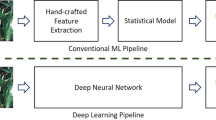
Abstract
Weed detection and classification are considered one of the most vital tools in identifying and recognizing plants in agricultural fields. Recently, machine learning techniques have been rapidly growing in the precision farming area related to plants, as well as weed detection and classification techniques. In digital agricultural analysis, these techniques have played and will continue to play a vital role in mitigating health, agricultural, and environmental impacts, improving sustainability, and reducing herbicides. Deep learning-based models are employed to solve the more sophisticated agricultural issues using individual CNN networks and hybrid models. Such models showed promising results. This paper highlights the major trends from the particular review of detection and classification approaches for weed plants. This review elaborates on the aspects of using traditional methods and deep learning-based methods to solve weed detection problems. It provides an overview of various methods for weed detection in recent years, analyzes the benefits and limitations of existing machine learning techniques, including deep learning techniques, and introduces several related plant leaves, weed datasets, and weeding machinery. Evaluation of the existing techniques has been compared, taking into account the real-world dataset used, images’ capacity, and shortcomings. Furthermore, this study helps to introduce the promising results and identify critically the remaining challenges in achieving robust weed detection, which could support noteworthy agricultural problems and assist researchers in the future. The significance of this study is to provide the potential techniques for solving illumination, overlapping, and occlusion issues of leafy plants, as well as other plant issues.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Detection of weed plants is an important step in facing the difficulties of the agricultural environment. In the agricultural field, weed plants, as shown in Fig. 1, are unwanted plants for their adverse impact on a crop’s productivity as they compete with crops for significant environmental resources (Adhikari et al. 2019; Hamuda et al. 2016). Its an aspect of plant or vegetation that grows in the incorrect location and obstructs the growth of crops, cultivating forests, or plantations. These weeds can slow the growth of the crop and reduce the farm’s yield. Therefore, these weeds should be identified and detected. Accurate weed detection assists in increasing crop productivity and income by improving the management system to extract a particular group of weeds (Wang et al. 2019a). Previously, the farmer used the manual hand-removal technique to remove the weeds manually. It detected the weeds by testing every region of the farm. Then, it plucked them out manually using their hands. Manual weeding presents numerous challenges, including time and cost constraints, labor-intensive detection work, and negative effects on crop productivity and the environment (Osorio et al. 2020).
Later, some automatic approaches commenced using advanced technology to remove the weeds, such as herbicides, chemical treatments for large-scale agricultural regions (Hamuda et al. 2016), and automatic machines that do not use image processing and pattern recognition techniques to detect the weeds. Recently, image processing and computer vision methods have been used to assist in site-specific weed management procedures to apply herbicide on a per-weed basis within a field (Etienne 2019). However, developing a weed detection system faces a major challenge, which requires annotated training data to differentiate between weeds and crops under various conditions, as illustrated in Fig. 2. Several studies focused on detecting multi-species of weeds by separating the weeds from grass (Binch and Fox 2017). Rumex obtusifolius L. (dockleaf) and Urtica are two significant broad-leaved weed plants that have spread across much of the globe (Kounalakis et al. 2018; Šeatović et al. 2008; Van Evert et al. 2009). These are serious weeds and necessitate further work to be conducted. Urtica has smaller leaves than Rumex, and their edges are serrated (Binch and Fox 2017). They are low in nutrients and contain a lot of oxalic acids. Oxalates can cause animal health problems if taken in large amounts by animals, and they can also degrade product quality (Kounalakis et al. 2019).
The main problem encountered by scientists in vegetation analysis is how to employ a method that can detect weed plants despite the presence of the illumination, overlapped, and occluded leaves of plants. Segmentation approaches for real-world images were insufficient in terms of accuracy and efficiency. Thus, it needs an efficient method, but with higher efficiency, to detect the optimum features that enhance the classification accuracy. Occluded leaves must be separated precisely and adequately without any distortion in the image. The previous works were implemented for an automated weed spraying system or weed control system. It helps the farmer reduce the time-consuming, hand-labor cost, and avoid herbicide pollution of the environment. Therefore, it is required to develop detection methods that are based on artificial intelligence techniques to optimize the use of agrochemicals. A variety of detection and classification algorithms have been employed to build automated weed maps from UAV data in recent studies (Herrmann et al. 2013; Tian et al. 2020; Wang et al. 2019a; Weis et al. 2008).
