Abstract
Key message
AtMYB31, a R2R3–MYB transcription factor that modulates wax biosynthesis in reproductive tissues, is involved in seed development in Arabidopsis.
Abstract
R2R3–MYB transcription factors play important roles in plant development; yet, the exact role of each of them remains to be resolved. Here we report that the Arabidopsis AtMYB31 is required for wax biosynthesis in epidermis of reproductive tissues, and is involved in seed development. AtMYB31 was ubiquitously expressed in both vegetative and reproductive tissues with higher expression levels in siliques and seeds, while AtMYB31 was localized to the nucleus and cytoplasm. Loss of function of AtMYB31 reduced wax accumulation in the epidermis of silique and flower tissues, disrupted seed coat epidermal wall development and mucilage production, altered seed proanthocyanidin and polyester content. AtMYB31 could direct activate expressions of several wax biosynthetic target genes. Altogether, AtMYB31, a R2R3–MYB transcription factor, regulates seed development in Arabidopsis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Arabidopsis seed development, consisting of early embryogenesis (until 7 DAP, days after pollination), maturation (8–16 DAP), and late maturation (17–20 DAP) stages (Baud et al. 2002), requires synchronized development of the embryo, the endosperm and the seed coat (Fornale et al. 2010). Seed development is driven by remarkable metabolic changes in both primary and secondary metabolites (Baud et al. 2002; Fait et al. 2006; Fornale et al. 2010). Primary metabolites include mainly carbohydrates, lipids and proteins, while secondary metabolites cover dominantly flavonoids, such as proanthocyanidins (PAs), and cuticular lipids, such as waxes and cutin. While the overall biosynthetic pathways have been characterized along seed development, regulations and interconnections of them remain largely unknown (Le et al. 2010; Haughn and Western, 2012; North et al. 2014). Transcription factors (TFs) play vital roles in almost every aspect of seed development including the development of the endosperm and the embryo (Le et al. 2010; Haughn and Western, 2012; North et al. 2014) and the development of seed coat (Shi et al. 2018). However, identities of those TFs and corresponding targets remain to be uncovered.
The plant cuticle plays important roles in plant reproduction including flower and seed development (Aharoni et al. 2004; Wang et al. 2020a). Developing Arabidopsis seeds are encircled by several layers of cuticles: 1) two cuticle layers situated in the inner and the outer epidermis of the silique, respectively and 2) four cuticle layers located at the embryo, the endosperm, the fusion zone between the inner and the outer seed coat integuments, and the outer cell layer of the seed coat (Ingram and Nawrath, 2017). 3) one cuticle layer (cutin-like polyester) presented in maternal seed coat (Panikashvili et al. 2009). These cuticle layers are believed to prevent organ fusion among the embryo, the endosperm and other tissues (Yang et al. 2008; Moussu et al. 2017), to protect the mature embryo from desiccation (Yang et al. 2008; Xing et al. 2013; Moussu et al. 2017), and to ensure proper seed development and germination. Albeit the abovementioned knowledge and the functional characterization of genes involved in modification of these cuticular structures, it is hard to distinguish the functionality of maternally derived seed coat cuticle and zygotically derived embryo and/or endosperm cuticle (Ingram and Nawrath, 2017). Moreover, the connection between the polysaccharide-rich mucilage and cuticle produced by the epidermal cells in seed is poorly known, neither are its underlying regulatory mechanisms (Nawrath et al. 2013; Shi et al. 2013).
R2R3–MYBs represent a large family of plant-specific TFs involving in growth, development, and responses to biotic and abiotic stresses (Dubos et al. 2010). Among identified R2R3–MYB TF in Arabidopsis (Stracke et al. 2001), five members in the same clade (AtMYB30, AtMYB31, AtMYB60, AtMYB94 and AtMYB96) are reported to be preferentially expressed in the epidermis of top and/or basal segments of stems (Suh et al. 2005), indicating the involvement of them in cuticle metabolism and related biological processes. Indeed, AtMYB30 modulates HR (hypersensitive response)-associated cell death via its function on the acyl-CoA elongase complex, generating VLCFAs (very long chain fatty acids) or their derivatives as signals (Raffaele et al. 2008). Both AtMYB94 and AtMYB96 are wax inducers in Arabidopsis leaves, directly activating the expression of wax biosynthetic genes (Seo et al. 2011; Lee and Suh, 2015; Lee et al. 2016). In addition, AtMYB96 is also required for wax biosynthesis responding to drought (Seo et al. 2011). Further studies indicated that AtMYB96 regulates seed dormancy and germination, seed lipid mobilization (Lee et al. 2015a; Lee and Seo, 2015), and seed TAG (triacylglycerol) accumulation (Lee et al. 2018). However, AtMYB60, a regulator of stomatal and root growth under abiotic stress (Oh et al. 2011), is not yet identified to be involved in cuticle metabolism, nor is the involvement of AtMYB31. It is reported that ZmMYB31 in maize is a regulator of lignin biosynthesis (Fornale et al. 2006, 2010), while SlMYB31 in tomato is a regulator of wax biosynthesis (Xiong et al. 2020). Here, we report that AtMYB31 specifically affects cuticle formation in reproductive tissues as well as metabolic pathways of polyester, seed coat and PAs in seeds. Our data implied a role of AtMYB31 in reproduction in general and in seed development in particular.
Materials and methods
Plant materials
All Arabidopsis mutant plants in the genetic background of Col-0, including two T-DNA insertion mutants (atmyb31-1and atmyb31-2), one SRDX mutant (atmyb31-3/AtMYB31-SRDX), and a CRISPR–Cas9 edited line (atmyb31–4), together with wild type Col-0, were grown at 20 °C with a 70% relative humidity under a 16/8-h light/dark photoperiod. T-DNA knock out lines were obtained from either ABRC or NASC.
Construction of phylogenetic trees and sequence alignments
The protein sequences were analyzed using ClustalA 2.0 (Larkin et al. 2007). The alignment editing was performed using GeneDoc. The multiple alignment parameters were as follows: gap opening set at 10 (default), gap extension set at 2.0, and the neighbor-joining method was used for calculating the tree. The bootstrapped tree was corrected for multiple substitutions. The scale bar of 0.1 is equal to 10% sequence divergence. The phylogenetic trees were constructed using the TreeView program.
Generation of transgenic plants
For pAtMYB31-GUS construct, the complete AtMYB31 5’ upstream region (2,110 bp; termed pAtMYB31) was amplified from wild-type plant genomic DNA, and subcloned into the pMAX vector containing the GUS coding sequence and the NOS terminator, and then cloned into the pBIN( +) binary vector. For 35S:AtMYB31-SRDX construct, a 990 bp AtMYB31 cDNA fragment without terminal stop codon was amplified from WT flower cDNA with the addition of a 36 bp nucleotide sequence of the SRDX (LDLDLELRLGFA), subcloned into the pFLAP100 vector, and then cloned into the pBIN( +) binary vector. The CRISPR–Cas9 transgenic vector was constructed and transformed as previously described (Xu et al. 2020). Agrobacterium mediated planta transformation was carried out via floral dipping as described (Bent and Clough, 1998). For 35S:AtMYB31:eGFP construct, a 990 bp AtMYB31 cDNA fragment without terminal stop codon was amplified from WT flower cDNA, subcloned into the pHB vector containing the eGFP coding sequence and the NOS terminator, the C-terminal fragment without stop codon of AtMYB31 was attached to eGFP through a short peptide linker (GGGGSGGGGS). For 35S:eGFP:AtMYB31 construct, a 993 bp AtMYB31 cDNA fragment was amplified from WT flower cDNA, subcloned into the m104 vector containing the eGFP coding sequence without terminal stop codon and the NOS terminator, the NH2-terminal fragment of AtMYB31 was attached to eGFP through a short peptide linker (GGGGSGGGGS). Both eGFP constructs were transformed into Agrobacterium strain GV3101 and infiltrated into 4-week-old tobacco (Nicotiana benthamiana) leaves (Sparkes et al. 2006) or transformed into protoplasts isolated from etiolated hypocotyl of Arabidopsis by polyethylene glycol-mediated transformation as described previously (Miao and Jiang, 2007). For transcription activation analysis, promoter sequences of the putative AtMYB31 target genes (about 2 kb upstream of the start codon) were amplified, subcloned into pGreen II 0800-LUC vector, and then transformed to Agrobacterium tumefaciens strain GV3101. All primers used in this section are listed in Table S1.
