Abstract
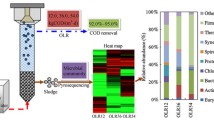
Upflow Anaerobic Sludge Blanket (UASB) reactors are alternatives in the anaerobic treatment of sanitary sewage in different parts of the world; however, in temperate environments, they are subject to strong seasonal influence. Understanding the dynamics of the microbial community in these systems is essential to propose operational alternatives, improve projects and increase the quality of treated effluents. In this study, for one year, high-performance sequencing, associated with bioinformatics tools for taxonomic annotation and functional prediction was used to characterize the microbial community present in the sludge of biodigesters on full-scale, treating domestic sewage at ambient temperature. Among the most representative phyla stood out Desulfobacterota (20.21–28.64%), Proteobacteria (7.48–24.90%), Bacteroidota (10.05–18.37%), Caldisericota (9.49–17.20%), and Halobacterota (3.23–6.55%). By performing a Canonical Correspondence Analysis (CCA), Methanolinea was correlated to the efficiency in removing Chemical Oxygen Demand (COD), Bacteroidetes_VadinHA17 to the production of volatile fatty acids (VFAs), and CI75cm.2.12 at temperature. On the other hand, Desulfovibrio, Spirochaetaceae_uncultured, Methanosaeta, Lentimicrobiaceae_unclassified, and ADurb.Bin063-1 were relevant in shaping the microbial community in a co-occurrence network. Diversity analyses showed greater richness and evenness for the colder seasons, possibly, due to the lesser influence of dominant taxa. Among the principal metabolic functions associated with the community, the metabolism of proteins and amino acids stood out (7.74–8.00%), and the genes related to the synthesis of VFAs presented higher relative abundance for the autumn and winter. Despite the differences in diversity and taxonomic composition, no significant changes were observed in the efficiency of the biodigesters.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The rational use of water resources and the proper treatment of wastewater originating from human activities have been widely stimulated in recent years, due to the scarcity of water in several parts of the world [1]. However, despite efforts, large volumes of untreated sewage still end up reaching water bodies—recent estimates suggest that only 56% of domestic sewage in the world is properly treated—causing damage to the delicate balance of aquatic ecosystems and socioeconomic impacts on the affect population [2]. In this context, the purification of sewage in Wastewater Treatment Plants (WWTPs) is extremely important and several technologies are used to remove these waters from their polluting potential, including anaerobic digestion units [3].
The use of anaerobic digestion (AD) in the treatment of solid and liquid waste has been widely applied in several parts of the world for over a century [4, 5]. The word digestion is used in the context of stabilizing organic matter through the action of microorganisms in conditions that favor their growth and reproduction [6]. AD under ideal conditions is processed in 4 main steps: hydrolysis, acidogenesis, acetogenesis, and methanogenesis, where a wide diversity of species acts on the degradation of organic matter, generating methane gas (CH4) and carbon dioxide (CO2) as a final product [7]. These microbiological processes commonly occur in specialized units known as biodigesters or reactors. Among the range of biodigesters used, the UASB reactors stand out for their broad spectrum of use, showing a good capacity to remove Biochemical Oxygen Demand (BOD), COD, and Total Suspended Solids (TSS), presenting low sludge production and resistance to organic overload, requiring low operating and maintenance costs [8, 9]. Especially in tropical climates, they operate under ideal temperature conditions, requiring no additional heating spending, and are viable alternatives in developing countries [10, 11].
The UASB reactors are essentially characterized by the presence of a sludge blanket consisting of active biomass that forms granules, responsible for the degradation of the incoming organic matter, and a three-phase separator system, which enables solids retention, biogas collection, and consequent release of the liquid fraction of sewage, then properly treated [9]. UASB digesters enable the treatment of wastewater originating from several polluting sources, such as sugar, paper and cellulose, beverage, chemical, dairy, slaughterhouse, and sanitary sewage systems [12]. Sludge blanket reactors are widespread throughout the world. In Brazil, about 40% of WWTPs located only in the South, Southeast, and Midwest regions employ UASB reactor technology in the secondary treatment [10, 13, 14].
Used in the treatment of domestic sewage, UASB reactors are subject to a series of interferences that change the characteristics of the incoming wastewater, such as seasonal effects, socioeconomic level of the population served, infiltrations in the network, and type of sewage collection system (mixed or with separator) [15,16,17]. Such disturbances can mainly lead to changes in factors such as temperature, pH, flow, hydraulic retention time, and applied organic load, which can trigger imbalances in crucial stages of anaerobic digestion [18,19,20]. Under the influence of these variables, the microbiota present in these systems tends to suffer impacts, resulting in losses of efficiency and quality of the treated sewage [21].
Several studies have focused on the response of microbial communities to variations in the operating conditions of anaerobic reactors, notably using molecular biology and bioinformatics techniques [22,23,24,25]. However, these studies do not jointly address seasonal aspects, full-scale anaerobic reactors, systems with low influent organic load, and domestic sewage treatment. In this context, it is crucial to understand the dynamics of the microbiota subject to environmental contingencies in real-scale systems to provide subsidies for the elaboration of projects, monitoring, and more efficient operational control of these units in the process of removing associated pollutants.
From this perspective, the present study aimed to analyze and characterize the microbial community of UASB reactors for one year, using high-performance sequencing (NGS) and bioinformatics tools, to determine the taxonomic profile as well as to predict the main metabolic functions associated with the groups found, relating seasonal conditions of temperature, physical–chemical aspects and efficiency parameters evaluated in WWTPs.
Materials and methods
Study area
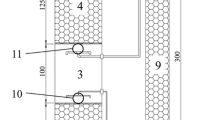
The collections of sludge, influent sewage, and effluent were carried out at three full-scale anaerobic reactors of the UASB-type (UASB-B, UASB-P, and UASB-T) while treating mixed sewage (rain and sewer) received by three different WWTPs (Fig. S1 and Table S1), located in the municipality of Caxias do Sul (51°10′06″ W; 29°10′05″ S), Brazil. The UASB-T reactor has a volume of 3990 m3, the others 2226 m3, and both were designed to operate continuously at ambient temperature and with a minimum hydraulic retention time (HRT) of 10 h. Considering that the reactors are inserted in the same context in the WWTPs where they are located and have similar characteristics regarding the sewage received, these were treated as replicates in the present study, noting that they were not subjected to significant changes in the operational process. The climate of the region is classified as Cfb (humid temperate) characterized by well-distributed rainfall and an average temperature of the hottest month below 22ºC, considering the Köppen (1931) classification [26, 27].
Physicochemical analysis
Physicochemical analyses were carried out to characterize the influent and effluent sewage as well as the sludge used in the specific methanogenic activity (SMA) test during the period from October 2020 to October 2021. Measurements of Flow (Q), pH, COD, Total Kjeldahl Nitrogen (TKN), Ammoniacal Nitrogen (NH3-N), and Total Phosphorus (PO4-P) were carried out, referring to sewage and evaluation of Total Solids (TS), Fixed Solids (FS) and Volatile Solids (VS) related to the sludge collected from the biodigesters. The aforementioned analyses were performed according to the methods described in Standard Methods [28]. VFAs analyses were determined using a gas chromatograph (GC-Shimadzu – QP2010 Ultra) equipped with a DN–FFAP column (30 mm × 0.32 mm × 0.25), Flame Ionization Detector (FID), with Helium (He) gas as carrier, as well as synthetic air and nitrogen (N2) as auxiliary gases [29]. The temperature assessment was performed daily and included measurements of the air, inlet, and outlet of the reactors. The SMA test was conducted according to the methodology proposed by Chernicharo [30]; however, the assembly of the apparatus to quantify the produced biogas followed the model suggested by Drosg et al. [31]. Data related to precipitation and other meteorological factors, referring to the study period, were acquired from the National Institute of Meteorology (INMET) [32].
Molecular analysis
Sampling, DNA extraction, and NGS sequencing
In all, 12 sludge collections were carried out for the months of January (summer), May (autumn), August (winter) and October (spring) of 2021 for analysis and characterization of the microbial community of the UASB reactors. The sludge was collected from valves installed on the side of the reactors at a height of 1.70 m and, when present, collected at the upper points of the blanket, eventually resulting in more than one sample per reactor and period, totaling 15 samples obtained. The samples were placed in sterilized vials and subsequently stored at 4 °C for one day before subsequent DNA extraction. DNA was extracted from the sampled sludge using the PowerSoil DNA Isolation Kit (MO BIO Laboratories Inc.) and later quantified with the Invitrogen Qubit Fluorimeter (Thermo Scientific), both processes being carried out according to the methodology proposed by the manufacturer.
The DNA samples resulting from the extraction process were submitted to the library preparation and sequencing steps. The preparation of libraries was carried out by PCR amplification of the V3/V4 gene regions of 16S rRNA, using primers 341F (CCTACGGGRSGCAGCAG) [33] and 806R (GGACTACHVGGGTWTCTAAT) [34]. In addition to the target region, a partial Illumina adapter, based on the TruSeq structure, was attached to the amplicons. After, the sequencing of the libraries was conducted using paired-end runs of 500 cycles, using V2 × 500 sequencing kits (Illumina, USA) with 100,000 coverage reads. The readings generated from the raw data were deposited in the database of the National Center for Biotechnology Information (NCBI) using the Sequence Read Archive (SRA) platform under the code BioProject PRJNA821214.
Bioinformatics analysis of the generated data
The bioinformatics analyses were initially processed using the program FastQC v.0.11.9 [35] to evaluate the quality of the sequenced nucleotides. The import and processing of the obtained sequences were conducted through the Quantitative Insights Into Microbial Ecology pipeline (QIIME2 version 2021.2) [36]. Sequences from primers and adapters were removed using the q2-cutadapt plugin with subsequent removal of noise, chimeras, and other sequencing errors as well as the merge of forward and reverse reads—considering the overlapping regions—using the Divisive Amplicon Denoising Algorithm (DADA2) [37] via the q2-dada2 plugin. Any singletons obtained in the process were discarded through the q2-feature-table plugin. The resulting Amplicon Sequence Variant (ASVs), fasta sequences, and the relative abundance values obtained were used to perform the taxonomic signature, functional prediction, construction of rarefaction curves, alpha, and beta diversity analyses.
The taxonomic annotation was processed using the feature-classifier plugin from the classify-sklearn method, using the SILVA database (v. 138) [38] obtained, processed, and trained for use in the QIIME2 program—considering the Naïve Bayes method—using the RESCRIPt plugin [39]. The taxonomic classification for the phylum category was based on the nomenclature proposed by Whitman et al. [40].
