These tumors generally have good prognoses. The second group is IDHm with intact 1p/19q and alterations in ATRX and TP53, corresponding to astrocytomas. In contrast, IDH wild type (IDHw) hLGG are molecularly and clinically distinct [2, 13, 14, 55].
In adults, IDHw gliomas may contain several pediatric-type tumors [31]; these include biologically more favorable glial and glioneuronal tumors with BRAF alterations, those gliomas with altered MYB-/MYBL-1, and high-grade gliomas with H3F3A mutation or RTK2. The other IDHw hLGG, which mainly arise from the adult supratentorial brain, frequently have molecular features of glioblastomas; TERT promoter mutation (pTERTm), epidermal growth factor receptor amplification (EGFRamp), or chromosome 7 gain and 10 loss (CH7/10). New WHO guidelines have categorized such groups as glioblastoma, IDH wild type [11]. IDHw hLGGs with molecular features of glioblastomas are typically known as molecular glioblastoma (mGBM).
Recent studies have revealed that mGBM has specific clinical features; tumors often have diffuse infiltration patterns (gliomatosis), multicentric loci or gyriform spread [39, 57]. Consequently, patients are likely to undergo biopsy rather than resection. Furthermore, they were less likely to receive adjuvant radiochemotherapy. Thus, while most studies have suggested the prognostic correspondence between histological glioblastoma (hGBM) and mGBM, the equivalence has not been sufficiently demonstrated. Reports have indicated that mGBM may reflect under-sampling phenomena or early-stage hGBM [49, 65]. For the former, treatment outcomes are worse if adjuvant radiochemotherapy is withheld, while for the latter, the outcomes depend on lead-time to hGBM development.
Molecular marker frequency in IDHw hLGGs differ extensively between studies. pTERTm rates vary from 15 to > 70% in IDHw hLGGs [33, 57], while EGFRamp rates vary from 10 to > 50% [19, 34]. It is unclear if such differences are caused by methodological or other factors, however, differences may affect tumor classification and prognostication.
As IDHw hLGGs are relatively rare, large-scale studies are scarce. We systematically reviewed IDHw hLGG studies and performed a meta-analysis to explore the variability underlying molecular marker frequency and their prognostic implications for mGBM.
Methods
We conducted this study according to Preferred Reporting Items for Systematic Reviews and Meta-Analysis (PRISMA) guidelines. It was not registered on The International Prospective Register of Systematic Reviews.
Literature search and data extraction
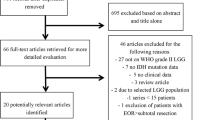
A PRISMA flow diagram is outlined (Fig. 1). From 2009 onward, we searched for relevant English studies using the keywords “glioma,” “astrocytoma,” AND “IDH” OR “isocitrate dehydrogenase” AND “TERT promoter” OR “EGFR” OR “chromosome 7” OR “chromosome 10” in PubMed, Ovid MEDLINR, Cochrane library and Scopus. Literature searches and data extraction were independently conducted by two study authors. EndoNote® (version 20) was used for data processing. Adult hemispheric IDHw LGG studies including ≥ 20 cases were incorporated if molecular testing in at least one of the followings was included; EGFRamp, pTERTm or CH7/10. Exclusion criteria: (1) molecular data could not be extracted, (2) studies with IDH1 data alone, (3) public database studies (e.g. The Cancer Genome Atlas, etc.), or (4) familial or pediatric cancers. For studies from the same institute, newer or relevant studies were selected. Finally, studies were selected by authors through discussion and consensus.
We collected patient age (mean or median) and sex information, molecular data, treatment-related factors, including tumor removal degree and adjuvant therapy, and overall survival (OS). Molecular diagnosis methods and tissue types (formalin-fixed paraffin-embedded (FFPE) or fresh-frozen (FF)) were also recorded. Studies were discriminated into Asian or non-Asian regions. When only median values and ranges were available, the mean and standard deviation were calculated using the method of Hozo et al. [26]. Individual patient data (IPD) were recorded if available. We re-calculated relevant statistical data from IPD, whereas tumors with H3F3A or BRAF mutations were excluded where possible. We used hazard ratio (HR) to determine treatment effects. When HRs were unavailable, we calculated values according to the method by Tierney et al. [59] or from Kaplan–Meier curve data. We used WebPlotDigitizer (https://automeris.io/WebPlotDigitizer/) to extract survival data from Kaplan–Meier curves [43, 44].
Statistical analysis
We used the R software program (v4.03) (https://www.r-project.org/) to perform statistical analyses (Meta, Metafor and EZR packages). An inverse-variance approach, with a random-effects method, was applied. Studies were assessed for heterogeneity (I2 statistic). A single-arm meta-analysis was performed to assess specific mutation rates using the metaprop function with a generalized linear mixed model. Meta-regression analyses were performed to identify heterogeneity related factors. Analyses were performed using the rma.uni function in Meta software package. In Kaplan–Meier curves from reconstructed data, log-rank and Cox-proportional hazard tests were used for survival analyses. Covariates of age, sex, type of surgery, adjuvant-treatment type, Karnofsky performance status score (KPS), and promoter methylation status of O6-methylguanine-DNA methyltransferase (MGMT), in addition to molecular status, were included in multivariate analyses when applicable. Two-sided P < 0.05 values were statistically significant.
