Abstract
Main conclusion
WOX family gene WOX2 is highly expressed during seed development, which functions redundantly with WOX1 and WOX4 to positively regulate seed germination.
Abstract
WOX (WUSCHEL-related homeobox) is a family of transcription factors in plants. They play essential roles in the regulation of plant growth and development, but their function in seed germination is not well understood. In this report, we show that WOX1, WOX2, and WOX4 are close homologues in Arabidopsis. WOX2 has a redundant function with WOX1 and WOX4, respectively, in seed germination. WOX2 is highly expressed during seed development, from the globular embryonic stage to mature dry seeds, and its expression is decreased after germination. Loss of function single mutant wox2, and double mutants wox1 wox2 and wox2 wox4-1 show decreased germination speed. WOX2 and WOX4 are essential for hypocotyl–radicle zone elongation during germination, potentially by promoting the expression of cell wall-related genes. We also found that WOX2 and WOX4 regulate germination through the gibberellin (GA) pathway. These results suggest that WOX2 and WOX4 integrate the GA pathway and downstream cell wall-related genes during germination.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The WUSCHEL (WUS)-related homeobox (WOX) family is a group of plant-specific transcription factors. Its members share a highly conserved DNA-binding domain composed of about 60 amino acids, known as the homeodomain (van der Graaff et al. 2009). In Arabidopsis thaliana, the WOX family contains 15 members (Tvorogova et al. 2021). They can be divided into three distinct clades based on phylogenetic analysis: the ancient clade (WOX10, WOX13, and WOX14), the intermediate clade (WOX8, WOX9, WOX11, and WOX12), and the modern clade (WOX1-7 and WUS) (Wu et al. 2019).
WOXs play important roles in many aspects of developmental processes. The WUS gene of the modern clade has a wide range of functions (Jha et al. 2020). It regulates the maintenance of shoot meristems, flower development, and somatic embryogenesis (Mayer et al. 1998; Zuo et al. 2002; Cao et al. 2015; Sun et al. 2019). The WOX1 and WOX3/PRESSED FLOWER (PRS) act redundantly to control leaf blade development and adaxial–abaxial leaf polarity (Nakata et al. 2012). In Medicago truncatula, when WOX1 ortholog, the STENOFOLIA (STF) gene is loss of function, the mutant displays multiple defects in leaf lamina, leaf vasculature, and flower development (Tadege et al. 2011). WOX2, WOX8, and WOX9 regulate early embryogenesis (Ueda et al. 2011; Palovaara et al. 2016). After fertilization and the first zygotic division, WOX2 expression is restricted to the apical cell, and WOX8 is active exclusively in the basal cell. The differential expression patterns of WOX2 and WOX8 mediate apical-basal embryo polarity formation through localized auxin responses (Breuninger et al. 2008). Furthermore, WOX2 and WOX8 function redundantly to enhance the expression of the CUP-SHAPED COTYLEDON (CUC) genes, influencing the development of the cotyledon boundary (Lie et al. 2012). Previous studies have shown that WOX4 plays versatile roles in plant growth and development (Kucukoglu et al. 2017; Yasui et al. 2018). WOX4 works redundantly with WOX14 downstream of the PHLOEM INTERCALATED WITH XYLEM (PXY) signaling pathway, promoting vascular cell proliferation (Etchells et al. 2013). In addition, WOX4 is directly suppressed by AUXIN RESPONSE FACTOR 5 (ARF5), which participates in auxin-mediated vascular cambium development (Brackmann et al. 2018). Studies in Oryza sativa show that OsWOX4 also has an important role in early leaf development and shoot apical meristem maintenance (Ohmori et al. 2013). WOX5 is a central regulator of stem cell activity in the root system (Sarkar et al. 2007). WOX5, which is specifically expressed in the root quiescent center (QC), restrains the division of QC cells and represses the differentiation of stem cells (Kong et al. 2015). The molecular mechanisms underlying the WOX5 regulating root system are complicated but have been comprehensively studied (Zhang et al. 2015; Zhou et al. 2015; Burkart et al. 2022). WOX5 inhibits the activity of D-type cyclins (CYCD) in QC, establishing quiescence (Forzani et al. 2014). Meanwhile, WOX5 protein transfers from QC into the columella stem cells to inhibit the expression of CYCLING DOF FACTOR 4 (CDF4) and then maintains stem cell state. TOPLESS/TOPLESS-RELATED (TPL/TPR) co-repressors and the HISTONE DEACETYLASE 19 (HDA19) are involved in the inhibition of WOX5 on CDF4 (Pi et al. 2015). The main function of WOX6 is to regulate the development of ovules (Park et al. 2005; Pillitteri et al. 2007). In addition, WOX6 enhances cold tolerance in Arabidopsis (Zhu et al. 2004). WOX7 mediates the inhibition of sugar on lateral root formation. WOX7 binds directly to the promoter of the cell cycle gene CYCLIND6;1 (CYCD6;1) and represses the expression of CYCD6;1 (Kong et al. 2016). WOX9 is essential for stimulating cell division and preventing premature differentiation during embryogenesis and post-embryonic development (Wu et al. 2005, 2007). WOX11/12 participates in callus and adventitious root formation. With the increase of auxin levels in explants, upregulated WOX11/12 promotes the expression of LATERAL ORGAN BOUNDARIES DOMAIN16 (LBD16) and LBD29, resulting in the first-step cell fate transition (Liu et al. 2014). WOX11/12 directly upregulates WOX5/7, which promotes root primordia initiation (Hu and Xu 2016). WOX11/12 has been found to regulate seed dormancy downstream of the phytochrome B (PHYB) signal pathway in Arabidopsis (Liao et al. 2022). WOX13 and WOX14, from the ancient clade, are found to participate in the regulation of the development of flowers, fruits, and vascular tissues (Deveaux et al. 2008; Romera-Branchat et al. 2013; Petzold et al. 2018; Smit et al. 2020). Collectively, the WOXs are involved in a wide variety of aspects of plant development by interacting with their targets, regulators, and partners.
Seed dormancy and germination are important stages in the plant life cycle (Penfield 2017). Dormancy can ensure that plants germinate under suitable environmental conditions and complete the whole life cycle. Germination is the initiation of a seedling establishment, which is a vital feature for the propagation of plant species (Bassel 2016). Therefore, it is of great theoretical and biological significance to reveal the mechanisms of seed dormancy and germination. WOX members are involved in the regulation of plant growth and development, but their roles in dormancy and germination remain largely unknown in Arabidopsis.
In this work, we identified the important functions of WOX1/2/4 in seed germination. Thus, we hypothesize that they act a redundant function in promoting the growth of the hypocotyl–radicle zone through the gibberellin (GA) pathway and the downstream cell wall-related genes. Our results offer more insights into the effects of embryonic growth on seed germination and the functions of the WOX gene family.
Materials and methods
Plant materials and growth conditions
Arabidopsis thaliana ecotype Columbia-0 (Col) was used as a wild-type plant. All of the Arabidopsis T-DNA insertion mutant lines wox1 (SALK_148070), wox2 (SALK_114607), wox2-3 (SALK_050488C), and wox4-1 (SALK_210239) were obtained from the Arabidopsis Biological Resource Center (ABRC). Homozygous mutants were identified using PCR-based screening. Gene-specific primers were designed using the SIGnAL T-DNA primer design tool and were used in combination with T-DNA left border primers (http://signal.salk.edu/tdnaprimers.2.html). The PCR primers for mutant screening are listed in Supplementary Table S1. The double mutants wox1 wox2 and wox2 wox4-1 were generated by crossing. Homozygous lines were confirmed by PCR and utilized for phenotyping. Seeds were imbibed at 4 °C in darkness for 3 days for stratification and then sown into the soil. Plants were grown in a growth chamber at 20 ± 2 °C, 50–60% humidity, and a 16-h light/8-h dark cycle.
