Abstract
This review provides a comprehensive overview of the key aspects of the natural metabolite production by endophytic fungi, which has attracted significant attention due to its diverse biological activities and wide range of applications. Synthesized by various fungal species, these metabolites encompass compounds with therapeutic, agricultural, and commercial significance. We delved into strategies and advancements aimed at optimizing fungal metabolite production. Fungal cultivation, especially by Aspergillus, Penicillium, and Fusarium, plays a pivotal role in metabolite biosynthesis, and researchers have explored both submerged and solid-state cultivation processes to harness the full potential of fungal species. Nutrient optimization, pH, and temperature control are critical factors in ensuring high yields of the targeted bioactive metabolites especially for scaling up processes. Analytical methods that includes High-Performance Liquid Chromatography (HPLC), Liquid Chromatography–Mass Spectrometry (LC–MS), Gas Chromatography–Mass Spectrometry (GC–MS), Nuclear Magnetic Resonance (NMR), and Mass Spectrometry (MS), are indispensable for the identification and quantification of the compounds. Moreover, genetic engineering and metabolic pathway manipulation have emerged as powerful tools to enhance metabolite production and develop novel fungal strains with increased yields. Regulation and control mechanisms at the genetic, epigenetic, and metabolic levels are explored to fine-tune the biosynthesis of fungal metabolites. Ongoing research aims to overcome the complexity of the steps involved to ensure the efficient production and utilization of fungal metabolites.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Endophytic fungi (EFs) are a hyperdiverse group of organisms that during part or all of their life cycle colonize plant tissues (e.g., stem, flowers, leaves, fruits, roots) intra and/or extracellularly without causing symptoms of disease (Jia et al. 2016; dos Reis et al. 2022). These microorganisms are present in almost all plant species in natural ecosystems, where they play key roles in the plant micro-ecosystem, mainly under conditions of biotic or abiotic stress (Rho et al. 2018; Dastogeer 2018; Molina-Montenegro et al. 2023). EFs have co-evolved with host plants synthesizing numerous bioactive compounds that contribute to plant-fungus interactions, providing fitness benefits to host plants (Jia et al. 2016; Rho et al. 2018; Dastogeer 2018; Molina-Montenegro et al. 2023). These characteristics make these microorganisms one of the largest underexploited natural resources for the discovery of novel bioactive metabolites.
The EFs are capable of synthesizing a wide diversity of chemically different metabolites, which include alkaloids, benzopyrones, cytochalasins, steroids, phenols, isocoumarins, terpenoids, xanthones, etc. (Schulz and Boyle 2005; Manganyi and Ateba 2020; Hashem et al. 2023; Shen et al. 2023). Many of these metabolites are bioactive and may have antimicrobial, antioxidant, antiviral, anti-inflammatory, cytotoxic, and immunosuppressive activities (Manganyi and Ateba 2020; Mousa et al. 2021; Mohamed et al. 2022). Furthermore, EFs are capable of synthesizing metabolites similar to those found in plants, which drives the plant-fungus relationship comprehension (Kim et al. 2016). However, numerous challenges arise when studying the diversity of metabolites produced by EFs due to the complexity of crude extracts and the fact that classical methods of extracting and isolating metabolites are time-consuming and laborious. In addition, some bioactive metabolites of interest are only detected in trace amounts (Alhadrami et al. 2021). As an alternative to classical methodologies, metabolomics has emerged as a powerful tool to allow the comprehensive characterization of complex crude extracts and the chemical diversity of EFs metabolites (Alhadrami et al. 2021; Sayed et al. 2022), including small molecules.
Metabolomics is an “omics” technology defined as the study of all metabolites or small molecules in biological systems under specific conditions (Bundy et al. 2008; Patti et al. 2012; Marchev et al. 2021). This approach uses technological advances in analytical chemistry such as mass spectrometry (MS) (Rampler et al. 2021), to measure and compare the metabolites and small molecules present in the systems (Martin et al. 2019). However, although metabolomics provides a more holistic view of the chemical diversity produced by EFs, this approach is recent and depends on expensive equipments and a series of critical steps to obtain and analyze the data, including raw data preprocessing, peak annotation and multivariate statistical analysis.
Studying and understanding the chemical diversity of metabolites synthesized by EFs is extremely important to comprehend their biology, evolution, ecology, and possible biotechnological applications, which has been allowed by metabolomics in a holistic way as an emerging technology. However, critical steps ranging from formulation of hypotheses and objectives to data collection and analysis should be better addressed to study the chemical complexity produced by EFs successfully. In this review, we discussed the chemical diversity of metabolites produced by EFs and its applications, the use of metabolomics to explore chemical diversity in EFs, the inherent challenges, perspectives, limitations, tips and strategies for studying metabolite-producing EFs.
Biosynthesis of primary and secondary metabolites by endophytic fungi
Primary and secondary metabolites are two broad categories of chemical compounds produced by many organisms, including endophytic fungi, which are highly important for their overall fitness and survival. Involved in the basic life processes (i.e., growth, development, and energy production), primary metabolites are essential compounds that comprises carbohydrates, proteins, lipids, and nucleic acids, as well as key intermediates in metabolic pathways like glucose, amino acids, and ATP (adenosine triphosphate) (Alam et al. 2021). Secondary metabolites are not directly involved in primary metabolic processes, but often have specialized functions mainly associated with the defense systems or adaptation of organisms to environmental stresses due to the production of alkaloids, terpenoids, phenolics, and various other classes of compounds with diverse roles (Rashmi and Venkateswara Sarma 2018; Alam et al. 2021). Primary metabolites typically occur in all cells and tissues, and their production is tightly regulated to meet the organism’s basic metabolic needs. However, some key steps of the primary metabolism provide precursors for the synthesis of secondary metabolites (Alam et al. 2021). Mostly synthesized in limited quantities, secondary metabolites are generally produced in specific tissues or under particular conditions in response to environmental factors (e.g., stress, infection, or competition) (Sumarah and Miller 2009). The production of primary metabolites is generally continuous and essentially controlled to maintain the basic metabolic processes of organisms. The synthesis of secondary metabolites is more flexible and responsive to external factors, and its production is often induced by specific cues, such as pathogen attacks or environmental stressors, varying in response to changing conditions, and not produced continuously necessarily (Sumarah and Miller 2009). Moreover, primary metabolites are usually more uniform in structure and function as they play essential roles in the basic cellular processes. On the other hand, secondary metabolites exhibit a wide range of structural diversity and have various functions specific to a particular organism or ecological niche (Schneider et al. 2008; Bielecka et al. 2022). In addition, genomic studies have shown that endophytic fungi possess a larger number of biosynthetic gene clusters than ever expected for the secondary metabolite production, since most gene clusters are silent under laboratory conditions (Rashmi and Venkateswara Sarma 2018).
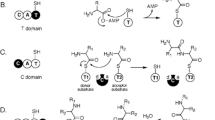
The main biochemical pathways that guide the biosynthesis of secondary metabolites by endophytic fungi involve polyketide synthases (PKSs), non-ribosomal peptide synthetases (NRPSs), hybrids (PKS-NRPS), terpene synthases (TPSs), terpene cyclases (TCs), and prenyltransferases (PTs) or combinations thereof (Rashmi and Venkateswara Sarma 2018). A relevant chemical group of secondary metabolites produced by endophytic fungi comprises non-ribosomal peptides synthesized by NRPSs enzymes (Yang et al. 2019) that possess modular structure and incorporate various building blocks into the growing peptide chain without the ribosome’s need. Non ribosomal peptide synthesis is an alternative pathway that allows production of polypeptides other than through the traditional translation mechanism. Despite the chemical diversity produced by NRPSs, the standard NRPS structure is composed of three canonical domains: adenylation (A), thiolation (T) or peptidyl carrier protein (PCP), and condensation (C) domains, which constitute a module within NRPS (Creamer et al. 2021). Each module is responsible for the recognition (A domain) and incorporation of a single amino acid into the growing peptide product. Generally, NRPSs possess more than one module, which terminates in a condensation-like (CT) domain that releases the peptide. Occasionally, epimerase (E) and N-methyltransferase (M) domains that convert L- to D-amino acids and N-methylate peptide bonds, respectively, are found within NRPSs. Deviations of the classical NRPS composition are present in hybrid PKS/NRPS (Creamer et al. 2021) and stand-alone monomodular NRPS-like enzymes, since not all canonical domains are presente. In addition, terpenes are an abundant and chemically diverse group of natural products synthesized by endophytic fungi and others, which is biosynthesized from isoprene units derived through the mevalonate and/or methylerythritol pathways that ranges from simple linear hydrocarbon chains to highly complex ring structures (Nazari et al. 2023).
Due to the diversity and complexity of biosynthetic pathways responsible for the production of secondary metabolites from endophytic fungi, Table 1 summarizes some chemically characterized secondary metabolites and putative associated biosynthetic genes/key enzymes determined by sequencing, including genes expressed/underexpressed in specific or mutational conditions.
Endophytic fungi as “hotspots” for discovering bioactive compounds
The discovery of the endophytic fungus Taxomyces andreanae as a taxol producer and related compounds, an anticancer used in the treatment of several types of tumors, placed EFs in the spotlight of search for bioactive metabolites similar to those found in plants (Strobel et al. 1996; Manganyi and Ateba 2020; Zhang et al. 2022; Hashem et al. 2023). It is currently well-known that EFs are a prophylactic source of metabolites from numerous chemical classes and with the most varied applications (Table 2), which characteristics make them “hotspots” of chemical diversity with different biological activities (Fig. 1). In ecology, the term ‘hotspots’ can be defined as geographic areas rich in biodiversity and threatened by habitat loss (Thompson et al. 2021). Here, “biodiversity hotspots” was adapted to “chemical diversity hotspots” in reference to EFs, since these microorganisms represent an important genetic, biological, and biotechnological resource for the discovery of novel biologically active compounds, with unprecedented chemical diversity, to be used in a wide variety of applications from medicine to agriculture, and beyond.
Guttation produced by endophytic fungi. Guttation is a phenomenon that involves the exudation of water and metabolites produced by fungal cells. These exudates are rich sources of mycotoxins, antimicrobials, insecticides, bioherbicides, antiviral, and anticancer agents. In a, b, the release of red/orange exudates from two strains of Epicoccum sp. cultured on potato peptone dextrose agar at 28 °C are shown; In c, brown exudates produced by colonies of Diaporthe sp. grown on potato dextrose agar (PDA) at 30 °C. In d, citrine yellow exudates produced by Penicillium sp. In e, it is shown the diffusion in the culture medium (PDA) of reddish-colored compounds produced by Chaetomium sp. In f, green colored exudate produced by Trichoderma sp. is highlighted. In g, h, and k–o, exudates from different endophytic Fusarium species grown in PDA at 25 °C are shown. In i, j, exudates produced by Diaporthe spp. In p, translucent exudates produced by colonies of Penicillium sp.
Different approaches have been employed to study the diversity of compounds from EFs (Mohamed et al. 2021, 2022; Hassane et al. 2022). Most protocols involve axenic cultivation, followed by obtaining a crude extract, screening based on bioassays (e.g., antimicrobial, antioxidant, antiparasitic activities, among others), isolation and purification steps of the target molecules (Orfali et al. 2017; Farooq et al. 2020; Liu et al. 2021; He et al. 2021). However, fungal extracts are complex and often contain tens to thousands of metabolites (Nischitha and Shivanna 2021a), which complicates the purification process. Studying metabolites produced by EFs in a specific condition simultaneously using classical approaches is quite laborious due to the complexity of the extracts, methodological limitations, laboriousness, and the time required to obtain pure compounds.
Metabolomic based methodologies is an alternative to classical methods and has become a powerfull tool to annotate EFs metabolites (González-Menéndez et al. 2016; Qadri et al. 2017; Toghueo et al. 2020). This new “omics” has been used to study the chemical diversity produced by fungi from the most varied ecosystems and lifestyles, enabling the discovery of new molecules, effects of epigenetic regulation on the metabolome, knowledge of the chemical diversity produced by these microorganisms etc. (Asai et al. 2012a, b, c; Zutz et al. 2013; Qadri et al. 2017; Triastuti et al. 2019; de Amorim et al. 2020; Ameen et al. 2020; Makhwitine et al. 2023). In this sense, we will address strategies in metabolomics research on EFs below: experimental designs, cultivation regimes, extraction methods, analytical methodologies and, data analysis.
Metabolomics applied to studies of endophytic fungi
Metabolomics comprises the study of all metabolites (< 1500 Da) in biological systems (e.g., cell, tissue, organ, organisms) in a specific physiological state (Bundy et al. 2008; Johnson et al. 2016; Marchev et al. 2021). This approach requires modern instrumental analytical methods of high throughput, sensitivity, and resolution, such as mass spectrometry (MS), combined with chemometric methods to measure and compare the metabolites (Johnson et al. 2016; Rampler et al. 2021; Martin et al. 2019). Metabolomics makes it possible to quickly measure thousands of metabolites simultaneously from minimal quantities of samples (Johnson et al. 2016; Rampler et al. 2021; Martin et al. 2019), and, traditionally, can be divided into metabolomics targeted and untargeted metabolomics (Fig. 2) (Roberts et al. 2012). In general, targeted metabolomics aims to qualitatively and quantitatively measure a predefined set of metabolites (Roberts et al. 2012). Recently, targeted metabolomics has been subdivided into widely targeted metabolomics, pseudo-targeted metabolomics, and quasi-targeted metabolomics (Lee et al. 2019; Sun et al. 2021; Wang et al. 2023). Untargeted metabolomics, on the other hand, analyzes all measurable metabolites in a given sample (Lippa et al. 2022).