Risk of bias
We used the Joanna Briggs Institute’s (JBI) critical appraisal checklist for case series (Supplementary Table 1) (https://jbi.global/critical-appraisal-tools) to evaluate risk of bias. Publication bias was evaluated using a funnel plot including > 10 studies. A linear regression analysis was also performed.
Ethics approval and informed consent
This review did not involve direct human investigations; therefore, no informed consent was required.
Results
We performed our literature search on September 4th, 2022 and retrieved 212 studies out of 688 records for full-text assessment (Fig. 1). We excluded 163 studies (lack of relevant data = 64, < 20 case = 29, public database only = 27, review articles = 9, hGBM only = 8, overlapping data = 19, and other reasons = 7). Finally, 49 studies were selected: 6 studies for comparing mGBM with hGBM [3, 6, 15, 37, 50, 58], 33 studies for calculating molecular marker frequency [2, 4, 7,8,9,10, 12, 16,17,18,19,20, 23, 24, 29, 30, 33,34,35,36, 38,39,40, 45, 48, 51,52,53,54, 56, 60, 61, 63, 66], and 8 studies for both [25, 28, 32, 42, 46, 47, 57, 64]. Two studies were utilized for analyses of prognostic factor in mGBM [21, 62]. IPD were collected from 12 studies [2, 4, 6, 15, 19, 21, 23, 25, 51, 61, 62, 64].
Risk of bias
Almost all studies were retrospective case series’ with low evidence levels. The JBI checklist for case series showed that most of them had one or two deficits (Supplementary Table 1).
mGBM marker frequency in IDHw hLGGs
In our analyses, 33 studies reported pTERTm rates in IDHw hLGG (Supplementary Table 2) [2,3,4, 8,9,10, 12, 18, 19, 24, 25, 28,29,30, 32, 33, 36, 38,39,40, 42, 45,46,47,48, 51, 52, 56, 57, 61, 63, 64, 66]. The pooled pTERTm proportion was 47.5% ([95% confidential interval (CI) 39.8–55.3]) (N = 2044, I2 = 86.2%) (Table 1) (Fig. 2A). A funnel plot showed no asymmetry (P = 0.34) (Fig. 2B). In subgroup analyses, the pooled proportion was significantly lower in studies from Asia when compared with non-Asian regions (36.2% [CI: 29.8–43.1] vs. 56.2% [CI: 45.2–66.5], P = 0.002), and significantly higher in studies using next generation sequencing (NGS) when compared with other methods (Sanger sequencing, Pyrosequencing or others) (68% [CI: 48.5–82.8] vs. 42.3% [CI: 35.5–49.3], P = 0.015), and significantly higher in studies using FFPE when compared with other methods (FF or either FFPE or FF) (30 studies; N = 1904: 56.2% [CI: 45.1–66.7] vs. 38.7% [CI: 28.5–50.0], P = 0.03). Meta-regression analyses also showed that the pooled pTERTm proportion increased by mean patient age (P = 0.02) and was possibly affected by grade III tumor rates (G3rate) (P = 0.068) (Table 1). Multiple meta-regression analyses showed that only regions (Asia or non-Asia) reached significance (P = 0.03) but FFPE exhibited borderline significance (P = 0.08). However, multiple meta-regression analyses still showed high residual heterogeneity (I2 = 84.8%). We hypothesized that undetermined factors may have affected our results. Such factors may have affected pTERTm rate in 1p/19q co-deleted gliomas (Codel). Indeed, we identified a significant correlation between pTERTm rates in IDHw LGG and Codel tumors (R = 0.49, P = 0.012) (Supplementary Fig.S1A). Moreover, a quantile–quantile (Q-Q) plot showed that distribution of pTERTm rates in Codel was normal when we excluded cases with rate < 90% (Supplementary Fig.S1B). For this reason, we selected 19 articles from institutes which reported ≥ 90% pTERTm rates in Codel for multiple meta-regressions. Then, the geographical region (P < 0.001) and method (NGS vs others, P = 0.002) were significant for the pTERTm proportion, and heterogeneity disappeared (I2 = 0%). The pooled pTERTm rate in IDHw hLGG was 41.1% [CI: 33.4–49.4] (7 studies, N = 335) in Asia but 64.1% [CI: 54.1–73.0] (12 studies, N = 588) in non-Asia regions (P < 0.001).
