Abstract
Variations at morphological, cytogenetical, cytochemical, biochemical, and molecular levels have been reported in cell, callus cultures, clonally propagated plants, and in regenerated plants in some plant species. Metabolic instability in the genetically manipulated transgenic cell lines with respect to secondary metabolite production has been reported by different authors in long-term in vitro culture, although in a few cases the transgenic nature of the cell lines was retained. Transgenic hairy root cultures are another promising way of production of commercially valuable secondary metabolites in vitro, which open up a new dimension of the role of plant tissue culture in secondary metabolite production. Hairy root cultures and the plants regenerated from transformed roots are well known for their cytogenetical, morphological, and biochemical stability when compared to cell suspension cultures and callus cultures. But there are very few studies on the stability of hairy roots under long-term cultural condition. Variability in Ri-transformed root cultures and regenerated Ri-transformed plants has also been reported in a few species. In the present review, the stability/instability of hairy root cultures and Ri-plants maintained in vitro for long term is discussed in detail.
Access provided by CONRICYT-eBooks. Download reference work entry PDF
Similar content being viewed by others
Keywords
- Agrobacterium rhizogenes
- Genetic stability
- Cytological stability
- Clonal fidelity
- In vitro culture
- Ri-transformed roots
- Ri-transformed plants
1 Introduction
Genetic instability is common in the cell cultures of many species [1–5]. Variations at the morphological level, cytological level (chromosome number and structure), cytochemical level (genome size), biochemical level (secondary metabolites and isozymes), and molecular levels in cells, callus cultures, and in regenerated plants have been reported [6–12]. Variation in clonally propagated plants as well as in plants regenerated directly from explants is also demonstrating such variation [8, 13, 14]. In few species, stability of callus cultures and plants regenerated from callus has been reported in long-term cultures [15–18]. Cytological abnormalities include changes in chromosome number, chromosome rearrangements, deletions, duplications, sequence change play major roles in the development of such variations [4, 6, 7, 19]. The role of gene activation and silencing, epigenetic changes in the development of variants in long-term tissue culture has been reported [13, 20].
Plant cell and tissue cultures are promising renewable alternative source of commercially valuable secondary metabolites, generally for those complex molecules for which chemical synthesis is not viable economically. Interest regarding these biotechnological techniques increased when it was found that undifferentiated cell suspension cultures or differentiated organ cultures synthesize and accumulate specific secondary metabolites at a similar or higher amount as compared to parent plants [21, 22]. However, for production of secondary metabolites in commercially useful amounts, optimization of culture conditions, selection of fast growing high-yielding cell lines, precursor feeding, elicitation, immobilization, and genetic transformation techniques have been utilized in different species [23, 24].
Plant cell suspension cultures of medicinally important species have been used as a promising alternative “chemical factory” for the production of industrially important pharmaceuticals. The major drawback of cell suspension culture is the spontaneous changes in the physiological, morphological, biochemical, and the cytological behavior of cells that leads to instability in secondary metabolite production during long-term culture [25, 26]. Cytological heterogeneity may be represented by mixture of euploid and aneuploid cell s with normal cells in cell suspension culture and callus cultures [5, 27–29]. Variations may arise at the structural level of chromosomes without affecting chromosome number. Detection of such kind of cytological instability is comparatively difficult as the culture cells show normal chromosome number. The high degree of such change has been documented in long-term tissue culture. Karyotypic changes due to chromosomal translocations have been observed [4, 30]. Culture induced phenotypic, biochemical, and genetic variations were observed in regenerated plants and progenies of the regenerated plants. Various analyses have revealed that these variants show genetic behavior similar to the naturally occurring mutant [4, 31]. Regeneration of plants may involve de-differentiation and redifferentiated during which different variables can develop.
Transgenic cell cultures of Vitis amurensis and Catharanthus have been developed by genetic manipulation for production of target secondary metabolites [32, 33]. Instability in transgenic cell cultures with respect to secondary metabolite production is reported by different authors [32, 33]. Metabolic instability of such several transgenic cell lines of Catharanthus overexpressing key enzymes of TIA pathway was observed in the long-term in vitro culture when studied for 30 months, although the transgenic nature of the cell lines was retained [32]. Drastic decrease in metabolite content in the transformed cell lines is also reported in grapes by Dubrovina and Kiselev [33]. rolB transformed cell lines were established in V. amurensis accumulating high levels of resveratrol showed a decline in the resveratrol content in long-term culture with regular subculturing. Stable expression of rolB gene in the transgenic cell cultures has been reported after 5 years of transformation by the same group [34]. In in vitro micropropagated clones of transgenic birch, decrease in expression of foreign genes with increase in number of subcultures has been reported [35]. Instability of the foreign gene is not restricted to transgenic cell cultures but also reported in some hairy root cultures and transgenic plants [36]. Integration of foreign gene/genes into the host plant genome might cause alteration in its structure and thus affecting the host plant and/or expression of the transgene by gene silencing or negative influence of the flanking plant DNA and location in the chromosome [37, 38].
Transgenic hairy root cultures have opened up a new dimension to application of organ cultures for synthesis, accumulation, and regulation of secondary metabolites production in vitro due to their rapid growth in simple media without phytohormone and easier maintenance in long-term culture. Agrobacterium rhizogenes , a Gram negative soil bacterium is responsible for causing hairy root disease in higher plants by transfer of T-DNA present in Ri-plasmid of the bacteria to the plant host genome. Root loci A-D (rol A-D) of the T-DNA are responsible for the development of hairy root phenotype [22, 39]. Such hairy roots are also capable of spontaneously regenerating plants (Ri-transformed plants) in a number of species [22, 40]. The hairy roots and Ri-transformed plants derived from them are known to synthesize important plant secondary metabolites in enhanced or similar levels to the nontransformed plants [22]. Bulgakov et al. [41] suggested that rol genes (rolB and rolC) of Ri plasmid are potential activators of secondary metabolism, activate phytoalexin production and suppress intracellular ROS level, the combination of defense responses and effect of ROS suppression affects sensitivity towards auxin, growth and metabolism of the transformed tissue [41].
Hairy root cultures are well known for their cytogenetical, morphological, and biochemical stability compared to cell suspension cultures and callus cultures. But there are very few studies on the stability of hairy roots under long-term cultural condition. In this chapter, we have discussed regarding the stability and variability of hairy root culture s and regenerated plants of different species on the basis of morphological, biochemical, and cytological analysis reported as far as possible. Genetic stability on the basis of integration and expression of T-DNA genes in transgenic cultures and clonal fidelity of hairy root cultures and Ri-transformed plants are also discussed in this review.
2 Characterization of Ri-transformed Roots
Hairy root cultures are characterized by profuse lateral branching and rapid root tip elongation with plagiotropic growth in hormone free medium [42]. Apart from the wild type transformed hairy roots, foreign genes can be also inserted into the hairy roots in many plants [43]. The altered morphological characteristics of the hairy root cultures enable marker free selection of the transformed root lines which are advantageous over use of A. tumefaciens-mediated transformation [40, 42].
A. rhizogenes-mediated transformation and transformed root cultures have been established in numerous plant species, including many medicinally important plants [22, 44] as an alternative source for the production of important plant secondary metabolites [22]. Transformed root lines are reported to vary in morphology, biomass accumulation, integration of T-DNA genes, and biosynthetic ability in a species [45–52]. Such variations in between the hairy root lines of the same plant species are suggested to be due to variation in the integration of T-DNA genes of Ri-plasmid into the plant genome [50, 53, 54].
Several reports are available on the effects of the TR and TL T-DNAs on the growth, morphology, and secondary metabolite production in transformed roots [46–50, 55, 56]. For example, in Catharanthus roseus, rolAB + /ags + and rolAB + /ags − root clones belonged to four different morphological types, whereas rolAB − /ags + root clones were of slow growing and callusing morphology [46]. In coffee hairy roots, rolB and rolC genes were reported to be systematically integrated; however, the presence of rolA and rolD genes could not be related to the morphological variability. The correlation could not be found between the presence of T-DNA genes and hairy root lines showing altered morphology in Coffea arabica [49]. Similarly, in Tylophora indica no direct correlation was found between the presence of T-DNA genes and transformed root morphology [52]. Differential loss of T-DNA genes in hairy root lines of C. roseus and the effect of such loss on the morphology and the biosynthetic ability of the root lines are reported by Taneja et al. [50]. Ten Ri-transformed root lines studied for the presence of 23 ORFs of the T-DNA showed loss of few ORFs that drastically affected the growth, morphology, and alkaloid biosynthesis in transformed root lines [50]. The induction and development of the hairy root phenotypes are affected by synergistic activity of the rol genes expressed simultaneously [57–59].
