Abstract
Purpose
MicroRNAs were identified as molecules that participate in gene regulation; alterations in their expression characterize central nervous system (CNS). Information in pediatrics is scarce, so the objective of this work was to determine and then compare the patterns of expression of microRNAs in astrocytomas, ependymomas, and medulloblastomas, as well as in non-neoplastic brain.
Methods
Low-density arrays were utilized to evaluate 756 microRNAs in three samples of each type of tumor and non-neoplastic brain. The relative expression was calculated in order to identify the three microRNAs whose expression was modified notably. This was verified using RT-qPCR in more number of tumor samples.
Results
The microRNAs selected for testing were miR-100-5p, miR-195-5p, and miR-770-5p. A higher expression of miR-100-5p was observed in the astrocytomas and ependymomas compared to the medulloblastomas: on average 3.8 times (p < 0.05). MiR-770-5p was expressed less in medulloblastomas compared to astrocytomas four times (p = 0.0162). MiR-195-5p had a low expression in medulloblastomas compared to non-neoplastic cerebellum (p = 0.049). In all three tumor types, expression of miR-770-5p was lower than in non-neoplastic brain (p < 0.001).
Conclusions
These microRNAs may represent potential markers in these tumors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Central nervous system tumors, CNST, belong to a particularly large and diverse group and almost 100 different types that have been described [7]. These tumors are still classified by anatomical location, clinical status, and the histopathological characteristics of the lesion [29]. Gliomas, for example, have characteristic genetic alterations are now allowing researchers to subdivide patients according to molecular patterns associated with greater survival [11]. In medulloblastomas, in addition to the histopathological classification, the molecular categorization based on different pathways—such as wingless (Wnt) or sonic hedgehog (Shh)—allows them to be recognized as tumors that best respond to treatment [8], thus opening research areas for the development of drugs that can inhibit these pathways [30]. The panorama in ependymomas is even less clear because of their marked genetic heterogeneity and the absence of distinguishable groups of genetic alterations that could orient patient management [31]. These facts emphasize the need to design and conduct more profound studies of the molecular biology of these types of tumors, especially in children [7]. For years, work has focused at the genetic level, but it is now clear that the epigenetic level exercises significant regulation. One such epigenetic regulator that acts at the level of the cytoplasm is performed by microRNAs. MicroRNAs are recognized as a relatively “new” regulating mechanism and one of the most abundant forms of post-transcriptional regulation [22]. Why are microRNAs important for the CNS? Because by 2009, at least 70% of all microRNAs detected and reported were found in the cerebral tissue of mammals with different neuronal subtypes with the highest concentration in the cerebral cortex and cerebellum [10, 21]. In brain tumors, several and different techniques in order to reveal global expression patterns of microRNAs and their differences against non-tumoral brain have been used [4]. The MiRNAome can be analyzed in formalin-fixed and paraffin-embedded tissue. First studies have been reported down regulation of 192 microRNAs in high-grade astrocytomas compared to non-tumoral brain [35]. Analysis of genome-wide mRNA (messenger RNA) expression by microarray analysis on 12 brain tumor samples detected a highly correlation between microRNA -mRNA, and they were validated by over-expression or suppressing the microRNA and the presumable targeted mRNA levels [27]. From an extensive microarray analysis carried on 252 tumor samples including 15 brain primary tumor samples, 187 non-brain primary tumors, and 50 brain metastases from different origins, microRNAs could also be used as biomarkers for brain tumors, since some microRNAs previously found to be over-expressed in brain stem cells and during neuronal development are significantly over-expressed in primary brain tumors samples, but not in metastatic brain tumor samples [34]. Also, from the analysis of microRNA signatures, they can differentiate secondary glioblastoma and primary glioblastoma from anaplastic astrocytoma with 94.8% of accuracy, secondary glioblastoma from primary glioblastoma, and progressive astrocytoma from de novo glioblastoma [38] This is the same case as in medulloblastoma: analysis of microRNAs has revealed dysregulation on this tumor when microRNAs are compared with non-neoplastic brain [45]. With TaqMan low-density arrays (TLDAs), some authors have been able to identify a signature of microRNAs that can distinguish normal ependymal cells from ependymomas and some associated to both time to relapse and overall survival [5]
It is known that tumors which develop in the CNS of children have molecular mechanisms distinct from their equivalents in adults [41]. Since the number of studies of microRNA alterations in children is scarce, we present a descriptive study of altered microRNAs in tumors in children and compare them against non-neoplastic brain.
Methods
Characteristics of patients and samples
Biological samples from the three most frequent tumors of the CNS in Mexican patients treated at the Hospital Infantil de Mexico Federico Gómez-Instituto Nacional de Salud were collected between 2000 and 2013 at the Pathology Department. Materials included formalin-fixed paraffin-embedded (FFPE) sections and some frozen tissues. All samples were reviewed to identify those that contained only tumor tissue and, in the case of the paraffin blocks, at least 80% of the tumor lesion. Applying this first inclusion criterion to a total of 166 samples, we determined that only 68 biopsies satisfied study conditions. At the end of this re-evaluation, the groups consisted of the following samples: 28 cases of low- and high-grade astrocytomas (23 FFPE, 5 frozen), 20 medulloblastomas (14 FFPE, 6 frozen), and 20 ependymomas (11 FFPE, 9 frozen). Characteristics of patients are shown in Table 1.