A Forest plot of proportion for TERT promoter mutation (pTERTm) in IDH wild-type lower-grade gliomas. Subgroup analyses showing a higher frequency in Asian regions (P = 0.0024). B Funnel plot showing no asymmetry. C Forest plot showing molecular glioblastomas (mGBM) proportions for pTERTm rates. Subgroup analysis presenting a higher frequency in Asian regions (P = 0.002). D Kaplan–Meier curves from individual patient data showing overall survival in patients with mGBM and IDH wild-type lower-grade gliomas “not elsewhere classified” (NEC), in Asian and non-Asian regions
EGFRamp rates in IDHw hLGGs (N = 2225) (Table 1) were reported by 27 studies [2,3,4, 7, 9, 16, 17, 19, 20, 25, 28, 32, 34, 36, 38, 39, 42, 45, 47, 51,52,53,54, 56, 57, 60, 61]. The pooled EGFRamp proportion was 23.6% [CI: 18.6–29.4] (I2 = 83.1%). We did not identify publication bias (P = 0.28). It was low in Asia studies when compared with other regions (15.9% [CI: 10.6–23.2] vs. 28.8% [CI: 22.9–35.5], P = 0.008). Methodologies (NGS vs others (FISH, MLPA, array CGH etc.)) did not affect result (P = 0.43), but tissue type did and was significant (FFPE 29.6% [CI: 22.2–38.3] vs FF and FF or FFPE 17.1% [CI: 10.3–27.1], P = 0.047). Meta-regression analyses showed that G3rates were significant factors (P = 0.035), but mean age was not (P = 0.23). Multiple meta-regression analyses also revealed that geographical region (P = 0.01), tissue type (P = 0.028) and G3rates (P = 0.037) were significant factors impacting EGFRamp rates (I2 = 67.6%).
CH7/10 status was evaluated in 13 studies (N = 1194) (Table 1) [4, 9, 19, 23, 32, 35, 36, 38, 51, 54, 56, 57, 61], although definition were different (e.g. whole CH7/10, 7p( +)/10q(−), 7( +)/10q(−), or EGFR gain/PTEN loss) (Supplementary Table 1). The pooled proportion in IDHw hLGG was 33.2% [CI: 21.5–47.4] (I2 = 89.8%). A Funnel plot showed no publication bias (P = 0.20). The rate was also lower in Asian regions (15.0% [CI: 6.7–30.4]) when compared with non-Asian regions (50.9% [CI 42.9–58.9]) (P = 0.0003). Methodologies (NGS vs. others) did not affect rates (P = 0.07), but tissue type did (FFPE 53.9% [CI: 44.6–63.0] vs. FF and FFPE or FF 24.7% [CI: 12.4–43.4] (P = 0.007). Meta-regression analyses showed that mean age (P = 0.03) and G3rate (P = 0.046) were significant factors. Multiple meta-regression analyses also revealed that geographical region (P < 0.0001), tissue type (P < 0.0001) and G3rate (P = 0.0028) significantly affected the CH7/10 rates. When evaluation was limited to studies using whole CH7/10 [9, 23, 32, 35, 36, 54, 56, 57], pooled proportion in IDHw hLGG was 34.8% [CI: 20.0–53.2], which is comparable to the abovementioned value. In this case, subgroup and meta-regression analyses did not reach significance except in terms of mean age (P < 0.001); however, they showed a tendency similar to the described analyses.
The prognostic implications of difference in mGBM marker rates
We calculated the pooled mGBM rate in IDHw hLGG studies examining pTERTm. The pooled mGBM rate in Asian regions (38.9% [CI: 33.4–44.8], 14 studies, N = 868) was lower than that in non-Asia regions (61.9% [CI: 49.0–73.2], 19 studies, N = 1215) (P = 0.0015) (Fig. 2C). Interestingly, the rate increased in studies examining multiple molecular markers (three markers 79.0% [CI: 68.9–86.5], two markers = 78.0% [CI: 50.3–92.5] vs. one marker (pTERTm) = 45.1% [CI: 34.2–56.4]) in non-Asian regions (P < 0.0001), while rates did not change in Asian regions (three markers = 39.6% [CI: 28.2–52.2], two markers = 32.0% [CI: 25.4–39.4], and one marker (pTERTm) = 41.1% [CI: 34.3–48.3]) (P = 0.38). This was true when we analyzed studies showing > 90% pTERTm rates in Codel tumors (Asian regions: P = 0.94 and non-Asian regions: P < 0.0001). Therefore, pTERT wild-type hLGGs in Asian regions rarely expressed other GBM molecular markers (4.1% when analyzed in IPD), whereas those in non-Asian did.
As molecular marker analyses showed, studies using FFPE showed a significantly higher mGBM rates (62.5% [CI: 49.5–74.0]) when compared with those using other tissue (FF or either FF or FFPE) (41.4% [CI: 31.4–52.1]) (P = 0.014).
