Abstract
Anaerobic systems for domestic sewage treatment, like septic tanks and anaerobic filters, are used in developing countries due to favorable economic and functional features. The anaerobic filter is used for the treatment of the septic tank effluent, to improve the COD removal efficiency of the system. The microbial composition and diversity of the microbiome from two wastewater treatment systems (factory and rural school) were compared through 16S rRNA gene sequencing using MiSeq 2 × 250 bp Illumina sequencing platform. Additionally, 16S rRNA data were used to predict the functional profile of the microbial communities using PICRUSt2. Results indicated that hydrogenotrophic methanogens, like Methanobacterium, were found in higher abundance in both systems compared to acetotrophic methanogens belonging to Methanosaeta genus. Also, important syntrophic microorganisms (Smithella, Syntrophus, Syntrophobacter) were found in the factory and rural school wastewater treatment systems. Microbial communities were also compared between stages (septic tank and anaerobic filter) of each wastewater treatment stage, revealing that, in the case of the rural school, both microbial communities were quite similar most likely due to hydraulic short-circuit issues. Meanwhile, in the factory, microbial communities from the septic tank and anaerobic filter were different. The school system showed lower COD removal rates (2–30%), which were probably related to a higher abundance of Firmicutes members in addition to the hydraulic short-circuit and low abundance of Chloroflexi members. On the other hand, the fiberglass factory presented higher COD removal rates (60–83%), harboring phyla reported as the core microbiome of anaerobic digesters (Bacteroidetes, Chloroflexi, and Proteobacteria phyla). The knowledge of the structure and composition of wastewater treatment systems may provide support for the improvement of the pollutant removal in anaerobic process.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Septic tanks are the oldest, simplest, and common primary systems for the onsite treatment of domestic wastewater. These anaerobic systems are more used as decentralized alternatives for treatment and are very attractive due to some favorable economic and functional reasons like low-cost, easy installation, operation and maintenance, being widely used to meet residence and small agglomeration demands. This kind of wastewater treatment system is favored in tropical countries by the predominance of a warm climate (Silva et al. 2013).
Conventional septic tanks can provide suspended solids removal, solid storage and can function as an anaerobic bioreactor for digestion of the organic matter. However, there are several inherent problems associated to such systems, the most significant one being its low treatment efficiency (25–35% of COD) with limited removal efficiency (Cullimore and Viraraghavan 1994; Nasr and Mikhaeil 2013). Also, onsite treatment systems are not effective either in removing nitrate and phosphorus compounds or in reducing the abundance of pathogenic microorganisms (Cullimore and Viraraghavan 1994; Metcalf et al. 2014). To overcome these problems, it is necessary to improve the existing design and/or to add post-treatment stages. There are several options for the post-treatment of septic tank effluent (Panswad and Komolmethee 1997; Wu et al. 2011), the most recommended is the use of an anaerobic filter (Agency 2002; Raman and Chakladar 1972). The efficiency values of septic tanks can increase between 70 and 80% (COD) when combined with anaerobic filter (Von Sperling 2007).
The monitoring of wastewater treatment systems is crucial for determining the quality of effluent water and the efficiency of the process. In addition to the monitoring of physicochemical parameters, such as COD, pH, and total suspended solids, among others, the bio-monitoring or biological monitoring can reveal important information about the process (Senthil Kumar et al. 2018).
The septic tank and anaerobic filter treatment units are based on anaerobic digestion processes performed by microorganisms, and their efficiency is directly linked to microbial community diversity. Thus, the microbial communities in these systems are crucial for well-performing processes. However, monitoring and maintaining wastewater treatment systems are still based on empirical relationships between physicochemical and operational parameters, which are not enough for reliable performance (Liu et al. 2016b). A systematic understanding of the microorganisms present in the wastewater treatment systems as a function of environmental factors and how they influence the performance is important to improve process stability and efficiency and to provide important guidance in diagnosis and prognosis (Liu et al. 2016b).
In addition to the traditional cultivation-dependent microbiological procedures, other techniques allow a more precise investigation of the composition of microbial communities, including 16 rRNA gene clone libraries, community profiling techniques like denaturing gradient gel electrophoresis (DGGE), fluorescence in situ hybridization (FISH) as well as high throughput sequencing of 16S rRNA genes and whole metagenome (Shu et al. 2015). Large scale sequencing of 16S rRNA gene amplicons is a powerful tool to investigate the ecology of microbial-mediated processes (Senthil Kumar et al. 2018). This technique allows one to analyze the structure, composition, and diversity of the microbiome present in a given environment. Liu et al. (2016a; b) demonstrated seasonal variability in microbial communities in various wastewater treatment systems in China using high throughput sequencing of the 16S rRNA gene. Based on the biological information, the researchers were able to create a satisfactory model to predict the effluent BOD, suspended solids, and total nitrogen values, providing an alternative for the assessment of the performance of the wastewater treatment system (Liu et al. 2016b). Despite the availability of these tools to survey microbial communities, the treatment by septic tanks and anaerobic filters are poorly evaluated. Connelly et al. (2019) carried out large-scale sequencing of two septic tanks (conventional and solar) and observed the presence of key taxonomic groups that were robust over time. Additionally, the authors speculated that the microbiology of septic tanks is highly variable as a function of wastewater features. Thus, the survey of treatment systems with septic tanks and anaerobic filters could provide data for the improvement of anaerobic process stability (Connelly et al. 2019).
In this study, 16S rRNA gene high throughput sequencing was used to investigate the microbial diversity from two different systems composed by septic tank and anaerobic filter applied in the wastewater treatment in a factory and rural school. The aims of the present study were to (i) compare the taxonomic and functional prediction profiles between the two wastewater treatment systems (factory and rural school); (ii) compare the taxonomic and functional prediction profiles between the two stages (septic tank and anaerobic filter); and (iii) correlate the biological information with the physicochemical parameters.
There is a big gap in the literature on taxonomic profiles of wastewater treatment systems object of this study, which likely represents a barrier in achieving higher efficiency values of the processes.
Material and methods
Sampling details and site description
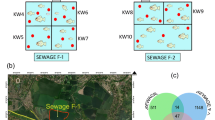
In this study, 14 sludge samples were collected from two domestic wastewater treatment systems composed of a septic tank combined with anaerobic filter (STANF). The samples were collected from the bottom of the tanks using sampler and 500 mL flasks previously “washed” with the sludge. One of the treatment systems studied is located at a fiberglass product manufacturing factory (22° 34′ 21.0″ S 47° 28′ 19.2″ W) and the other one at the rural school Professor Dorivaldo Damm (22° 32′ 16.1″ S 47° 16′ 48.0″ W), both in Limeira, SP (Fig. S1). Samples were collected in duplicate and different seasons of the year.
Description of the domestic wastewater treatment system: factory
The fiberglass manufacturing factory is located on the border of the municipalities Limeira and Iracemápolis, São Paulo, Brazil. Considering that the factory employs eight workers (sewage contribution estimated: 50–70 L person−1 day−1), the wastewater treatment system receives approximately 400–560 L day−1. The STANF is composed of a septic tank combined with an anaerobic filter and discharges the final effluent into a sinkhole (Fig. S2).
Description of the domestic wastewater treatment system: rural school
The rural school is located in the rural area of the municipality of Limeira, São Paulo, Brazil. The rural school has 677 students and 50–90 employees. The treatment system is composed of a grease trap, a septic tank, four anaerobic filters, and eight sinkholes (Fig. S3). The sewage generated is estimated between 36.500 and 38.800 L day−1 (sewage contribution estimated: 50 L person−1 day−1).
Sludge samples
At the rural school, liquid samples were collected in three points (Fig. S3), input and output of the anaerobic filter, and input of the septic tank. Sampling was performed in three seasons of 2018, autumn (June), winter (August), and spring (October). At the factory, liquid samples were collected in two points of the wastewater treatment system (Fig. S2), the input of the septic tank and output of the anaerobic filter, in September (winter) and October (spring) 2018 and January (summer) and April (autumn) 2019. All sludge samples were sampled in duplicate and kept on ice immediately after sampling. Samples were further stored at − 20 °C until DNA extraction.
Physicochemical analyses
Analysis of pH (4500B), fixed solids—FSS (2540E), volatile solids—VSS (2540E), alkalinity (2320B), nitrogen ammonia (4500-NH3 B) and chemical oxygen demand (5220D) were determined according to the Standard Methods for Examination of Water and Wastewater (APHA AWWA-WPCF 2005).