Analysis of alpha, beta diversity, rarefaction curves as well as PERMANOVA and Principal Coordinates (PCoA) were designed using the q2-phylogeny and q2-diversity plugins. For that, the Shannon–Wiener and Pielou Evenness indices were applied for alpha diversity, and Weighted Unifrac considered beta diversity [41,42,43,44].
Functional inferences were made using the Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt2) software [45], in the predefined configurations (default), using the ASVs generated in the previous step for sequence alignment (HMMER) and insertion into a phylogenetic tree of reference [46, 47]. The generated data were normalized by the 16S rRNA copy number, while the prediction of genes, enzymes, and metabolic functions was conducted using the Kyoto Encyclopedia of Genes and Genomes (KEGG) Orthology (KO) [48], Enzyme Commission Number (EC number), and MetaCyc [49]. The identified gene families were classified, considering the second and third functional level of classification (KEGG BRITE) and their respective relative abundance values obtained through the online software BURRITO [50].
Statistical analysis
The taxonomic data generated were submitted to differential abundance analysis using the R DESeq2 package [51], based on the extreme seasons (winter and summer). The resulting matrices were used to generate a Volcano-type plot, built using the ggplot2 package [52]. The representative identified taxa (equal to or greater than 1%) and their respective relative abundance values were used to generate a correlation matrix—using Spearman’s metric – and the subsequent construction of a network (graph), respectively, using the Past (version 4.06b) [53] and Gephi (version 0.9.2) software [54]. The taxonomic composition was also crossed with the physicochemical data and correlated through Canonical Correspondence Analysis (CCA) using the Past program (version 4.06b). The metabolic functions and the abundance values predicted in the PICRUSt2 and BURRITO software from the generated ASVs were used for the analysis of differential abundance, considering all seasons, using the Stamp software (version 2.1.3) [55], applying the one-way ANOVA statistical test, post-hoc test Tukey–Kramer. Other statistical analyses, including normality tests (Kolmogorov–Smirnov and Shapiro–Wilk), descriptive statistics, ANOVA, and correlation, were conducted in the SPSS program (version 20) [56].
Results and Discussion
Physicochemical analyses and meteorological conditions
Influent sewage, effluent, and sludge from UASB reactors were monitored from October 2020 to October 2021. The values obtained for the different physical–chemical parameters analyzed as well as the performance characteristics of the biodigesters are presented in Table 1, divided by seasons. The biodigesters operated at ambient temperature in the psychrophilic (< 20 °C) or mesophilic (20–40 °C) [57, 58] range during the study period, as verified by the temperature values of the influent sewage and the sludge from the biodigesters (see Table 1). It was verified that the ambient temperature was statistically different between the seasons (Kruskal–Wallis p = 0.00; significance ≤ 0.01), promoting significant changes in the temperature of the influent sewage (ANOVA p = 0.00; significance ≤ 0.01), except for autumn and winter periods, which did not differ (Dunnett T3 p = 0.745; significance ≤ 0.05). Temperature values in the psychrophilic range pose challenges for anaerobic digestion, since chemical reactions, which would normally take place in mesophilic conditions under lower energy expenditure, demand higher costs under these circumstances, thus some microorganisms, normally psychotrophic, tend to stand out for thermodynamic reasons, such as archaea and hydrogenotrophic bacteria, usually sulfate-reducing [59].
The evaluation of the parameters COD, PO4-P, TKN, and NH3-N (see Table 1) allowed us to classify the influent sewage from the biodigesters as low to medium concentrated, and the results are compatible with the usual concentration ranges found for domestic sewage [60,61,62]. Despite the low influent organic load during the study period, the reactors showed a good COD removal efficiency between 48.89 and 88.84% (above 80% during autumn and below 50% during summer). These results are satisfactory considering that UASB reactors were designed to treat wastewater with a high organic load [9]. In monitoring full-scale UASB reactors, Gaur et al. [63] found good COD removal capacity (51 ± 13.56%) treating domestic sewage with low organic loading rate (OLR); however, the values obtained were, on average, lower than those presented here.
The temperature and other variables related to seasonal changes did not promote significant changes in the efficiency of the reactors, in terms of COD removal (ANOVA p = 0.215; significance ≤ 0.05) between different seasons. In a study based on pilot-scale UASB reactors, Zhang et al. [22] also did not find significant changes in COD removal efficiency, operating at temperatures between 12 and 20 °C, thus obtaining average efficiency of 60%. The same author found a loss of efficiency, due to insufficient methanogenic capacity, only when the temperature decreased to 10ºC.
The results obtained for the SMA test at 37ºC revealed a low sludge activity of CH4 (0.017–0.096 gCODCH4.gVS−1.d−1) compared to average values for sludge from UASB digesters fed with domestic sewage situated between 0.10 to 0.20 gCODCH4.gVS−1.d−1 [64]. However, other works found numbers close to those described in the present study for UASB reactors. Bertolino et al. [65], treating sanitary sewage and using volatile suspended solids (VSS) values in the calculation, obtained a mean SMA of 0.0579 gCODCH4.gVSS−1.d−1, Agrawal et al. [66] numbers equivalent to 0.040 gCODCH4.gVSS−1.d−1 and employing a solution containing different organic acids—acetic, propionic, and butyric acid—Alves et al. [67] reached values in the range of 0.040 gCODCH4.gVS−1.d−1 for sludge from swine manure.
Regarding rainfall (Fig. S2), despite the reactors receiving mixed sewage—domestic sewage plus rainwater—no significant correlations were found between the average inflow of the system and the accumulated precipitation for the study period (Spearman = 0.495; p = 0.086; significance: ≤ 0.05), which suggests that the effects triggered by precipitation were punctual and not lasting to the point of influencing the average flow, HRT, as well as the efficiency in the COD removal process; However, it should be noted that the accumulated precipitation for months of study was low by regional standards. Punctual changes promoted by precipitation were also observed by Oliveira et al. [68], triggering a slight increase in the influent and effluent BOD, changes in pH, and alkalinity, although such changes did not impair the treatment efficiency, similar to the results found here. Other authors, however, have verified negative effects of precipitation, such as changes in the influent organic load, oxidation–reduction potential of the medium, conductivity and temperature decrease, parameters equally relevant for the anaerobic digestion process [69].
Characterization of the prokaryotic microbiota present in UASB biodigesters
The NGS sequencing analyses of the sludge samples collected from the 3 UASB reactors operating in full scale, from amplicons from the V3 and V4 regions of the 16S rRNA gene, resulted in a total of 1,451,602 reads recovered after pre-processing and denoising. In total, 3,384 ASVs, represented by more than one sequence, were found after applying the DADA2 algorithm, thus achieving stabilization in the rarefaction curves for all sample areas (Fig. S3). The number of ASVs obtained was higher than that found in other studies carried out in anaerobic reactors, operating at full scale and treating domestic sewage [70].
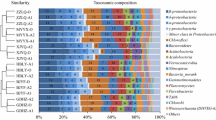
The taxonomic annotation from all ASVs recovered allowed for the identification of 50 phyla, of which 10 did not present a defined taxonomy (phylum candidate). Of the total number of phyla found, only 12 presented relative abundance greater than or equal to 1% in at least one season of the year (Fig. 1a). Some phyla showed visible seasonal variation in relative abundance, highlighting: Desulfobacterota (20.21–28.64%), Proteobacteria (7.48–24.90%), Bacteroidota (10.05–18.37%), and Caldisericota (9.49–17.20%). The only representative phylum inserted in the Archaea domain with an abundance greater than or equal to 1% was Halobacterota (3.23–6.55%) for the analyzed periods.
In anaerobic digesters, operating at full-scale and treating domestic sewage, other studies have identified the phyla Proteobacteria, Chloroflexi, Firmicutes, and Bacteroidota (Bacteroidetes) among the most abundant [70,71,72,73]. The exceptions presented were the phyla Acidobacteriota, Caldisericota and Desulfobacterota. The phylum Acidobacteriota is made up of members with an important role in the degradation of carbohydrates as well as in the acetogenic step of anaerobic digestion, notably, using aromatic compounds for this purpose [74,75,76]. Concerning the phylum Caldisericota, it currently has only one representative genus (Caldisericum) related to the reduction of sulfur compounds [77, 78]. Desulfobacterota is a new nomenclatural proposition, considering that some members of this group had phylogenetic inconsistencies, which did not support their classification in the previous phylum, Proteobacteria [79]. Representatives of this group are associated with the reductive activity of sulfur compounds, but some members subsist in anaerobic environments, where they produce acetate and butyrate from propionate in syntrophy with hydrogenotrophic methanogenic archaea [80, 81]. Regarding the phylum Halobacterota, recent studies reassessing phylogenetic aspects within the Archaea domain found problems in maintaining the phylum Euryarchaeota as a monophyletic clade, proposing its division into 3 new phyla, where the phylum Halobacterota (Halobacteriota) was created [82]. Members of this phylum are associated with methanogenesis (hydrogenotrophic, acetoclastic, and methylotrophic metabolic pathways), sulfur-reducing activity and some groups are present in hypersaline environments [83,84,85,86].
Within the taxonomic level of the genus, 655 distinct groups were found, 641 belonging to the Bacteria domain and 14 to the Archaea domain. Evaluating the total number of groups found, 46.87% received a taxonomic signature of the genus, and only 20 presented relative abundance greater than or equal to 1%, 10 of which were unclassified or not cultivable, considering at least one season of the year (Fig. 1b). Among the most abundant genera prevailed CI75cm.2.12 (5.34–22.45%), Smithella (9.60–17.36%), Caldisericum (9.49–17.20%), Bacteroidetes_vadinHA17 (6.12–9.46%), Methanolinea (2.51–4.81%) and Longilinea (2.21–3.93%).
CI75cm.2.12 (Family Nitrosococcaceae) was present in all samples, but with higher relative abundance for the summer and spring periods. The Nitrosococcaceae family is related to the nitrogen cycle, which in anaerobic environments may indicate the incidence of denitrification processes with reduction of NO3− to N2 [87, 88]. Considering that the studied reactors are followed by Submerged Aerated Biofilters (SABs) units—to oxidize the ammonia (NH3) present in the sewage to nitrate (NO3−) and that the sludge produced is recycled to the UASB reactors—the prevalence of the group in question could be explained, due to the availability of the electron acceptor nitrate for anaerobic respiration by these microorganisms. However, the group in question decreased for the autumn and winter periods, a fact that may be related to temperature since biological processes of nitrification are impaired with lower values for this variable, thus making NO3− unavailable for use by this organism [89,90,91,92,93].
Bacteroidetes_vadinHA17 belongs to the order Bacteroidales and is related to the metabolism of proteins and amino acids, in addition to being of great importance in UASB reactors in the degradation of complex organic compounds from different types of effluents, playing a relevant role in the stages of hydrolysis and acidogenesis [94,95,96,97].