When dealing with complicated data, however, recent state-of-the-art works have been proven (Alam et al. 2020; Gao et al. 2018; Liakos et al. 2018) that machine learning methods are more accurate and efficient than traditional parametric algorithms. Among these machine learning algorithms, support vector machines have been popularly used for weed and crop classification (Abouzahir et al. 2018; Bakhshipour and Jafari 2018; Brinkhoff et al. 2020; Zhang et al. 2019). On the other hand, Kazmi et al. (2015) employed the KNN algorithm to detect creeping thistle in sugar beet farms. This paper aims to investigate real-world data problems that face weed detection in precision farming. The following are the study's contributions:
-
Study the effects of using real-world data for detecting broad-leafed weed plants.
-
Investigate the machine learning techniques that were used to deal with real-world scenarios.
-
A discussion of current methods for addressing features, as well as the implications of transfer learning and additional contextual information on the accuracy of these classifiers, as well as the accompanying augmentation strategies.
-
A description of recent research on publicly available plant datasets, as well as a review of the data augmentation structures.
The following sections constitute this paper: The real-world dataset and its properties are described in Sect. 2. In Sect. 3, we go through the present state of weed detection and show how recent advances in detection and classification methods have been demonstrated in real-world weed images. The challenges of weed detection are highlighted in Sect. 3. Section 4 covers the discussion and future work, pointing to outstanding problems and future research possibilities connected to weed identification, as well as how to use the results to benefit from categorization in smart agriculture.
Real-world data
Real-world data refers to the data that is captured under various circumstances of illumination, overlapping, occlusion, diseases, and growth-stage regardless of the visibility of the whole leaf or plant and noises. Initially, real-world conditions are a worth-mentioning issue in vegetation that has a significant impact on weed detection. Several cases face weed detection in their natural environment, including illumination, occlusion, and overlapping, which are inherent issues in real-world data as shown in Fig. 3. Partial occlusion is one of the real-world conditions that change the size and shape of the leaf, which reduces the accuracy of classification (Wang et al. 2019a) by more than 5% (Hall 2018).
Table 1 depicts the specifications of the dataset that is utilized in the methods of various researchers. Occlusion means that some parts of a leaf are totally or partially covered by another leaf. It occurs due to various conditions, such as growth stages and the orientation of the camera during capturing (Hamuda et al. 2018; Lin 2010). Another reason for occlusion is the planting process, when the farmer plants extra seeds at the same site to increase the potential rate of germination. Additionally, weed can cause occlusion when it covers the crop’s leaves (Tian 1995). Usually, the potential for occlusion and overlapping occurrences increases in late growth stages (Espejo-Garcia et al. 2020; Fernández-Quintanilla et al. 2018).
An overview of weed detection systems
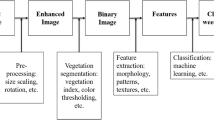
Both RGB and infrared (IR) imaging sensors have been employed in the field to acquire images for weed detection. The collected images are then fed as inputs to the processing techniques. Pre-processing, segmentation, feature extraction, and classification are common procedures in image processing (Weis and Sökefeld 2010). Figure 4 illustrates the general workflow and the input and output of each processing procedure.
Pre-processing phase
The goal of the pre-processing phase is to improve the quality of the obtained pictures and make the ROI segmentation phase easier. The key goal of this phase is to improve segmentation and classification accuracy by removing background noise and increasing object visibility. Noise can occur due to various causes, such as low-resolution images, various illumination, inaccurate camera sensors, and undesired objects like soil, plastic, and other residues (Hamuda et al. 2017; Kamilaris and Prenafeta-Boldú 2018). Koščević et al. (2020) stated three reasons for image illumination, including real farm images were taken under a natural lighting source; real-world images were taken under unreal lighting, and artificial images were taken under unreal lighting. Several factors have an impact on the image pre-processing, such as various illuminations, plant distribution in the field, the growth stage of the plants, and overlapping or occlusion leaf. Despite the multi-capturing image techniques for weed identification, contrast is still considered one of the most arduous challenges in image processing that necessitates an enhancement due to the various illumination conditions that cause noise.