The study of stability of such variant Ri-transformed root lines in the long-term in vitro culture is reported only in a few plant species. The details of such reports on morphological, biosynthetic, cytological, and molecular stability of A. rhizogenes-transformed root cultures in various species are discussed in detail in the following section of this review.
2.1 Morphological Stability of Ri-transformed Roots
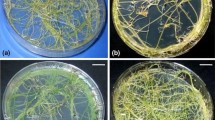
There are contradictory reports regarding the extent of stability in transformed roots maintained in vitro during prolonged culture. Stability in growth and morphological characters over prolonged culture is reported in different species like – Lycopersicon esculentum [60], Cinchona officinalis “Ledgeriana” [61], Aesculus hippocastanum L. [62], Pimpinella anisum [63], Coffee [49], T. indica [52], Arachis hypogaea [51], Plumbago zeylanica L. [64], Rauvolfia serpentina [65, 66], etc. Morphology of different Ri-transformed root lines of different plant species is known to vary to a large extent (Fig. 1).
Ri-transformed root lines of different plant species transformed with wild type Agrobacterium rhizogenes strains, maintained for long term in vitro culture for more than 5–8 years (Bar = 1 cm) – (a, c–g) LBA 9402 transformed root cultures of Arabidopsis thaliana (Photo P. Paul), Convolvulus arvensis (Photo A. Majumder), Digitalis purpurea (Photo A. Basu), Nicotiana tabacum (Photo M. Halder), Plumbago zyelanica (Photo A. Basu), and Rauvolfia serpentina (Photo Smita Ray), respectively. (b, h, i) A4 transformed root culture of Arachis hypogaea var. JL-24 (Photo M. Halder), Tylophora indica (Photo D. Roychowdhury), and Withania somnifera (Photo Swagata Ray), respectively
Long-term stability of Ri-transformed root cultures in L. esculentum is reported by Lipp Jao and Brown [60]. Transformed root clones established with A. rhizogenes strain R1601 were maintained in solid and liquid media. The hairy roots were found to retain their growth and characteristic phenotype for 50 passages over 25 months in liquid culture and for 12 passages over 12 months in solid culture. The growth rate of these root clones was not affected by the presence or absence of selective agent in the solid media, thus not necessary for maintaining the transformed state in long-term culture as suggested by the authors [60].
Stability in growth and morphology of hairy root cultures of C. officinalis “Ledgeriana” after 1 year of in vitro culture is reported [61]. Similar unchanged growth rate of hairy root cultures of A. hippocastanum L. after 4 years of in vitro culture is reported by Zdravković-Korać et al. [62].
Santos et al. [63] studied the morphological stability of hairy root lines of P. anisum in four different media. Root lines growing on SH medium showed stable morphological characters over the period of study compared to the other three media where marked changes were observed [63].
Alpizar et al. [49] reported that exogenous auxin supplementation was essential for proliferation of hairy root lines of coffee. Sixty two root clones varied significantly in two growth parameters (total root length and frequency of fine roots), and the characteristic phenotypes were stable over the subcultures for over 3 years [49].
In T. indica, four morphologically distinct phenotypes observed in A. rhizogenes strain A4 transformed root lines were stable when maintained for more than 4 years on hormone free MS basal medium [52].
In peanut (A. hypogaea) cv. JL-24, Halder and Jha [51] reported strain independent variability among 30 Ri-transformed root lines on the basis of morphology, biomass accumulation, and trans-resveratrol content. Root lines and clones maintained in vitro for over 3 years showed stable morphological characters as well as growth index value [51].
Growth and morphology of LBA9402 transformed root lines of P. zeylanica cultured on solid modified MS medium were observed to be stable for 2 years in in vitro culture [64].
Ray et al. [66] reported retention of stable phenotype of Ri-transformed root lines of R. serpentina over 3 years of in vitro culture. The phenotype of transformed root lines included creamish roots with a high degree of branching, plagiotropic growth, and devoid of extensive root hairs [66]. Long-term stability of hairy root cultures of R. serpentina is also reported by Pandey et al. [65] for more than 6 years of in vitro culture. The growth kinetic analysis of R. serpentina hairy roots exhibited higher growth potential following long-term cultivation with either of the two carbohydrates, viz., sucrose and table sugar [65].
However, instability in morphological phenotype and growth kinetics of hairy roots in prolonged in vitro culture is reported in certain species like carrot [36], Brugmansia candida [67], Duboisia myoporoides [68], Hyoscyamus muticus [69], etc.
Instability in phenotype and expression of transgenes were observed in hairy root lines of Daucus carota in long-term culture [36]. Different phenotypes and growth patterns were observed between hairy root clones of carrot and between the subcultures of single root clones [36]. Significant decrease in growth of hairy root cultures is reported in B. candida on prolonged subculture for 5 years [67]. Yukimune et al. [68] performed repeated selection in hairy root cultures of D. myoporoides and observed that the morphology of the transformed roots with improved scopolamine content differed after the repeated selection, viz., fine root lines with extensive lateral branching. In H. muticus, variation was observed between the different hairy root lines and hairy root clones derived from protoplast culture [69].
Thus, it is evident from the above reports that the majority of the plant species studied to observe the long-term morphological characters and growth of hairy roots in culture conditions showed stability. And only certain species are found to behave differently in the long-term in vitro cultures with a decrease in growth kinetics and change in morphology. Type of growth media can be a deciding factor for the phenotypic stability of A. rhizogenes-transformed root cultures as observed in P. anisum [63].
2.2 Biochemical Stability of Ri-transformed Roots
The utility of hairy root culture for synthesizing important secondary metabolites greatly depends on the biochemical stability of the high yielding roots in the long-term in vitro culture. The stable production of important plant metabolites by the Ri-transformed hairy roots after prolonged culture period is reported in different plants like Beta vulgaris, Nicotiana rustica [70], Datura stramonium [71, 72], D. innoxia [73], H. muticus [69, 74], P. anisum [63], T. indica [52], R. serpentina [65], etc.
Biosynthetic stability of hairy roots of B. vulgaris and N. rustica after long-term cryopreservation is reported by Benson and Hamill [70]. The total betalain (total betaxanthin and betacyanin pigments) in B. vulgaris and total alkaloid production in N. rustica was found to be stable after the recovery of the cryopreserved hairy roots suggesting biosynthetic stability of these Ri-transformed root lines [70].
Biosynthetic stability of hairy root lines in the long-term in vitro culture is also reported in D. stramonium [71]. Very high stability in total alkaloid production for over 5 years, i.e., over 75 subcultures, was observed in the high alkaloid producing root lines [71]. This finding was advantageous over the high level of instability observed in normal root cultures of D. stramonium upon subculturing [75]. Baíza et al. [72] also reported biochemical stability of the hairy root lines of D. stramonium over a period of 6 years. On contrary, only one root line of D. stramonium showed great instability in alkaloid production through time, with regard to the production of both hyoscyamine and scopolamine [72, 75]. Dechaux and Boitel-Conti [73] reported a decrease in scopolamine levels similar to control levels after 1 year of subculture in hairy roots of D. innoxia overexpressing H6H gene from H. niger.
Stable secondary metabolite production after 5 years of in vitro culture is reported in hairy roots of H. muticus. All the studied Ri-transformed root lines of H. muticus showed stable alkaloid production during the long-term culture [69]. Stability of secondary metabolite production in hairy roots of H. muticus during two and a half years of culture is also reported by Jouhikainen et al. [74].
Hairy root cultures of P. anisum, maintained in the SH medium, both under darkness and photoperiod conditions showed stable essential oil production [63].
Hairy root cultures of T. indica established by using A. rhizogenes strain A4 showed biochemical stability over a period of 4 years. The transformed root lines of different phenotypes accumulated tylophorine at higher levels as compared to the to nontransformed roots, and the potential of enhanced growth and tylophorine content was stably maintained in long-term in vitro culture [52].