Metabolomics has played crucial roles in elucidating physiological processes in numerous areas of research and development to discover disease state markers, stress response, identification of metabolic profiles, among others (Zutz et al. 2013; Triastuti et al. 2019; Aldholmi et al. 2020; Wei et al. 2020; Ameen et al. 2020; Zhu et al. 2021; Letertre et al. 2021; Katam et al. 2022). In recent decades, this approach has also been successfully employed to explore the chemical diversity of metabolites produced by fungi, including EFs (Table 3).
Design of experiments to study the chemical diversity of metabolites produced by endophytic fungi
According to the hypotheses and objectives of the research, different workflows are employed to evaluate the chemical diversity and consequently the bioactive compounds produced by endophytic fungi (Fig. 3). As hypotheses and/or objectives precede and guide the choice of the scientific experimentation methods, these workflows are generally complex. In metabolomic studies to explore the chemical diversity or to search for bioactive metabolites produced by EFs, the experimental design requires contextualization of the objectives at all stages, as each stage directly interferes with the results obtained and their interpretations. If the objective of the study aims to compare how the metabolic profile of EFs changes depending on different concentrations of heavy metals, the experimental design may contextualize its objective at each stage as follows: (1) the sample size should be sufficient to reveal differences if they occur; (2) all treatments should preferably possess the same sample number; (3) cultures should be maintained consistently under the same conditions; (4) the extraction methods and solvents used should be capable of extracting the greatest possible amount of metabolites; (5) the chosen analytical method should be capable of highlighting differences if they occur. If any of these steps are not carefully taken into consideration, the objective is not achieved, and biased results are generally generated. If temperature varies between different metal concentrations, for instance, differences in the metabolite profile may arise not only from the effect of metal concentration, but also from temperature variation.
The experimental design is, therefore, a critical step to be well structured according to the study’s objectives. Considering the study’s objectives or hypothesis to explore the chemical diversity produced by EFs using metabolomics, four main steps may be still summarized: (1) culture conditions, (2) sample preparation, (3) data collection and processing, 4) and data analysis. Since each of these steps is crucial to the final explanation, different strategies and tools that may be adopted to increase the sample number of metabolites from optimizing cultivation conditions, sample extraction, and data acquisition to choosing analytical methods and bioinformatics platforms, are discussed next.
Enhancement of metabolites production from endophytic fungi
Endophytic fungi are known to synthesize a wide variety of compounds. However, there is an inconsistency between the actual number of biosynthetic gene clusters (BGCs) present in the genome of these microorganisms and the number of compounds detected in any fungal strain under laboratory cultivation conditions (Fisch et al. 2009; Qadri et al. 2017; Ul-Hassan et al. 2012; González-Menéndez et al. 2016; Pillay et al. 2022; Xue et al. 2023). This divergence between the actual number of BGCs and the number of chemical molecules produced by any given fungal strain is attributed to the fact that the majority of BGCs remain silenced, low in expression, or not expressed at all under laboratory conditions (González-Menéndez et al. 2016; Pillay et al. 2022; Xue et al. 2023). Furthermore, the activation of these BGCs is likely to discover a greater number of compounds, including the discovery of new metabolites (Ding et al. 2020). As a result of this fact, there is a need to use different techniques that can induce the activation of these biosynthetic pathways, such as co-culture, One Strain-Many Compounds (OSMAC), epigenetic and molecular modification methods, thus increasing the biosynthetic capacity of these microorganisms (Fig. 4) (Bode et al. 2002; Cichewicz 2009; González-Menéndez et al. 2016; Pillay et al. 2022; Xue et al. 2023).
In natural ecossystems, the different species that make up microbial communities establish a series of ecological relationships (e.g., mutualism, commensalism, antagonism, and parasitism) with other microbial species or organisms (e.g., plants, animals, among others) that are mediated mainly by the production of compounds and small molecules (Toghueo et al. 2020; Koza et al. 2022) similarly to metabolite production expected by co-cultivation (Fig. 5). Thus, numerous BGCs are dependent on microbe-microbe or microbe-host interactions to be activated (Toghueo et al. 2020; Koza et al. 2022). The co-cultivation method (Fig. 5) aims to simulate interactions that occur in the environment naturally between microorganisms from the same or different ecological niches (Kim et al. 2021; Boruta et al. 2023), since it consists in growing one or more microbial species together. In addition to simulating ecological stress, including nutrient depletion during competition between species, the co-cultivation method allows the monitoring of metabolites produced over time through visual morphological changes in the species growth (Kim et al. 2021; Boruta et al. 2023) (Fig. 5). In most cases, this approach leads to changes in the biosynthetic profile of the strains analysed, resulting in production of unusual compounds not found in monocultures (Kim et al. 2021; Boruta et al. 2023). Competition for iron, for instance, triggers antibiotic biosynthesis in Streptomyces coelicolor (bacteria) during co-cultivation with Myxococcus xanthus (bacteria) (Lee et al. 2020).
Co-cultivation of different species of endophytic fungi on Potato Dextrose Agar (PDA). In a-c, the co-cultivation of six species of endophytic fungi isolated from Ouratea hexasperma is shown. The white arrows demonstrate the increased production of red pigment by a specific colony as a result of interaction with adjacent colonies. In d, both sides of a specific endophytic fungus colony are shown. In e, f, co-cultivation of this fungus (edge colony) with another endophytic fungus (middle colony) is shown. It is possible to observe that the fungus increases the production of yellow pigments (white arrows in e) and starts to produce a red pigment at the edge of the colony next (white arrows in f). In g, growth inhibition of an endophytic fungus due to the interaction with a saprophytic fungus is shown. In h, co-cultivation of Fusarium sp. (pink colony) and Diaporthe sp. (gray colony) is presented. It is possible to observe exudation of yellow metabolites by the Diaporthe where the colonies meet. In i, co-cultivation of Penicillium sp. and Eremothecium coryli showing the exudation of yellow compounds into the culture medium is indicated by the white arrow
Co-cultivation does not require knowledge in genetics or molecular biology, nor laborious methods, which makes it an advantageous method because of the simplicity and practicality. Different strategies for co-cultivation that includes co-cultivation in solid or liquid media (mixed fermentation), and the choice of the co-cultivated microorganisms (e.g., fungus-fungus, fungus-bacteria, and bacteria-bacteria) that may or not belong to the same ecological niche, are widely explored.
Similar to co-culture, the OSMAC approach is a relatively simple and effective technique used to explore the biosynthetic potential of microorganisms from the most diverse habitats (Gao et al. 2020; Schwarz et al. 2021; Pinedo-Rivilla et al. 2022; Hebra et al. 2022), including endophytic fungi (Gao et al. 2020; Wei et al. 2021; da Silva et al. 2023). As the OSMAC’s main idea, each microbial strain has the potential to produce many compounds, but subsets of these compounds are produced only under specific conditions (Bode et al. 2002). Thus, OSMAC involves combination and alteration of one or more cultivation parameters (e.g., carbon, nitrogen, and micronutrient sources; pH; temperature; light regime; addition of enzyme inhibitors; incubation time; O2 and CO2 tension; addition of solvents, heavy metals, precursors and other small molecule elicitors). Changes in cultivation conditions cause activation of silent BGCs or overexpression of BGCs that were underexpressed, resulting in production of a greater diversity of compounds (Bode et al. 2002). The efficiency of this approach was demonstrated by the increased production of griseofulvin derivatives produced by endophytic fungi from the host plant Moquiniastrum polymorphism when cultivated in culture media of malt peptone extract and Wickerham broth (Farinella et al. 2021). A species of endophytic Penicillium was capable of producing different types of compounds under combination of different cultivation media, many of which were specific to each combination evaluated (da Silva et al. 2023). Co-cultivation combined with OSMAC increased the biosynthetic production of compounds by microbial strains under environmental conditions capable of influencing the microbial interactions prior established, activating biosynthetic pathways that were initially silenced (Zutz et al. 2013).
Among the inducing strategies, epigenetic chemical regulation is considered a powerful approach to generate a greater number of compounds biosynthesized by fungi (González-Menéndez et al. 2016; Qadri et al. 2017; Toghueo et al. 2020). Epigenetic regulation involves alteration of the level of chromatin condensation to express silenced genes, which does not require prior knowledge in genomic content, resulting in the biosynthesis of previously unexpressed compounds. Generally, small molecules act as inhibitors of methyltransferase, histone deacetylase, and histone acetyltransferase, generating alterations in the structure of DNA, histones, and proteasomes, which activates cryptic BGCs (Table 4) (González-Menéndez et al. 2016; Qadri et al. 2017; Toghueo et al. 2020). Fermentation by Aspergillus calidoustus and Aspergillus westerdijkiae with vorinostat, an inhibitor of histone deacetylases (HDACs), for instance, induced changes in the metabolic profile of both species, with induction and repression of the biosynthesis of specific metabolites (Aldholmi et al. 2020). Moreover, addition of hydroxamic suberoylanilide (SAHA) and sodium valproate (VS), histone deacetylase (HDACs) inhibitors, caused different responses in the biosynthetic profile of the endophyte Botryosphaeria mamane (Triastuti et al. 2019). SAHA induced the production of eight metabolites, while VS induced the biosynthesis of two metabolites. In addition to the simplicity of the method, epigenetic regulation mediated by regulatory molecules can be used in conjunction with other strategies to stimulate the synthesis of metabolites, including coculture, OSMAC, and transformation methods (Zutz et al. 2013; Triastuti et al. 2019; Aldholmi et al. 2020; Ameen et al. 2020). However, alteration in the metabolic profile vary depending on the fungal species, type of regulatory molecule, and concentration (Zutz et al. 2013; Triastuti et al. 2019; Aldholmi et al. 2020; Ameen et al. 2020).
In addition, molecular approaches that include gene deletion or induction of mutants are valuable strategies to stimulate the production of compounds and activate BGCs of interest in fungi (Guzman-Chavez et al. 2018; Ding et al. 2020; Jo et al. 2023). Deletion of the hdaA gene in Penicillium chrysogenum strain Fes1701 induced a significant change in its metabolic profile, resulting in the bioactive indole alkaloid meleagrin synthesis (Ding et al. 2020). In other strain of P. chrysogenum, deletion of the hdaA gene caused a decrease in pigment production and overexpression of the sorbicillinoid biosynthetic gene cluster, producing overproduction of associated compounds (Guzman-Chavez et al. 2018). The plasma-induced mutant of Aspergillus unguis was able to synthesize four compounds not observed previously in the wild-type strain, one of them characterized as a novel aspergillusidone G (Yang et al. 2018). Thus, different molecular approaches have been efficiently employed to stimulate the biosynthesis and search for novel compounds. However, genomic knowledge, laborious methodologies and expensive equipments are essential.
Sample preparation and obtaining crude extracts
In metabolomics studies to investigate fungal metabolites, the sample preparation method is considered one of the most important steps due to its intrinsic sensitivity, since even small changes in procedures may deeply influence on the recovery of types and levels of metabolites, and in the biological interpretation of data consequently (Mohd et al. 2022). The choice of sample preparation and extraction method is crucial to define the number, type, and abundance of metabolites detected (Fig. 6). Therefore, an efficient, robust, simple, and reproducible method to demonstrate real occurrences and/or changes in the cellular metabolism is desirable. Generally, the workflow for sample preparation involves: (1) metabolic arrest by quenching; (2) sample collection, and separation of intra- and extracellular metabolites, and (3) extraction of metabolites.
Variation in the metabolomic profile according to different extraction methods employed. Each extraction method reflects the metabolic profile accessed effectively. The combination of different extraction strategies, solvents, temperature, sonication conditions, and extraction time result in different metabolic profiles. Figure created in BioRender software and Canva.com
Quenching is a rapid and sudden disruption of various metabolic pathways within cells on a timescale, maintaining stable metabolic contents, and reducing the degradation rates of investigated metabolites (Mohd et al. 2022). Briefly, quenching basically consists of cooling the samples in order to reduce or interrupt cellular metabolism. In microbial metabolomics research, the main quenching method uses ice-cold methanol (Mohd et al. 2022) by adding this solvent (60%–80%) (−80 to −20 °C) to the medium in which cells are grown to rapid stop the metabolic processes (Mohd et al. 2022). However, this approach is mainly used in bacterial metabolomic studies (Mohd et al. 2022), and hardly employed in metabolomic studies of filamentous fungi (Li et al. 2022). After disruption of metabolic processes, the next step comprises cell separation from the culture medium followed by metabolite extraction. To separate cells from the culture medium, centrifugation or gauze filtration methods are mainly used (Phan and Blank 2020; Mohd et al. 2022; Nzimande et al. 2022; Makhwitine et al. 2023).
The physical properties and chemical structure of the metabolites investigated should not be altered by extraction methods, which also should maximize the recovery of the maximum amount of metabolites. These methods are grouped into physical and chemical methods (Mohd et al. 2022; Li et al. 2022; Gopčević et al. 2022). Physical methods include the use of an ultrasonic bath, maceration with glass beads, freezing and thawing cycles, among others. (Mohd et al. 2022). Chemical methods mainly use polar and non-polar organic solvents, non-aqueous inorganic solvents, and combinations of both ones (Asai et al. 2012b, c; Qadri et al. 2017; Triastuti et al. 2019; Pacheco-Tapia et al. 2022). To increase extraction efficiency, physical and chemical methods can be combined (Mohd et al. 2022; Li et al. 2022; Gopčević et al. 2022; Makhwitine et al. 2023). In Table 5 are summarized some extraction methods commonly used in fungal metabolomics studies.