Other important genera found, Longilinea and Leptolinea (0.95–1.77%), metabolize carbohydrates, the former in syntrophy with hydrogenotrophic archaea, both producing acetate, a substrate for the production of CH4 by the acetoclastic pathway [98, 99]. In addition to this fundamental association, Longilinea (Anaerolineaceae) seems to contribute to the formation of granules in reactors, together with acetoclastic archaea of the genus Methanosaeta (0.61–1.80%), helping in the efficiency, maintenance of sludge blankets and the anaerobic digestion process as a whole [95, 96, 100]. Also, acetogenic, the genera Syntrophorhabdus (1.46–1.98%) and Syntrophobacter (1.06–1.83%) were identified in the samples. However, the first is linked to the degradation of phenolic compounds, and the second uses propionic acid as substrate, both in syntrophy with hydrogenotrophic archaea [101, 102].
In the methanogenic stage, only the genera Methanolinea and Methanosaeta presented relative abundance above 1%, standing out from the other archaea found: Methanospirillum, Methanobacterium, Methanoregula, Methanosphaera, Methanomethylovorans, and Methanomassiliicoccus. The genus Methanolinea is associated with syntrophic systems where there is the degradation of propionic acid and production of H2 [103], corroborated by the strong presence of the Smithella acetogenic bacterium that metabolizes this substrate producing acetate and H2 [104]. Methanosaeta, on the other hand, is related to the production of methane by the acetoclastic pathway [105]. Zhang et al. [81] also reported the predominance of these genera in samples collected from reactors operating at full-scale treating domestic sewage. The prevalence of these genera is related to the syntrophic associations already reported. However, the more expressive abundance of a hydrogenotrophic genus (Methanolinea) can be explained by the greater susceptibility of acetotrophic archaea to environmental disturbances, often acting as indicators of potential failures in the anaerobic digestion process when in small numbers, since most of the CH4 produced in these systems occurs, under optimal conditions, by the acetoclastic pathway [6]. In addition, lower temperatures tend to favor the occurrence of hydrogenotrophic groups [25, 106, 107] and McKeown et al. [108] also reported the formation of microbial consortia in psychrophilic bands, where there is propionic acid oxidation in syntrophy with this class of archaea, in agreement with the results observed here. In the present work, a low occurrence of the genus Methanosaeta was observed, with greater abundance in the winter period, indicating a possible suppression of this taxon by factors not linked to temperature, such as: competition for substrate (acetate) or by the high concentration of inhibitory chemical compounds (H2S, NH3+, and Na+) [109]. It is worth mentioning that despite the low proportion of methanogenic organisms and the reduced values of SMA, no extreme changes were observed in the maintenance of the efficiency of COD removal from the system, indicating that organic matter removal occurred through other metabolic pathways.
The analyses also revealed the presence of other groups that are important for the operation of the biodigestion units. The genera Desulfovibrio (1.44–3.33%) and Caldisericum, which presents sulfidogenic activity, were found in considerable abundance in the analyzed samples. In a study conducted in UASB reactors, verifying the dynamics of the microbial community in the methanogenesis-sulfidogenesis transition process, Fernández-Palacios et al. [110] found that Desulfovibrio seems to be linked to a considerable impairment of methanogenesis via the acetoclastic pathway, promoting an accumulation of acetate in the biodigesters in the presence of sulfate. However, Borja [111] explained that the genus, in the absence of SO42− is associated with acetogenic activity, therefore not causing damage but rather contributing to the production of CH4. Sulfate-reducing bacteria can compete for carbon sources with methanogenic archaea. This process occurs when the COD/SO42− ratio is equal to or less than 1 or through the generated hydrogen sulfide (H2S), which is toxic to these organisms, hindering their growth and reducing the contribution of methane (CH4) to the biogas produced [112,113,114]. In addition to the biological aspect, excess H2S, as it is a highly corrosive gas, compromises pipes, structures, and equipment used in the construction of treatment units and eventual energy use of the biogas produced [64].
Differential abundance of representative groups
Regarding the differential abundance, considering the extreme seasons (summer and winter), Fig. 2 highlights the groups at the genus/ASV level that showed significant changes in their relative abundances through the use of the DESeq2 algorithm. Among the representative groups (Fig. 1b) that were statistically different, Methanolinea, CI75cm.2.12, Caldisericum, Lentimicrobiaceae_unclassified, and Arcobacter are present. Except for Arcobacter and Lentimicrobiaceae, whose abundance was higher in the winter, the other groups showed higher abundance in the summer.
Volcano plot showing differential abundance by genus/ASV considering winter and summer seasons. The red circles represent the groups that presented significant differential abundance (p < 0.05), while the triangles non-significant groups (p > 0.05). The values of the y-axis are expressed in -log10 of the p value and the x-axis in log2FoldChange, relative to the abundance of the genera, considering the winter and summer period
The groups reported with the highest occurrence for the summer period have some species whose optimal growth conditions are related to mesophilic and thermophilic environments. CI75cm.2.12, as already discussed, a possible link with nitrogen metabolism (denitrification), occurring in environments with NO3− availability, therefore dependent on nitrification processes that operate in aerobic environments under higher temperatures [88]. On the other hand, the groups reported with higher abundance for winter are linked to psychrophilic-mesophilic conditions, with species of the genus Arcobacter being described as pathogenic to humans [115, 116], but recent studies have revealed that some members of the group are related to acetate consumption and H2S oxidation in anaerobic environments and psychrophilic conditions [117]. The other representative genera (Fig. 1b), which did not show significant differential abundance, indicate that the UASB reactors studied are capable of maintaining a relatively stable microbial community throughout the year, even under the influence of environmental variations imposed by seasonal factors.
Relationship between representative taxonomic groups in the community
In anaerobic systems, the co-occurrence of certain groups of microorganisms can indicate the degree of mutual dependence to which they are subjected. Figure 3 presents the most representative genera and their correlations in the samples from the different UASB reactors. In the presented network, it is possible to observe interesting associations between members of the Anaerolineaceae family (phylum Chloroflexi), highlighted by the positive correlation between the Longilinea, Leptolinea, and Anaerolineaceae_unclassified, which play a relevant role in the structuring of sludge granules in anaerobic digesters [118, 119]. This family has been associated with the presence of the methanogenic genus Methanosaeta in anaerobic systems [120]; however, in the present study no significant correlations were identified, with the before-mentioned taxa.
Co-occurrence network showing the correlation between the most representative groups present in the sludge samples. The network was built using data from a Spearman correlation matrix and presents only connections whose significance level was ≤ 0.05. The connections in grey represent positive correlations, while in orange, negative correlations. Dark grey nodes have higher values for Closeness Centrality, while larger nodes have higher Betweenness Centrality. Correlation analyses were performed using the Past 4.06 program, and the network was built using the Gephi 0.9.2 program
The genus Methanosaeta was correlated with Syntrophobacter, Smithella (with expressive abundance in the samples), and Lentimicrobiaceae_unclassified – relevance in the production of acetate and VFA fermenting carbohydrates [121]—but highlighting the strong correlation with Desulfovibrio (Spearman = 0.935; p = 0.000; significance ≤ 0.01) which in the absence of sulfate can ferment several amino acids to acetate [122]. Methanosaeta still correlates with Spirochaetaceae_uncultured, a family which has some members with possible carbohydrate fermentative activity—producing acetate in syntrophy with hydrogenotrophic archaea—but groups with oxidative acetate metabolism have also been reported [123,124,125].
Among the hydrolytic and acidogenic groups, establishing several connections in the network, Bacteroidetes_VadinHA17 stands out, showing a correlation with several acetogenic groups in this system. As for the association observed between the Smithella genus and Bacteriodetes_vadinHA17, both with expressive abundance in the samples, it is feasible to consider that the fermentative capacity of the latter would result in the production of substrates capable of being used by the former, for example, the fermentation of some amino acids, such as methionine and threonine, which can be metabolized to propionic acid, a substrate used by some species of the Smithella genus [79].
Regarding the genus Caldisericum and the absence of significant correlations in the network with other groups, despite its expressive abundance, it is imperative to point out that its role in the metabolism of organic compounds in anaerobic systems remains unclear; however, some studies propose its participation in the amino acid degradation, ammonia assimilation, butyric acid consumption and even autotrophic growth using enzymes linked to the reverse citric acid cycle [126,127,128].
Another group with outstanding abundance in the study, CI75cm.2.12 (family Nitrosococcaceae), presents a positive correlation with the acetogenic elements Syntrophorhabdus and Syntrophales_uncultured, but several negative associations with other groups of importance for the functioning of the reactors, including Methanosaeta (Spearman = − 0.728; p = 0.002; significance ≤ 0.01), which would show possible inhibitory effects. Wang et al. [129] highlighted that some denitrifying groups could use acetate as an electron donor, which corroborates the negative relationships observed.
In the network, it is still possible to verify a positive link between the archaea Methanolinea with the genus Syntrophorhabdus, TPD58, and Holophagaceae_uncultured (family that has some genera with outstanding syntrophic importance in anaerobic systems, such as Holophaga and Geothrix). Holophaga is related to the production of acetate from aromatic compounds through the homoacetogenic pathway, while Geothrix is related to the oxidation of acetate, using the iron ion as an electron acceptor [75]. TPD58 (family Thermoanaerobaculaceae) is possibly associated with carbohydrate fermentation [130] and related to syntrophy with the mentioned archaea.
The calculations of the topological properties of the network allowed us to verify the importance of some taxa in the network (see Table S2). Considering the Closeness Centrality (CC), Betweeness Centrality (BC), and degree together, Bacteriodetes_VadinHA17 (CC = 0.8; BC = 6.25; degree = 9), Spirochaetaceae_uncultured (CC = 0.83; BC = 19.16; degree = 8), and Desulfovibrio (CC = 1; BC = 14.25; degree = 11) stood out. Besides these taxa, other groups showed high values for degree and low values for BC: ADurb.Bin063-1 (BC = 1.66; degree = 8), Methanosaeta (BC = 1.25; degree = 10), Lentimicrobiaceae_unclassified (BC = 4.41; degree = 8), and CI75cm.2.12 (BC = 0; degree = 10). In this context, groups that present higher numbers for the three coefficients may present synergistic or competitive relevance to others in the microbial community, while taxa showing lower values for BC and higher for degree may be considered keys, determining the structure of the network and promoting an increase or decrease in the diversity of the system [131,132,133,134].