Molecular analyses
DNA extraction and sequencing of 16S rRNA genes
DNeasy Power Soil Pro Kit (Qiagen) was used for DNA extraction according to the manufacturer’s instructions. DNA integrity was checked through electrophoresis in 0.8% agarose gel. DNA quality was assessed by a NanoDrop1000 spectrophotometer (Thermo Fischer Scientific, Waltham, MA, USA).
The hypervariable V4 region of 16S rRNA gene from the microbial community of sludge samples was sequenced using Illumina MiSeq platform 2 × 250 bp by NGS Soluções Genômicas (Piracicaba, SP, Brazil) using the modified primers 515F (5′ GTGYCAGCMGCCGCGGTAA) and 806R (5′ GGACTACNVGGGTWTCTAAT) (Caporaso et al. 2010). In these primers, additional degeneracy was added to reduce bias against Crenarchaeota/Thaumarchaeota and the SAR11 clade (Apprill et al. 2015; Parada et al. 2016).
Taxonomic profiles
Quality control of sequencing reads was performed using FASTQ v.0.11.5 (Andrews 2010). Trimmomatic v. 0.39 (Bolger et al. 2014) was used to trim reads with quality lower than 30 (Phred score) and length smaller than 100 bp. Further bioinformatics analyses were carried out using Quantitative Insights into Microbial Ecology (QIIME2, version 2019.4, https://docs.qiime2.org/2019.4) (Bolyen et al. 2018) and its plugins. The ‘cutadapt’ plugin (Martin 2011) was used for primer trimming. “Qiime tools import” plugin was used for the import of demultiplexed quality filtered paired-end reads and the creation of the “artifact” file (type ‘SequencesWithQuality’) followed by denoising, chimera removal (consensus method) and clustering into representative sequences, amplicon sequence variants (ASVs), using “dada2” plugin (Callahan et al. 2016) (via q2-dada2). All ASVs were aligned with mafft (Katoh et al. 2002) (via q2-alignment) and used for phylogenetic reconstruction with fasttree2 (Price et al. 2010) (via q2-phylogeny). The taxonomic assignment of ASVs was done using q2-feature-classifier plugin (Bokulich et al. 2018) and classify-consensus-vsearch (Rognes et al. 2016) against the SILVA ribosomal RNA gene database version 132 (Quast et al. 2012).
Functional prediction profiles
Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt2) is a software for inferring metagenome functional content from 16S rRNA gene sequencing data. The output ASV table from taxonomic analysis (section 'Taxonomic profiles') was used as input into PICRUSt2 QIIME2 plugin (q2-picrust2) (Douglas et al. 2019) (https://github.com/picrust/picrust2/wiki/q2-picrust2-Tutorial). The KEGG ortholog database (Kanehisa et al. 2016) was used for inference from PICRUSt results. These results were normalized calculating the relative abundance of each KEGG ID. Specific metabolisms were explored, like carbon and energy metabolisms (KEGG level 2).
Statistical analyses
The downstream statistical analyses were developed in the R statistical environment (v. 3.6.1) (R Core Team 2013). The taxonomic and count tables and metadata files were imported as phyloseq object using the R package Phyloseq (McMurdie and Holmes 2013). The technical replicates were combined using the geometric mean. Principal coordinates analysis (PCoA) based on Bray–Curtis (Bray and Curtis 1957) and Weighted Unifrac (Lozupone et al. 2011) distances were performed to evaluate how divergent the technical replicates and the samples were before and after combining the replicates. Sequencing coverage was evaluated by rarefaction analysis. Alpha diversity indices, including Chao1 richness estimator (Chao 1984), observed species, Shannon–Wiener index H′ and Simpson evenness index J (Magurran 2013), were calculated using Phyloseq package in R. Statistical differences in alpha diversity indexes between both wastewater treatment systems were determined by the one-way ANOVA test. Before, data normality was assessed by the Shapiro–Wilk’s test. Shared families among the wastewater treatment systems were shown with a Venn diagram (Bardou et al. 2014). Microbial compositions were expressed as relative abundances at the levels of phylum, family, and genus. DESeq2-phyloseq was used to analyze the genera differently abundant (p < 0.05) between the two domestic wastewater treatment systems (Love et al. 2014; McMurdie and Holmes 2014). Sequences yielded in this study were deposited at the European Nucleotide Archive (ENA) database under the project accession number PRJEB36453.
Results and discussion
Physicochemical characterization
Eight samples were taken in the factory and nine samples in the rural school, at the initial, intermediate (only at the school) and final treatment stage, along the four seasons of the year, aiming at the physicochemical characterization of the sludge (Figs. S2 and S3).
The pH is an essential parameter for maintaining anaerobic metabolism in the treatment systems (Bodkhe 2009). The pH values (Table 1) of input and output in the STANF of the factory were in the range of 6.7–8.3 and 7.7–8.1, respectively, along the different seasons. Meanwhile, at the rural school, pH values were between 6.8–7.1 and 6.9–8.1 for the input and output of the septic tank, respectively, and 6.9–7.2 at the end of the treatment system (output of anaerobic filter). The alkalinity of influent and treated effluent at the factory was in the range of 235–644 mg/L and 240–610 mg/L, respectively, as CaCO3 (Table 1). Meanwhile, at the rural school the alkalinity of input and treated sludge was in the range of 509–685 mg/L and 593–836 mg/L, respectively, as CaCO3. The ratio of intermediate to total alkalinity is a simple measure of the stability of an anaerobic process. This ratio indicates a balance between acidogenesis and methanogenesis in the treatment system, and it should be maintained well below 0.5 for optimum functioning (Bodkhe 2008). In this work, these ratios varied from 0.13 to 0.28 at the factory and from 0.14 to 0.78 at the rural school.
Influent COD ranges were 195–254 and 742–962 at the factory and the rural school, respectively. Figure S4 shows the COD removal efficiency in the two treatment systems along the different seasons. COD reduction varied from 60 to 83% among the seasons at the factory (Fig. S4, green bars). Meanwhile, at the rural school, the COD removal rate was lower than 30% (Fig. S4, blue bars). This may be attributed to (i) issues with hydraulic short-circuit in the septic tank; (ii) unequal effluent distribution between the septic tank and the anaerobic filters (Fig. S3b); and (iii) a higher input COD concentration, which was possibly due to a higher organic load related to food preparation. Although there were two steps of treatment in the rural school, the low COD removal rate in the school was equivalent to the observed in septic tanks (Von Sperling 2007), indicating that hydraulic issues may have hindered organic removal.
The concentration of ammonia in the influent varied in the range of 45–154 mg/L at the factory and 101–225 mg/L at the rural school. At the factory, 61.9% of the ammonia was incorporated in spring and 31.5% in summer. While at the rural school, 26.3% of the ammonia was incorporated in spring and 25% in winter. Despite a high concentration of ammonia that can inhibit the anaerobic process (Von Sperling and de Lemos Chernicharo 2005), the concentrations observed in the two treatment systems were below the levels reported inhibiting microbial growth (> 1500 mg/L) (McCarty 1964).
Microbiome composition of sludge samples
Sequencing statistics
Twenty-eight sequence datasets were obtained, derived from 16 rRNA gene sequencing of 14 samples in duplicate. The 16S rRNA gene sequencing yielded 9.8–16.8 × 104 sequences per sample as raw data, with an average length of 271 bp. On average, after trimming, denoising, and chimera removal, 30.98% of sequences were removed due to low quality, and the average lengths were reduced to 100–225 bp (Table S.1). The average GC-content was 53%.