The choice of the extraction method depends on the study’s objective, which may include a combination of different strategies for extracting metabolites efficiently. In general, the main extraction method used for metabolomics studies in fungi comprises the liquid–liquid extraction employing ethyl acetate (EtOAc) (Asai et al. 2012b, c; Qadri et al. 2017; Triastuti et al. 2019; Pacheco-Tapia et al. 2022). However, other solvents and combinations of solvents are used according to the objectives established (Phan and Blank 2020; Makhwitine et al. 2023) as shown in Fig. 7. EtOAc, for instance, was used to extract the extracellular metabolites from a fermented broth by the endophytic fungus A. unguis, while methanol was used to extract intracellular metabolites (Yang et al. 2018). Other approaches include combining solvents with physical extraction methods. Using ethanol (EtOH:H2O; 7:2), methanol (MeOH:H2O; 7:2), and a chloroform/MeOH/water mixture (2:5:2) at different temperatures and sonication conditions, Phan and Blank (2020) quantified intracellular metabolites from Ustilago maydis. To study the metabolome of the endophyte P. chrysogenum, the extraction method selected consisted of adding methanol to the fermented broth, followed by incubation overnight on a rotary shaker at 150 rpm (Makhwitine et al. 2023). Currently, there is no standard extraction method to be recommended, since they vary greatly and depend on the sample nature and research objectives. However, the selected extraction method should prioritize the main compounds to meet the study’s objectives effectively.
Instrumental analysis methods used for metabolomic studies of endophytic fungi
The main analytical methods used in metabolomics studies of fungi extracts are gas chromatography/mass spectrometry (GC–MS), liquid chromatography/mass spectrometry (LC–MS), nuclear magnetic resonance spectroscopy (NMR), and nuclear magnetic resonance spectroscopy/mass spectrometry (NMR–MS) (David 2008; Zutz et al. 2013; Asai et al. 2012c; Qadri et al. 2017). Table 6 summarizes some of the advantages and limitations of these analytical platforms. However, capillary electrophoresis/mass spectrometry (CE–MS) (Ibrahim et al. 2016a) and matrix-assisted laser desorption ionization mass spectrometry (MALDI–MS) (Chen et al. 2021) have also been employed.
GC–MS is used to analyze volatile metabolites or metabolites that are easily volatilized after derivatization reaction (Qadri et al. 2017; Nzimande et al. 2022; Makhwitine et al. 2023). This approach possesses good repeatability and reproducibility and, is used in metabolomic analysis of various components that includes alcohols, aldehydes, amino acids, fatty acids, among others (Fiehn 2016; Qadri et al. 2017; Nzimande et al. 2022; Makhwitine et al. 2023). GS–MS was successfully employed to analyze fractions of crude extracts obtained from the fermented broth by the endophyte Alternaria alternata, and revealed the presence of 48 compounds (Nzimande et al. 2022). The main compounds were cyclotrisiloxane octamethyl, propaninitrile, pyrrolol[1,2-a]pyrazine-1,4-dione, hexahydro-3-(2-methyl propyl), diethylethoxy(2-ethoxyethyloxy), coumarin, 3,4-dihydro-4,5,7-trimethyl-4,5,7-trimethyl-2-chromanone, and 1,2-cyclobutanedicarbonitrile (Nzimande et al. 2022). Also, GC–MS was used to study volatile organic compounds (VOCs) produced by Muscodor yucatanensis, and demonstrated the presence of more than 40 compounds, including trans-3-dodecene, tau-gurjunene, benzene propanoic acid, 3,5-bis(1,1-dimet), 1-chloroheptacosane, and 2-hexyl-1-octanol as the main ones (Qadri et al. 2017). Moreover, GC–MS analysis demonstrated the presence of more than 110 compounds synthesized by the endophyte Penicillium chrysogenum, with some of the identified compounds known to exhibit antiviral activity (Makhwitine et al. 2023). However, GC–MS analysis has some disadvantages, since non-volatile compounds require derivatization (Bollenbach and Tsikas 2022).
LC–MS possesses numerous advantages, including simple sample preparation, high sensitivity, and high qualitative and quantitative capabilities (Gathungu et al. 2020). Thousands of peaks are detected by LC–MS in metabolomics studies (Nischitha and Shivanna 2021a). Each of the peaks can correspond to a detected ion with a mass-to-charge ratio (m/z) and a certain retention time that characterizes the metabolite. Moreover, with the development of high-performance liquid chromatography (HPLC) and ultra-performance liquid chromatography (UHPLC), peak resolution has been widely improved (Behnoush et al. 2015). Due to the high-resolution generated to detect most metabolites, LC–MS is the main technique to study the metabolic profile of fungi (Zutz et al. 2013; Mafezoli et al. 2018; Pacheco-Tapia et al. 2022). Four crude extracts of EFs from the host plant Alisma orientale analyzed by LC–MS revealed the presence of more than 50 compounds per extract, with a predominance of metabolites belonging to different chemical classes (flavonoids, non-flavonoids, phenolic acids, and flavonolignans). LC–MS analysis performed on the EFs Curvularia protuberata and Penicillium citrinum demonstrated the presence of 2352 and 2500 compounds, respectively, which were separated by positive and negative ion modes (Nischitha and Shivanna 2021a). Furthermore, LC–MS enabled the identification of some compounds known to exhibit antimicrobial activities. It is worth noting that despite the high resolution and sensitivity of this technique, overlapping peaks may occur, making decovolution necessary during the data processing stages.
NMR and NMR-MS are generally the analytical methods of choice when structural characterization of unknown compounds is required (Bingol and Brüschweiler 2016; Markley et al. 2017; Grienke et al. 2019; Gathungu et al. 2020). This analytical method is based on the interaction of a magnetically active nuclei with an applied external magnetic field, and is highly advantageous because of the high reproducibility, accurate quantification, simple preparation, measurable analytes in various solvents, clear identification of unknown metabolites, and complete detection of metabolites (Bingol and Brüschweiler 2016; Markley et al. 2017; Gathungu et al. 2020). NMR, for instance, was used to elucidate the structure of bipolarisenol produced by the endophyte Bipolaris sorokiniana (Khan et al. 2015), and four new chromium derivatives produced by the endophyte Phomopsis sp. (Huang et al. 2016). However, NMR-based methods have low sensitivity, which limits their applications in metabolomics (Markley et al. 2017).
Finally, the advantages and disadvantages of the different analytical tools should be considered in metabolomics research. A single tool cannot characterize and quantify efficiently thousands of metabolites produced by the fungal metabolism, requiring in some cases the combination of different methods to achieve the research objectives. In addition, other factors must be considered, which include errors associated with the platform used, implementation of internal standards, blanks (culture medium and solvents), quality controls (QC), which in metabolomics experiments of crude fungal extracts can be a mixture of all samples to be tested, and random sample injections.
Processing and analysis of metabolomics data from endophytic fungi
Data obtained in metabolomics experiments contains a large amount of important biological information from biomolecules (González-Menéndez et al. 2016; Nischitha and Shivanna 2021a). Interpreting these data involves inferring the mass and abundance of the biomolecules injected into the device. However, raw data may not provide a clean and reliable metabolite spectra (Lommen 2009; Katajamaa et al. 2006; Smith et al. 2006). Therefore, the raw data should be preprocessed to reduce the noise and to promote the baseline correction, peak detection and deconvolution, and data normalization as summarized in Fig. 8. Table 7 provides some online and open-source software as well as workflows for data analysis in metabolomics studies.
Briefly, tandem MS datasets are captured into binary files or databases by the software that controls the instruments (Chambers et al. 2012). However, the file format provided by different analytical platforms differs depending on the supplier (Kessner et al. 2008; Holman et al. 2014; Chambers et al. 2012). Thus, access to primary data can critically affect subsequent steps and the comparability of analytical platforms because some tools and workflows are designed for specific types of file formats. Therefore, in metabolomics studies, one of the first steps is file format conversion. To this end, numerous tools have been developed (Sturm et al. 2008; Kessner et al. 2008; Chambers et al. 2012; Holman et al. 2014), with the MS Convert tool, available on the ProteoWizard platform (Kessner et al. 2008; Holman et al. 2014), as the most tool employed. MS Convert is a command-line tool used for mass spectrometry data format conversion (Holman et al. 2014) that accepts raw data from several commercial companies and developers, including AB SCIEX (file format: WIFF; T2D), Agilent (file format: MassHunter;.d directories), Bruker (file format: fid;.d directories; xmassa; xml), Thermo (file format: raw), Waters (file format: raw directories), HUPO PSI (file format: mzML), ISB Seattle Proteome Center (file format: mzXML), Matrix Science ( file format: mgf), Yates/MacCoss Laboratories (file format: ms2; cms2; bms2), and Steen & Steen Laboratory (file format: mz5). Output files include mzML, mzXML, mz5, mgf, text, ms1, cms1, ms2, and cms2 formats.
After the file format conversion, the next step is the data preprocessing. Among the software developed, the XCMS (Smith et al. 2006) and MZmine (Katajamaa et al. 2006) are considered powerful tools for pre-processing, including deconvolution of analytical signals, noise filtering, detection and alignment of chromatographic peaks, baseline correction and gap filling and quantification of data from experiments analyzed by LC–MS and GC–MS. However, the processing in this software appears to underestimate GC–MS data (Ma and Qi 2021), with the MSDIAL software as an interesting alternative to be considered (Tsugawa et al. 2015).
Next, statistical analysis is applied after data pre-processing. Single variable or multivariable analyses are commonly used to evaluate groups as classical methods, with multivariate analysis as the most used in metabolomic studies of fungi (van Tilburg Bernardes et al. 2020; Swift et al. 2021; Castaño et al. 2022; Kandasamy et al. 2023). These analyses are generally employed to evaluate changes in the metabolic profiles between groups, and are divided into supervised and unsupervised analyzes (van Tilburg Bernardes et al. 2020; Swift et al. 2021; Castaño et al. 2022; Kandasamy et al. 2023). The major unsupervised analysis comprise Principal Component Analysis (PCA) and Hierarchical Clustering Analysis (HCA), and the supervised ones include Partial Least Squares Discriminant Analysis (PLS-DA) and Orthogonal Partial Least Squares Discriminant Analysis that is based on orthogonal signal correction (OPLS-DA) (Xia et al. 2015). PCA, for example, can be used to discriminate the metabolic profile between samples under different treatments (Du et al. 2021). PLS-DA and OPLS-DA can be employed to distinguish samples under different treatments and show the potential metabolites that contribute to these differences (Du et al. 2021; Maserumule et al. 2023; Singh et al. 2023). The main platform used for this type of analysis in metabolomics studies is MetaboAnalyst (Xia et al. 2015). In addition to these analyses, other statistical analysis (one factor), biomarker analysis, enrichment analysis, pathway analysis, functional meta-analysis of MS peaks, MS peaks to pathways, and network analysis, are included in this platform. Moreover, other tools are summarized in Table 7 in addition to MetaboAnalyst, as previously shown.
The Molecular Networking (MN) is a useful data analysis workflow for untargeted MS/MS-based metabolomics studies, since it provides means of identifying known compounds/molecular families, putatively novel molecular families, and evaluate differences associated with changes in culture conditions, among others (Ernst et al. 2019; Fan et al. 2019; Xu et al. 2021; Beniddir et al. 2021). MN organizes the MS/MS spectra of metabolites present in an extract according to their fragment similarities, which can be visualized by suitable software such as Cytoscape (Fan et al. 2019). MN uses the GNPS platform (Wang et al. 2016) to integrate a publicly available spectral library and the experimental MS/MS spectra for comparisons. GNPS offers a user-friendly workflow and numerous tools for analyzing MS/MS data, including MolNetEnhancer and Spec2Vec (Ernst et al. 2019; Huber et al. 2021). MolNetEnhancer integrates GNPS molecular network results, molecular mining tools (MN and MS2LDA), in silico annotation (NAP and DEREPLICATOR), and automated chemical annotation (Ernst et al. 2019). Thus, this approach allows annotation of multiple compounds, providing a comprehensive view of chemodiversity within a metabolome (Liu et al. 2022). It is worth noting that the MN created in the GNPS workflow are based on cosine scoring. Cosine-based methods are very good at revealing almost identical spectra, but are not as suitable for dealing with molecules with multiple local chemical modifications (Huber et al. 2021). The Spec2Vec, integrated into the GNPS workflow, can be an alternative, as it considers fragmentary relationships within a spectral dataset to derive abstract spectral embeddings that can be used to assess spectral similarities (Huber et al. 2021). There is also the SIMILE tool, which produces structural connections inferred from spectral alignment in MN (Treen et al. 2022). This tool makes it possible to classify spectral alignments based on p-values in order to explore the structural relationships between compounds and improve the chemical connectivity obtained with MN (Treen et al. 2022).
Other tools can also be used to exploit the data obtained from metabolomic studies of endophytic fungi or molecules isolated from these microorganisms, such as in silico bioactivity prediction tools. This type of approach has been considered promising for rapid, low-cost screening to discover potentially bioactive molecules and their mechanisms of action (Keiser et al. 2007; Gfeller et al. 2014; Gu et al. 2023). Currently, there are numerous protocols used to predict bioactivity in silico, which can be classified as approaches based on molecular similarity (Nickel et al. 2014), network-based models (Wang and Zeng 2013), and advanced machine learning methods (Pahikkala et al. 2015). An example of such a tools is the Swiss Target Prediction web server, which makes it possible to combine 2D and 3D similarity measurements of molecules with known ligands and map predictions between and within organisms based on target homology (Gfeller et al. 2014). There are also other in silico prediction tools, such as ChemMapper (Gong et al. 2013), SuperPred (Nickel et al. 2014), PharmMapper (Wang et al. 2017), and DrugBank (Wishart et al. 2006). Each of these tools has advantages and disadvantages, and the choice of tool depends on the objectives of the study. Furthermore, in silico bioactivity prediction does not replace biological assays, but serves as a quick guide for the search for new compounds.
