Abstract
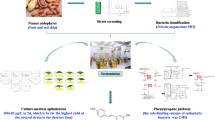
To identify the substrates and enzymes related to resveratrol biosynthesis in Alternaria sp. MG1, different substrates were used to produce resveratrol, and their influence on resveratrol production was analyzed using high performance liquid chromatography (HPLC). Formation of resveratrol and related intermediates was identified using mass spectrum. During the biotransformation, activities of related enzymes, including phenylalanine ammonia-lyase (PAL), trans-cinnamate 4-hydroxylase (C4H), and 4-coumarate-CoA ligase (4CL), were analyzed and tracked. The reaction system contained 100 mL 0.2 mol/L phosphate buffer (pH 6.5), 120 g/L Alternaria sp. MG1 cells, 0.1 g/L MgSO4, and 0.2 g/L CaSO4 and different substrates according to the experimental design. The biotransformation was carried out for 21 h at 28 °C and 120 rpm. Resveratrol formation was identified when phenylalanine, tyrosine, cinnamic acid, and p-coumaric acid were separately used as the only substrate. Accumulation of cinnamic acid, p-coumaric acid, and resveratrol and the activities of PAL, C4H, and 4CL were identified and changed in different trends during transformation with phenylalanine as the only substrate. The addition of carbohydrates and the increase of phenylalanine concentration promoted resveratrol production and yielded the highest value (4.57 μg/L) when 2 g/L glucose, 1 g/L cyclodextrin, and phenylalanine (4.7 mmol/L) were used simultaneously.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Resveratrol (3,5,4′-trihydroxystilbene) is a plant stilbene having multiple functions and is in high demand in food and medical industries (Bradamante et al. 2004; Baur and Sinclair 2006; Dudley et al. 2009; Gresele et al. 2011; Howitz et al. 2003; Lee et al. 2011). Biosynthesis of resveratrol using plant cells and genetically modified microorganisms is recently explored to overcome the difficulties caused by shortage of plant sources from which resveratrol is extracted (Beihadj et al. 2008; Kiselev et al. 2013; Donnez et al. 2011). However, these techniques normally need long time or expensive precursors in cultivation. For example, 120 h is required for resveratrol production using plant cell cultures (Lucas-Abellan et al. 2007; Lijavetzky et al. 2008), and p-coumaric acid is needed to produce resveratrol using genetically modified organisms (Watts et al. 2006; Shin et al. 2011). Nongenetically modified Alternaria sp. MG1 produced resveratrol in a glucose medium (Shi et al. 2012; Lou et al. 2013) and a simple medium only containing phosphate buffer and phenylalanine within 21 h (Zhang et al. 2013), hence showing potential in resveratrol production with low cost and short time. However, the resveratrol production of this strain was only 422 μg/L in the glucose medium and 1.38 μg/L in the phenylalanine medium (Shi et al. 2012; Zhang et al. 2013), much lower than that produced using grape cell suspension cultures (5 g/L; Lijavetzky et al. 2008) and genetically modified yeast (5.8 mg/L; Shin et al. 2011) and Escherichia coli (100 mg/L; Watts et al. 2006). Metabolic regulation based on resveratrol biosynthesis pathway is sought to enhance the resveratrol production in cultures using Alternaria sp. MG1 since this method has been widely proven as the efficient way to improve metabolite accumulation by microorganisms.
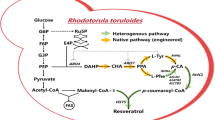
The resveratrol biosynthesis pathway in plants has been identified and widely accepted as the phenylpropanoid pathway and summarized as shown in Fig. 1 (Beekwilder et al. 2006; Jeandet et al. 2012; Watts et al. 2006). In the phenylpropanoid pathway, phenylalanine and tyrosine are converted into resveratrol through many steps with cinnamic acid, p-coumaric acid, cinnamoyl-CoA, and 4-coumaroyl-CoA as intermediates. Phenylalanine/tyrosine ammonia lyase (PAL/TAL), trans-cinnamate 4-hydroxylase (C4H), 4-coumarate-CoA ligase (4CL), and stilbene synthase (STS) are the key enzymes that determine resveratrol biosynthesis by the phenylpropanoid pathway (Ro and Douglas 2004; Shumakova et al. 2011; Marienhagen and Bott 2013). Specifically, PAL converts phenylalanine to cinnamic acid, TAL converts tyrosine to p-coumaric acid, C4H converts cinnamic acid to p-coumaric acid and cinnamoyl-CoA to p-coumaroyl-CoA, and 4CL converts cinnamic acid to cinnamoyl-CoA and p-coumaric acid to p-coumaroyl-CoA with the aid of ATP and coenzyme A. Finally, STS converts p-coumaroyl-CoA and three molecules of malonyl-CoA to resveratrol (Jeandet et al. 2012) (Fig. 1). Based on this pathway, genetically modified resveratrol-producing microorganisms have been successfully constructed by introducing the entire pathway or several plant-specific genes (Becker et al. 2003; Dai et al. 2012; Liu et al. 2011). Comparatively, the latter case is easier to achieve and has been extensively studied using p-coumaric acid as the substrate (Jeandet et al. 2012; Lim et al. 2011; Wang and Yu 2012). However, the resveratrol biosynthesis pathway in microorganisms has not been reported.
The trans-resveratrol biosynthesis pathway. Trans-resveratrol is synthesized starting with either phenylalanine or tyrosine; both pathways giving rise to 4-coumaroyl-CoA through several enzymes, including phenylalanine ammonia lyase (PAL), tyrosine ammonia lyase (TAL), cinnamate 4-hydroxylase (C4H), and 4-coumaroyl-CoA ligase (4CL). Finally, 4-coumaroyl-CoA is converted to trans-resveratrol in a reaction catalyzed by stilbene synthase (STS)
Based on the resveratrol biosynthesis pathway in plants and the fact that Alternaria sp. MG1 can convert phenylalanine to resveratrol, we speculated that there may be a similar phenylpropanoid pathway in this strain. In order to verify this speculation, the current study tested the capability of Alternaria sp. MG1 to convert different precursors and intermediates of the phenylpropanoid pathway to resveratrol, mainly including different amino acids, cinnamic acid, and p-coumaric acid. The activities of the key enzymes and the accumulation of intermediates in the phenylpropanoid pathway were also identified and monitored. Finally, the addition of glucose, soluble starch, and cyclodextrin to the medium was tried to improve the resveratrol biosynthesis from phenylalanine by Alternaria sp. MG1 cells.