We compared OS rates of IDHw hLGG tumors between Asian and non-Asian studies using IPD. The median OS rate in Asian mGBM tumors (21.1 months [CI: 16.1–24.7], N = 185) was similar to in non-Asian tumors (20.8 months [CI: 17.9–22.7], N = 275) (P = 0.45) (Fig. 2D). The median OS rate of IDHw hLGG tumors, “not elsewhere classified” (NEC) in Asian regions (38.1 months [30.4–46.8], N = 304), was also similar to non-Asian regions (35.2 months [CI: 26.4–53.3], N = 123) (P = 0.6).
Differences between mGBM and hGBM
mGBM and hGBM survival data were provided by 14 studies (Table 2) [3, 6, 15, 25, 28, 32, 37, 42, 46, 47, 50, 57, 58, 64]. Five showed that biopsied tumors were more frequent for mGBM when compared with hGBM [6, 37, 50, 57, 58] (P < 0.001 in each study, Fisher’s exact tests), while two studies reported comparative total removal rates between the two [15, 32]. Eight studies described adjuvant radiochemotherapy frequency rates [6, 15, 32, 37, 42, 50, 57, 58]. Four studies reported that the patients with mGBM underwent radiotherapy less often when compared with patients with hGBM. Three studies reported that the patients with mGBM underwent chemotherapy less often when compared with patients with hGBM. Treatments frequency differences in other studies failed to reach significance.
While most studies showed that mGBM patients had comparable OS rates when compared with hGBM patients, the pooled HR value of mGBM against hGBM was 0.82 in univariate analyses ([CI: 0.69–0.98], I2 = 46%) (P = 0.028) (Fig. 3A). No publication bias was detected (P = 0.92) (Fig. 3B). Moderate heterogeneity improved in the pooled HR in multivariate analyses (including covariates of age (five studies), sex (four studies), surgery (four studies), radiotherapy and/or chemotherapy (three studies), KPS (three studies), MGMT (two studies) and CDKN2A/B (one study)), which showed a significantly better OS rate in mGBM (HR 0.61 [CI: 0.50–0.74], I2 = 0%) (P < 0.0001) (Fig. 3C). We identified no differences in pooled HR values between Asian and non-Asian regions (P = 0.80). Kaplan–Meier curves from IPD showed an identical OS rate between hGBM and mGBM with grade III histology (P = 0.39), but better OS for mGBM with grade II histology (P = 0.014) (Fig. 3D).
A Forest plot showing hazard ratio (HR) values from overall survival analyses (univariate) between patients with molecular glioblastoma (mGBM) and histological glioblastoma (hGBM). B Funnel plot showing no asymmetry. C Forest plot (from multivariate analyses) showing significantly lower HR values in mGBM against hGBM (P < 0.0001), with low heterogeneity. *data from individual patient data (IPD):**extracted data from Kaplan–Meier curves: †data from https://www.surgery.cuhk.edu.hk/BTC/HSBC/molecular_grading_paper_data.pdf. D Kaplan- Meier curves of IPD showing better OS rates for mGBM with grade II histology when compared with mGBM with grade III histology or hGBM
Prognostic factors in mGBM
We meta-analyzed prognostic factors related to OS rates in mGBM (16 studies) (Table 3) [3, 4, 9, 15, 19, 21, 25, 32, 38, 46, 47, 50, 51, 61, 62, 64]. A younger age (HR1.028 [CI: 1.011–1.045], P = 0.001) and extent of surgery (HR 0.643 [CI: 0.446–0.926], P = 0.018) were identified as significantly better prognostic factors. Although no molecular markers reached significance, patients with histologically grade III gliomas had worse OS rates when compared with grade II tumors in pooled results from both univariate (HR 1.58 [CI: 1.02–2.429], P = 0.003) and multivariate analyses (HR 1.314 [CI: 1.041–1.660], P = 0.022) (Supplementary Fig. S2A, B).
Discussion
mGBM has significantly better OS rates when compared with hGBM. Additionally, mGBM with grade II histological features had better OS rates when compared with that with grade III histological features. Although mGBM rates in IDHw hLGG varied extensively among studies, differences were partly explained by geographical regions. Rates in Asia were significantly lower when compared with non-Asian regions. Interestingly, pTERT wild-type tumors in Asia rarely expressed other molecular markers, however, despite these differences, mGBM in Asian and non-Asian regions showed identical OS rates using IPD.
Diagnosis and frequency of mGBM
We identified one reason which possibly explained mGBM rate differences across studies, i.e. the geographical region. While low pTERTm frequency rates in glioblastomas in Asia were previously reported [5], this was also true for IDHw LGGs. Not only pTERTm, but also EGFRamp and CH7/10 also occurred less frequently in Asian regions. As lower-grade glioma incidences in Asia were lower when compared with eastern countries [41], mGBM incidence was assumed to be lower. We observed differences in molecular markers combination patterns between Asian and non-Asian regions. Fujimoto et al. [19] reported that pTERT mutation status was necessary and sufficient to diagnose IDHw hLGG with molecular features of glioblastoma. This appeared to be true for Asian but not in non-Asian regions. However, if only the test for pTERTm is used, approximately 4.1% of mGBM may be missed from the analyses even in Asia.