Ali et al. (2017) and VijayaLakshmi and Mohan (2016) utilized Histogram Equalization (HE) to achieve three advantages, enhance the image contrast, remove background information, and provide the facility to process the redundant information and hidden details. Husham et al. (2016), Kumar and Prema (2016) and Li et al. (2016) employed the Adaptive Median Filter technique for noise removal and alleviating image distortion instead of the standard median filter. AMF overcame the lack in removing the tiny details, which cannot be filtered out via the normal median filter. Then, morphological operations, including dilation and erosion, are performed to eliminate the small-sized weeds and extract the soybean crop images. Their results revealed that the proposed technique can detect or identify weeds of similar size and color. In another work, Sathesh and Rasitha (2010) mentioned that the limitations of AMF are that it is considered expensive and rough to compute. Adams et al. (2020) utilized size reduction and data augmentation in deep learning. They used an image resizing (size reduction) technique to scale down the image resolution. It converts the input image to a standard small square size of 256 × 256, 224 × 224, 128 × 128, 96 × 96, and 60 × 60 pixels. Espejo-Garcia et al. (2020) claimed that 224 × 224 is the optimal resolution for VGG-16, DenseNet, and MobileNet networks due to the high performance that was registered with this size. The benefit of their technique is that it minimizes the computing time.
Adhikari et al. (2019), dos Santos Ferreira et al. (2019), Hari et al. (2019), Krizhevsky et al. (2012), Lin et al. (2020) and Olsen et al. (2019) used data augmentation technique to address the short-scale dataset images and reduce the overfitting. They used such a technique to increase the training account of synthetic images to be adjusted and generalized in real-world issues (Chen et al. 2017; Kamilaris and Prenafeta-Boldú 2018; Mortensen et al. 2016; Namin et al. 2018; Sladojevic et al. 2016; Sørensen et al. 2017). Generalization is the process of testing or predicting the trained network on new data that has never been seen before (Sharma et al. 2020). Nonetheless, Wang et al. (2019a) state that the problem with training large-scale data is not effective due to its being time-consuming in calculating and annotating. Table 2 summarizes the use of the most prevalent pre-processing algorithms for vegetation imagery in machine learning and deep learning. In deep learning, the more training images fed into a deep learning network, the more accurate the predicted outcome images will be. In precision farming, weed segmentation is a challenging case for real-time applications. Mis-segmentation affects classification accuracy and causes low performance in weed detection (Hamuda et al. 2016). Occlusion and natural illumination contribute to a lack of segmentation and reduce classification accuracy (Slaughter et al. 2008). On the other hand, high segmentation performance plays a crucial role in enhancing precision farming and effective herbicide treatment (Hamuda et al. 2016).
Weed segmentation phase
According to Chen et al. (2014); Sørensen et al. (2017), both crops and weeds have a similar green color, making the differentiation process quite tricky. They used image segmentation to separate an image into various distinguishable regions by highlighting the ROI. Generally, the vital benefits of segmentation images are stated below:
-
To simplify the learning process,
-
to facilitate the selection of optimum features,
-
to detect the weeds.
For image segmentation, there are two types of methodologies covered under computer vision: machine learning-based and deep learning-based.
Color index-based machine learning methods
According to Wang et al. (2019a), the color index is classified as one of the segmentation methods since it is used to separate the foreground from the background. Many works utilized color space transformation or color-based to separate the three standard channels of any color, Red, Green, and Blue (RGB) as a first stage to identify crops from the background. Woebbecke et al. (1995) invented Excessive Green color (ExG), ExG-ExR by Meyer and Neto (2008), Modified ExG (MExG) by Guo et al. (2013) and Burgos-Artizzu et al. (2009) implemented threshold algorithms in precision farming to recognize between the two classes, soil background class, and plant vegetation class depend on vegetation segmentation. Few studies utilized color space-based methods as an end-to-end for the distinguishing (Slaughter et al. 2008). The process of transforming a gray image into a binary image is known as thresholding. Otsu (1979) is the first algorithm applied to determine the threshold value. Additionally, the threshold technique is based on both color space and leaf shape. To identify the class, thresholds were mainly applied to a transformation of the original image; for example, several color indices-based systems were discussed and employed either zero thresholds or a threshold based on Otsu's method. Selecting the proper threshold value can play an essential role in segmentation. For example, if the threshold value is set to be extremely high, some interesting areas would be combined with other areas to form one big area.