R. serpentina hairy roots (Ri-A4 transformed roots) produce considerably higher reserpine content than the normal roots after 10 weeks of growth [76]. Ray et al. [66] reported higher reserpine (approx 3 mg g−1DW) content in the LBA 9402 root lines. Pandey et al.[65] reported the production of all three major terpene alkaloid in R. serpentina, viz., reserpine, ajmaline , yohimbine since the establishment of cultures with yohimbine concentration being highest followed by ajmaline and reserpine. After 6 years of maintenance on both sucrose and table sugar containing media, the selected root culture of R. serpentina maintained similar trend of production, yohimbine appeared to be the major alkaloid. Root cultures showed differential yields of the alkaloids in the two carbon sources used. The hairy roots of R. serpentina showed 193.8% more reserpine production in the table sugar supplemented media compared to media supplemented with sucrose. However, the yield of yohimbine and ajmaline was 38.32% and 61.98% higher in the sucrose supplemented media than the table sugar supplemented one. These hairy root cultures of R. serpentina showed escalation of secondary metabolite production in long-term culture over the 6 years of study [65].
Increase in secondary metabolite production over a long period of subculture of Ri-transformed roots is reported in plant species like B. candida [67] and D. myoporoides [68]. Marconi et al. [67] reported increasing secondary metabolite production after 5 years of in vitro culture of B. candida hairy roots, possibly due to the prolonged in vitro culture mediated stress. In the long-term in vitro culture, a pronounced increase in the production of scopolamine was observed in such root lines [67]. In D. myoporoides, repeated selection in transformed root cultures was done and it was observed that scopolamine content of the hairy root lines obtained at each selection increased with the number of selections [68].
On the contrary, complete loss of capacity to accumulate alkaloids after 1 year of in vitro culture of hairy roots is reported in C. officinalis “Ledgeriana” [61]. These transgenic hairy roots containing tryptophan decarboxylase (Tdc) and strictosidine synthase (Str) from C. roseus, two key enzymes in terpenoid indole and quinoline alkaloid biosynthesis, showed increased levels of quinine and quinidine initially after transformation [61].
2.3 Cytological Stability of Ri-transformed Roots
Several reports suggested that long-term in vitro tissue culture causes somaclonal variations and chromosomal abnormality , including both numerical and structural alterations [1]. Unorganized cultures are more prone to chromosomal instability – aneuploidy and polyploidy [4, 10, 25] – than organized cultures. There are very few reports regarding the chromosomal status (genetic stability ) of long-termed hairy root culture.
Ambros et al. [77] reported chromosomal localization by in situ hybridization of Ri-T-DNA in five different Crepis capillaries (2n = 6) transformed root lines, and all the root lines showed normal diploid chromosome number. The first report of chromosomal stability in A. rhizogenes-transformed root cultures was by Aird et al. [78]. The hairy root lines were studied after 6–18 months of in vitro culture and were found to show stable, normal chromosome number in all the seven plant species, viz., C. roseus (2n = 16), D. stramonium (2n = 24), N. rustica (2n = 48), N. umbratica (2n = 46), N. africana (2n = 46), Phaseolus vulgaris (2n = 22), and B. vulgaris (2n = 18) [78]. Transformed root cultures of D. stramonium showed typical karyotype and chromosome number same as in the control plants [72]. Baíza et al. [72] studied the karyotypic stability of three lines of hairy roots with stable secondary metabolite production of D. stramonium compared to instability of nontransformed root cultures. The transformed root cultures growing in the absence of any phytohormones exclusively contained diploid cells with 2n = 24, whereas the nontransformed root cultures whose growth needs exogenous hormone supplementation showed presence of mixoploidy and aneusomaty . This cytological stability of the hairy root lines of D. stramonium was maintained irrespective of the age of the transformed cultures [72]. In Swainsona galegifolia, untransformed root cultures were found to contain 90% diploid (2n = 32) cells, while A. rhizogenes transformed roots showed stability of diploid chromosome number [79].
Chromosomal stability in transformed hairy root cultures of Artemisia annua L. with normal diploid chromosome number 2n = 18 in all the LBA9402 transformed clones studied was reported by Mukherjee et al. [80]. Addition of growth regulators induced disorganization and dedifferentiation accompanied by loss of chromosomal stability in such cultures. Redifferentiation and rhizogenesis could be induced in such cultures in phytohormone free media and 90% of the regenerated roots showed diploid chromosome number (2n = 18) [80]. In R. serpentina, chromosome and karyotype analysis in transformed root lines maintained over 3 years in vitro was reported by Ray et al. [66] concluding that the karyotype of transformed root lines was similar to the roots of parent R. serpentina plants [66].
Cytological instability in hairy root cultures has been observed in some plant species such as Trifolium pretense, T. repens, Lotus corniculatus [81], Artemisia cina [82], Vicia faba [83], Lycopersicon peruvianum [84], potato [85], and Onobrychis viciaefolia [86].
Webb et al. [81] reported instability in anatomy, morphology, and cytology of Ri-transformed root lines of three legume species, namely T. pretense, T. repens, and L. corniculatus established using wild type strain C58C1 with pRi15834 of A. rhizogenes.
In A. cina, hairy root cultures established with three different A. rhizogenes strains showed instability in chromosome numbers along with the nontransformed ones after 6–12 months in culture. Nontransformed roots had a diploid chromosome number 2n = 32 in 53.7% of the cells, while the rest 46.3% cells showed chromosome number ranging from 2n = 22 to 64. The chromosome numbers of transformed roots were affected by different strains of A. rhizogenes used. Hairy roots established with strain 07–20001 showed the highest normal chromosome number (62.4%) followed by strain ATCC15834 (61.9%) and strain A4 (43%). The chromosome number range in transformed root lines was 2n = 11 to 2n = 66 [82].
In V. faba, 65 transformed root clones were studied, out of which 50% were polyploids and 6% were aneuploids or showed structural rearrangements. Polyploid root clones included octaploids, i.e., 2n = 8× = 48 [83]. Detailed karyotypic analysis of L. peruvianum hairy roots showed a diploid chromosome number with structural rearrangements [84]. Vries-Uijtewaal et al. [85] showed that hairy root clones developed from mono-haploid or di-haploid genotype were either diploid or tetraploid in potato. Ri-transformed roots of O. viciaefolia showed normal chromosome number 2n = 4× = 28 and spontaneously formed shoot buds developing into plants. However, the percentage of hairy root cells with a normal chromosome number reduced drastically with time, i.e., from 85% to 23% after 4 months and only 4% after 8 months. After 12 months of maintenance, roots with a normal chromosome number 2n = 28 was scarcely found with an increase in cells with 14 chromosomes [86]. The elimination of the normal chromosome number in hairy roots of O. viciaefolia was accompanied with loss of regeneration potential.
2.4 Integration and Expression of T-DNA Genes in Ri-transformed Roots
The T-DNA genes that are integrated and expressed during prolonged culture is a measure for assessment of transgenic nature of the hairy roots of different plants after repeated subcultures. Ri-transformed roots of different plant species have been reported to show the genetic stability of the Ri-T-DNA genes in long-term in vitro culture like in B. vulgaris, N. rustica [70], L. esculentum [60], C. officinalis “Ledgeriana”[61], A. hippocastanum L. [62], D. innoxia [73], coffee [49], T. indica [52], A. hypogaea [51], P. zeylanica L. [64], R. serpentina [65], etc.
Post-freeze molecular stability of T-DNA genes has been reported in Ri-transformed roots of B. vulgaris and N. rustica after short term and long-term recovery from cryopreservation [70]. The TL-DNA fragment was stably retained in hairy roots of both the species recovered from cryopreservation [70].
Molecular stability of hairy root lines of tomato is reported by Lipp Jao and Brown [60]. Transformed root lines of L. esculentum maintained for 50 passages in liquid culture and 12 passages in solid culture were reported to show the presence of the nptII gene by PCR and dot blot hybridization analyses. These root lines were also found to show NPTII enzyme activity in the long-term cultures [60].