One substantial molecular diagnostic issue in this field is that TERT promotor areas have high guanine-cytosine content and easily form secondary structures which lead to a poor amplification [27]. Thus, a diagnosis may be misleading when tumor cell density is low. While it is generally accepted that FF tissues are better materials, our meta-regression analysis showed that pTERTm rates were higher in FFPE samples. EGFRamp and CH7/10 detection rates had similar frequencies. This was most likely due to sampling selection spots for molecular diagnostics. IDHw hLGGs sometimes grow invasively and adopt gliomatosis cerebri forms. In such cases, tumor cells may be sparse in tumor areas by preoperative imaging. A more appropriate area for molecular diagnoses may be selected in FFPE rather than FF samples. While different methodologies have different molecular diagnostic sensitivities [1, 22], specimen condition, especially tumor cell density, appears to be more important. To exclude the problems in material and methodological issues, we re-analyzed pTERTm data in studies that showed pTERTm > 90% of Codel tumors. This generated a small change in the pTERTm rates in mGBM but our overall conclusions were unaltered.
Prognostic factors in mGBM
Although mGBM in most studies exhibited similar OS rates to hGBM, pooled HR analyses showed better OS rates in mGBM when compared with hGBM. Furthermore, pooled HR values from multivariate analyses, including a covariate analysis of age, sex and surgery, showed a much better OS rates with lower heterogeneity (HR 0.61 [CI: 0.50–0.74], I2 = 0%). Berzero et al. [9] reported better OS rates in mGBM with grade II histology when compared with grade III, although their study lacked multivariate analyses. As mGBMs with grade III features showed worse OS rates when compared with grade II (HR 1.582 [CI: 1.030–2.429], P = 0.003), differences in OS rates between mGBM and hGBM appeared to be due to the grade II subgroup in mGBM, as indicated by Kaplan–Meier curves (Fig. 3D).
However, even mGBM in grade II histology sometimes shows rapid growth with new ring-like contrast enhancement by magnetic resonance imaging [28, 39], and hGBM transformation at recurrence [49, 52]. Thus far, specific molecular abnormalities showed no significant difference in OS rates [21, 50]. We also showed that no molecular markers significantly affected prognoses other than histological grade. The prognostic significance of the extent of mGBM surgery remains controversial. Ruda et al. [52] identified no survival benefit from gross total resection of mGBM with grade II histology (they did not include stereotactic biopsy cases). Ramos-Fresnedo et al. [50] did not demonstrate OS differences between biopsy and GTR (P = 0.079). Nevertheless, we showed that pooled HR favored more extensive surgery (HR 0.644 [CI: 0.420–0.990], P = 0.045). Zhang et al. [65] reported that the prospective identification of mGBM resulted in more aggressive patient management and improved clinical outcomes when compared with a biologically matched historical control patient cohort receiving standard-of-care therapy based on histomorphologic diagnoses alone. However, the effects of early aggressive treatment for mGBM must be examined in future studies.
Study limitations
Our study had several limitations. Firstly, most studies were retrospective and had low evidence levels. Consequently, heterogeneity was high in pooled data and multiple meta-regression analyses were required to identify reasons for this. Secondly, while we largely excluded tumors with BRAF or H3F3A mutation, some studies presented no information. Most of the studies had no data regarding other pediatric-type gene changes, such as altered MYB/MYB-L1. Therefore, NEC numbers in IDHw hLGG may have been overestimated in some studies. Thirdly, in some cases, we extracted survival data from Kaplan–Meier curves in figures of articles. Although we confirmed overlapping of original and extracted curves, measurement errors in graph line thickness may have inadvertently occurred. Fourthly, some multi-institutional studies may have had overlapping data with other studies. While we carefully selected studies, some duplicates data may have been included. Potential data overlaps are recorded in supplementary Table 1.
Conclusion
Patients with mGBM have better OS rates when compared with patients with hGBM, especially when grade II histology is indicated. We could not identify other molecular markers that differentiated mGBM prognosis, although patient age and surgical extent were prognostic factors. We identified differences between Asian and non-Asian regions in terms of molecular marker frequency pattern for mGBM in IDHw hLGG. However, OS rates in patients with mGBM were concordant between Asian and non-Asian cohorts. Although a considerable number of “NEC” in IDHw hLGGs were identified especially in Asian patients, further studies are warranted for their classification. Controversies continue to persist in the new WHO classification of IDHw hLGG. DNA methylation profiling is an evolving method that will further facilitate classification [31]. Prospective clinical studies in IDHw hLGG with DNA methylation profiling are necessary in the future.