Eco-Metabolomics of interaction between endophytic fungi and host plant
Endophytic fungi are an important component of the plant microecosystem and perform key functions for physiological, biological, and adaptive processes of host plants (Baron and Rigobelo 2021). The mechanisms of interaction between endophytes and host plants are complex and generally involve production of compounds by both fungus and plant, which act as effectors of a multidimensional interaction (Alam et al. 2021). Therefore, the colonization of plant tissue by endophytic fungi does not occur merely through chemotaxis, but depends on a complex and specific relationship, which is the result of the coevolution of these two groups of organisms (Alam et al. 2021). Plants secrete chemical compounds that are recognized by endophytic fungi (Tripathi et al. 2022; Hashem et al. 2023). This most often results in the production of specialized enzymes and compounds by endophytic fungi that are recognized by the host plant and act to mediate the colonization process (Tripathi et al. 2022; Hashem et al. 2023). Once inside the plant tissue, endophytic fungi can act directly on the fitness and physiology of their host through the production of numerous secondary metabolites (Alam et al. 2021; Tripathi et al. 2022). The plant, in turn, offers the fungus a habitat with stable environmental conditions and nutrients (Alam et al. 2021). However, it is worth highlighting that most of the metabolic pathways and compounds that mediate and/or originate from this interaction are unknown mainly due to methodological limitations.
Recently, a new disciplinary area within metabolomics has emerged, eco-metabolomics. This area of knowledge is focused on the application of metabolomics techniques to ecology aiming at characterizing the biochemical interactions of organisms at different spatio-temporal scales (Nagler et al. 2018; Peters et al. 2018; Wong et al. 2020). Therefore, the application of metabolomics to study the different interactions between species provides not only a comprehensive view of metabolic pathways involved, but also helps explain the mechanisms underlying the interactions (Gupta et al. 2022).
Many studies on interaction between EFs and host plant to decipher and/or understand ecological relationships have been published using metabolomics (Szűcs et al. 2018; Plaszkó et al. 2022; Poveda et al. 2022; Ma et al. 2023). The use of untargeted metabolomics (LC–MS) combined with metagenomics correlated the structure of the root mycobiome of Armoracia rusticana with significant changes in its metabolome (Plaszkó et al. 2022). Also, the concentration of kaempferol flavonoid glycosides positively correlated with the abundance of specific fungal taxa, while indole and glutathione isothiocyanate phytoalexins were negatively correlated with other fungal taxa (Plaszkó et al. 2022). Ma et al. (2023) used metabolomics to understand the symbiotic relationship between soil–plant–fungi and secondary metabolites in Fagopyrum dibotrys, demonstrating positive and negative correlations between certain fungal taxonomic groups with specific classes of compounds. These findings reinforce the importance of metabolomics to study the interaction between endophytes and host plants, aiming at elucidating and understanding the processes of endophytic community assembly, the environmental filter effect exerted by the host plant, and the effects of the EFs community on plant physiology, among other ecological aspects.
Final considerations
The exploration of metabolite production by EFs holds tremendous potential for a wide range of applications from medicine to agriculture, and beyond. Currently, EFs, with their vast genetic diversity and ability to produce an array of bioactive compounds, are an invaluable resource for addressing some of the most pressing challenges worldwide. The potential benefits from these multifaceted microorganims comprise life-saving antibiotics and antifungal agents, and new solutions to combat drug-resistant pathogens, with sustainable alternatives to traditional chemical pesticides and fertilizers in agriculture, enabling environmentally friendly and resilient practices. Furthermore, the commercial applications of fungal metabolites extend to various biotechnological innovations, offering solutions for a cleaner environment and reduced carbon footprint. As promising research continues to advance, the metabolite production potential by EFs tends to increase consistently. However, it is essential to focus on some associated challenges that includes optimization of the production methods in adittion to ensure sustainable sourcing. Finally, the EFs metabolite-producing capabilitiy represent a new era of scientific discovery and innovation to minimize environmental damages coupled with new drugs for a wide variety of human and animals needs.
Data availability
No datasets were generated or analysed during the current study.
References
Alam B, Lǐ J, Gě Q, Khan MA, Gōng J, Mehmood S, Yuán Y, Gǒng W (2021) Endophytic fungi: from symbiosis to secondary metabolite communications or vice versa? Front Plant Sci 17(12):791033. https://doi.org/10.3389/fpls.2021.791033
Aldholmi M, Wilkinson B, Ganesanm A (2020) Epigenetic modulation of secondary metabolite profiles in Aspergillus calidoustus and Aspergillus westerdijkiae through histone deacetylase (HDAC) inhibition by vorinostat. J Antibiot (tokyo) 73(6):410–413. https://doi.org/10.1038/s41429-020-0286-5
Alhadrami HA, Sayed AM, El-Gendy AO, Shamikh YI (2021) A metabolomic approach to target antimalarial metabolites in the Artemisia annua fungal endophytes. Sci Rep 11(1):2770. https://doi.org/10.1038/s41598-021-82201-8
Ameen F, Almansob A, Al Tami M, Al-Enazi N, Al-Sabri A, Orfali R (2020) Epigenetic modifiers affect the bioactive compounds secreted by an endophyte of the tropical plant Piper longum. Molecules 26(1):29. https://doi.org/10.3390/molecules26010029
Asai T, Morita S, Shirata N, Taniguchi T, Monde K, Sakurai H, Ozeki T, Oshima Y (2012a) Structural diversity of new C13-polyketides produced by Chaetomium mollipilium cultivated in the presence of a NAD(+)-dependent histone deacetylase inhibitor. Org Lett 14(21):5456–5459. https://doi.org/10.1021/ol302539s
Asai T, Yamamoto T, Oshima Y (2012b) Aromatic polyketide production in Cordyceps indigotica, an entomopathogenic fungus, induced by exposure to a histone deacetylase inhibitor. Org Lett 14(8):2006–2009. https://doi.org/10.1021/ol3005062
Asai T, Chung YM, Sakurai H, Ozeki T, Chang FR, Wu YC, Yamashita K, Oshima Y (2012c) Highly oxidized ergosterols and isariotin analogs from an entomopathogenic fungus, Gibellula formosana, cultivated in the presence of epigenetic modifying agents. Tetrahedron Lett 68:5817–5823. https://doi.org/10.1016/j.tet.2012.05.020
Asai T, Morita S, Taniguchi T, Monde K, Oshima Y (2016) Epigenetic stimulation of polyketide production in Chaetomium cancroideum by an NAD(+)-dependent HDAC inhibitor. Org Biomol Chem 14(2):646–651. https://doi.org/10.1039/c5ob01595b
Baron NC, Rigobelo EC (2021) Endophytic fungi: a tool for plant growth promotion and sustainable agriculture. Mycology 13(1):39–55. https://doi.org/10.1080/21501203.2021.1945699
Beale DJ, Pinu FR, Kouremenos KA, Poojary MM, Narayana VK, Boughton BA, Kanojia K, Dayalan S, Jones OAH, Dias DA (2018) Review of recent developments in GC-MS approaches to metabolomics-based research. Metabolomics 14(11):152. https://doi.org/10.1007/s11306-018-1449-2
Behnoush B, Sheikhazadi A, Bazmi E, Fattahi A, Sheikhazadi E, Saberi Anary SH (2015) Comparison of UHPLC and HPLC in benzodiazepines analysis of postmortem samples: a case-control study. Medicine 94(14):e640. https://doi.org/10.1097/MD.0000000000000640
Ben Mefteh F, Daoud A, Bouket AC, Thissera B, Kadri Y, Cherif-Silini H, Eshelli M, Alenezi FN, Vallat A, Oszako T et al (2018) Date palm trees root-derived endophytes as fungal cell factories for diverse bioactive metabolites. Int J Mol Sci 19:1986
Beniddir MA, Kang KB, Genta-Jouve G, Huber F, Rogers S, van der Hooft JJJ (2021) Advances in decomposing complex metabolite mixtures using substructure- and network-based computational metabolomics approaches. Nat Prod Rep 38(11):1967–1993. https://doi.org/10.1039/d1np00023c
Bielecka M, Pencakowski B, Nicoletti R (2022) Using next-generation sequencing technology to explore genetic pathways in endophytic fungi in the syntheses of plant bioactive metabolites. Agriculture 12:187. https://doi.org/10.3390/agriculture12020187
Bingol K, Brüschweiler R (2016) Knowns and unknowns in metabolomics identified by multidimensional NMR and hybrid MS/NMR methods. Curr Opin Biotechnol 43:17–24. https://doi.org/10.1016/j.copbio.2016.07.006
Bode HB, Bethe B, Höfs R, Zeeck A (2002) Big effects from small changes: possible ways to explore nature’s chemical diversity. ChemBioChem 3(7):619–627. https://doi.org/10.1002/1439-7633(20020703)3:7%3c619::AID-CBIC619%3e3.0.CO;2-9
Bollenbach A, Tsikas D (2022) GC-MS studies on the conversion and derivatization of γ-glutamyl peptides to pyroglutamate (5-oxo-proline) methyl ester pentafluoropropione amide derivatives. Molecules 27(18):6020. https://doi.org/10.3390/molecules27186020
Boruta T, Ścigaczewska A, Ruda A, Bizukojć M (2023) Effects of the coculture initiation method on the production of secondary metabolites in bioreactor cocultures of Penicillium rubens and Streptomyces rimosus. Molecules 28(16):6044. https://doi.org/10.3390/molecules28166044
Bundy JG, Davey MP, Viant MR (2008) Environmental metabolomics: a critical review and future perspectives. Metabolomics 5(1):3. https://doi.org/10.1007/s11306-008-0152-0
Cadamuro RD, Bastos IMADS, de Freitas ACO, Rosa MDS, Costa GO, da Silva IT, Robl D, Stoco PH, Sandjo LP, Treichel H, Steindel M, Fongaro G (2023) Bioactivity screening and chemical characterization of biocompound from endophytic Neofusicoccum parvum and Buergenerula spartinae isolated from mangrove ecosystem. Microorganisms 11(6):1599. https://doi.org/10.3390/microorganisms11061599
Castaño JD, Muñoz-Muñoz N, Kim YM, Liu J, Yang L, Schilling JS (2022) Metabolomics highlights different life history strategies of white and brown rot wood-degrading fungi. mSphere 7(6):e0054522. https://doi.org/10.1128/msphere.00545-22
Chambers MC, Maclean B, Burke R, Amodei D, Ruderman DL, Neumann S, Gatto L, Fischer B, Pratt B, Egertson J, Hoff K, Kessner D, Tasman N, Shulman N et al (2012) A cross-platform toolkit for mass spectrometry and proteomics. Nat Biotechnol 30(10):918–920. https://doi.org/10.1038/nbt.2377
Chapla VM, Zeraik ML, Ximenes VF, Zanardi LM, Lopes MN, Cavalheiro AJ, Silva DH, Young MC, Fonseca LM, Bolzani VS, Araújo AR (2014) Bioactive secondary metabolites from Phomopsis sp., an endophytic fungus from Senna spectabilis. Molecules 19(5):6597–6608. https://doi.org/10.3390/molecules19056597
Chatterjee S, Ghosh R, Mandal NC (2019) Production of bioactive compounds with bactericidal and antioxidant potential by endophytic fungus Alternaria alternata AE1 isolated from Azadirachta indica A. Juss. PLoS ONE 14(4):e0214744. https://doi.org/10.1371/journal.pone.0214744
Che J, Shi J, Gao Z, Zhang Y (2016) Transcriptome analysis reveals the genetic basis of the resveratrol biosynthesis pathway in an endophytic fungus (Alternaria sp. MG1) isolated from Vitis vinifera. Front Microbiol 7:1257
Chen XF, Hou X, Xiao M, Zhang L, Cheng JW, Zhou ML, Huang JJ, Zhang JJ, Xu YC, Hsueh PR (2021) Matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF MS) analysis for the identification of pathogenic microorganisms: a review. Microorganisms 9(7):1536. https://doi.org/10.3390/microorganisms9071536