Germination assays
The seeds used for all germination comparison assays in this work were harvested in the same batch of plants. For seed dormancy tests, freshly harvested seeds were immediately used for experiments or stored at room temperature for after-ripening treatment. About 50 to 100 seeds harvested from individual plants were plated onto filter paper saturated with distilled water in Petri dishes and incubated in a growth chamber (16-h light/8-h dark, 22 °C). After 7 days, germination was scored as radicle emergence. Six to eight individual plants of each genotype were used in the dormancy phenotyping.
For ABA and paclobutrazol (PAC) sensitivity tests, seeds were stored for 3 months to be fully released from dormancy and sown on filter paper moistened with water containing different concentrations of ABA (0.5, 1.5, and 3 μΜ) or PAC (0.5 and 1 μΜ). The seeds were stratified for 3 days at 4 °C in darkness and then transferred to a growth chamber at 22 °C. The germination was recorded at different time points of incubation. For GA treatment, freshly harvested seeds were imbibed with water containing 2.5, 5, and 10 μΜ GA4+7 and incubated at 22 °C in a growth chamber. Each germination test was done with at least three replicates from eight independent plants. Stock solutions of ABA, PAC, and GA4+7 were dissolved in ethanol. When imbibed with chemicals, the control (0 μΜ) corresponds to 0.01% (v/v) ethanol in water.
Hypocotyl and radicle lengths analysis
The freshly harvested seeds were stored at room temperature for 3 months to break dormancy. Seeds were imbibed in distilled water at 4 °C for 3 days for stratification and then transferred to a growth chamber (16-h light/8-h dark, 22 °C). The seeds germinated at different time points and were photographed by stereo microscope (Nikon 80i). The lengths of hypocotyl and radicle and the transition zone were quantified with ImageJ software (https://imagej.net/ij/index.html).
Cotyledon phenotype analysis
The seeds of wild type (Col) and wox2 wox4-1 were sown on half‐strength Murashige and Skoog (1/2 MS) medium containing 0.8% (m/v) agar and 1% (m/v) sucrose. Subsequently, the seeds were stratified at 4 °C in the dark for 3 days and then incubated in a growth chamber (16-h light/8-h dark, 22 °C) for 6 days. The images of seedlings were taken by a Nikon 80i digital camera (https://www.nikon.com).
Construction of transgenic lines
A 2000 bp (− 2000/0) promoter region of WOX2 (AT5G59340) was fused to the β-glucuronidase (GUS) reporter gene. PCR was performed using DNA extracted from Arabidopsis Col young leaves as a template, and primer pairs proWOX2-F/-R (Supplementary Table S1) were used to amplify the WOX2 promoter fragment. The forward primer contained a HindIII site, and the reverse primer contained a SalI site. The fragment was purified and then inserted into the pBI101 vector between the HindIII and SalI restriction sites. Sequencing confirmed all of the DNA constructs used in this study. WOX2pro::GUS fusion was introduced into Arabidopsis wild-type Col using Agrobacterium-mediated transformation (Clough and Bent 1998). The seeds of transformants were collected, surface sterilized, and then selected based on their ability to survive for 7 days on 1/2 MS medium containing 25 mg L−1 hygromycin. Transgenic lines with a single copy insertion of a T-DNA cassette were confirmed by segregation ratio analysis. The homozygous T3 generation lines were used for subsequent analysis.
RNA extraction and RT-qPCR
Total RNA was extracted from dry seeds or seeds with different times of stratification or germination with Plant Total RNA Purification Kit (GeneMark, Taiwan, China) according to the manufacturer’s instructions and then reverse transcribed using the FastKing RT Kit with gDNA Remover (TIANGEN, Beijing, China). The qPCR was performed on an Eppendorf Mastercycler RealPlex real-time system using Taq Pro Universal SYBR qPCR Master Mix (Vazyme, Nanjing, China). The qPCR primer pairs were designed based on the qPCR Primer Database (https://biodb.swu.edu.cn/qprimerdb/) and are listed in Supplementary Table S1. The gene expression data were analyzed using the 2−△△CT or 2−△CT method using the Arabidopsis ACTIN8 and UBIQUITIN5 as reference genes (Gutierrez et al. 2008). Each analysis had at least three biological replicates.
RNA-seq analysis
The after-ripened seeds of Col and wox2 wox4-1 were cold stratified for 3 days and germinated at 22 °C. Total RNA was extracted from seeds that germinated for 18 h. There were three biological replicates of the experiment. The RNA samples were sequenced using an Illumina HiSeq 2000 by BGI in Shenzhen, China. Total RNA-seq reads were mapped to the Arabidopsis Information Resource (TAIR) 10 genome. Genes with fold change > 1 and false discovery rate (FDR)-adjusted P values < 0.05 were assigned as differentially expressed compared with the wild-type Col. Heatmap was plotted by https://www.bioinformatics.com.cn (last accessed on July 10, 2023), an online platform for data analysis and visualization.
GUS analysis
T3 transgenic homozygous plants containing the WOX2pro::GUS construct were selected for GUS analysis. During embryogenesis, immature embryos were dissected from green siliques 3, 4, 5, and 10 days after pollination (DAP) under a dissecting microscope using a fine forceps and a tungsten tip needle. Silique was collected at 14 DAP. After the seeds were mature, embryos were dissected from freshly harvested dry seeds, and the seeds were stratified for 24 h and 3 days. Cotyledons were from seedlings that grew for 36 h. The shoot apical meristem and the first true leaf were from 5- and 10-day-old seedlings, respectively. Inflorescence, shoot, lateral root, and primary root were collected from 20-day-old plants. The samples at different developmental stages were incubated in the GUS staining solution (50 mM sodium phosphate buffer, pH 7.0, 1 mM EDTA, 0.1% [v/v] Triton X-100, 1 mg mL−1 5-bromo-4-chloro-3-indolyl-β-d-glucuronide, 2 mM ferricyanide, and 2 mM ferrocyanide) at 37 °C in the dark for 12 h. Samples were transferred to 70% (v/v) ethanol and then washed several times to remove chlorophyll. Photographs were taken using a Nikon 80i stereomicroscope.
Statistical analysis
For data analysis, the results are expressed as the means ± standard error (SE). Student’s t-test was performed to see if there were significant differences between the samples using SPSS version 26 (Statistical Product and Service Solutions, IBM). The difference was considered statistically significant when P < 0.05.
Accession numbers
Sequence data from this article can be found in TAIR under the following accession numbers: WOX1 (AT3G18010), WOX2 (AT5G59340), WOX3 (AT2G28610), WOX4 (AT1G46480), WOX5 (AT3G11260), WOX6 (AT2G01500), WOX7 (AT5G05770), WOX8 (AT5G45980), WOX9 (AT2G33880), WOX10 (AT1G20710), WOX11 (AT3G03660), WOX12 (AT5G17810), WOX13 (AT4G35550), WOX14 (AT1G20700), WUSCHEL (AT2G17950), XTH7 (AT4G37800), XTH24 (AT4G30270), XTH33 (AT1G10550), ARF2 (AT5G62000), IAA9 (AT5G65670), KAO2 (AT2G32440), FCLY (AT5G63910), NAC1 (AT1G56010), ACTIN8 (AT1G49240), and UBIQUITIN5 (AT3G62250).