Toluidine blue (TB) staining
Mature siliques (13–14 DAP) were valve peeled along valve margin, put into a 2 ml centrifuge tube containing an aqueous solution of 0.05% (w/v) TB, which had been filtered through a fiber media filter. Plant tissues were submerged and the centrifuge tube was gently shaken (up and down) for 2 min. After being washed with water, tissues (silique and seeds) were photographed. Cuticle permeability examination by toluidine blue was performed as previously described (Tanaka et al. 2010).
Histological observations
For Ruthenium red staining, dry seeds were immersed in 0.05% ruthenium red (Sigma-Aldrich) for 20 min under room temperature, washed with double distilled water and observed using binocular. For vanillin staining, the seeds were immersed in 1% (w/v) vanillin/6 N HCl solution for 10 min to more than 1 h, washed with double distilled water and observed using binocular (Debeaujon 2000). For developing embryo and endosperm observations, immature seeds were dissected from developing siliques, cleared with Hoyer’s solution overnight at 4 °C, and photographed with a Leica DM2500 and Nicon DsRi1 digital cameras (Zhang et al. 2013).
GUS staining
GUS activity was determined as described previously (Jefferson et al. 1987).
Electron microscopy
For SEM (Scanning Electron Microscopy), siliques, seeds, and stems were collected, fixed with glutaraldehyde using standard SEM protocol (Weigel and Glazebrook, 2002), dried using CPD (critical point drying), mounted on aluminum stubs and sputter-coated with gold. SEM was performed using an XL30 ESEM FEG microscope (FEI) at 5–10 kV.
RNA extraction and qualitative reverse transcription PCR (qRT-PCR) analysis
Total RNA was extracted from leaf, stem, root, flower, and developing silique from WT and/or atmyb31-1 plants using RNeasy Plant Mini Kit (Qiagen) with an on-column DNase treatment. The subsequent qRT-PCR analysis was performed as described previously (Shi et al. 2011) using gene-specific primers (Table S1). The experiment was performed with three independent biological replicates, each with a pooled samples from 5 to 6 individual plants. UBC21 (ubiquitin-conjugating enzyme 21, AT5G25760) was used as an internal control for normalizing the level of tested genes.
Chemical analysis
Cuticular waxes were exhaustively extracted by dipping intact tissues (leaf, flower, silique and seed) into 20 ml of chloroform for 1 min at room temperature. Tetracosane, heneicosanoic acid, and heptadecan-1-ol were added as internal standards. The extracts were dried under N2 gas to about 100 μl, and derived with 20 μl of bis-N,N-(trimethylsilyl)trifluoroacetamide and 20 μl of pyridine at 70 °C for 2 h. These derivatized samples were then analyzed by GC–FID (gas chromatography–flame ionization detector) and GC–MS (GC–mass spectrometer). The GC program used was identical to a previous report (Aharoni et al. 2004). Each wax components were quantified against the internal standard through manually integrating peak areas.
Leaf and flower tissues that had been used for wax extraction were used for cutin analyses, while liquid-N2 ground and lyophilized seed fine powder was used for seed polyester analysis. Samples were delipidated exhaustively with chloroform:methanol (1:1, v/v) for two weeks (change dilapidation solvent everyday) at room temperature. Delipidated tissues were dried over silica gels at room temperature and trans-esterified in 1 N methanolic HCl for 2 h at 80 °C. After the addition of saturated NaCl to stop the reaction, dotriacontane, heneicosanoic acid, and heptadecan-1-ol were added as internal standards, and the hydrophobic monomers were extracted three times with hexane. The organic phases were dried under a stream of N2 gas, and the remaining samples were derivatized and analyzed with GC–MS and GC–FID analyses for cutin and with GC–QQQ–MS (GC–triple quadrupole–MS) for polyester as did for the wax analysis (Panikashvili et al. 2009). For seed polyester analysis, atmyb31-1 seeds including normal-looking and abnormal seeds were pooled together.
Dual luciferase assay
For dual luciferase assay, promoter plasmids were individually and co-infiltrated with the plasmid of 35S:AtMYB31 in the pBIN Plus to young tobacco leaf. 3 days later, LUC/REN (Luciferase/Renilla) ratio was measured in leaf discs using Modulus Microplate Luminometer (Turner Biosystems, Sunnyvale, CA) as described previously (Shi et al. 2011). Background controls were run with AtMYB31 in pBIN Plus alone, promoter–LUC alone, and pBIN Plus empty vector alone, and the signal of pBIN Plus empty vector was chosen for background control due to its stable induction of luciferase activity.
Statistical analysis
Student's t test in SPSS 17.0 was performed for statistical analysis for all comparisons. Each treatment contained at least 3 biological repeats, 5–6 individual plants each repeat.
Accession numbers
Genetic information is available in The Arabidopsis Information Resource database (https://www.arabidopsis.org), and accession numbers of all genes discussed in this study are: ACC1, AT1G36160; APLM2, AT1G60780; AtMYB30, AT3G28910; AtMYB31, AT1G74650; AtMYB60, AT1G08810; AtMYB94, AT3G47600; AtMYB96, AT5G62470; CER1, AT1G02205; CER2, AT4G24510; CER10, AT3G55360; COBL2, AT3G29810; CSLA2, AT5G22740; DCR, AT5G23940; FAR3, AT4G33790; FLY1, AT4G28370; FLY2, AT2G20650; GAUT11, AT1G18580; GPAT5, AT3G11430; KCR1, AT1G67730; KCS1, AT1G01120; KCS2, AT1G04220; KCS5, AT1G25450; KCS6, AT1G68530; KCS12, AT2G28630; KNAT7, AT1G62990; LTP3, AT5G59320; MAH1, AT1G57750; MUCI70, AT1G28240; MUM4, AT1G53500; PAS2, AT5G10480; RUBY, AT1G19900; UBC21, AT5G25760; UUAT1, AT5G04160; WAX2/CER3, AT5G57800; WSD1, AT5G37300.