Low-density TaqMan arrays to reveal global expression of microRNAs
To determine the global pattern of expression for each tumor type, three samples each from astrocytomas, classic histological medulloblastomas, and classic histological ependymomas were selected. All these were frozen samples. Non-neoplastic brain samples came from fresh tissue obtained from three autopsies performed in the Pathology Department on patients who died of non-oncological causes. Total RNA was extracted with Trizol reagent (Invitrogen, Carlsbad, CA). All samples received treatment with DNase, and those that showed an A260/A280 ratio of 1.7 to 2.1 were included in this study. The concentration and quality of the RNA from each sample were verified using a Nanodrop, ND-1000 (Thermo Fisher Scientific, Waltham, MA) and 2100 Bioanalyzer (Agilent, Palo Alto, CA). Complementary DNA (cDNA) was synthetized from 1 μg of RNA using the Megaplex Kit (Applied Biosystems, Foster City, CA). For each sample, we evaluated a total of 756 microRNAs, divided into reactions A and B. For real-time PCR, the study used TLDA (Applied Biosystems, Foster City, CA), divided in two arrays (plates A v2.0 and B v3.0). Amplifications were performed in a Viia7 thermocycler (Applied Biosystems, Foster City, CA). All microRNAs with a Ct > 35 were excluded and levels of expression were normalized with endogenous RNU44 and RNU48. Next, the relative quantification method was utilized through the 2−ΔCt formula [36]. The expression values obtained for each tumor type were compared to the samples of normal brain and among the three types. The microRNAs were amplified only in control brains, and only all three types of tumors were identified. Normalized expression values were then used to elaborate heat maps using the MultiExperiment Viewer program 4.6 (MeV) (Dana-Farber Cancer Institute, Boston MA), and significant microRNAs were obtained with significance analysis of microarrays (SAM). Regarding the expression of the microRNAs in the different tumors, this aspect was analyzed by applying a Student’s t test and an ANOVA. Differences between groups were considered significant when the value of p was < 0.05.
Confirmation of the altered expression of the selected microRNAs by RT-qPCR
Based on the results of microRNAs expressed in the three kind of tumors from the TLDA analysis, the microRNAs miR-100-5p, miR-195-5p, and miR-770-5p were chosen, and their expression was evaluated in 28 astrocytomas, 20 ependymomas, and 20 medulloblastomas to corroborate the results of the TLDA analysis. This step involved TaqMan probes (Applied Biosystem, Foster City, CA) from each microRNA to determine relative quantification (Table 2). The expression values of all microRNAs were averaged for each tumor type and normal tissues and then compared using a Student’s t test. Differences were considered significant when the p value was < 0.05.
Expression analysis of the mRNA of the EPDR1 gene using RT-qPCR and of the protein by immunohistochemistry in tumor and normal tissues
An analysis at www.targetscan.org revealed that the mRNA of ependymin-related protein 1, EPDR1 gene, has 14 nucleotides conserved in the seed 3′UTR region. With this information, quantification of the levels of expression of the mRNA of EPDR1 was performed by RT-qPCR, with primers with the following sequences: EPDR1-forw 5′-CCTACGACGGGCTCAACC-3′, EPDR1-rev 5′-GGATCAGCGCCTTCCTCTC-3′. The cDNA synthesis was performed with 100 ng of mRNA. The real-time reaction conditions were as follows: initial denaturalization at 95 °C for 10 min, then 40 cycles of denaturalization at 95 °C for 15 s, and alignment/extension at 60 °C for 1 min. A Stratagene Mx2100 thermocycler was used (Palo Alto, CA). Real-time quantification was performed using SYBR green as the principal fluorochrome, and ROX as the passive stain reference, both included in the Maxima SYBR green qPCR Master Mix (Thermo Scientific, Waltham, MA). No-template controls (NTC) were included, and the samples in which a Ct (threshold cycle) ≥ 35 cycles were eliminated.
Immunohistochemistry was performed with 5-μm sections mounted on charged slides. Blocking of endogenous peroxidase was done with 3% hydrogen peroxide, the antigen was recovered with citrate buffer and EDTA at pH 9, and protein-blocking was conducted with normal horse serum. The primary human antibody anti-EPDR1 (Santa Cruz Biotechnology, Santa Cruz, CA) was diluted to 1:40 in normal horse serum (dil. 1:10) in a PBS solution at pH 7.0 for 24 h. After it was incubated with the secondary antibody (multilink) for 3 h, diaminobenzidine was added and, finally, counter-staining of nuclei was performed with Hill’s hematoxylin. Then, the number of positive and negative cases for the presence of the EPDR1 protein was recorded.
Levels of expression of the microRNAs in relation to patient data
Expression of the three microRNAs selected was analyzed to determine a relation to tumor grade (in astrocytomas), to histological classification (in medulloblastomas), and to the anatomical location of the tumor, survival rates, and recurrence. Differences between the levels of expression of these microRNAs in relation to the groups were analyzed by a Student’s t test. Differences were considered significant when the value of p was < 0.05.