Materials and methods
Microorganism and reagents
Alternaria sp. MG1, obtained from the China Center for Type Culture Collection (Wuhan, China) (code: Alternaria sp. CCTCC M 2011348), was used in this study. Chromatographically pure glutamic acid (Glu), arginine (Arg), phenylalanine (Phe), tyrosine (Tyr), tryptophan (Trp), histidine (His), and proline (Pro) (Sigma, St Louis, MO, USA) and cinnamic acid and p-coumaric acid (98 %; Aladdin, Shanghai, China) were used as substrates in the bioconversion to resveratrol. These substances and trans-resveratrol (≥99 %; National Institutes for Food and Drug Control, Beijing, China) were also used as the standards (dissolved by methyl alcohol) in the measurements. CoA-SH (Sigma) and glucose-6-phosphate sodium salt (G-6-PNa2) and ATP (MP Biomedicals, Santa Ana, CA, USA) were used in the enzyme reactions to detect enzyme activity.
Preparation of Alternaria sp. MG1 cells
Alternaria sp. MG1 was grown at 28 °C on potato dextrose agar (PDA) plates for 5 days and then prepared into a spore suspension of 1 × 107 spores/mL (measured using a hemacytometer) by washing the culture with sterile water. A 2-mL aliquot of the spore suspension was inoculated into 100 mL liquid potato dextrose broth (PDB) in a 250-mL flask and cultivated at 28 °C in a rotary shaker (120 rpm). After 4 days, the cells were collected by centrifugation at 1,136×g for 10 min at 4 °C using a refrigerated centrifuge (HC-3018R, Anhui USTC Zonkia Scientific Instruments Co., Ltd., Anhui, China). The cells were washed twice with sterile water and used to produce resveratrol throughout the study.
Conditions for resveratrol biosynthesis
The medium for bioconversion contained 0.2 mol/L phosphate buffer (pH 6.5), 0.1 g/L MgSO4, and 0.2 g/L CaSO4. For bioconversion, the prepared Alternaria sp. MG1 cells were added at 12 g cells (wet weight) per 100 mL medium (in a 250-mL flask). The bioconversion was carried out for 21 h at 28 °C and 120 rpm in a ZHWY-2102 constant temperature shaker (Shanghai WitCity Analyzing Instrument Manufactory Co., Ltd., Shanghai, China). After bioconversion, cells and medium were separately collected from the culture by filtration with intermediate speed qualitative filter paper (GB/T1914-2007, 102, Hangzhou Whatman-Xinhua Filter Paper Co., Ltd, Hangzhou, China) and used for further research on enzyme activities and the accumulation of cinnamic acid, p-coumaric acid, and resveratrol.
Bioconversion of different amino acids to resveratrol
In the bioconversion medium, Alternaria sp. MG1 cells and different amino acids (4 mmol/L) were added and resveratrol production was analyzed after the bioconversion. The amino acids used were the aliphatic amino acids glutamic acid (Glu) and arginine (Arg), the aromatic amino acids phenylalanine (Phe) and tyrosine (Tyr), and the heterocyclic amino acids tryptophan (Trp), histidine (His), and proline (Pro). A medium containing only Alternaria sp. MG1 cells was used as the control. Each treatment was conducted in duplicate and mean values and standard deviations are presented.
Intermediates accumulated during bioconversion of phenylalanine to resveratrol
In the bioconversion medium, 4.7 mmol/L phenylalanine and Alternaria sp. MG1 cells were used. During bioconversion, accumulation of cinnamic acid and p-coumaric acid inside and outside the cells was monitored at 8, 16, and 22 h, as well as the effects of different concentrations of phenylalanine (3, 4, 5, and 6 mmol/L) after 21 h. At the same time, resveratrol production outside the cells was also monitored. Each treatment was conducted in duplicate. The mean values are presented.
Biosynthesis of resveratrol from phenylalanine and different phenylpropanoid pathway intermediates
In the bioconversion medium containing Alternaria sp. MG1 cells, phenylalanine, cinnamic acid, and p-coumaric acid were separately added at different levels to start the bioconversion. The added levels were 2, 3, 4, 5, and 6 mmol/L of phenylalanine and 0.5, 1.0, 1.5, 2.0, and 3.0 mmol/L of cinnamic acid and p-coumaric acid. After 21 h, resveratrol accumulation outside the cells in the culture was determined. A medium containing only Alternaria sp. MG1 cells was used as the control. Each treatment was conducted in duplicate.
Distribution of enzymes bioconverting phenylalanine to resveratrol in Alternaria sp. MG1 culture
The distribution of phenylpropanoid pathway enzymes in Alternaria sp. MG1 culture was estimated according to the overall capability to bioconvert phenylalanine to resveratrol by crude enzyme extracts from inside and outside the cells. Namely, the bioconversion culture (100 mL) with phenylalanine as the substrate was collected after 21 h and separated to cells and liquid phase by filtration through a quantitative filter paper. Ammonium sulfate was added into the liquid phase, up to 75 % saturation. The obtained mixture was incubated overnight at 4 °C, and then the precipitated protein was collected by centrifugation for 10 min at 13,363×g. After washing twice with a 0.2 mol/L phosphate buffer (pH 7.0), the protein sediment, which represents the extracellular enzyme extracts, was dissolved in 8 mL phosphate buffer (pH 7.0). The cells were washed twice with phosphate buffer (pH 7.0) and homogenized with 6 mL phosphate buffer (pH 7.0) in an ice bath using a mortar. After centrifugation at 3,340×g for 10 min, the supernatant, which represents the intracellular enzyme extract, was collected and made up to 8 mL with phosphate buffer.
The overall activity of the extracted enzymes that bioconvert phenylalanine to resveratrol was analyzed by adding 5 mL enzyme extracts into 100 mL medium containing 4.7 mmol/L phenylalanine. After 5 h, the bioconversion was immediately stopped by cooling the culture from 28 to 4 °C. A bioconversion medium containing 4.7 mmol/L phenylalanine and 5 mL distilled water, instead of the enzyme extracts, was used as the blank control.