The hairy root lines of C. officinalis “Ledgeriana” showing stable growth and morphology were found to completely lose the capacity of alkaloid production after 1 year. These root lines, however, showed the transgenic nature on PCR analysis, gus assay, and southern hybridization of the transgenes (Tdc and Str probes from C. roseus) after 1 year of maintenance. In Northern blot analysis, it was found that Tdc gene was not being expressed; only signal was detected for the GUS probe [61]. Therefore, hairy roots of C. officinalis showed stable integration of transgenes after 1 year, but the expression of one transgene was lost in long-term culture [61].
Transformed root lines of A. hippocastanum showed stable integration of rolA, rolB, rolC, and rolD genes by PCR analysis after 4 years of in vitro culture [62]. In D. innoxia, stable transgene expression is reported after 1 year of the subculture of the hairy roots by Dechaux and Boite-Conti [73].
Stable integration of rol genes has been reported in Ri-transformed root lines of coffee [49]. Fifty five Ri-transformed root lines maintained for over 3 years in vitro and characterized at the molecular level to study intra- and inter-clonal variability of the root lines showed to be TL+/TR− [49].
In T. indica transformed root lines were of two types; majority (87%) were TL+/TR−, while rest were TL+/TR+. Stable integration and expression of T-DNA genes of each clone were maintained in long-term in vitro culture [52].
Stable integration and expression of different rol genes were observed in 30 Ri-transformed root lines of A. hypogaea induced following infection with different strains of A. rhizogenes, after 2 years of maintenance in basal medium establishing genetic stability root lines and clones [51].
In Ri-transformed root lines of P. zeylanica, the rolA, rolB, rolC, and rolD genes were stably integrated, retained, and expressed at the transcription level as revealed by PCR and RT‐PCR analysis of 12–18-month-old cultures [64].
In R. serpentina, stable integration and expression of Ri-T-DNA genes have been reported in long-term in vitro culture by Pandey et al. [65]. The hairy root lines showed positive results for rolB and rolC after 6 years of maintenance on either of the carbon source used in media, viz., sucrose or table sugar [65].
Contradictorily, in hairy roots of certain plant species, instability of integrated transgene is reported. For example, instability in expression of transgene in in vitro culture of carrot hairy roots was reported by Guivarc’h et al. [36]. Reversible inactivation of the transgene expression was noted in one of the hairy root clones studied with a high copy number of the transgene [36]. In potato, frequent spontaneous deletion of Ri-T-DNA in hairy roots and regenerated plants was observed [87].
The study on the clonal fidelity of hairy root cultures in long-term in vitro culture can be an important criterion to assess the genetic stability of the transformed cultures. DNA fingerprinting profiles of 15 hairy root lines of T. indica along with nontransformed roots generated with 11 OPA primers after every 1 year for 4 years showed genetic stability and clonal fidelity of the clones of root lines in the long-term in vitro culture [52]. The study showed similarity in between the transformed roots and nontransformed ones at the genetic level in terms of the primers used, and the fingerprinting profiles for each primer did not vary with age [52].
3 Characterization of Ri-transformed Plants
Regeneration of Ri-transformed plants from transformed roots has been reported in many plant species, including medicinally important plants as reviewed earlier [22, 40]. These Ri-transformed plants exhibit unique characteristic features distinctly different from the nontransformed plants known as the “hairy root syndrome ” [88]. Ri-transformed plants are morphologically characterized by stunted growth with reduced shoot and internodes length, accompanied with the increase in the number of nodes, internodes and leaves (Fig. 2). However, the morphology of leaves is different from the nontransformed plants, i.e., the leaves are smaller in size and wrinkled in appearance. The decrease in shoot length is compensated by an increase in number of axillary branches. The root system is pronounced with decrease in the length of main root and an increase in the number of lateral branches giving rise to highly branched and extensive root system in the Ri-transformed plants. The roots are plageotropic and often seem to grow above the surface of the media. Presence of adventitious roots in the Ri-transformed plants is another feature noted. Apart from these characters, in some species, altered floral morphology, early flowering, and reduced pollen and seed production have also been reported in the transgenic plants regenerated from hairy roots. Additionally, conversion of biennial species to annuals was observed in Ri-transformed plants [42, 89, 90]. Hairy root syndromes of Ri-transformed plants are due to the insertion and expression of rol genes of the TL-DNA of Ri-plasmid. The alterations in morphological characters are caused by rolA, rolB, and rolC genes [42, 88].
Spontaneous regeneration of plants from root cultures of Nicotiana tabacum on phytohormone-free MS medium (Photos M. Halder) (a) nontransformed root culture showing spontaneous regeneration of shoot (Bar = 1 cm; arrow indicating regenerating shoot), (b) regenerated nontransformed plant growing on MS basal medium maintained for more than 3 years by regular subculture (Bar = 1 cm), (c) Ri-transformed root culture showing spontaneous regeneration of shoot (Bar = 1 cm, arrow indicating regenerating shoot), (d) regenerated Ri-transformed plant growing on MS basal medium maintained for more than 3 years by regular subculture (Bar = 1 cm)
Biochemically, the Ri-transformed plants are capable of synthesizing all the important secondary metabolites at a level comparable to or higher than the parent plants. Enhanced biomass of the transformed plants itself makes the use of Ri-transformed plants advantageous over nontransformed plants for the production of biologically active plant secondary metabolites. Many species are reported to produce medicinally important metabolites at higher concentration compared to their nontransformed counterparts, in addition to higher production of target metabolite due to increased biomass. Such plants are often reported retaining these desired characters, even after transfer to the field. However, the genetic stability of transformed plants has been reported in a very few species as discussed in the following segments of this review.
3.1 Morphological Stability of Ri-transformed Plants
Studies on stability of phenotype of transformed plants regenerated from transformed roots are very few in number, although, we have seen quite a number of reports are available for the hairy root cultures as discussed above. The altered morphology of the Ri-transformed plants are reported to be stable under long-term in vitro culture in very few species like Kiwi [91], T. indica [92, 93], Bacopa monnieri [94], and C. roseus [95].
Morphological stability of rolABC transformed plants and rolB transformed plants of Kiwi for more than 6 years has been reported by Rugini et al. [91]. The rolB transformed plants of Kiwi were morphologically similar to the control plants, whereas the rolABC plants showed a typical hairy root phenotype . These agronomic traits were maintained for over 6 years and in 50% of the plants of T1 generation [91].
Ri-transformed plants of T. indica, established by spontaneous regeneration from A4-transformed roots, have been reported to show typical Ri-transformed phenotype [96]. The plants showed a stable transformed phenotype in long-term culture (6 years) [92]. However, a detailed study on the fate of integrated T-DNA rol genes during regeneration via somatic embryogenesis in T. indica showed 19 out of 23 Ri-transformed plants with typical Ri-transformed phenotype and rest four plants with morphology similar to the nontransformed plants. These 23 plants with typical or variant morphology retained their characteristic phenotype for more than 3 years of in vitro maintenance [93].
In B. monnieri (Linn.), most of the clones of Ri-transformed plants were morphologically stable and did not show any alteration in the morphological characters including flowering [94].
Transgenic C. roseus plants are characterized with broad dark green leaves, shorter internodal length, rooting from its lower nodes, and flower with filamentous corolla compared to nontransformed plants [95, 97]. During 5 years of maintenance of these transgenic plants , drastic changes were observed: like reduction in plant height up to 1/4th of the initial height, highly proliferating root system, and absence of flowering and narrower yellowish leaves [95].
In several other plant species, the alteration in phenotype in transformed plants have been retained after transfer to the greenhouse, such as in Datura arborea [98], Limonium hybrid [99], and Kalanchoe blossfeldiana [100]. After 6 months of growing in the greenhouse, height of some clones of D. arborea was similar to untransformed plants [98]. In Ri-transformed plants of T. indica [92], Pelargonium graveolens [101], and Plumbago rosea [102], leaf wrinkling was not observed after transfer to the field.