Results
Analysis of expression of the microRNAs in tumors of the CNS
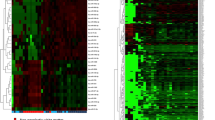
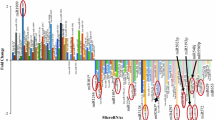
Of the 756 microRNAs evaluated, 51% were expressed in astrocytomas and ependymomas, 53% in medulloblastomas, and 37% in normal tissues. This indicates, first, that the tumors express under half of all the microRNAs analyzed, and more than those found in normal tissues. One group of highly expressed microRNAs was seen to be shared by all three tumor types: miR-195-5p, miR-652-3p, miR-100-5p, miR-20a-5p, and miR-532-5p. Similarly, four microRNAs with low expression—miR-770-5p, miR-330-3p, miR-1254, and miR-605-5p—were also shared (Table 3). The heat maps show the microRNAs that had a significant expression in each tumor type. The microRNAs that were in area of higher concentration (red) and the lower (green) were included (Fig. 1). These data show that not only astrocytomas, ependymomas, and medulloblastomas can share a group of microRNAs (Table 3), but also each tumor has their own microRNAs that acts above them (Table 4). It is important to highlight that miR-93-5p, miR-19b-3p, miR-21-5p, and miR-25-3p are important because they were also identified as being shared in other reports that studied low- and high-grade astrocytomas and medulloblastomas. Also, analysis of the heat maps (Fig. 1) showed that miR-100-5p and miR-195-5p were abundant in gliomas, though this was not the case of miR-195-5p in the medulloblastomas. Regarding miR-770-5p, a reduction in its expression was seen in all three tumor types, but this was most evident in astrocytomas and medulloblastomas (p < 0.0001). As a result, these three microRNAs were chosen to evaluate their expression in a larger number of tumors of the CNS. Moreover, microRNAs only expressed in non-neoplastic brain were miR-622, miR-519a, and miR-518e; however, only miR-622 showed a value of expression of 19. They represented the 0.45%. On the other hand, we found microRNAs that were expressed in tumors but not in non-neoplastic brain. In these cases, differences in expression were observed between the three types of tumors. The miR-28 had a greater expression in the astrocytomas (eight times more) compared to the medulloblastomas and two times more than in the ependymomas. The miR-1290 was expressed 7-fold more in ependymomas compared to medulloblastomas, and the miR-888 had greater expression in the ependymomas (five times more) compared to the medulloblastomas. In total, they were 11 microRNAs that represented 1.67%.
Real-time PCR expression profiling of mature microRNAs in a astrocytomas, b ependymomas, and c medulloblastomas, using arrays in TLDA plates. Heat map generated from the log of 2−ΔCt. The microRNAs showed significant changes in each tumor with respect to controls using SAM (significance analysis of microarrays). The color range starts at − 10 and corresponds to green color and represents microRNAs with lower expression. Value 1 corresponds to red color and microRNAs with high expression. Black color shows microRNAs without any change in their expression
Evaluation of miR-100-5p, miR-195-5p, and miR-770-5p expression in tumors
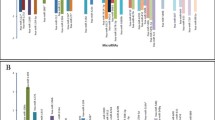
The most highly expressed microRNA in the three tumors was miR-100-5p. In contrast, miR-770-5p appeared less in the tumors than control (brain tissue and cerebellum for gliomas and medulloblastomas, respectively, p < 0.0001 (Fig. 2). MiR-195-5p was expressed more often in astrocytomas and ependymomas (11 times) than controls (brain tissue), while in medulloblastomas, miR-195-5p decreased 2-fold with respect to its control (cerebellum) (p = 0.04) (Fig. 2c). Comparing the expression values, we found that expression of miR-195-5p was highest in the astrocytomas and lowest in the medulloblastomas. The largest difference was seen in the expression of miR-770-5p, which was four times lower in medulloblastomas than astrocytomas (0.00692 vs 0.0289, respectively) (p = 0.016).
Expression of microRNAs in TSNC. a, b In the case of astrocytomas and ependymomas, expression was compared to non-tumorous brain tissue. c In the case of the medulloblastomas, against non-tumorous cerebellum tissue. In white bars are tumor expressions, and in gray bars are control expressions. *p ≤ 0.0001; **p = 0.049
Expression of the mRNA of the EPDR1 gene and protein in tumor tissues by immunohistochemistry
Given that miR-100-5p increased its expression 3.8 times in astrocytomas and ependymomas and 2.9 times in medulloblastomas, the next step was to determine the expression of one of its main targets described in TargetScan, for example, EPDR1. Results showed a higher, though discrete, expression of the mRNA of EPDR1 in tumor tissues, compared to that observed in cerebellum and normal brain tissues (Fig. 3). Finally, staining of EPDR1 by immunohistochemistry in cases of low-grade astrocytoma (seven cases), high-grade astrocytoma (four cases), ependymoma (five cases), medulloblastoma (five cases), brain (five cases), and cerebellum (five cases) was performed. Findings showed a strong, significant expression of EPDR1 in the brain sections that was also important in the cerebellum (Fig. 4). This pattern contrasts sharply with the absence of staining in sections from high-grade astrocytoma, ependymoma, and medulloblastoma; only a few cases of low-grade astrocytoma were positive to staining (five of seven cases) (Fig. 4).