Results
Identification of candidate WOXs genes involved in seed germination
We previously found that two histone deacetylase-binding factors, SWI-INDEPENDENT3 (SIN3)-LIKE1 (SNL1) and SNL2, negatively regulate radicle protrusion during seed germination (Wang et al. 2016). Transcriptome analysis showed that the expression of several WOX family members, such as WOX2/3/5/11/12 and WUS, were identified significantly changed in the imbibed seeds of the snl1 snl2 double mutant (Wang et al. 2016). We therefore speculate that some WOXs may be involved in seed germination regulation. To investigate the expression pattern of WOX genes in seed, we performed the RT-qPCR test on dry seeds, cold stratified seeds, germinating seeds, and seedlings of ecotype Col at seven different time periods.
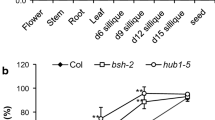
The results showed that four WOX genes (WOX4, WOX6, WOX7, and WOX8) were expressed at relatively low levels at all the tested stages (Fig. 1a–d; Fig. S1a–c); WOX5, WOX9, WOX11, WOX12, and WOX13 transcripts were highly accumulated during stratification and then decreased to a quite low level after germination (Fig. 1e–i; Fig. S1d–h); the expression levels of WOX1, WOX2, WOX10, and WOX14 were high in dry seeds and decreased to a much lower level after stratification and germination (Fig. 1j–m; Fig. S1i–l); WOX3 and WUS were undetectable throughout all the time points when ACTIN8 and UBIQUITIN5 were used as control, while WOX8 was not detected when UBIQUITIN5 as control. The expression pattern of WOX2 in seeds is much like that of SNL1/SNL2. Furthermore, the expression of WOX2 was altered in the germinating seeds of the snl1 snl2 mutant (Wang et al. 2016). These results suggest a possible role for WOX2 in seed germination.
Relative transcript abundance of the WOX family genes in dry, stratified, and germinating seeds and seedlings. a–m The expression levels of WOX4, WOX6, WOX7, WOX8, WOX5, WOX9, WOX11, WOX12, WOX13, WOX1, WOX2, WOX10, and WOX14 in dry, stratified, and germinating seeds, and seedlings. ACTIN8 was used as an internal control for the normalization of gene expression. Values are the means of three biological replicates. The error bars represent SE. Dry seeds, seeds stratified for 24 h, 48 h, and 72 h, and seeds germinated for 8 h, 16 h, and seedlings germinated for 24 h, were collected for total RNA extraction
WOX2 is highly expressed in seed embryos
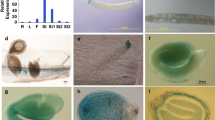
To further monitor WOX2 expression pattern in different tissues, we generated stable transgenic lines in the Col background by introducing a WOX2pro::GUS construct. The histochemical GUS staining showed that WOX2 is highly expressed during embryonic stages, such as the globular stage at 3 DAP under our growth conditions (Fig. 2a), the transitional stage from the globular stage to the heart stage at 4 DAP (Fig. 2b), and the heart stage at 5 DAP (Fig. 2c). GUS signals were subsequently observed in embryos at the mature green stage at 10 DAP (Fig. 2d) and the post-mature green stage at 14 DAP in the silique (Fig. 2e). When the seed became mature and desiccated, GUS expression was detected with strong signals in the whole embryo (Fig. 2f). The GUS signals decreased in embryos from seed stratified for 24 h to that stratified for 3 days (Fig. 2g, h) and showed weaker staining toward germination (Fig. 2i). In addition, GUS staining signals were also observed in other tissues, including the veins of the cotyledons (Fig. 2j), the first true leaves and mature leaves (Fig. 2k, l). By contrast, no GUS signals were detected in inflorescence (Fig. 2l), anthers (Fig. 2m), and stigma (Fig. 2n). GUS signals were further found in the shoot apical meristem (Fig. 2o), shoot (Fig. 2p), lateral root primordia (Fig. 2q), lateral root (Fig. 2r), and primary root (Fig. 2s). The WOX2 promoter activity in embryos indicates that WOX2 may have a function in seed development or germination.
WOX2 promoter-driven GUS expression during seed development, seed germination, and in other various tissues. a Globular stage of embryonic development at 3 days after pollination (DAP). b Transitional stage from globular to heart stage at 4 DAP. c Heart stage at 5 DAP. d Post-embryonic stage at 10 DAP. e Silique at 14 DAP. f Freshly harvested dry seed. g, h Seed cold stratified for 24 h (g) and 3 days (h). i Seedling germinated for 24 h. j Cotyledons. k First true leaf of the 10-day-old seedling. l Leaves and inflorescence of 20-day-old plants. m Anthers. n Stigma. o Shoot apical meristem of 5-day-old seedling. p Shoot of 20-day-old plants. q Lateral root primordium of 20-day-old plants. r Lateral root of 20-day-old plants. s Primary root of 20-day-old plants. Experiments were repeated three times, and representative images were displayed. Bars = 100 μm (a–d, f–j, o–q, s), 200 μm (m, n, r), 1 mm (e, k, l)
WOX2 acts redundantly with WOX1 and WOX4 to positively regulate seed germination
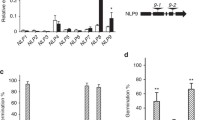
The phylogenetic analysis of the Arabidopsis WOX family showed that WOX2 is closely related to WOX1 and WOX4 (Fig. S2), implying that they have similar functions. To investigate the molecular function of the three members, we ordered T-DNA insertion mutant lines wox1, wox2, wox2-3, and wox4-1 from the ABRC. By genome PCR assay, we found T-DNA is inserted in the third intron of WOX1 as for wox1 mutant, and the second exons of both WOX2 and WOX4 as for the wox2, wox2-3, and wox4-1 mutants, respectively (Fig. 3a). To investigate whether WOX1/2/4 is involved in seed germination regulation, we first examined the dormancy phenotype of these single null mutants. The results showed that freshly harvested wox2 and wox2-3 single mutant seeds showed enhanced seed dormancy (Fig. S3a), whereas the loss function of WOX1 or WOX4 did not affect seed dormancy (Fig. S3b, c). We then analyzed the germination speed of these single mutants using after-ripened (dormancy-released) seeds. As shown in Fig. 3b, wox2 and wox2-3 seeds germinated slightly more slowly than the wild-type Col after cold stratification. In contrast, there was no significant difference in germination rates between the single mutant wox1 and Col (Fig. 3c), wox4-1 and Col (Fig. 3d). Because WOX1, WOX2, and WOX4 are closely related and their transcripts were all detected in seeds (Fig. S2; Fig. 1), we tested their redundancy by constructing two double mutants, wox1 wox2 and wox2 wox4-1. The homozygous double mutants were confirmed by genome PCR before phenotypic analysis. As shown in Fig. S3b, c, both wox1 wox2 and wox2 wox4-1 double mutants showed enhanced seed dormancy compared to the wild type. Furthermore, both double mutants exhibited strong phenotypes of reduced germination speed (Fig. 3c, d). At 20 h following stratification, seeds of the wild type germinated over 80%, whereas wox1 wox2 and wox2 wox4-1 seeds germinated about 30% and 10%, respectively (Fig. 3c, d). Interestingly, we also found that the double mutant wox2 wox4-1 displayed defects in cotyledon initiation (Fig. S4), but wox1 wox2 had no difference from the wild type. The results of germination assays indicate that WOX2 functions redundantly with WOX1 and WOX4 in the positive regulation of seed germination.
