Abstract
Trans-resveratrol (trans-3,5,4′-trihydroxystilbene) is one of the most promising stilbenes, a type of natural phenol that is produced naturally by some plant species in response to stress. Resveratrol exhibits multiple bioactivities and is used in the agriculture, medical, food, and cosmetic industries due to its antitumor, anti-inflammatory, cardioprotective, and antioxidant properties. Due to the increasing demand, an active area of investigation is the use of plant cell culture and metabolic engineering techniques to produce large quantities of active resveratrol. However, most recent studies have focused on the efficiency of synthesizing resveratrol in vitro, but have not investigated the contributions of the transcriptional activities of the genes encoding the related enzymes in the biosynthesis pathway. This article reviews recently developed methods for the biosynthesis of resveratrol and comprehensively reviews the current state of knowledge of the function of the key pathway enzymes in resveratrol synthesis. Approaches for enhancing resveratrol production, such as introducing non-pathway genes and co-localizing enzymes are described in detail.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Resveratrol (trans-3,5,4′-trihydroxystilbene) is a phenolic compound that belongs to the stilbene class and was first isolated from white hellebore in 1940 (Takaoka 1940). It is widely found in diverse plants, including Japanese knotweed, peanuts, grapes, blueberries, cranberries, and even ferns (Jeandet et al. 2012; Li et al. 2015a; Lim et al. 2011). In nature, resveratrol exists in two geometric isoforms, trans- and cis-resveratrol (Fig. 1). The trans-isomer is more stable than the cis-isomer due to the steric hindrance between aromatic rings but can be transformed to cis by UV treatment (Hasan et al. 2012). Trans-resveratrol is the major product of stilbene synthase (STS), and the latter is an important target enzyme that is widely used in metabolic engineering and plant cell cultures (Chong et al. 2009; Hasan et al. 2012). In addition, glycosylation is a common modification of resveratrol that is catalyzed by glucosyltransferases and generates two corresponding piceid, trans- and cis-piceid (Fig. 1). This modification allows subsequent storage in vacuoles and may both protect plant cells from potentially toxic effects caused by resveratrol and protect the resveratrol from oxidation and enzymatic degradation (Giovinazzo et al. 2012).
Resveratrol is a phytoalexin in plants and is synthesized by phenylpropanoid pathway in response to biotic and abiotic stresses including pathogen infection, UV radiation, and mechanical wounding. In the resveratrol biosynthesis pathway in plants, resveratrol is bioconversed from one p-coumaroyl-CoA and three malonyl CoA by stilbene synthase (STS). Over-expression of STS gene in transgenic plants is an efficient way to increase their resveratrol content as well as their antioxidative properties and resistance to pathogens (Delaunois et al. 2009). Dabauza et al. (2015) over-expressed stilbene synthase gene (Vst1) in grapevine (Vitis vinifera L.) and found that the concentration of resveratrol in the genetically modified plant was up to 7.5-fold higher compared to the control and resulted in increased resistance to Botrytis cinerea. Resveratrol has received widespread attention for its promising pharmaceutical and nutritional values (Belchi-Navarro et al. 2012). Over the last 50 years, numerous studies have found that resveratrol can provide anticancer, cardioprotective, antioxidant, and neuroprotective activities for human health and may also exert lifespan-extending properties (Shi et al. 2014). Resveratrol has been found effective in the inhibition of various tumor cells (Jeandet et al. 2012). It exhibits antitumor activity by altering gene expression by controlling the expression of a small RNA and through the regulation of a series of transcription factors (Mei et al. 2015). Resveratrol also alters many cellular processes, including cell apoptosis, cell proliferation, and cell cycle progression (Leifer and Barberio 2016). Some studies have reported resveratrol has an anti-aging effect on mice, yeast, nematodes, and fruit flies (Lancon et al. 2012; Whitlock and Baek 2012). The mechanism of this effect is thought to be the ability of resveratrol to specifically activate sirtuin-like protein deacetylases. Sirtuins are redox-sensing enzymes that regulate a wide range of important biological processes, modulate the cellular response to environmental stresses, and allow a reduction in general metabolic activity, delayed apoptosis, and increased longevity (Halls and Yu 2008).
Currently, there is high market demand for resveratrol and the natural content of resveratrol in plants is insufficient to meet the demand of over 100 t/year (Mei et al. 2015). One option to meet the needs of the nutritional supplement industry is to utilize resveratrol derivatives (piceatannol, resveratrodehydes, and pterostilbene) in addition to plant-produced resveratrol since they may have equivalent bioactivities (Leifer and Barberio 2016; Lin et al. 2014; Martinez-Marquez et al. 2016; Wang et al. 2014). Another option is to use chemical synthesis, but this approach presents challenges because it requires a complex procedure and requires the use of toxic organic solvents and heavy metals which can result in environmental contamination (Suzuki et al. 2014). As an alternative to chemical synthesis, plant cell culture, hairy roots tissues, and metabolic engineering can be combined to biosynthesize resveratrol on a large scale and with high-yield. These methods have been shown to allow production of a wide spectrum of plant secondary metabolites with multiple bioactivities (Almagro et al. 2015; Becker et al. 2003; Krivoruchko and Nielsen 2015; Lin et al. 2014). Efforts have been made to improve the efficacy of the in vitro platform, such as the use of elicitors, introducing specific genes with different promoters, using different hosts, codon optimization, altering culture conditions, and regulating metabolic flux (Jeandet et al. 2012). Codon optimization can be performed to modify the sequence of the exogenous gene to conform its codon usage to that of the specific host (Klein-Marcuschamer et al. 2010). Compared to the synthesis of protein containing rare codons, codon-optimized sequences are more abundantly expressed in specific hosts, allowing efficient expression of exogenous gene and improved yields of metabolites (Anthony et al. 2009; Chang et al. 2007; Engels et al. 2008; Wang et al. 2011b). In this review, we highlight the latest studies on the enhancement of resveratrol bioproduction and the expression of related enzymes and discuss the contributions of different factors that contribute to maximizing production.
Biosynthesis of resveratrol in plants and microorganisms
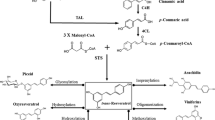
The biosynthesis pathway of resveratrol in plants is the phenylpropanoid pathway (Fig. 2). It begins with the synthesis of phenylpropanoic acids from aromatic amino acid phenylalanine (Phe) or tyrosine (Tyr) via the shikimate pathway (Lim et al. 2011). In the following steps, there are two branches. TAL/PTAL catalyzes the desamination reaction of Tyr to form p-coumaric acid. Next, 4CL combines p-coumaric acid and coenzyme A (CoA) to produce 4-coumaroyl-CoA. In the separate branch, cinnamic aid is formed as the product of Phe catalysis by PAL/PTAL (Becker et al. 2003; Jeandet et al. 2012; Mei et al. 2015). In some species such as Zea mays L., PAL can also use tyrosine as a substrate to produce p-coumaric acid (Mei et al. 2015). Cinnamic acid is then catalyzed by C4H and 4CL in either order to form 4-coumaroyl-CoA. Finally, stilbene synthase (STS) acts to catalyze the condensation reaction of successive units of malonyl CoA with 4-coumaroyl-CoA. STS plays two roles in the pathway. First, STS combines 3 malonyl CoA to form a linear polyketide molecule skeleton. Second, STS directs a C2 to C7 aldol cyclization of the polyketide to form resveratrol (Chong et al. 2009; Delaunois et al. 2009; Flores-Sanchez and Verpoorte 2009).
Biosynthesis of resveratrol via phenylpropanoid pathway in plants and engineered microbes. PAL phenylalanine ammonia lyase, PTAL phenylalanine hydroxylase, TAL tyrosine ammonia lyase, C4H cinnamate 4-hydroxylase, 4CL 4-coumarate coenzyme A ligase, STS stilbene synthase, PDA phenyl acrylic acid decarboxylase. Increasing the pool of malonyl-CoA can be realized either by over-expressing ACC (acetyl-CoA carboxylase), MatB (malonyl-CoA synthetase) and MatC (malonate carrier protein) or inhibiting fabB-fabF of fatty acid synthesis using cerulenin
Some endophytic fungi have the capability to produce high value bioactive molecules like their host-plants (Lou et al. 2013; Wang et al. 2014). This phenomenon may be the result of horizontal gene transfer (HGT) that occurred between plants and microorganisms during their long period of co-evolution (Venugopalan and Srivastava 2015). Some fungal biosynthetic enzymes are homologous to their host counterparts, consistent with the HGT hypothesis. Recently, Zhang et al. (2013b)) identified the resveratrol synthesis pathway of Alternaria sp. MG1 (isolated from grape) by the detection of intermediates and the biosynthesis of resveratrol upon addition of Phe and Tyr, similar to the phenylpropanoid pathway in plants. However, in order to confirm the metabolic flow of resveratrol, further analysis of Alternaria sp. MG1 including genomic and transcriptomic studies are required.
