Abstract
The stability of integration and expression level of transgenes in long-term micropropagation clones of transgenic birch (Betula platyphylla Suk.) was examined. Multiplexed PCR and reverse primer PCR demonstrated stable integration of transgenes into regenerated plants. Expression levels of the bgt and gus genes among shoot plantlets, subcultured 4, 7, 9 and 15 times, were significantly different. The transcriptional expression level of extraneous genes in regenerated plants decreased with increasing subculture number. Transcriptional gene silencing (TGS) occured in regenerated transgenic lines. The silencing rate of GUS in the 5th subculture plants was 22–65%. TGS in regenerated plants could be reactivated with 5-azacytidine (Azac) at 50–200 μM. GUS and BGT protein expression was reactivated in the micropropagated transgenic birch plants when treated with Azac. A decrease in expression level with increasing number of subcultures is thus associated with DNA methylation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Once a gene can be successfully transferred, its usefulness is then determined by the ability to regenerate many transgenic plants from cultured tissues or cells of the desired tree species. Fixed integration of foreign genes into the genome of forest trees, and their subsequent stable expression, are essential for the further use of transgenic trees in forest tree breeding programs. However, significant variation occurs in the degree of transgene expression in regenerated plants derived from callus (Schween and Schwenkel 2003; Al-Zahim et al. 1999; Nakano et al. 2003). These tissue culture-induced variations, collectively called somaclonal variations, may be stable or unstable, reversible or irreversible, meiotically reset or transgenerationally transmitted (Krizova et al. 2009; Phillips et al. 1994). This variation hinders the application of a tissue culture techniques for propagation of transgenic plants.
Studies on this type of variation have, however, shown conflicting results. Transgenes conferring kanamycin resistance in tobacco are stable through the de-differentiation/differentiation cycle and the expression of the transgene remains high (Bellini et al. 1992). Foreign genes are stable in transgenic poplar plants through two full cycles of de-differentiation and differentiation (Wang et al. 1996). In contrast, although the cry1Ac and cry1C genes were both maintained (Cao and Earle 2003), ELISA showed that most propagated clones had significantly lower levels of Cry1C protein than the original plant. These variations in gene expression were believed to be mediated by epigenetic mechanisms involving methylation of DNA and chromatin modifications. Shifts from parental methylation states have been observed in many cell cultures and in the subsequently regenerated plants, at both global and local levels (Jaligot et al. 2000; Kubis et al. 2003; Schellenbaum et al. 2008; Smulders et al. 1995).
This variation highlights the importance of evaluating transgene expression in transgenic clones propagated in vitro over the long term. In the case of tree species, few studies have demonstrated that long-term tissue culture propagation does affect the structural integrity or expression of transgenes. Thus, stability of transgene integration and expression levels in transgenic birch lines is an important consideration for cultivar development programs. The aim of this study was therefore to establish an efficient system for the in vitro propagation of transgenic birch plants using buds and leaves as the initial explants and to analyse the stability of integration and expression level of transgenes during long-term propagation of regenerated clones.
Materials and methods
Characterisation of transformed plants
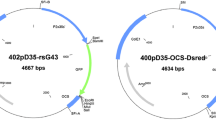
Betula platyphylla was transformed with Agrobacterium (LBA4404) containing a chimeric gene of a spider insecticidal peptide gene and the C-peptide sequence of Bt gene (Fig. 1). Four independent transformants were selected and cultured on WPM medium supplemented with 50 mg kanamycin l−1. Transgenic and non-transformed control plants were propagated in vitro and grown in a greenhouse under natural daylight conditions for 5 years.
Schematic representation of the T-DNA, restriction sites and probes. PstI restriction endonuclease was applied to analyse the copy number of bgt and gus. P35S, promoter from the cauliflower mosaic virus; nptII, bacterial neomycin phosphotransferase II gene; bgt, the chimeric gene of a spider insecticidal peptide gene and the C-peptide sequence of Bt gene; gus, β–glucuronidase gene; LB, T-DNA left border; RB, T-DNA right border
Tissue culture
Buds from transgenic plants were rinsed in running tap water and disinfected with 70% (v/v) ethanol for 30 s, 10% (w/v) Ca(ClO)2 for 10 min, then rinsed five times with sterile distilled water. The buds were cultured on WPM media supplemented with 1 mg BA l−1. After 1 month, leaves (1–2 cm) from the sterile shoots were explanted onto WPM media with 0.8 mg BA l−1 and 0.2 mg NAA l−1 for callus induction. WPM media containing 0.4 mg BA l−1 supplemented with 50 mg kanamycin l−1 were used for callus differentiation and subculture at 16/8 h day/night cycle at 25 ± 2°C and irradiance of 60 mM m−2 s−1 provided by cool-white fluorescent tubes.