In field evaluation of transgenic plants in Brassica, 40% plants showed severe Ri-phenotype, 40% showed moderate phenotype, however 20% of the transgenics showed normal phenotype [103]. Similarly, Ri-transformed plants regenerated spontaneously from the hairy roots of R. serpentina showed varied morphological characters. Out of 30 successfully acclimatized transgenic plants, 90% plants showed phenotypic similarity with nontransformed controls, and only 10% plants showed stunted growth [76]. The phenotypic variation may be due to the presence of multiple copies of the transgenes as revealed by Southern hybridization in R. serpentina [76]. Transformed plants exhibiting normal phenotypes regenerated from hairy roots are reported in many other species like tomato [104], Stylosanthes humilis [105], tobacco [106], Brassica napus [107], L. corniculatus [81], Cauliflower [108, 109], and Broccoli [109]. The absence of the transformed phenotype may not be correlated with the absence of specific rol gene(s), but it can also occur due to independent segregation or co-segregation of T-DNAs. In addition, the expression of transgene can be influenced synergistically by the number of integration, site of integration (repetitive DNA or transcriptionally active region of the host genome), and orientation of multiple copy insertion of foreign gene [76, 110–112].
Segregation of phenotypic characteristics has been reported in S. tuberosum among Ri-transformed plants regenerated through callus formation, producing phenotypes similar to the controls [87, 113]. Deletion of a single copy of TL-DNA in two shoot lines showed disappearance of Ri-characters [87]. Similar phenotypes were observed in the transformed sister shoot lines of potato [113].
In H. muticus L., somaclonal variation among plants regenerated from a single hairy root clone via protoplast culture has been reported [114, 115]. Half of regenerated plants showed typical Ri-transformed phenotype, while the rest of the plant lines were morphologically similar to nontransformed controls [114, 115].
3.2 Biochemical Stability of Ri-transformed Plants
Reports on analysis of secondary metabolites in plants are comparatively very few compared to the extensive reports available for hairy roots [22, 94]. Stable secondary metabolite production in Ri-transformed plants maintained in vitro have been reported in some plant species like B. monnieri [94],C. roseus [95], P. graveolens [101], R. serpentina [76], and T. indica [92, 93].
In P. graveolens, the Ri-transformed plants are reported to synthesize essential oils after 5 months of transferring to the soil [101].
In T. indica, Ri-transformed plants showed augmented production of the major alkaloid tylophorine compared to nontransformed plants [96], retaining their ability to synthesize improved level of tylophorine after 6 years of maintenance in vitro and there after 1 year of field transfer [92, 93].
Hairy root lines of R. serpentina and plants regenerated from such roots contain high reserpine [76]. Reserpine content of Ri-transformed plant was considerably higher compared to the control nontransformed plants grown in fields and was stably maintained over more than 2 years of cultivation [76].
In C. roseus transgenic plants, the tryptophan and alkaloid profile are reported to be almost constant during the 5 years of maintenance [95]. These transgenic plants were found to show better tryptophan content (357.5 ± 25.6 μg g−1DW in Ri-transformed plants compared to 114.5 ± 7.5 μg g−1DW in control) and alkaloid profile compared to nontransformed plants [95].
Biochemical stability of Ri-transformed plants of B. monnieri long-term in vitro culture is reported by Paul et al. [94] in Ri-crypt co-transformed plants. The plants showed stable, high content of bacosides compared to nontransformed plants for 4 years of in vitro culture [94].
3.3 Cytological Stability of Ri-transformed Plants
There is a lacuna in i nformation on chromosomal analysis of plants regenerated from transformed roots as compared to hairy roots in long-term culture. Among the few reports available, T. indica transformed plants showed chromosomal stability after 6 years of maintenance in vitro with a normal chromosome number 2n = 22 as parent plant [92, 116]. On the other hand, Webb et al. [81] reported changes in cytology along with morphology and physiology in plants regenerated from hairy roots of L. corniculatus.
3.4 Integration and Expression of T-DNA Genes in Ri-transformed Plants
Among the handful of reports available on long-term genetic stability of Ri-transformed plants, genetic stability have been reported in kiwi [91], T. indica [92, 93], B. monnieri [94], and C. roseus [95]. Genetic stability in transgenic kiwi plants transformed with rolABC and rolB has been reported by Rugini et al. [91]. In T. indica, most of the plant lines are reported to show genetic stability over 3–6 years of in vitro culture and after 1 year of field transfer, however, some variants are also reported [92, 93, 96]. Transgenic plants of B. monnieri showed stable integration as well as stable expression of rolA, rolB, rolC, and rolD genes after 4 years of maintenance in vitro [94].
Stable retention of three important rol genes – rolA, rolB, and rolC genes – have been reported in Ri-transformed plants of C. roseus even in the fifth year, indicating the stable nature of the Ri-transgenes [95].
Instability in the integration or expression of transgene in transformed plant lines regenerated from hairy roots is reported in S. tuberosum L. cv. Bintje [87], H. muticus [114, 115], and T. indica [93].
In S. tuberosum L. cv. Bintje, it was suggested that spontaneous deletions of TL-DNA and TR-DNA can occur during long-term root culture and regeneration of Ri-transformed plants [87].
In H. muticus, 50% of the Ri-transformed plants regenerated from the root line via protoplast culture showed the presence of rolA, rolB, and rolC similar to the root line from which they regenerated. Whereas, the latter 50% plants regenerated from the same root line were found rolA−/rolB−/rolC−. The transgenic morphology of the regenerated plants of H. muticus could be directly correlated with the presence of rol genes [114, 115].
Four variant plant lines of T. indica showing morphology similar to nontransformed plants were observed to show the presence and expression of only rolA gene. Such plant lines were regenerated through somatic embryogenesis from callus lines spontaneously developed from root lines with stable integration and expression of all the four rol genes . The variant plant lines showed the integration and expression of only the rolA gene throughout the period of study [93]. Loss of the rol genes in H. muticus and T. indica could be due to the deletion of the rol genes during regeneration [87, 117, 118].
Studies on the clonal fidelity of Ri-transformed plants are very few. Marker studies performed with 15 ISSR primers and cluster analysis in transgenic C. roseus plants revealed the relationship between the Ri-plants, Ti-plants, and their respective controls. The Dhawal Ti-plants were more close to Nirmal control plants followed by Nirmal Ri-plants [95].
4 Conclusions
Stability of Ri-transformed roots and regenerated plants is an important requirement if such cultures are to be used commercially. From the different reports it is evident that a detailed study on the extent of stability of transformed phenotype and genotype is far from adequate. The majority of the Ri-transformed cultures reported so far in different species show morphological, biosynthetic and genetic stability (in terms of T-DNA integration, expression, and cytological studies) when maintained in vitro by regular subculture. In some species, this stability is not obtained in Ri-transformed root and Ri-transformed plants. Apart from the variation between different clones of hairy roots and Ri-plants of a single species, instability in morphology, growth kinetics, biosynthetic potential, and loss of integrated T-DNA gene or its expression are reported to take place during long-term in vitro culture. Thus, transgenic root lines and regenerated plants should be analyzed for stability of the desired characters in long-term in vitro culture before being considered for scale up studies.