EPDR1 expression (ependymin-related protein 1) in CNST and brain and cerebellum tissue. The mean expression values of astrocytomas (Ast) and ependymomas (Epe) were compared to non-tumorous brain tissue, and the mean expression values of the expression of medulloblastomas (Med) were compared to non-tumorous cerebellum tissue. In white bars are tumor expressions, and in gray bars are control expressions. The endogenous control was HPRT (hypoxanthine phosphoribosyltransferase). *p = 0.005
Immunohistochemistry of the EPDR1 protein in sections of tumors of the CNS and normal tissues. a Astrocytoma GI positive, × 10. b Astrocytoma GII negative, × 10. c Astrocytoma GIV negative, × 10. d Ependymoma negative, × 10. e Medulloblastoma negative, × 10. f Cerebral cortex positive, × 10. g Cerebellum positive; positive Purkinje cell and glial cells, × 10
Expression levels of microRNAs and their relation to patient data
The analysis of the astrocytomas divided by tumor grade, location, survival, and recurrence, as well as the levels of expression of the three microRNAs (Fig. 5) revealed a significant reduction of miR-770-5p, as reported previously in the results of the low-density arrays (Fig. 1; Table 3); this decrease was more evident in patients with low grade and live tumors; however, it was not significant. For miR-100-5p and miR-195-5p, there are evident upwards tendencies in their levels in the low-grade tumors that are infratentorial. Moreover, for miR-100-5p and miR-195-5p, an increase was seen in the groups of live patients (survival) and without recurrence (Fig. 5). Statistical analysis showed that the increase in the expression values of the low-grade infratentorial tumors was significant for miR-100-5p compared to high-grade supratentorial tumors (p < 0.05). In the case of miR-195-5p, there were no significant differences. Of course, the number of cases in these two groups differed, but the tendency was evaluated by Welch correction for uneven groups in the number of cases.
Values of expression of miR-100-5p, 195-5p, and 770-5p in astrocytomas. MiR-100-5p and miR-195-5p showed the same trend. a Expression of miR-100-5p, miR-195-5p and miR-770-5p in low (white bars) and high-grade (gray bars) astrocytomas; in low-grade tumors, the expression of miR-100-5p and miR-195-5p was high, whereas the expression of miR-770-5p was low; only significant differences were observed in the case of miR-100-5p, *p = 0.0384. b Expression of microRNAs in astrocytomas according to anatomical location. Comparison of infratentorial (white bars) vs supratentorial tumors (gray bars), differences were significant in the case of miR-100 (*p = 0.0274). c Expression of microRNA in patients with tumors according to no recurrence (white bars) and recurrence (gray bars). d Expression of microRNAs by current patient status, live (white bars) and death (gray bars). In the case of miR-770-5p, no significant differences were observed between groups. Student’s t test with Welch correction
In relation to the patients with ependymomas and medulloblastomas, we analyzed the expression of the three microRNAs selected according to survival and recurrence data, and for the histological classification of medulloblastomas, but no important data or significant differences were found.
Discussion
MicroRNAs obtained from analyses of low-density arrangements
MicroRNAs regulate diverse biological processes including cancer. The present study analyzed low-density arrays of 784 microRNAs in the three most common tumors of the CNST in Mexican children. This work proves that tumor tissues express a higher percentage of microRNAs than non-tumor tissues, indicating, albeit indirectly, that many more cellular functions are altered than in normal tissues. This analysis does allow us to identify microRNAs that characterize tumors and, therefore, relate them to the characteristics of patients. Meanwhile, Wang (2012) and Miele (2014) examined pediatric gliomas and showed miR-20a with high expression [33, 48]. This result is consistent with our study; we reported high expression of this microRNA, but in the three types of tumors. Finally, Jia (2013) reported to miR-19b with high expression in gliomas [19], but in our study, the expression was low in astrocytomas. As can be seen, despite the fact that Miele and cols. (2014), for example, utilized the same TLDAs as was done in this study and analyzed pediatric gliomas, only miR-20a and miR-19b corresponded to that found in our study, miR-20a was the only one that corresponded to those most highly expressed in the three types of tumors we examined. This may reflect two issues: that the differences are due to elements inherent in the techniques used in different platforms or that they reflect the particularity observed in all tumors studied by microarray platforms, which would support the notion that each tumor represents an individual pathological entity, despite sharing a common histopathological pattern. The results from these different study groups, using microRNA analysis, are important because they confirm the complexity of molecular alterations in tumors and their multi-clonal origin. As Table 3 shows, our study demonstrated that miR-100-5p, miR-195-5p, and miR-770-5p are the main microRNAs whose expression is altered, while others were proven to be exclusive to individual tumor types (Table 4). For example, miR-21 is one microRNA that has been reported by other microarray platforms, where the constant result is its increase [12, 15, 16, 26, 53].