DNA isolation and multiplex PCR analyses
Genomic DNA was isolated from fresh leaf material using the CTAB method. Multiplex PCR was performed in a reaction volume of 30 μl containing 50 ng DNA, 0.5 μM from each primer (see Supplementary Table 1 for details), 200 μM dNTPs and 1 U Taq DNA polymerase (TAKARA). PCR reactions was performed with the following amplification program: 94°C for 3 min; 35 cycles of 30 s at 94°C, 40 s at 58°C, and 1 min at 72°C, followed by 72°C for 10 min. The amplification products were subjected to electrophoresis using a 0.8% (w/v) agarose gel.
Reverse primer PCR for T-DNA structure
T-DNA structure for inserts with multiple copies was evaluated by reverse primer PCR (RP-PCR) (Kumar and Fladung 2000). Six different primers located along the T-DNA sequence were used for PCR amplification (see Supplementary Table 2 and Supplementary Fig. 1 for details). T–DNA structure was inferred by the presence and size of amplified bands. PCR conditions were Long Taq DNA polymerase (TaKaRa) in a volume of 20 μl containing 2 μl 10× LA PCR Buffer II (Mg2+ plus), 0.2 mM dNTP Mixture, 0.5 μM Primer 1, 0.5 μM Primer 2, 50 ng DNA, 1 U LA Taq DNA Polymerase. The PCR cycles were: 95°C for 2 min; 30 cycles of: 95°C 20 s, 68°C 15 min; 72°C for 7 min.
Nucleic acid isolation and northern blot analysis
Total RNA was isolated from fresh leaf material using the CTAB method. For northern blots, total RNA (10 μg) was denatured with 1 M glyoxal and 50% (v/v) DMSO in 10 mM sodium phosphate (pH 7.0) for 1 h at 50°C, separated in 1% (w/v) agarose-formaldehyde denaturing gels, blotted onto a Hybond-N+ nylon membrane (Roche) in 20×SSC transfer buffer, and fixed by UV cross-linking. The DIG labelling and detection system (Roche) was used following the manufacturer’s instructions. After the hybridisation of the bgt probe, membrane was stripped twice for 1 h at 80°C in formamide containing 1% (v/v) SDS to remove the DIG-labelled bgt probe, rinsed for 5 min in 2×SSC, and stored in 2×SSC buffer. Stripped membrane was then prehybridised and hybridised with the gus probe using the same procedures as for the bgt probe.
Protein isolation and ELISA
Protein was extracted from leaf tissues or callus. BGT protein was quantified by double-Ab sandwich ELISA using protein G-purified polyclonal rabbit antibodies and HRP-linked antibody, by the sodium periodate method.
5-Azacytidine (Azac) treatment
Explants from silenced transgenic birch plants were cultured Azac upto 0.2 mM. The explant callus and shoots analysed histochemically for RNA, GUS activity and BGT protein.
GUS assay
Expression of the gus gene was examined histochemically in leaves or callus from regenerated plants at different subculture times as previously described (Jefferson et al. 1987).
Results
Analyse of GUS activity of micropropagated plantlets
GUS analysis of 30 randomly selected transformed plants showed that foreign genes were stable through a cycle of dedifferentiation and differentiation and were maintained long term. As shown in Table 1, GUS activity in regenerated plants of the fourth subculture was similar to that of the original plants. No silenced plants were found, indicating that expression level of gus had stabilised within four subcultures. However, the rates of decreased gus activity were ranged from 15 to 50% of total micropropagated plantlets by the fifth subculture, corresponding to a silencing of 22.5–65%.
Structural integrity of the transgenes
Multiplex PCR results for regenerated plants subcultured 15 times, using primers specific to bgt, gus and npt II genes, are shown in Fig. 2. Gene specific bands appeared in multiplex PCR products from the original plant and all of the propagated clones, including those clones that showed decreased production and silencing of the gus protein. The non-transgenic control lacked either specific band. The transgenes were therefore stably transmitted and maintained over the long term. Reverse primer PCR analysis using genomic DNA from the clones in which the gus gene was silent showed amplified bands with primer pairs (1 + 6, 1 + 2, 1 + 5, 1 + 2 and 2 + 6), and the size of PCR products was consistent with predictions (Fig. 3). The presence of entire gus and bgt genes and ligations between T-DNAs were therefore indicated and no structural rearrangements or gene loss had occurred.