Activities of phenylalanine ammonia lyase (PAL, E.C.4.3.1.5) or tyrosine ammonia lyase (TAL, EC 4.3.1.25), cinnamate 4-hydroxylase (C4H, EC1.14.13.11), and 4-coumarate-CoA ligase (4CL, EC 6.2.1.12) in the crude intracellular enzyme extract were also tested to identify the presence of these enzymes in Alternaria sp. MG1. Each treatment was conducted in duplicate. The mean values and standard deviations are presented.
Activity of phenylpropanoid pathway enzymes during bioconversion of phenylalanine to resveratrol
At 6, 12, 18, and 24 h during bioconversion of 4.7 mmol/L phenylalanine to resveratrol, crude enzymes were extracted from Alternaria sp. MG1 cells and the activities of three key phenylpropanoid pathway enzymes, PAL, C4H, and 4CL, were measured. The effects of phenylalanine concentration on the enzyme activities of PAL, C4H, and 4CL were also investigated after 21 h at 1, 2, 3, 4, 5, and 6 mmol/L. Each treatment was conducted in duplicate.
Effects of carbon nutrients on bioconversion from phenylalanine to resveratrol
Under the consideration that phenylalanine and tryptophan can be produced from glucose through the shikimic acid pathway and glucose can be produced from hydrolysis of disaccharides and polysaccharides, the addition of these carbohydrates was supposed to increase resveratrol production. Therefore, six different carbon sources (glucose, fructose, sucrose, lactose, soluble starch, and cyclodextrin) were added at a final concentration of 1 g/L in the bioconversion system, and their effects on cell growth and resveratrol production were tested. Three carbohydrates, glucose, soluble starch, and cyclodextrin, were also tested at different levels, 0.5, 1.0, 2.0, and 3.0 g/L. In addition, the combination of different carbohydrates was also tried as shown in Table 4. After bioconversion for 21 h, resveratrol production was measured, and dry cell weight of Alternaria sp. MG1 was measured by drying cells at 60 °C for 48 h. All experiments were carried out in the reaction system containing 4.7 mmol/L phenylalanine, and the medium without any carbohydrate addition was used as the control. Each treatment was carried out in duplicate. The mean values are presented together with standard deviation.
Measurement of enzyme activities
Enzyme activities were measured using the prepared enzyme extracts in 0.2 mol/L phosphate buffer, pH 7.0. Protein concentration of the enzyme extracts was determined using the visible spectrophotometer, UVmini-1240 (Shimadzu, Kyoto, Japan) according to the Bradford method with bovine serum albumin (BSA) as the standard. All enzyme assays were conducted twice using freshly prepared extracts. At the end of the assays, the reaction system was centrifuged at 13,363×g for 10 min and the absorbance (OD value) of the supernatant was determined using the visible spectrophotometer, UVmini-1240. One unit (U) of the enzyme was defined as an increase of 0.01 of OD value per hour, and the enzyme activity was expressed as the units of enzyme per milligram protein (U/mg). The presented results are averages of duplicates.
PAL/TAL activity was measured using the methods reported by Beaudoin-Eagan and Thorpe (1985) and Xue et al. (2007). Briefly, PAL activity was assayed by following cinnamic acid formation at 290 nm in 5 mL of 50 mmol/L Tris–HCl buffer (pH 8.9) containing 5 mmol/L phenylalanine and 1.0 mL enzyme extract. TAL activity was similarly measured using tyrosine as the substrate in the same buffer containing 2 mmol/L tyrosine and 1.0 mL enzyme extract, and p-coumaric acid production was followed at 315 nm. As a control, a 1-min boiled enzyme extract was treated with the same conditions. The assays ran for 1 h in a 37 °C water bath and terminated by the addition of 200 μL 16 mol/L HCl.
C4H activity was assayed at 340 nm according to the method reported by Lamb and Rubery (1975). Briefly, the measurement was carried out in 2.5 mL of 50 mmol/L Tris–HCl buffer (pH 8.9), containing 2 μM cinnamic acid, 2 μM NADPNa2, 5 μM G-6-PNa2, and 1.0 mL enzyme extract. This system was incubated for 30 min in a water bath at 30 °C and then terminated by the addition of 200 μL 16 mol/L HCl. The control using double distilled water (ddH2O) instead of the enzyme extract was treated with the same conditions.
4CL activity was measured at 333 nm according to previously reported methods (Knobloch and Hahlbrock 1977). The enzyme reaction system contained 0.45 mL of 15 mmol/L MgSO4⋅7H2O, 0.15 mL of 5 mmol/L p-coumaric acid, 0.15 mL of 1 mmol/L CoA-SH, 0.15 mL of 50 mmol/L ATP, and 1.1 mL of 0.2 mol/L Tris–HCl buffer (pH 8.9). This enzyme reaction was incubated for 10 min in a water bath at 40 °C and terminated by the addition of 200 μL 16 mol/L HCl. The ddH2O substituting the crude enzyme solution was treated similarly and taken as the control.
Measurement and identification of cinnamic acid, p-coumaric acid, and resveratrol
For all treatments, resveratrol measurements were carried out only on the cell-free liquid phase, because when resveratrol accumulation was measured in previous studies inside and outside the cells, it was mainly found in the cell-free liquid phase (Zhang et al. 2013). Concentrations of cinnamic acid and p-coumaric acid were measured both inside and outside the cells.
Before high performance liquid chromatography (HPLC) measurements, cells were crushed with quartz sand for 10 min in an ice bath and then mixed with the cell-free culture filtrate for extraction of cinnamic acid and p-coumaric acid. Samples for measurements of resveratrol, cinnamic acid, and p-coumaric acid were all extracted three times (10 h each time) with ethyl acetate at a ratio of 100 mL samples to 150 mL ethyl acetate (50 mL each time). The ethyl acetate phases were combined and washed three times with 20 mL of 3 % NaHCO3 and then dried under vacuum at 40 °C. The dried residue was dissolved in 2 mL methanol (chromatographic grade; Sigma), filtered through a Millex®-HV filter membrane (0.45 μM, 13 mm diameter; Millipore, Billerica, MA, USA), and then directly subjected to HPLC measurement. The concentrations of resveratrol, cinnamic acid, and p-coumaric acid in the reaction system were calculated by dividing the reading values obtained according to the standard curves in HPLC by 50, because the sample was concentrated by 50 times before the HPLC measurements.