Correspondence of abiotic factors with representative groups
Analyzing the graph obtained through the CCA (Fig. 4), it is possible to verify that the Spirochaetaceae_uncultured, Smithella, Methanosaeta, Desulfovibrio, and Arcobacter are associated with an increase in COD and OLR values. In anaerobic systems where low-concentrated sewage predominates, small increases in organic load would probably be able to sustain a greater diversity of species due to the addition of resources and consequent decrease in interspecific competition as well as a more favorable C/N ratio [135]. Reshmi et al. [136], studying estuarine environments, reported that a greater contribution of organic matter favored the occurrence of acetotrophic groups, such as the genus Methanosaeta, similar to the data presented in the present study.
CCA between environmental variables and representative taxonomic groups. SMA: Specific Methanogenic Activity. HRT: Hydraulic Retention Time. OLR: Organic Loading Rate. COD: Chemical Oxygen Demand. TKN: Total Kjedahl Nitrogen. Q: Flow. inf-T: Influent Temperature (ºC). air-T: Air Temperature (°C). Environmental variables are shown in red and green arrows to illustrate the weight of correlation with microbial genera, while taxonomic groups are represented in blue. The proximity between environmental variables and microorganisms reflects their correlation
The genus Methanolinea, the most abundant archaea in the samples, is correlated with an increase in the efficiency of the COD removal process and with higher levels of TKN. According to Bandara et al. [137], hydrogenotrophic archaea play a significant role in COD removal in biodigesters operating under psychrophilic conditions, and Ruiz-Sánchez et al. [138] also found an association between this group of archaea with increased nitrogen concentration. This taxon is extremely relevant, considering that secondary treatment in WWTPs aims to remove organic matter from the sewage [12].
The Longilinea, Leptolinea and Anaerolinaceae_unclassified showed a close correlation with the sludge SMA, contrary to what was expected if we consider the methanogenic groups. This factor may be related to the ability that members of the Anaerolineaceae family have in the granular structuring of the sludge. They form filamentous colonies, allowing syntrophic relationships to be processed more efficiently, in addition to acting in the production of acetate from carbohydrate fermentation [99, 100, 139].
As for VFAs, it is possible to identify that SAR324, Syntrophobacter, and Bacteroidetes_vadinHA17 are correlated to the concentration of propionic and valeric acids. Syntrophorhabdus and CI75cm.2.12 are related to acetic acid. In this context, the importance of Bacteroidetes_VadinHA17 in the production of VFAs is evident, mainly its hydrolytic and acidogenic role in the metabolism of proteins, amino acids, and carbohydrates. Syntrophobacter has a relevant acetogenic role, metabolizing propionic acid and SAR324 is linked to the degradation of VFAs and oxidation of sulfur compounds [97, 140, 141]. Syntrophorhabdus is also related to the production of acetic acid [102] and CI75cm.2.12 may be linked to the use of this substrate as an electron donor for NO3− reduction reactions [129], which would represent a potential competitor in methanogenic systems.
The increase in temperature (air and influent), an important factor in seasonal conditions, did not seem to exert a preponderant influence on the abundance of most of the identified taxa, with the exception of the CI75cm.2.12, which explains its massive presence in the summer period (Fig. 1b). Conversely, lower temperatures are correlated with a higher richness of representative taxa found, as highlighted in the CCA.
Diversity analysis for the sample periods studied
The alpha diversity analysis using Shannon index and Pielou evenness revealed greater diversity for winter, followed by spring/autumn and summer, showing significant differences between all seasons in relation to summer (Fig. S4). The greater diversity for the colder periods can be explained by the decrease in dominant taxa (greater equitability) that could inhibit or suppress competition other groups in a favorable climate situation (temperature), either by dispute related to resources or by the generation of by-products with potential toxicity [142]. From this perspective, toxic compounds, such as H2S, can act as growth inhibitors for many microorganisms [143] and a decrease in the relative abundance of the sulfidogenic taxon Caldisericum, as well as CI75cm.2.12 can be observed during the winter period, which could have promoted the increase in diversity for the colder season.
These results contradict other studies, which commonly associate the increase in temperature with a greater richness of Operational Taxonomic Units (OTUs) or ASVs [91]. However, the increase in alpha diversity related to the decrease in sludge temperature could be attested by the strong negative correlation (Spearman = 0.787; p = 0.000; significance ≤ 0.01) between the values obtained for the Shannon index and the variable in question. Another relevant aspect refers to the progressive increase in the number of rarer ASVs (summer to winter), verified by the greater presence of taxa with abundance lower than 1% (Others) (Fig. 1b), contributing to the increase in richness and consequently of the diversity in the samples [91]. In general, higher levels of diversity are desirable in biological systems since this would represent a greater repertoire of available redundant metabolic pathways, allowing the microbiota to adapt to eventual disturbances and maintain the integrity of the system [144].
As for the beta diversity, the application of the Weighted Unifrac metric, concomitant with the Principal Coordinate Analysis (PCoA), evaluating the proximity relationships between the microbial communities of the four seasons of the year, revealed more dissimilarity between winter-summer and autumn-spring, most of the difference explained by the x-axis (43.87%) of the PCoA graph. The mentioned similarity relations were supported by the Permutation Analysis of Variance (PERMANOVA), using 999 permutations and a significance level ≤ 0.05 (Fig. S5). These combined analyses showed differences between the microbial communities of the UASB reactors in detriment to seasonal conditions, especially considering the extreme seasons (summer and winter). This fact, possibly related to temperature, was also verified by other authors studying microbial communities in WWTPs [16, 91, 145]. However, despite the differences observed, there were no significant changes in the efficiency of the biodigesters, possibly indicating an overlap in the functions performed by taxa within the community. This fact was also verified by Resende et al. [146], but evaluating the microbiota from the biodigestion of bovine manure, considering pilot scale reactors, under ambient temperature conditions and aspects related to seasonal changes.
Functional prediction
The functional prediction analysis using the PICRUSt2 software, by comparing with the KO, EC, and MetaCyc databases, revealed the presence of 6,853 genes, 2,085 enzymes, and 373 metabolic pathways associated with sequences of ASVs obtained from sludge samples from UASB reactors for the periods evaluated. Considering the total number of genes identified (KO) 92.57%, 89.63%, 96.11%, and 95.40% were found for summer, autumn, winter, and spring.
The prediction evaluation was assessed using the Nearest Sequenced Taxon Index (NSTI) values calculated using the PICRUSt2 software for each ASV, with the numbers obtained being in the range of 0.352 to 0.434 (weighted NSTI) considering the seasons. Of the total number of ASVs obtained (3,384), only two were removed because they exceeded the cutoff limit stipulated as standard by the program (2.0). The NSTI values achieved are within the standard range configured in the PICRUSt2 software. However, they suggest low coverage – since poorly characterized environments present less availability of genomes in the database for phylogenetic comparisons performed by the program, thus not resulting in more accurate approximations [147].
From the genes found, using the BURRITO software and the KEGG BRITE classification hierarchy, it was possible to identify the main functions associated with the microbial community for the period studied. The most representative functions obtained for the SuperPathway category are linked to cellular mechanisms of translation (16.94–17.59%), membrane transport (14.48–14.93%), amino acid metabolism (7.74–8 0.00%), as well as DNA replication and repair (6.76–6.82%). In general, at this level of classification, there were no evident differences in the functions identified between the seasons. However, when considering more specific categories (SubPathway) differential abundance was observed (Fig. S6). The Alanine, aspartate, and glutamate metabolism and Transport functions were more abundant for the winter and autumn periods, differing statistically from the other seasons of the year. DNA replication proteins, Glyoxylate and dicarboxylate, and Terpenoid backbone biosynthesis showed differential abundance only between autumn and spring, while Protein folding and associated processing showed a statistical difference between spring-winter and spring–autumn.
From these data, it was possible to identify that some taxonomic groups were correlated with the above metabolic functions, possibly promoting the differences observed between the seasons (see Table 2). Thus, the correlation between Desulfovibrio, Methanosaeta, and Spirochaetaceae_uncultured with amino acid metabolism is highlighted, and Arcobacter, Bacteroidetes_VadinHA17, Lentimicrobiaceae_unclassified and Smithella in addition to this function, also associated with membrane transport and structural biosynthesis of terpenes. Except for Arcobacter, Lentimicrobiaceae_unclassified, Smithella, and Methanosaeta, the others have already been reported in the literature for presenting fermentative activity from amino acids [97, 122, 125]. Concerning terpene synthesis, there is a need to study the role of these taxa in their metabolism, assuming that these compounds have biotechnological applications, mainly in the production of drugs, insecticides, antimicrobials, and alternative fuels [148]. In the present work, the PWY-5910, PWY-922, and PWY-6174 pathways (mevalonate pathway) were found associated with the identified genes, related in prokaryotes to the synthesis of essential components of cell walls, membranes, electron transporters, and compounds used in the conversion from light into energy [149].
Syntrophales_uncultured and CI75cm.2.12 were correlated to the glyoxylate and dicarboxylate cycle. The glyoxylate cycle in prokaryotes is linked to the synthesis of carbohydrates from fatty acids and acetate – mainly under conditions of oxidative stress, infections, and antibiotic resistance – and has even been associated with cellular respiration and denitrification processes [150]. In the present study, the aceA (K01637) and aceB/glcB (K01638) genes which, respectively, encode the enzymes isocitrate lyase and malate synthase, were found in greater abundance for the summer and spring samples, periods in which the taxa were described in greater prevalence. These results, added to the data presented in Fig. 4, demonstrated that the CI75cm.2.12 probably uses acetate as an electron donor or carbon source, competing with acetoclastic archaea for this substrate and performing some type of function related to nitrogen metabolism, possibly acting in denitrification processes.
The expressive number of genes with functions related to the metabolism of amino acids and proteins (Fig. S6a), which in addition to having an essential cellular function, reflects the composition of domestic sewage, since these biomolecules are present in high concentration, reaching together with fibers, the largest portion of the organic matter present in these residues [151, 152]. This characteristic highlights another problem, the excess of hydrogen sulfide, considering that a large portion of the H2S generated in anaerobic systems treating domestic effluents derives from the metabolism of proteins and amino acids that have the thiol group (-SH) in their constitution [153]. On the other hand, the low relative abundance of functions associated with lipid metabolism may be related to the small concentration of these macromolecules in the domestic effluent that supplies the reactors, due to the installation of fat retaining boxes in homes and commercial establishments in the city, which prevent the arrival of excess of these compounds to the sewage system, consequently reducing the proliferation of microorganisms capable of processing these substrates. There is also a large number of unpredicted or unknown functions associated with the genes found, an important fact, as it is indicative that despite the high number of studies addressing microbial communities in anaerobic systems, it is still necessary to know the functions performed by these microorganisms in the environment.