Key enzymes contributing to resveratrol biosynthesis
PAL/TAL
Phenylalanine ammonia lyase (PAL, EC 4.3.1.24) is a key enzyme catalyzing the first step in the phenylpropane pathway converting Phe into trans-cinnamic, and thus channeling the carbon flow to synthesize important secondary metabolites such as stilbenes, flavonoids, coumarins, pinoresinol diglucoside, and lignin (Kong 2015). Its activity may increase under biotic and abiotic stresses in vivo (Świeca 2016). PAL has been found, isolated, and characterized in a variety of species including plants, fungi, and prokaryotes, but not in animals (Kong 2015; MacDonald et al. 2016). PAL also exhibits activity in the reverse reaction that catalyzes trans-cinnamic to form Phe, which provides pools of precursor during metabolic engineering (D’cunha et al. 1994; Qi et al. 2007; Vannelli et al. 2007). PAL is present as gene families in most plants, but the number of PAL varies considerably among different species. 40–50 and 16 genes were found in potato and grapevine, respectively, but only two were found in lemon balm and raspberry (Kumar and Ellis 2001; Weitzel and Petersen 2010).
Studies have indicated that some versions of PALs possess nonoxidative deamination activity that can utilize either phenylalanine or tyrosine as substrates; these are referred to as PTAL, or phenylalanine/tyrosine ammonia lyase (EC 4.3.1.26) (Kong et al. 2014; Nishiyama et al. 2010). Tyrosine ammonia lyase (TAL, EC 4.3.1.23) can directly produce p-coumaroyl-CoA by deamination of tyrosine, reducing the number of reaction steps involving C4H in the resveratrol synthesis pathway (Ferrer et al. 2008). TAL was first discovered in Rhodobacter capsulatus by genome analysis in 2001. It has only been identified in prokaryotes, including Streptomyces globisporus and Saccharothrix espanaensis (Watts et al. 2006). PAL and TAL are members the same family of enzymes with lyase activity which can transform amino acids into α, β-unsaturated acids by the removal of ammonia. These enzymes also have a mutase activity that can form β-amino acids (Jendresen et al. 2015). Interestingly, the substrate specificity of PAL and TAL is controlled by a single amino acid. TAL is unable to use Phe as its substrate, but when its His89 residue is replaced by Phe, its substrate selectivity completely switches from tyrosine to phenylalanine; the single point mutation similarly can switch the substrate specificity of PAL (Pinto et al. 2015).
C4H
Cinnamate 4-hydroxylase (C4H, EC 1.14.13.11), the key enzyme catalyzing the second step in the phenylpropanoid pathway, combines with partner reductase (CPR) and subsequently catalyzes the hydroxylation of trans-cinnamic acid to p-coumaric acid (Luo et al. 2015). It is part of the CYP73 family of cytochrome P450 monooxygenases and is involved in the biosynthesis of a large and diverse variety of secondary metabolites (Singh et al. 2009). After it was discovered and isolated, the full-length cDNA encoding this enzyme was identified and cloned from many plants, including Jerusalem artichoke, mung bean, and alfalfa. C4H activity can be induced by a number of stresses, including chemical treatment and mechanical wound (Batard et al. 1997; Schilmiller et al. 2009).
In the phenylpropanoid metabolic pathway, C4H may be the rate-limiting step. Reducing the C4H transcriptional level leads to a 2.5-fold decrease of 4CL and an eightfold decrease of PAL activities in tobacco (Li et al. 2015a). Additionally, C4H fails to efficiently express in E. coli but can be successfully expressed in S. cerevisiae (Achnine et al. 2004; Lee et al. 2016). Because of these limitations, C4H is rarely used in the construction of a heterogenous expression system of resveratrol. Instead, strategies of introducing TAL bypass the C4H catalytic step to directly produce p-coumaric acid.
4CL
4-Coumarate coenzyme A ligase (4CL, EC 6.2.1.12) was first found and identified in 1981 (Ragg et al. 1981). Subsequently, a series of 4CLs were purified and characterized in rice, Arabidopsis thaliana, aspen, soybean, and other plant species (Li et al. 2014; Zhang et al. 2015a). The 4CL gene family is much smaller compared with STS and PAL (Chen et al. 2014a). 4CL plays an important role in catalyzing diverse aromatic substrates like p-coumaric acids, cinnamic acids, caffeic acids, and ferulic acids into corresponding CoA thioesters. Subsequently, these products participate in the biosynthesis of numerous secondary metabolites (Gao et al. 2015). The 4CL genes are classified into two types: Type I and Type II. The amino acid sequences of Type I genes are more conserved and are mainly associated with lignin accumulation and Type II regulates flavonoid metabolism (Rao et al. 2014; Sun et al. 2015). Crystal structures of 4CL showed that 4CLs exhibit different isoform distribution patterns allowing recognition of specific substrates for the synthesis of different products (Gao et al. 2015; Li and Nair 2015b).
STS
Stilbene synthase (STS,EC 2.3.1.95) is a key enzyme that catalyzes the final step in the resveratrol biosynthesis pathway. STS belongs to the polyketide synthase (PKS) super family and was first purified from stressed cell suspension cultures of peanuts (Arachis hypogaea) (Chong et al. 2009; Delaunois et al. 2009; Vannozzi et al. 2012). It is a dimer of approximately 90 kDa molecular weight with an isoelectric point of 4.8. The genes encoding STS have been identified and isolated from many plant species including grapevine, peanut, pine, and sorghum. It appears to exist as a family of closely related genes in most plants except for sorghum, in which only one STS member has been identified (Kiselev et al. 2013b). Genomic analysis of grapevine revealed 48 STS genes and at least 33 full-length sequences encoding STS protein were obtained. These sequences were classified into three clusters based on phylogenetic tree analysis and predicted amino acid sequences (Vannozzi et al. 2012). STS families share 75–90 % similarity in amino acid sequence but significantly differ in the promoter regions and in the hydrogen-binding domain adjacent to the downstream regions (Duan et al. 2015; Chong et al. 2009; Lim et al. 2011; Shi et al. 2014). STS is predominantly located in vesicles or close to the cell wall in plant tissues and is associated with resveratrol excretion (Fornara et al. 2008).
Three residues in the active site (Cys164, His303, and Asn336) are conserved in all known STSs. The active site is composed of a substrate-binding pocket, a CoA-binding tunnel, and a cyclization pocket (Flores-Sanchez and Verpoorte 2009). This specific protein conformation can catalyze diversified cyclization reactions. In flavonoid biosynthesis, the C6 and C1 of backbone are connected directly through a Claisen condensation reaction by chalcone synthase (CHS). However, for stilbenes, cyclization occurs as an aldol type of condensation between C2 and C7 with an additional decarboxylative loss of CO2 in the position of C1. The mechanistic differences between these two reactions can be explicated by their crystal structures. The presence of a thiolase fold and a cryptic thioesterase activity switches the cyclization reaction from Claisen condensation (CHS) to Aldol condensation (STS) (Ferrer et al. 2008). Overall, knowledge of resveratrol metabolism and the function of key enzymes in the pathway will lay the foundation for enhancing resveratrol biosynthesis efficiency via metabolic engineering.
Metabolic engineering—a promising method for the biosynthesis of resveratrol in vitro
In the past five decades, extensive studies of the synthesis and metabolism of resveratrol have led to multiple strategies for the production of resveratrol, including plant and microorganism resources, chemosynthesis, plant cell culture, metabolic engineering, and other methods (Almagro et al. 2015; Cai et al. 2012b; Kang et al. 2015; Krivoruchko and Nielsen 2015; Zhang et al. 2013a; Zhang et al. 2013b). Plants are the major natural resources for producing resveratrol, but its biosynthesis is limited to only a few species such as Polygonum cuspidatum, Vitis vinifera, peanut, and berry fruits. These plants contain extremely low content of resveratrol and typically produce resveratrol only under some biotic and abiotic stresses (Chung et al. 2003; Zamboni et al. 2006). In grapes, resveratrol is mainly accumulated in the skin of grape berries at a level of approximately 1.5–7.8 mg/g fresh weight (FW) (Mei et al. 2015), while the detected concentration is 2 and 50.61 μg/g dry weight (DW) in peanut and mulberry fruit, respectively (Hasan et al. 2012; Nopo-Olazabal et al. 2013; Sales and Resurreccion 2014; Shrikanta et al. 2015). Additionally, trace amounts of resveratrol have been found in some food products, such as chocolate and cocoa liquor (Counet et al. 2006; Jeffrey Hurst et al. 2008; Jerkovic et al. 2010; Pimentel et al. 2010). Although some new technologies have been developed to improve the extraction of resveratrol from plants, it remains inefficient, costly, and unable to meet the market demand (He et al. 2015; Lu et al. 2015; Soural et al. 2015).
The chemical synthesis of resveratrol may be able to achieve the goal of relatively high yields. However, it presents disadvantages in the production of byproducts during the complex process and the requirement for toxic organic solvents which can result in environmental contamination (Du et al. 2011; Fan et al. 2010).