Concentrations of resveratrol, cinnamic acid, and p-coumaric acid were simultaneously determined using a Shimadzu Essentia LC-15C analytical HPLC system (Shimadzu) equipped with an LC-15C pump, a SIL-10AF automated sample injector, an SPD-15C dual-wavelength detector, a Shimadzu Wondasil C18-column (250 × 4.6 mm), and LC Solution software (Shimadzu). The column was operated at 35 °C. The mobile phase consisted of acetonitrile (chromatographic grade; Sigma) (solvent A) and ddH2O (solvent B). A multistep gradient was used for all analyses as follows: 0–28 min, 95 % (v/v) B; 28–33 min, 40 % B; 33–40 min, 15 % B; 40–50 min, 95 % B. The flow rate was 1 mL/min and the sample injection volume was 20 μL. The detection wavelength was 306 nm.
Standard cinnamic acid, p-coumaric acid, and trans-resveratrol (chromatographic grade; Sigma) were prepared in methanol solution with different concentrations ranging from 50 to 450 μg/L, from 50 to 250 μg/L, and from 25 to 325 μg/L with the lowest detection limits of 18.88, 21.03, and 9.11 μg/L, respectively. The obtained standard curves and linear regression equations were provided in the supplementary materials (Table S1and Fig. S1).
Production of cinnamic acid, p-coumaric acid, and resveratrol was identified using electrospray ionization mass spectrometry (ESI−) method in a Thermo LTQ XL ion trap HPLC-MS (Thermo Fisher Scientific Inc., USA) at capillary temperature 320 °C and sheath gas flow of 35 L/h. Positive polarity was set as source voltage of 4 kV, source current of 100 μA, and capillary voltage of 40 V. Negative polarity was set as source voltage of 4.5 kV, source current of 100 μA, and capillary voltage of −20 V. The run time was 60 min at a flow rate of 500 μL/min and the collision gas was argon. Negative ion mode selected multireaction monitoring (MRM) mode was used in the operation. In order to enhance the detection sensitivity, molecular ions with m/z = 146.5–147.5, m/z = 162.5–163.5, and m/z = 226.5–227.5 were focused in the analysis according to the molecular weight of cinnamic acid (148.17), p-coumaric acid (164.16), and resveratrol (228.24), respectively. Total ion current chromatogram and primary and secondary mass spectra were obtained for the standard and the detected compounds in the samples. This method has been successfully used by Pavlovic et al. (2013) and Wu et al. (2013) in the measurement of metabolites.
Statistical analysis
Data were analyzed using DPS software (Data Processing System: Statistics Version 6.55, 2005). Significance of differences between the control and treatments (α = 0.05) was tested using Duncan's new multiple range test (DMRT).
Results
Identification of cinnamic acid, p-coumaric acid, and resveratrol production
As shown in Figs. 2, 3, and 4, the mass spectra of the compounds in the samples were consistent with those of the corresponding standards in aspects of characteristic absorption peak and retention time in total ion current chromatogram, molecular weight ion in primary mass spectrum, and major daughter ions in secondary mass spectrum. The molecular weight and major daughter ions of the compounds detected in samples were obtained at m/z = 147.14 and m/z = 103.01 for cinnamic acid (Fig. 2), m/z = 163.09 and m/z = 119.01 for p-coumaric acid (Fig. 3), and m/z = 227.02 and m/z = 185.01, 183.00, 159.05, 156.93, and 143.03 for resveratrol (Fig. 4), being with the data obtained from the corresponding standards. Therefore, cinnamic acid, p-coumaric acid, and resveratrol detected in the samples were the same with the standard ones.
Bioconversion of different amino acids into resveratrol
In plants, resveratrol is synthesized via the phenylpropanoid pathway from phenylalanine or tyrosine. As expected, biosynthesis of resveratrol was found when the aromatic amino acids, phenylalanine and tyrosine, were used as the substrates, but not when aliphatic amino acids or heterocyclic amino acids were used as substrates (Table 1). Therefore, phenylalanine was selected as the main substrate to explore the resveratrol biosynthesis pathway.
Intermediates accumulated during bioconversion of phenylalanine into resveratrol
In the first step of the phenylpropanoid pathway, phenylalanine is converted into cinnamic acid by PAL (Donnez et al. 2009; Fig. 1). As shown in Fig. 5a, accumulation of cinnamic acid and p-coumaric acid was found in the bioconversion system with phenylalanine as the substrate, which is consistent with the phenylpropanoid pathway. During the bioconversion, cinnamic acid kept increasing, while p-coumaric acid increased and then decreased, and resveratrol increased continuously. Especially, after 16 h, there was a rapid increase in cinnamic acid and resveratrol, corresponding to a rapid decrease in p-coumaric acid, indicating the relationship between the biosynthesis of these compounds. Comparatively, cinnamic acid accumulation was much higher than p-coumaric acid and resveratrol, which is consistent with the material flow in the phenylpropanoid pathway, indicating a low efficiency in converting cinnamic acid to p-coumaric acid. The similar change rate of p-coumaric acid and resveratrol implied a similar efficiency of bioconversion of p-coumaric acid to p-coumaric acyl-CoA compared with that of p-coumaric acyl-CoA to resveratrol.
The accumulation of cinnamic acid, p-coumaric acid, and resveratrol during reaction in the resting cell system with phenylalanine as the substrate. The used condition is 4.7 mmol/L phenylalanine and reaction time of 21 h in a and b, respectively. Signals: cinnamic acid, triangle; p-coumaric acid, square; trans-resveratrol, diamond
The effects of phenylalanine concentration on the accumulation of cinnamic acid, p-coumaric acid, and resveratrol were also investigated. As shown in Fig. 5b, cinnamic acid still had the highest accumulation level, p-coumaric acid was second, and resveratrol was the lowest at all phenylalanine concentrations. The highest accumulation of cinnamic acid and resveratrol occurred at 5 mmol/L phenylalanine and that of p-coumaric acid at 4 mmol/L. The accumulation of cinnamic acid, resveratrol, and p-coumaric acid was inhibited when phenylalanine concentration was higher and lower than these levels, indicating that high phenylalanine concentration inhibits the material flow in the pathway.