Assessment of microbial community composition and structure in the two wastewater treatment systems
Microbial community analysis of sludge samples based on 16S rRNA gene sequencing allowed clustering of sequences into ASVs and subsequent alpha diversity calculations (Table 2 and Fig. 1). The number of ASVs obtained ranged from 402 to 773 and 848 to 1013, for factory and rural school datasets, respectively (Table 2). Rarefaction analyses showed that all samples reached the plateau, indicating that sequencing effort was enough to cover the microbial diversity in these treated and untreated sludge samples (Fig. 1a). In Table 2, Chao1 (species richness estimator) values indicated that species richness was higher in all rural school sludge samples, especially in the septic tank. The highest species richness was observed in the autumn sample (rural school, Septic Tank Autumn/18). ANOVA analyses (p < 0.01) showed a significant effect of the wastewater treatment system (factory or rural school) on the observed ASVs (p-value 0.00006), Chao1 (p-value 0.00006) and Shannon (p-value 0.00903) index (Table 2), confirming higher abundance and diversity in the samples from the rural school.
a Rarefaction curves of species richness as a function of sequencing effort of microbial communities from sludge samples of the two domestic wastewater treatment systems; b Multidimensional scaling analysis based on pairwise Bray–Curtis (the first two coordinates are shown, representing 74.7% of the total variance), and c Weighted UniFrac (representing 80.4% of the total variance) distance matrices of the bacterial communities; colors represent samples and shapes represent stage of treatment (circle—septic tank, square—anaerobic filter)
MDS analysis of microbial communities of all samples based on Bray–Curtis and Weighted UniFrac distances matrices allowed clustering of samples by treatment system (factory and rural school). In addition, samples of the factory were separated into two groups according to the stage of treatment (septic tank and anaerobic filter) (Fig. 1b), indicating that microbial communities are different between the two units of treatment in the factory. On the other hand, microbial communities from the two stages of treatment at the rural school were more similar. This similarity observed in samples from the septic tank and anaerobic filter at the rural school corroborates the possibility of hydraulic short-circuit. Nevertheless, Weighted UniFrac analysis (Fig. 1c), which takes the phylogenetic distance between ASVs into account, showed that microbial communities from all samples are phylogenetically close to each other. Despite the variation of the mean temperature of the region (ranging from 17 to 26 °C when the samples were collected), no impact in the microbial community related to the different seasons was observed. Most likely, the heat generated by the biological process and the placement of treatment units reduced the susceptibility to external temperature variations. The anaerobic filter and septic tank in both systems were placed buried in the soil, reducing the heat exchange with the external environment.
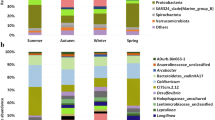
Sequencing of 16S rRNA genes in all sludge samples detected 836 ASVs. These ASVs were classified into 56 phyla, 470 families, and 836 genera, with 78.3% classified at the family level and 74.6% at the genus level. Taxonomic affiliation revealed that bacteria accounted for 88–96% and 92–97% and archaea for 3.8–12% and 2.4–7.7% of the total community in the factory and the rural school, respectively. The use of methodologies such as metagenomics and metatranscriptomics, aiming to scrutinize microbial community composition, structure, and function, confirmed that members of the Bacteria domain are largely responsible for carbon removal in activated sludge from municipal wastewater treatment system (Sánchez et al. 2011; Yu and Zhang 2012) while the Archaea domain seems to be less relevant (Ferrera and Sánchez 2016). At the phylum level (Fig. 2a), more than 50 distinct phyla were observed. Proteobacteria, Bacteroidetes, Chloroflexi, and Firmicutes were the most representative phyla in the factory sludge samples; whereas Proteobacteria, followed by Firmicutes and Bacteroidetes were the most abundant phyla in the rural school. Studies of microbial community composition and structure in different wastewater treatment plants have demonstrated the dominance of Proteobacteria members, followed by other phyla, like Bacteroidetes, Chloroflexi, Firmicutes, among others, depending on the prevalent conditions (Boon et al. 2002; Snaidr et al. 1997; Wagner et al. 1994; Wang et al. 2011). Chloroflexi, Proteobacteria, Bacteroidetes and Synergistetes phyla belong to a core group of common phylotypes from anaerobic digesters (Riviere et al. 2009). In the present study, samples from the fiberglass factory were the ones that showed microbial community structure most similar to the core group. Proteobacteria and Firmicutes are important groups of microbes in anaerobic treatment systems because members of the classes β-proteobacteria and γ-proteobacteria are involved in the first steps of the degradation process (Riviere et al. 2009) and are propionate, butyrate, and acetate-utilizers, and Firmicutes members correspond to the butyrate-utilizing microbes (Ariesyady et al. 2007), including syntrophic bacteria that can degrade volatile fatty acids (Riviere et al. 2009). These products can be degraded to hydrogen, which is then degraded by hydrogenotrophic methanogens (Riviere et al. 2009). The Chloroflexi phylum comprises a fermenting bacterial group frequently reported in anaerobic reactors that may have a role in the granulation due to their filamentous morphology (Bovio et al. 2019; Demirel and Scherer 2008; Fernández et al. 2008; Riviere et al. 2009). Bacteroidetes are known as proteolytic bacteria (Kindaichi et al. 2004), able to degrade proteins and then to ferment amino acids to acetate (Riviere et al. 2009).
The Euryarchaeota phylum (2.4–11.6% relative abundance) was the most abundant archaeal group in the sludge samples, followed by Omnitrophicaeota (0.03–1.4% relative abundance). The methanogenic archaea belong to the Euryarchaeota phylum, mainly present in anaerobic processes, generating methane as a value-added by-product (Ferrera and Sánchez 2016). Methanogenic archaea are active at the end of the anaerobic degradation process and are very specialized because they can degrade a very limited number of substrates (Demirel and Scherer 2008; Fernández et al. 2008).
In the factory wastewater treatment system, the most representative families (Fig. 2b) were Anaerolineaceae, Bacteroidetes VadinHA17, Methanobacteriaceae, and Syntrophaceae. Meanwhile, Eubacteriaceae was the most abundant family in the rural school treatment system, followed by Moraxellaceae, Methanobacteriaceae, and Syntrophaceae (Fig. 2b). Members of the Bacteroidetes environmental group VadinHA17 were isolated from an anaerobic digester treating winery wastewater (Baldwin et al. 2015).
The predominant genera in the factory were Smithella (5.8%), Methanobacterium (3.2%), Syntrophohabdus (3.0%), Syntrophus (2.4%), and Syntrophobacter (1.7%). Meanwhile, Acinetobacter (7.7%), Smithella (4.4%), Arcobacter (2.9%), Methanobacterium (2.89%), and Syntrophobacter (2.4%) were the top five most abundant genera in the rural school. Acinetobacter has been related to fecal contamination (McLellan et al. 2010). The genera Smithella and Syntrophobacter are considered typical obligate anaerobic syntrophic bacteria, able to oxidize propionate and fatty acid under methanogenic, mesophilic temperature conditions in association with hydrogen scavengers such as members of Methanomicrobiaceae or Methanobacteriaceae (Embree et al. 2015; Ju et al. 2017; Kato and Watanabe 2009; McInerney et al. 2015) via hydrogenotrophic methanogenesis pathway (Werner et al. 2011). The study of the sewage community composition mainly focuses on syntrophic microorganisms associated with methanogenic archaea (McInerney et al. 2009; Narihiro et al. 2015). Functional redundancy, especially in hydrogen production, and the high diversity among syntrophs of the same genus, make them more resistant and resilient to disturbances, which is essential to maintain robust anaerobic treatment systems over time (Allison and Martiny 2008; Fujimoto et al. 2019). The proportion of syntrophic bacteria (Smithella, Syntrophus, Syntrophobacter, Syntrophohabdus) was higher in the factory than in the rural school. Most likely, the higher proportion of syntrophic bacteria in the factory could be due to a stimulus caused by the presence of solvents or other toxic compounds, used in fiberglass manufacturing, in the sewage. Torres et al. (2018) used an upflow anaerobic sludge blanket reactors (UASB) to treat wastewater polluted with organic solvents, like ethanol, ethyl acetate and 1-ethyoxy-2-propanol. Analysis of the microbial community revealed that the syntrophic microorganisms predominated the reactor and were able to produce acetate and propionate from ethanol (Torres et al. 2018).