Regarding the analysis by arrays of microRNAs in pediatric ependymomas, the only data are from work by Costa and cols. (2011), which used the TLDA system. They reported increase in the expression of miR-135a and miR-17-5p and a reduction in the expression of miR-383 and 485-5p [5]. Our study also reports an important elevation of miR-135a, as did the study by Costa and cols. Finally, with respect to the pattern of expression in medulloblastomas, Ferreti and cols. (2009) found that miR-18a, miR-19a, miR-20a, miR-21, miR-25, miR-106a, miR-106b, and miR-137 are most highly expressed in tumor tissue, compared to normal tissue [9]. Of these microRNAs, our study found that miR-25 and miR-592 also increased in medulloblastomas. It is important to mention that the expression level of miR-592 has allowed differentiate molecular groups of medulloblastomas [20, 23]. On the other hand, miR-330-3p had a reduced expression in our study; this microRNA in cell lines of GBM, promoted cell proliferation, migration, and inhibiting apoptosis by negatively regulating SH3GL2 [37]. In this regard, Visani et al. (2014) reported low expression of miR-330-3p in 60 GBM [46]. The alteration in the expression of this microRNA in distinct tumors means that this microRNA may constitute a new target of therapies. In this study, its expression was reported as low in all three tumor types, especially the medulloblastomas.
MicroRNAS only expressed in control brain were miR-622, miR-519a, and miR-518e; however, only miR-622 showed high expression. This microRNA is described as tumor suppressor in several tumors; in gliomas tissue and cell lines miR-622, it is downregulated and its over-expression suppressed tumor cell proliferation, migration, and invasion; this microRNA can be considered potential therapeutic target for the treatment of patients with malignant gliomas [47, 55]. On the other hand, we found microRNAs that were expressed in tumors but not in control brain. In these cases, differences in expression were observed between the three types of tumors. Within this group, miR-10a, miR-28, and miR-92 have been reported in gliomas with high expression [32, 43, 51]; miR-23b and miR-181c with low expression [14, 25] and miR-15a reported with variable expressions [3, 32, 50]. No doubt these microRNAs contribute or are altered with the development of the tumor.
Importance of microRNAs with altered expression in the three tumor types
Of the microRNAs whose expression was modified in all three tumor types, miR-195-5p has been most intensely studied in glioblastoma cells, its decrease showed a significant effect of cell death [44]. MiR-195 has also been associated as a marker; in the U87 MG cell line of gliomas, studies show that therapy with cyclosporine A induces a reduction in the expression of miR-195 improves the effect of the cyclosporine in these cells [52]. Finally, in relation to the cell cycle, miR-195 has a significant participation because its targets include important regulators in glioma cells [18, 54]. In relation to miR-770-5p, there are no reports suggesting its possible participation in tumors of the CNS; however, its targets can activate signaling of the Wnt/β-catenin pathway [49].
Another microRNA reported to show increased expression in the three tumor types was miR-100-5p, as Liu and cols. (2009) found this in pediatric medulloblastomas, in a form similar to our findings [28]. The expression patterns and effects of miR-100 in tumor progression have not been fully elucidated, depending on tumor types and microenvironment and could be a good candidate for use as a prognostic biomarker and a potential therapeutic target for tumors of CNS [1]. The first report of an increase of miR-100, at least in medulloblastomas, was by Liu and cols. (2009) [28]. There are no reports documenting an increase of miR-100-5p in astrocytomas and ependymomas, so our work represents the first study. It is important to mention that miR-100-5p forms part of the microRNAs that play an important role in determining the fate of cells during differentiation of the CNS [42], so it is possible that their high expression in tumor cells from astrocytomas, ependymomas, and medulloblastomas could be related to reduced differentiation. According to scarce information about miR-100-5p and its relevant expression in three types of tumors (Table 2), we decided to study one of its targets, EPDR1, described on three website sites of targets: www.targetscan.org; www.microrna.org, and www.mirdb.org.
EPDR1
The mRNA of EPDR1 gene has 14 nucleotides conserved in the seed 3′UTR region. The key characteristic of these proteins is that they bind calcium ions, thanks to the presence of modifications of sialic acid [17], which also can form part of the blood-brain barrier and in neural regeneration [40]. In relation to its expression, it has been reported that the brain is one of the human tissues with the highest expression of EPDR1 [2, 39]. The protein expression was significantly reduced in high-grade astrocytomas, ependymomas and medulloblastomas (Fig. 4), despite the increased levels mRNA of EPDR1 (Fig. 3). The control of expression by microRNAs is performed at the level of the translation which produces a decrease of the protein. Most plant cells microRNAs show perfect complementarity to their 3′UTR and cause endonucleolytic mRNA cleavage to the 10th and 11th nucleotides of the microRNA target and a decreasing notable of it [6]. In animal cells, microRNAs repress protein levels with little or no influence on mRNA levels. The microRNA cause mRNA deadenylation which promotes a de-capping and with the time mRNA turnover. These two modifications produce destabilization usually modest which represses the translation without detectable mRNA changes because there is no mRNA endonucleolytic cleavage as does in plants [13]. This could explain the contrary expression pattern of EPDR1 protein and mRNA in tumoral tissues.
MicroRNA expression and its relation to patient data
Some studies conducted with miR-195 have produced contradictory results. On the one hand, a tumor-suppressor effect has been shown since miR-195 has important targets, including cyclin D1 and E1 [18], as well as E2F3 and CCND3 [54], such that an increase in its expression induces the reduction of these proteins that participate very significantly in the cell cycle. In this sense, the elevated miR-195-5p levels in our study also suggest a role as a tumor suppressor; although the differences were not statistically significant, we did find higher levels in the patients with low-grade infratentorial astrocytomas without recurrence. However, this finding needs to be tested with a larger number of cases.