The data generated also made it possible to determine the presence of several genes linked to the production of the main VFAs found, acetate, butyrate, and propionate. Regarding acetate synthesis, was verified the occurrence of folD (K01491) (Wood–Ljungdahl pathway), metF (K00297), and pta (K13788) genes, respectively, related to the enzymes methylenetetrahydrofolate dehydrogenase, methylenetetrahydrofolate reductase, and phosphate acetyltransferase [154,155,156]. About propionate, were found the genes PCCB (K01966), sucC and sucD (K01902, K01903), MUT (K01847), and MCEE (K05606) responsible for encoding the enzymes propionyl-CoA carboxylase, succinyl-CoA synthetase, methylmalonyl-CoA mutase and methylmalonyl-CoA epimerase [157]. And for butyrate, the genes buk (K00929), bcd (K00248), fadB (K00074), abfD (K14534) were predicted, in order, linked to the enzymes butyrate kinase, butyryl-CoA dehydrogenase, 3-hydroxybutyryl-CoA dehydrogenase and 4 -hydroxybutyryl-CoA dehydratase/vinylacetyl-CoA-Delta-isomerase [158]. Except for genes sucC and sucD, which presented higher values for the autumn and summer periods, the others were found in greater abundance for the autumn and winter periods (Fig. 5), coinciding with the greater presence of taxonomic groups linked to acidogenic and acetogenic processes as well as a greater diversity of ASVs verified (Fig. 1b, e Fig. S4).
Considering the methanogenic step, the main genes identified were associated with the acetotrophic (METH-ACETATE-PWY) and hydrogenotrophic (METHANOGENESIS-PWY) pathways, respectively presenting relative abundances in the range of 0.215–0.290% and 0.049–0.092% for the periods evaluated, the first being more abundant for the autumn–winter period and the second autumn-summer. These periods also coincide with the highest values of relative abundance recorded for the Methane and Energy metabolism functions (Fig. S6a) linked to CH4 biosynthesis. It surprising that the acetotrophic methanogenic pathway was more abundant, bearing in mind that the most representative methanogenic taxon identified was the genus Methanolinea (hydrogenotrophic). This fact was also verified by other authors [159] and may be linked to the overlap of functions presented by some taxa that have genes coding for enzymes related to both pathways, such as Formylmethanofuran dehydrogenase (K11261 gene; fwdE), identified in the present study.
Conclusion
The seasonal study of the sludge from the UASB reactors revealed the presence of a diverse microbial community represented by members related to the different stages of the anaerobic digestion process. Among them stood out Bacteroidetes_VadinHA17, with hydrolytic and acidogenic importance, the acetogenic bacterium Smithella and the methanogenic hydrogenotrophic archaea Methanolinea. The correlation between taxa revealed some already known syntrophic interactions and possible inhibitory effects caused by interspecific competition within the community. Methanolinea was linked to reactors efficiency and TKN; Anaerolineaceae family members to SMA, and other groups related to the increase of influent organic matter. However, some groups need to be better characterized to confirm their real function within the microbiome, among them Caldisericum, described in the literature as sulfidogenic, but not showing significant correlations with any taxon in the constructed co-occurrence network (Fig. 3) and CI75cm. 2.12 (family Nitrosococaceae) related to the nitrogen cycle, identified in this work, strongly related to temperature increase, acetate concentration and negatively correlated to several members of the microbial community. The present study also showed that the microbiota present in the digesters varied between seasons, showing high alpha diversity, including higher levels for the winter period.
The prediction of metabolic pathways revealed stability of the main functions at the SuperPathway categorical level and small variations in SubPathway for the seasons. Functions linked to amino acid and protein metabolism, membrane transport, glyoxylate cycle, and terpene synthesis showed differential abundance. The expressive number of genes linked to the metabolism of amino acids and proteins highlighted the importance of groups capable of metabolizing these substrates due to the composition of the sewage, highlighting the Bacteroidetes_VadinHA17. Surprisingly, a more fraction of the genes linked to the synthesis of the main VFAs found in the samples showed higher relative abundance for the autumn and winter periods.
The sludge from the reactors showed a low methane production capacity (< 0.10 gDQOCH4.gSV−1.day−1); however, the efficiency in COD removal remained stable even under psychrophilic circumstances. These facts allowed us to glimpse potential applications of sludge as inoculum in biodigesters, used in the treatment of domestic sewage and subjected to similar climatic conditions, although they do not support its viability for the production of CH4, for energy purposes.
Studies of this nature are important for understanding the dynamics of the microbiota under unfavorable situations and its relationship with the functioning of biodigesters, mainly to guide operational processes and possible modulations in these microbiomes. The importance of further studies with similar themes in anaerobic systems operated on a real scale is highlighted, remembering that in uncontrolled environments, abiotic conditions are heterogeneous and are subject to daily, monthly, and annual changes, unlike simulated experiments in laboratories. Thus, given the historical differences in seasonal characteristics in the region, further research addressing this issue should be carried out to confirm or refute the patterns and results obtained in the present work.
Data availability
The data used to support the findings of this study is available upon request.
References
UNESCO (2019) The United Nations World Water Development Report 2019: Leaving no one behind. UNESCO, Paris
UN-Habitat, WHO (2021) Progress on wastewater treatment: Global status and acceleration needs for SDG indicator 6.3.1. UN Habitat and WHO, Geneva
Sikosana ML, Sikhwivhilu K, Moutloali R, Madyira DM (2019) Municipal wastewater treatment technologies: a review. Procedia Manuf 35:1018–1024. https://doi.org/10.1016/j.promfg.2019.06.051
Abbasi T, Tauseef SM, Abbasi SA (2012) Biogas energy. Biogas Energy. https://doi.org/10.1007/978-1-4614-1040-9
Diamantis V, Eftaxias A, Stamatelatou K et al (2021) Bioenergy in the era of circular economy: anaerobic digestion technological solutions to produce biogas from lipid-rich wastes. Renew Energy 168:438–447. https://doi.org/10.1016/j.renene.2020.12.034
Rehman MLU, Iqbal A, Chang C-C et al (2019) Anaerobic digestion. Water Environ Res 91:1253–1271. https://doi.org/10.1002/wer.1219
Amin FR, Khalid H, El-Mashad HM et al (2021) Functions of bacteria and archaea participating in the bioconversion of organic waste for methane production. Sci Total Environ 763:1–21. https://doi.org/10.1016/j.scitotenv.2020.143007
Mainardis M, Buttazzoni M, Goi D (2020) Up-flow anaerobic sludge blanket (Uasb) technology for energy recovery: a review on state-of-the-art and recent technological advances. Bioengineering 7:1–29. https://doi.org/10.3390/bioengineering7020043
Daud MK, Rizvi H, Akram MF et al (2018) Review of upflow anaerobic sludge blanket reactor technology: effect of different parameters and developments for domestic wastewater treatment. J Chem 2018:1–13. https://doi.org/10.1155/2018/1596319
de Chernicharo CA, Ribeiro TB, Pegorini ES et al (2018) Contribuição para o aprimoramento de projeto, construção e operação de reatores UASB aplicados ao tratamento de esgoto sanitário - Parte 1: Tópicos de Interesse. Rev DAE 66:5–16. https://doi.org/10.4322/dae.2018.038
Cecconet D, Callegari A, Capodaglio AG (2022) UASB performance and perspectives in Urban wastewater treatment at sub-mesophilic operating temperature. Water (Switzerland) 14:1–13. https://doi.org/10.3390/w14010115
Metcalf & Eddy Inc (2014) Wastewater engineering: treatment and resource recovery, 15th edn. McGraw Hill, London
van Lier JB, van der Zee FP, Frijters CTMJ, Ersahin ME (2015) Celebrating 40 years anaerobic sludge bed reactors for industrial wastewater treatment. Rev Environ Sci Bio/Technology 14:681–702. https://doi.org/10.1007/s11157-015-9375-5
ANA (2020) Atualização da Base de Dados de Estações de Tratamento de Esgotos no Brasil. In: ANA (ed) Atlas Esgotos, 1st ed. ANA, Brasília, p. 47
Henze M, Comeau Y (2008) Wastewater Characterization. In: Henze M, van Loosdrecht MCM, Ekama GA, Brdjanovic D (eds) Biological wastewater treatment: principles, modelling and design, 1st edn. IWA Publishing, London, pp 33–53
Zhang B, Yu Q, Yan G et al (2018) Seasonal bacterial community succession in four typical wastewater treatment plants: Correlations between core microbes and process performance. Sci Rep 8:1–11. https://doi.org/10.1038/s41598-018-22683-1
Rashid SS, Liu YQ (2020) Assessing environmental impacts of large centralized wastewater treatment plants with combined or separate sewer systems in dry/wet seasons by using LCA. Environ Sci Pollut Res 27:15674–15690. https://doi.org/10.1007/s11356-020-08038-2
Leitão RC, Van Haandel AC, Zeeman G, Lettinga G (2006) The effects of operational and environmental variations on anaerobic wastewater treatment systems: a review. Bioresour Technol 97:1105–1118. https://doi.org/10.1016/j.biortech.2004.12.007
Marcos A, Al-Kassir A, López F et al (2012) Environmental treatment of slaughterhouse wastes in a continuously stirred anaerobic reactor: Effect of flow rate variation on biogas production. Fuel Process Technol 103:178–182. https://doi.org/10.1016/j.fuproc.2011.12.035