References
Bayliss MW (1980) Chromosomal variation in plant tissues in culture. Int Rev Cytol Suppl 11A:113–144
Scowcroft WR (1985) Somaclonal variation: the myth of clonal uniformity. In: Hohn B, Dennis ES (eds) Genetic flux in plants. Springer Verlag, Berlin
Scowcroft WR, Brettell RIS, Ryan SA, Davies PA, Pallotta MA (1987) Somaclonal variation and genomic flux. In: Green CE, Somers DA, Hackett WP, Biesboer DD (eds) Plant tissue and cell culture. Alan R Liss, New York
Winfield M, Davey MR, Karp A (1993) A comparison of chromosome instability in cell suspensions of diploid, tetraploid and hexaploid wheats. Heredity 70:187–194
Wilson SA, Roberts SC (2012) Recent advances towards development and commercialization of plant cell culture processes for the synthesis of biomolecules. Plant Biotechnol J 10:249–268
Jha S, Sen S (1987) Karyotype variability in regenerated plants of Urginea indica Kunth. Cytologia 52:615–626
Jha S (1989) Cytological analysis of embryogenic callus and regenerated plants of Urginea indica Kunth. Indian squill. Caryologia 42:165–173
Rani V, Raina SN (2000) Genetic fidelity of organized meristem-derived micropropagated plants: a critical reappraisal. In Vitro Cell Dev Biol Plant 36:319–330
Hao YJ, Deng XX (2002) Occurrence of chromosomal variations and plant regeneration from long-term-cultured Citrus callus. In Vitro Cell Dev Biol Plant 38:472–476
Kumar PS, Mathur VL (2004) Chromosomal instability in callus culture of Pisum sativum. Plant Cell Tiss Org Cult 78:267–271
Orbovic V, Calovic M, Viloria Z, Nielsen B, Gmitter FG Jr, Castle WS, Grosser JW (2008) Analysis of genetic variability in various tissue culture-derived lemon plant populations using RAPD and flow cytometry. Euphytica 161:329–335
Jain SM, Ahloowalia BS, Brar DS (2013) Somaclonal variation and induced mutations in crop improvement. Springer Science & Business Media, Dordrecht
Kaeppler SM, Kaeppler HF, Rhee Y (2000) Epigenetic aspects of somaclonal variation in plants. Plant Mol Biol 43:179–188
Devi SP, Kumaria S, Rao SR, Tandon P (2015) Genetic fidelity assessment in micropropagated plants using cytogenetical analysis and heterochromatin distribution: a case study with Nepenthes khasiana Hook f. Protoplasma 252:1305–1312
Sengupta J, Jha S, Sen S (1988) Karyotype stability in long-term callus derived plants of Crepis tectorum L. Biol Plant 30:247–251
Franklin CI, Mott RL, Vuke TM (1989) Stable ploidy levels in long-term callus cultures of loblolly pine. Plant Cell Rep 8:101–104
Jha S, Sen J, Sen S (1989) Stable regenerants from long-term callus cultures of Ruscus hypophyllum L. Cytologia 54:687–691
Nayak S, Sen S (1995) Rapid and stable propagation of Ornithogalum umbellatum L. in long term culture. Plant Cell Rep 15:150–153
Jha TB, Jha S (1989) In vitro regeneration and cytological study of Allium hookeri Thw. Indian J Exp Biol 27:363–365
Landey RB, Cenci A, Guyot R, Bertrand B, Georget F, Dechamp E, Herrera JC, Aribi J, Lashermes P, Etienne H (2015) Assessment of genetic and epigenetic changes during cell culture ageing and relations with somaclonal variation in Coffea arabica. Plant Cell Tiss Org Cult 122:517–531
Hamill JD, Lidgett AJ (1997) Hairy root cultures- opportunities and key protocols for studies in metabolic engineering. In: Doran PM (ed) Hairy roots. Gordon and Breach/Harwood Academic, London
Roychowdhury D, Majumder A, Jha S (2013) Agrobacterium rhizogenes-mediated transformation in medicinal plants: prospects and challenges. In: Chandra S et al (eds) Biotechnology for medicinal plants. Springer, Berlin/Heidelberg
DiCosmo F, Misawa M (1995) Plant cell and tissue culture: alternatives for metabolite production. Biotechnol Adv 13:425–435
Rao SR, Ravishankar GA (2002) Plant cell cultures: chemical factories of secondary metabolites. Biotechnol Adv 20:101–153
Deus-Neumann B, Zenk MH (1984) Instability of indole alkaloid production in Catharanthus roseus cell suspension cultures. Planta Med 50:427–431
Holden MA, Holden PR, Yeoman MM (1988) Elicitation of secondary product formation in Capsicum frutescens cultures. In: Robins RJ, Rhodes MJC (eds) Manipulating secondary metabolism in culture. Cambridge University Press, Cambridge
Evans DA, Gamborg OL (1982) Chromosome stability of cell suspension cultures of Nicotiana spp. Plant Cell Rep 1:104–107
Karp A, Wu QS, Maddock SE, Jones MGK (1990) Chromosome instability in bread wheat (Triticum aestivum) cell suspensions and their dividing protoplasts. In: Bajaj YPS (ed) Wheat, vol 13. Springer, Berlin/Heidelberg
Anrini M, Jha S (2009) Characterization of podophyllotoxin yielding cell lines of Podophyllum hexandrum. Caryologia 62:220–235
Lapitan NLV, Sears RG, Gill BS (1984) Translocations and other karyotypic structural changes in wheat x rye hybrids regenerated from tissue culture. Theor Appl Genet 68:547–554
Phillips RL, Kaeppler SM, Olhoft P (1994) Genetic instability of plant tissue cultures: breakdown of normal controls. Proc Natl Acad Sci 91:5222–5226
Whitmer S, Canel C, Heijden RVD, Verpoorte R (2003) Long-term instability of alkaloid production by stably transformed cell lines of Catharanthus roseus. Plant Cell Tiss Org Cult 74:73–80
Dubrovina AS, Kiselev KV (2012) Effect of long-term cultivation on resveratrol accumulation in a high-producing cell culture of Vitis amurensis. Acta Physiol Plant 34:1101–1106
Kiselev KV, Dubrovina AS, Bulgakov VP (2009) Phenylalanine ammonia-lyase and stilbene synthase gene expression in rolB transgenic cell cultures of Vitis amurensis. Appl Microbiol Biotechnol 82:647–655
Zeng F, Qian J, Luo W, Zhan Y, Xin Y, Yang C (2010) Stabilityof transgenes in long-termmicropropagation of plants of transgenicbirch (Betula platyphylla). Biotechnol Lett 32:151–156
Guivarc’h A, Boccara M, Prouteau M, Chriqui D (1999) Instability of phenotype and gene expression in long-term culture of carrot hairy root clones. Plant Cell Rep 19:43–50
Kooter JM, Matzke MA, Meyer P (1999) Listening to the silent genes: transgene silencing, gene regulation and pathogencontrol. Trends Plant Sci 4:340–347
Mette MF, Van der Winden J, Matzke MA, Matzke AJM (1999) Production of aberrant promoter transcripts contributes to methylation and silencing of unlinked homologous promoters in trans. EMBO J 18:241–248
Slightom JL, Durand-Tardif M, Jouanin L, Tepfer D (1986) Nucleotide sequence analysis of TL-DNA of Agrobacterium rhizogenes agropine type plasmid. J Biol Chem 261:108–121
Christey MC (2001) Use of Ri-mediated transformation for production of transgenic plants. In Vitro Cell Dev Biol Plant 37:687–700
Bulgakov VP, Shkryl YN, Veremeichik GN, Gorpenchenko TY, Inyushkina YV (2011) Application of Agrobacterium rol genes in plant biotechnology: a natural phenomenon of secondary metabolism regulation. In: Alvarez M (ed) Genetic transformation. INTECH Open Access Publisher
Tepfer D (1984) Genetic transformation of several species of higher plants by Agrobacterium rhizogenes: phenotypic consequences and sexual transmission of the transformed genotype and phenotype. Cell 37:959–967
Hamill JD, Evans D, Robins RJ, Rhodes MJC, Prescott A, Martin C (1988) Foreign gene insertion into hairy roots with binary vectors and Agrobacterium rhizogenes: potential for genetic manipulation of plant secondary metabolism in culture. In: Robins RJ, Rhodes MJC (eds) Manipulating secondary metabolism in culture. Cambridge University Press, Cambridge
Sevón N, Oksman-Caldentey KM (2002) Agrobacterium rhizogenes-mediated transformation: root cultures as a source of alkaloids. Planta Med 68:859–868