In relation to miR-100-5p, it has been reported to have tumor-suppressor activity in glioblastoma cells [1]. In this study, we observed higher levels in patients with low-grade astrocytomas and infratentorial localization without recurrence; in this case, the differences were statistically significant. A report in this regard, conducted by Lakomy and cols. (2011), described that the tumor-suppressor effect of miR-100 was associated with high survival rates in patients with GBM and, in fact, had a better correlation than the state of expression of O-6-methylguanine-DNA methyltransferase [24].
Conclusions
Tumor tissues of astrocytoma, ependymoma, and medulloblastoma express higher quantities of microRNAs. MiR-100-5p and miR-195-5p are expressed at high levels and occurred in all three tumor types. MiR-770-5p is expressed at low levels, a feature it shares with all three tumor types analyzed. The levels of expression of the mRNA, the protein of EPDR1, and miR-100-5p will be analyzed in a larger number of cases of different tumors of the CNS in order to identify whether the presence of the protein can indicate this protein as a new tumor marker.
References
Alrfaei B, Vemuganti R, Kuo J (2013) MicroRNA-100 targets SMRT/NCOR2, reduces proliferation, and improves survival in glioblastoma animal models. PLoS One 8:1–13
Apostolopoulos J, Sparrow R, McLeod J, Collier F, Darcy P, Slater H, Ngu C, Gregorio-King C, Kirkland M (2004) Identification and characterization of a novel family of mammalian ependymin-related proteins (MERPs) in hematopoietic, nonhematopoietic, and malignant tissues. DNA Cell Biol 20:625–635
Braoudaki M, Lambrou G, Giannikou K, Papadodima S, Lykoudi A, Stefanaki K, Sfakianos G, Kolialexi A, Tzortzatou-Stathopoulou F, Tzetis M, Kitsiou-Tzeli S, Kanavakis E (2016) miR-15a and miR-24-1 as putative prognostic microRNA signatures for pediatric pilocytic astrocytomas and ependymomas. Tumor Biol. doi:10.1007/s13277-016-4903-7
Catania A, Maira F, Skarmoutsou E, D’amico F, Abounader R, Mazzarino C (2012) Insight into the role of micrornas in brain tumors. Inter J Oncol 40:605–624
Costa F, Bischof M, Vanin F, Lulla R, Wang M, Sredni T, Rajaram V, Bonaldo F, Wang D, Goldman S, Tomita T, Soares B (2011) Identification of MicroRNAs as potential prognostic markers in ependymoma. PLoS One 6:e25114
Dalmay T (2013) Mechanism of miRNA-mediated repression of mRNA translation. Essays Biochem 54:29–38
Dubuc M, Mack S, Unterberger A, Northcott P, Taylor M (2012) The epigenetics of brain tumors. In Dumitrescu R, Verma M(eds). CancerEpigenetics. Humana Press Toronto, pp139–153
Ellison D (2010) Childhood medulloblastoma: novel approaches to the classification of a heterogeneous disease. Acta Neuropathol 120:305–316
Ferretti E, De Smaele E, Po A, Marcotullio L, Tosi E, Espinola M, Di Rocco C, Riccardi R, Giangaspero F, Farcomeni A, Nofroni I, Laneve P, Gioia U, Caffarelli E, Bozzoni I, Screpanti I, Gulino A (2009) MicroRNA profiling in human medulloblastoma. Int J Cancer 124:568–577
Fineberg K, Kosik S, Davidson L (2009) MicroRNAs potentiate neural development. Neuron 64:303–309
Freije A, Castro-Vargas E, Fang Z, Horvath S, Cloughesy T, Liau L, Mischel P, Nelson S (2004) Gene expression profiling of gliomas strongly predicts survival. Cancer Res 64:6503–6510
Gaur A, Holbeck S, Colburn N, Israel M (2011) Downregulation of Pdcd4 by mir-21 facilitates glioblastoma proliferation in vivo. Neuro-Oncology 13:580–590
Guo H, Ingolia N, Weissman J, Bartel D (2010) Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466:835–840
He X, Liu Z, Peng Y, Yu C (2016) MicroRNA-181c inhibits glioblastoma cell invasion, migration and mesenchymal transition by targeting TGF-β pathway. Biochem Biophys Res Commun 469:1041–1048
Hermansen S, Nielsen B, Aaberg-Jessen C, Kristensen B (2016) MiR-21 is linked to glioma angiogenesis: a co-localization study. J Histochem Cytochem 64:138–148
Ho C, Bar E, Giannini C, Marchionni L, Karajannis M, Zagzag D, Gutmann D, Charles G, Eberhart C, Rodriguez F (2013) MicroRNA profiling in pediatric pilocytic astrocytoma reveals biologically relevant targets, including PBX3, NFIB, and METAP2. Neuro-Oncology 15:69–82
Hoffmann W (1994) Ependymins and their potential role in neuro-plasticity and regeneration: calcium-binding meningeal glycoproteins of the cerebrospinal fluid and extracellular matrix. Int J BioChemiPhysics 26:607–619