Loss of function of WOX2 and WOX4 results in slower germination in Arabidopsis. a Schematic representation of the T-DNA insertion in wox1, wox2, wox2-3, and wox4-1 mutants. The black box represents exons, the horizontal line represents introns, and the inverted triangle represents insertion sites. ATG indicates the start codon, and TAA (or TGA) indicates the stop codon. b Germination phenotypes of wild type (Col), wox2, and wox2-3. c Germination phenotypes of Col, wox1, wox2, and wox1 wox2. d Germination phenotypes of Col, wox2, wox4-1, and wox2 wox4-1. In b to d, seeds were stratified for 3 days at 4 °C before the germination assays. Percentages of seed germination are the means ± SE based on the seeds from six to eight individual plants for each genotype. Asterisks indicate significant differences compared to the wild type as determined by Student’s t-test (**P < 0.01 and *P < 0.05)
The hypocotyl and radicle growth of wox2 wox4-1 was weakened
As GUS activity driven by the WOX2 promoter was observed in hypocotyl and radicle in stratified seeds, germinating seeds, and seedlings (Fig. 2g–i), we hypothesized that the retarded germination of the wox2 wox4-1 double mutant is caused by hypocotyl and radicle development. To verify this hypothesis, we measured the hypocotyl and radicle lengths of the wild-type and wox2 wox4-1 seeds during germination. As shown in Fig. 4a, b, the hypocotyl and radicle of wox2 wox4-1 were smaller than those of the wild type when imbibed for 5 h (before radicle emergence). When seeds were imbibed in water for 5 h, 12 h, and 24 h, the hypocotyl and radicle lengths of the wox2 wox4-1 double mutant were significantly shorter than those of Col, while they showed no difference at 8 h (Fig. 4c). Similarly, the wox2 wox4-1 mutant also showed a significantly shorter hypocotyl–radicle transition zone in seeds imbibed for 0 h, 5 h, and 12 h, and seedlings germinated for 24 h (Fig. 4d). These results supported that the hypocotyl and radicle combinatorial parts are smaller in the wox2 wox4-1 double mutant, implying that WOX2 and WOX4 accelerate seed germination by promoting the hypocotyl elongation and radicle protrusion.
The growth of hypocotyl and radicle of wox2 wox4-1 is weakened. a, b The hypocotyl and radicle morphology of wild type (Col) (a) and wox2 wox4-1 (b) at 5 h after imbibition. Black triangles indicate the length of the hypocotyl–radicle transition zone. Green triangles indicate the hypocotyl and radicle length. Bars = 100 μm. c, d Lengths of hypocotyl and radicle (c) and hypocotyl–radicle transition zone (d) of Col and wox2 wox4-1 at different imbibition time points, respectively. Seeds were stratified for 3 days at 4 °C before the germination assays. Values are the means ± SE based on 12 seeds from eight individual plants for each genotype. The double asterisk and single asterisk indicate significant differences from the wild type using Student’s t-test (**P < 0.01 and *P < 0.05)
Transcriptome analysis of wox2 wox4-1
To further understand the downstream pathways affected by WOX2 and WOX4 mutations during seed germination, we performed RNA-seq. We used wild-type Col and wox2 wox4-1 seeds imbibed at 25 °C for 18 h when the two materials showed a significant difference in germination (Fig. S5). A total of 514 differentially expressed genes (DEGs) were identified (Supplementary Table S2). Among them, 410 DEGs were upregulated, and 104 DEGs were downregulated. Gene Ontology (GO) enrichment analysis showed that these genes were enriched in cell wall biogenesis, and organization, and response to phytohormones (Fig. 5a). By analyzing the DEGs in the highlighted pathways in Fig. 5a, we found that the Xyloglucan endotransglucosylase/hydrolase (XTH) family genes were significantly enriched in processes such as cell wall biogenesis, and xyloglucan metabolic process, and xyloglucosyl transferase activity (Fig. S6). The heatmap showed that the expression of seven XTH genes decreased in wox2 wox4-1 (Fig. 5b). In addition, the expression of some genes involved in phytohormone signaling pathways was also altered (Fig. S6).
Transcriptomic analysis of gene expression profiles in wild-type (Col) and wox2 wox4-1 germinating seeds. a Gene Ontology (GO) enrichment analysis of all differentially expressed genes in wox2 wox4-1 versus wild type (Col). The GO category biological processes are shown. Processes related to the cell wall are highlighted with red boxes. Phytohormone-related pathways are marked with green boxes. The dot size is proportional to the gene number. The color scale indicates the significance level. b The expression heatmap of the XTH family genes analyzed by RNA-seq. The color scale indicates the log2 FPKM value
Furthermore, we examined the DEGs by RT-qPCR using dry seeds, stratified seeds, and germinating seeds. Eight genes involved in the cell wall, phytohormones, and root growth were selected. As shown in Fig. 6a, expression of XTH7 was decreased in the double mutant seeds compared with the wild type when stratified for 72 h and germinated for 18 h. XTH24 transcript level decreased in dry seeds and 24 h stratified seeds, while it showed no differences in seeds germinated for 5 h and 18 h (Fig. 6b). XTH33 was upregulated in both dry and 72-h stratified seeds but downregulated in seeds germinated for 18 h, in the double mutant compared to that of the wild type (Fig. 6c). In addition, the expression levels of some phytohormone-related genes were also affected by WOX2 and WOX4 mutations, including auxin signaling pathway-related genes ARF2 and Indole-3-acetic acid 9 (IAA9) (Fig. 6d, e), GA synthesis gene Ent-kaurenoic acid oxidase 2 (KAO2) (Fig. 6f), and ABA signaling pathway gene Farnesylcysteine lyase (FCLY) (Fig. 6g). We also found that the NAC family transcription factor gene NAC1, involved in root development regulation (Xie et al. 2000), was upregulated in wox2 wox4-1 mutant seeds germinated for 5 h and 18 h (Fig. 6h). These results suggest that WOX2 and WOX4 regulate seed germination through the phytohormone pathway and then the downstream XTH family genes.