References
Adachi JI, Shirahata M, Suzuki T et al (2021) Droplet digital PCR assay for detecting TERT promoter mutations in patients with glioma. Brain Tumor Pathol 38:201–209
Aibaidula A, Chan AK, Shi Z et al (2017) Adult IDH wild-type lower-grade gliomas should be further stratified. Neuro Oncol 19:1327–1337
Akyerli CB, Yuksel S, Can O et al (2018) Use of telomerase promoter mutations to mark specific molecular subsets with reciprocal clinical behavior in IDH mutant and IDH wild-type diffuse gliomas. J Neurosurg 128:1102–1114
Aoki K, Nakamura H, Suzuki H et al (2018) Prognostic relevance of genetic alterations in diffuse lower-grade gliomas. Neuro Oncol 20:66–77
Arita H, Ichimura K (2022) Prognostic significance of TERT promoter mutations in adult-type diffuse gliomas. Brain Tumor Pathol 39:121–129
Arita H, Yamasaki K, Matsushita Y et al (2016) A combination of TERT promoter mutation and MGMT methylation status predicts clinically relevant subgroups of newly diagnosed glioblastomas. Acta Neuropathol Commun 4:79. https://doi.org/10.1186/s40478-016-0351-2
Bale TA, Jordan JT, Rapalino O et al (2019) Financially effective test algorithm to identify an aggressive, EGFR-amplified variant of IDH-wildtype, lower-grade diffuse glioma. Neuro Oncol 21:596–605
Bell EH, Zhang P, Shaw EG et al (2020) Comprehensive genomic analysis in NRG oncology/RTOG 9802: A Phase III trial of radiation versus radiation plus procarbazine, lomustine (CCNU), and vincristine in high-risk low-grade glioma. J Clin Oncol 38:3407–3417
Berzero G, Di Stefano AL, Ronchi S et al (2021) IDH-wildtype lower-grade diffuse gliomas: the importance of histological grade and molecular assessment for prognostic stratification. Neuro Oncol 23:955–966
Bieńkowski M, Wöhrer A, Moser P et al (2018) Molecular diagnostic testing of diffuse gliomas in the real-life setting: a practical approach. Clin Neuropathol 37:166–177
Brat DJ Ed, Figarella-Branger D, et al. (2021) Gliomas, glioneuronal tumours, and neuronal tumours. In: Board Wcote (editor) Central nervous system tumours 5th edn. International Agency for Research on Cancer pp 15–187
Brito C, Azevedo A, Esteves S et al (2019) Clinical insights gained by refining the 2016 WHO classification of diffuse gliomas with: EGFR amplification, TERT mutations, PTEN deletion and MGMT methylation. BMC Cancer 19:968. https://doi.org/10.1186/s12885-019-6177-0
Cancer Genome Atlas Research Network, Brat DJ, Verhaak RG, Aldape KD et al (2015) Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med 372:2481–2498
Ceccarelli M, Barthel FP, Malta TM et al (2016) Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164:550–563
Chan AK, Shi ZF, Li KK et al (2022) Combinations of single-gene biomarkers can precisely stratify 1028 adult gliomas for prognostication. Front Oncol 12:839302. https://doi.org/10.3389/fonc.2022.839302
Di Stefano AL, Enciso-Mora V, Marie Y et al (2013) Association between glioma susceptibility loci and tumour pathology defines specific molecular etiologies. Neuro Oncol 15:542–547
Eckel-Passow JE, Lachance DH, Molinaro AM et al (2015) Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. N Engl J Med 372:2499–2508
Fleming JL, Pugh SL, Fisher BJ et al (2021) Long-term report of a comprehensive molecular and genomic analysis in NRG oncology/RTOG 0424: a phase II study of radiation and temozolomide in high-risk grade II glioma. JCO Precis Oncol. https://doi.org/10.1200/po.21.00112
Fujimoto K, Arita H, Satomi K et al (2021) TERT promoter mutation status is necessary and sufficient to diagnose IDH-wildtype diffuse astrocytic glioma with molecular features of glioblastoma. Acta Neuropathol 142:323–338
Gorovets D, Kannan K, Shen R et al (2012) IDH mutation and neuroglial developmental features define clinically distinct subclasses of lower grade diffuse astrocytic glioma. Clin Cancer Res 18:2490–2501
Grogan D, Bray DP, Cosgrove M et al (2022) Clinical and radiographic characteristics of diffuse astrocytic glioma, IDH-wildtype, with molecular features of glioblastoma: a single institution review. J Neurooncol 157:187–195
Hasanau T, Pisarev E, Kisil O et al (2022) Detection of TERT promoter mutations as a prognostic biomarker in gliomas: methodology, prospects, and advances. Biomedicines. https://doi.org/10.3390/biomedicines10030728
Hattori N, Hirose Y, Sasaki H et al (2016) World health organization grade II-III astrocytomas consist of genetically distinct tumor lineages. Cancer Sci 107:1159–1164