Chen CJ, Lee DY, Yu J, Lin YN, Lin TM (2023) Recent advances in LC-MS-based metabolomics for clinical biomarker discovery. Mass Spectrom Rev 42(6):2349–2378. https://doi.org/10.1002/mas.21785
Chester K, Zahiruddin S, Ahmad A, Khan W, Paliwal S, Ahmad S (2017) Fusaristerol A: a new cytotoxic and antifungal ergosterol fatty acid ester from the endophytic fungus Fusarium sp. associated with Mentha longifolia roots. Phcog Mag 13:179–188. https://doi.org/10.4103/pm.pm
Cichewicz RH (2009) Epigenome manipulation as a pathway to new natural product scaffolds and their congeners. Nat Prod Rep 27(1):11–22. https://doi.org/10.1039/b920860g
Cook D, Donzelli B, Creamer R, Baucom DL, Gardner DR, Pan J, Moore N, Krasnoff SB, Jaromczyk JW, Schardl CL (2017) Swainsonine biosynthesis genes in diverse symbiotic and pathogenic fungi. Genes Genomes Genet 7:1791–1797
Creamer R, Hille DB, Neyaz M, Nusayr T, Schardl CL, Cook D (2021) Genetic relationships in the toxin-producing fungal endophyte, Alternaria oxytropis using polyketide synthase and non-ribosomal peptide synthase genes. J Fungi 7:538. https://doi.org/10.3390/jof70705
Crook AA, Powers R (2020) Quantitative NMR-based biomedical metabolomics: current status and applications. Molecules 25(21):5128. https://doi.org/10.3390/molecules25215128
da Silva FMR, Paggi GM, Brust FR, Macedo AJ, Silva DB (2023) Metabolomic strategies to improve chemical information from OSMAC studies of endophytic fungi. Metabolites 13(2):236. https://doi.org/10.3390/metabo13020236
Dastogeer KMG (2018) Influence of fungal endophytes on plant physiology is more pronounced under stress than well-watered conditions: a meta-analysis. Planta 248(6):1403–1416. https://doi.org/10.1007/s00425-018-2982-y
David S (2008) Wishart, quantitative metabolomics using NMR. Trends Anal Chem 27(3):228–237. https://doi.org/10.1016/j.trac.2007.12.001
de Amorim MR, Wijeratne EMK, Zhou S, Arnold AE, Batista ANL, Batista JMJ, Dos Santos LC, Gunatilaka AAL (2020) An epigenetic modifier induces production of 3-(4-oxopyrano)-chromen-2-ones in Aspergillus sp. AST0006, an endophytic fungus of Astragalus lentiginosus. Tetrahedron Lett 76(43):131525. https://doi.org/10.1016/j.tet.2020.131525
Ding X, Liu K, Zhang Y, Liu F (2017) De novo transcriptome assembly and characterization of the 10-hydroxycamptothecin-producing Xylaria sp. M71 following salicylic acid treatment. J Microbiol 55:871–876
Ding Z, Zhou H, Wang X, Huang H, Wang H, Zhang R, Wang Z, Han J (2020) Deletion of the histone deacetylase HdaA in endophytic fungus Penicillium chrysogenum Fes1701 induces the complex response of multiple bioactive secondary metabolite production and relevant gene cluster expression. Molecules 25(16):3657. https://doi.org/10.3390/molecules25163657
Dos Reis JBA, Lorenzi AS, do Vale HMM (2022) Methods used for the study of endophytic fungi: a review on methodologies and challenges, and associated tips. Arch Microbiol 204(11):675. https://doi.org/10.1007/s00203-022-03283-0
Du J, Ma W, Li Y, Lu X, Geng Z, Huang H, Yuan Y, Liu Y, Wang X, Wang J (2021) UPLC-MS-based non-targeted analysis of endogenous metabolite changes in the leaves of Scabiosa tschiliensis Grüning induced by 6-benzylaminopurine and kinetin. Front Plant Sci 21(12):700623. https://doi.org/10.3389/fpls.2021.700623
Duan R, Zhou H, Yang Y, Li H, Dong J, Li X, Chen G, Zhao L, Ding Z (2016) Antimicrobial meroterpenoids from the endophytic fungus Penicillium sp. T2–8 associated with Gastrodia elata. Phytochem Lett 18:197–201. https://doi.org/10.1016/j.phytol.2016.10.013
Ernst M, Kang KB, Caraballo-Rodríguez AM, Nothias LF, Wandy J, Chen C, Wang M, Rogers S, Medema MH, Dorrestein PC, van der Hooft JJJ (2019) MolNetEnhancer: enhanced molecular networks by integrating metabolome mining and annotation tools. Metabolites 9(7):144. https://doi.org/10.3390/metabo9070144
Fan B, Parrot D, Blümel M, Labes A, Tasdemir D (2019) Influence of OSMAC-based cultivation in metabolome and anticancer activity of fungi associated with the brown alga Fucus vesiculosus. Mar Drugs 17(1):67. https://doi.org/10.3390/md17010067
Farinella VF, Kawafune ES, Tangerina MMP, Domingos HV, Costa-Lotufo LV, Ferreira MJP (2021) OSMAC strategy integrated with molecular networking for accessing griseofulvin derivatives from endophytic fungi of Moquiniastrum polymorphum (Asteraceae). Molecules 26(23):7316. https://doi.org/10.3390/molecules26237316
Farooq S, Qayum A, Nalli Y, Lauro G, Chini MG, Bifulco G, Chaubey A, Singh SK, Riyaz-Ul-Hassan S, Ali A (2020) Discovery of a secalonic acid derivative from Aspergillus aculeatus, an endophyte of Rosa damascena Mill, Triggers Apoptosis in MDA-MB-231 triple negative breast cancer cells. ACS Omega 5(38):24296–24310. https://doi.org/10.1021/acsomega.0c02505
Fiehn O (2016) Metabolomics by gas chromatography-mass spectrometry: combined targeted and untargeted profiling. Curr Protoc Mol Biol 114:30.4.1-30.4.32. https://doi.org/10.1002/0471142727.mb3004s114
Firrincieli A, Eotillar R, Esalamov A, Eschmutz J, Ekhan Z, Redman RS, Fleck ND, Elindquist E, Grigoriev IV, Doty SL (2015) Genome sequence of the plant growth promoting endophytic yeast Rhodotorula graminis WP1. Front Microbiol 6:978
Fisch KM, Gillaspy AF, Gipson M, Henrikson JC, Hoover AR, Jackson L, Najar FZ, Wägele H, Cichewicz RH (2009) Chemical induction of silent biosynthetic pathway transcription in Aspergillus niger. J Ind Microbiol Biotechnol 36(9):1199–1213. https://doi.org/10.1007/s10295-009-0601-4
Gao Y, Stuhldreier F, Schmitt L, Wesselborg S, Guo Z, Zou K, Mándi A, Kurtán T, Liu Z, Proksch P (2020) Induction of new lactam derivatives from the endophytic fungus Aplosporella javeedii through an OSMAC approach. Front Microbiol 4(11):600983. https://doi.org/10.3389/fmicb.2020.600983
Gathungu RM, Kautz R, Kristal BS, Bird SS, Vouros P (2020) The integration of LC-MS and NMR for the analysis of low molecular weight trace analytes in complex matrices. Mass Spectrom Rev 39(1–2):35–54. https://doi.org/10.1002/mas.21575
Gfeller D, Grosdidier A, Wirth M, Daina A, Michielin O, Zoete V (2014) SwissTargetPrediction: a web server for target prediction of bioactive small molecules. Nucleic Acids Res 42(Web Server issue):W32–W38. https://doi.org/10.1093/nar/gku293
Gong J, Cai C, Liu X, Ku X, Jiang H, Gao D, Li H (2013) ChemMapper: a versatile web server for exploring pharmacology and chemical structure association based on molecular 3D similarity method. Bioinformatics 29(14):1827–1829. https://doi.org/10.1093/bioinformatics/btt270
González-Menéndez V, Pérez-Bonilla M, Pérez-Victoria I, Martín J, Muñoz F, Reyes F, Tormo JR, Genilloud O (2016) Multicomponent analysis of the differential induction of secondary metabolite profiles in fungal endophytes. Molecules 21(2):234. https://doi.org/10.3390/molecules21020234
Gopčević K, Grujić S, Arsenijević J, Džamić A, Veličković I, Izrael-Živković L, Medić A, Mudrić J, Soković M, Đurić A (2022) Bioactivity and phenolics profile of aqueous and ethyl acetate extracts of Satureja kitaibelii Wierzb. ex Heuff. obtained by ultrasound-assisted extraction. Sci Rep 12(1):21221. https://doi.org/10.1038/s41598-022-25668-3
Grienke U, Foster PA, Zwirchmayr J, Tahir A, Rollinger JM, Mikros E (2019) 1H NMR-MS-based heterocovariance as a drug discovery tool for fishing bioactive compounds out of a complex mixture of structural analogues. Sci Rep 9(1):11113. https://doi.org/10.1038/s41598-019-47434-8
Gu Y, Li J, Kang H, Zhang B, Zheng S (2023) Employing molecular conformations for ligand-based virtual screening with equivariant graph neural network and deep multiple instance learning. Molecules 28(16):5982. https://doi.org/10.3390/molecules28165982
Gupta S, Schillaci M, Roessner U (2022) Metabolomics as an emerging tool to study plant-microbe interactions. Emerg Top Life Sci 6(2):175–183. https://doi.org/10.1042/ETLS20210262
Guzman-Chavez F, Salo O, Samol M, Ries M, Kuipers J, Bovenberg RAL, Vreeken RJ, Driessen AJM (2018) Deregulation of secondary metabolism in a histone deacetylase mutant of Penicillium chrysogenum. Microbiologyopen 7(5):e00598. https://doi.org/10.1002/mbo3.598
Hashem AH, Attia MS, Kandil EK, Fawzi MM, Abdelrahman AS, Khader MS, Khodaira MA, Emam AE, Goma MA, Abdelaziz AM (2023) Bioactive compounds and biomedical applications of endophytic fungi: a recent review. Microb Cell Fact 22(1):107. https://doi.org/10.1186/s12934-023-02118-x
Hassane AMA, Taha TM, Awad MF, Mohamed H, Melebari M (2022) Radical scavenging potency, HPLC profiling and phylogenetic analysis of endophytic fungi isolated from selected medicinal plants of Saudi Arabia. Electron J Biotechnol 58:37–45. https://doi.org/10.1016/j.ejbt.2022.05.001
Haug K, Salek RM, Conesa P, Hastings J, de Matos P, Rijnbeek M, Mahendraker T, Williams M, Neumann S, Rocca-Serra P, Maguire E, González-Beltrán A, Sansone SA, Griffin JL, Steinbeck C (2012) MetaboLights–an open-access general-purpose repository for metabolomics studies and associated meta-data. Nucleic Acids Res 41(Database issue):D781–D786. https://doi.org/10.1093/nar/gks1004
He W, Xu Y, Wu D, Wang D, Gao H, Wang L, Zhu W (2021) New alkaloids from the diversity-enhanced extracts of an endophytic fungus Aspergillus flavus GZWMJZ-288. Bioorg Chem 107:104623. https://doi.org/10.1016/j.bioorg.2020.104623
Hebra T, Pollet N, Touboul D, Eparvier V (2022) Combining OSMAC, metabolomic and genomic methods for the production and annotation of halogenated azaphilones and ilicicolins in termite symbiotic fungi. Sci Rep 12(1):17310. https://doi.org/10.1038/s41598-022-22256-3
Holman JD, Tabb DL, Mallick P (2014) Employing ProteoWizard to convert raw mass spectrometry data. Curr Protoc Bioinform 46:13.24.1-13.24.9. https://doi.org/10.1002/0471250953.bi1324s46
Horai H, Arita M, Kanaya S, Nihei Y, Ikeda T, Suwa K, Ojima Y, Tanaka K, Tanaka S, Aoshima K, Oda Y, Kakazu Y (2010) MassBank: a public repository for sharing mass spectral data for life sciences. J Mass Spectrom 45(7):703–714. https://doi.org/10.1002/jms.1777
Huang M, Li J, Liu L, Yin S, Wang J, Lin Y (2016) Phomopsichin A-D; four new chromone derivatives from mangrove endophytic fungus Phomopsis sp. 33. Mar Drugs 14(11):215. https://doi.org/10.3390/md14110215
Huber F, Ridder L, Verhoeven S, Spaaks JH, Diblen F, Rogers S, van der Hooft JJJ (2021) Spec2Vec: improved mass spectral similarity scoring through learning of structural relationships. Plos Comput Biol 17(2):e1008724. https://doi.org/10.1371/journal.pcbi.1008724
Ibrahim M, Gahoual R, Enkler L, Becker HD, Chicher J, Hammann P, François YN, Kuhn L, Leize-Wagner E (2016a) Improvement of mitochondria extract from Saccharomyces cerevisiae characterization in Shotgun proteomics using sheathless capillary electrophoresis coupled to tandem mass spectrometry. J Chromatogr Sci 54(4):653–663. https://doi.org/10.1093/chromsci/bmw005
Ibrahim SRM, Elkhayat ES, Mohamed GAA, Fat’hi SM, Ross SA (2016b) Fusarithioamide A, a new antimicrobial and cytotoxic benzamide derivative from the endophytic fungus Fusarium chlamydosporium. Biochem Biophys Res Commun 479(2):211–216. https://doi.org/10.1016/j.bbrc.2016.09.041
Jahn L, Schafhauser T, Wibberg D, Rückert C, Winkler A, Kulik A, Weber T, Flor L, van Pée KH, Kalinowski J et al (2017) Linking secondary metabolites to biosynthesis genes in the fungal endophyte Cyanodermella asteris: the anti-cancer bisan-thraquinone skyrin. J Biotechnol 257:233–239
Jaitly N, Mayampurath A, Littlefield K, Adkins JN, Anderson GA, Smith RD (2009) Decon2LS: an open-source software package for automated processing and visualization of high resolution mass spectrometry data. BMC Bioinform 17(10):87. https://doi.org/10.1186/1471-2105-10-87