Many sequence analyses of methanogenic sludge have shown that archaeal communities are mainly composed of Methanoregulaceae (hydrogenotrophic), Methanosarcinaceae (hydrogenotrophic/acetotrophic/methylotrophic) and Methanobacteriaceae (hydrogenotrophic) (Ferrera and Sánchez 2016). The hydrogen solubility is higher at low temperatures, thus the hydrogenotrophic methanogenesis pathway is favored, because the substrate availability is increased, resulting in the availability of free energy for microbial growth (Kotsyurbenko 2005). During anaerobic treatment of various wastewaters at different temperatures, the hydrogenotrophic methanogenesis pathway was found to be dominant (Álvarez et al. 2008; Bialek et al. 2012; Connaughton et al. 2006; McHugh et al. 2004). In the present study, the hydrogenotrophic methanogen Methanobacterium was found in higher abundance in both systems compared to the acetotrophic methanogen Methanosaeta. In a previous study on the treatment of long-chain fatty acids (LCFA) at mesophilic conditions, Methanobacterium members were found at a higher relative abundance than Methanosaeta (Duarte et al. 2018). It was also found that Methanobacterium spp. were less sensitive to hydraulic and organic shocks than Methanosaeta (Kundu et al. 2013). Significant enrichment of Methanobacterium could be produced by the increase in the hydrogen concentration (Wu et al. 2019) because this methanogenic archaea can produce methane by reducing CO2 with H2 and formic acid and was often observed in organic waste anaerobic treatments and phenol-degrading enrichment cultures (Chen et al. 2008; Sierra et al. 2018; Zhang et al. 2018). In other studies, the predominant genera were acetotrophic methanogens such as Methanosaeta and Methanosarcina (Ariesyady et al. 2007; Fredriksson et al. 2012; Stets et al. 2014). Herein, Methanosaeta was found in higher relative abundance in the factory than in the school (data not shown) and Methanosarcina was practically absent in both systems. Methanosaeta requires a lower concentration of acetate than Methanosarcina and is more resistant to environmental changes (Liu and Whitman 2008). In the absence of acetotrophic methanogens, acetate can be converted to hydrogen by syntrophic acetate-oxidizing bacteria, thus stimulating the hydrogenotrophic pathway (Karakashev et al. 2006). Members of the Clostridia class (i.e. Clostridiaceae, Eubacteriaceae, Ruminococcaceae) can perform syntrophic acetate oxidation (De Vrieze et al. 2015; Müller et al. 2016). A high relative abundance of Eubacteriaceae was observed at the rural school, and Clostridiaceae and Ruminococcaceae in the septic tank and the anaerobic filter at the factory, respectively, suggesting that the hydrogenotrophic methanogenesis pathway is stimulated by the syntrophic association in both sites. However, it is known that syntrophic acetate oxidizers are slow producers, and thus less competitive than acetotrophic methanogens (Westerholm et al. 2016), as observed in the differential efficiency between the factory and the rural school wastewater treatment systems. The hydrogenotrophic methanogenic pathway seems to be feasible in both sites. However, it is more efficient in the factory, probably due to community composition, syntrophic consortia, and the lack of issues of hydraulic short-circuit.
Diverse studies about wastewater treatment show that the most common methanogen species found depend on the type of wastewater input. Dairy wastes are characterized by high levels of free ammonia (3.6–6.1 mg/L) (Karakashev et al. 2005) and volatile fatty acids, and members of Methanosarcinaceae, Methanomicrobiales, and Methanobacterium thermoautotrophicum are present as dominant methanogens (Ahring 1995; Baloch et al. 2007; Karakashev et al. 2005; Koster and Lettinga 1984). Meanwhile, cheese whey wastewater is characterized by the presence of long-chain fatty acids, mainly oleic acid, and species such as Methanosaeta spp. and Methanobrevibacter spp. (Demirel and Yenigün 2006; Ince et al. 2003; Perle et al. 1995; Uyanik et al. 2002). In the pulp and paper wastewaters, toxic and recalcitrant compounds (i.e. lignins, resins, tannins, and highly chlorinated organics, such as chlorophenolic compounds) are commonly found and Methanosarcina spp. and Methanobacterium spp. occur as dominant methanogenic archaea (Bolaños et al. 2005; Buzzini and Pires 2002; Buzzini et al. 2006; Demirel and Scherer 2008; Kortekaas et al. 1998; Ney et al. 1991; Roest et al. 2005; Sierra-Alvarez 1990). As already mentioned, at the rural school, food preparation is a daily activity. The methanogen Methanobrevibacter was found in much higher relative abundance in the rural school system than in the factory one. And Methanobacterium is slightly more abundant in the factory than in the rural school. However, the relative abundances of such phylotypes are low (~ 0.4%) in the total microbial community.
Madigou et al. (2019) evaluated the consequences of abrupt temperature modification on the microbiota of anaerobic digestion (Madigou et al. 2019). They operated two different size reactors which were subjected to temperature increase and decrease, first 35 ºC followed by an abrupt increase to 55 ºC and finally a decrease to 35 °C. They observed microbial shifts after an increase or decrease of temperature and concluded that the bacteria Ruminococcaceae and the archaea Methanobacterium were able to recover their initial abundances after abrupt temperature modifications, suggesting that they were key drivers of bioreactor resilience after return to the mesophilic condition. Mesophilic processes are more stable and robust, requiring less energy and with less probability of inhibition by long-chain fatty acids, thus resulting in a more efficient system (Fernández-Rodríguez et al. 2015). Ruminococcaceae family was found as the predominant phylotype in a methanogenic culture enriched with Methyl tert-butyl ether (MTBE) (Liu et al. 2016a). All members of this family are obligate anaerobes and can produce acetate from hydrogen and carbon dioxide using the acetyl-CoA pathway (Drake et al. 2006).
The different stages (septic tank and anaerobic filter) of each treatment system (factory and rural school) showed a taxonomic overlap of microbial families (Fig. S6). In total, 96 families were shared between both systems. The number of families shared between each stage in the rural school (46) was higher than in the factory (9). In the rural school, 22 and 23 families were exclusive in the septic tank and the anaerobic filter, respectively. In the factory, 11 and 18 families were exclusive in the septic tank and anaerobic filter, respectively. These results are congruent with Bray–Curtis analyses, which showed that in the factory there was higher speciation between the two stages of the treatment compared to the rural school (Fig. 1b). The anaerobic treatment process is composed of four stages (i.e. hydrolysis, acidogenesis, acetogenesis, and methanogenesis) (Angelidaki and Batstone 2011). The acidogens and methanogens have different growth rates, nutritional requirements, and resistance to environmental changes, such as variation in temperature and input sewage characteristics (Shi et al. 2019). Thus, the two-stage or two-phase anaerobic system treatment, such as septic tank-anaerobic filter, has been proposed to optimize conditions for the growth of both kinds of microorganisms and to improve efficiency (Azbar and Speece 2001; Pohland and Ghosh 1971; Shi et al. 2019). The lack of differentiation in the microbial species between the two phases of the rural school wastewater treatment is likely a result of hydraulic short-circuit issues, contributing to a low removal of COD.
The volcano plot (Fig. 3) showed a differential abundance of genera in each site. In the factory wastewater treatment system, genera like Ruminococcaceae UCG-010, Methylobacter, Methanomethylovorans, Syntrophus, Leptolinea, and Methanolinea, among others, had higher abundance compared with the rural school treatment system. Petriglieri et al. (2018) characterized microbial communities in more than 30 anaerobic digesters and observed that Leptolinea was the dominant Chloroflexi member (Petriglieri et al. 2018). Leptolinea is a filamentous and strict anaerobic bacterium, commonly found on the surface of granular sludge (Delforno et al. 2014; Yamada et al. 2006). Zhang et al. (2019) studied the microbial communities present in different full-scale anaerobic digesters and linked them to the digester performance. They found that Leptolinea, belonging to the Anaerolineaceae family, was positively correlated with Methanosaeta (Zhang et al. 2019). Anaerolineaceae and Methanosaeta (acetotrophic methanogen) are syntrophs in the acetotrophic pathway (Liang et al. 2015; McIlroy et al. 2017; Zhao et al. 2016). Syntrophus is a strictly anaerobic bacterium that can degrade fatty acids and benzoate in syntrophic association with hydrogen/formate consumers (Jackson et al. 1999; McInerney et al. 2007). A positive correlation between Syntrophus and Methanolinea was observed previously (Zhang et al. 2019). These two genera were found in higher abundance in the factory compared to the school, reflecting a syntrophic community able to efficiently utilize complex carbon substrates in the factory wastewater treatment. Methanomethylovorans, another genus with higher abundance in the factory, is a mesophilic, methylotrophic, methanogenic archaea, able to use methanol, mono-, di- and trimethylamine, dimethylsulfide, and methanethiol as catabolic and methanogenic substrates (Cha et al. 2013). Particularly, fiberglass production involves a binder that could produce odor due to the presence of trimethylamine (Miele 1994). On the other hand, Achromobacter, Acinetobacter, Thermovirga, Bacteroides, and Lactivibrio showed higher abundance in the rural school treatment system. Dahle and Birkeland (2006) found a species of Thermovirga able to use protein substrates, some single amino acids, and few organic acids (Dahle and Birkeland 2006). This type of specialized metabolism is expected in the rural school wastewater treatment, considering that there were food residues derived from the preparation of meals.