Hui W, Yuntao L, Lun L, WenSheng L, ChaoFeng L, HaiYong H, Yueyang B (2013) MicroRNA-195 inhibits the proliferation of human glioma cells by directly targeting cyclin D1 and cyclin E1. PLoS One 8:e54932
Jia Z, Wang K, Zhang A, Wang G, Kang C, Han L, Pu P (2013) MiR-19a and miR-19b overexpression in gliomas. Pathol Oncol Res 13:847–853
Kaur K, Kakkar A, Kumar A, Mallick S, Julka P, Gupta D, Ashish Suri A, Suri V, Sharma M, Sarkar C (2015) Integrating molecular subclassification of medulloblastomas into routine clinical practice: a simplified approach. Brain Pathol. doi:10.1111/bpa.12293
Krichevsky A, King K, Donahue C, Khrapko K, Kosik K (2003) A microRNA array reveals extensive regulation of microRNAs during brain development. RNA 9:1274–1281
Krol J, Loedige I, Filipowicz W (2010) The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet 11:597–610
Kunder R, Jalali R, Sridhar E, Moiyadi A, Goel N, Goel A, Gupta T, Krishnatry R, Kannan S, Kurkure P, Deopujari C, Shetty P, Biyani N, Korshunov A, Pfister S, Northcott P, Shirsat N (2013) A real-time PCR assay based on the differential expression of microRNAs and protein-coding genes for molecular classification of formalin-fixed paraffin embedded medulloblastomas. Neuro-Oncol 15:1644–1651
Lakomy R, Sana J, Hankeova S, Fadrus P, Kren L, Lzicarova E, Svoboda M, Dolezelova H, Smrcka M, Vyzula R, Michalek J, Hajduch M, Slaby O (2011) MiR-195, miR-196b, miR-181c, miR-21 expression levels and O-6-methylguanine-DNA methyltransferase methylation status are associated with clinical outcome in glioblastoma patients. Cancer Sci 102:2186–2190
Li R, Li X, Ning S, Ye J, Han L, Kang C, Li X (2014) Identification of a Core miRNA-pathway regulatory network in glioma by therapeutically targeting miR-181d, miR-21, miR-23b, b-catenin, CBP, and STAT3. PLoS One 9:1–13
Liu F, Xiong Y, Zhao Y, Tao L, Zhang Z, Zhang H, Liu Y, Feng G, Li B, He L, Qin S, Yang Y (2013) Identification of aberrant microRNA expression pattern in pediatric gliomas by microarray. Diagnostic Pathol 8:158–168
Liu T, Papagiannakopoulos T, Puskar K, Qi S, Santiago F, Clay W, Lao K, Lee Y, Nelson F, Kornblum I, Doyle F, Petzold L, Shraiman B, Kosik S (2007) Detection of a microRNA signal in an in vivo expression set of mRNAs. PLoS One 2:804
Liu W, Gong Y, Chao T, Peng X, Yuan J, Ma Z, Jia G, Zhao J (2009) Identification of differentially expressed microRNAs by microarray: a possible role for microRNAs gene in medulloblastomas. Chin Med J 20:2405–2411
Louis D, Ohgaki H, Wiestler O, Cavenee W, Burger P, Jouvet A, Scheithauer B, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114:97–109
Low A, De Sauvage F (2010) Clinical experience with Hedgehog pathway inhibitors. J Clin Oncol 28:5321–5326
Mack C, Taylor D (2009) The genetic and epigenetic basis of ependymoma. Childs Nerv Syst 25:1195–1201
Malzkorn B, Wolter M, Liesenberg F, Grzendowski M, Meyer H, Reifenberger G (2009) Identification and functional characterization of miRNAs involved in the malignant progression of gliomas. Brain Pathol. doi:10.1111/j.1750-3639.2009.00328.x
Miele E, Romana F, Arcella A, Begalli F, Garg N,Silvano M, Po A, Baldi C, Carissimo G, Antonelli M, Spinelli G, Capalbo C, Donofrio V, Morra I, Nozza P, Gulino A, Giangaspero F, Ferretti E (2014) High-throughput microRNA profiling of pediatric high-grade gliomas. Neuro-Oncol 16:228-40
Nass D, Rosenwald S, Meiri E, Gilad S, Tabibian-Keissar H, Schlosberg A, Kuker H, Sion-Vardy N, Tobar A, Kharenko O, Sitbon E, Lithwick G, Elyakim E, Cholakh H, Gibori H, Spector Y, Bentwich Z, Barshack I, Rosenfeld N (2009) MiR-92b and miR-9/9* are specifically expressed in brain primary tumors and can be used to differentiate primary from metastatic brain tumors. Brain Pathol 19:375–383
Negrini M, Maira G, Croce C, Farace M (2005) Extensive modulation of a set of microRNAs in primary glioblastoma. Biochem Biophys Res Commun 334:1351–1358
Pfaffl M (2001) A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res 29:e45
Qu S, Yao Y, Shang C, Xue Y, Ma J, Li Z, Liu Y (2012) MicroRNA-330 is an oncogenic factor in glioblastoma cells by regulating SH3GL2 gene. PLoS One 7:e46010
Rao A, Santosh V, Somasundaram K (2010) Genome-wide expression profiling identifies deregulated miRNAs in malignant astro-cytoma. Mod Pathol 23:1404–1417