Bioconversion of phenylalanine and different phenylpropanoid pathway intermediates into resveratrol
Resveratrol production was found in the bioconversion system with Alternaria sp. MG1 cells when only cinnamic acid or p-coumaric acid was used as the substrate (Fig. 6). Resveratrol production first increased with the increase of the concentration of these substrates and then decreased significantly, indicating substrate inhibition effects of these compounds as well as phenylalanine (Fig. 5b, the signals of diamond) on resveratrol production. Specifically, the highest resveratrol production was observed at 5.0 mmol/L phenylalanine, 1.0 mmol/L cinnamic acid, and 1.5 mmol/L p-coumaric acid. Comparing the highest bioconversion of different substrates to resveratrol revealed that resveratrol biosynthesis from p-coumaric acid was the highest (4.3 μg/L), followed by cinnamic acid (2.5 μg/L) and phenylalanine (1.35 μg/L). This is consistent with the fact that there are more steps and inhibition factors involved in the biosynthesis of resveratrol from phenylalanine and cinnamic acid compared with the biosynthesis from p-coumaric acid, which is in agreement with the material flow in the phenylpropanoid pathway.
Distribution of enzymes bioconverting phenylalanine to resveratrol in Alternaria sp. MG1 culture
The results of the substrate effects on resveratrol production indicated that the resveratrol biosynthesis pathway of Alternaria sp. MG1 may be similar to the reported phenylpropanoid pathway. To confirm this speculation, key enzymes in the phenylpropanoid pathway were examined.
As shown in Table 2, proteins were detected both inside and outside the Alternaria sp. MG1 cells in the bioconversion system. The protein concentration in the enzyme extracts obtained outside the cells (0.066 ± 0.011 mg/mL) was lower than that obtained inside the cells (0.341 ± 0.018 mg/mL). Furthermore, no activity of resveratrol synthesis was detected in the extracellular enzyme extracts in the reaction with phenylalanine as the substrate, whereas the intracellular enzyme extracts converted phenylalanine to 1.17 μg/L resveratrol, similar to the production using Alternaria sp. MG1 cells (1.24 μg/L, the control shown in Table 3). Therefore, the enzymes with the activity of resveratrol synthesis mainly accumulated inside Alternaria sp. MG1 cells.
In this study, PAL, TAL, C4H, and 4CL activities were determined because they are the key enzymes in the phenylpropanoid pathway. As shown in Table 2, the crude intracellular enzyme extract produced resveratrol; furthermore, the activities of PAL, TAL, C4H, and 4CL in the intracellular enzyme extract were 121.30 ± 5.65, 76.16 ± 3.36, 71.88 ± 1.39, and 325.21 ± 3.05 U/mg, respectively, indicating enzymatic evidence of the phenylpropanoid pathway in Alternaria sp. MG1.
Activity of phenylpropanoid pathway enzymes during bioconversion of phenylalanine to resveratrol
The changes in PAL, C4H, and 4CL activities with time during bioconversion of 4.7 mmol/L phenylalanine to resveratrol and after 21 h with different concentrations of phenylalanine are shown in Fig. 7. We found that PAL and 4CL activities decreased during the bioconversion, while C4H activity increased (Fig. 7a). The changes in activities were more rapid after 18 h. At 21 h, we observed that PAL and 4CL activities decreased with the increase of phenylalanine concentration, especially when the phenylalanine concentration was higher than 4.0 mmol/L. In contrast, C4H activity slightly decreased at phenylalanine concentration lower than 3 mmol/L, but increased at phenylalanine concentration higher than 4.0 mmol/L (Fig. 7b).
Effects of carbohydrate addition on bioconversion of phenylalanine to resveratrol
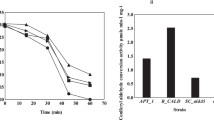
Coenzyme A, ATP, and malonyl-CoA are also important substrates of enzymatic reactions that produce resveratrol. These compounds are normally produced by glycometabolism using glucose as the starter (Murata et al. 1981). Therefore, carbon nutrients as energy sources that can be transformed into glucose were considered to be capable of improving resveratrol production in Alternaria sp. MG1. As expected, we found that biosynthesis of resveratrol from phenylalanine was significantly improved by adding different carbohydrates (Table 3). Cell weight also increased when these carbon sources were added, probably because they were used as carbon sources for cell growth. Comparatively, glucose resulted in the highest resveratrol production followed by soluble starch, while fructose, sucrose, and lactose did not have a significant effect on resveratrol production and cell growth. Dividing resveratrol production by cell weight revealed that the resveratrol produced per gram of cells was 0.53 and 0.47 μg/g when soluble starch and glucose were added, respectively, which was higher than that produced without their addition (control, 0.39 μg/g), indicating that the resveratrol synthesis efficiency increased by the addition of these carbon sources.
To optimize the conditions for bioconversion of phenylalanine to resveratrol by Alternaria sp. MG1 cells, different carbon sources were added at different concentrations as shown in Fig. 8 and, on that basis, different combinations with appropriate concentration as indicated in Table 4. Cyclodextrin was also tested because it has been reported to improve resveratrol production in cultured plant cells (Bru et al. 2006; Charmila et al. 2012). The results showed that the overall resveratrol production significantly increased by increasing the carbon sources' concentration within a certain level. The highest resveratrol production was achieved with 2 g/L glucose (3.21 ± 0.31 μg/L), 1 g/L soluble starch (2.17 ± 0.17 μg/L), and 1 g/L cyclodextrin (2.55 ± 0.18 μg/L), when these were added separately. Finally, the highest resveratrol production (4.57 ± 0.32 μg/L) was achieved when 2 g/L glucose and 1 g/L cyclodextrin were added together to the bioconversion system containing 4.7 mmol/L phenylalanine.