Enhanced volcano plot demonstrating differential abundance of genera (p < 0.05) between the two domestic wastewater treatment system sites. Scattered points represent genera: the x-axis is the log2 fold change for the genera abundance in the factory (left) vs. rural school (right), whereas the y-axis is the − Log10P (where P is the probability that a genus has statistical significance in its differential abundance). Red dots are thus genera significantly over-abundant in each wastewater treatment system, and green dots are genera without significant abundance between both systems
Functional prediction using Picrust
The average values calculated from all samples of the first level in the metabolisms of KEGG orthology were 09100 Metabolism (38.12%), 09120 Genetic Information Processing (24.71%), 09130 Environmental Information Processing (14.78%), Cellular Processes (5.75%) and Unclassified (14.45%). Inside the 09100 Metabolism category, the carbohydrate metabolism (level 2) accounted for ~ 7.0% of all sequences. Pathways such as pyruvate, glycolysis, and gluconeogenesis metabolism belong to it (Fig. 4—section VII). Genes like aceE, pdhA, pdhB, and aceF are present in the acetogenesis pathways from pyruvate. This pathway was more abundant in the dataset from the rural school treatment system. Aldehyde dehydrogenase, yiaY, frmA, adh_1 genes are involved in the synthesis of ethanol from acetate. The alcohol dehydrogenase codified by yiaY gene was more abundant in the rural school than in the factory (Fig. 4).
Functional inference from phylogenetic data. Genes from specific metabolism (i) methanogenesis from methylated compounds; (ii) methanogenesis from methanol; iii) hydrogenotrophic methanogenesis; (iv) acetotrophic methanogenesis; (v) methanogenesis; (vi) dissimilatory nitrate reduction; (vii) carbon metabolism; (viii) assimilatory sulfate reduction. Numbers represent the percentage of the total dataset
The methanogenesis pathway (level 1 metabolism/level 2 energy metabolism) is divided in methanogenesis from mono-, di-, tri-methylamines (Fig. 4, section I), methanogenesis from methanol (Fig. 4, section II), hydrogenotrophic methanogenesis (Fig. 4, section III), acetotrophic methanogenesis (Fig. 4, section IV) and general methanogenesis, that includes the common genes for all methanogenesis pathways (Fig. 4, section VI). The methanogenesis from methylated compounds was more representative in the rural school treatment system. The abundance of other methane metabolisms was very similar between the two sites. Based on the results, the most prevalent methanogenesis pathway was the acetotrophic one, followed by hydrogenotrophic, methanol, and methylotrophic pathways. These results suggest that these genes play a key role in the generation of methane in anaerobic treatment systems. The prevalence of the acetotrophic pathway of methanogenesis in anaerobic wastewater treatments has also been observed by other authors (Guo et al. 2015; Yu et al. 2005). Nevertheless, the abundance of hydrogenotrophic methanogenic pathways was not consistent with taxonomic results, which indicated that the hydrogenotrophic genus Methanobacterium (~ 2.9–3.2%) was the most abundant methanogenic group in both treatment systems. On the other hand, the acetotrophic Methanosaeta showed ~ 0.7% and 1.7% of relative abundance in the rural school and factory treatment systems, respectively. These discrepancies between taxonomic and functional inference can be explained by the overlapping metabolic functions in many taxa, i.e. Methanosarcinaceae family can produce methane by hydrogenotrophic and acetotrophic pathways.
Genes of assimilatory sulfate reduction (Fig. 4, section VIII) and dissimilatory nitrate reduction (Fig. 4, section VI) were observed in the two treatment systems with similar abundances, but these metabolisms were not prevalent.
Influence of physicochemical parameters on microbiome structure
The treatment system of the fiberglass factory presented the highest organic matter removal (measured as COD) and the more adequate core of phylotypes. Connelly et al. (2019) evaluated two (a conventional and a solar-type) septic tanks which presented COD removal rate of 70–89% and predominance of members of Synergistetes, Firmicutes, Bacteroidetes and Euryarchaeota phyla. The COD removal rates obtained in the fiberglass factory (60–83%) were lower than the values reported by Connelly et al. (2019), who had an improvement of removal rate in the solar septic tank due to the control of temperature. The differences in the microbiome structure were related to the greater abundance of Synergistetes and Euryarchaeota: Synergistetes phylum corresponds to one from the core group present in anaerobic digestion bioreactors (Riviere et al. 2009); Euryarchaeota phylum corresponds to the archaeal members responsible for methanogenesis, the last step of anaerobic digestion. Furthermore, hydrogenotrophic methanogenic pathways were more abundantly inferred in the fiberglass factory in comparison with acetotrophic methanogenic pathways. Connelly et al. (2019) suggested that balanced methanogenic pathways could improve the performance of organic matter removal.
Comparing the similarities of the microbiome structure between the sample from the fiberglass factory and the one reported by Connelly et al. (2019), both presented reads affiliated to Firmicutes and Bacteroidetes. Additionally, a high abundance of Chloroflexi and Proteobacteria phyla were detected by Connelly et al. (2019). Except for the Firmicutes phylum, the other phyla belong to the core group of anaerobic digestion postulated by Riviere et al. (2009). These authors studied samples from anaerobic digesters receiving municipal sewage sludge and mixed by biogas reinjection whereas septic tanks and anaerobic filters do not have mixing, which could influence the phylotype composition. Connelly et al. (2019) correlated a low COD removal rate with the presence of Firmicutes in the effluent. These observations emphasize that Firmicutes phylum does not belong to the bacterial core group of anaerobic digestion, however reinforce the fact that this phylum can be frequently found in septic tanks and anaerobic filters.
The low removal rate of organic matter in the treatment system of school could be related to design, especially the hydraulic short-circuit, which resulted in a microbiome structure with high abundance of Firmicutes members. A high abundance of Firmicutes (~ 37%) was detected in the sample from the anaerobic filter of school collected in Autumn of 2018, which corresponds to the lowest COD removal rate observed (2%). As aforementioned, the presence of Firmicutes phylum in the effluent of septic tanks has been previously related to a low COD removal rate (Connelly et al. 2019). Compared to the factory treatment system, a lower abundance of Chloroflexi members was detected in the school treatment. The low presence of Chloroflexi phylum could be related to the hydraulic short-circuit, since this issue hinders the cellular retention and this phylum comprises filamentous bacteria related to granulation (Bovio et al. 2019).
Conclusion
In summary, by combining 16S rRNA gene high-throughput sequencing and physicochemical analyses, we were able to identify the main microorganisms, predict metabolisms involved in the anaerobic digestion and characterize the operational parameters for two different size two-stage wastewater treatment systems. Some of the phyla that represent the microbial core of anaerobic digesters were found in samples from the fiberglass factory (Bacteroidetes, Chloroflexi and Proteobacteria). Conversely, in the school treatment the predominance of Firmicutes members was related to a low removal of COD, and the low abundance of Chloroflexi members was related to the system design, which seems to impair cellular retention. Characterizing and understanding the microbiome of sewage treatment systems allow to infer the metabolic processes that may take place in the systems. Thus, knowledge on microbiome structure, function and dynamics allows one to propose changes in order to improve the efficiency of pollutant removal.