The use of plant cell cultures has been widely applied for the production of resveratrol due to the low-cost of medium (Jeandet et al. 2014). In fact, this method has inherent advantages that allow the constitutive production of resveratrol and the yield can be increased in response to stresses (Donnez et al. 2009). A grapevine V. vinifera L. cv. Monastrell Albino cell suspension produced 5207 mg/L resveratrol when randomly methylated-beta-cyclodextrin was added as an elicitor (Martinez and Garcia 2007). The use of V. vinifera cell suspension with elicitors-cyclodextrin or cyclodextrin derivatives could allow synthesis of resveratrol in a range of 700–5000 mg/L (Kiselev 2011). However, the main limiting factor of plant cell culture is genomic instability which can give rise to silencing of some gene clusters (Venugopalan and Srivastava 2015). Moreover, products released in the culture media may be susceptible to enzyme degradation (Cai et al. 2012a).
Considering the above technologies, metabolic engineering may be the most promising and eco-friendly approach. E. coli and S. cerevisiae are popular choices for in vitro platforms for resveratrol synthesis and have advantages including fast growth rate, simple and inexpensive culture medium, and ease of genetic manipulation (Wang et al. 2016). The S. cerevisiae expression system provides a number of distinct advantages over that in E. coli. First of all, S. cerevisiae is a eukaryotic organism and thus expresses some complex enzymes (C4H) more efficiently and stably than E. coli. Additionally, S. cerevisiae exhibits higher resistance to low pH and high osmotic stress (Wang et al. 2011a). Finally, it belongs to the class of generally regarded as safe (GRAS) microorganisms, allowing its use in food fermentation to improve nutritional value (Kong 2015; Wang et al. 2016). The development of metabolic engineering tools allowed the successful reconstitution of resveratrol heterologous pathway in both E. coli and S. cerevisiae. Engineered strains can produce resveratrol with yields of 531.41 and 2300 mg/L in S. cerevisiae and E. coli, respectively (Li et al. 2015a; Lim et al. 2011). Notably, the engineered plasmid can also be transformed into other hosts in addition to the two commonly used above. Kang et al. (2015) first transformed an expression vector with 4CL and the resveratrol synthase gene (RS) into bTf (blastospore of Tremella fuciformis) by LiAc/PEG-mediated transformation. The transformant can produce 0.92 μg/g DW resveratrol upon the addition of p-coumaric acid as a substrate. Tantong et al. (2016) successfully selected a strain of cyanobacterium Synechocystis PCC 6803 to engineer a heterologous pathway of resveratrol with TAL, 4CL, and STS genes. In addition to introducing exogenous genes, gene replacement into a specific location was reported to allow successful synthesis of resveratrol in E. coli. Liu et al. (2016) utilized the site-specific replacement strategy to integrate STS, TAL, and 4CL into the loci of genes tyrR and trpED in the chromosome of E. coli BW25113 to produce 4.612 mg/L resveratrol.
Recently, there are reports of resveratrol synthesis using novel approaches. Zhang et al. (2013a) used resting cells of a non-genetically modified strain, Alternaria sp. MG1, to produce 1.376 μg/L resveratrol from phenylalanine. Another new approach is enzymatic transformation that allows synthesis of resveratrol from polydatin by glucosidase (Chen et al. 2014b; Zhang et al. 2014).
Strategies for enhancing resveratrol production and pathway enzymes
Introducing the whole metabolic pathway or specific enzyme-encoding genes
In order to improve resveratrol production and upregulate the expression of key enzymes, it is crucial to construct a resveratrol biosynthesis pathway via introduction of the entire pathway or specific genes with the initial addition of corresponding substrates (phenylalanine and tyrosine). Shin et al. (2012) obtained 3.3 mg/L resveratrol by using recombinant yeast with PAL, C4H, 4CL, and STS genes grown in YP medium (1 % yeast extract and 2 % peptone) containing 2 % galactose. The yield was further increased to 5.8 mg/L when tyrosine was added. Similarly, Wu et al. (2013) established an E. coli system by adding the whole pathway genes to obtain resveratrol at a level of 35.02 mg/L. Interestingly, the capacity of synthesizing resveratrol from different exogenous expressing systems varies considerably due to several factors, including gene resources, the choice of promoters, host strains, vector designs, addition of substrates, and the optimization of culture growth conditions (Lim et al. 2011; Shi et al. 2014; Sydor et al. 2010; Wu et al. 2013). For example, a recombinant yeast strain transformed with STS and 4CL genes produced 6 mg/L resveratrol after 144 h of incubation in SD medium (6.7 g/L yeast nitrogen base with amino acids, 20 g/L glucose) supplemented with 5 mM p-coumarate. Then after switching to rich YEPD medium (10 g/L yeast extract, 20 g/L bacterial peptone, 20 g/L glucose) with geneticin and hygromycin supplemented with 10 mM p-coumarate, the strain was able to achieve 262 mg/L resveratrol (Sydor et al. 2010). However, it may be easier to enhance the resveratrol content in plant cells (including hairy root culture) as this approach would not require genetic engineering.
Use of elicitors
Elicitors are signaling molecules that activate plant defense mechanisms and function through a massive reprogramming of gene expression that allows both the activation of resveratrol biosynthesis and the expression of related proteins (Almagro et al. 2015). Elicitors can be classified as biotic and abiotic, and the former includes polysaccharides, proteins, and oral secretions from insects and the latter includes UV-light, metal ions, and chemical compounds (Cai et al. 2013; Laura et al. 2007; Yang et al. 2015). The improvement of resveratrol synthesis in plant cell cultures triggered by elicitors is well-documented (Almagro et al. 2015; Belchi-Navarro et al. 2012; Keskin and Kunter 2008). Here, we discuss the latest findings about elicitors and their action mechanisms. The improvements in resveratrol content and the expression of key enzymes by different elicitors are presented in Table 1.
Cyclodextrins, jasmonic acid (JA), and its methyl ester, methyl jasmonate (MeJa) are effective elicitors involved in the signal transduction pathway and induce the biosynthesis of resveratrol as a component of plant defense reactions (Belchi-Navarro et al. 2012; Dubrovina and Kiselev 2011; Vuong et al. 2014). With a similar mechanism to that of cyclodextrins and JA, coronatine (including its analog indanoyl-isoleucine) and salicylic acid (SA) can also induce synthesis of secondary metabolites and upregulate the expression of pathway enzymes (Almagro et al. 2015; Kiselev et al. 2013a). Kiselev et al. (2015) found that resveratrol production increased by almost fivefold with the use of SA, resulting in increased expression of all STS genes except for STS7. Other studies revealed that elevated levels of cytoplasmic calcium could stimulate resveratrol production. A calcium ionophore, A23187 was found to increase resveratrol content and the expression of STS by influencing calcium influx (Kiselev et al. 2012).
Actually, biosynthetic gene clusters are often transcriptionally controlled by epigenetic regulation including methylation and acetylation (Venugopalan and Srivastava 2015). They are associated with gene silence and expression, thereby involving the synthesis of secondary metabolites. Recently, a study found that when 5-azacytidine (azaC) and chitosan were used to treat plant cells, the amount of resveratrol increased two to three times and the expression of biosynthetic genes upregulated greatly. AzaC is a demethylation agent that can significantly decrease the methylation level of DNA and chitosan can alter the acetylation level (Kiselev et al. 2013b; Xu et al. 2015a). Interestingly, SA also decreased the methylation of STS2 and STS10 genes when it was used to enhance resveratrol production (Kiselev et al. 2015). Additionally, UV radiance, well-known to induce resveratrol synthesis in plants, is thought to be connected with the demethylation of STS genes as well (Tyunin and Kiselev 2015).
Metal ions act as abiotic elicitors and induce biosynthesis of resveratrol in plant cell cultures. Co2+ and Ag+ at different concentrations (5, 25, and 50 μM) were employed to stimulate resveratrol production in V. vinifera, resulting in 1.3–1.5-fold increase in the yields of resveratrol for all treatments (Cai et al. 2013). Ag+ also can promote other secondary metabolites, such as tanshinones (Li et al. 2016; Zhao et al. 2010). Additionally, Cd2+ has a significant effect in improving resveratrol production and is widely used to treat grapevine cell cultures. Cd2+ can increase the yields of resveratrol 3.5–4.2-fold (Cetin et al. 2014; Çetin and Baydar 2016). However, these heavy metals had a negative effect on cell growth and inhibited cell viability remarkably except for Co2+ (Cai et al. 2013; Cetin et al. 2014). It has been demonstrated that cadmium inhibits and disturbs various physiological and biochemical processes such as respiration, photosynthesis, cell elongation, nitrogen metabolism and mineral nutrition when it is excess in plants, resulting in cell death and inhibition of growth (Çetin and Baydar 2016). Although little is known about the defense mechanism of grapevine cell cultures responding to metal iron stresses, it has been found that Cd2+ significantly increased PAL activity more than 25-fold on day 3 when compared to control cells (Cetin et al. 2014). Similarly, Ag+ treatment remarkably upregulated the expression of key enzymes of the phenylpropanoid pathway including PAL, C4H, and 4CL almost 5–30-fold (Xing et al. 2015). Therefore, we speculate that metal irons may be involved in the phenylpropanoid pathway to cause enhancement of resveratrol production.