Aoki T, Matsumoto H, Asako Y, Matsunaga Y, Shimomura K (1997) Variation of alkaloid productivity among several clones of hairy roots and regenerated plants of Atropa belladonna transformed with Agrobacterium rhizogenes 15834. Plant Cell Rep 16:282–286
Batra J, Dutta A, Singh D, Kumar S, Sen J (2004) Growth and terpenoid indole alkaloid production in Catharanthus roseus hairy root clones in relation to left- and right-termini-linked Ri T-DNA gene integration. Plant Cell Rep 23:148–154
Chaudhuri KN, Ghosh B, Tepfer D, Jha S (2005) Genetic transformation of Tylophora indica with Agrobacterium rhizogenes A4: growth and tylophorine productivity in different transformed root clones. Plant Cell Rep 24:25–35
Bandyopadhyay M, Jha S, Tepfer D (2007) Changes in morphological phenotypes and withanolide composition of Ri-transformed roots of Withania somnifera. Plant Cell Rep 26:599–609
Alpizar E, Dechamp E, Lapeyre-Montes F, Guilhaumon C, Bertrand B, Jourdan C, Lashermes P, Etienne H (2008) Agrobacterium rhizogenes-transformed roots of Coffee (Coffea arabica): conditions for long-term proliferation and morphological and molecular characterization. Ann Bot 101:929–940
Taneja J, Jaggi M, Wankhede DP, Sinha AK (2010) Effect of loss of T-DNA genes on MIA biosynthetic pathway gene regulation and alkaloid accumulation in Catharanthus roseus hairy roots. Plant Cell Rep 29:1119–1129
Halder M, Jha S (2016) Enhanced trans-resveratrol production in genetically transformed root cultures of Peanut (Arachis hypogaea L.). Plant Cell Tiss Org Cult 124:555–572
Roychowdhury D, Basu A, Jha S (2015) Morphological and molecular variation in Ri-transformed root lines are stable in long term cultures of Tylophora indica. Plant Growth Regul 75:443–453
Mano Y, Ohkawa H, Yamada Y (1989) Production of tropane alkaloids by hairy root cultures of Duboisia leichhardtii transformed by Agrobacterium rhizogenes. Plant Sci 59:191–201
Jouanin L, Guerche D, Pamboukdjian N, Tourneur C, Casse-Delbart F, Tourneur J (1987) Structure of T-DNA in plants regenerated from roots transformed by Agrobacterium rhizogenes strain A4. Mol Gen Genet 206:387–392
Piñol MT, Palazòn J, Serrano M (1996) Effects of Ri T-DNA from Agrobacterium rhizogenes on growth and hyoscyamine production in Datura stramonium root cultures. Bot Acta 109:133–138
Bulgakov VP, Khodakovskaya MV, Labetskaya NV, Chernoded GK, Zhuravlev YN (1998) The impact of plant rolC oncogene on ginsenoside production by ginseng hairy root cultures. Phytochemistry 49:1929–1934
Spena A, Schmülling T, Koncz C, Schell J (1987) Independent and synergistic activity of Rol A, B and C loci in stimulating abnormal growth in plants. EMBO J 6:3891–3899
Schmülling T, Schell J, Spena A (1988) Single genes from Agrobacterium rhizogenes influence plant development. EMBO J 7:2621–2629
Spanó L, Mariotti D, Cardarelli M, Branca C, Constantino P (1988) Morphogenesis and auxin sensitivity of transgenic tobacco with different complements of Ri T-DNA. Plant Physiol 87:479–483
Lipp Joao KH, Brown TA (1994) Long-term stability of root cultures of tomato transformed with Agrobacterium rhizogenes R1601. J Exp Bot 45:641–647
Geerlings A, Hallard D, Martinez Caballero A, Lopes Cardoso I, van der Heijden R, Verpoorte R (1999) Alkaloid production by a Cinchona officinalis ‘ledgeriana’ hairy root culture containing constitutive expression constructs of tryptophan decarboxylase and strictosidine synthase cDNA from Catharanthus roseus. Plant Cell Rep 19:191–196
Zdravković-Korać S, Ćalić D, Druart PH, Radojević L (2003) The horse chestnut lines harboring the rol genes. Biol Plant 47:487–491
Santos PM, Figueiredo AC, Oliveira MM, Barroso JG, Pedro LG, Deans SG, Younus AKM, Scheffer JJC (1999) Morphological stability of Pimpinella anisum hairy root cultures and time-course study of their essential oils. Biotechnol Lett 21:859–864
Basu A, Joshi RK, Jha S (2015) Genetic transformation of Plumbago zeylanica with Agrobacterium rhizogenes strain LBA 9402 and characterization of transformed root lines. Plant Tiss Cult Biotechnol 25:21–35
Pandey P, Kaur R, Singh S, Chattopadhyay SK, Srivastava SK, Banerjee S (2014) Long-term stability in biomass and production of terpene indole alkaloids by hairy root culture of Rauvolfia serpentina and cost approximation to endorse commercial realism. Biotechnol Lett 36:1523–1528
Ray S, Samanta T, Majumder A, Bandyopadhyay M, Jha S (2014) Cytogenetic characterization of Agrobacterium rhizogenes transformed root lines of Rauvolfia serpentina. Nucleus 57:105–112
Marconi PL, Setten LM, Cálcena EN, Alvarez MA et al (2008) Changes in growth and tropane alkaloid production in long-term culture of hairy roots of Brugmansia candida. J Integr Biosci 3:38–44
Yukimune Y, Hara Y, Yamada Y (1994) Tropane alkaloid production in root cultures of Duboisia myoporoides obtained by repeated selection. Biosci Biotechnol Biochem 58:1443–1446
Sevón N, Hiltunen R, Oksman-Caldentey KM (1998) Somaclonal variation in transformed roots and protoplast-derived hairy root clones of Hyoscyamus muticus. Planta Med 64:37–41
Benson EE, Hamill JD (1991) Cryopreservation and post freeze molecular and biosynthetic stability in transformed roots of Beta vulgaris and Nicotiana rustica. Plant Cell Tiss Org Cult 24:163–172
Maldonado-Mendoza IE, Ayora-Talavera T, Loyola-Vargas VM (1993) Establishment of hairy root cultures of Datura stramonium characterization and stability of tropane alkaloid production during long periods of subculturing. Plant Cell Tiss Org Cult 33:321–329
Baíza AM, Quiroz-Moreno A, Ruíz JA, Loyola-Vargas VM (1999) Genetic stability of hairy root cultures of Datura stramonium. Plant Cell Tiss Org Cult 59:9–17
Dechaux C, Boite-Conti M (2005) A strategy for overaccumulation of scopolamine in Datura innoxia hairy root cultures. Acta Biol Cracov Bot 47:101–107
Jouhikainen K, Lindgren L, Jokelainen T, Hiltunen R, Teeri TH, Oksman-Caldentey KM (1999) Enhancement of scopolamine production in Hyoscyamus muticus L. hairy root culture by genetic engineering. Planta 208:545–551
Maldonado-Mendoza IE, Ayora-Talavera TRD, Loyola-Vargas VM (1992) Tropane alkaloid production in Datura stramonium root cultures. In Vitro Cell Dev Biol Plant 28:67–72
Mehrotra S, Goel MK, Rahman LU, Kukreja AK (2013) Molecular and chemical characterization of plants regenerated from Ri-mediated hairy root cultures of Rauwolfia serpentina. Plant Cell Tiss Org Cult 114:31–38
Ambros PF, Matzke AJM, Matzke MA (1986) Localization of Agrobacterium rhizogenes T-DNA in plant chromosomes by in situ hybridization. EMBO J 5:2073–2077
Aird ELH, Hamill JD, Rhodes MJC (1988) Cytogenetic analysis of hairy root cultures from a number of plant species transformed by Agrobacterium rhizogenes. Plant Cell Tiss Org Cult 15:47–57
Ermayanti TM, McComb JA, O’Brien PA (1992) Cytological analysis of seedling roots, transformed root cultures and roots regenerated from callus of Swainsona galegifolia (Andr.) R. Br. J Exp Bot 44:375–380
Mukherjee S, Das S, Jha S (1994) Chromosomal stability in transformed hairy root cultures of Artemissia annua L. Cell Chromos Res 17:71–76
Webb KJ, Jones S, Robbins MP, Minchin FR (1990) Characterization of transgenic root cultures of Trifolium repens, Trifolium pratense and Lotus corniculatus and transgenic plants of Lotus corniculatus. Plant Sci 70:243–254
Ermayanti TM, Octavia Y, Hafizh EA (2004) Cytological analysis of root cultures of Artemissia cina. Ann Bogoriensesns 9:50–58
Ramsay G, Kumar A (1990) Transformation of Vicia faba cotyledon and stem tissues by Agrobacterium rhizogenes: infectivity and cytological studies. J Exp Bot 41:841–847