Results
AtMYB31 is ubiquitously expressed but with specific expression patterns along seed development
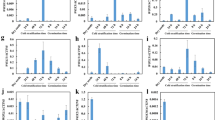
Phylogenetically, AtMYB31 is more closely related to AtMYB94 and AtMYB96 (Fig. S1a, b) in the clade. Although all five genes are reported to be preferably expressed in stem epidermis (Suh et al. 2005), the exact spatial and temporal expression pattern of AtMYB31 has not been reported yet. We first performed in silico expression analysis of AtMYB31 using eFP Brower database (Winter et al. 2007), and found that AtMYB31 is constitutively expressed in both vegetative and reproductive tissues, with the highest expression level in the stem and the pedicel, and relative higher expression level in developing seeds and early developing siliques (Supplementary Fig. S1c). We then carried out qRT-PCR analysis and confirmed the ubiquitous expression pattern of AtMYB31; in which AtMYB31 exhibited the highest expression in the stem, relative higher expression in flowers and in early developing siliques, and low expression in other vegetative tissues, such as roots and leaves (Fig. 1a). To further explore the expression patterns of AtMYB31, we detected the GUS signals in pAtMYB31-GUS transgenic plants, which verified the qRT-PCR result; the strongest GUS signals were found in stems and strong GUS signals were detected also in flowers (Fig. 1a) and in both outside and inside of the early developing siliques (Fig. 1b–-e; Supplementary Fig. S2). Although GUS signals could be detected ubiquitously in both vegetative and reproductive tissues (Supplementary Fig. S2a–2i), they were detected mainly in certain specific sites of reproductive tissues. In flowers, GUS signals were detected in early florescence including pistils (Supplementary Fig. S2e, f), mature sepals (Supplementary Fig. S2g, n), anthers including anther filament and pollens (Supplementary Fig. S2h, n). In developing silique, GUS signals were detected in the abscission zone, the gynophore region, the valve margin (Fig. 1b; Supplementary Fig. S2i, o), the replum or the central ridge (Fig. 1c), the septum (Fig. 1d), the epidermis (Supplementary Fig. S2p), the vascular region inside the silique (Supplementary Fig. S2q), and the top of funiculus and chalaza region (Fig. 1e). In developing seeds, GUS signals were detected in developing embryo (Fig. 1f, g) and seed coat (outer and inner integument) (Fig. 1h, i).
Spatial–temporal expression patterns of AtMYB31. a qRT-PCR analysis of the expression patterns of AtMYB31. R root, L leaf, F flower, Sil1 3 DAP. Silique, Sil2 6 DAP silique, Sil3 8 DAP silique, St stem. Error bars represent standard deviation (n = 3). b–e GUS signals detected in young (b) and mature (c–e). siliques. f, g GUS signals detected in developing embryos (bent and mature stage, respectively). h GUS signal detected in a mature seed (mature stage). i GUS signal detected in the cross section of a developing seed. Scale bars: b 200 μm; c 2 mm; d–i 50 μm
AtMYB31 localizes not only in the nucleus
Because AtMYB31 is supposed to be a transcription factor, we expected its localization to the nucleus. To our surprise, AtMYB31 was found to be localized not only to the nucleus but also to the cytoplasm in the epidermis of transient transformed tobacco leaves (Fig. 2a), irrespective of the fusion of eGFP to the N or C terminal of AtMYB31 (Supplementary Fig. S3). In transient transformed Arabidopsis protoplasts, AtMYB31 was also found to be localized to the nucleus and the cytoplasm (Fig. 2b), which confirmed the localization results of AtMYB31 as observed in transient transformed tobacco leaves (Fig. 2a). In both experiments, AtMYB96 and AtMYB30 were used as positive nucleus-localization controls, and indeed, AtMYB96 and AtMYB30 was found to be localized only to the nucleus (Data not shown).
Loss of function of AtMYB31 produces defective seeds
To further elucidate in planta function of AtMYB31, we characterized and obtained two homozygous mutants atmyb31-1 from SALK_109402 and atmyb31-2 from Sail_168_B10, respectively. In addition, we generated a SRDX mutant atmyb31-3 (AtMYB31–SRDX) and created a CRISPR–Cas9 edited AtMYB31 line atmyb31-4 (25 bp deletion in the third exon) (Supplementary Fig. S4a). Although expression levels of AtMYB31 in flower tissues of these mutants differed dramatically (Supplementary Fig. S4b), all mutants showed, to different extends, abnormal seeds in developing siliques (Figs. 3a–-c; Supplementary Fig. S4c–e), and smaller, darker and shrunken mature seeds (Fig. 3d, e; Supplementary Figs. S4f–4 h), implying the involvement of AtMYB31 in seed development. Based on this similar phenotype, and that the complementation of any of them with AtMYB31 did not work (likely due to maternal effects discovered later in below or redundancy among different clade members), atmyb31-1 mutant was chosen for further analyses.
Phenotypic analysis on seed development in wild type (WT) and atmyb31 plants. a Statistic data of defective seed rates in WT and various atmyb31 mutants. b, c Opened WT (b) and atmyb31-1 (c) silique. Note seed abortion inside the mutant silique. d, e Dry seeds of WT (d) and atmyb31-1 (e). f Opened F1 silique from the cross of WT × atmyb31-1 (upper) and atmyb31 × WT (lower), respectively. g Statistic data of defective seeds in F1 siliques derived from reciprocal cross. Scale bars: b–d 2 mm; f 2 mm
The effect of AtMYB31 on seed development is maternally inherited
Because the complementation of any loss of function atmyb31 mutants with AtMYB31 did not work, we performed reciprocal crosses, and the results demonstrated that the function of AtMYB31 on seed development is inherited in a maternal manner. When WT pistils were pollinated with pollens of atmyb31-1, seeds of F1 generation were as normal as WT seeds. In contrast, when atmyb31-1 pistils were pollinated with WT pollens, seeds of F1 generation exhibited aborted seeds as did in atmyb31-1 mutant; the rate of defective seeds reached up to about 47.66% (Fig. 3f–-g). This effect explained, at least partially, our failures in the complementation experiments.
Loss of function of AtMYB31 partially arrests embryo development
To elucidate the mechanisms underlying the defective seed development in atmyb31-1, we first investigated pollen viability, stigma acceptance, in vivo pollen germination, and pollen tube guidance in WT and atmyb31-1 plants, and found that all these processes in mutants are as normal as those in WT (Supplementary Figs. S5a–S5l). Therefore, we subsequently examined the seed development process using the Hoyer's solution. Embryos in atmyb31-1 developed normally until transition to heart stage (5–6 DAP); at that stage, embryos development were partially arrested (Fig. 4, the lower panel). While seeds bearing these retarded embryos continued to enlarge afterward, even got larger size (the bottom panel in Fig. 4), they finally aborted (Supplementary Fig. S5m–p).
Embryo development process as observed in developing WT (top) and atmyb31-1 (middle and bottom corresponding normal and abnormal seeds, respectively) seeds after being cleared with the Hoyer's solution. DAP day after pollination. Black and red starts point to normal and arrested embryos, respectively. All images were taken under the same magnification. Scale bars: 100 μm
Loss of function of AtMYB31 affects mucilage and flavonoid metabolism
To closely examine the defectiveness of seed development, we used scanning electronic microscopy (SEM) to observe the morphological changes in the seed epidermis of atmyb31-1, and revealed remarkable impairments in the mucilage-producing seed epidermal cells of mutants. The columella, a volcano-shaped secondary cell wall, did not form properly in either developing (Fig. 5a, b) or mature seeds (Fig. 5c, d), neither did outer-tangential cell walls in developing (Fig. 5a, b) or dry mature seeds (Fig. 5c, d). These results indicated that loss of function of AtMYB31 alters seed surface.