Finlay (2012) employed other data transformation techniques such as CIE Lab, Hue, CIE XYZ, Cyan, Majenta, Yellow, and Black (CMYK). ExG was deployed by Liu et al. (2016) to separate five seedlings species and five densities in a wheat field. Their proposed method counted the number of overlapped patches using a chain code-based skeleton optimization method. The performance of their ExG was 89.94% accuracy as an average to compute the number of wheat seedlings. Li et al. (2016) used the HSI (Hue, Saturation, and Intensity) model for analyzing colors to simulate the way that the human eyes can recognize the colors’ contrast. They elucidated the principal fact that despite the diversity in plants’ species colors, crops and weeds almost share the green color to various degrees. This contrast was observed in the hue and saturation bands. The intensity band represented by I band in HSI depends on changing the illumination under real light conditions. On the other hand, illumination has a low effect on the hue and saturation bands. The proposed method attained 68, 83, 97, and 99% accuracy to discriminate celery cabbage, amaranthus tricolor, broccoli, and Chenopodium, respectively.
To decrease the overall noise of the dataset produced by the impact of illumination variations, the Normalized Difference Vegetation Index (NDVI) masks were used as one of the background reduction approaches (Haug et al. 2014; Lee et al. 2015; Prasanna Mohanty et al. 2016). Kumar and Prema (2016) employed CIE Lab color space in their proposed Wrapping Curvelet Transformation-Based Angular Texture Pattern (WCTATP) Extraction technique for weed identification in the carrot field. The CIE Lab adjusts or balances the optimum contrast through the spectral feature. Firstly, Red, Green, and Blue (RGB) color space was transformed into absolute sRGB color space to set the contrast and correct the balance. After that the sRGB color was converted to CIE XYZ values, after which they were transformed into linear sRGB values. Once the CIE XYZ value has been obtained, the D65 standard illuminant algorithm is performed to convert it to CIE Lab. Finally, the pixels were grouped into clusters depending on the dominant color of each pixel using the K-means method to separate the soil background, and plant vegetation.
Hamuda et al. (2017) proposed HSV for recognizing weeds, soil, and cauliflower crops using a threshold value for each band to identify the ROI. After which, morphological operations of erosion and dilation were applied. Their method achieved high performance with low detection error using controlled light conditions. The drawback with such a method was the sensitivity of HSV to the color, which is affected by leaf diseases and various illumination conditions that make sense. The author claims that the HSV color space is effective in addressing the illumination variations. The HSV color space, on the other hand, is more matched with human color perception and durable to illumination change, according to Hamuda et al. (2017).
Ali et al. (2017) used Hue Saturation Value (HSV) color space to obtain various illumination spaces. These transformations aim to produce a superior predictive model than using the original form of the input image. Concerning contrast and noise issues, several methods are conducted to eliminate undesired noises and enhance the image's deformations. Wang et al. (2019a) utilized background removal, foreground pixel extraction, or non-green pixel removal like soil, and other residues. Background removal is one of three factors, besides the transfer learning scheme and pooling category, affecting the performance of the fine-tuning technique for the pre-trained model (Espejo-Garcia et al. 2020).
Rangarajan and Purushothaman (2020) proposed three color spaces for (YCbCr): Hue Saturation Intensity (HSV), grayscale, and Green (Y), Blue (Cb), and Red (Cr). The proposed work used the images obtained from the previous color spaces with the Visual Geometry Group16 (VGG-16) model to classify the eggplant diseases. The experimental results showed a high accuracy in classification for the images handled by the YCbCr color space. The author focused on five various diseases that threaten the Eggplant. Notably, all the images were generated manually due to the nonexistence of the related images. The extracted features were classified using the multi-class support vector machine (MSVM).
Espejo-Garcia et al. (2020) and Kumar and Prema (2016) utilized the normalization or contrast scheme was used for contrast enhancement or color adjustment. Contrast means that the distribution of the pixel’s intensity value varies from the contiguous pixels, or it denotes the image resolution. Several factors have a great effect on image contrast, such as the distribution of black and white, boundary severity, and the duration of pattern recurrence. The normalization algorithm was used to eliminate the impact of the various illumination conditions like light, and shadow of the color channels. Moreover, the normalization algorithms can be used for background removal. This algorithm normalized the image in its green channel. Table 3 depicts the common color spaces in the vegetation field. As a result, Rangarajan and Purushothaman (2020) found that using gray and HSV color space with ResNet-101, GoogLeNet, AlexNet, and DensNet-201 classifiers resulted in low accuracy. However, ResNet-101 and GoogLeNet achieved higher results with RGB images than AlexNet and DensNet-201.