Banerjee-Chattopadhyay S, Schwemmin AM, Schwemmin DJ (1985) A study of karyotypes and their alterations in cultured and Agrobacterium transformed roots of Lycopersicon peruvianum Mill. Theor Appl Genet 71:258–262
Vries-Uijtewaal ED, Gilissen LJW, Flipse E, Sree Ramulu K, De Groot B (1988) Characterization of root clones obtained after transformation of monohaploid and diploid potato genotypes with hairy root inducing strains of Agrobacterium. Plant Sci 58:193–202
Xu Z-Q, Jia J-F (1996) The reduction of chromosome number and the loss of generation ability during subculture of hairy root cultures of Onobrychis viciaefolia transformed by Agrobacterium rhizogenes. Plant Sci 120:107–112
Hänisch ten Cate CH, Loonen AE, Ottaviani MP, Ennik L, van Eldik G, Stiekema WJ (1990) Frequent spontaneous deletions of Ri T-DNA in Agrobacterium rhizogenes transformed potato roots and regenerated plants. Plant Mol Biol 14:735–741
Nilsson O, Olsson O (1997) Getting to the root: the role of the Agrobacterium rhizogenes rol genes in the formation of hairy roots. Physiol Plant 100:463–473
Kamada H, Saitou T, Harada H (1992) No requirement of vernalization for flower formation in Ri-transformed Cichorium plants. Plant Tiss Cult Lett 9:206–208
Sun LY, Touraud G, Charbonnier C, Tepfer D (1991) Modification of phenotype in Belgian endive (Cichorium intybus) through genetic transformation by Agrobacterium rhizogenes: conversion from biennial to annual flowering. Transgenic Res 1:14–22
Rugini E, Caricato G, Muganu M, Taratufolo C, Camilli M, Cammilli C (1997) Genetic stability and agronomic evaluation of six year-old transgenic kiwi plants for rolABC and rolB gene. Acta Hortic 447:609–610
Roychowdhury D, Ghosh B, Chaubey B, Jha S (2013) Genetic and morphological stability of six-year-old transgenic Tylophora indica plants. Nucleus 56:81–89
Roychowdhury D, Chaubey B, Jha S (2015) The fate of integrated Ri T-DNA rol genes during regeneration via somatic embryogenesis in Tylophora indica. J Bot 2015:16
Paul P, Sarkar S, Jha S (2015) Effects associated with insertion of cryptogein gene utilizing Ri and Ti plasmids on morphology and secondary metabolites are stable in Bacopa monnieri transformed plants grown in vitro and ex vitro. Plant Biotechnol Rep 9:231–245
Verma P, Sharma A, Khan SA, Mathur AK, Shanker K (2015) Morphogenetic and chemical stability of long-term maintained Agrobacterium-mediated transgenic Catharanthus roseus plants. Nat Prod Res 29:315–320
Chaudhuri KN, Ghosh B, Tepfer D, Jha S (2006) Spontaneous plant regeneration in transformed roots and calli from Tylophora indica: changes in morphological phenotype and tylophorine accumulation associated with transformation by Agrobacterium rhizogenes. Plant Cell Rep 25:1059–1066
Choi PS, Kim YD, Choi KM, Chung HJ, Choi DW Jr (2004) Plant regeneration from hairy-root cultures transformed by infection with Agrobacterium rhizogenes in Catharanthus roseus. Plant Cell Rep 22:828–831
Giovannini A, Pecchioni N, Rabaglio M, Allavena A (1997) Characterization of ornamental Datura plants transformed by Agrobacterium rhizogenes. In Vitro Cell Dev Biol Plant 33:101–106
Mercuri A, Bruna S, Benedetti LD, Burchi G, Schiva T (2001) Modification of plant architecture in Limonium spp. induced by rol genes. Plant Cell Tiss Org Cult 65:247–253
Christensen B, Sriskandarajah S, Müller R (2009) Biomass distribution in Kalanchoe blossfeldiana transformed with rol-genes of Agrobacterium rhizogenes. Hortic Sci 44:1233–1237
Saxena G, Banerjee S, ur Rahman L, Verma PC, Mallavarapu GR, Kumar S (2007) Rose-scented geranium (Pelargonium sp.) generated by Agrobacterium rhizogenes mediated Ri-insertion for improved essential oil quality. Plant Cell Tiss Org Cult 90:215–223
Satheeshkumar K, Jose B, Soniya EV, Seeni S (2009) Isolation of morphovariants through plant regeneration in Agrobacterium rhizogenes induced hairy root cultures of Plumbago rosea L. Indian J Biotechnol 8:435–441
Christey MC, Braun RH, Reader JK (1999) Field performance of transgenic vegetable Brassicas (Brassica oleracea and B. rapa) transformed with Agrobacterium rhizogenes. SABRAO J Breed Genet 31:93–108
Shahin EA, Sukhapinda K, Simpson RB, Spivey R (1986) Transformation of cultivated tomato by a binary vector in Agrobacterium rhizogenes: transgenic plants with normal phenotypes harbour binary vector T-DNA, but no Ri-plasmid T-DNA. Theor Appl Genet 72:770–777
Manners JM, Way H (1989) Efficient transformation with regeneration of the tropical pasture legume Stylosanthes humilis using Agrobacterium rhizogenes and a Ti plasmid binary vector system. Plant Cell Rep 8:341–345
Hatamoto H, Boulter ME, Shirsat AH, Croy EJ, Ellis JR (1990) Recovery of morphologically normal transgenic tobacco from hairy roots co-transformed with Agrobacterium rhizogenes and a binary vector plasmid. Plant Cell Rep 9:88–92
Boulter ME, Croy E, Simpson P, Shields R, Croy RRD, Shirsat AH (1990) Transformation of Brassica napus L. (oilseed rape) using Agrobacterium tumefaciens and Agrobacterium rhizogenes- a comparison. Plant Sci 70:91–99
Braun RH, Reader JK, Christey MC (2000) Evaluation of cauliflower transgenic for resistance to Xanthomonas campestris pv. campestris. Acta Hortic 539:137–143
Puddephat IJ, Robinson HT, Fenning TM, Barbara DJ, Morton A, Pink DAC (2001) Recovery of phenotypically normal transgenic plants of Brassica oleracea upon Agrobacterium rhizogenes mediated cotransformation and selection of transformed hairy roots by GUS assay. Mol Breed 7:229–242
Stam M, Joseph NMM, Kooter JM (1997) The silence of genes in transgenic plants. Ann Bot 79:3–12
Li XG, Chen SB, Lu ZX, Chang TJ, Zeng QC, Zhu Z (2002) Impact of copy number on transgene expression in tobacco. Acta Bot Sin 44:120–123
Tang W, Ronald JN, Douglas AW (2007) Genetic transformation and gene silencing mediated by multiple copies of a transgene in eastern white pine. J Exp Bot 58:545–554
Hänisch ten Cate CH, Ennik E, Roest S, Sree Ramulu K, Dijkhuis P, De Groot B (1988) Regeneration and characterization of plants from potato root lines transformed by Agrobacterium rhizogenes. Theor Appl Genet 75:452–459
Sevón N, Oksman-Caldentey KM, Hiltunen R (1995) Efficient plant regeneration from hairy root-derived protoplasts of Hyoscyamus muticus. Plant Cell Rep 14:738–742
Sevón N, Dräger B, Hiltunen R, Oksman-Caldentey KM (1997) Characterization of transgenic plants derived from hairy roots of Hyoscyamus muticus. Plant Cell Rep 16:605–611
Samaddar T, Nath S, Halder M, Sil B, Roychowdhury D, Sen S et al (2012) Karyotype analysis of three important traditional Indian medicinal plants, Bacopa monnieri, Tylophora indica and Withania somnifera. Nucleus 55:17–20
Trulson AJ, Simpson RB, Shahin EA (1986) Transformation of cucumber (Cucumis sativus L.) plants with Agrobacterium rhizogenes. Theor Appl Genet 73:11–15
Atkinson RG, Gardner R (1991) Agrobacterium-mediated transformation of pepino and regeneration of transgenic plants. Plant Cell Rep 10:208–212
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing Switzerland
About this entry
Cite this entry
Roychowdhury, D., Halder, M., Jha, S. (2017). Agrobacterium rhizogenes-Mediated Transformation in Medicinal Plants: Genetic Stability in Long-Term Culture. In: Jha, S. (eds) Transgenesis and Secondary Metabolism. Reference Series in Phytochemistry. Springer, Cham. https://doi.org/10.1007/978-3-319-28669-3_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-28669-3_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-28668-6
Online ISBN: 978-3-319-28669-3
eBook Packages: Chemistry and Materials ScienceReference Module Physical and Materials ScienceReference Module Chemistry, Materials and Physics