Phenotypic analysis in seeds of wild type (WT) and atmyb31 plants. a–d SEM images of developing (a and b) and dry (c and d) seed surfaces of WT (a and c) and atmyb31-1 (b and d). Black arrows point to columella, while white arrows to outer-tangential cell wall. e–j WT (e and f) and atmyb31-1 (g and h, normal; i and j, abnormal) seeds after 1 h ruthenium red staining, numbers after x indicate the magnification numbers used for microscopic observation. k–n WT (k and m) and atmyb31-1 (l and n) seeds without (k and l) or with (m and n) 1 h vanillin staining, all photos were taken under the same magnification. Black arrows point to aborted seeds. Scale bars: a–d 50 μm
When mature WT seeds were imbibed in ruthenium red, a pectin staining dye, a gel-like capsule surrounding the seeds could be observed (Fig. 5e, f); however, upon staining with ruthenium red solution, normal-looking seeds of atmyb31-1 displayed a 10–15% thinner mucilage as compared with that of WT seeds (Fig. 5g, h), while abnormal seeds of atmyb31-1 showed much reduced or almost absent of mucilage (Fig. 5i, j). This result implied that mutation of AtMYB31 affects mucilage/biosynthesis or extrusion.
In contrast, when stained with vanillin, a PA specific dye, defective seeds of atmyb31-1 quickly turned to red color (Fig. 5k–n), indicating more accumulation of PAs, a specific flavonoid, in defective mutant seeds. This result also indicated that atmyb31-1 seeds are more permeable to vanillin, exhibiting a reduced capacity to protect the oxidation of PA.
Loss of function of AtMYB31 increases seed polyester profiles
Because of the alteration of surface permeability in atmyb31-1 seeds, which is supposed to be associated with changes in seed polyesters (Molina et al. 2008; Panikashvili et al. 2009), we employed GC–QQQ–MS to compare the changes of seed polyester between WT and atmyb31-1 mutant. Surprisingly, GC–QQQ–MS analysis showed that levels of total polyester in atmyb31-1 seeds were about twofold of that of wild type, which was contributed by increases of almost all detected monomers of seed polyesters (Fig. 6a). Previous studies using promoter–reporter gene fusions demonstrated the existence of cutin and suberin in the inner and outer integument, respectively, and that lipid polyesters, but not waxes, influence seed coat permeability (Molina et al. 2008); therefore, the altered polyester profile in atmyb31-1 could lead to defective seed coat permeability. Toluidine blue staining of developing seeds confirmed the defective seed coat permeability in atmyb31-1 seeds, in which atmyb31-1 seeds and funiculi were dottily and completely stained, respectively, while neither wild type seeds nor wild type funiculi were stained, even at the tip of the funiculus, where the seed was accidently removed (Supplementary Fig. S6).
Chemical analysis of wild type (WT) and atmyb31-1 plants. a Seed polyester profile and total seed polyester (inserted) of mature seeds. DW dry weight. b Silique wax profile and total silique wax (inserted). FA fatty acids, ALK alkane, 1-OL primary alcohol, KET ketone, DIOL dihydroxyl alcohol, ALD aldehyde, 2 HFA 2-hydroxylated FA, DHA di-hydroxylated FA, DFA dicarboxylic FA, ωHFA ω-hydroxylated FA. * and **, significant at 5% and 1% level from t-student test, respectively. Values are presented by mean ± SD (n = 4)
Loss of function of AtMYB31 reduces cuticular waxes in reproductive tissues
To ascertain the function of AtMYB31 in cuticle metabolism, we performed SEM analysis on silique and stem waxes, and GC–FID/GC–MS analyses on waxes and cutin monomers in different vegetative and reproductive tissues of WT and atmyb31-1 plants. SEM observations showed that, as compared with WT, total wax on the epidermis of mutant silique were mildly reduced (Supplementary Figs. S7a–7b), which was verified by chemical measurements, in which total wax in the outer epidermis of atmyb31-1 siliques was reduced by about 20.54%, and the reduction in C29 alkane and some primary alcohols contribute mainly to such a reduction (Fig. 6b). A similar reduction (about 21.75%) in total wax was also found in atmyb31-1 flowers (Supplementary Fig. S8a). Notably, the content of cutin monomers in atmyb31-1 flowers was not significantly changed (Supplementary Fig. S7b). In addition, neither total wax nor total cutin amounts were significantly altered in leaves of atmyb31-1 plants (Supplementary Fig. S8c, d). Furthermore, SEM observation results did not show remarkable reduction of wax on atmyb31-1 stem epidermis (Supplementary Figs. S7c–7f). Altogether, above results implied that AtMYB31 is a positive cuticle regulator on epidermis of reproductive tissues, such as flower, silique, and seed.
Characterization of the putative AtMYB31 target genes for wax biosynthesis in reproductive tissue epidermis
Because of the reduction of wax accumulations in both silique and flower tissues in atmyb31-1 plants, qRT-PCR was employed to explore putative target genes of AtMYB31, in which we tested most of reported wax genes (Kunst and Samuels, 2009) in flowers (easier for RNA extraction than siliques). Loss of function of AtMYB31 remarkably down-regulated expression levels of ACC1 (acetyl-CoA carboxylase 1) (Lu et al. 2011), CER1 (eceriferum1), CER10, KCS5 (ketoacyl-CoA synthase 5), KCS12, MAH1 (mid-chain alkane hydroxylase 1) (Greer et al. 2007), PAS2 (Pasticcino 2) (Bach et al., 2008) and WAX2 (Chen et al. 2003; Rowland et al. 2007) (Fig. 7a). We subsequently measured the activation of promoters of AtMYB31 putative target genes by AtMYB31 using a dual luciferase assay system (Shi et al. 2011). Among seven putative targets examined, promoters of four out of them, including ACC1, KCS5, CER3/WAX2, and CER1, were significantly activated by AtMYB31 (Fig. 7b), all four genes are known to be involved in wax biosynthesis. These results indicated that AtMYB31 regulates wax biosynthesis in reproductive tissues.
Identification of target genes of AtMYB31. a qRT-PCR analysis. Expression analysis on wax biosynthetic genes in WT and atmyb31-1 flowers. * and **, significant at 5% and 1% level from t-student test, respectively. Values are presented by mean ± SD (n = 3). Relative expression is calculated comparing with control gene UBC (AT5G25760). b Transient expression assays of AtMYB31 transcription factor putative target gene promoter regions. Vectors containing those promoters (about 2 kb upstream of ATG) were infiltrated alone and co-infiltrated with vectors containing AtMYB31 transcription factor fused to the 35S promoter. 35S:AtMYB31 and pBin ( +) were controls. LUC/REN (firefly luciferase/renilla luciferase) values are presented by mean ± SD (n = 4). * and **, significant at 5% and 1% level from t test (compared with signals from those infiltrated with 35S:AtMYB31 or individual promoters), respectively
Discussion
AtMYB31 belongs to a small clade consisting of only five members. Previous studies have reported that AtMYB30, AtMYB94 and AtMYB96 in the same clade are wax biosynthesis regulators of vegetative tissue (leaf) under both normal and stressful conditions (Raffaele et al. 2008; Seo et al. 2011; Lee and Suh, 2015; Lee et al. 2016). AtMYB96 also involves in seed dormancy, germination (Lee and Seo, 2015; Lee et al. 2015c) and seed TAG biosynthesis (Lee et al. 2015b, 2018), while AtMYB94 and AtMYB96 additively inhibit callus formation (Dai et al. 2020). AtMYB30 is a key regulator that links systemic ROS signaling with systemic acquired acclimation (Fichman et al. 2020) and increased levels of AtMYB30 in the phloem accelerates flowering (Liu et al. 2014). Abovementioned results indicate that members of this clade participate in both plant development and stress response. Nevertheless, the function of AtMYB31 remains unknown. Despite studies in other plants revealed possible function of MYB31 in primary and secondary metabolisms via overexpression analysis, such as overexpression of ZmMYB31 in Arabidopsis (Fornale et al. 2010) and in sugarcane (Poovaiah et al. 2016), in planta functional characterizations of MYB31 with loss-of-function mutants are still missing. Our results in this study indicated that AtMYB31 is a wax biosynthesis regulator in reproductive tissues, such as flower, silique and seed, and that AtMYB31 is involved in seed development in Arabidopsis.