Another approach to increase resveratrol production is feeding the plant cells precursors of resveratrol. These precursors are key components of the phenylalanine pathway, important for synthesizing resveratrol (Shin et al. 2011). However, the influence of precursors on resveratrol biosynthesis is concentration-dependent (Kiselev et al. 2013a; Shumakova et al. 2011).
Engineer of non-pathway exogenous genes
The aim of engineering a tailored vector with non-pathway genes is to elevate the intracellular concentration of precursors, regulating synthetic pathway, increasing the activity of key enzymes, and promoting resveratrol release and accumulation. Successful examples of enhancing resveratrol production using these kinds of strategies are summarized in Table 2.
The agrobacterium rolB gene is a powerful tool for induction of resveratrol synthesis in transgenic strains and cultured plant cells. The over-expression of rolB gene resulted in more than a 100-fold increase in resveratrol production in V. amurensis Rupr. callus culture compared to the control (Kiselev et al. 2007). The positive influence of rolB gene on resveratrol accumulation was demonstrated to be Ca2+dependent. Dubrovina et al. (2009) added calcium channel blockers, LaCl3, verapamil, and niflumic acid to treat rolB transgenic cultures and found that resveratrol production decreased significantly. Taken together, these reports suggest that the flux of Ca2+ and signal transduction mediated by Ca2+ are necessary to increase of resveratrol production. When VaCPK20 gene, encoding a calcium-dependent protein kinase (CDPK) was transformed into five cell lines of V. amurensis under the control of CaMV 35S promoter, it resulted in 9–68 times improvement in resveratrol production and the upregulation of the STS7 gene (Aleynova-Shumakova et al. 2014). However, not all members of the CDPK multigene family have promoting effect on resveratrol accumulation. Aleynova et al. (2015) investigated the influence of overexpression of four CDPK genes (VaCPK9, VaCPK13, VaCPK21, and VaCPK29) on resveratrol biosynthesis and found that only VaCPK29 overexpression increased resveratrol content 1.6–2.4-fold but there was no obvious effect on resveratrol biosynthesis of overexpression of the other genes. Because CDPKs are major sensors of Ca2+, the high level of intracellular calcium plays an important role in resveratrol synthesis.
In order to increase precursor levels of phenylpropanoid pathway and accelerate resveratrol synthesis, the introduction of non-pathway genes is very important (Fig. 2). Acetyl-CoA carboxylase (ACC) is a key enzyme in the acetate assimilation pathway which catalyzes acetyl-CoA into malonyl-CoA and subsequently elevates the cytosolic malonyl-CoA pool. However, the over-expression of ACC in S. cerevisiae only resulted in a slight improvement in resveratrol production, one to twofold (Li et al. 2015a; Shin et al. 2012). This may be because malonyl-CoA is also the precursor of other secondary metabolites, such as flavonoids. Enhancement of resveratrol content may require overexpression of both malonyl-CoA synthetase (matB) and malonate carrier protein (matC) with addition of malonate to increase the intracellular levels of malonyl-CoA and inhibiting fatty acid synthesis using cerulenin (Lim et al. 2011; Wang et al. 2011a). Over-expression of upstream genes DAHP-synthase (ARO4) and chorismate mutase (ARO7) in the prephenic acid pathway increased the amounts of phenylalanine precursor. Applying this strategy allowed resveratrol production to increase to 4.85 mg/L (Li et al. 2015a). Surprisingly, knockout of the phenyl acrylic acid decarboxylase (PAD1) gene did not improve the conversion of p-coumaric acid to resveratrol; further studies are required to understand this effect (Shin et al. 2011).
V-myb myeloblastosis viral oncogene homolog (MYB) transcriptional factors act as key regulators of the synthesis of phenylpropanoid-derived compounds (Liu et al. 2015). MYB proteins are classified into four groups: 1R, 2R, 3R, and 4R. 2R proteins are mainly involved in secondary metabolism and the cellular response to biotic and abiotic stresses. Höll et al. (2013) identified two 2R-MYB transcriptional factors (TFs) from grapevine, MYB14 and MYB15, which were demonstrated to specifically activate the STS promoters. Additionally, the ectopic expression of MYB15 in grapevine hairy roots resulted in increased STS expression and the accumulation of resveratrol. Moreover, the average expression of the PAL gene in MYB15 trangenic lines was 13-fold higher than that of the control. Recently, the role of MYB TFs to regulate the resveratrol biosynthetic pathway was further confirmed by Fang et al. (2014). Transient overexpression of MYB14 in young “Jingzaojing” (V. vinifera) leaf could promote the expression of STS 15-fold. They also found that MYB14 proteins could recognize the promoter of STS and bind to the Box-L5 motif (gagttggtgaga), which was identified within the promoter of STS. Therefore, MYB14 is directly involved in the regulation of the expression of STS.
Spatial organization of enzymes
Generally, most exogenous genes encoding enzymes show relatively low or no activity on their natural substrates in genetically modified strains (Lim et al. 2011). An emerging strategy to combat this problem is to co-localize pathway enzymes into multi-protein complexes through artificially scaffolded proteins or unnatural protein fusion. The advantages of this strategy include channeling intermediates and bringing active sites together, thereby increasing enzyme efficiency and improving the yield of products. For fusion protein technology, two or more genes can be connected with an amino acids linker. The activity of the fusion enzyme directly depends on the type and length of linker (Wang et al. 2011c). Zhang et al. (2015b) used a 15-amino acid flexible linker to connect the 4CL gene (Arabidopsis thaliana) and RS gene (Arachis hypogaea) and produced 80.524 mg/L resveratrol when 1 mM p-coumaric acid was supplied. Similarly, a 15-fold increase of resveratrol was produced from an expression vector encoding a fusion protein of 4CL (Arabidopsis thaliana) and STS (Vitis vinifera) connected with a three-amino acid liker (Zhang et al. 2006). An additional advantage of a fusion enzyme strategy is that it can reduce the number of vectors required in a heterologous expression system. However, protein scaffold may be more effective in increasing resveratrol synthesis and enzyme activity than the use of protein fusions. Wang and Yu (2012) cloned nine scaffold configurations consisting of the GTPase binding domain (GBD), the Src homology 3 domain (SH3), and the PSD95/DlgA/Zo-1 domain (PDZ) of different numbers into nine yeast expression vectors. These protein interaction domains were linked together by flexible nine-residue glycine-serine linkers (Dueber et al. 2009). The domains SH3 and PDZ were used to recruit 4CL and STS, respectively. By optimizing the number of repeats of these domains, they obtained a more than fivefold improvement of resveratrol (6.7 mg/L) relative to the control via using the optimal scaffold (GBD1SH32PDZ4). Additionally, they found a 2.7-fold increase over the previous results using a fusion protein approach. These results indicate that optimization of the organization of enzymes in metabolic engineering is a promising strategy allowing improved production.
Fungal interspecific interactions
The interspecific interactions between different fungi could enhance biosynthesis of phenylpropanoid metabolites. This effect may depend on the signaling pathway mediated by nitric oxide (NO) that is generated in the co-culture system. Zhao et al. (2015) co-cultured two white-rot fungi, Phellinus morii and Inonotus obliquus, and the system induced NO synthesis that was followed by an increase in the transcriptional level of PAL and 4CL. When the NO synthase (NOS) selective inhibitor aminoguanidine (AG) was added into the culture, NO production was significantly inhibited, eliminating transcription of PAL and 4CL. Additionally, they detected upregulated biosynthesis of styrylpyrone polyphenols. Notably, the function of NO is also confirmed by Vandelle et al. (2006). They found that the production of NO that was induced in endopolygalacturonase 1 (BcPG1)-treated grapevine cells activated the transcription of both PAL and STS genes. When they added 2-4-carboxyphenyl-4,4,5,5-tetramethylimidazoline-1-oxyl-3-oxide (cPTIO) into culture system to scavenge NO, reductions of the transcript accumulation of two genes were observed after a 3-h treatment. Taken together, NO involves in the expression of resveratrol synthesis genes. Therefore, we speculate that the co-culture of resveratrol producing endophytic fungus like Alternaria and other fungi species may generate a similar effect.
Conclusions
Many approaches have been developed for improving resveratrol production by over-expressing key enzymes using an in vitro platform. In plant cell cultures (including hairy root cultures), rolB gene transformation and the addition of elicitors can enhance resveratrol production. However, metabolic engineering in E. coli and S. cerevisiae is relatively complicated and requires the construction of the whole resveratrol biosynthesis pathway or introduction of specific gene with the initial addition of precursors. To increase the efficiency of recombinant microorganism, host strains, vectors, promoters and culturing conditions should be optimized. The introduction of non-pathway genes, increasing the proximity of enzymes, and promoting fungal interspecific interactions are additional strategies to enhance resveratrol production by improving enzyme activity and regulating metabolic flux.