Jia M, Chen L, Xin HL, Zheng CJ, Rahman K, Han T, Qin LP (2016) A friendly relationship between endophytic fungi and medicinal plants: a systematic review. Front Microbiol 9(7):906. https://doi.org/10.3389/fmicb.2016.00906
Jo C, Zhang J, Tam JM, Church GM, Khalil AS, Segrè D, Tang TC (2023) Unlocking the magic in mycelium: using synthetic biology to optimize filamentous fungi for biomanufacturing and sustainability. Mater Today Bio 21(19):100560. https://doi.org/10.1016/j.mtbio.2023.100560
Johnson CH, Ivanisevic J, Siuzdak G (2016) Metabolomics: beyond biomarkers and towards mechanisms. Nat Ver Mol Cell Biol 17(7):451–459. https://doi.org/10.1038/nrm.2016.25
Kalimuthu AK, Pavadai P, Panneerselvam T, Babkiewicz E, Pijanowska J, Mrówka P, Rajagopal G, Deepak V, Sundar K, Maszczyk P, Kunjiappan S (2022) Cytotoxic potential of bioactive compounds from Aspergillus flavus, an endophytic fungus isolated from Cynodon dactylon, against breast cancer: experimental and computational approach. Molecules 27(24):8814. https://doi.org/10.3390/molecules27248814
Kandasamy D, Zaman R, Nakamura Y, Zhao T, Hartmann H, Andersson MN, Hammerbacher A, Gershenzon J (2023) Conifer-killing bark beetles locate fungal symbionts by detecting volatile fungal metabolites of host tree resin monoterpenes. Plos Biol 21(2):e3001887. https://doi.org/10.1371/journal.pbio.3001887
Kang X, Liu C, Shen P, Hu L, Lin R, Ling J, Xiong X, Xie B, Liu D (2019) Genomic characterization provides new insights into the biosynthesis of the secondary metabolite huperzine A in the endophyte Colletotrichum gloeosporioides Cg01. Front Microbiol 9:3237
Katajamaa M, Miettinen J, Oresic M (2006) MZmine: toolbox for processing and visualization of mass spectrometry based molecular profile data. Bioinformatics 22(5):634–636. https://doi.org/10.1093/bioinformatics/btk039
Katam R, Lin C, Grant K, Katam CS, Chen S (2022) Advances in plant metabolomics and its applications in stress and single-cell biology. Int J Mol Sci 23(13):6985. https://doi.org/10.3390/ijms23136985
Keiser MJ, Roth BL, Armbruster BN, Ernsberger P, Irwin JJ, Shoichet BK (2007) Relating protein pharmacology by ligand chemistry. Nat Biotech 25(2):197–206
Kessner D, Chambers M, Burke R, Agus D, Mallick P (2008) ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24(21):2534–2536. https://doi.org/10.1093/bioinformatics/btn323
Khan AL, Ali L, Hussain J, Rizvi TS, Al-Harrasi A, Lee IJ (2015) Enzyme inhibitory radicinol derivative from endophytic fungus Bipolaris sorokiniana LK12, associated with Rhazya stricta. Molecules 20(7):12198–12208. https://doi.org/10.3390/molecules200712198
Kim W, Peever TL, Park JJ, Park CM, Gang DR, Xian M, Davidson JA, Infantino A, Kaiser WJ, Chen W (2016) Use of metabolomics for the chemotaxonomy of legume-associated Ascochyta and allied genera. Sci Rep 5(6):20192. https://doi.org/10.1038/srep20192
Kim JH, Lee N, Hwang S, Kim W, Lee Y, Cho S, Palsson BO, Cho BK (2021) Discovery of novel secondary metabolites encoded in actinomycete genomes through coculture. J Ind Microbiol Biotechnol 48(3–4):kuaa001. https://doi.org/10.1093/jimb/kuaa001
Koza NA, Adedayo AA, Babalola OO, Kappo AP (2022) Microorganisms in plant growth and development: roles in abiotic stress tolerance and secondary metabolites secretion. Microorganisms 10(8):1528. https://doi.org/10.3390/microorganisms10081528
Krishna T, Yashwanti K, Surendra U, Ateet S, Kumar KM (2020) Piperine production from endophytic fungi of Piper nigrum l and in Silico approach for anti-inflammatory and anti-mycobacterial potential. Int Res J Biol Sci 8:8–15
Lee HJ, Kremer DM, Sajjakulnukit P, Zhang L, Lyssiotis CA (2019) A large-scale analysis of targeted metabolomics data from heterogeneous biological samples provides insights into metabolite dynamics. Metabolomics 15(7):103. https://doi.org/10.1007/s11306-019-1564-8
Lee N, Kim W, Chung J, Lee Y, Cho S, Jang KS, Kim SC, Palsson B, Cho BK (2020) Iron competition triggers antibiotic biosynthesis in Streptomyces coelicolor during coculture with Myxococcus xanthus. ISME J 14(5):1111–1124. https://doi.org/10.1038/s41396-020-0594-6
Lee S, Nguyen QN, Phung HM, Shim SH, Kim D, Hwang GS, Kang KS (2021) Preventive effects of anthraquinones isolated from an endophytic fungus, Colletotrichum sp. JS-0367 in tumor necrosis factor-α-stimulated damage of human dermal fibroblasts. Antioxidants (basel) 10(2):200. https://doi.org/10.3390/antiox10020200
Letertre MPM, Giraudeau P, de Tullio P (2021) Nuclear magnetic resonance spectroscopy in clinical metabolomics and personalized medicine: current challenges and perspectives. Front Mol Biosci 20(8):698337. https://doi.org/10.3389/fmolb.2021.698337
Li X, Lu P (2019) Transcriptome profiles of Alternaria oxytropis provides insights into swainsonine biosynthesis. Sci Rep 2019(9):6021
Li G, Jian T, Liu X, Lv Q, Zhang G, Ling J (2022) Application of metabolomics in fungal research. Molecules 27(21):7365. https://doi.org/10.3390/molecules27217365
Lippa KA, Aristizabal-Henao JJ, Beger RD, Bowden JA, Broeckling C, Beecher C, Clay Davis W, Dunn WB, Flores R, Goodacre R, Gouveia GJ (2022) Reference materials for MS-based untargeted metabolomics and lipidomics: a review by the metabolomics quality assurance and quality control consortium (mQACC). Metabolomics 18(4):24. https://doi.org/10.1007/s11306-021-01848-6
Liu DZ, Liang BW, Li XF, Liu Q (2014) Induced production of new diterpenoids in the fungus Penicillium funiculosum. Nat Prod Commun 9(5):607–608
Liu W, Liu Y, Yang F, Han S, Zhang J, Yang H, Cheng Z, Li Q (2021) Asperflaloids A and B from Aspergillus flavipes DZ-3, an endophytic fungus of Eucommia ulmoides oliver. Molecules 26(12):3514. https://doi.org/10.3390/molecules26123514
Liu SW, Jadambaa N, Nikandrova AA, Osterman IA, Sun CH (2022) Exploring the diversity and antibacterial potentiality of cultivable actinobacteria from the soil of the saxaul forest in southern Gobi desert in Mongolia. Microorganisms 10(5):989. https://doi.org/10.3390/microorganisms10050989
Lommen A (2009) MetAlign: interface-driven, versatile metabolomics tool for hyphenated full-scan mass spectrometry data preprocessing. Anal Chem 81(8):3079–3086. https://doi.org/10.1021/ac900036d
Lu H, Quan H, Ren Z, Wang S, Xue R, Zhao B (2016) The genome of Undifilum oxytropis provides insights into swainsonine biosynthesis and locoism. Sci Rep 6:30760
Lu Y, Ye C, Che J, Xu X, Shao D, Jiang C, Liu Y, Shi J (2019) Genomic sequencing, genome-scale metabolic network reconstruction, and in silico flux analysis of the grape endophytic fungus Alternaria sp. MG1. Microb Cell Fact 18:13
Ma A, Qi X (2021) Mining plant metabolomes: methods, applications, and perspectives. Plant Commun 2(5):100238. https://doi.org/10.1016/j.xplc.2021.100238
Ma N, Yin D, Liu Y, Gao Z, Cao Y, Chen T, Huang Z, Jia Q, Wang D (2023) Succession of endophytic fungi and rhizosphere soil fungi and their correlation with secondary metabolites in Fagopyrum dibotrys. Front Microbiol 1(14):1220431. https://doi.org/10.3389/fmicb.2023.1220431
Mafezoli J, Xu YM, Hilário F, Freidhof B, Espinosa-Artiles P, Dos Santos LC, de Oliveira MCF, Gunatilaka AAL (2018) Modulation of polyketide biosynthetic pathway of the endophytic fungus, Anteaglonium sp. FL0768, by copper (II) and anacardic acid. Phytochem Lett 28:157–163. https://doi.org/10.1016/j.phytol.2018.10.011
Makhwitine JP, Kumalo HM, Ndlovu SI, Mkhwanazi NP (2023) Epigenetic induction of secondary metabolites production in endophytic fungi Penicillium chrysogenum and GC-MS analysis of crude metabolites with anti-HIV-1 activity. Microorganisms 11(6):1404. https://doi.org/10.3390/microorganisms11061404
Manganyi MC, Ateba CN (2020) Untapped potentials of endophytic fungi: a review of novel bioactive compounds with biological applications. Microorganisms 8(12):1934. https://doi.org/10.3390/microorganisms8121934
Marchev AS, Vasileva LV, Amirova KM, Savova MS, Balcheva-Sivenova ZP, Georgiev MI (2021) Metabolomics and health: from nutritional crops and plant-based pharmaceuticals to profiling of human biofluids. Cell Mol Life Sci 78(19–20):6487–6503. https://doi.org/10.1007/s00018-021-03918-3
Markley JL, Brüschweiler R, Edison AS, Eghbalnia HR, Powers R, Raftery D, Wishart DS (2017) The future of NMR-based metabolomics. Curr Opin Biotechnol 43:34–40. https://doi.org/10.1016/j.copbio.2016.08.001
Martin HC, Ibáñez R, Nothias LF, Boya PCA, Reinert LK, Rollins-Smith LA, Dorrestein PC, Gutiérrez M (2019) Viscosin-like lipopeptides from frog skin bacteria inhibit Aspergillus fumigatus and Batrachochytrium dendrobatidis detected by imaging mass spectrometry and molecular networking. Sci Rep 9(1):3019. https://doi.org/10.1038/s41598-019-39583-7
Maserumule M, Rauwane M, Madala NE, Ncube E, Figlan S (2023) Defence-related metabolic changes in wheat (Triticum aestivum L.) seedlings in response to infection by Puccinia graminis f. sp. tritici. Plant Sci 14:1166813. https://doi.org/10.3389/fpls.2023.1166813
Miao LY, Mo XC, Xi XY, Zhou L, De G, Ke YS, Liu P, Song FJ, Jin WW, Zhang P (2018) Transcriptome analysis of a taxol-producing endophytic fungus Cladosporium cladosporioides MD2. AMB Expr 8:41
Mingxun W, Jeremy J, Carver VV et al (2016) (2016) Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat Biotech 34(8):828
Mohamed H, Hassane A, Atta O, Song Y (2021) Deep learning strategies for active secondary metabolites biosynthesis from fungi: harnessing artificial manipulation and application. Biocatal Agric Biotechnol 38:102195. https://doi.org/10.1016/j.bcab.2021.102195
Mohamed H, Ebrahim W, El-Neketi M, Awad MF, Zhang H, Zhang Y, Song Y (2022) In vitro phytobiological investigation of bioactive secondary metabolites from the malus domestica-derived endophytic fungus Aspergillus tubingensis strain AN103. Molecules 27:3762. https://doi.org/10.3390/molecules27123762
Mohd KK, Mahamad Maifiah MH, Abdul Rahim N, Hashim YZH, Abdullah Sani MS, Azizan KA (2022) Bacterial metabolomics: sample preparation methods. Biochem Res Int 12(2022):9186536. https://doi.org/10.1155/2022/9186536
Molina-Montenegro MA, Escobedo VM, Atala C (2023) Inoculation with extreme endophytes improves performance and nutritional quality in crop species grown under exoplanetary conditions. Front Plant Sci 22(14):1139704. https://doi.org/10.3389/fpls.2023.1139704
Mousa AAA, Mohamed H, Hassane AMA et al (2021) Antimicrobial and cytotoxic potential of an endophytic fungus Alternaria tenuissima AUMC14342 isolated from Artemisia judaica L. growing in Saudi Arabia. J King Saud Univ Sci 33(5):101462. https://doi.org/10.1016/j.jksus.2021.101462
Nagler M, Nägele T, Gilli C, Fragner L, Korte A, Platzer A, Farlow A, Nordborg M, Weckwerth W (2018) Eco-metabolomics and metabolic modeling: making the leap from model systems in the lab to native populations in the field. Front Plant Sci 6(9):1556. https://doi.org/10.3389/fpls.2018.01556
Nazari J, Payamnoor V, Kavosi MR, Asadi J (2023) Increasing terpenes in bark endophytic fungi of Betula pendula Roth as an anticancer potential source by cellulose nanofibers and sodium nitrate. S Afr J Bot 157:592–601. https://doi.org/10.1016/j.sajb.2023.04.032
Nickel J, Gohlke BO, Erehman J, Banerjee P, Rong WW, Goede A, Dunkel M, Preissner R (2014) SuperPred: update on drug classification and target prediction. Nucleic Acids Res 42(Web Server issue):W26–W31. https://doi.org/10.1093/nar/gku477
Nischitha R, Shivanna MB (2021a) Antimicrobial activity and metabolite profiling of endophytic fungi in Digitaria bicornis (Lam) Roem. and Schult. and Paspalidium flavidum (Retz) A. Camus. Biotech 11(2):53. https://doi.org/10.1007/s13205-020-02590-x
Nischitha R, Shivanna MB (2021b) Metabolite fingerprinting, in vitro antimicrobial and antioxidant activities and in-silico docking in Alloteropsis cimicina and its endophytic fungus Penicillium pinophilum. Mol Biol Rep 48:4021–4037. https://doi.org/10.1007/s11033-021-06410-0