Heidenreich B, Rachakonda PS, Hosen I et al (2015) TERT promoter mutations and telomere length in adult malignant gliomas and recurrences. Oncotarget 6:10617–10633
Higa N, Akahane T, Yokoyama S et al (2022) Molecular genetic profile of 300 Japanese patients with diffuse gliomas using a glioma-tailored gene panel. Neurol Med Chir (Tokyo) 62:391–399. https://doi.org/10.2176/jns-nmc.2022-0103
Hozo SP, Djulbegovic B, Hozo I (2005) Estimating the mean and variance from the median, range, and the size of a sample. BMC Med Res Methodol 5:13. https://doi.org/10.1186/1471-2288-5-13
Ichimura K (2019) TERT promoter mutation as a diagnostic marker for diffuse gliomas. Neuro Oncol 21:417–418
Izquierdo C, Barritault M, Poncet D et al (2019) Radiological characteristics and natural history of adult IDH-wildtype astrocytomas with TERT promoter mutations. Neurosurgery 85:E448–E456. https://doi.org/10.1093/neuros/nyy513
Jiang C, Kong Z, Zhang Y et al (2020) Conventional magnetic resonance imaging-based radiomic signature predicts telomerase reverse transcriptase promoter mutation status in grade II and III gliomas. Neuroradiology 62:803–813
Killela PJ, Pirozzi PC, Healy P et al (2014) Mutations in IDH1, IDH2, and in the TERT promoter define clinically distinct subgroups of adult malignant gliomas. Oncotarget 5:1515–1525
Komori T (2023) Update of the 2021 WHO classification of tumors of the central nervous system: adult diffuse gliomas. Brain Tumor Pathol 40:1–3. https://doi.org/10.1007/s10014-022-00446-1
Kumari K, Dandapath I, Singh J et al (2022) Molecular characterization of IDH wild-type diffuse astrocytomas: the potential of cIMPACT-NOW guidelines. App Immunohistochem Mol Morphol 30:410–417
Kuwahara K, Ohba S, Nakae S et al (2019) Clinical, histopathological, and molecular analyses of IDH-wild-type WHO grade II-III gliomas to establish genetic predictors of poor prognosis. Brain Tumor Pathol 36:135–143
Lasica AB, Jaunmuktane Z, Fersht N et al (2021) Genomic prognosticators and extent of resection in molecularly subtyped World Health Organization grade II and III gliomas-A single-institution, nine-year data. World Neurosurg 151:e217–e233
Li J, Xue Y, Wenger A et al (2019) Individual assignment of adult diffuse gliomas into the EM/PM molecular subtypes using a TaqMan low-density array. Clin Cancer Res 25:7068–7077
Lin AL, Rosenblum M, Mellinghoff IK et al (2020) Prognostic and radiographic correlates of a prospectively collected molecularly profiled cohort of IDH1/2-wildtype astrocytomas. Brain Pathol 30:653–660
Ma S, Rudra S, Campian JL et al (2020) Prognostic impact of CDKN2A/B deletion, TERT mutation, and EGFR amplification on histological and molecular IDH-wildtype glioblastoma. Neurooncol Adv 2:vdaa26. https://doi.org/10.1093/noajnl/vdaa126
Makino Y, Arakawa Y, Yoshioka E et al (2021) Prognostic stratification for IDH-wild-type lower-grade astrocytoma by sanger sequencing and copy-number alteration analysis with MLPA. Sci Rep 11:14408
Mesny E, Barritault M, Izquierdo C et al (2022) Gyriform infiltration as imaging biomarker for molecular glioblastomas. J Neurooncol 157:511–521
Mizoguchi M, Hata N, Kuga D et al (2021) Clinical implications of molecular analysis in diffuse glioma stratification. Brain Tumor Pathol 38:210–217
Mo Z, Xin J, Chai R et al (2022) Epidemiological characteristics and genetic alterations in adult diffuse glioma in East Asian populations. Cancer Biol Med 19:1440–1459
Mortensen D, Ulhoi BP, Lukacova S et al (2022) Impact of new molecular criteria on diagnosis and survival of adult glioma patients. IBRO Neurosci Rep 13:299–305
Nakasu S, Nakasu Y (2022) Malignant progression of diffuse low-grade gliomas: a systematic review and meta-analysis on incidence and related factors. Neurol Med Chir (Tokyo) 62:177–185
Nakasu S, Nakasu Y, Tsuji A et al (2023) Incidental diffuse low-grade gliomas: a systematic review and meta-analysis of treatment results with correction of lead-time and length-time biases. Neuro-Oncol Pract 10:113–125. https://doi.org/10.1093/nop/npac073
Park YW, Kim S, Park CJ et al (2022) Adding radiomics to the 2021 WHO updates may improve prognostic prediction for current IDH-wildtype histological lower-grade gliomas with known EGFR amplification and TERT promoter mutation status. Eur Radiol 32:8089–8098. https://doi.org/10.1007/s00330-022-08941-x
Pekmezci M, Rice T, Molinaro AM et al (2017) Adult infiltrating gliomas with WHO 2016 integrated diagnosis: additional prognostic roles of ATRX and TERT. Acta Neuropathol 133:1001–1016