References
Achnine L, Blancaflor EB, Rasmussen S, Dixon RA (2004) Colocalization of L-phenylalanine ammonia-lyase and cinnamate 4-hydroxylase for metabolic channeling in phenylpropanoid biosynthesis. Plant Cell 16:3098–3109. doi:10.1105/tpc.104.024406
Aleynova OA, Dubrovina AS, Manyakhin AY, Karetin YA, Kiselev KV (2015) Regulation of resveratrol production in Vitis amurensis cell cultures by calcium-dependent protein kinases. Appl Biochem Biotechnol 175:1460–1476. doi:10.1007/s12010-014-1384-2
Aleynova-Shumakova OA, Dubrovina AS, Manyakhin AY, Karetin YA, Kiselev KV (2014) VaCPK20 gene overexpression significantly increased resveratrol content and expression of stilbene synthase genes in cell cultures of Vitis amurensis Rupr. Appl Microbiol Biotechnol 98:5541–5549. doi:10.1007/s00253-014-5625-7
Almagro L, Belchi-Navarro S, Martinez-Marquez A, Bru R, Pedreno MA (2015) Enhanced extracellular production of trans-resveratrol in Vitis vinifera suspension cultured cells by using cyclodextrins and coronatine. Plant Physiol Biochem 97:361–367. doi:10.1016/j.plaphy.2015.10.025
Anthony JR, Anthony LC, Nowroozi F, Kwon G, Newman JD, Keasling JD (2009) Optimization of the mevalonate-based isoprenoid biosynthetic pathway in Escherichia coli for production of the anti-malarial drug precursor amorpha-4,11-diene. Metab Eng 11:13–19. doi:10.1016/j.ymben.2008.07.007
Batard Y, Schalk M, Pierrel M, Zimmerlin A, Durst F, Werck-Reichhart D (1997) Regulation of the cinnamate 4-hydroxylase (CYP73Al) in Jerusalem artichoke tubers in response to wounding and chemical treatments. Plant Physiol 113:951–959. doi:10.1104/pp.113.3.951
Becker JVW, Armstrong GO, van der Merwe MJ, Lambrechts MG, Vivier MA, Pretorius IS (2003) Metabolic engineering of Saccharomyces cerevisiae for the synthesis of the wine-related antioxidant resveratrol. FEMS Yeast Res 4:79–85. doi:10.1016/s1567-1356(03)00157-0
Belchi-Navarro S, Almagro L, Lijavetzky D, Bru R, Pedreno MA (2012) Enhanced extracellular production of trans-resveratrol in Vitis vinifera suspension cultured cells by using cyclodextrins and methyljasmonate. Plant Cell Rep 31:81–89. doi:10.1007/s00299-011-1141-8
Cai ZZ, Kastell A, Knorr D, Smetanska I (2012a) Exudation: an expanding technique for continuous production and release of secondary metabolites from plant cell suspension and hairy root cultures. Plant Cell Rep 31:461–477. doi:10.1007/s00299-011-1165-0
Cai ZZ, Kastell A, Speiser C, Smetanska I (2013) Enhanced resveratrol production in Vitis vinifera cell suspension cultures by heavy metals without loss of cell viability. Appl Biochem Biotechnol 171:330–340. doi:10.1007/s12010-013-0354-4
Cai ZZ, Knorr D, Smetanska I (2012b) Enhanced anthocyanins and resveratrol production in Vitis vinifera cell suspension culture by indanoyl-isoleucine, N-linolenoyl-L-glutamine and insect saliva. Enzym Microb Technol 50:29–34. doi:10.1016/j.enzmictec.2011.09.001
Çetin ES, Baydar NG (2016) Elicitor applications to cell suspension culture for production of phenolic compounds in grapevine. Tarım Bilimleri Dergisi 22:42–53
Cetin ES, Babalik Z, Hallac-Turk F, Gokturk-Baydar N (2014) The effects of cadmium chloride on secondary metabolite production in Vitis vinifera cv. cell suspension cultures. Biol Res 47:47. doi:10.1186/0717-6287-47-47
Chang MC, Eachus RA, Trieu W, Ro DK, Keasling JD (2007) Engineering Escherichia coli for production of functionalized terpenoids using plant P450s. Nat Chem Biol 3:274–277. doi:10.1038/nchembio875
Chen M, Li D, Gao ZQ, Zhang CZ (2014b) Enzymatic transformation of polydatin to resveratrol by piceid-β-d-glucosidase from Aspergillus oryzae. Bioprocess Biosyst Eng 37:1411–1416. doi:10.1007/s00449-013-1113-1
Chen HC, Song J, Wang JP, Lin YC, Ducoste J, Shuford CM, Liu J, Li Q, Shi R, Nepomuceno A, Isik F, Muddiman DC, Williams C, Sederoff RR, Chiang VL (2014a) Systems biology of lignin biosynthesis in Populus trichocarpa: heteromeric 4-coumaric acid:coenzyme A ligase protein complex formation, regulation, and numerical modeling. Plant Cell 26:876–893. doi:10.1105/tpc.113.119685
Chong JL, Poutaraud A, Hugueney P (2009) Metabolism and roles of stilbenes in plants. Plant Sci 177:143–155. doi:10.1016/j.plantsci.2009.05.012
Chung IM, Park MR, Chun JC, Yun SJ (2003) Resveratrol accumulation and resveratrol synthase gene expression in response to abiotic stresses and hormones in peanut plants. Plant Sci 164:103–109. doi:10.1016/S0168-9452(02)00341-2
Counet C, Callemien D, Sonia C (2006) Chocolate and cocoa: new sources of trans-resveratrol and trans-piceid. Food Chem 98:649–657. doi:10.1016/j.foodchem.2005.06.030
Dabauza M, Velasco L, Pazos-Navarro M, Perez-Benito E, Hellin P, Flores P, Gomez-Garay A, Martinez MC, Lacasa A (2015) Enhanced resistance to Botrytis cinerea in genetically-modified Vitis vinifera L. plants over-expressing the grapevine stilbene synthase gene. Plant Cell Organ Cult 120:229–238. doi:10.1007/s11240-014-0598-x
D’cunha GB, Satyanarayan V, Nair PM (1994) Novel direct synthesis of L-phenylalanine methyl ester by using Rhodotorula glutinis phenylalanine ammonia lyase in an organic-aqueous biphasic system. Enzym Microb Technol 16:318–322. doi:10.1016/0141-0229(94)90173-2
Delaunois B, Cordelier S, Conreux A, Clement C, Jeandet P (2009) Molecular engineering of resveratrol in plants. Plant Biotechnol J 7:2–12. doi:10.1111/j.1467-7652.2008.00377.x
Donnez D, Jeandet P, Clement C, Courot E (2009) Bioproduction of resveratrol and stilbene derivatives by plant cells and microorganisms. Trends Biotechnol 27:706–713. doi:10.1016/j.tibtech.2009.09.005
Du J, Shao ZY, Zhao HM (2011) Engineering microbial factories for synthesis of value-added products. J Ind Microbiol Biot 38:873–890. doi:10.1007/s10295-011-0970-3
Duan D, Halter D, Baltenweck R, Tisch C, Troster V, Kortekamp A, Hugueney P, Nick P (2015) Genetic diversity of stilbene metabolism in Vitis sylvestris. J Exp Bot 66:3243–3257. doi:10.1093/jxb/erv137
Dubrovina AS, Kiselev KV (2011) Effect of long-term cultivation on resveratrol accumulation in a high-producing cell culture of Vitis amurensis. Acta Physiol Plant 34:1101–1106. doi:10.1007/s11738-011-0907-5
Dubrovina AS, Kiselev KV, Veselova MV, Isaeva GA, Fedoreyev SA, Zhuravlev YN (2009) Enhanced resveratrol accumulation in rolB transgenic cultures of Vitis amurensis correlates with unusual changes in CDPK gene expression. J Plant Physiol 166:1194–1206. doi:10.1016/j.jplph.2009.01.006
Dueber JE, Wu GC, Malmirchegini GR, Moon TS, Petzold CJ, Ullal AV, Prather KL, Keasling JD (2009) Synthetic protein scaffolds provide modular control over metabolic flux. Nat Biotechnol 27:753–759. doi:10.1038/nbt.1557
Engels B, Dahm P, Jennewein S (2008) Metabolic engineering of taxadiene biosynthesis in yeast as a first step towards Taxol (paclitaxel) production. Metab Eng 10:201–206. doi:10.1016/j.ymben.2008.03.001
Fan EG, Zhang K, Zhu MZ, Wang Q (2010) Obtaining resveratrol: from chemical synthesis to biotechnological production. Mini-Rev Org Chem 7:272–281. doi:10.2174/157019310792246454
Fang LC, Hou YL, Wang LJ, Xin HP, Wang N, Li SH (2014) Myb14, a direct activator of STS, is associated with resveratrol content variation in berry skin in two grape cultivars. Plant Cell Rep 33:1629–1640. doi:10.1007/s00299-014-1642-3
Ferrer JL, Austin MB, Stewart Jr C, Noel JP (2008) Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant Physiol Biochem 46:356–370. doi:10.1016/j.plaphy.2007.12.009