Relative expression of XTHs, phytohormones-related genes, and NAC1 in dry seeds, cold stratified seeds, and germinating seeds for wild-type Col and wox2 wox4-1 mutant. a–c RT-qPCR validation of the expression levels of XTH7, XTH24, and XTH33. d–g RT-qPCR analysis of the expression levels of ARF2, IAA9, KAO2, and FCLY. h RT-qPCR analysis of the expression level of root development-related gene NAC1. Total RNAs were isolated from freshly harvested dry seeds, 24-h and 72-h cold stratified seeds, and 5-h and 18-h germinating seeds. Expression levels were normalized to ACTIN8. Values are means of three biological replicates. The error bars represent SE. Asterisks indicate significant differences compared to the wild type as determined by Student’s t-test (**P < 0.01 and *P < 0.05)
Seed germination sensitivity of wox2 wox4-1 to ABA, PAC, and GA4+7
The balance between ABA and GA plays a pivotal role in regulating seed germination (Shu et al. 2016). Our transcriptome data showed that the expression of GA and ABA-related genes was affected (Fig. 5a). We, therefore, hypothesized that the retarded germination of wox2 wox4-1 might be caused by an enhanced ABA pathway or an attenuated GA pathway. We then examined the seed germination in response to ABA, GA4+7, and PAC (an inhibitor of GA biosynthesis). In the presence of PAC, fully after-ripened seeds of wox2 wox4-1 showed enhanced sensitivity to PAC compared with the wild type. The results indicate that the GA pathway is affected in wox2 wox4-1 (Fig. 7a). GA4+7 treatment could partially restore the weakened seed germination phenotype of wox2 wox4-1 (Fig. 7b). However, wox2 wox4-1 seeds showed a similar sensitivity to ABA as the wild type, suggesting that the ABA signaling pathway is not affected in wox2 wox4-1 (Fig. 7c). These results suggest that WOX2 and WOX4 are involved in the GA pathway to control seed germination.
Germination of wild-type and wox2 wox4-1 mutant seeds in the presence of different concentrations of PAC, ABA, and GA4+7. a Germination of wild-type Col and wox2 wox4-1 mutant seeds treated with different concentrations of PAC (0, 0.5, and 1 μΜ). b Germination of Col and wox2 wox4-1 mutant seeds treated with different concentrations of GA4+7 (0, 2.5, 5, and 10 μΜ). c Germination of Col and wox2 wox4-1 mutant seeds treated with different concentrations of ABA (0, 0.5, 1.5, and 3 μΜ). Seeds were cold stratified for 3 days before being treated with PAC or ABA. Freshly harvested seeds were used for GA4+7 treatment. At least 50 seeds for each genotype were used in three independent biological replicates. Data are means (± SE). Significant differences compared with the wild type were determined using Student’s t-test (**P < 0.01 and *P < 0.05)
Discussion
In this study, we showed that the WOX family gene WOX2 is highly expressed during seed development and acts redundantly with WOX1 and WOX4 to positively regulate seed germination. We hypothesize that WOX2 and WOX4 regulate seed germination by stimulating cell elongation in the hypocotyl–radicle region via the GA pathway and downstream cell wall-related genes (Fig. 8).
A model for the roles of WOX1/2/4 in the control of seed germination. Under favorable conditions, GA is synthesized in seeds, which may induce or stabilize WOX1/2/4 proteins. WOXs positively regulate the expression of XTH genes (directly or indirectly), and then promote cell elongation of the hypocotyl–radicle region. Consequently, seed germination is facilitated. Arrows indicate positive effects, and the dashed line indicates indirect or unknown regulation. The graphic was created with BioRender.com
The mature Arabidopsis seed consists of two parts: the embryo and the surrounding tissues (single-cell endosperm layer and testa). When the seed dormancy release is complete and the external environment is suitable for germination, the seed will enter the germination state (Holdsworth et al. 2008). The completion of seed germination depends on the balance of two opposing forces: the growth potential of the embryo and the restraint of the enclosing structures. In Arabidopsis, the hypocotyl–radicle transition zone and lower hypocotyl, as growing tissues, increase the embryo growth potential to exceed the restraint and then complete seed germination (Sliwinska et al. 2009; Steinbrecher and Leubner-Metzger 2017). The molecular genetic networks that regulate seed germination have been gradually unraveled (Rajjou et al. 2012). In this study, we preliminarily investigate the function of WOX genes on seed germination. The double mutant wox2 wox4-1 showed reduced seed germination, and its growth of the hypocotyl–radicle zone was weakened (Figs. 3, 4), which led us to speculate that WOX2 and WOX4 may promote seed germination by increasing the embryo growth potential. The GA signaling pathway plays an important role during seed germination. In Arabidopsis, root apical meristem is an important site for GA production. GA signals moved from the radicle to the hypocotyl, promoting the elongation of the hypocotyl–radicle zone by regulating the expression of downstream genes, including the XTH genes XTH9/19 and the expansin genes EXPA1/8/15 (Becnel et al. 2006; Bassel et al. 2014). The GA biosynthetic mutant ga1-3 could not germinate. After exogenous application of GA, the hypocotyl cells can elongate normally, and the mutant seeds can eventually germinate (Sun et al. 1992; Bassel et al. 2014). Therefore, GA signals are crucial for hypocotyl–radicle zone elongation. In this study, GO enrichment analysis of the DEGs revealed that “response to gibberellin” is one of the significantly enriched GO terms (Fig. 5; Fig. S6). Seed germination sensitivity to PAC and GA4+7 indicated that the GA pathway was affected by WOX2 and WOX4 mutations (Fig. 7a, b). RT-qPCR assay also confirmed that the GA biosynthesis gene KAO2 was downregulated in wox2 wox4-1 (Fig. 6f). These results suggest that WOX2 and WOX4 may regulate seed germination through the GA pathway.
The structure of the hypocotyl in Arabidopsis is relatively simple. Normally, the number of hypocotyl cells in Arabidopsis is determined during embryogenesis (Gendreau et al. 1997). Previous studies have shown that hypocotyl–radicle zone elongation is the result of cell expansion rather than cell division (Sliwinska et al. 2009). The prerequisite for cell expansion is cell wall loosening (Cosgrove 2005). In Arabidopsis, the XTH family contains 33 members (Becnel et al. 2006). XTHs encode cell wall modification enzymes, which can catalyze the hydrolysis or transfer of xyloglucan molecules, thereby loosening the structure of the cell wall (Van Sandt et al. 2007; Maris et al. 2009; Miedes et al. 2013). Research has shown that XTH is involved in multiple processes of plant growth and development, such as root elongation, leaf vein differentiation, fruit ripening, and petal senescence (Matsui et al. 2005; Liu et al. 2007; Singh et al. 2011; Tsuchiya et al. 2015). XTH7 was reported to regulate hypocotyl elongation in Arabidopsis by ethylene and brassinosteroid signaling pathways (Liu et al. 2018). XTH24 works downstream of SHORT-ROOT (SHR) to regulate hypocotyl cell elongation (Dhar et al. 2022). XTH33 is involved in root growth in response to ethylene (Kong et al. 2018). Furthermore, they were also expressed in the hypocotyl, cotyledon, or elongation zone, which highly overlapped with the expression pattern of WOX2 (Becnel et al. 2006; Fig. 2). These data suggest that XTH7, XTH24, and XTH33 are the downstream genes of WOX2 and WOX4. We hypothesize that the two transcription factors may act redundantly in hypocotyl–radicle cells to promote the expression of XTH genes, increasing cell wall extensibility, cell elongation, and, eventually, seed germination. The induction of WOX2 and WOX4 on XTH genes requires further confirmation by a transcriptional activation assay. Whether the cell expansion in the wox2 wox4-1 hypocotyl–radicle zone is retarded still needs further cell morphological analysis.
In conclusion, our research identified a novel function of WOX2 and WOX4 in seed germination. Genetic and transcriptomic analyses provide a framework for how WOX2 and WOX4 regulate seed germination. We suggest that WOX2 and WOX4 increase the growth of the hypocotyl–radicle zone through the downstream XTH genes (XTH7/24/33) during germination, hence increasing germination potential. Our results provide insights into the regulatory mechanism of WOX genes in seed germination.