Zhao H, Li J, Li J et al (2013) Organic loading rate shock impact on operation and microbial communities in different anaerobic fixed-bed reactors. Bioresour Technol 140:211–219. https://doi.org/10.1016/j.biortech.2013.04.027
Ketheesan B, Stuckey DC (2015) Effects of hydraulic/organic shock/transient loads in anaerobic wastewater treatment: a review. Crit Rev Environ Sci Technol 45:2693–2727. https://doi.org/10.1080/10643389.2015.1046771
Zhang L, De Vrieze J, Hendrickx TLG et al (2018) Anaerobic treatment of raw domestic wastewater in a UASB-digester at 10 °C and microbial community dynamics. Chem Eng J 334:2088–2097. https://doi.org/10.1016/j.cej.2017.11.073
Callejas C, Fernández A, Passeggi M et al (2019) Microbiota adaptation after an alkaline pH perturbation in a full-scale UASB anaerobic reactor treating dairy wastewater. Bioprocess Biosyst Eng 42:2035–2046. https://doi.org/10.1007/s00449-019-02198-3
Delforno TP, Lacerda Júnior GV, Noronha MF et al (2017) Microbial diversity of a full-scale UASB reactor applied to poultry slaughterhouse wastewater treatment: integration of 16S rRNA gene amplicon and shotgun metagenomic sequencing. Microbiologyopen 6:1–12. https://doi.org/10.1002/mbo3.443
Seib MD, Berg KJ, Zitomer DH (2016) Influent wastewater microbiota and temperature influence anaerobic membrane bioreactor microbial community. Bioresour Technol 216:446–452. https://doi.org/10.1016/j.biortech.2016.05.098
Kuinchtner A, Buriol G (2001) Clima do estado do Rio Grande do Sul segundo a classificação climática de Köppen e Thornthwaite. Discip Sci Série Ciências Exatas 2:171–182
Alvares CA, Stape JL, Sentelhas PC et al (2014) Köppen’s climate classification map for Brazil. Meteorol Zeitschrift 22:711–728. https://doi.org/10.1127/0941-2948/2013/0507
APHA et al (2017) Standard Methods for Examination of Water and Wastewater, 23rd ed. American Public Health Association, Washington
Magrini FE, de Almeida GM, da Maia SD et al (2020) Effect of different heat treatments of inoculum on the production of hydrogen and volatile fatty acids by dark fermentation of sugarcane vinasse. Biomass Convers Biorefinery 11:2443–2456. https://doi.org/10.1007/s13399-020-00687-0
Chernicharo CA de L (2007) Anaerobic Reactors. In: Biological Wastewater Treatment Series, 1st ed. IWA Publishing, London, p. 190
Drosg B, Braun R, Bochmann G, Al Saedi T (2013) Analysis and characterisation of biogas feedstocks. In: Wellinger A, Murphy J, Baxter D (eds) The Biogas Handbook. Elsevier, pp 52–84
INMET (2022) Instituto Nacional de Meteorologia. In: INMET. https://tempo.inmet.gov.br/. Accessed 19 Mar 2022
Caporaso JG, Lauber CL, Walters WA et al (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. https://doi.org/10.1038/ismej.2012.8
Wang Y, Qian PY (2009) Conservative fragments in bacterial 16S rRNA genes and primer design for 16S ribosomal DNA amplicons in metagenomic studies. PLoS ONE 4:1–9. https://doi.org/10.1371/journal.pone.0007401
Andrews S, Lindenbaum P, Howard B, Ewels P (2010) FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
Robeson MS, O’Rourke DR, Kaehler BD et al (2021) RESCRIPt: reproducible sequence taxonomy reference database management. PLoS Comput Biol 17:1–37. https://doi.org/10.1371/journal.pcbi.1009581
Whitman WB, Oren A, Chuvochina M et al (2018) Proposal of the suffix –ota to denote phyla. Addendum to ‘proposal to include the rank of phylum in the international code of nomenclature of prokaryotes.’ Int J Syst Evol Microbiol 68:967–969. https://doi.org/10.1099/ijsem.0.002593
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27:379–423. https://doi.org/10.1002/j.1538-7305.1948.tb01338.x
Pielou EC (1966) The measurement of diversity in different types of biological collections. J Theor Biol 13:131–144. https://doi.org/10.1016/0022-5193(66)90013-0
Lozupone C, Lladser ME, Knights D et al (2011) UniFrac: an effective distance metric for microbial community comparison. ISME J 5:169–172. https://doi.org/10.1038/ismej.2010.133
Knight R, Vrbanac A, Taylor BC et al (2018) Best practices for analysing microbiomes. Nat Rev Microbiol 16:410–422. https://doi.org/10.1038/s41579-018-0029-9
Douglas GM, Maffei VJ, Zaneveld JR et al (2020) PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 38:685–688. https://doi.org/10.1038/s41587-020-0548-6
Czech L, Barbera P, Stamatakis A (2020) Genesis and Gappa: Processing, analyzing and visualizing phylogenetic (placement) data. Bioinformatics 36:3263–3265. https://doi.org/10.1093/bioinformatics/btaa070
Barbera P, Kozlov AM, Czech L et al (2019) EPA-ng: massively parallel evolutionary placement of genetic sequences. Syst Biol 68:365–369. https://doi.org/10.1093/sysbio/syy054
Kanehisa M, Sato Y, Kawashima M et al (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44:457–462. https://doi.org/10.1093/nar/gkv1070
Caspi R, Billington R, Ferrer L et al (2016) The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res 44:471–480. https://doi.org/10.1093/nar/gkv1164
McNally CP, Eng A, Noecker C et al (2018) BURRITO: an interactive multi-omic tool for visualizing taxa-function relationships in microbiome data. Front Microbiol 9:1–11. https://doi.org/10.3389/fmicb.2018.00365
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:1–21. https://doi.org/10.1186/s13059-014-0550-8
Wickham H (2009) ggplot2: elegant graphics for data analysis, 1st edn. Springer, London
Hammer Ø, Harper DAT, Ryan PD (2001) Past: paleontological statistics software package for education and data analysis. Palaentologia Electron 4:1–9
Bastian M, Heymann S, Jacomy M (2009) Gephi: An open source software for exploring and manipulating networks. Int. AAAI Conf. Weblogs Soc. Media 1–2
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124. https://doi.org/10.1093/bioinformatics/btu494
IBM Corp. (2011) IBM SPSS Statistics for Windows, Version 20.0
Moyer CL, Eric Collins R, Morita RY (2017) Psychrophiles and psychrotrophs. Ref Modul Life Sci. https://doi.org/10.1016/b978-0-12-809633-8.02282-2
Saavedra O, Escalera R, Heredia G et al (2019) Evaluation of a domestic wastewater treatment plant at an intermediate city in Cochabamba, Bolivia. Water Pract Technol 14:908–920. https://doi.org/10.2166/wpt.2019.071
Lettinga G, Rebac S, Zeeman G (2001) Challenge of psychrophilic anaerobic wastewater treatment. Trends Biotechnol 19:363–370. https://doi.org/10.1016/S0167-7799(01)01701-2
Jordão EP, Pessôa CA (1995) Tratamento de Esgotos Domésticos, 1st edn. ABES, Rio de Janeiro
Von Sperling M (2007) Biological wastewater treatment series: wastewater characteristics, treatment and disposal, 1st edn. IWA Publishing, London
Kang W, Chai H, Xiang Y et al (2017) Assessment of low concentration wastewater treatment operations with dewatered alum sludge-based sequencing batch constructed wetland system. Sci Rep 7:1–7. https://doi.org/10.1038/s41598-017-17783-3
Gaur RZ, Khan AA, Lew B et al (2017) Performance of full-scale uasb reactors treating low or medium strength municipal wastewater. Environ Process 4:137–146. https://doi.org/10.1007/s40710-017-0208-0
da Lobato LC, S, Ribeiro TB, Silva BS da, et al (2018) Contribuição para o aprimoramento de projeto, construção e operação de reatores UASB aplicados ao tratamento de esgoto sanitário - Parte 3: Gerenciamento de lodo e escuma. Rev DAE 66:30–55. https://doi.org/10.4322/dae.2018.040
Bertolino SM, Carvalho CF, Aquino SF (2008) Caracterização e biodegradabilidade aeróbia e anaeróbia dos esgotos produzidos em Campus universitário. Eng sanit Ambient 13:271–277. https://doi.org/10.1590/s1413-41522008000300005
Agrawal LK, Harada H, Okui H (1997) Treatment of dilute wastewater in a UASB reactor at a moderate temperature: Performance aspects. J Ferment Bioeng 83:179–184. https://doi.org/10.1016/S0922-338X(97)83579-9
Alves RGCM, Belli Filho P, Philippi LS, et al (2005) Digestores anaeróbios para tratamento de dejetos suínos: avaliação de partida para diferentes configurações de reatores. In: 23° Congresso Brasileiro de Engenharia Sanitária e Ambiental. ABES, Campo Grande, pp 1–7
Oliveira DBC, Soares WA, Holanda MACR (2020) Effects of rainwater intrusion on an activated sludge sewer treatment system. Rev Ambient e Agua 15:1–12. https://doi.org/10.4136/ambi-agua.2497
Wilén BM, Lumley D, Mattsson A, Mino T (2006) Rain events and their effect on effluent quality studied at a full scale activated sludge treatment plant. Water Sci Technol 54:201–208. https://doi.org/10.2166/wst.2006.721
Tonanzi B, Crognale S, Gianico A et al (2021) Microbial community successional changes in a full-scale mesophilic anaerobic digester from the start-up to the steady-state conditions. Microorganisms 9:1–15. https://doi.org/10.3390/microorganisms9122581
de Lucena RM, Gavazza S, Florencio L et al (2011) Study of the microbial diversity in a full-scale UASB reactor treating domestic wastewater. World J Microbiol Biotechnol 27:2893–2902. https://doi.org/10.1007/s11274-011-0771-x
Calusinska M, Goux X, Fossépré M et al (2018) A year of monitoring 20 mesophilic full-scale bioreactors reveals the existence of stable but different core microbiomes in bio-waste and wastewater anaerobic digestion systems. Biotechnol Biofuels 11:1–19. https://doi.org/10.1186/s13068-018-1195-8
Jiang C, Peces M, Andersen MH et al (2021) Characterizing the growing microorganisms at species level in 46 anaerobic digesters at Danish wastewater treatment plants: a six-year survey on microbial community structure and key drivers. Water Res 193:1–13. https://doi.org/10.1016/j.watres.2021.116871
Liesack W, Bak F, Stackebrandt J-UKE (1994) Holophaga foetida gen. nov., sp. nov., a new, homoacetogenic bacterium degrading methoxylated aromatic compounds. Arch Microbiol 162:85–90. https://doi.org/10.1007/s002030050106
Fukunaga Y, Ichikawa N (2014) The Class Holophagaceae. In: Rosenberg E, DeLong EF, Lory S et al (eds) The prokaryotes: other major lineages of bacteria and the archaea, 4th edn. Springer-Verlag, Berlin Heidelberg, pp 683–687