AtMYB31 is closely associated with wax biosynthesis in reproductive tissues
AtMYB31 likely regulates wax accumulation in reproductive tissues, functioning in plant reproduction. First, loss of function of AtMYB31 did not reduce wax accumulation in vegetative tissues including leaves (Supplementary Fig. S8c) and stems (Supplementary Figs. S7c–7f) but it did reduce wax accumulation in reproductive tissues including flowers (Supplementary Fig. S8a) and siliques (Fig. 5b; Supplementary Fig. 7a, b). Second, loss of function of AtMYB31 did not alter the cutin profiles in both leaf and flower tissues (Supplementary Fig. S8b, d). Due to the difficulties in the calculation of the surface area of the inner and outer epidermis of the un-uniformed siliques, we did not perform cutin measurement in siliques. Nevertheless, AtMYB1’s roles in wax biosynthesis in reproductive tissues corresponded well with its relative higher expression level in reproductive tissues (Fig. 1; Supplementary Figs. S1c and S2g–2q), particularly in both the outer and the inner silique epidermis, the embryo and the endosperm epidermises, the fusion zone between the inner and the outer integument, and the outer cell layer of the seed coat (Fig. 1; Supplementary Fig. S2). qRT-PCR performed with flowers confirmed the regulatory role of AtMYB31 in wax biosynthesis, because expression levels of four genes with known functions in wax biosynthesis in vegetative tissues were significantly reduced in the flower tissues of atmyb31 (Fig. 7a). Among them, ACC1 is necessary for the elongation of VLCFAs (Baud et al. 2010), while both CER1 and CER3/WAX2 are required for alkane biosynthesis; both are important components of waxes (Aarts et al. 1995; Chen et al. 2003; Rowland et al. 2007). KCS5 encodes an endoplasmic reticulum-associated fatty acid elongase that catalyzes the elongation of VLCFAs in Saccharomyces cerevisae, thus, is involved in wax biosynthesis (Tresch et al., 2012), although its function in planta remains uncharacterized, the presence of the Skn-1 motif in KCS5 promoter might indicate its involvement in embryonic cuticle during seed development (Singh et al., 2020). Notably, all these four genes showed similar expression patterns to that of AtMYB31, particularly in reproductive tissues. ACC1 is highly expressed in siliques (Lu et al. 2011), CER1 in flowers, stems and siliques (Aarts et al. 1995), and CER3/WAX2 in siliques, flowers and stems (Chen et al. 2003). KCS5 is greatly expressed in flowers, young developing siliques and early developing seeds (Winter et al. 2007).
Dual luciferase assay (Fig. 7b), together with data from the in silico motif analysis using PlantCARE (Lescot et al. 2002) (Supplementary Fig. S9), further implied that AtMYB31 regulates wax biosynthesis through either direct binding to the promoters of KCS5 and CER1 or indirect acting on ACC1 and CER3/WAX2. Compared with reported target genes of AtMYB94 (WSD1-wax ester synthase acyl coenzyme A: diacylglycerol acyltransferase 1, KCS2, CER1, CER2, FAR3-alcohol-forming fatty acyl CoA reductase 3 and CER10) (Lee and Suh, 2015) and AtMYB96 (KCS1, KCS2, KCS6, KCR1-beta ketoacyl reductase 1, CER3, WSD1 and LTP3) (Seo et al. 2011; Guo et al. 2013) in vegetative tissues, AtMYB31 seemed to regulate a unique set of wax biosynthetic genes in reproductive tissues. Therefore, AtMYB31 could function differently from other clade members. Nevertheless, more lines of evidence from additional studies are needed to conclude AtMYB31’s function in the wax biosynthesis in reproductive tissues.
AtMYB31 participates in seed development
AtMYB31 regulates seed cuticle formation, thus, participating in seed development. Decreased wax accumulation on the outer epidermis of atmyb31-1 siliques (Fig. 6b; Supplementary Figs. S7a–7b) provided the first line of evidence for AtMYB31’s role in the biosynthesis of the first lipidic protective layer for developing seeds. GUS signals detected in developing embryo and endosperm epidermis (Figs. 1f–-h; Supplementary Fig. S2q) provided the second line of evidence of the involvement of AtMYB31 in cuticle formation during seed development. Result of toluidine blue staining provided the third evidence of the involvement of AtMYB31 in cuticle formation during seed development, in which mutant seed and funiculus were more permeable (Supplementary Fig. S6). This changed seed permeability could be attributed to the disrupted polyester metabolism, which is supposed to affect seed coat permeability (Molina et al. 2008).
AtMYB31 modulates the expression of wax-biosynthetic genes, thus, affecting seed development. Among them, ACC1 is necessary for embryo development (Baud et al. 2010); its weak allele mutant produces a few seed-bearing siliques (Lu et al. 2011), while its strong alleles are lethal (Baud et al. 2010). CER1 mutant shows reduced wax deposition on silique surface and exhibits conditional sterile (Aarts et al. 1995). CER3/WAX2 mutant displays small and nearly seedless siliques (Chen et al. 2003; Rowland et al. 2007). Although there is no functional report of KCS5 in seed development, its high expression in siliques and seeds (Winter et al. 2007) implied that KCS5 is likely involved in seed development as well.
AtMYB31 also regulates seed coat development
Our results showed clearly that AtMYB31 regulates seed coat development. First, the effect of AtMYB31 on seed development is maternal inherited, a typical feature of most seed coat genes (Mizzotti et al. 2014). The observed misshaped epidermal cell and columnar structure in developing mutant seeds (Fig. 5b), deformed seed epidermal cell and impaired seed coat production and secretion (Fig. 5e–j), and the fact that AtMYB31 is highly expressed in developing silique at the globular stage (3–4 DAP) (Fig. 1a) that earlier than the initiation of mucilage biosynthesis and accumulation at the linear stage (7 DAP) (Western and Haughn, 2000; Le et al. 2010), strongly supported the involvement of AtMYB31 in mucilage production. Different from those mutants of mucilage biosynthesis, secretion or regulation, such as muci70 (mucilage related 70), gaut11 (galacturonosyltransferase 11) (Voiniciuc et al. 2018), csla2 (cellulose synthase-like 2) (Yu et al. 2014), uuat1 (UDP-uronic acid transporter 1) (Saez-Aguayo et al. 2017), ap1m2 (adaptor protein-1 mu-adaptin 2) (Shimada et al. 2018), ruby (ruby particles in mucilage) (Sola et al. 2019), fly1fly2 (flying saucer1flying saucer2) (Kunieda et al. 2020), mum4 (mucilage-modified 4) (Western et al. 2004), knat7 (knotted-like homeobox 7) (Wang et al. 2020b), myb52 (Shi et al. 2018) and cobl2 (cobra-like protein 2) (Ben-Tov et al. 2018), atmyb31 showed defectiveness in embryo development (Fig. 4). This result implied AtMYB31’s unique function in seed coat development. Second, AtMYB31 regulates synthesis of PAs that synthesized in the inner most layer of seed coat cells (inner integument 1, II1) with characteristics of maternal inheritance (Wang et al. 2014), thus affecting seed coat development. The vanillin staining and toluidine staining in aborted atmyb31-1 seeds indicated the accumulation of PAs in mutants (Fig. 5k–n) and reduction of cuticular lipids in mutant seeds (Supplementary Fig. S6), respectively, reflecting opposite changes of PAs and cuticular lipids. This result was consistent with previous studies that the content of PAs in seed coat is negatively correlated with the level of fatty acids in embryo (Arsovski et al. 2010; Wang et al. 2014; Xuan et al. 2018). Third, AtMYB31 controls the biosynthesis of seed polyester that is supposed to exist in the inner integument of the seed coat (Molina et al. 2008; Panikashvili et al. 2009), therefore, affecting seed coat development. Reduced polyester profiles in seeds of mutants, such as in gpat5 (glycerol-3-phosphate 2-O-acyltransferase 5) (Beisson et al. 2007) and dcr (defective in cuticular ridges) (Panikashvili et al. 2009), lead to a defective seed epidermis. Different from those two mutants, polyesters in atmyb31-1 seeds increased as compared with those in wild type seeds. These results indicated that the integrity of the cuticle layer from seed polyester is essential for seed development, which merits further investigations.
