Abstract
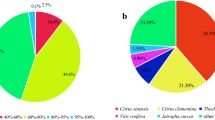
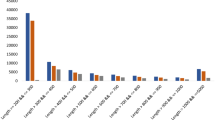
In the present study, we identified genes that are putatively involved in the production of fungal 10-hydroxycamptothecin via transcriptome sequencing and characterization of the Xylaria sp. M71 treated with salicylic acid (SA). A total of 60,664,200 raw reads were assembled into 26,044 unigenes. BLAST assigned 8,767 (33.7%) and 10,840 (41.6%) unigenes to 40 Gene Ontology (GO) annotations and 108 Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, respectively. A total of 3,713 unigenes comprising 1,504 upregulated and 2,209 downregulated unigenes were found to be differentially expressed between SA-induced and control fungi. Based on the camptothecin biosynthesis pathway in plants, 13 functional genes of Xylaria sp. M71 were mapped to the mevalonate (MVA) pathway, suggesting that the fungal 10-hydroxycamptothecin is produced via the MVA pathway. In summary, analysis of the Xylaria sp. M71 transcriptome allowed the identification of unigenes that are putatively involved in 10-hydroxycamptothecin biosynthesis in fungi.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Burlat, V., Oudin, A., Courtois, M., Rideau, M., and St-Pierre, B. 2004. Co-expression of three MEP pathway genes and geraniol 10-hydroxylase in internal phloem parenchyma of Catharanthus roseus implicates multicellular translocation of intermediates during the biosynthesis of monoterpene indole alkaloids and isoprenoid-derived primary metabolites. Plant J. 1, 131–141.
Cho, H.Y., Son, S.Y., Rhee, H.S., Yoon, S.Y., Lee-Parsons, C.W., and Park, J.M. 2008. Synergistic effects of sequential treatment with methyl jasmonate, salicylic acid and yeast extract on benzophenanthridine alkaloid accumulation and protein expression in Eschscholtzia californica suspension cultures. J. Biotechnol. 1, 117–122.
Conesa, A., Gotz, S., Garcia-Gomez, J.M., Terol, J., Talon, M., and Robles, M. 2005. Blast2GO: a universal tool for annotation, visualization and analysis in functionalgenomics research. Bioinformatics 18, 3674–3676.
Fox, A.R., Soto, G., Mozzicafreddo, M., Garcia, A.N., Cuccioloni, M., Angeletti, M., Salerno, J.C., and Ayub, N.D. 2014. Understanding the function of bacterial and eukaryotic thiolases II by integrating evolutionary and functional approaches. Gene 1, 5–10.
Haas, B.J., Papanicolaou, A., Yassour, M., Grabherr, M., Blood, P.D., Bowden, J., Couger, M.B., Eccles, D., Li, B., Lieber, M., et al. 2013. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512.
Hoang, M.H., Nguyen, C., Pham, H.Q., Nguyen, L.V., Duc, L.H., Son, L.V., Hai, T.N., Ha, C.H., Nhan, L.D., Anh, H.T.L., et al. 2016. Transcriptome sequencing and comparative analysis of Schizochytrium mangrovei PQ6 at different cultivation times. Biotechnol. Lett. 38, 1781–1789.
Kusari, S., Zühlke, S., and Spiteller, M. 2009. An endophytic fungus from Camptotheca acuminata that produces camptothecin and analogues. J. Nat. Prod. 1, 2–7.
Liu, K.H., Ding, X.W., Deng, B.W., and Chen, W.Q. 2010. 10-Hydroxycamptothecin produced by a new endophytic Xylaria sp., M20, from Camptotheca acuminata. Biotechnol. Lett. 5, 689–693.
Martín, J., García-Estrada, C., Rumbero, A., Recio, E., Albillos, S.M., Ullán, R.V., and Martín, J.F. 2011. Characterization of an autoinducer of penicillin biosynthesis in Penicillium chrysogenum. Appl. Environ. Microbiol. 16, 5688–5696.
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A.C., and Kanehisa, M. 2007. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, 182–185.
Robin, K.P. 2011. Small-molecule elicitation of microbial secondary metaboliites. Microb. Biotechnol. 4, 471–478.
Sun, J., Sun, X.X., Tang, P.W., and Yuan, Q.P. 2012. Molecular cloning and functional expression of two key carotene synthetic genes derived from Blakeslea trispora into E. coli for increased β-carotene production. Biotechnol. Lett. 11, 2077–2082.
Sun, Y.Z., Luo, H.M., Li, Y., Sun, C., Song, J.Y., Niu, Y.J., Zhu, Y.J., Dong, L., Lv, A. Tramontano, E., et al. 2011. Pyrosequencing of the Camptotheca acuminata transcriptome reveals putative genes involved in camptothecin biosynthesis and transport. BMC Genomics 12, 533.
Ye, J., Fang, L., Zheng, H.K., Zhang, Y., Chen, J., Zhang, Z., Wang, J., Li, S., Li, R., and Bolund, L. 2006. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res. 34, W293–W297.
Yu, G.J., Wang, M., Huang, J., Yin, Y.L., Chen, Y.J., Jiang, S., Jin, Y.X., Lan, X.Q., Wong, B.H., Liang, Y., et al. 2012. Deep insight into the Ganoderma lucidum by comprehensive analysis of its transcriptome. PLoS One 7, e44031.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Ding, X., Liu, K., Zhang, Y. et al. De novo transcriptome assembly and characterization of the 10-hydroxycamptothecin-producing Xylaria sp. M71 following salicylic acid treatment. J Microbiol. 55, 871–876 (2017). https://doi.org/10.1007/s12275-017-7173-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-017-7173-1