Flores-Sanchez IJ, Verpoorte R (2009) Plant polyketide synthases: a fascinating group of enzymes. Plant Physiol Biochem 47:167–174. doi:10.1016/j.plaphy.2008.11.005
Fornara V, Onelli E, Sparvoli F, Rossoni M, Aina R, Marino G, Citterio S (2008) Localization of stilbene synthase in Vitis vinifera L. during berry development. Protoplasma 233:83–93. doi:10.1007/s00709-008-0309-8
Gao S, HN Y, RX X, Cheng AX, Lou HX (2015) Cloning and functional characterization of a 4-coumarate CoA ligase from liverwort Plagiochasma appendiculatum. Phytochemistry 111:48–58. doi:10.1016/j.phytochem.2014.12.017
Giovinazzo G, Ingrosso I, Paradiso A, Gara LD, Santino A (2012) Resveratrol biosynthesis: plant metabolic engineering for nutritional improvement of food. Plant Foods Hum Nutr 67:191–199. doi:10.1007/s11130-012-0299-8
Gonzalez-Candelas L, Gil JV, Lamuela-Raventos RM, Ramon D (2000) The use of transgenic yeasts expressing a gene encoding a glycosyl-hydrolase as a tool to increase resveratrol content in wine. Int J Food Microbiol 59:179–183. doi:10.1016/S0168-1605(00)00354-8
Halls C, Yu O (2008) Potential for metabolic engineering of resveratrol biosynthesis. Trends Biotechnol 26:77–81. doi:10.1016/j.tibtech.2007.11.002
Hasan MM, Cha M, Bajpai VK, Baek KH (2012) Production of a major stilbene phytoalexin, resveratrol in peanut (Arachis hypogaea) and peanut products: a mini review. Rev Environ Sci Biotechnol 12:209–221. doi:10.1007/s11157-012-9294-7
He S, Shi Y, Zhang SY, Zhang ZY (2015) Extraction of resveratrol and emondin from Polygonum cuspidatum by supercritical CO2 with different solubilizers. Afr J Pharm Pharmacol 9:12–18. doi:10.5897/ajpp2013.3816
Höll J, Vannozzi A, Czemmel S, D’Onofrio C, Walker AR, Rausch T, Lucchin M, Boss PK, Dry IB, Bogs J (2013) The R2R3-MYB transcription factors MYB14 and MYB15 regulate stilbene biosynthesis in Vitis vinifera. Plant Cell 25:4135–4149. doi:10.1105/tpc.113.117127
Jeandet P, Clément C, Courot E (2014) Resveratrol production at large scale using plant cell suspensions. Eng Life Sci 14:622–632. doi:10.1002/elsc.201400022
Jeandet P, Delaunois B, Aziz A, Donnez D, Vasserot Y, Cordelier S, Courot E (2012) Metabolic engineering of yeast and plants for the production of the biologically active hydroxystilbene, resveratrol. J Biomed Biotechnol 2012:1–14. doi:10.1155/2012/579089
Jeffrey Hurst W, Glinski JA, Miller KB, Apgar J, Davey MH, Stuart DA (2008) Survey of the trans-resveratrol and trans-piceid content of cocoa-containing and chocolate products. J Agric Food Chem 56:8374–8378. doi:10.1021/jf801297w
Jendresen CB, Stahlhut SG, Li M, Gaspar P, Siedler S, Forster J, Maury J, Borodina I, Nielsen AT (2015) Novel highly active and specific tyrosine ammonia-lyases from diverse origins enable enhanced production of aromatic compounds in bacteria and yeast. Appl Environ Microbiol 81:4458–4476. doi:10.1128/AEM.00405-15
Jerkovic V, Bröhan M, Monnart E, Nguyen F, Nizet S, Collin S (2010) Stilbenic profile of cocoa liquors from different origins determined by RP-HPLC-APCI (+)-MS/MS. Detection of a new resveratrol hexoside. J Agric Food Chem 58:7067–7074. doi:10.1021/jf101114c
Kang LZ, Li Q, Lin JF, Guo LQ (2015) Biosynthesis of resveratrol in blastospore of the macrofungus tremella fuciformis. Mol Biotechnol 57:675–684. doi:10.1007/s12033-015-9858-1
Keskin N, Kunter B (2008) Production of trans-resveratrol in “cabernet sauvignon” (Vitis vinifera L.) callus culture in response to ultraviolet-C irradiation. Vitis 47:193–196
Kiselev KV (2011) Perspectives for production and application of resveratrol. Appl Microbiol Biotechnol 90:417–425. doi:10.1007/s00253-011-3184-8
Kiselev KV, Dubrovina AS, Veselova MV, Bulgakov VP, Fedoreyev SA, Zhuravlev YN (2007) The rolB gene-induced overproduction of resveratrol in Vitis amurensis transformed cells. J Biotechnol 128:681–692. doi:10.1016/j.jbiotec.2006.11.008
Kiselev KV, Shumakova OA, Manyakhin AY (2013a) Effects of the calmodulin antagonist W7 on resveratrol biosynthesis in Vitis amurensis Rupr. Plant Mol Biol Report 31:1569–1575. doi:10.1007/s11105-013-0620-1
Kiselev KV, Shumakova OA, Manyakhin AY, Mazeika AN (2012) Influence of calcium influx induced by the calcium ionophore, A23187on resveratrol content and the expression of CDPK and STS genes in the cell cultures of Vitis amurensis. Plant Growth Regul 68:371–381. doi:10.1007/s10725-012-9725-z
Kiselev KV, Tyunin AP, Karetin YA (2015) Salicylic acid induces alterations in the methylation pattern of the VaSTS1, VaSTS2, and VaSTS10 genes in Vitis amurensis Rupr. Cell cultures. Plant Cell Rep 34:311–320. doi:10.1007/s00299-014-1708-2
Kiselev KV, Tyunin AP, Zhuravlev YN (2013b) Involvement of DNA methylation in the regulation of STS10 gene expression in Vitis amurensis. Planta 237:933–941. doi:10.1007/s00425-012-1806-8
Klein-Marcuschamer D, Yadav VG, Ghaderi A, Stephanopoulos GN (2010) De Novo metabolic engineering and the promise of synthetic DNA. Adv Biochem Eng Biot 120:101–131. doi:10.1007/10_2009_52
Kong JQ (2015) Phenylalanine ammonia-lyase, a key component used for phenylpropanoids production by metabolic engineering. RSC Adv 5:62587–62603. doi:10.1039/c5ra08196c
Kong JQ, Lu D, Wang ZB (2014) Molecular cloning and yeast expression of cinnamate 4-hydroxylase from Ornithogalum saundersiae baker. Molecules 19:1608–1621. doi:10.3390/molecules19021608
Krivoruchko A, Nielsen J (2015) Production of natural products through metabolic engineering of Saccharomyces cerevisiae. Curr Opin Biotech 35:7–15. doi:10.1016/j.copbio.2014.12.004
Kumar A, Ellis BE (2001) The phenylalanine ammonia-lyase gene family in raspberry. Structure, expression, and evolution. Plant Physiol 127:230–239. doi:10.1104/pp.127.1.230
Lancon A, Kaminski J, Tili E, Michaille JJ, Latruffe N (2012) Control of microRNA expression as a new way for resveratrol to deliver its beneficial effects. J Agric Food Chem 60:8783–8789. doi:10.1021/jf301479v
Laura R, Franceschetti M, Ferri M, Tassoni A, Bagni N (2007) Resveratrol production in Vitis vinifera cell suspensions treated with several elicitors. Caryologia 60:169–171. doi:10.1080/00087114.2007.10589568
Lee D, Lloyd NDR, Pretorius IS, Borneman AR (2016) Heterologous production of raspberry ketone in the wine yeast Saccharomyces cerevisiae via pathway engineering and synthetic enzyme fusion. Microb Cell Factories 15:1–7. doi:10.1186/s12934-016-0446-2
Leifer A, Barberio DM (2016) Direct ingestion method for enhancing production and bioavailability of resveratrol and other phytoalexins in Vitis vinifera. Med Hypotheses 88:1–5. doi:10.1016/j.mehy.2015.12.008
Li B, Wang BQ, Li HY, Peng L, Ru M, Liang ZS, Yan XJ, Zhu YH (2016) Establishment of Salvia castanea Diels F. tomentosa Stib. Hairy root cultures and the promotion of tanshinone accumulation and gene expression with Ag+, methyl jasmonate, and yeast extract elicitation. Protoplasma 253:87–100. doi:10.1007/s00709-015-0790-9
Li MJ, Kildegaard KR, Chen Y, Rodriguez A, Borodina I, Nielsen J (2015a) De novo production of resveratrol from glucose or ethanol by engineered Saccharomyces cerevisiae. Metab Eng 32:1–11. doi:10.1016/j.ymben.2015.08.007
Li Z, Nair SK (2015b) Structural basis for specificity and flexibility in a plant 4-coumarate: CoA ligase. Structure 23:2032–2042. doi:10.1016/j.str.2015.08.012
Li ZB, Li CF, Li J, Zhang YS (2014) Molecular cloning and functional characterization of two divergent 4-coumarate: coenzyme A ligases from kudzu (Pueraria lobata). Biol Pharm Bull 37:113–122. doi:10.1248/bpb.b13-00633