In sum, AtMYB31 plays an important role in maternally controlled mucilage production, PAs biosynthesis and seed cuticle formation, which is important for reproductive development in general and seed development in particular. Future studies should focus on the understanding of regulatory mechanisms of AtMYB31 on the allocation of carbon source for the biosynthesis of cuticle, mucilage, columella cell wall, and PA, and for seed development.
Author contribution statement JS designed and supervised the research; LS, YC, and GS conducted the experiments; LS, YC, JH, LS, HC and JS analyzed the data; LS, YC, and JH drafted the manuscript, DZ participated the discussion, HC, AA and JS revised the manuscript. All authors read the final version of the manuscript.
Data availability statement
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Aarts MGM, Keijzer CJ, Stiekema WJ, Pereira A (1995) Molecular characterization of the CER1 gene of Arabidopsis involved in epicuticular wax biosynthesis and pollen fertility. Plant Cell 7:2115–2127. https://doi.org/10.2307/3870155
Aharoni A, Dixit S, Jetter R, Thoenes E, Van Arkel G, Pereira A (2004) The SHINE clade of AP2 domain transcription factors activates wax biosynthesis, alters cuticle properties, and confers drought tolerance when overexpressed in Arabidopsis. Plant Cell 16:2463–2480. https://doi.org/10.1105/tpc.104.022897
Arsovski AA, Haughn GW, Western TL (2010) Seed coat mucilage cells of Arabidopsis thaliana as a model for plant cell wall research. Plant Signal Behav 5:796–801. https://doi.org/10.4161/psb.5.7.11773
Bach L, Michaelson LV, Haslam R, Bellec Y, Gissot L, Marion J, Da Costa M, Boutin JP, Miquel M, Tellier F, Domergue F, Markham JE, Beaudoin F, Napier JA, Faure JD (2008) The very-long-chain hydroxy fatty acyl-CoA dehydratase PASTICCINO2 is essential and limiting for plant development. Proc Natl Acad Sci U S A 105(38):14727–14731. https://doi.org/10.1073/pnas.0805089105
Baud S, Boutin JP, Miquel M, Lepiniec L, Rochat C (2002) An integrated overview of seed development in Arabidopsis thaliana ecotype WS. Plant Physiol Biochem 40:151–160. https://doi.org/10.1016/S0981-9428(01)01350-X
Baud S, Guyon V, Kronenberger J, Wuillème S, Rochat C (2010) Multifunctional acetyl-CoA carboxylase 1 is essential for very long chain fatty acid elongation and embryo development in Arabidopsis. Plant J 33:75–86. https://doi.org/10.1046/j.1365-313X.2003.016010.x
Beisson F, Li Y, Bonaventure G, Pollard M, Ohlrogge JB (2007) The acyltransferase GPAT5 is required for the synthesis of suberin in seed coat and root of Arabidopsis. Plant Cell 19:351–368. https://doi.org/10.1105/tpc.106.048033
Bent AF, Clough SJ (1998) Agrobacterium germ-line transformation: transformation of Arabidopsis without tissue culture. In: Gelvin SB, Schilperoort RA (eds) Plant Molecular Biology Manual. Springer, Dordrecht, pp 17–30
Ben-Tov D, Idan-Molakandov A, Hugge A et al (2018) The role of COBRA-LIKE 2 function, as part of the complex network of interacting pathways regulating Arabidopsis seed mucilage polysaccharide matrix organization. Plant J 94:497–512. https://doi.org/10.1111/tpj.13871
Chen X, Goodwin SM, Boroff VL, Liu X, Jenks MA (2003) Cloning and characterization of the WAX2 gene of Arabidopsis involved in cuticle membrane and wax production. Plant Cell 15:1170–1185. https://doi.org/10.1105/tpc.010926
Dai X, Liu N, Wang L et al (2020) MYB94 and MYB96 additively inhibit callus formation via directly repressing LBD29 expression in Arabidopsis thaliana. Plant Sci 293:110323. https://doi.org/10.1016/j.plantsci.2019.110323
Debeaujon I (2000) Influence of the testa on seed dormancy, germination, and longevity in Arabidopsis. Plant Physiol 122:403–414. https://doi.org/10.1104/pp.122.2.403
Dubos C, Stracke R, Grotewold E, Weisshaa B, Martin C, Lepiniec LC (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581. https://doi.org/10.1016/j.tplants.2010.06.005
Fait A, Angelovici R, Less H et al (2006) Arabidopsis seed development and germination is associated with temporally distinct metabolic switches. Plant Physiol 142:839–854. https://doi.org/10.1104/pp.106.086694
Fichman Y, Zandalinas SI, Sengupta S, Burks D, Myers RJ, Azad RK, Mittler R (2020) MYB30 orchestrates systemic reactive oxygen signaling and plant acclimation. Plant Physiol 184:666–675
Fornale S, Sonbol FM, Maes T, Capellades M, Puigdomenech P, Rigau J, Caparros-Ruiz D (2006) Down-regulation of the maize and Arabidopsis thaliana Caffeic acid O-methyltransferase genes by two new maize R2R3-MYB transcription factors. Plant Mol Biol 62:809–823. https://doi.org/10.1007/s11103-006-9058-2
Fornale S, Shi X, Chai C et al (2010) ZmMYB31 directly represses maize lignin genes and redirects the phenylpropanoid metabolic flux. Plant J 64:633–644. https://doi.org/10.1111/j.1365-313X.2010.04363.x
Greer S, Wen M, Bird D, Wu X, Jetter R (2007) The cytochrome P450 enzyme CYP96A15 is the midchain alkane hydroxylase responsible for formation of secondary alcohols and ketones in stem cuticular wax of Arabidopsis. Plant Physiol 145:653–667. https://doi.org/10.1104/pp.107.107300
Guo L, Yang H, Zhang X, Yang S (2013) Lipid transfer protein 3 as a target of MYB96 mediates freezing and drought stress in Arabidopsis. J Exp Bot 64:1755–1767. https://doi.org/10.1093/jxb/ert040
Haughn GW, Western TL (2012) Arabidopsis seed coat mucilage is a specialized cell wall that can be used as a model for genetic analysis of plant cell wall structure and function. Front Plant Sci 3:64. https://doi.org/10.3389/fpls.2012.00064
Ingram G, Nawrath C (2017) The roles of the cuticle in plant development: organ adhesions and beyond. J Exp Bot 68:5307–5321. https://doi.org/10.1093/jxb/erx313
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: betaglucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907. https://doi.org/10.1002/j.1460-2075.1987.tb02730.x
Kunieda T, Hara-Nishimura I, Demura T, Haughn GW (2020) Arabidopsis FLYING SAUCER 2 functions redundantly with FLY1 to establish normal seed coat mucilage. Plant Cell Physiol 61:308–317. https://doi.org/10.1093/pcp/pcz195