Petersen JK, Boldt HB, Sorensen MD et al (2021) Targeted next-generation sequencing of adult gliomas for retrospective prognostic evaluation and up-front diagnostics. Neuropathol Appl Neurobiol 47:108–126
Poulen G, Gozé C, Rigau V, Duffau H (2018) Huge heterogeneity in survival in a subset of adult patients with resected, wild-type isocitrate dehydrogenase status, WHO grade II astrocytomas. J Neurosurg 130:1289–1298
Ramos-Fresnedo A, Domingo RA, Perez-Vega C et al (2022) The early infiltrative phase of GBM hypothesis: are molecular glioblastomas histological glioblastomas in the making? A preliminary multicenter study. J Neurooncol 158:497–506
Ramos-Fresnedo A, Pullen MW, Perez-Vega C et al (2022) The survival outcomes of molecular glioblastoma IDH-wildtype: a multicenter study. J Neurooncol 157:177–185
Reuss DE, Kratz A, Sahm F et al (2015) Adult IDH wild type astrocytomas biologically and clinically resolve into other tumor entities. Acta Neuropathol 130:407–417
Ruda R, Bruno F, Ius T et al (2022) IDH wild-type grade 2 diffuse astrocytomas: prognostic factors and impact of treatments within molecular subgroups. Neuro Oncol 24:809–820
Shibahara I, Sonoda Y, Kanamori M et al (2012) IDH1/2 gene status defines the prognosis and molecular profiles in patients with grade III gliomas. Int J Clin Oncol 17:551–561
Stichel D, Ebrahimi A, Reuss D et al (2018) Distribution of EGFR amplification, combined chromosome 7 gain and chromosome 10 loss, and TERT promoter mutation in brain tumors and their potential for the reclassification of IDHwt astrocytoma to glioblastoma. Acta Neuropathol 136:793–803
Suzuki H, Aoki K, Chiba K et al (2015) Mutational landscape and clonal architecture in grade II and III gliomas. Nat Genet 47:458–468
Tabouret E, Nguyen AT, Dehais C et al (2016) Prognostic impact of the 2016 WHO classification of diffuse gliomas in the French POLA cohort. Acta Neuropathol 132:625–634
Tesileanu CMS, Dirven L, Wijnenga MMJ et al (2020) Survival of diffuse astrocytic glioma, IDH1/2 wildtype, with molecular features of glioblastoma, WHO grade IV: a confirmation of the cIMPACT-NOW criteria. Neuro Oncol 22:515–523
Teske N, Karschnia P, Weller J et al (2022) Extent, pattern, and prognostic value of MGMT promotor methylation: does it differ between glioblastoma and IDH-wildtype/TERT-mutated astrocytoma? J Neurooncol 156:317–327
Tierney JF, Stewart LA, Ghersi D et al (2007) Practical methods for incorporating summary time-to-event data into meta-analysis. Trials 8:16. https://doi.org/10.1186/1745-6215-8-16
Wang P, Luo C, Hong PJ et al (2021) The role of surgery in IDH-wild-type lower-grade gliomas: threshold at a high extent of resection should be pursued. Neurosurgery 88:1136–1144
Weller M, Weber RG, Willscher E et al (2015) Molecular classification of diffuse cerebral WHO grade II/III gliomas using genome- and transcriptome-wide profiling improves stratification of prognostically distinct patient groups. Acta Neuropathol 129:679–693
Wijnenga MMJ, Dubbink HJ, French PJ et al (2017) Molecular and clinical heterogeneity of adult diffuse low-grade IDH wild-type gliomas: assessment of TERT promoter mutation and chromosome 7 and 10 copy number status allows superior prognostic stratification. Acta Neuropathol 134:957–959
Yang P, Cai J, Yan W et al (2016) Classification based on mutations of TERT promoter and IDH characterizes subtypes in grade II/III gliomas. Neuro Oncol 18:1099–1108
Yang Z, Ling F, Ruan S et al (2021) Clinical and Prognostic Implications of 1p/19q, IDH, BRAF, MGMT Promoter, and TERT Promoter Alterations, and Expression of Ki-67 and p53 in Human Gliomas. Cancer Manag Res 13:8755–8765
Zhang Y, Lucas CG, Young JS et al (2022) Prospective genomically guided identification of “early/evolving” and “undersampled” IDH-wildtype glioblastoma leads to improved clinical outcomes. Neuro Oncol 24:1749–1762
Zhang ZY, Zhan YB, Zhang FJ et al (2019) Prognostic value of preoperative hematological markers combined with molecular pathology in patients with diffuse gliomas. Aging (Albany NY) 11:6252–6272
Funding
No specific funding was received for this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare none.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nakasu, S., Deguchi, S. & Nakasu, Y. IDH wild-type lower-grade gliomas with glioblastoma molecular features: a systematic review and meta-analysis. Brain Tumor Pathol 40, 143–157 (2023). https://doi.org/10.1007/s10014-023-00463-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-023-00463-8