Effects of glucose (solid bars), soluble starch (shaded bars), and cyclodextrin (open bars) addition on resveratrol production. The data were obtained at reaction time of 21 h. Phenylalanine concentration used in the reaction was 4.7 mmol/L. Signals in the figure indicate addition of glucose (solid bars), soluble starch (shaded bars), and cyclodextrin (open bars) and the control (dotted bars) without any carbohydrate addition treatment
Discussion
The phenylpropanoid pathway in plants has been extensively studied and widely accepted as the pathway that biosynthesizes resveratrol (Fig. 1). Key enzymes in this pathway have been widely and successfully used to engineer resveratrol producing microorganisms and plants. However, so far, the entire phenylpropanoid pathway has not been found in one microorganism, although the key enzymes in this pathway have been separately found in different microorganisms. For example, PAL has been found in Rhodotorula glutinis and Rhodosporidium toruloids (Vannelli et al. 2007a; Wu et al. 2009); TAL in R. glutinis, Trichosporon cutaneum, and Saccharothrix espanaensis (Vannelli et al. 2007b; Berner et al. 2006); and 4CL in Streptomyces coelicolor (Park et al. 2009; Kaneko et al. 2003). To date, only plant STS has been used in genetically modified resveratrol-producing microorganisms. The phenylpropanoid pathway, STS, and chalcone synthase (CHS) have been found specifically taken from plants, until an STS-like domain was found in the complete genome sequence of Nocardiopsis dassonvillei (IMRU 509) (Sun et al. 2010). A strain of Bacillus sp. N had been found to produce stilbene (Kumar et al. 2012). However, this is the first report providing evidence of a resveratrol-producing phenylpropanoid pathway in microorganisms.
In the current study, no resveratrol biosynthesis was found when aliphatic amino acids or heterocyclic amino acids were used as substrates. This indicates that the presence of the benzene ring in aromatic amino acids may be the required moiety for resveratrol biosynthesis by Alternaria sp. MG1. There was no resveratrol accumulation in resting medium containing tryptophan as the sole substrate, although it contains a benzene ring. This may be because tryptophan contains an indole structure in the side chain, which is different from that in phenylalanine and tyrosine.
In plants, resveratrol accumulation has mainly been found inside cells (Kondan et al. 2001; Ferri et al. 2011). Using genetically modified yeast and E. coli produced resveratrol outside the cells (Horinouchi 2009; Beekwilder et al. 2006). In this study, accumulation of resveratrol was mainly found outside Alternaria sp. MG1 cells, which is consistent with the genetically modified microorganisms. Crude enzymes extracted from these cells also showed good capability to synthesize resveratrol, indicating the integrity and regulation systems in the enzyme mixtures and thus implying a potential to produce resveratrol by immobilizing the crude enzyme extracts or the whole cells. If it is efficient enough, biosynthesis of resveratrol will change from fermentation to enzymatic reactions.
Comprehensive analysis of intermediates accumulation (Fig. 5) and enzyme activities (Fig. 7) revealed that there was a correlation between the accumulation of phenylpropanoid pathway intermediates and the change in related enzyme activities, especially when the bioconversion time was longer than 18 h and the phenylalanine concentration was higher than 4.0 mmol/L. The relationship between the enzyme activities and the accumulation of phenylpropanoid pathway intermediates could be explained by product inhibition of metabolic regulation. Specifically, the strong correlation between the significant decrease in PAL activity and the increase in cinnamic acid accumulation after 18 h and at phenylalanine concentrations higher than 4.0 mmol/L indicated that there may be an inhibition regulation of PAL activity by high cinnamic acid concentration. Similarly, the decrease in 4CL activity after 18 h and at phenylalanine concentration higher than 4.0 mmol/L may be caused by the increase in resveratrol under these conditions. Based on this logic, the increase in C4H activity may be related to the decrease in p-coumaric acid accumulation in the bioconversion of phenylalanine to resveratrol.
However, further study is still needed to confirm the phenylalanine pathway in Alternaria sp. MG1 and to elucidate the mechanism and metabolic regulation involved. Although the occurrence of phenylpropanoid pathway intermediates and enzymes has been found in the bioconversion of phenylalanine to resveratrol by Alternaria sp. MG1, the exact material flow still needs to be identified by tracing the conversion of substrates during the bioconversion. This problem can be solved by monitoring the intermediates' sequence of occurrence during bioconversion and tracing the material flow of isotope-labeled phenylalanine and tyrosine. Furthermore, to identify the intact pathway, the properties and regulation of the enzymes involved, especially STS, need to be elucidated.
Finally, we found that carbon sources significantly improve the resveratrol biosynthesis by Alternaria sp. MG1 cells. This can be explained by the relationship between glycometabolism and the phenylpropanoid pathway. Glucose is the starter of glycometabolism. It can be metabolized to phenylalanine by the shikimic acid pathway, and thus, the relative phenylalanine content increases in the bioconversion system. At the same time, glucose can also be metabolized to other substrates essential for resveratrol biosynthesis, such as ATP, coenzyme A, and malonyl-CoA, by the Embden–Meyerhof–Parnas pathway (EMP), the hexose monophophate pathway (HMP), and the tricarboxylic acid cycle (TCA) (Altekar and Rangaswamy 1990; Ronimus and Morgan 2003; Yimga et al. 2006; Xu et al. 2012). In addition, glucose can increase the total enzyme content in the bioconversion system by increasing cell growth, as shown in this study. Therefore, glucose showed the most significant improvement of resveratrol production when different carbon sources were separately added at the same concentration. Soluble starch also significantly improved resveratrol production because it slowly metabolizes into glucose. The inability of fructose, sucrose, and lactose to improve resveratrol production and cell growth might be caused by the low efficiency of Alternaria sp. MG1 in metabolizing these carbon sources to glucose. The increase in resveratrol production by cyclodextrin may be because it serves as a carbon source in microorganism cultures (Fernandes et al. 2003), and it acts as an elicitor of resveratrol biosynthesis (Lucas-Abellan et al. 2007).
In conclusion, the current study provides evidence of a phenylalanine pathway in Alternaria sp. MG1 from three aspects: first, the capability of Alternaria sp. MG1 to biosynthesize resveratrol from phenylalanine, cinnamic acid, and p-coumaric acid when they were used as the sole substrate in the bioconversion system; second, the occurrence of key phenylalanine pathway enzymes in Alternaria sp. MG1; and third, the correlation between intermediates' accumulation and the changes in the related enzymes' activities. All the results obtained in the study strongly support the hypothesis of resveratrol biosynthesis by the phenylalanine pathway. Furthermore, consistent with the relationship between glycometabolism and the phenylpropanoid pathway, resveratrol production was significantly increased when carbon sources were added.