Nischitha R, Shivanna MB (2022) Screening of secondary metabolites and antioxidant potential of endophytic fungus Penicillium citrinum and host Digitaria bicornis by spectrophotometric and electrochemical methods. Arch Microbiol 204(4):206. https://doi.org/10.1007/s00203-022-02795-z
Nischitha R, Vasanthkumari MM, Kumaraswamy BE, Shivanna MB (2020) Antimicrobial and antioxidant activities and chemical profiling of Curvularia tsudae endophytic in Cynodon dactylon (L.) Pers. Biotech 10(7):300. https://doi.org/10.1007/s13205-020-02250-0
Nishad JH, Singh A, Bharti R, Prajapati P, Sharma VK, Gupta VK, Kharwar RN (2021) Effect of the histone methyltransferase specific probe BRD4770 on metabolic profiling of the endophytic fungus Diaporthe longicolla. Front Microbiol 17(12):725463. https://doi.org/10.3389/fmicb.2021.725463
Ntana F, Bhat W, Johnson S, Jørgensen H, Collinge D, Jensen B, Hamberger B (2021) A Sesquiterpene synthase from the endophytic fungus Serendipita indica catalyzes formation of viridiflorol. Biomolecules 11:898
Nzimande B, Kumalo HM, Ndlovu SI, Mkhwanazi NP (2022) Secondary metabolites produced by endophytic fungi, Alternaria alternata, as potential inhibitors of the human immunodeficiency virus. Front Genet 13(13):1077159. https://doi.org/10.3389/fgene.2022.1077159
Orfali RS, Ebrahim W, El-Shafae AM (2017) Secondary metabolites from Alternaria sp., a fungal endophyte isolated from the seeds of Ziziphus jujuba. Chem Nat Com 53:878–880. https://doi.org/10.1007/s10600-017-2195-9
Pacheco-Tapia R, Vásquez-Ocmín P, Duthen S, Ortíz S, Jargeat P, Amasifuen C, Haddad M, Vansteelandt M (2022) Chemical modulation of the metabolism of an endophytic fungal strain of Cophinforma mamane using epigenetic modifiers and amino-acids. Fungal Biol 126(5):385–394. https://doi.org/10.1016/j.funbio.2022.02.005
Pahikkala T, Airola A, Pietilä S, Shakyawar S, Szwajda A, Tang J, Aittokallio T (2015) Toward more realistic drug-target interaction predictions. Brief Bioinf 16(2):325–337. https://doi.org/10.1093/bib/bbu010
Pang XJ, Zhang SB, Chen HL, Zhao WT, Yang DF, Xian PJ, Xu LL, Tao YD, Fu HY, Yang XL (2018) Emericelactones A-D: four novel polyketides produced by Emericella sp. XL 029, a fungus associated the leaves of Panax notoginseng. Tetrahedron Lett 59:4566–4570. https://doi.org/10.1016/j.tetlet.2018.11.032
Patti GJ, Yanes O, Siuzdak G (2012) Innovation: metabolomics: the apogee of the omics trilogy. Nat Rev Mol Cell Biol 13(4):263–269. https://doi.org/10.1038/nrm3314
Peng W, You F, Li XL, Jia M, Zheng CJ, Han T, Qin LP (2013) A new diphenyl ether from the endophytic fungus Verticillium sp. isolated from Rehmannia glutinosa. Chin J Nat Med 11(6):673–675. https://doi.org/10.1016/S1875-5364(13)60078-3
Peters K, Worrich A, Weinhold A, Alka O, Balcke G, Birkemeyer C, Bruelheide H, Calf OW, Dietz S, Dührkop K, Gaquerel E, Heinig U, Kücklich M et al (2018) Current challenges in plant eco-metabolomics. Int J Mol Sci 19(5):1385. https://doi.org/10.3390/ijms19051385
Phan ANT, Blank LM (2020) GC-MS-based metabolomics for the smut fungus Ustilago maydis: a comprehensive method optimization to quantify intracellular metabolites. Front Mol Biosci 19(7):211. https://doi.org/10.3389/fmolb.2020.00211
Pietzke M, Vazquez A (2020) Metabolite AutoPlotter—an application to process and visualise metabolite data in the web browser. Cancer Metab 10(8):15. https://doi.org/10.1186/s40170-020-00220-x
Pillay LC, Nekati L, Makhwitine PJ, Ndlovu SI (2022) Epigenetic activation of silent biosynthetic gene clusters in endophytic fungi using small molecular modifiers. Front Microbiol 14(13):815008. https://doi.org/10.3389/fmicb.2022.815008
Pinedo-Rivilla C, Aleu J, Durán-Patrón R (2022) Cryptic metabolites from marine-derived microorganisms using OSMAC and epigenetic approaches. Mar Drugs 20(2):84. https://doi.org/10.3390/md20020084
Plaszkó T, Szűcs Z, Cziáky Z, Ács-Szabó L, Csoma H, Géczi L, Vasas G, Gonda S (2022) Correlations between the metabolome and the endophytic fungal metagenome suggests importance of various metabolite classes in community assembly in horseradish (Armoracia rusticana, Brassicaceae) roots. Front Plant Sci 17(13):921008. https://doi.org/10.3389/fpls.2022.921008
Pluskal T, Castillo S, Villar-Briones A, Oresic M (2010) MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform 23(11):395. https://doi.org/10.1186/1471-2105-11-395
Poveda J, Rodríguez VM, Díaz-Urbano M, Sklenář F, Saati-Santamaría Z, Menéndez E, Velasco P (2022) Endophytic fungi from kale (Brassica oleracea var. acephala) modify roots-glucosinolate profile and promote plant growth in cultivated Brassica species. First description of Pyrenophora gallaeciana. Front Microbiol 13:981507. https://doi.org/10.3389/fmicb.2022.981507
Qadri M, Nalli Y, Jain SK, Chaubey A, Ali A, Strobel GA, Vishwakarma RA, Riyaz-Ul-Hassan S (2017) An insight into the secondary metabolism of Muscodor yucatanensis: small-molecule epigenetic modifiers induce expression of secondary metabolism-related genes and production of new metabolites in the endophyte. Microb Ecol 73(4):954–965. https://doi.org/10.1007/s00248-016-0901-y
Qiao W, Tang T, Ling F (2020) Comparative transcriptome analysis of a taxol-producing endophytic fungus, Aspergillus aculeatinus Tax-6, and its mutant strain. Sci Rep 10:10558
Rampler E, Abiead YE, Schoeny H, Rusz M, Hildebrand F, Fitz V, Koellensperger G (2021) Recurrent topics in mass spectrometry-based metabolomics and lipidomics-standardization, coverage, and throughput. Anal Chem 93(1):519–545. https://doi.org/10.1021/acs.analchem.0c04698
Rashmi M, Venkateswara Sarma V (2018) Secondary metabolite production by endophytic fungi: the gene clusters, nature, and expression. In: Jha S (ed) Endophytes and secondary metabolites. Reference series in phytochemistry. Springer, Cham, pp 1–16. https://doi.org/10.1007/978-3-319-76900-4_20-1
Rho H, Hsieh M, Kandel SL, Cantillo J, Doty SL, Kim SH (2018) Do endophytes promote growth of host plants under stress? A meta-analysis on plant stress mitigation by endophytes. Microb Ecol 75(2):407–418. https://doi.org/10.1007/s00248-017-1054-3
Roberts LD, Souza AL, Gerszten RE, Clish CB (2012) Targeted metabolomics. Curr Protoc Mol Biol 98:30.2.1-30.2.24. https://doi.org/10.1002/0471142727.mb3002s98
Ruan BH, Yu ZF, Yang XQ, Yang YB, Hu M, Zhang ZX, Zhou QY, Zhou H, Ding ZT (2018) New bioactive compounds from aquatic endophyte Chaetomium globosum. Nat Prod Res 32(9):1050–1055. https://doi.org/10.1080/14786419.2017.1378210
Sawada Y, Nakabayashi R, Yamada Y, Suzuki M, Sato M, Sakata A, Akiyama K, Sakurai T, Matsuda F, Aoki T, Hirai MY, Saito K (2012) RIKEN tandem mass spectral database (ReSpect) for phytochemicals: a plant-specific MS/MS-based data resource and database. Phytochem 82:38–45. https://doi.org/10.1016/j.phytochem.2012.07.007
Sayed AM, Sherif NH, El-Gendy AO, Shamikh YI, Ali AT, Attia EZ, El-Katatny MH, Khalifa BA, Hassan HM, Abdelmohsen UR (2022) Metabolomic profiling and antioxidant potential of three fungal endophytes derived from Artemisia annua and Medicago sativa. Nat Prod Res 36(9):2404–2408. https://doi.org/10.1080/14786419.2020.1831495
Schneider P, Misiek M, Hoffmeister D (2008) In vivo and in vitro production options for fungal secondary metabolites. Mol Pharm 5:234–242. https://doi.org/10.1021/mp7001544
Schulz B, Boyle C (2005) The endophytic continuum. Mycol Res 109(Pt 6):661–686. https://doi.org/10.1017/s095375620500273x
Schwarz J, Hubmann G, Rosenthal K, Lütz S (2021) Triaging of culture conditions for enhanced secondary metabolite diversity from different bacteria. Biomolecules 11(2):193. https://doi.org/10.3390/biom11020193
Seger C (2012) Usage and limitations of liquid chromatography-tandem mass spectrometry (LC-MS/MS) in clinical routine laboratories. Wien Med Wochenschr 162(21–22):499–504. https://doi.org/10.1007/s10354-012-0147-3
Shaw JJ, Berbasova T, Sasaki T, Jefferson-George K, Spakowicz D, Dunican BF, Portero CE, Narváez-Trujillo A, Strobel SA (2015) Identification of a fungal 1,8-cineole synthase from Hypoxylon sp. with specificity determinants in common with the plant synthases. J Biol Chem 290:8511–8526
Shen N, Chen Z, Cheng G, Lin W, Qin Y, Xiao Y, Chen H, Tang Z, Li Q, Yuan M, Bu T (2023) Diversity, chemical constituents and biological activities of endophytic fungi from Alisma orientale (Sam.) Juzep. Front Microbiol 14:1190624. https://doi.org/10.3389/fmicb.2023.1190624
Siless GE, Gallardo GL, Rodriguez MA, Rincón YA, Godeas AM, Cabrera GM (2018) Metabolites from the dark septate endophyte Drechslera sp. evaluation by LC/MS and principal component analysis of culture extracts with histone deacetylase inhibitors. Chem Biodivers 15(8):e1800133. https://doi.org/10.1002/cbdv.201800133
Singh DP, Maurya S, Yerasu SR et al (2023) Metabolomics of early blight (Alternaria solani) susceptible tomato (Solanum lycopersicum) unfolds key biomarker metabolites and involved metabolic pathways. Sci Rep 13:21023. https://doi.org/10.1038/s41598-023-48269-0
Smith CA, Want EJ, O’Maille G, Abagyan R, Siuzdak G (2006) XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal Chem 78(3):779–787. https://doi.org/10.1021/ac051437y
Sobreira ACM, Pinto FDCL, Florêncio KGD, Wilke DV, Staats CC, Streit RDAS, Freire FDCDO, Pessoa ODL, Trindade-Silva AE, Canuto KM (2018) Endophytic fungus Pseudofusicoccum stromaticum produces cyclopeptides and plant-related bioactive rotenoids. RSC Adv 8:35575–35586
Strobel G, Yang X, Sears J, Kramer R, Sidhu RS, Hess W (1996) Taxol from Pestalotiopsis microspora, an endophytic fungus of Taxus wallachiana. Microbiology 142:435–440. https://doi.org/10.1099/13500872-142-2-435
Sturm M, Bertsch A, Gröpl C, Hildebrandt A, Hussong R, Lange E, Pfeifer N, Schulz-Trieglaff O, Zerck A, Reinert K, Kohlbacher O (2008) OpenMS—an open-source software framework for mass spectrometry. BMC Bioinform 26(9):163. https://doi.org/10.1186/1471-2105-9-163
Sud M, Fahy E, Cotter D, Azam K, Vadivelu I, Burant C, Edison A, Fiehn O, Higashi R, Nair KS, Sumner S, Subramaniam S (2016) Metabolomics Workbench: an international repository for metabolomics data and metadata, metabolite standards, protocols, tutorials and training, and analysis tools. Nucleic Acids Res 44(D1):D463–D470. https://doi.org/10.1093/nar/gkv1042
Sumarah MW, Miller JD (2009) Anti-insect secondary metabolites from fungal endophytes of conifer trees. Nat Prod Commun 4(11):1497–1504
Sun X, Wang N, Li P, Jiang Z, Liu X, Wang M, Su Z, Zhang C, Lin F, Liang Y (2019) Endophytic fungus Falciphora oryzae promotes lateral root growth by producing indole derivatives after sensing plant signals. Plant Cell Environ 43:358–373
Sun H, Chen N, Wang X, Li N, Wang S, Zhang Z, Zhou Y, Yang J (2021) The study on the pathogenesis of pediatric lymphoma based on the combination of pseudotargeted and targeted metabolomics. Biomed Res Int 25:9984357. https://doi.org/10.1155/2021/9984357
Swift CL, Louie KB, Bowen BP, Hooker CA, Solomon KV, Singan V, Daum C, Pennacchio CP, Barry K, Shutthanandan V, Evans JE, Grigoriev IV, Northen TR, O’Malley MA (2021) Cocultivation of anaerobic fungi with rumen bacteria establishes an antagonistic relationship. Mbio 12(4):e0144221. https://doi.org/10.1128/mBio.01442-21
Szűcs Z, Plaszkó T, Cziáky Z, Kiss-Szikszai A, Emri T, Bertóti R, Sinka LT, Vasas G, Gonda S (2018) Endophytic fungi from the roots of horseradish (Armoracia rusticana) and their interactions with the defensive metabolites of the glucosinolate - myrosinase - isothiocyanate system. BMC Plant Biol 18(1):85. https://doi.org/10.1186/s12870-018-1295-4