Molecular analysis for T-DNA repeats in transgenic line TP48. Lane M, 250 bp DNA ladder (250, 500, 750, 1000, 1500, 2250, 3000, 4500 bp); Lanes 1 + 6, 1 + 2, 1 + 5, 1 + 2 and 2 + 6 are PCR products amplified by each primer pair (see Table S1). Primer pairs produced PCR products consistent with predicted size. The results indicated two T-DNAs were arranged as direct repeats in one site
Transcription of bgt and gus genes in regenerated plants
The extent of expression of the bgt and gus genes varied among the different subcultured plants. As shown in Fig. 4, the transcription of the bgt and gus genes in fourth subculture shoots were similar in level to that of the original plant. However, the expression in propagated clones gradually weakened in the 7th, 9th and 15th subcultures. Overall, an increase in the number of subcultures resulted in weakening or even silencing of transgene expression.
Reactivation of expression of the silenced genes in micropropagated plants
Since PCR indicated no losses of the transgenes, the loss of expression after subculturing was most likely due to transgene methylation. To further test this, silenced propagated plants were cultured on WPM medium supplemented by different concentrations of Azac for Northern blot analysis.
An overall increase in gus gene expression in the callus tissues and shoots was seen in response to Azac for 20 days (Fig. 5). In all, 100% of the silenced plants from the 50 or 100 μM Azac treatment showed transgene reactivation. Expression of the gus gene in callus from 200 μM Azac treatment was higher than that from callus treated with 50 μM Azac. Spontaneous reactivation of gene expression was not found in any of the callus cultured on 25 μM Azac or on Azac-free medium.
After hybridisation of the gus probe, membranes were stripped using a DIG-labelled gus probe. The stripped membrane was then prehybridised and hybridised with the bgt probe. The bgt reactivation showed results consistent with those of the gus gene, indicating that gene silencing was at the transcriptional level in transgenic birch tissue cultures and that gene silencing could be reactivated by Azac treatment. DNA methylation most likely had increased with the number of subcultures undergone by transgenic birch tissues and this led to transgenic gene silencing (TGS).
Effect of Azac treatment on expression level of GUS and BGT protein
GUS protein expression was restored by Azac treatment of silenced callus cultures at 50 or 100 μM (Supplementary Fig. 2). However, only two out of the 30 callus cultures showed GUS protein expression in response to 25 μM Azac. No restoration of expression was seen in the absence of Azac. ELISA analysis of BGT proteins in Azac-treated callus showed protein expression similar to expected values. As shown in Fig. 6, high levels of BGT protein were seen in the callus from the 50 and 100 μM Azac treatments, indicating that the silenced bgt gene could be reactivated by Azac treatment.
ELISA analysis of BGT proteins in callus from TP23, TP27, TP46, TP48 following treatment with different concentrations of Azac. No restoration of expression was seen in the absence of Azac. Only TP27 showed BGT expression in response to 25 μM Azac. The contents of BGT protein were ranged from 0.027 to 0.36, 0.029 to 0.34 and 0.019 to 0.26% of total soluble protein in the calli of these transgenic plants treated by 50, 100 and 200 μM Azac, respectively
Discussion
The influence of tissue culture on gene expression and epigenetic modification has been discussed in several reports. Epigenetic changes such as methylation and chromatin modifications induced in tissue culture can have a direct effect on gene expression (e.g., activation or silencing), or can indirectly induce point mutations, excisions or insertions of activated transposable elements, or even lead to chromosome breakage (Filipecki and Malepszy 2006). Tissue culture induced DNA methylation polymorphism is often observed (Smulders et al. 1995). These changes occur in both directions, but hypomethylation is much more frequently reported (Bardini et al. 2003).