References
Altekar W, Rangaswamy V (1990) Indication of a modified EMP pathway for fructose breakdown in a halophilic archaebacterium. FEMS Microbiol Lett 69:139–143
Baur JA, Sinclair DA (2006) Therapeutic potential of resveratrol: the in vivo evidence. Nat Rev Drug Discov 5:493–506
Beaudoin-Eagan LD, Thorpe TA (1985) Tyrosine and phenylalanine ammonia lyase activities during shoot initiation in tobacco callus cultures. Plant Physiol 78:438–441
Becker JVW, Armstrong GO, Merwe MJ, Lambrechts MG, Vivier MA, Pretorius IS (2003) Metabolic engineering of Saccharomyces cerevisiae for the synthesis of the wine-related antioxidant resveratrol. FEMS Yeast Res 4:79–85
Beekwilder J, Wolswinkel R, Jonker H, Hall R, de Vos CH, Bovy A (2006) Production of resveratrol in recombinant microorganisms. Appl Environ Microb 72:5670–5672
Beihadj A, Telef N, Saigne C, Cluzet S, Barrieu F, Hamdi S, Merillon JM (2008) Effect of methyl jasmonate in combination with carbohydrates on gene expression of PR proteins, stilbene and anthocyanin accumulation in grapevine cell cultures. Plant Physiol Biochim 46:493–499
Berner M, Krug D, Bihlmaier C, Vente A, Muller R, Bechthold A (2006) Genes and enzymes involved in caffeic acid biosynthesis in the actinomycete Saccharothrix espanaensis. J Bacteriol 188:2666–2673
Bradamante S, Barenghi L, Villa A (2004) Cardiovascular protective effects of resveratrol. Cardiovasc Drug Rev 22:169–188
Bru R, Selles S, Casado-Vela J, Belchi-Navarro S, Pedren˜o MA, (2006) Modified cyclodextrins are chemically defined glucan inducers of defense responses in grapevine cell cultures. J Agric Food Chem 54:65–71
Charmila C, Ratnasooriya HP, Rupasinghe V (2012) Extraction of phenolic compounds from grapes and their pomace using β-cyclodextrin. Food Chem 134:625–631
Dai R, Ge H, Howard S, Qiu W (2012) Transcriptional expression of stilbene synthase genes are regulated developmentally and differentially in response to powdery mildew in Norton and Cabernet Sauvignon grapevine. Plant Sci 197:70–76
Donnez D, Jeandet P, Clément C, Courot E (2009) Bioproduction of resveratrol and stilbene derivatives by plant cells and microorganisms. Trends Biotechnol 27:706–713
Donnez D, Kim KH, Antoine S, Conreux A, de Luca V, Jeandet P, Clement C, Courot E (2011) Bioproduction of resveratrol and viniferins by an elicited grapevine cell culture in a 2 L stirred bioreactor. Process Biochem 46:1056–1062
Dudley J, Das S, Mukherjee S, Das DK (2009) Resveratrol, a unique phytoalexin present in red wine, delivers either survival signal or death signal to the ischemic myocardium depending on dose. J Nutr Biochem 20:443–452
Fernandes P, Cruz A, Angelova B, Pinheiro HM, Cabral JMS (2003) Microbial conversion of steroid compounds: recent developments. Enzym Microb Technol 32:688–705
Ferri M, Righetti L, Tassoni A (2011) Increasing sucrose concentrations promote phenylpropanoid biosynthesis in grapevine cell cultures. J Plant Physiol 168:189–195
Gresele P, Cerletti C, Guglielmini G, Pignatelli P, de Gaetano G, Violi F (2011) Effects of resveratrol and other wine polyphenols on vascular function: an update. J Nutr Biochem 22:201–211
Horinouchi S (2009) Combinatorial biosynthesis of plant medicinal polyketides by microorganisms. Curr Opin Chem Biol 13:197–204
Howitz KT, Bitterman KJ, Cohen HY, Lamming DW, Lavu S, Wood JG, Zipkin RE, Chung P, Kisielewski A, Zhang LL, Scherer B, Sinclair DA (2003) Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature 425:191–196
Jeandet P, Delaunois B, Aziz A, Donnez D, Vasserot Y, Cordelier S, Courot E (2012) Metabolic Engineering of yeast and plants for the production of the biologically active hydroxystilbene, resveratrol. J Biomed Biotechnol 2012:14. doi:10.1155/2012/579089
Kaneko M, Ohnishi Y, Horinouchi S (2003) Cinnamate: coenzyme A ligase from the filamentous bacterium Streptomyces coelicolor A 3(2). J Bacteriol 185:20–27
Kiselev KV, Tyunin AP, Zhuravlev YN (2013) Involvement of DNA methylation in the regulation of STS10 gene expression in Vitis amurensis. Planta 237:933–941
Knobloch KH, Hahlbrock K (1977) 4-Coumarate: CoA ligase from cell suspension culture of Petroselinum hortense Hoffm: partial purification, substrate specificity, and further properties. Arch Biochem Biophys 184:237–248
Kondan A, Kuroda H, Sakai F (2001) Simultaneous expression of stilbene synthase genes in Japanese red pine (Pinus densiflora). J Wood Sci 47:58–62
Kumar SN, Siji JV, Rajasekharan KN, Nambisan B, Mohandas C (2012) Bioactive stilbenes from a Bacillus sp. N strain associated with a novel rhabditid entomopathogenic nematode. Lett Appl Microbiol 54:410–417
Lamb CJ, Rubery PH (1975) A spectrophotometric assay for trans-cinnamic acid 4-hydroxylase activity. Anal Biochem 68:554–561
Lee SM, Yang H, Tartar DM, Gao B, Luo X, Ye SQ, Zaghouani H, Fang D (2011) Prevention and treatment of diabetes with resveratrol in a non-obese mouse model of type 1 diabetes. Diabetologia 54:1136–1146
Lijavetzky D, Almagro L, Belchi-Navarro S, Martínez-Zapater JM, Bru R, Pedreno MA (2008) Synergistic effect of methyljasmonate and cyclodextrin on stilbene biosynthesis pathway gene expression and resveratrol production in Monastrell grapevine cell cultures. BMC Res Notes 1:132
Lim CG, Fowler ZL, Hueller T, Schaffer S, Koffas MAG (2011) High-yield resveratrol production in engineered Escherichia coli. Appl Environ Microb 77:3451–3460