Kunst L, Samuels L (2009) Plant cuticles shine: advances in wax biosynthesis and export. Curt Opin Plant Biol 12:721–727. https://doi.org/10.1016/j.pbi.2009.09.009
Larkin MA, Blackshields G, Brown NP, Chenna R, Mcgettigan PA, Mcwilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Le BH, Cheng C, Bui AQ et al (2010) Global analysis of gene activity during Arabidopsis seed development and identification of seed-specific transcription factors. Proc Nat Acad Sci USA 107:8063–8070. https://doi.org/10.1073/pnas.1003530107
Lee K, Seo PJ (2015) Coordination of seed dormancy and germination processes by MYB96. Plant Signal Behav 10:e1056423. https://doi.org/10.1080/15592324.2015.1056423
Lee SB, Suh MC (2015) Cuticular wax biosynthesis is up-regulated by the MYB94 transcription factor in Arabidopsis. Plant Cell Physiol 5:48–60. https://doi.org/10.1093/pcp/pcu142
Lee HG, Suh KimMC Kim Seo HHUPG (2018) The MYB96 transcription factor regulates triacylglycerol accumulation by activating DGAT1 and PDAT1 expression in Arabidopsis seeds. Plant Cell Physiol 59:1432–1442. https://doi.org/10.1093/pcp/pcy073
Lee HG, Lee K, Seo PJ (2015a) The Arabidopsis MYB96 transcription factor plays a role in seed dormancy. Plant Mol Biol 87:371–381. https://doi.org/10.1007/s11103-015-0283-4
Lee HG, Park BY, Kim HU, Seo PJ (2015b) MYB96 stimulates C18 fatty acid elongation in Arabidopsis seeds. Plant Biotechnol Rep 9:161–166. https://doi.org/10.1007/s11816-015-0352-9
Lee K, Lee HG, Yoon S, Kim HU, Seo PJ (2015c) The Arabidopsis MYB96 transcription factor is a positive regulator of ABSCISIC ACID-INSENSITIVE4 in the control of seed germination. Plant Physiol 168:677–689. https://doi.org/10.1104/pp.15.00162
Lee SB, Kim HU, Suh MC (2016) MYB94 and MYB96 additively activate cuticular wax biosynthesis in Arabidopsis. Plant Cell Physiol 57:2300–2311. https://doi.org/10.1093/pcp/pcw147
Lescot M, Déhais P, Moreau Y, De Moor B, Rouzé P, Rombauts S (2002) PlantCARE: a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. https://doi.org/10.1093/nar/30.1.325%3epACC1
Liu L, Zhang J, Adrian J, Gissot L, Coupland G, Yu D, Turck F (2014) Elevated levels of MYB30 in the phloem accelerate flowering in Arabidopsis through the regulation of FLOWERING LOCUS T. PLoS ONE 9:e89799. https://doi.org/10.1371/journal.pone.0089799
Lu S, Zhao H, Parsons E et al (2011) The glossyhead1 allele of ACC1 reveals a principal role for multidomain acetyl-CoA carboxylase in the biosynthesis of cuticular waxes by Arabidopsis. Plant Physiol 157:1079–1092. https://doi.org/10.1104/pp.111.185132
Miao Y, Jiang L (2007) Transient expression of fluorescent fusion proteins in protoplasts of suspension cultured cells. Nat Protoc 2:2348–2353. https://doi.org/10.1038/nprot.2007.360
Mizzotti C, Ezquer I, Paolo D et al (2014) SEEDSTICK is a master regulator of development and metabolism in the Arabidopsis seed coat. PLoS Genet 10:e1004856. https://doi.org/10.1371/journal.pgen.1004856
Molina I, Ohlrogge JB, Pollard M (2008) Deposition and localization of lipid polyester in developing seeds of Brassica napus and Arabidopsis thaliana. Plant J 53:437–449. https://doi.org/10.1111/j.1365-313X.2007.03348.x
Moussu S, Doll NM, Chamot S et al (2017) ZHOUPI and KERBEROS mediate embryo/endosperm separation by promoting the formation of an extracuticular sheath at the embryo surface. Plant Cell 29:1642–1656. https://doi.org/10.1105/tpc.17.00016
Nawrath C, Schreiber L, Franke RB, Geldner N, Kunst L (2013) Apoplastic diffusion barriers in Arabidopsis. Arab Book 11:e0167. https://doi.org/10.1199/tab.0167
North HM, Berger A, Saez-Aguayo S, Ralet MC (2014) Understanding polysaccharide production and properties using seed coat mutants: future perspectives for the exploitation of natural variants. Ann Bot 114:1251–1263. https://doi.org/10.1093/aob/mcu011
Oh J, Noh H, Kwon Y, Hong SW, Kim JH, Le H (2011) A dual role for MYB60 in stomatal regulation and root growth of Arabidopsis thaliana under drought stress. Plant Mol Biol 77:91–103. https://doi.org/10.1007/s11103-011-9796-7
Panikashvili D, Shi JX, Aharoni A (2009) The Arabidopsis DCR encoding a soluble BAHD acyltransferase is required for cutin polyester formation and seed hydration properties. Plant Physiol 151:1773–1789. https://doi.org/10.1104/pp.109.143388
Poovaiah CR, Bewg WP, Lan W, Ralph J, Coleman HD (2016) Sugarcane transgenics expressing MYB transcription factors show improved glucose release. Biotechnol Biofuels 9:143. https://doi.org/10.1186/s13068-016-0559-1
Raffaele S, Vailleau F, Leger A et al (2008) A MYB transcription factor regulates very-long-chain fatty acid biosynthesis for activation of the hypersensitive cell death response in Arabidopsis. Plant Cell 20:752–767. https://doi.org/10.1105/tpc.107.054858
Rowland O, Lee R, Franke R, Schreibe L, Kunst L (2007) The CER3 wax biosynthetic gene from Arabidopsis thaliana is allelic to WAX2/YRE/FLP1. FEBS Let 581:3538–3544. https://doi.org/10.1016/j.febslet.2007.06.065
Saez-Aguayo S, Rautengarten C, Temple H et al (2017) UUAT1 is a golgi-localized UDP-uronic acid transporter that modulates the polysaccharide composition of Arabidopsis seed mucilage. Plant Cell 29:129–143. https://doi.org/10.1105/tpc.16.00465
Seo PJ, Lee SB, Suh MC, Park MJ, Go YS, Park CM (2011) The MYB96 transcription factor regulates cuticular wax biosynthesis under drought conditions in Arabidopsis. Plant Cell 23:1138–1152. https://doi.org/10.4161/psb.6.7.15606
Shi JX, Adato A, Alkan N et al (2013) The tomato SlSHINE3 transcription factor regulates fruit cuticle formation and epidermal patterning. New Phytol 197:468–480. https://doi.org/10.1111/nph.12032
Shi D, Ren A, Tang X et al (2018) MYB52 negatively regulates pectin demethylesterification in seed coat mucilage. Plant Physiol 176:2737–2749. https://doi.org/10.1104/pp.17.01771
Shi JX, Malitsky S, De Oliveira S et al (2011) SHINE transcription factors act redundantly to pattern the archetypal surface of Arabidopsis flower organs. PLoS Genet 7:e1001388
Shimada T, Kunieda T, Sumi S et al (2018) The AP-1 complex is required for proper mucilage formation in Arabidopsis seeds. Plant Cell Physiol 59:2331–2338. https://doi.org/10.1093/pcp/pcy158