Data availability
All data in this study are provided in this manuscript and supplementary data files. It will be provided upon a reasonable request.
Abbreviations
- DAP:
-
Days after pollination
- GA:
-
Gibberellin
- GO:
-
Gene Ontology
- GUS:
-
β-Glucuronidase
- PAC:
-
Paclobutrazol
- WOX:
-
WUSCHEL-related homeobox
- XTH:
-
Xyloglucan endotransglucosylase/hydrolase
References
Bassel GW (2016) To grow or not to grow? Trends Plant Sci 21:498–505. https://doi.org/10.1016/j.tplants.2016.02.001
Bassel GW, Stamm P, Mosca G, Barbier de Reuille P, Gibbs DJ, Winter R, Janka A, Holdsworth MJ, Smith RS (2014) Mechanical constraints imposed by 3D cellular geometry and arrangement modulate growth patterns in the Arabidopsis embryo. Proc Natl Acad Sci USA 111:8685–8690. https://doi.org/10.1073/pnas.1404616111
Becnel J, Natarajan M, Kipp A, Braam J (2006) Developmental expression patterns of Arabidopsis XTH genes reported by transgenes and Genevestigator. Plant Mol Biol 61:451–467. https://doi.org/10.1007/s11103-006-0021-z
Brackmann K, Qi J, Gebert M, Jouannet V, Schlamp T, Grunwald K, Wallner ES, Novikova DD, Levitsky VG, Agusti J, Sanchez P, Lohmann JU, Greb T (2018) Spatial specificity of auxin responses coordinates wood formation. Nat Commun 9:875. https://doi.org/10.1038/s41467-018-03256-2
Breuninger H, Rikirsch E, Hermann M, Ueda M, Laux T (2008) Differential expression of WOX genes mediates apical-basal axis formation in the Arabidopsis embryo. Dev Cell 14:867–876. https://doi.org/10.1016/j.devcel.2008.03.008
Burkart RC, Strotmann VI, Kirschner GK, Akinci A, Czempik L, Dolata A, Maizel A, Weidtkamp-Peters S, Stahl Y (2022) PLETHORA-WOX5 interaction and subnuclear localization control Arabidopsis root stem cell maintenance. EMBO Rep 23:e54105. https://doi.org/10.15252/embr.202154105
Cao XW, He ZS, Guo L, Liu XG (2015) Epigenetic mechanisms are critical for the regulation of WUSCHEL expression in floral meristems. Plant Physiol 168:1189–1196. https://doi.org/10.1104/pp.15.00230
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6:850–861. https://doi.org/10.1038/nrm1746
Deveaux Y, Toffano-Nioche C, Claisse G, Thareau V, Morin H, Laufs P, Moreau H, Kreis M, Lecharny A (2008) Genes of the most conserved WOX clade in plants affect root and flower development in Arabidopsis. BMC Evol Biol 8:291. https://doi.org/10.1186/1471-2148-8-291
Dhar S, Kim J, Yoon EK, Jang S, Ko K, Lim J (2022) SHORT-ROOT controls cell elongation in the etiolated Arabidopsis hypocotyl. Mol Cells 45:243–256. https://doi.org/10.14348/molcells.2021.5008
Etchells JP, Provost CM, Mishra L, Turner SR (2013) WOX4 and WOX14 act downstream of the PXY receptor kinase to regulate plant vascular proliferation independently of any role in vascular organisation. Development 140:2224–2234. https://doi.org/10.1242/dev.091314
Forzani C, Aichinger E, Sornay E, Willemsen V, Laux T, Dewitte W, Murray JA (2014) WOX5 suppresses CYCLIN D activity to establish quiescence at the center of the root stem cell niche. Curr Biol 24:1939–1944. https://doi.org/10.1016/j.cub.2014.07.019
Gendreau E, Traas J, Desnos T, Grandjean O, Caboche M, Höfte H (1997) Cellular basis of hypocotyl growth in Arabidopsis thaliana. Plant Physiol 114:295–305. https://doi.org/10.1104/pp.114.1.295
Gutierrez L, Mauriat M, Guénin S, Pelloux J, Lefebvre JF, Louvet R, Rusterucci C, Moritz T, Guerineau F, Bellini C, Van Wuytswinkel O (2008) The lack of a systematic validation of reference genes: a serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR) analysis in plants. Plant Biotechnol J 6:609–618. https://doi.org/10.1111/j.1467-7652.2008.00346.x
Holdsworth MJ, Bentsink L, Soppe WJJ (2008) Molecular networks regulating Arabidopsis seed maturation, after-ripening, dormancy and germination. New Phytol 179:33–54. https://doi.org/10.1111/j.1469-8137.2008.02437.x
Hu X, Xu L (2016) Transcription factors WOX11/12 directly activate WOX5/7 to promote root primordia initiation and organogenesis. Plant Physiol 172:2363–2373. https://doi.org/10.1104/pp.16.01067
Jha P, Ochatt SJ, Kumar V (2020) WUSCHEL: a master regulator in plant growth signaling. Plant Cell Rep 39:431–444. https://doi.org/10.1007/s00299-020-02511-5
Kong X, Lu S, Tian H, Ding Z (2015) WOX5 is shining in the root stem cell niche. Trends Plant Sci 20:601–603. https://doi.org/10.1016/j.tplants.2015.08.009
Kong D, Hao Y, Cui H (2016) The WUSCHEL related homeobox protein WOX7 regulates the sugar response of lateral root development in Arabidopsis thaliana. Mol Plant 9:261–270. https://doi.org/10.1016/j.molp.2015.11.006
Kong X, Li C, Zhang F, Yu Q, Gao S, Zhang M, Tian H, Zhang J, Yuan X, Ding Z (2018) Ethylene promotes cadmium-induced root growth inhibition through EIN3 controlled XTH33 and LSU1 expression in Arabidopsis. Plant Cell Environ 41:2449–2462. https://doi.org/10.1111/pce.13361
Kucukoglu M, Nilsson J, Zheng B, Chaabouni S, Nilsson O (2017) WUSCHEL-RELATED HOMEOBOX4 (WOX4)-like genes regulate cambial cell division activity and secondary growth in Populus trees. New Phytol 215:642–657. https://doi.org/10.1111/nph.14631
Liao JK, Deng B, Cai XY, Yang QX, Hu BP, Cong JJ, Zhang YX, Wang G, Xin GL, Li YT, Yang L, Zhang DZ, Zhang J, Liu BB (2022) Time-course transcriptome analysis reveals regulation of Arabidopsis seed dormancy by the transcription factors WOX11/12. J Exp Bot 74:1090–1106. https://doi.org/10.1093/jxb/erac457
Lie C, Kelsom C, Wu X (2012) WOX2 and STIMPY-LIKE/WOX8 promote cotyledon boundary formation in Arabidopsis. Plant J 72:674–682. https://doi.org/10.1111/j.1365-313X.2012.05113.x
Liu YB, Lu SM, Zhang JF, Liu S, Lu YT (2007) A xyloglucan endotransglucosylase/hydrolase involves in growth of primary root and alters the deposition of cellulose in Arabidopsis. Planta 226:1547–1560. https://doi.org/10.1007/s00425-007-0591-2
Liu J, Sheng L, Xu Y, Li J, Yang Z, Huang H, Xu L (2014) WOX11 and 12 are involved in the first-step cell fate transition during de novo root organogenesis in Arabidopsis. Plant Cell 26:1081–1093. https://doi.org/10.1105/tpc.114.122887
Liu K, Li Y, Chen X, Li L, Liu K, Zhao H, Wang Y, Han S (2018) ERF72 interacts with ARF6 and BZR1 to regulate hypocotyl elongation in Arabidopsis. J Exp Bot 69:3933–3947. https://doi.org/10.1093/jxb/ery220