Lim CG, Fowler ZL, Hueller T, Schaffer S, Koffas MAG (2011) High-yield resveratrol production in engineered Escherichia coli. Appl Environ Microbiol 77:3451–3460. doi:10.1128/AEM.02186-10
Lin YH, Jain R, Yan YJ (2014) Microbial production of antioxidant food ingredients via metabolic engineering. Curr Opin Biotechnol 26:71–78. doi:10.1016/j.copbio.2013.10.004
Liu JY, Osbourn A, Ma P (2015) MYB transcription factors as regulators of phenylpropanoid metabolism in plants. Mol Plant 8:689–708. doi:10.1016/j.molp.2015.03.012
Liu XL, Lin J, HF H, Zhou B, Zhu BQ (2016) De novo biosynthesis of resveratrol by site-specific integration of heterologous genes in Escherichia coli. FEMS Microbiol Lett. doi:10.1093/femsle/fnw061
Lou JF, LY F, Peng YL, Zhou LG (2013) Metabolites from Alternaria fungi and their bioactivities. Molecules 18:5891–5935. doi:10.3390/molecules18055891
Lu Q, Zhao Q, QW Y, Feng YQ (2015) Use of pollen solid-phase extraction for the determination of trans-resveratrol in peanut oils. J Agr Food Chem 63:4771–4776. doi:10.1021/jf505938w
Luo YZ, Li BZ, Liu D, Zhang L, Chen Y, Jia B, Zeng BX, Zhao HM, Yuan YJ (2015) Engineered biosynthesis of natural products in heterologous hosts. Chem Soc Rev 44:5265–5290. doi:10.1039/C5CS00025D
MacDonald MC, Arivalagan P, Barre DE, MacInnis JA, D’Cunha GB (2016) Rhodotorula glutinis Phenylalanine/tyrosine ammonia lyase enzyme catalyzed synthesis of the methyl ester of para-hydroxycinnamic acid and its potential antibacterial activity. Front Microbiol 7:1–11. doi:10.3389/fmicb.2016.00281
Martinez RB, Garcia MAP (2007) Method for the production of resveratrol in cell cultures. Google Patents, United States
Martinez-Marquez A, Morante-Carriel JA, Ramirez-Estrada K, Cusido RM, Palazon J, Bru-Martinez R (2016) Production of highly bioactive resveratrol analogues pterostilbene and piceatannol in metabolically engineered grapevine cell cultures. Plant Biotechnol J 10:1–13. doi:10.1111/pbi.12539
Mei YZ, Liu RX, Wang DP, Wang X, Dai CC (2015) Biocatalysis and biotransformation of resveratrol in microorganisms. Biotechnol Lett 37:9–18. doi:10.1007/s10529-014-1651-x
Nishiyama Y, Yun CS, Matsuda F, Sasaki T, Saito K, Tozawa Y (2010) Expression of bacterial tyrosine ammonia-lyase creates a novel p-coumaric acid pathway in the biosynthesis of phenylpropanoids in Arabidopsis. Planta 232:209–218. doi:10.1007/s00425-010-1166-1
Nopo-Olazabal C, Hubstenberger J, Nopo-Olazabal L, Medina-Bolivar F (2013) Antioxidant activity of selected stilbenoids and their bioproduction in hairy root cultures of muscadine grape (Vitis rotundifolia Michx.). J Agr Food Chem 61:11744–11758. doi:10.1021/jf400760k
Pimentel FA, Nitzke JA, Klipel CB, Jong EV (2010) Chocolate and red wine - a comparison between flavonoids content. Food Chem 120:109–112. doi:10.1016/j.foodchem.2009.09.078
Pinto GP, Ribeiro AJM, Ramos MJ, Fernandes PA, Toscano M, Russo N (2015) New insights in the catalytic mechanism of tyrosine ammonia-lyase given by QM/MM and QM cluster models. Arch Biochem Biophys 582:107–115. doi:10.1016/j.abb.2015.03.002
Qi WW, Vannelli T, Breinig S, Ben-Bassat A, Gatenby AA, Haynie SL, Sariaslani FS (2007) Functional expression of prokaryotic and eukaryotic genes in Escherichia coli for conversion of glucose to p-hydroxystyrene. Metab Eng 9:268–276. doi:10.1016/j.ymben.2007.01.002
Ragg H, Kuhn DN, Hahlbrock K (1981) Coordinated regulation of 4-coumarate: CoA ligase and phenylalanine ammonia-lyase mRNAs in cultured plant cells. J Biol Chem 256:10061–10065
Rao GD, Pan X, Xu F, Zhang YZ, Cao S, Jiang XN, Lu H (2014) Divergent and overlapping function of five 4-coumarate/coenzyme A ligases from Populus tomentosa. Plant Mol Biol Rep 33:841–854. doi:10.1007/s11105-014-0803-4
Ruhmann S, Pfeiffer J, Brunner P, Szankowski I, Fischer TC, Forkmann G, Treutter D (2013) Induction of stilbene phytoalexins in grapevine (Vitis vinifera) and transgenic stilbene synthase-apple plants (Malus domestica) by a culture filtrate of Aureobasidium pullulans. Plant Physiol Biochem 72:62–71. doi:10.1016/j.plaphy.2013.03.011
Sales JM, Resurreccion AV (2014) Resveratrol in peanuts. Crit Rev Food Sci Nutr 54:734–770. doi:10.1080/10408398.2011.606928
Schilmiller AL, Stout J, Weng JK, Humphreys J, Ruegger MO, Chapple C (2009) Mutations in the cinnamate 4-hydroxylase gene impact metabolism, growth and development in Arabidopsis. Plant J 60:771–782. doi:10.1111/j.1365-313X.2009.03996.x
Shi JL, He MY, Cao JL, Wang H, Ding JH, Jiao YT, Li RM, He J, Wang D, Wang YJ (2014) The comparative analysis of the potential relationship between resveratrol and stilbene synthase gene family in the development stages of grapes (Vitis quinquangularis and Vitis vinifera). Plant Physiol Biochem 74:24–32. doi:10.1016/j.plaphy.2013.10.021
Shin SY, Han NS, Park YC, Kim MD, Seo JH (2011) Production of resveratrol from p-coumaric acid in recombinant Saccharomyces cerevisiae expressing 4-coumarate:coenzyme A ligase and stilbene synthase genes. Enzym Microb Technol 48:48–53. doi:10.1016/j.enzmictec.2010.09.004
Shin SY, Jung SM, Kim MD, Han NS, Seo JH (2012) Production of resveratrol from tyrosine in metabolically engineered Saccharomyces cerevisiae. Enzym Microb Technol 51:211–216. doi:10.1016/j.enzmictec.2012.06.005
Shrikanta A, Kumar A, Govindaswamy V (2015) Resveratrol content and antioxidant properties of underutilized fruits. J Food Sci Tech 52:383–390. doi:10.1007/s13197-013-0993-z
Shumakova OA, Manyakhin AY, Kiselev KV (2011) Resveratrol content and expression of phenylalanine ammonia-lyase and stilbene synthase genes in cell cultures of Vitis amurensis treated with coumaric acid. Appl Biochem Biotechnol 165:1427–1436. doi:10.1007/s12010-011-9361-5
Singh K, Kumar S, Rani A, Gulati A, Ahuja PS (2009) Phenylalanine ammonia-lyase (PAL) and cinnamate 4-hydroxylase (C4H) and catechins (flavan-3-ols) accumulation in tea. Funct Integr Genomics 9:125–134. doi:10.1007/s10142-008-0092-9
Soural I, Vrchotová N, Tříska J, Balík J, Horník Š, Cuřínová P, Sýkora J (2015) Various extraction methods for obtaining stilbenes from grape cane of Vitis vinifera L. Molecules 20:6093–6112. doi:10.3390/molecules20046093
Sun HY, Guo K, Feng SQ, Zou WH, Li Y, Fan CF, Peng LC (2015) Positive selection drives adaptive diversification of the 4-coumarate: CoA ligase (4CL) gene in angiosperms. Ecol Evol 5:3413–3420. doi:10.1002/ece3.1613
Suzuki S, Koeduka T, Sugiyama A, Yazaki K, Umezawa T (2014) Microbial production of plant specialized metabolites. Plant Biol 31:465–482. doi:10.5511/plantbiotechnology.14.1003a
Świeca M (2016) Hydrogen peroxide treatment and the phenylpropanoid pathway precursors feeding improve phenolics and antioxidant capacity of quinoa sprouts via an induction of L-tyrosine and L-phenylalanine ammonia-lyases activities. J Chem 2016:1–7. doi:10.1155/2016/1936516
Sydor T, Schaffer S, Boles E (2010) Considerable increase in resveratrol production by recombinant industrial yeast strains with use of rich medium. Appl Environ Microbiol 76:3361–3363. doi:10.1128/AEM.02796-09
Takaoka M (1940) Of the phenolic substances of white hellebore (Veratrum grandiflorum Loes. fil.). J Faculty Sci Hokkaido Imperial University 3:1–16. doi:10.1246/nikkashi1921.60.1261
Tantong S, Incharoensakdi A, Sirikantaramas S, Lindblad P (2016) Potential of Synechocystis PCC 6803 as a novel cyanobacterial chassis for heterologous expression of enzymes in the trans-resveratrol biosynthetic pathway. Protein Expr Purif 121:163–168. doi:10.1016/j.pep.2016.01.020