References
Agency UEP (2002) Onsite wastewater treatment systems manual. Office of Research and Development, Washington, DC
Ahring BK (1995) Methanogenesis in thermophilic biogas reactors. Antonie Van Leeuwenhoek 67:91–102. https://doi.org/10.1007/BF00872197
Allison SD, Martiny JB (2008) Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci USA 105:11512–11519. https://doi.org/10.1073/pnas.0801925105
Álvarez JA, Armstrong E, Gómez M, Soto M (2008) Anaerobic treatment of low-strength municipal wastewater by a two-stage pilot plant under psychrophilic conditions. Bioresour Technol 99:7051–7062. https://doi.org/10.1016/j.biortech.2008.01.013
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/. Accessed 7 Sept 2020
Angelidaki I, Batstone DJ (2011) Anaerobic digestion: process. Solid waste technology and management. Wiley, Hoboken, pp 583–600. https://doi.org/10.1002/9780470666883.ch37
Apprill A, McNally S, Parsons R, Weber L (2015) Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat Microb Ecol 75:129–137. https://doi.org/10.3354/ame01753
Ariesyady HD, Ito T, Okabe S (2007) Functional bacterial and archaeal community structures of major trophic groups in a full-scale anaerobic sludge digester. Water Res 41:1554–1568. https://doi.org/10.1016/j.watres.2006.12.036
APHA AWWA-WPCF (2005) WEF, 2005. Standard methods for the examination of water and wastewater 21:258–259
Azbar N, Speece RE (2001) Two-phase, two-stage, and single-stage anaerobic process comparison. J Environ Eng 127:240–248. https://doi.org/10.1061/(ASCE)0733-9372(2001)127:3(240)
Baldwin SA, Khoshnoodi M, Rezadehbashi M, Taupp M, Hallam S, Mattes A, Sanei H (2015) The microbial community of a passive biochemical reactor treating arsenic, zinc, and sulfate-rich seepage. Front Bioeng Biotechnol 3:27–27. https://doi.org/10.3389/fbioe.2015.00027
Baloch M, Akunna JC, Collier PJ (2007) The performance of a phase separated granular bed bioreactor treating brewery wastewater. Bioresour Technol 98:1849–1855. https://doi.org/10.1016/j.biortech.2006.06.014
Bardou P, Mariette J, Escudié F, Djemiel C, Klopp C (2014) jvenn: an interactive Venn diagram viewer. BMC Bioinform 15:293. https://doi.org/10.1186/1471-2105-15-293
Bialek K, Kumar A, Mahony T, Lens PNL, O’ Flaherty V (2012) Microbial community structure and dynamics in anaerobic fluidized-bed and granular sludge-bed reactors: influence of operational temperature and reactor configuration. Microb Biotechnol 5:738–752. https://doi.org/10.1111/j.1751-7915.2012.00364.x
Bodkhe S (2008) Development of an improved anaerobic filter for municipal wastewater treatment. Bioresour Technol 99:222–226. https://doi.org/10.1016/j.biortech.2006.11.026
Bodkhe S (2009) A modified anaerobic baffled reactor for municipal wastewater treatment. J Environ Manag 90:2488–2493. https://doi.org/10.1016/j.jenvman.2009.01.007
Bokulich NA et al (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6:90. https://doi.org/10.1186/s40168-018-0470-z
Bolaños R, Damianovic MHRZ, Zaiat M, Foresti E (2005) Assessment of the ability of sludge to degrade PCP under anaerobic conditions. Braz J Chem Eng 22:611–617. https://doi.org/10.1590/S0104-66322005000400014
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. https://doi.org/10.1093/bioinformatics/btu170
Bolyen E et al (2018) QIIME 2: reproducible, interactive, scalable, and extensible microbiome data science. PeerJ Preprints. https://doi.org/10.1038/s41587-019-0209-9
Boon N, De Windt W, Verstraete W, Top E (2002) Evaluation of nested PCR-DGGE with group-specific 16S rRNA primers for the analysis of bacterial communities from different wastewater treatment plants. FEMS Microbiol Ecol 39:101–112. https://doi.org/10.1016/S0168-6496(01)00198-2
Bovio P, Cabezas A, Etchebehere C (2019) Preliminary analysis of Chloroflexi populations in full-scale UASB methanogenic reactors. J Appl Microbiol 126:667–683. https://doi.org/10.1111/jam.14115
Bray J, Curtis J (1957) An ordination of the upland forest communities of southern Wisconsin. Ecol Monogr 27:325–349. https://doi.org/10.2307/1942268
Buzzini A, Pires E (2002) Cellulose pulp mill effluent treatment in an upflow anaerobic sludge blanket reactor. Process Biochem 38:707–713. https://doi.org/10.1016/S0032-9592(02)00190-5
Buzzini AP, Sakamoto IK, Varesche M, Pires EC (2006) Evaluation of the microbial diversity in an UASB reactor treating wastewater from an unbleached pulp plant. Process Biochem 41:168–176. https://doi.org/10.1016/j.procbio.2005.06.009
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581. https://doi.org/10.1038/nmeth.3869
Caporaso JG et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods. https://doi.org/10.1038/nmeth.f.303
Chao A (1984) Nonparametric estimation of the number of classes in a population. Scand J Stat. https://doi.org/10.2307/4615964
Cha I-T, Min U-G, Kim S-J, Yim KJ, Roh SW, Rhee S-K (2013) Methanomethylovorans uponensis sp. nov., a methylotrophic methanogen isolated from wetland sediment. Antonie Van Leeuwenhoek 104:1005–1012. https://doi.org/10.1007/s10482-013-0020-4
Chen C-L, Wu J-H, Liu W-T (2008) Identification of important microbial populations in the mesophilic and thermophilic phenol-degrading methanogenic consortia. Water Res 42:1963–1976. https://doi.org/10.1016/j.watres.2007.11.037
Connaughton S, Collins G, O’Flaherty V (2006) Development of microbial community structure and activity in a high-rate anaerobic bioreactor at 18 °C. Water Res 40:1009–1017. https://doi.org/10.1016/j.watres.2005.12.026
Connelly S et al (2019) Solar septic tank: next generation sequencing reveals effluent microbial community composition as a useful index of system performance. Water 11:2660. https://doi.org/10.3390/w11122660
Cullimore D, Viraraghavan T (1994) Microbiological aspects of anaerobic filter treatment of septic tank effluent at low temperatures. Environ Technol 15:165–173. https://doi.org/10.1080/09593339409385416
Dahle H, Birkeland N-K (2006) Thermovirga lienii gen. nov., sp. nov., a novel moderately thermophilic, anaerobic, amino-acid-degrading bacterium isolated from a North Sea oil well. Int J Syst Evol Microbiol 56:1539–1545. https://doi.org/10.1099/ijs.0.63894-0
Delforno TP, Moura AGL, Okada DY, Varesche MBA (2014) Effect of biomass adaptation to the degradation of anionic surfactants in laundry wastewater using EGSB reactors. Bioresour Technol 154:114–121. https://doi.org/10.1016/j.biortech.2013.11.102
Demirel B, Yenigün O (2006) Changes in microbial ecology in an anaerobic reactor. Bioresour Technol 97:1201–1208. https://doi.org/10.1016/j.biortech.2005.05.009
Demirel B, Scherer P (2008) The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Rev Environ Sci Bio/Technol 7:173–190. https://doi.org/10.1007/s11157-008-9131-1
De Vrieze J, Saunders AM, He Y, Fang J, Nielsen PH, Verstraete W, Boon N (2015) Ammonia and temperature determine potential clustering in the anaerobic digestion microbiome. Water Res 75:312–323. https://doi.org/10.1016/j.watres.2015.02.025
Douglas GM et al (2019) PICRUSt2: an improved and extensible approach for metagenome inference. bioRxiv. https://doi.org/10.1101/672295
Drake HL, Küsel K, Matthies C (2006) Acetogenic prokaryotes. The prokaryotes: volume 2: ecophysiology and biochemistry. Springer, Berlin, pp 354–420
Duarte MS, Silva SA, Salvador AF, Cavaleiro AJ, Stams AJM, Alves MM, Pereira MA (2018) Insight into the role of facultative bacteria stimulated by microaeration in continuous bioreactors converting LCFA to methane. Environ Sci Technol 52:6497–6507. https://doi.org/10.1021/acs.est.8b00894
Embree M, Liu JK, Al-Bassam MM, Zengler K (2015) Networks of energetic and metabolic interactions define dynamics in microbial communities. Proc Natl Acad Sci USA 112:15450–15455. https://doi.org/10.1073/pnas.1506034112
Fernández-Rodríguez J, Pérez M, Romero LI (2015) Temperature-phased anaerobic digestion of industrial organic fraction of municipal solid waste: a batch study. Chem Eng J 270:597–604. https://doi.org/10.1016/j.cej.2015.02.060
Fernández N, Díaz EE, Amils R, Sanz JL (2008) Analysis of microbial community during biofilm development in an anaerobic wastewater treatment reactor. Microb Ecol 56:121–132. https://doi.org/10.1007/s00248-007-9330-2