Kielak AM, Barreto CC, Kowalchuk GA et al (2016) The ecology of acidobacteria: moving beyond genes and genomes. Front Microbiol 7:1–16. https://doi.org/10.3389/fmicb.2016.00744
Mori K, Yamaguchi K, Sakiyama Y et al (2009) Caldisericum exile gen. nov., sp. nov., an anaerobic, thermophilic, filamentous bacterium of a novel bacterial phylum, Caldiserica phyl. nov., originally called the candidate phylum OP5, and description of Caldisericaceae fam. nov., Caldisericales ord. no. Int J Syst Evol Microbiol 59:2894–2898. https://doi.org/10.1099/ijs.0.010033-0
Aida AA, Hatamoto M, Yamamoto M et al (2014) Molecular characterization of anaerobic sulfur-oxidizing microbial communities in up-flow anaerobic sludge blanket reactor treating municipal sewage. J Biosci Bioeng 118:540–545. https://doi.org/10.1016/j.jbiosc.2014.04.011
Waite DW, Chuvochina M, Pelikan C et al (2020) Proposal to reclassify the proteobacterial classes deltaproteobacteria and oligoflexia, and the phylum thermodesulfobacteria into four phyla reflecting major functional capabilities. Int J Syst Evol Microbiol 70:5972–6016. https://doi.org/10.1099/ijsem.0.004213
Liu Y, Balkwill DL, Henry CA et al (1999) Characterization of the anaerobic propionate- degrading syntrophs Smithella propionica gen. nov., sp. nov. and Syntrophobacter wolinii. Int J Syst Bacteriol 49:545–556
Zhang Q, Wang M, Ma X et al (2019) High variations of methanogenic microorganisms drive full-scale anaerobic digestion process. Environ Int 126:543–551. https://doi.org/10.1016/j.envint.2019.03.005
Rinke C, Chuvochina M, Mussig AJ et al (2021) A standardized archaeal taxonomy for the genome taxonomy database. Nat Microbiol 6:946–959. https://doi.org/10.1038/s41564-021-00918-8
Aouad M, Borrel G, Brochier-Armanet C, Gribaldo S (2019) Evolutionary placement of Methanonatronarchaeia. Nat Microbiol 4:558–559. https://doi.org/10.1038/s41564-019-0359-z
Aouad M, Taib N, Oudart A et al (2018) Extreme halophilic archaea derive from two distinct methanogen Class II lineages. Mol Phylogenet Evol 127:46–54. https://doi.org/10.1016/j.ympev.2018.04.011
Laso-Pérez R, Hahn C, van Vliet DM et al (2019) Anaerobic degradation of non-methane alkanes by “ Candidatus Methanoliparia” in hydrocarbon seeps of the Gulf of Mexico. MBio 10:1–20. https://doi.org/10.1128/mBio.01814-19
Rosenberg E, Delong EF, Lory S et al (2014) Other major lineages of bacteria and the archaea, 4th edn. Springer, Berlin
Barton LL, Northup DE (2011) Microbial ecology, 1st edn. Wiley, New Jersey
Taylor HB, Kurtz HD Jr (2019) Composition, diversity, and activity of aerobic ammonia-oxidizing Bacteria and Archaea in the intertidal sands of a grand strand South Carolina beach. Microbiol Open 9:1–18. https://doi.org/10.1002/mbo3.1011
Zhu S, Chen S (2002) The impact of temperature on nitrification rate in fixed film biofilters. Aquac Eng 26:221–237. https://doi.org/10.1016/S0144-8609(02)00022-5
Ju F, Guo F, Ye L et al (2014) Metagenomic analysis on seasonal microbial variations of activated sludge from a full-scale wastewater treatment plant over 4 years. Environ Microbiol Rep 6:80–89. https://doi.org/10.1111/1758-2229.12110
Liu F, Hu X, Zhao X et al (2018) Microbial community structures’ response to seasonal variation in a full-scale municipal wastewater treatment plant. Environ Eng Sci. https://doi.org/10.1089/ees.2018.0280
Johnston J, LaPara T, Behrens S (2019) Composition and dynamics of the activated sludge microbiome during seasonal nitrification failure. Sci Rep 9:1–15. https://doi.org/10.1038/s41598-019-40872-4
Zhang Q, Chen X, Luo W et al (2019) Effects of temperature on the characteristics of nitrogen removal and microbial community in post solid-phase denitrification biofilter process. Int J Environ Res Public Health 16:1–15. https://doi.org/10.3390/ijerph16224466
Baldwin SA, Khoshnoodi M, Rezadehbashi M et al (2015) The microbial community of a passive biochemical reactor treating arsenic, zinc, and sulfate-rich seepage. Front Bioeng Biotechnol 3:1–13. https://doi.org/10.3389/fbioe.2015.00027
Chen C, Liang J, Yoza BA et al (2017) Evaluation of an up-flow anaerobic sludge bed (UASB) reactor containing diatomite and maifanite for the improved treatment of petroleum wastewater. Bioresour Technol 243:620–627. https://doi.org/10.1016/j.biortech.2017.06.171
Wang Q, Liang J, Zhan Y et al (2018) Treatment of petroleum wastewater using an up-flow anaerobic sludge blanket (UASB) reactor and turf soil as a support material. J Chem Technol Biotechnol 93:3317–3325. https://doi.org/10.1002/jctb.5694
Mei R, Nobu MK, Narihiro T, Liu WT (2020) Metagenomic and metatranscriptomic analyses revealed uncultured bacteroidales populations as the dominant proteolytic amino acid degraders in anaerobic digesters. Front Microbiol 11:1–11. https://doi.org/10.3389/fmicb.2020.593006
Yamada T, Sekiguchi Y, Hanada S et al (2006) Anaerolinea thermolimosa sp. nov., Levilinea saccharolytica gen. nov., sp. nov. and Leptolinea tardivitalis gen. nov., sp. nov., novel filamentous anaerobes, and description of the new classes Anaerolineae classis nov. and Caldilineae classis nov. in the. Int J Syst Evol Microbiol 56:1331–1340. https://doi.org/10.1099/ijs.0.64169-0
Yamada T, Imachi H, Ohashi A et al (2007) Bellilinea caldifistulae gen. nov., sp. nov and Longilinea arvoryzae gen. nov., sp. nov., strictly anaerobic, filamentous bacteria of the phylum Chloroflexi isolated from methanogenic propionate-degrading consortia. Int J Syst Evol Microbiol 57:2299–2306. https://doi.org/10.1099/ijs.0.65098-0
Bovio-Winkler P, Cabezas A, Etchebehere C (2021) Database mining to unravel the ecology of the phylum chloroflexi in methanogenic full scale bioreactors. Front Microbiol 11:1–16. https://doi.org/10.3389/fmicb.2020.603234
Chen S, Liu X, Dong X (2005) Syntrophobacter sulfatireducens sp. nov., a novel syntrophic, propionate-oxidizing bacterium isolated from UASB reactors. Int J Syst Evol Microbiol 55:1319–1324. https://doi.org/10.1099/ijs.0.63565-0
Qiu YL, Hanada S, Ohashi A et al (2008) Syntrophorhabdus aromaticivorans gen. nov., sp. nov., the first cultured anaerobe capable of degrading phenol to acetate in obligate syntrophic associations with a hydrogenotrophic methanogen. Appl Environ Microbiol 74:2051–2058. https://doi.org/10.1128/AEM.02378-07
Imachi H, Sakai S, Sekiguchi Y et al (2008) Methanolinea tarda gen. nov., sp. nov. a methane-producing archaeon isolated from a methanogenic digester sludge. Int J Syst Evol Microbiol 58:294–301. https://doi.org/10.1099/ijs.0.65394-0
Puengrang P, Suraraksa B, Prommeenate P et al (2020) Diverse microbial community profiles of propionate-degrading cultures derived from different sludge sources of anaerobic wastewater treatment plants. Microorganisms 8:1–14. https://doi.org/10.3390/microorganisms8020277
Patel G, Sprott D (1990) Methanosaeta concilii characterization. Int J Syst Bacteriol 40:79–82. https://doi.org/10.1099/00207713-40-1-79
O’Reilly J, Lee C, Collins G et al (2009) Quantitative and qualitative analysis of methanogenic communities in mesophilically and psychrophilically cultivated anaerobic granular biofilims. Water Res 43:3365–3374. https://doi.org/10.1016/j.watres.2009.03.039
Siggins A, Enright AM, O’Flaherty V (2011) Low-temperature (7 °C) anaerobic treatment of a trichloroethylene-contaminated wastewater: Microbial community development. Water Res 45:4035–4046. https://doi.org/10.1016/j.watres.2011.05.013
McKeown RM, Scully C, Enright AM et al (2009) Psychrophilic methanogenic community development during long-term cultivation of anaerobic granular biofilms. ISME J 3:1231–1242. https://doi.org/10.1038/ismej.2009.67
Lee J, Hwang S (2019) Single and combined inhibition of Methanosaeta concilii by ammonia, sodium ion and hydrogen sulfide. Bioresour Technol 281:401–411. https://doi.org/10.1016/j.biortech.2019.02.106
Fernández-Palacios E, Zhou X, Mora M, Gabriel D (2021) Microbial diversity dynamics in a methanogenic-sulfidogenic uasb reactor. Int J Environ Res Public Health 18:1–16. https://doi.org/10.3390/ijerph18031305
Borja R (2011) Biogas production. In: Moo-Young M (ed) Comprehensive biotechnology, 2nd edn. Elsevier, pp 785–798
Hu Y, Jing Z, Sudo Y et al (2015) Effect of influent COD/SO42- ratios on UASB treatment of a synthetic sulfate-containing wastewater. Chemosphere 130:24–33. https://doi.org/10.1016/j.chemosphere.2015.02.019
Wu J, Niu Q, Li L et al (2018) A gradual change between methanogenesis and sulfidogenesis during a long-term UASB treatment of sulfate-rich chemical wastewater. Sci Total Environ 636:168–176. https://doi.org/10.1016/j.scitotenv.2018.04.172
Wu J, Liu Q, Feng B et al (2019) Temperature effects on the methanogenesis enhancement and sulfidogenesis suppression in the UASB treatment of sulfate-rich methanol wastewater. Int Biodeterior Biodegrad 142:182–190. https://doi.org/10.1016/j.ibiod.2019.05.013
Fisher JC, Levican A, Figueras MJ, McLellan SL (2014) Population dynamics and ecology of Arcobacter in sewage. Front Microbiol 5:1–9. https://doi.org/10.3389/fmicb.2014.00525
Rovetto F, Carlier A, Van Den Abeele AM et al (2017) Characterization of the emerging zoonotic pathogen Arcobacter thereius by whole genome sequencing and comparative genomics. PLoS ONE 12:1–27. https://doi.org/10.1371/journal.pone.0180493
Callbeck CM, Pelzer C, Lavik G et al (2019) Arcobacter peruensis sp. nov., a chemolithoheterotroph isolated from sulfide-and organic-rich coastal waters off Peru. Appl Environ Microbiol 85:1–17. https://doi.org/10.1128/AEM.01344-19
Yamada T, Sekiguchi Y (2009) Cultivation of uncultured Chloroflexi subphyla: Significance and ecophysiology of formerly uncultured Chloroflexi “subphylum i” with natural and biotechnological relevance. Microbes Environ 24:205–216. https://doi.org/10.1264/jsme2.ME09151S