Liu ZY, Zhuang CX, Sheng SJ, Shao L, Zhao W, Zhao SJ (2011) Overexpression of a resveratrol synthase gene (PcRS) from Polygonum cuspidatum in transgenic Arabidopsis caused the accumulation of trans-piceid with antifungal activity. Plant Cell Rep 30:2027–2036
Lou JF, Fu LY, Peng YL, Zhou L (2013) Metabolites from Alternaria fungi and their bioactivities. Molecules 18:5891–5935
Lucas-Abellan C, Fortea I, Lopez-Nicolas JM, Nunez-Delicado E (2007) Cyclodextrins as resveratrol carrier system. Food Chem 104:39–44
Marienhagen J, Bott M (2013) Metabolic engineering of microorganisms for the synthesis of plant natural products. J Biotechnol 163:166–178
Murata K, Tani K, Kato J, Chibate I (1981) Glycolytic pathway as an ATP generation system and its application to the production of glutathione and NADH. Enzyme Microb Technol 3:233–242
Park SR, Yoon JA, Paik JH, Park JW, Jung WS, Ban YH, Kim EJ, Yoo YJ, Han AR, Yoon YJ (2009) Engineering of plant-specific phenylpropanoids biosynthesis in Streptomyces venezuelae. J Biotechnol 141:181–188
Pavlovic R, Cannizzo FT, Panseri S, Biolatti B, Trutic N, Biondi PA, Chiesa L (2013) Tetrahydro-metabolites of cortisol and cortisone in bovine urine evaluated by HPLC-ESI-mass spectrometry. J Steroid Biochem 135:30–35
Ro DK, Douglas CJ (2004) Reconstitution of the entry point of plant phenylpropanoid metabolism in yeast (Saccharomyces cerevisiae): implications for control of metabolic flux into the phenylpropanoid pathway. J Biol Chem 279:2600–2607
Ronimus RS, Morgan HW (2003) Distribution and phylogenies of enzymes of the Embden–Meyerhof–Parnas pathway from archaea and hyperthermophilic bacteria support a gluconeogenic origin of metabolism. Archaea 1:199–221
Shi JL, Zeng Q, Liu YL, Pan ZL (2012) Alternaria sp. MG1, a resveratrol-producing fungus: isolation, identification, and optimal cultivation conditions for resveratrol production. Appl Microbiol Biotechnol 95:369–379
Shin SY, Han NS, Park YC, Kim MD, Seo JH (2011) Production of resveratrol from p-coumaric acid in recombinant Saccharomyces cerevisiae expressing 4-coumarate: coenzyme A ligase and stilbene synthase genes. Enzym Microb Technol 48:48–53
Shumakova OA, Manyakhin AY, Kiselev KV (2011) Resveratrol content and expression of phenylalanine ammonia-lyase and stilbene synthase genes in cell cultures of Vitis amurensis treated with coumaric acid. Appl Biochem Biotechnol 165:1427–1436
Sun H, Lapidus A, Nolan M, Lucas S, Del Rio TG, Tice H, Cheng JF, Tapia R, Han C, Goodwin L, Pitluck S, Pagani L, Lvanova N, Mavromatis K, Mikhailova N, Pati A, Chen A, Palaniappan K, Land M, Hauser L, Chang YJ, Jeffries CD, Djao ODN, Rohde M, Sikorski J, Goker M, Woyke T, Bristow J, Eisen JA, Markowitz V, Hugenholtz P, Kyrpides NC, Klenk HP (2010) Complete genome sequence of Nocardiopsis dassonvillei type strain (IMRU 509). Stand Genomic Sci 3:325–336
Vannelli T, Wei QW, Sweigard J, Gatenby AA, Sariaslani FS (2007a) Production of p-hydroxycinnamic acid from glucose in Saccharomyces cerevisiae and Escherichia coli by expression of heterologous genes from plants and fungi. Metab Eng 9:142–151
Vannelli T, Xue ZX, Breinig S, Qi WW, Sariaslani FS (2007b) Functional expression in Escherichia coli of the tyrosine-inducible tyrosine ammonia-lyase enzyme from yeast Trichosporon cutaneum for production of p-hydroxycinnamic acid. Enzym Microb Technol 41:413–422
Wang YC, Yu O (2012) Synthetic scaffolds increased resveratrol biosynthesis in engineered yeast cells. J Biotechnol 157:258–260
Watts KT, Lee PC, Schmidt-Dannert C (2006) Biosynthesis of plant-specific stilbene polyketides in metabolically engineered Escherichia coli. BMC Biotechnol 6:22
Wu L, Wang X, Xu W, Farzaneh F, Xu R (2009) The structure and pharmacological functions of coumarins and their derivatives. Curr Med Chem 16:4236–4260
Wu ZJ, Wang JH, Fang DM, Zhang GL (2013) Analysis of iridoid glucosides from Paederia scandens using HPLC-ESI-MS/MS. J Chromatogr B 923–924:54–64
Xu QY, Ma L, Xie XX, Che N, Wang J (2012) Impacts of sodium citrate on metabolic flux distributions of L-valine fermentation. Adv Mater Res 343–344:643–648
Xue Z, McCluskey M, Cantera K, Ben-Bassat A, Sariaslani FS, Huang LX (2007) Improved production of p-hydroxycinnamic acid from tyrosine using a novel thermostable phenylalanine/tyrosine ammonia lyase enzyme. Enzym Microb Technol 42(58):64
Yimga MT, Leatham MP, Allen JH, Laux DC, Conway T, Cohen PS (2006) Role of gluconeogenesis and the tricarboxylic acid cycle in the virulence of Salmonella enterica serovar typhimurium in BALB/c mice. Infect Immun 74:1130–1140
Zhang JH, Shi JL, Liu YL (2013) Bioconversion of resveratrol using resting cells of non genetically modified Alternaria sp. Biotechnol Appl Biochem 60:236–243
Acknowledgments
This work was supported by the Agriculture Department of China through Project no. CARS-30 and 201003021.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 820 kb)
Rights and permissions
About this article
Cite this article
Zhang, J., Shi, J. & Liu, Y. Substrates and enzyme activities related to biotransformation of resveratrol from phenylalanine by Alternaria sp. MG1. Appl Microbiol Biotechnol 97, 9941–9954 (2013). https://doi.org/10.1007/s00253-013-5212-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-5212-3