Tassoni A, Fornale S, Franceschetti M, Musiani F, Michael AJ, Perry B, Bagni N (2005) Jasmonates and Na-orthovanadate promote resveratrol production in Vitis vinifera cv. Barbera cell cultures. New Phytol 166:895–905. doi:10.1111/j.1469-8137.2005.01383.x
Tyunin AP, Kiselev KV (2015) Alternations in VaSTS gene cytosine methylation and t-resveratrol production in response to UV-C irradiation in Vitis amurensis Rupr. cells. Plant Cell Tiss Org 124:33–45. doi:10.1007/s11240-015-0872-6
Vandelle E, Poinssot B, Wendehenne D, Bentejac M, Pugin A (2006) Integrated signaling network involving calcium, nitric oxide, and active oxygen species but not mitogen-activated protein kinases in BcPG1-elicited grapevine defenses. Mol Plant-Microbe Interact 19:429–440. doi:10.1094/MPMI-19-0429
Vannelli T, Qi WW, Sweigard J, Gatenby AA, Sariaslani FS (2007) Production of p-hydroxycinnamic acid from glucose in Saccharomyces cerevisiae and Escherichia coli by expression of heterologous genes from plants and fungi. Metab Eng 9:142–151. doi:10.1016/j.ymben.2006.11.001
Vannozzi A, Dry IB, Fasoli M, Zenoni S, Lucchin M (2012) Genome-wide analysis of the grapevine stilbene synthase multigenic family: genomic organization and expression profiles upon biotic and abiotic stresses. BMC Plant Biol 12:1–22. doi:10.1186/1471-2229-12-130
Venugopalan A, Srivastava S (2015) Endophytes as in vitro production platforms of high value plant secondary metabolites. Biotechnol Adv 33:873–887. doi:10.1016/j.biotechadv.2015.07.004
Vuong TV, Franco C, Zhang W (2014) Treatment strategies for high resveratrol induction in Vitis vinifera L. cell suspension culture. Biol Reprod 1-2:15–21. doi:10.1016/j.btre.2014.04.002
Wang J, Guleria S, Koffas MA, Yan YJ (2016) Microbial production of value-added nutraceuticals. Curr Opin Biotechnol 37:97–104. doi:10.1016/j.copbio.2015.11.003
Wang JH, Cox DG, Ding WJ, Huang GH, Lin YC, Li CY (2014) Three new resveratrol derivatives from the mangrove endophytic fungus Alternaria sp. Mar Drugs 12:2840–2850. doi:10.3390/md12052840
Wang YC, Yu O (2012) Synthetic scaffolds increased resveratrol biosynthesis in engineered yeast cells. J Biotechnol 157:258–260. doi:10.1016/j.jbiotec.2011.11.003
Wang YC, Chen S, Yu O (2011a) Metabolic engineering of flavonoids in plants and microorganisms. Appl Microbiol Biotechnol 91:949–956. doi:10.1007/s00253-011-3449-2
Wang YC, Halls C, Zhang J, Matsuno M, Zhang YS, Yu O (2011b) Stepwise increase of resveratrol biosynthesis in yeast Saccharomyces cerevisiae by metabolic engineering. Metab Eng 13:455–463. doi:10.1016/j.ymben.2011.04.005
Wang YC, Yi H, Wang M, Yu O, Jez JM (2011c) Structural and kinetic analysis of the unnatural fusion protein 4-coumaroyl-CoA ligase::stilbene synthase. J Am Chem Soc 133:20684–20687. doi:10.1021/ja2085993
Watts KT, Mijts BN, Lee PC, Manning AJ, Schmidt-Dannert C (2006) Discovery of a substrate selectivity switch in tyrosine ammonia-lyase, a member of the aromatic amino acid lyase family. Chem Biol 13:1317–1326. doi:10.1016/j.chembiol.2006.10.008
Weitzel C, Petersen M (2010) Enzymes of phenylpropanoid metabolism in the important medicinal plant Melissa officinalis L. Planta 232:731–742. doi:10.1007/s00425-010-1206-x
Whitlock NC, Baek SJ (2012) The anticancer effects of resveratrol: modulation of transcription factors. Nutr Cancer 64:493–502. doi:10.1080/01635581.2012.667862
Wu JJ, Liu P, Fan YM, Bao H, GC D, Zhou JW, Chen J (2013) Multivariate modular metabolic engineering of Escherichia coli to produce resveratrol from L-tyrosine. J Biotechnol 167:404–411. doi:10.1016/j.jbiotec.2013.07.030
Xing BC, Yang DF, Guo WL, Liang ZS, Yan XJ, Zhu YH, Liu Y (2015) Ag+ as a more effective elicitor for production of tanshinones than phenolic acids in Salvia miltiorrhiza hairy roots. Molecules 20:309–324. doi:10.3390/molecules20010309
Xu A, Zhan JC, Huang WD (2015a) Combined elicitation of chitosan and ultraviolet C enhanced stilbene production and expression of chitinase and β-1,3-glucanase in Vitis vinifera cell suspension cultures. Plant Cell Tiss Org 124:105–117. doi:10.1007/s11240-015-0879-z
Xu A, Zhan JC, Huang WD (2015b) Oligochitosan and sodium alginate enhance stilbene production and induce defense responses in Vinifera cell suspension. Acta Physiol Plant 37:144–157. doi:10.1007/s11738-015-1900-1
Yang TH, Fang LL, Nopo-Olazabal C, Condori J, Nopo-Olazabal L, Balmaceda C, Medina-Bolivar F (2015) Enhanced production of resveratrol, piceatannol, arachidin-1, and arachidin-3 in hairy root cultures of peanut co-treated with methyl jasmonate and cyclodextrin. J Agric Food Chem 63:3942–3950. doi:10.1021/jf5050266
Zamboni A, Vrhovsek U, Kassemeyer HH, Mattivi F, Velasco R (2006) Elicitor-induced resveratrol production in cell cultures of different grape genotypes (Vitis spp.). Vitis 45:63–68
Zhang CH, Ma T, Luo WC, JM X, Liu JQ, Wan DS (2015a) Identification of 4CL genes in desert poplars and their changes in expression in response to salt stress. Genes 6:901–917. doi:10.3390/genes6030901
Zhang EH, Guo XF, Meng ZF, Wang J, Sun J, Yao X, Xun H (2015b) Construction, expression, and characterization of Arabidopsis thaliana 4CL and Arachis hypogaea RS fusion gene 4CL::RS in Escherichia coli. World J Microbiol Biotechnol 31:1379–1385. doi:10.1007/s11274-015-1889-z
Zhang J, Wang D, Pan JC, Wang J, Zhao HB, Li Q, Zhou XH (2014) Efficient resveratrol production by immobilized β-glucosidase on cross-linked chitosan microsphere modified by l-lysine. J Mol Catal B-Enzym 104:29–34. doi:10.1016/j.molcatb.2014.03.003
Zhang JH, Shi JL, Liu YL (2013a) Bioconversion of resveratrol using resting cells of non-genetically modified Alternaria sp. Biotechnol Appl Bioc 10:236–243. doi:10.1002/bab.1060
Zhang JH, Shi JL, Liu YL (2013b) Substrates and enzyme activities related to biotransformation of resveratrol from phenylalanine by Alternaria sp. MG1. Appl Microbiol Biotechnol 97:9941–9954. doi:10.1007/s00253-013-5212-3
Zhang YS, Li SZ, Li J, Pan XQ, Cahoon RE, Jaworski JG, Wang XM, Jez JM, Chen F, Yu O (2006) Using unnatural protein fusions to engineer resveratrol biosynthesis in yeast and mammalian cells. J Am Chem Soc 128:13030–13031. doi:10.1021/ja0622094
Zhao JL, Zhou LG, Wu JY (2010) Effects of biotic and abiotic elicitors on cell growth and tanshinone accumulation in Salvia miltiorrhiza cell cultures. Appl Microbiol Biotechnol 87:137–144. doi:10.1007/s00253-010-2443-4
Zhao YX, Xi Q, Xu Q, He MH, Ding JN, Dai YC, Keller NP, Zheng WF (2015) Correlation of nitric oxide produced by an inducible nitric oxide synthase-like protein with enhanced expression of the phenylpropanoid pathway in Inonotus obliquus cocultured with Phellinus morii. Appl Microbiol Biotechnol 99:4361–4372. doi:10.1007/s00253-014-6367-2
Acknowledgment
This review is supported by the National Key Technology R&D Program (No. 2015BAD16B02), the National Natural Science Fund (Grant No. 31471718), the Agriculture Department of China (Grant No. CARS-30), and the Northwestern Polytechnical University (No. 3102014JCQ15011 and No. 3102014GEKY1010).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Lu, Y., Shao, D., Shi, J. et al. Strategies for enhancing resveratrol production and the expression of pathway enzymes. Appl Microbiol Biotechnol 100, 7407–7421 (2016). https://doi.org/10.1007/s00253-016-7723-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7723-1