Ferrera I, Sánchez O (2016) Insights into microbial diversity in wastewater treatment systems: how far have we come? Biotechnol Adv 34:790–802. https://doi.org/10.1016/j.biotechadv.2016.04.003
Fredriksson NJ, Hermansson M, Wilén B-M (2012) Diversity and dynamics of Archaea in an activated sludge wastewater treatment plant. BMC Microbiol 12:140. https://doi.org/10.1186/1471-2180-12-140
Fujimoto M, Carey DE, Zitomer DH, McNamara PJ (2019) Syntroph diversity and abundance in anaerobic digestion revealed through a comparative core microbiome approach. Appl Microbiol Biotechnol 103:6353–6367. https://doi.org/10.1007/s00253-019-09862-4
Guo J, Peng Y, Ni B-J, Han X, Fan L, Yuan Z (2015) Dissecting microbial community structure and methane-producing pathways of a full-scale anaerobic reactor digesting activated sludge from wastewater treatment by metagenomic sequencing. Microb Cell Fact 14:33. https://doi.org/10.1186/s12934-015-0218-4
Ince BK, Ince O, Oz NA (2003) Changes in acetoclastic methanogenic activity and microbial composition in an upflow anaerobic filter. Water Air Soil Pollut 144:301–315. https://doi.org/10.1023/A:1022917217474
Jackson BE, Bhupathiraju VK, Tanner RS, Woese CR, McInerney MJ (1999) Syntrophus aciditrophicus sp. nov., a new anaerobic bacterium that degrades fatty acids and benzoate in syntrophic association with hydrogen-using microorganisms. Arch Microbiol 171:107–114. https://doi.org/10.1007/s002030050685
Ju F, Lau F, Zhang T (2017) Linking microbial community, environmental variables, and methanogenesis in anaerobic biogas digesters of chemically enhanced primary treatment sludge. Environ Sci Technol 51:3982–3992. https://doi.org/10.1021/acs.est.6b06344
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44:D457-462. https://doi.org/10.1093/nar/gkv1070
Karakashev D, Batstone DJ, Angelidaki I (2005) Influence of environmental conditions on methanogenic compositions in anaerobic biogas reactors. Appl Environ Microbiol 71:331–338. https://doi.org/10.1128/AEM.71.1.331-338.2005
Karakashev D, Batstone DJ, Trably E, Angelidaki I (2006) Acetate oxidation is the dominant methanogenic pathway from acetate in the absence of Methanosaetaceae. Appl Environ Microbiol 72:5138–5141. https://doi.org/10.1128/aem.00489-06
Katoh K, Misawa K, Ki K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf436
Kato S, Watanabe K (2009) Ecological and evolutionary interactions in syntrophic methanogenic consortia. Microbes Environ. https://doi.org/10.1264/jsme2.ME10122
Kindaichi T, Ito T, Okabe S (2004) Ecophysiological interaction between nitrifying bacteria and heterotrophic bacteria in autotrophic nitrifying biofilms as determined by microautoradiography-fluorescence in situ hybridization. Appl Environ Microbiol 70:1641–1650. https://doi.org/10.1128/AEM.70.3.1641-1650.2004
Kortekaas S, Vidal G, Yan-Ling H, Lettinga G, Field JA (1998) Anaerobic-aerobic treatment of toxic pulping black liquor with upfront effluent recirculation. J Ferment Bioeng 86:97–110. https://doi.org/10.1016/S0922-338X(98)80041-X
Koster I, Lettinga G (1984) The influence of ammonium-nitrogen on the specific activity of pelletized methanogenic sludge. Agric Wastes 9:205–216. https://doi.org/10.1016/0141-4607(84)90080-5
Kotsyurbenko OR (2005) Trophic interactions in the methanogenic microbial community of low-temperature terrestrial ecosystems. FEMS Microbiol Ecol 53:3–13. https://doi.org/10.1016/j.femsec.2004.12.009
Kundu K, Bergmann I, Hahnke S, Klocke M, Sharma S, Sreekrishnan TR (2013) Carbon source—a strong determinant of microbial community structure and performance of an anaerobic reactor. J Biotechnol 168:616–624. https://doi.org/10.1016/j.jbiotec.2013.08.023
Liang B, Wang L-Y, Mbadinga SM, Liu J-F, Yang S-Z, Gu J-D, Mu B-Z (2015) Anaerolineaceae and Methanosaeta turned to be the dominant microorganisms in alkanes-dependent methanogenic culture after long-term of incubation. Amb Express 5:37. https://doi.org/10.1186/s13568-015-0117-4
Liu Y, Whitman WB (2008) Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Ann N Y Acad Sci 1125:171–189. https://doi.org/10.1196/annals.1419.019
Liu T, Ahn H, Sun W, McGuinness LR, Kerkhof LJ, Häggblom MM (2016a) Identification of a Ruminococcaceae species as the methyl tert-butyl ether (MTBE) degrading bacterium in a methanogenic consortium. Environ Sci Technol 50:1455–1464. https://doi.org/10.1021/acs.est.5b04731
Liu T, Liu S, Zheng M, Chen Q, Ni J (2016b) Performance assessment of full-scale wastewater treatment plants based on seasonal variability of microbial communities via high-throughput sequencing. PLoS ONE 11:e0152998. https://doi.org/10.1371/journal.pone.0152998
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R (2011) UniFrac: an effective distance metric for microbial community comparison. ISME J 5:169. https://doi.org/10.1038/ismej.2010.133
Madigou C, Lê Cao K-A, Bureau C, Mazéas L, Déjean S, Chapleur O (2019) Ecological consequences of abrupt temperature changes in anaerobic digesters. Chem Eng J 361:266–277. https://doi.org/10.1016/j.cej.2018.12.003
Magurran AE (2013) Measuring biological diversity. Wiley, Hoboken
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
McCarty PL (1964) Anaerobic waste treatment fundamentals, part III, toxic materials and their control. Public Works 95:91–94
McHugh S, Carton M, Collins G, O’Flaherty V (2004) Reactor performance and microbial community dynamics during anaerobic biological treatment of wastewaters at 16–37 °C. FEMS Microbiol Ecol 48:369–378. https://doi.org/10.1016/j.femsec.2004.02.012
McIlroy SJ, Kirkegaard RH, Dueholm MS, Fernando E, Karst SM, Albertsen M, Nielsen PH (2017) Culture-independent analyses reveal novel anaerolineaceae as abundant primary fermenters in anaerobic digesters treating waste activated sludge. Front Microbiol 8:1134. https://doi.org/10.3389/fmicb.2017.01134
McInerney MJ, Sieber JR, Gunsalus RP (2009) Syntrophy in anaerobic global carbon cycles. Curr Opin Biotechnol 20:623–632. https://doi.org/10.1016/j.copbio.2009.10.001
McInerney MJ, Stams AJ, Boone DR (2015) Syntrophobacter. In: Whitman WB, Rainey F, Kämpfer P, Trujillo M, Chun J, DeVos P, Hedlund B, Dedysh S (eds) Bergey’s manual of systematics of archaea and bacteria. Wiley, Hoboken. https://doi.org/10.1002/9781118960608.gbm01068
McInerney MJ et al (2007) The genome of Syntrophus aciditrophicus: life at the thermodynamic limit of microbial growth. Proc Natl Acad Sci USA 104:7600–7605. https://doi.org/10.1073/pnas.0610456104
McLellan SL, Huse SM, Mueller-Spitz SR, Andreishcheva EN, Sogin ML (2010) Diversity and population structure of sewage-derived microorganisms in wastewater treatment plant influent. Environ Microbiol 12:378–392. https://doi.org/10.1111/j.1462-2920.2009.02075.x
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. https://doi.org/10.1371/journal.pone.0061217
McMurdie PJ, Holmes S (2014) Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 10:e1003531. https://doi.org/10.1371/journal.pcbi.1003531
Metcalf L, Eddy HP, Tchobanoglous G (2014) Wastewater engineering: treatment, disposal, and reuse, vol 4, 5th edn. McGraw-Hill NewYork, New York
Miele PF (1994) Low odor binder for molded fiberglass. SAE Trans. https://doi.org/10.2307/44581258
Müller B, Sun L, Westerholm M, Schnürer A (2016) Bacterial community composition and fhs profiles of low-and high-ammonia biogas digesters reveal novel syntrophic acetate-oxidising bacteria. Biotechnol Biofuels 9:48. https://doi.org/10.1186/s13068-016-0454-9
Narihiro T, Nobu MK, Kim N-K, Kamagata Y, Liu W-T (2015) The nexus of syntrophy-associated microbiota in anaerobic digestion revealed by long-term enrichment and community survey. Environ Microbiol 17:1707–1720. https://doi.org/10.1111/1462-2920.12616
Nasr FA, Mikhaeil B (2013) Treatment of domestic wastewater using conventional and baffled septic tanks. Environ Technol 34:2337–2343. https://doi.org/10.1080/09593330.2013.767285
Ney U, Schoberth S, Sahm H (1991) Anaerobic degradation of sulphite evaporator condensate in a fixed-bed loop reactor by a defined bacterial consortium. Appl Microbiol Biotechnol 34:818–822. https://doi.org/10.1007/BF00169357