Maris A, Suslov D, Fry SC, Verbelen JP, Vissenberg K (2009) Enzymic characterization of two recombinant xyloglucan endotransglucosylase/hydrolase (XTH) proteins of Arabidopsis and their effect on root growth and cell wall extension. J Exp Bot 60:3959–3972. https://doi.org/10.1093/jxb/erp229
Matsui A, Yokoyama R, Seki M, Ito T, Shinozaki K, Takahashi T, Komeda Y, Nishitani K (2005) AtXTH27 plays an essential role in cell wall modification during the development of tracheary elements. Plant J 42:525–534. https://doi.org/10.1111/j.1365-313X.2005.02395.x
Mayer KF, Schoof H, Haecker A, Lenhard M, Jurgens G, Laux T (1998) Role of WUSCHEL in regulating stem cell fate in the Arabidopsis shoot meristem. Cell 95:805–815. https://doi.org/10.1016/S0092-8674(00)81703-1
Miedes E, Suslov D, Vandenbussche F, Kenobi K, Ivakov A, Van Der Straeten D, Lorences EP, Mellerowicz EJ, Verbelen JP, Vissenberg K (2013) Xyloglucan endotransglucosylase/hydrolase (XTH) overexpression affects growth and cell wall mechanics in etiolated Arabidopsis hypocotyls. J Exp Bot 64:2481–2497. https://doi.org/10.1093/jxb/ert107
Nakata M, Matsumoto N, Tsugeki R, Rikirsch E, Laux T, Okada K (2012) Roles of the middle domain-specific WUSCHEL-RELATED HOMEOBOX genes in early development of leaves in Arabidopsis. Plant Cell 24:519–535. https://doi.org/10.1105/tpc.111.092858
Ohmori Y, Tanaka W, Kojima M, Sakakibara H, Hirano HY (2013) WUSCHEL-RELATED HOMEOBOX4 is involved in meristem maintenance and is negatively regulated by the CLE gene FCP1 in rice. Plant Cell 25:229–241. https://doi.org/10.1105/tpc.112.103432
Palovaara J, de Zeeuw T, Weijers D (2016) Tissue and organ initiation in the plant embryo: a first time for everything. Annu Rev Cell Dev Biol 32:47–75. https://doi.org/10.1146/annurev-cellbio-111315-124929
Park SO, Zheng Z, Oppenheimer DG, Hauser BA (2005) The PRETTY FEW SEEDS2 gene encodes an Arabidopsis homeodomain protein that regulates ovule development. Development 132:841–849. https://doi.org/10.1242/dev.01654
Penfield S (2017) Seed dormancy and germination. Curr Biol 27:R874–R878. https://doi.org/10.1016/j.cub.2017.05.050
Petzold HE, Chanda B, Zhao C, Rigoulot SB, Beers EP, Brunner AM (2018) DIVARICATA AND RADIALIS INTERACTING FACTOR (DRIF) also interacts with WOX and KNOX proteins associated with wood formation in Populus trichocarpa. Plant J 93:1076–1087. https://doi.org/10.1111/tpj.13831
Pi L, Aichinger E, van der Graaff E, Llavata-Peris CI, Weijers D, Hennig L, Groot E, Laux T (2015) Organizer-derived WOX5 signal maintains root columella stem cells through chromatin-mediated repression of CDF4 expression. Dev Cell 33:576–588. https://doi.org/10.1016/j.devcel.2015.04.024
Pillitteri LJ, Bemis SM, Shpak ED, Torii KU (2007) Haploinsufficiency after successive loss of signaling reveals a role for ERECTA-family genes in Arabidopsis ovule development. Development 134:3099–3109. https://doi.org/10.1242/dev.004788
Rajjou L, Duval M, Gallardo K, Catusse J, Bally J, Job C, Job D (2012) Seed germination and vigor. Annu Rev Plant Biol 63:507–533. https://doi.org/10.1146/annurev-arplant-042811-105550
Romera-Branchat M, Ripoll JJ, Yanofsky MF, Pelaz S (2013) The WOX13 homeobox gene promotes replum formation in the Arabidopsis thaliana fruit. Plant J 73:37–49. https://doi.org/10.1111/tpj.12010
Sarkar AK, Luijten M, Miyashima S, Lenhard M, Hashimoto T, Nakajima K, Scheres B, Heidstra R, Laux T (2007) Conserved factors regulate signaling in Arabidopsis thaliana shoot and root stem cell organizers. Nature 446:811–814. https://doi.org/10.1038/nature05703
Shu K, Liu XD, Xie Q, He ZH (2016) Two faces of one seed: hormonal regulation of dormancy and germination. Mol Plant 9:34–45. https://doi.org/10.1016/j.molp.2015.08.010
Singh AP, Tripathi SK, Nath P, Sane AP (2011) Petal abscission in rose is associated with the differential expression of two ethylene-responsive xyloglucan endotransglucosylase/hydrolase genes, RbXTH1 and RbXTH2. J Exp Bot 62:5091–5103. https://doi.org/10.1093/jxb/err209
Sliwinska E, Bassel GW, Bewley JD (2009) Germination of Arabidopsis thaliana seeds is not completed as a result of elongation of the radicle but of the adjacent transition zone and lower hypocotyl. J Exp Bot 60:3587–3594. https://doi.org/10.1093/jxb/erp203
Smit ME, McGregor SR, Sun H, Gough C, Bagman AM, Soyars CL, Kroon JT, Gaudinier A, Williams CJ, Yang XY, Nimchuk ZL, Weijers D, Turner SR, Brady SM, Etchells JP (2020) A PXY-mediated transcriptional network integrates signaling mechanisms to control vascular development in Arabidopsis. Plant Cell 32:319–335. https://doi.org/10.1105/tpc.19.00562
Steinbrecher T, Leubner-Metzger G (2017) The biomechanics of seed germination. J Exp Bot 68:765–783. https://doi.org/10.1093/jxb/erw428
Sun T, Goodman HM, Ausubel FM (1992) Cloning the Arabidopsis GA1 locus by genomic subtraction. Plant Cell 4:119–128. https://doi.org/10.1105/tpc.4.2.119
Sun B, Zhou YY, Cai J, Shang EL, Yamaguchi N, Xiao J, Looi LS, Wee WY, Gao XY, Wagner D, Ito T (2019) Integration of transcriptional repression and polycomb-mediated silencing of WUSCHEL in floral meristems. Plant Cell 31:1488–1505. https://doi.org/10.1105/tpc.18.00450
Tadege M, Lin H, Bedair M, Berbel A, Wen J, Rojas CM, Niu L, Tang Y, Sumner L, Ratet P, McHale NA, Madueno F, Mysore KS (2011) STENOFOLIA regulates blade outgrowth and leaf vascular patterning in Medicago truncatula and Nicotiana sylvestris. Plant Cell 23:2125–2142. https://doi.org/10.1105/tpc.111.085340
Tsuchiya M, Satoh S, Iwai H (2015) Distribution of XTH, expansin, and secondary-wall-related CesA in floral and fruit abscission zones during fruit development in tomato (Solanum lycopersicum). Front Plant Sci 6:323. https://doi.org/10.3389/fpls.2015.00323
Tvorogova VE, Krasnoperova EY, Potsenkovskaia EA, Kudriashov AA, Dodueva IE, Lutova LA (2021) What does the WOX say? Review of regulators, targets, partners. Mol Biol 55:311–337. https://doi.org/10.1134/S002689332102031X
Ueda M, Zhang Z, Laux T (2011) Transcriptional activation of Arabidopsis axis patterning genes WOX8/9 links zygote polarity to embryo development. Dev Cell 20:264–270. https://doi.org/10.1016/j.devcel.2011.01.009