Tautenhahn R, Patti GJ, Rinehart D, Siuzdak G (2012) XCMS Online: a web-based platform to process untargeted metabolomic data. Anal Chem 84(11):5035–5039. https://doi.org/10.1021/ac300698c
Taware R, Abnave P, Patil D, Rajamohananan PR, Raja R, Soundararajan G, Kundu GC, Kharat MKD, Pai K, Ahmad A (2015) Trichothecin from endophytic fungus Trichothecium sp. and its anticancer effect on murine melanoma and breast cancer cell lines. Curr Biochem Eng 2:73–80. https://doi.org/10.2174/2212711901666140804220845
Thompson MSA, Couce E, Webb TJ, Grace M, Cooper KM, Schratzberger M (2021) What’s hot and what’s not: making sense of biodiversity ‘hotspots.’ Glob Chang Biol 27(3):521–535. https://doi.org/10.1111/gcb.15443
Toghueo RMK, Sahal D, Boyom FF (2020) Recent advances in inducing endophytic fungal specialized metabolites using small molecule elicitors including epigenetic modifiers. Phytochem 174:112338. https://doi.org/10.1016/j.phytochem.2020.112338
Treen DGC, Wang M, Xing S et al (2022) (2022) SIMILE enables alignment of tandem mass spectra with statistical significance. Nat Commun 13:2510. https://doi.org/10.1038/s41467-022-30118-9
Triastuti A, Vansteelandt M, Barakat F, Trinel M, Jargeat P, Fabre N, Amasifuen Guerra CA, Mejia K, Valentin A, Haddad M (2019) How histone deacetylase inhibitors alter the secondary metabolites of Botryosphaeria mamane, an endophytic fungus isolated from Bixa orellana. Chem Biodivers 16(4):e1800485. https://doi.org/10.1002/cbdv.201800485
Tripathi A, Pandey P, Tripathi SN, Kalra A (2022) Perspectives and potential applications of endophytic microorganisms in cultivation of medicinal and aromatic plants. Front Plant Sci 29(13):985429. https://doi.org/10.3389/fpls.2022.985429
Tsugawa H, Cajka T, Kind T, Ma Y, Higgins B, Ikeda K, Kanazawa M, VanderGheynst J, Fiehn O, Arita M (2015) MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat Methods 12(6):523–526. https://doi.org/10.1038/nmeth.3393
Ul-Hassan SR, Strobel GA, Booth E, Knighton B, Floerchinger C, Sears J (2012) Modulation of volatile organic compound formation in the Mycodiesel-producing endophyte Hypoxylon sp. CI-4. Microbiology 158(Pt 2):465–473. https://doi.org/10.1099/mic.0.054643-0
van Tilburg BE, Pettersen VK, Gutierrez MW, Laforest-Lapointe I et al (2020) Intestinal fungi are causally implicated in microbiome assembly and immune development in mice. Nat Commun 11(1):2577. https://doi.org/10.1038/s41467-020-16431-1
Wang Y, Zeng J (2013) Predicting drug-target interactions using restricted boltzmann machines. Bioinform 29(13):i126–i134. https://doi.org/10.1093/bioinformatics/btt234
Wang X, SenaFilho JG, Hoover AR, King JB, Ellis TK, Powell DR, Cichewicz RH (2010) Chemical epigenetics alters the secondary metabolite composition of guttate excreted by an atlantic-forest-soil-derived Penicillium citreonigrum. J Nat Prod 73(5):942–948. https://doi.org/10.1021/np100142h
Wang M, Carver JJ, Phelan VV, Sanchez LM, Garg N, Peng Y, Nguyen DD, Watrous J, Kapono CA, Luzzatto-Knaan T, Porto C, Bouslimani A, Melnik AV, Meehan MJ (2016) Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat Biotechnol 34(8):828–837. https://doi.org/10.1038/nbt.3597
Wang X, Shen Y, Wang S, Li S, Zhang W, Liu X, Lai L, Pei J, Li H (2017) PharmMapper 2017 update: a web server for potential drug target identification with a comprehensive target pharmacophore database. Nucleic Acids Res 45(W1):W356–W360. https://doi.org/10.1093/nar/gkx374
Wang J, Xu Z, Hu X, Yang Y, Su J, Liu Y, Zhou L, Qin J, Zhang D, Yu H (2020) Epoxycytochalasin H: an endophytic Phomopsis compound induces apoptosis in A2780 cells through mitochondrial damage and endoplasmic reticulum stress. Onco Targets Ther 3(13):4987–4997. https://doi.org/10.2147/OTT.S253716
Wang X, Liu Y, Kang N, Xu G (2023) Wide identification of chemical constituents in fermented licorice and explore its efficacy of anti-neurodegeneration by combining quasi-targeted metabolomics and in-depth bioinformatics. Front Neurosci 18(17):1156037. https://doi.org/10.3389/fnins.2023.1156037
Wei J, Chen F, Liu Y, Abudoukerimu A, Zheng Q, Zhang X, Sun Y, Yimiti D (2020) Comparative metabolomics revealed the potential antitumor characteristics of four endophytic fungi of Brassica rapa L. ACS Omega 5(11):5939–5950. https://doi.org/10.1021/acsomega.9b04258
Wei Q, Bai J, Yan D, Bao X, Li W, Liu B, Zhang D, Qi X, Yu D, Hu Y (2021) Genome mining combined metabolic shunting and OSMAC strategy of an endophytic fungus leads to the production of diverse natural products. Acta Pharm Sin B 11(2):572–587. https://doi.org/10.1016/j.apsb.2020.07.020
Wen B, Mei Z, Zeng C, Liu S (2017) metaX: a flexible and comprehensive software for processing metabolomics data. BMC Bioinform 18(1):183. https://doi.org/10.1186/s12859-017-1579-y
Wishart DS (2008) Quantitative metabolomics using NMR. TrAC Trends Anal Chem 27(3):228–237. https://doi.org/10.1016/j.trac.2007.12.001
Wishart DS, Knox C, Guo AC, Shrivastava S, Hassanali M, Stothard P, Chang Z, Woolsey J (2006) DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res 34(Database issue):D668–D672. https://doi.org/10.1093/nar/gkj067
Wong C, Ling YS, Wee JLS, Mujahid A, Müller M (2020) A comparative UHPLC-Q/TOF-MS-based eco-metabolomics approach reveals temperature adaptation of four Nepenthes species. Sci Rep 10(1):21861. https://doi.org/10.1038/s41598-020-78873-3
Wu LS, Hu CL, Han T, Zheng CJ, Ma XQ, Rahman K, Qin LP (2013) Cytotoxic metabolites from Perenniporia tephropora, an endophytic fungus from Taxus chinensis var. mairei. Appl Microbiol Biotechnol 97(1):305–315. https://doi.org/10.1007/s00253-012-4189-7
Wu LS, Jia M, Chen L, Zhu B, Dong HX, Si JP, Peng W, Han T (2015) Cytotoxic and antifungal constituents isolated from the metabolites of endophytic fungus DO14 from Dendrobium officinale. Molecules 21(1):E14. https://doi.org/10.3390/molecules21010014
Xia J, Psychogios N, Young N, Wishart DS (2009) MetaboAnalyst: a web server for metabolomic data analysis and interpretation. Nucleic Acids Res 37(Web Server issue):W652–W660. https://doi.org/10.1093/nar/gkp356
Xia J, Sinelnikov IV, Han B, Wishart DS (2015) MetaboAnalyst 3.0–making metabolomics more meaningful. Nucleic Acids Res 43(W1):W251–W257. https://doi.org/10.1093/nar/gkv380
Xie J, Wu YY, Zhang TY, Zhang MY, Zhu WW, Gullen EA, Wang ZJ, Cheng YC, Zhang YX (2017) New and bioactive natural products from an endophyte of Panax notoginseng. RSC Adv 7:38100–38109. https://doi.org/10.1039/c7ra07060h
Xu D, Zhang BY, Yang XL (2016) Antifungal monoterpene derivatives from the plant endophytic fungus Pestalotiopsis foedan. Chem Biodivers 13(10):1422–1425. https://doi.org/10.1002/cbdv.201600114
Xu R, Lee J, Chen L, Zhu J (2021) Enhanced detection and annotation of small molecules in metabolomics using molecular-network-oriented parameter optimization. Mol Omics 17(5):665–676. https://doi.org/10.1039/d1mo00005e
Xue M, Hou X, Fu J, Zhang J, Wang J, Zhao Z, Xu D, Lai D, Zhou L (2023) Recent advances in search of bioactive secondary metabolites from fungi triggered by chemical epigenetic modifiers. J Fungi 9(2):172. https://doi.org/10.3390/jof9020172
Yang H, Wang Y, Zhang Z, Yan R, Zhu D (2014a) Whole-genome shotgun assembly and analysis of the genome of Shiraia sp. strain Slf14, a novel endophytic fungus producing huperzine A and hypocrellin A. Genome Announc 2:e00011–e00014
Yang Y, Zhao H, Barrero RA, Zhang B, Sun G, Wilson IW, Xie F, Walker KD, Parks JW, Bruce R et al (2014b) Genome sequencing and analysis of the paclitaxel-producing endophytic fungus Penicillium aurantiogriseum NRRL 62431. BMC Genom 15:69
Yang WC, Bao HY, Liu YY, Nie YY, Yang JM, Hong PZ, Zhang Y (2018) Depsidone derivatives and a cyclopeptide produced by marine fungus Aspergillus unguis under chemical induction and by its plasma induced mutant. Molecules 23(9):2245. https://doi.org/10.3390/molecules23092245
Yang Y, Liu X, Cai J, Chen Y, Li B, Guo Z, Huang G (2019) Genomic characteristics and comparative genomics analysis of the endophytic fungus Sarocladium brachiariae. BMC Genomics 20(1):782. https://doi.org/10.1186/s12864-019-6095-1
Zaher AM, Makboul MA, Moharram AM, Tekwani BL, Calderón AI (2015) A new enniatin antibiotic from the endophyte Fusarium tricinctum Corda. J Antibiot 68:197–200. https://doi.org/10.1038/ja.2014.129
Zhang G, Wang W, Zhang X, Xia Q, Zhao X, Ahn Y, Ahmed N, Cosoveanu A, Wang M, Wang J et al (2015) De novo RNA sequencing and transcriptome analysis of Colletotrichum gloeosporioides ES026 reveal genes related to biosynthesis of huperzine A. PLoS ONE 10:e0120809
Zhang X, Wang Z, Jan S, Yang Q, Wang M (2017) Expression and functional analysis of the lysine decarboxylase and copper amine oxidase genes from the endophytic fungus Colletotrichum gloeosporioides ES026. Sci Rep 7:2766
Zhang J, Zhu Y, Si J, Wu L (2022) Metabolites of medicine food homology-derived endophytic fungi and their activities. Curr Res Food Sci 7(5):1882–1896. https://doi.org/10.1016/j.crfs.2022.10.006
Zhou J, Zhong L (2022) Applications of liquid chromatography-mass spectrometry based metabolomics in predictive and personalized medicine. Front Mol Biosci 3(9):1049016. https://doi.org/10.3389/fmolb.2022.1049016
Zhou B, Xiao JF, Tuli L, Ressom HW (2012) LC-MS-based metabolomics. Mol Biosyst 8(2):470–481. https://doi.org/10.1039/c1mb05350g
Zhu W, Han H, Liu A, Guan Q, Kang J, David L, Dufresne C, Chen S, Tian J (2021) Combined ultraviolet and darkness regulation of medicinal metabolites in Mahonia bealei revealed by proteomics and metabolomics. J Proteomics 20(233):104081. https://doi.org/10.1016/j.jprot.2020.104081
Zou K, Liu X, Hu Q, Zhang D, Fu S, Zhang S, Huang H, Lei F, Zhang G, Miao B et al (2021) Root endophytes and Ginkgo biloba are likely to share and compensate secondary metabolic processes, and potentially exchange genetic information by LTR-RTs. Front Plant Sci 12:704985
Zutz C, Gacek A, Sulyok M, Wagner M, Strauss J, Rychli K (2013) Small chemical chromatin effectors alter secondary metabolite production in Aspergillus clavatus. Toxins 5(10):1723–1741. https://doi.org/10.3390/toxins5101723
Acknowledgements
We would like to thank Dr. Andrei Stecca Steindorff and M.Sc. Leila Lorenço Furtado for their excellent advice during the preparation of this work. We are grateful to the Coordination for the Improvement of Higher Education Personnel—Brazil (CAPES)—Financing Code 001, and University of Brasília DPG Nº 0010/2023.
Funding
This study was sponsored by grants from the Fundação Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brazil (CAPES)—Funding code 001, and University of Brasília DPG Nº 0010/2023.
Author information
Authors and Affiliations
Contributions
All authors contributed to the preparation of the work. Jefferson Brendon Almeida dos Reis conceived the idea for the manuscript. The first version of the manuscript and all the figures were prepared by Jefferson Brendon Almeida dos Reis. Patrícia Cardoso Cortelo and Adriana Sturion Lorenzi read the first version and commented. Helson Mario Martins do Vale and Danilo Batista Pinho read subsequent versions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that the research was conducted in the absence of any commercial or financial relationship that could be construed as a potential conflict of interest.
Additional information
Communicated by Yusuf Akhter.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
dos Reis, J.B.A., Lorenzi, A.S., Pinho, D.B. et al. The hidden treasures in endophytic fungi: a comprehensive review on the diversity of fungal bioactive metabolites, usual analytical methodologies, and applications. Arch Microbiol 206, 185 (2024). https://doi.org/10.1007/s00203-024-03911-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-024-03911-x