Structural integrity and stability of transgene expression following long term in vitro propagation of transgenic tree species are important. In this study, transgenic birch was propagated by shoot proliferation and callus induction. The silencing of GUS activity was observed after long-term propagation of the plants. We postulate that this instability is due to the number of subcultures performed during propagation. To validate this hypothesis, the effects of increasing number of subcultures on the expression level during 15 continuous subculture cycles were investigated. As the number of subcultures increased, the expression of transgenes in regenerated plants gradually weakened, or was even silenced. Multiplex PCR showed that the transgenes themselves were not lost, which suggested some type of epigenetic effect was occurring during vegetative propagation. This idea was supported by the restoration of transgene activity in response to Azac, which indicated that DNA methylation had induced transgene silencing in our clones after long-term micropropagation.
In conclusion, this study documents a successful protocol for the efficient propagation of clones derived from transgenic birch plants. Structures of the transgenes remained stable in clones propagated in vitro for 15 subcultures. Although expression of the transgenes decreased significantly in most of the long-term clones, reactivation of the transgenes could be obtained by Azac treatment.
References
Al-Zahim MA, Ford-Lloyd BV, Newbury HJ (1999) Detection of somaclonal variation in garlic (Allium sativum L.) using RAPD and cytological analysis. Plant Cell Rep 18:473–477
Bardini M, Labra M, Winfield M et al (2003) Antibiotic-induced DNA methylation changes in calluses of Arabidopsis thaliana. Plant Cell Tissue Organ Cult 72:157–162
Bellini C, Giordani C, Lupotto E et al (1992) Stability of a foreign gene in transgenic Nicotiana tabacum L. plants during a cycle of dedifferentiation/differentiation. Plant Sci 82:193–200
Cao J, Earle ED (2003) Transgene expression in broccoli (Brassica oleracea var.) clones propagated in vitro via leaf explants. Plant Cell Rep 21:789–796
Filipecki M, Malepszy S (2006) Unintended consequences of plant transformation: a molecular insight. J Appl Genet 47:277–286
Jaligot E, Rival A, Beule T et al (2000) Somaclonal variation in oil palm (Elaesis guineensis Jacg.): the DNA methylation hypothesis. Plant Cell Rep 19:684–690
Jefferson RA, Kavanagh TA, Beven MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Krizova K, Fojtova M, Depicker A et al (2009) Cell culture-induced gradual and frequent epigenetic reprogramming of invertedly repeated tobacco transgene epialleles. Plant Physiol 149:1493–1504
Kubis SE, Castilho AM, Vershinin A et al (2003) Retroelements, transposons and methylation status in the genome of oil palm (Elaeis guineensis) and the relationship to somaclonal variation. Plant Mol Biol 52:69–79
Kumar S, Fladung M (2000) Transgene repeats in aspen: molecular characterization suggests simultaneous integration of independent T-DNAs into receptive hotspots in the host genome. Mol Gen Genet 264:20–28
Nakano M, Tanaka S, Oota M et al (2003) Regeneration of diploid and tetraploid plants from callus-derived protoplasts of Agapanthus praecox ssp. orientalis (Leighton) Leighton. Plant Cell Tissue Organ Cult 72:63–69
Phillips RL, Kaeppler SM, Olhoft P (1994) Genetic instability of plant tissue cultures: breakdown of normal controls. Proc Natl Acad Sci USA 91:5222–5226
Schellenbaum P, Mohler V, Wenzel G et al (2008) Variation in DNA methylation patterns of grapevine somaclones (Vitis vinifera L.). BMC Plant Biol 8:78–87
Schween G, Schwenkel HG (2003) Effect of genotype on callus induction, shoot regeneration, and phenotypic stability of regenerated plants in the greenhouse of Primula ssp. Plant Cell Tissue Organ Cult 72:53–61
Smulders MJM, Rus-Kortekaas W, Vosman B (1995) Tissue culture-induced DNA methylation polymorphism in repetitive DNA of tomato calli and regenerated plants. Theor Appl Genet 91:1257–1264
Wang G, Castiglione S, Chen Y et al (1996) Poplar (Populus nigra L.) plants transformed with a Bacillus thuringiensis toxin gene: insecticidal activity and genomic analysis. Transgenic Res 5:289–301
Acknowledgment
This work was supported by the National Natural Science Foundation of China (NO: 30872045) and Specialized Research Fund for the Doctoral Program of Higher Education (NO: 200802251038).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zeng, F., Qian, J., Luo, W. et al. Stability of transgenes in long-term micropropagation of plants of transgenic birch (Betula platyphylla). Biotechnol Lett 32, 151–156 (2010). https://doi.org/10.1007/s10529-009-0120-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-009-0120-4