Schmidt R, Brysch W, Rother S, Schlingensiepen K (1995) Inhibition of memory consolidation after active avoidance conditioning by antisense intervention with ependymin gene expression. J Neurochem 65:1465–1471
Shashoua V (1991) Ependymin, a brain extracellular glycoprotein, and CNS plasticity. Ann N Y Acad Sci 627:94–114
Sintupisut N, Liu P, Yeang C (2013) An integrative characterization of recurrent molecular aberrations in glioblastoma genomes. Nucleic Acids Res 41:8803–8821
Sokol N (2012) Small temporal RNAs in animal development. Curr Opin Genet Dev 22:368–373
Son J, Jeong H, Park D, Gyoon S, Lee E, Lee J, Chung H (2017) miR-10a and miR-204 as a potential prognostic indicator in low-grade gliomas. Cancer Informat 16:1–6
Ujifuku K, Mitsutake N, Takakura S, Matsuse M, Saenko V, Suzuki K, Hayashi K, Matsuo T, Kamada K, Nagata I, Yamashita S (2010) MiR-195, miR-455-3p and miR-10a are implicated in acquired temozolomide resistance in glioblastoma multiforme cells. Cancer Lett 28:241–248
Venkataraman S, Birks K, Balakrishnan I, Alimova I, Harris S, Patel R, Handler H, Dubuc A, Taylor D, Foreman K, Vibhakar R (2013) MicroRNA 218 acts as a tumor suppressor by targeting multiple cancer phenotype-associated genes in medulloblastoma. J Biol Chem 288:1918–1928
Visani M, de Biase D, Marucci G, Cerasoli S, Nigrisoli E, Bacchi M, Albani F, Baruzzi A, Pession A (2014) Expression of 19 microRNAs in glioblastoma and comparison with other brain neoplasia of grades I-III. Mol Oncol 8:417–430 Erratum in: MolOncol 2015;9:334
Wang X, Xin Z, Xu Y, Ma Y (2015) Upregulated miRNA-622 inhibited cell proliferation, motility, and invasion via repressing Kirsten rat sarcoma in glioblastoma. Tumor Biol. doi:10.1007/s13277-015-4455-2
Wang X, Zhang H, Zhang A, Han L, Wang K, Liu R, Yang S, Pu P, Shen C, Kang C, Yu C (2012) Upregulation of miR-20a and miR-106b is involved in the acquisition of malignancy of pediatric brainstem gliomas. Oncol Reports 28:1293–1300
Wu W, Shi J, Hu G, Yu X, Lu H, Yang M, Liu B, Wu Z (2016) Wnt/β-catenin signaling inhibits FBXW7 expression by upregulation of microRNA-770 in hepatocellular carcinoma. Tumor Biol 37:2313–2320
Xie T, Liu P, Chen L, Chen Z, Luo Y, Chen X, Feng Y, Luo X (2014) MicroRNA-15a down-regulation is associated with adverse prognosis in human glioma. Clin Transl Oncol. doi:10.1007/s12094-014-1265-8
Yan Y, Wang Q, Yan X, Zhang Y, Li W, Tang F, Li X, Yang P (2015) miR-10a controls glioma migration and invasion through regulating epithelial–mesenchymal transition via EphA8. FEBS Lett 589:756–765
Yilmaz S, Biray A, Dodurga Y, Dogan O, Oktar N, Gunduz C (2015) Downregulation of miR-195 via cyclosporin A in human glioblastoma cells. J Buon 20:1337–1340
Zhang K, Han L, Chen L, Shi Z, Yang M, Ren Y (2014) Blockage of a miR-21/EGFR regulatory feedback loop augments anti-EGFR therapy in glioblastomas. Cancer Lett 342:139–149
Zhang Q, Xu H, Huang M, Ma L, Huang Q, Yao Q, Zhou H, Qu L (2012) MicroRNA-195 plays a tumor-suppressor role in human glioblastoma cells by targeting signaling pathways involved in cellular proliferation and invasion. Neuro-Oncology 14:278–287
Zhang R, Luo H, Wang S, Chen Z, Hua L, Wang H, Chen W, Yuan Y, Zhou Y, Li D, Shen S, Jiang T, You Y, Liu N, Wang H (2015) MiR-622 suppresses proliferation, invasion and migration by directly targeting activating transcription factor 2 in glioma cells. J Neuro-Oncol 121:63–72
Acknowledgements
The authors acknowledge Keintel-Márquez Alejandro, Departamento de Patología, Hospital General de México Dr. Eduardo Liceaga.
Funding information
Statement of financial support: FOSISS-CONACYT, Salud-2012-01-181456, and Fondos Federales Hospital Infantil de México Federico Gómez, HIM/2012/005.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The Ethics Committee of Hospital Infantil de Mexico Federico Gómez approved this study.
Conflict of interest
The authors declare that they have no conflict of interests.
Ethical approval
For this type of study, formal consent is not required.
Rights and permissions
About this article
Cite this article
Eguía-Aguilar, P., Gutiérrez-Castillo, L., Pérezpeña-Díazconti, M. et al. Expression of microRNAs in tumors of the central nervous system in pediatric patients in México. Childs Nerv Syst 33, 2117–2128 (2017). https://doi.org/10.1007/s00381-017-3569-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00381-017-3569-9