Singh S, Geeta R, Das S (2020) Comparative sequence analysis across Brassicaceae, regulatory diversity in KCS5 and KCS6 homologs from Arabidopsis thaliana and Brassica juncea, and intronic fragment as a negative transcriptional regulator. Gene Exp Patterns 38:119146. https://doi.org/10.1016/j.gep.2020.119146
Sola K, Gilchrist EJ, Ropartz D et al (2019) RUBY, a putative galactose oxidase, influences pectin properties and promotes cell-to-cell adhesion in the seed coat epidermis of Arabidopsis. Plant Cell 31:809–831. https://doi.org/10.1105/tpc.18.00954
Sparkes IA, Runions J, Kearns A, Hawes C (2006) Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat Protoc 1:2019–2025. https://doi.org/10.1038/nprot.2006.286
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Cur Opin Plant Biol 4:447–456. https://doi.org/10.1016/S1369-5266(00)00199-0
Suh MC, Samuels AL, Jetter R et al (2005) Cuticular lipid composition, surface structure, and gene expression in Arabidopsis stem epidermis. Plant Physiol 139:1649–1665. https://doi.org/10.1104/pp.105.070805
Tanaka H, Onouchi H, Kondo M, Hara-Nishimura I, Machida Y (2001) A subtilisin-like serine protease is required for epidermal surface formation in Arabidopsis embryos and juvenile plants. Development 128:4681–4689. https://doi.org/10.1007/s429-001-8007-y
Tanaka T, Tanaka H, Machida C, Watanabe M, Machida Y (2010) A new method for rapid visualization of defects in leaf cuticle reveals five intrinsic patterns of surface defects in Arabidopsis. Plant J 37:139–146. https://doi.org/10.1046/j.1365-313X.2003.01946.x
Tresch S, Heilmann M, Christiansen N, Looser R, Grossmann K (2012) Inhibition of saturated very-long-chain fatty acid biosynthesis by mefluidide and perfluidone, selective inhibitors of 3-ketoacyl-CoA synthases. Phytochemistry 76:162–171. https://doi.org/10.1016/j.phytochem.2011.12.023
Voiniciuc C, Engle KA, Gunl M et al (2018) Identification of key enzymes for pectin synthesis in seed mucilage. Plant Physiol 178:1045–1064. https://doi.org/10.1104/pp.18.00584
Wang Z, Chen M, Chen T et al (2014) TRANSPARENT TESTA2 regulates embryonic fatty acid biosynthesis by targeting FUSCA3 during the early developmental stage of Arabidopsis seeds. Plant J 77:757–769. https://doi.org/10.1111/tpj.12426
Wang X, Kong L, Zhi P, Chang C (2020a) Update on cuticular wax biosynthesis and its roles in plant disease resistance. Int J Mol Sci 21:5514. https://doi.org/10.3390/ijms21155514
Wang Y, Xu Y, Pei S et al (2020b) KNAT7 regulates xylan biosynthesis in Arabidopsis seed-coat mucilage. J Exp Bot 71:4125–4139. https://doi.org/10.1093/jxb/eraa189
Weigel D, Glazebrook J (2002) Arabidopsis: A Laboratory Manual. Cold Spring Harbor Laboratory Press, New York, USA. https://doi.org/10.1177/1368431007080699
Western TL, Haughn SGW (2000) Differentiation of mucilage secretory cells of the Arabidopsis seed coat. Plant Physiol 122:345–355. https://doi.org/10.1186/1756-0500-5-156
Western TL, Young DS, Dean GH, Tan WL, Samuels AL, Haughn GW (2004) MUCILAGE-MODIFIED4 encodes a putative pectin biosynthetic enzyme developmentally regulated by APETALA2, TRANSPARENT TESTA GLABRA1, and GLABRA2 in the Arabidopsis seed coat. Plant Physiol 134:296–306. https://doi.org/10.1104/pp.103.035519
Winter D, Vinegar B, Nahal H, Ammar R, Wilson GV, Provart NJ (2007) An “electronic fluorescent pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2:e718. https://doi.org/10.1371/journal.pone.0000718
Xing Q, Creff A, Waters A, Tanaka H, Goodrich J, Ingram GC (2013) ZHOUPI controls embryonic cuticle formation via a signalling pathway involving the subtilisin protease ABNORMAL LEAF-SHAPE1 and the receptor kinases GASSHO1 and GASSHO2. Development 140:770–779. https://doi.org/10.1242/dev.088898
Xiong C, Xie Q, Yang Q, Sun P, Gao S, Li H, Zhang J, Wang T, Ye Z, Yang C (2020) WOOLLY, interacting with MYB transcription factor MYB31, regulates cuticular wax biosynthesis by modulating CER6 expression in tomato. Plant J 103:323–337. https://doi.org/10.1111/tpj.14733
Xu D, Mondol PC, Ishiguro S, Shi J, Zhang D, Liang W (2020) NERD1 is required for primexine formation and plasma membrane undulation during microsporogenesis in Arabidopsis thaliana. aBIOTECH 1:205–218. https://doi.org/10.1007/s42994-020-00022-1
Xuan L, Zhang C, Yan T et al (2018) TRANSPARENT TESTA 4-mediated flavonoids negatively affect embryonic fatty acid biosynthesis in Arabidopsis. Plant Cell Environ 41:2773–2790. https://doi.org/10.1111/pce.13402
Yang S, Johnston N, Talideh E et al (2008) The endosperm-specific ZHOUPI gene of Arabidopsis thaliana regulates endosperm breakdown and embryonic epidermal development. Development 135:3501–3509. https://doi.org/10.1242/dev.026708
Yu L, Shi D, Li J et al (2014) CELLULOSE SYNTHASE-LIKE A2, a glucomannan synthase, is involved in maintaining adherent mucilage structure in Arabidopsis seed. Plant Physiol 164:1842–1856. https://doi.org/10.1104/pp.114.236596
Zhang Y, Liang W, Shi J, Xu J, Zhang D (2013) MYB56 encoding a R2R3 MYB transcription factor regulates seed size in Arabidopsis thaliana. J Integr Plant Biol 55:1166–1178. https://doi.org/10.1111/jipb.12094
Acknowledgements
This study was in part supported by the National Natural Science Foundation of China (Grant No. 31971907, 31671511 and 31461143001), the Programme of Introducing Talents of Discipline to Universities (111 Project, B14016), and SJTU Global Strategic Partnership Fund (2021 SJTU-HUJI).
Funding
National Natural Science Foundation of China, 31971907, Jianxin Shi, 31671511, Jianxin Shi, 31461143001, Jianxin Shi, Project 211,B14016, Dabing Zhang, SJTU Global Strategic Partnership Fund, 2021 SJTU-HUJI, Jianxin Shi.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Communicated by Anastasios Melis.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shi, L., Chen, Y., Hong, J. et al. AtMYB31 is a wax regulator associated with reproductive development in Arabidopsis. Planta 256, 28 (2022). https://doi.org/10.1007/s00425-022-03945-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-022-03945-9