Owusu-Agyeman I, Eyice Ö, Cetecioglu Z, Plaza E (2019) The study of structure of anaerobic granules and methane producing pathways of pilot-scale UASB reactors treating municipal wastewater under sub-mesophilic conditions. Bioresour Technol 290:1–9. https://doi.org/10.1016/j.biortech.2019.121733
Liang B, Wang LY, Mbadinga SM et al (2015) Anaerolineaceae and Methanosaeta turned to be the dominant microorganisms in alkanes-dependent methanogenic culture after long-term of incubation. AMB Express 5:1–13. https://doi.org/10.1186/s13568-015-0117-4
Sun L, Toyonaga M, Ohashi A et al (2016) Lentimicrobium saccharophilum gen. nov., sp. nov., a strictly anaerobic bacterium representing a new family in the phylum bacteroidetes, and proposal of lentimicrobiaceae fam. nov. Int J Syst Evol Microbiol 66:2635–2642. https://doi.org/10.1099/ijsem.0.001103
Baena S, Fardeau ML, Labat M et al (1998) Desulfovibrio aminophilus sp. nov., a novel amino acid degrading and sulfate reducing bacterium from an anaerobic dairy wastewater lagoon. Syst Appl Microbiol 21:498–504. https://doi.org/10.1016/S0723-2020(98)80061-1
Lee SH, Park JH, Kang HJ et al (2013) Distribution and abundance of Spirochaetes in full-scale anaerobic digesters. Bioresour Technol 145:25–32. https://doi.org/10.1016/j.biortech.2013.02.070
Karami A, Sarshar M, Ranjbar R, Zanjani RS (2014) The Phylum Spirochaetaceae Ali. In: Rosenberg E, De Long EF, Lory S et al (eds) The prokaryotes: other major lineages of bacteria and the archaea. Springer, Berlin, pp 915–925
Delbès C, Moletta R, Godon JJ (2000) Monitoring of activity dynamics of an anaerobic digester bacterial community using 16S rRNA polymerase chain reaction- single-strand conformation polymorphism analysis. Environ Microbiol 2:506–515. https://doi.org/10.1046/j.1462-2920.2000.00132.x
Nobu MK, Narihiro T, Rinke C et al (2015) Microbial dark matter ecogenomics reveals complex synergistic networks in a methanogenic bioreactor. ISME J 9:1710–1722. https://doi.org/10.1038/ismej.2014.256
Probst AJ, Castelle CJ, Singh A et al (2017) Genomic resolution of a cold subsurface aquifer community provides metabolic insights for novel microbes adapted to high CO 2 concentrations. Environ Microbiol 19:459–474. https://doi.org/10.1111/1462-2920.13362
Martinez MA, Woodcroft BJ, Ignacio Espinoza JC et al (2019) Discovery and ecogenomic context of a global Caldiserica-related phylum active in thawing permafrost, Candidatus Cryosericota phylum nov., Ca. Cryosericia class nov., Ca. Cryosericales ord. nov., Ca. Cryosericaceae fam. nov., comprising the four species C. Syst Appl Microbiol 42:54–66. https://doi.org/10.1016/j.syapm.2018.12.003
Wang D, Huang Q, Wang C et al (2007) The effects of different electron donors on anaerobic nitrogen transformations and denitrification processes in Lake Taihu sediments. Hydrobiologia 581:71–77. https://doi.org/10.1007/s10750-006-0499-z
Dedysh SN, Losey NA, Lawson P (2020) Thermoanaerobaculaceae. In: Bergey’s Manual of Systematics of Archaea and Bacteria. John Wiley & Sons, Inc., pp 1–3
Liu Z, Ma A, Mathé E et al (2021) Network analyses in microbiome based on high-throughput multi-omics data. Brief Bioinform 22:1639–1655. https://doi.org/10.1093/bib/bbaa005
Ma B, Wang H, Dsouza M et al (2016) Geographic patterns of co-occurrence network topological features for soil microbiota at continental scale in eastern China. ISME J 10:1891–1901. https://doi.org/10.1038/ismej.2015.261
Agler MT, Ruhe J, Kroll S et al (2016) Microbial Hub Taxa Link host and abiotic factors to plant microbiome variation. PLoS Biol 14:1–31. https://doi.org/10.1371/journal.pbio.1002352
Berry D, Widder S (2014) Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front Microbiol 5:1–14. https://doi.org/10.3389/fmicb.2014.00219
Wang X, Lu X, Li F, Yang G (2014) Effects of temperature and Carbon-Nitrogen (C/N) ratio on the performance of anaerobic co-digestion of dairy manure, chicken manure and rice straw: focusing on ammonia inhibition. PLoS ONE 9:1–7. https://doi.org/10.1371/journal.pone.0097265
Reshmi RR, Deepa Nair K, Zachariah EJ, Vincent SGT (2014) Methanogenesis: seasonal changes in human impacted regions of Ashtamudi estuary (Kerala, South India). Estuar Coast Shelf Sci 156:144–154. https://doi.org/10.1016/j.ecss.2014.11.031
Bandara WMKRTW, Kindaichi T, Satoh H et al (2012) Anaerobic treatment of municipal wastewater at ambient temperature: Analysis of archaeal community structure and recovery of dissolved methane. Water Res 46:5756–5764. https://doi.org/10.1016/j.watres.2012.07.061
Ruiz-Sánchez J, Guivernau M, Fernández B et al (2019) Functional biodiversity and plasticity of methanogenic biomass from a full-scale mesophilic anaerobic digester treating nitrogen-rich agricultural wastes. Sci Total Environ 649:760–769. https://doi.org/10.1016/j.scitotenv.2018.08.165
Pierangeli GMF, Domingues MR, de Jesus TA et al (2021) Higher abundance of sediment methanogens and methanotrophs do not predict the atmospheric methane and carbon dioxide flows in eutrophic tropical freshwater reservoirs. Front Microbiol 12:1–15. https://doi.org/10.3389/fmicb.2021.647921
Sheik CS, Jain S, Dick GJ (2013) Metabolic flexibility of enigmatic SAR324 revealed through metagenomics and metatranscriptomics. Environ Microbiol 16:1–14. https://doi.org/10.1111/1462-2920.12165
Wang HZ, Lv XM, Yi Y et al (2019) Using DNA-based stable isotope probing to reveal novel propionate- and acetate-oxidizing bacteria in propionate-fed mesophilic anaerobic chemostats. Sci Rep 9:1–12. https://doi.org/10.1038/s41598-019-53849-0
Hibbing ME, Fuqua C, Parsek MR, Peterson SB (2010) Bacterial competition: surviving and thriving in the microbial jungle. Nat Rev Microbiol 8:15–25. https://doi.org/10.1038/nrmicro2259
Paulo LM, Stams AJM, Sousa DZ (2015) Methanogens, sulphate and heavy metals: a complex system. Rev Environ Sci Biotechnol 14:537–553. https://doi.org/10.1007/s11157-015-9387-1
Werner JJ, Knights D, Garcia ML et al (2011) Bacterial community structures are unique and resilient in full-scale bioenergy systems. Proc Natl Acad Sci USA 108:4158–4163. https://doi.org/10.1073/pnas.1015676108
Kang X-H, Leng Y, O MM, et al (2020) The seasonal changes of core bacterial community decide sewage purification in sub-plateau municipal sewage treatment plants. Bioprocess Biosyst Eng 43:1609–1617. https://doi.org/10.1007/s00449-020-02352-2
Resende JA, Godon JJ, Bonnafous A et al (2016) Seasonal Variation on microbial community and methane production during anaerobic digestion of cattle manure in Brazil. Microb Ecol 71:735–746. https://doi.org/10.1007/s00248-015-0647-y
Langille MGI, Zaneveld J, Caporaso JG et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Reddy GK, Leferink NGH, Umemura M et al (2020) Exploring novel bacterial terpene synthases. PLoS ONE 15:1–20. https://doi.org/10.1371/journal.pone.0232220
Boronat A, Rodríguez-Concepción M (2015) Terpenoid biosynthesis in prokaryotes. In: Schrader J, Bohlmann J (eds) Biotechnology of isoprenoids, 1st edn. Springer, Switzerland, pp 3–18
Ahn S, Jung J, Jang IA et al (2016) Role of glyoxylate shunt in oxidative stress response. J Biol Chem 291:11928–11938. https://doi.org/10.1074/jbc.M115.708149
Huang MH, Li YM, Gu GW (2010) Chemical composition of organic matters in domestic wastewater. Desalination 262:36–42. https://doi.org/10.1016/j.desal.2010.05.037
Rose C, Parker A, Jefferson B, Cartmell E (2015) The characterization of feces and urine: a review of the literature to inform advanced treatment technology. Crit Rev Environ Sci Technol 45:1827–1879. https://doi.org/10.1080/10643389.2014.1000761
Tian G, Xi J, Yeung M, Ren G (2020) Characteristics and mechanisms of H2S production in anaerobic digestion of food waste. Sci Total Environ 724:137977. https://doi.org/10.1016/j.scitotenv.2020.137977
Zhao C, Dong H, Zhang Y, Li Y (2019) Discovery of potential genes contributing to the biosynthesis of short-chain fatty acids and lactate in gut microbiota from systematic investigation in E. coli. NPJ Biofilms Microbiomes 5:1–8. https://doi.org/10.1038/s41522-019-0092-7
Lovell CR, Leaphart AB (2005) Community-level analysis: Key genes of CO2-reductive acetogenesis. Methods Enzymol 397:454–469. https://doi.org/10.1016/S0076-6879(05)97028-6
Breton-Deval L, Salinas-Peralta I, Aguirre JSA et al (2021) Taxonomic binning approaches and functional characteristics of the microbial community during the anaerobic digestion of hydrolyzed corncob. Energies 14:1–14. https://doi.org/10.3390/en14010066
Yang T, Mbadinga SM, Zhou L et al (2017) Propionate metabolism and diversity of relevant functional genes by in silico analysis and detection in subsurface petroleum reservoirs. World J Microbiol Biotechnol 33:1–10. https://doi.org/10.1007/s11274-017-2350-2
Kircher B, Woltemate S, Gutzki F et al (2022) Predicting butyrate- and propionate-forming bacteria of gut microbiota from sequencing data. bioRxiv. https://doi.org/10.1101/2022.03.06.48315
Hidalgo KJ, Saito T, Silva RS et al (2020) Microbiome taxonomic and functional profiles of two domestic sewage treatment systems. Biodegradation 32:17–36. https://doi.org/10.1007/s10532-020-09921-y
Acknowledgements
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES)—Finance Code 001.
Funding
Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, 001
Author information
Authors and Affiliations
Contributions
JG designed the study, carried out the experiments, analyzed the data, and wrote the paper; NLL carried out the experiments; JI carried out the experiments; FEM supervised the research; SP supervised the research. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gaio, J., Lora, N.L., Iltchenco, J. et al. Seasonal characterization of the prokaryotic microbiota of full-scale anaerobic UASB reactors treating domestic sewage in southern Brazil. Bioprocess Biosyst Eng 46, 69–87 (2023). https://doi.org/10.1007/s00449-022-02814-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-022-02814-9