Panswad T, Komolmethee L (1997) Effects of hydraulic shock loads on small on-site sewage treatment unit. Water Sci Technol 35:145–152. https://doi.org/10.1016/S0273-1223(97)00161-3
Parada AE, Needham DM, Fuhrman JA (2016) Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol 18:1403–1414. https://doi.org/10.1111/1462-2920.13023
Perle M, Kimchie S, Shelef G (1995) Some biochemical aspects of the anaerobic degradation of dairy wastewater. Water Res 29:1549–1554. https://doi.org/10.1016/0043-1354(94)00248-6
Petriglieri F, Nierychlo M, McIlroy SJ, Nielsen PH The in situ characterization of the Chloroflexi communities of full-scale anaerobic digesters and the influence of immigration. In: Danish Microbiological Society (DMS) congress 2017, 2018
Pohland FG, Ghosh S (1971) Developments in anaerobic stabilization of organic wastes—the two-phase concept. Environ Lett 1:255–266. https://doi.org/10.1080/00139307109434990
Price MN, Dehal PS, Arkin AP (2010) FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS ONE 5:e9490. https://doi.org/10.1371/journal.pone.0009490
Quast C et al (2012) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
R Core Team (2014). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Raman V, Chakladar N (1972) Upflow filters for septic tank effluents. Journal (Water Pollut Control Fed) 44:1552–1560. https://doi.org/10.2307/25037574
Riviere D et al (2009) Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J 3:700. https://doi.org/10.1038/ismej.2009.2
Roest K, Heilig HG, Smidt H, de Vos WM, Stams AJ, Akkermans AD (2005) Community analysis of a full-scale anaerobic bioreactor treating paper mill wastewater. Syst Appl Microbiol 28:175–185. https://doi.org/10.1016/j.syapm.2004.10.006
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Sánchez O, Garrido L, Forn I, Massana R, Maldonado MI, Mas J (2011) Molecular characterization of activated sludge from a seawater-processing wastewater treatment plant. Microb Biotechnol 4:628–642. https://doi.org/10.1111/j.1751-7915.2011.00256.x
Senthil Kumar P, Suganya S, Varjani SJ (2018) Evaluation of next-generation sequencing technologies for environmental monitoring in wastewater abatement. In: Varjani SJ, Agarwal AK, Gnansounou E, Gurunathan B (eds) Bioremediation: applications for environmental protection and management. Springer Singapore, Singapore, pp 29–52. https://doi.org/10.1007/978-981-10-7485-1_3
Shi X et al (2019) Genomic dynamics of full-scale temperature-phased anaerobic digestion treating waste activated sludge: focusing on temperature differentiation. Waste Manag 87:621–628. https://doi.org/10.1016/j.wasman.2019.02.041
Shu D, He Y, Yue H, Wang Q (2015) Microbial structures and community functions of anaerobic sludge in six full-scale wastewater treatment plants as revealed by 454 high-throughput pyrosequencing. Bioresour Technol 186:163–172. https://doi.org/10.1016/j.biortech.2015.03.072
Sierra-Alvarez R (1990) The role of natural wood constituents on the anaerobic treatability of forest industry wastewaters. Dissertation, Wageningen University & Research
Sierra JDM, Oosterkamp MJ, Wang W, Spanjers H, van Lier JB (2018) Impact of long-term salinity exposure in anaerobic membrane bioreactors treating phenolic wastewater: performance robustness and endured microbial community. Water Res 141:172–184. https://doi.org/10.1016/j.watres.2018.05.006
Silva FJAd, Lima MGS, Mendonça LAR, Gomes MJTL (2013) Septic tank combined with anaerobic filter and conventional UASB: results from full scale plants. Braz J Chem Eng 30:133–140. https://doi.org/10.1590/S0104-66322013000100015
Snaidr J, Amann R, Huber I, Ludwig W, Schleifer K-H (1997) Phylogenetic analysis and in situ identification of bacteria in activated sludge. Appl Environ Microbiol 63:2884–2896
Stets M, Etto R, Galvão C, Ayub R, Cruz L, Steffens M, Barana A (2014) Microbial community and performance of slaughterhouse wastewater treatment filters. Genet Mol Res 13:4444–4455. https://doi.org/10.4238/2014.June.16.3
Torres K, Álvarez-Hornos FJ, San-Valero P, Gabaldón C, Marzal P (2018) Granulation and microbial community dynamics in the chitosan-supplemented anaerobic treatment of wastewater polluted with organic solvents. Water Res 130:376–387. https://doi.org/10.1016/j.watres.2017.12.009
Uyanik S, Sallis P, Anderson G (2002) The effect of polymer addition on granulation in an anaerobic baffled reactor (ABR). Part I: process performance. Water Res 36:933–943. https://doi.org/10.1016/S0043-1354(01)00315-3
Von Sperling M (2007) Basic principles of wastewater treatment. IWA publishing, London
Von Sperling M, de Lemos Chernicharo CA (2005) Biological wastewater treatment in warm climate regions, vol 1. IWA publishing, London
Wagner M, Erhart R, Manz W, Amann R, Lemmer H, Wedi D, Schleifer K (1994) Development of an rRNA-targeted oligonucleotide probe specific for the genus Acinetobacter and its application for in situ monitoring in activated sludge. Appl Environ Microbiol 60:792–800. https://doi.org/10.1128/AEM.60.3.792-800.1994
Wang X, Wen X, Yan H, Ding K, Zhao F, Hu M (2011) Bacterial community dynamics in a functionally stable pilot-scale wastewater treatment plant. Bioresour Technol 102:2352–2357. https://doi.org/10.1016/j.biortech.2010.10.095
Werner JJ et al (2011) Bacterial community structures are unique and resilient in full-scale bioenergy systems. Proc Natl Acad Sci USA 108:4158–4163. https://doi.org/10.1073/pnas.1015676108
Westerholm M, Moestedt J, Schnürer A (2016) Biogas production through syntrophic acetate oxidation and deliberate operating strategies for improved digester performance. Appl Energy 179:124–135. https://doi.org/10.1016/j.apenergy.2016.06.061
Wu S, Austin D, Liu L, Dong R (2011) Performance of integrated household constructed wetland for domestic wastewater treatment in rural areas. Ecol Eng 37:948–954. https://doi.org/10.1016/j.ecoleng.2011.02.002
Wu B, He C, Yuan S, Hu Z, Wang W (2019) Hydrogen enrichment as a bioaugmentation tool to alleviate ammonia inhibition on anaerobic digestion of phenol-containing wastewater. Bioresour Technol 276:97–102. https://doi.org/10.1016/j.biortech.2018.12.099
Yamada T, Sekiguchi Y, Hanada S, Imachi H, Ohashi A, Harada H, Kamagata Y (2006) Anaerolinea thermolimosa sp. nov., Levilinea saccharolytica gen. nov., sp. nov. and Leptolinea tardivitalis gen. nov., sp. nov., novel filamentous anaerobes, and description of the new classes Anaerolineae classis nov. and Caldilineae classis nov. in the bacterial phylum Chloroflexi. Int J Syst Evol Microbiol 56:1331–1340. https://doi.org/10.1099/ijs.0.64169-0
Yu K, Zhang T (2012) Metagenomic and metatranscriptomic analysis of microbial community structure and gene expression of activated sludge. PLoS ONE 7:e38183. https://doi.org/10.1371/journal.pone.0038183
Yu Y, Lee C, Hwang S (2005) Analysis of community structures in anaerobic processes using a quantitative real-time PCR method. Water Sci Technol 52:85–91. https://doi.org/10.2166/wst.2005.0502
Zhang Z, Gao P, Cheng J, Liu G, Zhang X, Feng Y (2018) Enhancing anaerobic digestion and methane production of tetracycline wastewater in EGSB reactor with GAC/NZVI mediator. Water Res 136:54–63. https://doi.org/10.1016/j.watres.2018.02.025
Zhang Q et al (2019) High variations of methanogenic microorganisms drive full-scale anaerobic digestion process. Environ Int 126:543–551. https://doi.org/10.1016/j.envint.2019.03.005
Zhao Z, Zhang Y, Holmes DE, Dang Y, Woodard TL, Nevin KP, Lovley DR (2016) Potential enhancement of direct interspecies electron transfer for syntrophic metabolism of propionate and butyrate with biochar in up-flow anaerobic sludge blanket reactors. Bioresour Technol 209:148–156. https://doi.org/10.1016/j.biortech.2016.03.005
Funding
This study was supported by the Brazilian National Council for Scientific and Technological Development (CNPq—Grant Number 401720/2016-0). The authors thank the Fiberglass factory and Professor Dorivaldo Damm Rural School for providing the samples.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hidalgo, K.J., Saito, T., Silva, R.S. et al. Microbiome taxonomic and functional profiles of two domestic sewage treatment systems. Biodegradation 32, 17–36 (2021). https://doi.org/10.1007/s10532-020-09921-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-020-09921-y