van der Graaff E, Laux T, Rensing SA (2009) The WUS homeobox-containing (WOX) protein family. Genome Biol 10:248. https://doi.org/10.1186/gb-2009-10-12-248
Van Sandt VS, Suslov D, Verbelen JP, Vissenberg K (2007) Xyloglucan endotransglucosylase activity loosens a plant cell wall. Ann Bot 100:1467–1473. https://doi.org/10.1093/aob/mcm248
Wang Z, Chen F, Li X, Cao H, Ding M, Zhang C, Zuo J, Xu C, Xu J, Deng X, Xiang Y, Soppe WJ, Liu Y (2016) Arabidopsis seed germination speed is controlled by SNL histone deacetylase-binding factor-mediated regulation of AUX1. Nat Commun 7:13412. https://doi.org/10.1038/ncomms13412
Wu X, Dabi T, Weigel D (2005) Requirement of homeobox gene STIMPY/WOX9 for Arabidopsis meristem growth and maintenance. Curr Biol 15:436–440. https://doi.org/10.1016/j.cub.2004.12.079
Wu X, Chory J, Weigel D (2007) Combinations of WOX activities regulate tissue proliferation during Arabidopsis embryonic development. Dev Biol 309:306–316. https://doi.org/10.1016/j.ydbio.2007.07.019
Wu CC, Li FW, Kramer EM (2019) Large-scale phylogenomic analysis suggests three ancient superclades of the WUSCHEL-RELATED HOMEOBOX transcription factor family in plants. PLoS ONE 14:e0223521. https://doi.org/10.1371/journal.pone.0223521
Xie Q, Frugis G, Colgan D, Chua N-H (2000) Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development. Genes Dev 14:3024–3036. https://doi.org/10.1101/gad.852200
Yasui Y, Ohmori Y, Takebayashi Y, Sakakibara H, Hirano HY (2018) WUSCHEL-RELATED HOMEOBOX4 acts as a key regulator in early leaf development in rice. PLoS Genet 14:e1007365. https://doi.org/10.1371/journal.pgen.1007365
Zhang Y, Jiao Y, Liu Z, Zhu YX (2015) ROW1 maintains quiescent centre identity by confining WOX5 expression to specific cells. Nat Commun 6:6003. https://doi.org/10.1038/ncomms7003
Zhou Y, Liu X, Engstrom EM, Nimchuk ZL, Pruneda-Paz JL, Tarr PT, Yan A, Kay SA, Meyerowitz EM (2015) Control of plant stem cell function by conserved interacting transcriptional regulators. Nature 517:377–380. https://doi.org/10.1038/nature13853
Zhu J, Shi H, Lee BH, Damsz B, Cheng S, Stirm V, Zhu JK, Hasegawa PM, Bressan RA (2004) An Arabidopsis homeodomain transcription factor gene, HOS9, mediates cold tolerance through a CBF-independent pathway. Proc Natl Acad Sci USA 101:9873–9878. https://doi.org/10.1073/pnas.0403166101
Zuo J, Niu QW, Frugis G, Chua NH (2002) The WUSCHEL gene promotes vegetative-to-embryonic transition in Arabidopsis. Plant J 30:349–359. https://doi.org/10.1046/j.1365-313X.2002.01289.x
Acknowledgements
The authors thank Dr. Galal Mabrouk very much for his great help improve the manuscript. This work was supported by grants from the National Natural Science Foundation of China (31870305, 32270353, and 32170355).
Author information
Authors and Affiliations
Contributions
YXL and HC planned and designed the research. YY, ZR, and LL conducted the experiments. YL and YH contributed materials and analysis tools. YY, ZR, YXL, and HC analyzed the data. YY and HC wrote the manuscript. YXL and HC revised the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest about the work described in this manuscript.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
425_2024_4357_MOESM1_ESM.docx
Supplementary file 1—Supplementary Fig. S1. Relative transcript abundance of the WOX family genes in dry, stratified, and germinating seeds, and seedlings. a–l The expression levels of WOX4, WOX6, WOX7, WOX5, WOX9, WOX11, WOX12, WOX13, WOX1, WOX2, WOX10, and WOX14 in dry, stratified, and germinating seeds, and seedlings. UBIQUITIN5 was used as an internal control for the normalization of gene expression. Values are means of three biological replicates. The error bars represent SE. Dry seeds, seeds stratified for 24 h, 48 h, and 72 h, seeds germinated for 8 h, 16 h, and seedlings germinated for 24 h, were collected for total RNA extraction. Fig. S2. Phylogenetic analysis of the Arabidopsis WOX family. The unrooted trees were inferred by the neighbor-joining method after the alignment of the protein sequences of the 15 Arabidopsis WOXs. WOX1, WOX2, and WOX4 are highlighted with the red box. Evolutionary analysis was conducted in the TBtools program. Fig. S3. Seed dormancy phenotypes of wild-type Col and wox mutants. a Germination after different dry storage periods of wild type (Col), wox2, and wox2-3. b Germination after different dry storage periods of Col, wox1, wox2, and wox1 wox2. c Germination after different dry storage periods of Col, wox2, wox4-1, and wox2 wox4-1. Seeds were stored at room temperature for after-ripening. Percentages of seed germination are means (± SE) based on at least eight individual plants for each genotype. Fig. S4. wox2 wox4-1 double mutant seedlings display a range of cotyledon phenotypes. a Six-day-old wild-type (Col) seedling with two normal cotyledons. b–e Six-day-old wox2 wox4-1 seedlings with two cotyledons positioned at an angle (b), two cotyledons of unequal size (c), single cotyledon (d) and heart-shaped cotyledon (e). Bars = 1 mm. Fig. S5. Total RNA was isolated from seeds imbibed for 18 h of wild type and wox2 wox4-1 for RNA-seq. Imbibition for 18 h was the first time point at which the germination of wox2 wox4-1 was significantly lower than that of Col. Seeds were stratified at 4 °C for 3 days before the germination experiment. Values are the means ± SE based on the seeds from six to eight individual plants for each genotype. The double asterisk indicates a significant difference compared to the wild type as determined by Student’s t-test (**P < 0.01). Fig. S6. Pathway enrichment analysis of DEGs in the boxes-marked pathways in Fig. 5a. XTHs genes are highlighted with the red box. The dot size is proportional to the gene number. The color scale indicates the significance level. (DOCX 1885 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, Y., Ren, Z., Li, L. et al. WOX2 functions redundantly with WOX1 and WOX4 to positively regulate seed germination in Arabidopsis. Planta 259, 83 (2024). https://doi.org/10.1007/s00425-024-04357-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-024-04357-7