Learning-based segmentation methods
Although colored-spaced methods achieved promising results in the segmentation of green plants, their ability to deal with real illumination conditions such as sunny and cloudy days is limited. Colored-spaced methods achieved promising results in the segmentation of green plants, their ability to deal with real illumination conditions such as sunny and cloudy cases is limited. Two types of learning-based methods are stated under this section; the first one is the supervised-learning method, and the second one is the unsupervised-learning method.
Supervised learning-based segmentation methods
Tian and Slaughter (1998) presented the Environmentally Adaptive Segmentation Algorithm (EASA), a partially supervised learning algorithm, to segment cotyledon crops in outdoor plants under different lighting conditions. Their algorithm was based on the automatic generation of a look-up table (LUT) of the RGB images. A group of observations was considered self-learning for clustering to build the structure of similar pixels. Clustering is considered the key point of EASA. The advantage of this algorithm is the ability to process the daytime changing conditions from the environmental cases compared with other trained methods under sunny conditions. One of the drawbacks of such an approach is that it requires a large amount of data to train. Furthermore, it demonstrated low quality in detecting the cotyledon crop under overcast conditions. Bergasa et al. (2000) applied Gaussian Mixture Modeling (GMM) to work with real-world RGB images and multi-backgrounds. Meyer et al. (2004) developed Fuzzy Clustering (FC) to extract and recognize plants from soil and other residue using ExR and ExG color space for real-time application. The benefit of this method is the ability to differentiate the green plants from the background. The segmentation performance of such a method is decreased when the green pixels in the image are smaller than 10% due to insufficient color information.
Ruiz-Ruiz et al. (2009) utilized EASA to segment sunflower crops under real-world conditions. They adapted EASA to work with different color spaces instead of using RGB, they employed Hue saturation (HS) and Hue (H). The authors claimed that using RGB color space is an insufficient process due to the convergence in the properties of the green color on the gray scale between the background and foreground objects. For this reason, clustering is an inadequate process. There are two stages to their method. The first stage of the segmentation process is the training stage. The second stage is generating the LUT for generalizing the segmentation in the real-world. As a result, the EASA technique that used RGB color space (Tian and Slaughter 1998) is regarded as time-consuming compared to EASA that used Hue and saturation (HS) or Hue (H) individually (Ruiz-Ruiz et al. 2009).
Zheng et al. (2009) developed Mean-Shift and Back Propagation Neural Network (MS-BPNN) algorithm to enhance the segmentation performance. This method was implemented in the real-world to test various green plant species in different illumination conditions. To evaluate the segmentation quality of this method, two of the color-based methods CIVE, and ExG are employed for the benchmark. The mis-segmented areas are measured by manually labeling the foreground pixels with ones and the background pixels with zeros. Then, these values are compared to the segmented images to calculate the min, max, and average values of the mis-segmented rate. The overall results show that MS-BPNN outperformed both the CIVE and ExG in terms of segmentation quality and efficiency.
Zheng et al. (2010) proposed a different approach to enhancing the segmentation accuracy in soybean plants. They combined two methods, including MS with the Fisher Linear Discriminant (MS-FLD) approach. To evaluate the segmentation performance, they compared three color-based approaches, including CIVE, NDI, and, ExG with their proposed approach. The performance of the three color-indices showed acceptable results, but they were unstable for all tested images compared with MS-FLD. However, the computation time was also assessed with the tree vegetation color-based. The evaluation demonstrated that the running time for the three compared methods was less than the proposed approach by 0.0156 and 3.3906 s, respectively.
Guo et al. (2013) produced the Decision Tree Segmentation Model (DTSM) to segment wheat crops from the background for images captured under various illumination conditions. DTSM trained the extracted features using the Classification and Regression Trees (CART) classifier for creating the decision tree and noise removal. Unlike color-space methods, the threshold value is not considered in DTSM to be adjusted for each new image. However, the disadvantage of such a method is the use of training data. ExG, ExG-ExR, and MExG are three color indices that were utilized to evaluate the segmentation performance of the proposed model with these methods. The results showed that the DTSM outperformed the three color indices in segmenting the natural luminous and dim images.
Unsupervised learning-based segmentation methods
Yu et al. (2013) adopted a combination approach of the hue-intensity (HI) look-up table and the Affinity Propagation (AP) clustering algorithm called AP-HI to separate the maize crops from the background under natural illumination conditions. To verify the validity of this method, five different methods were used, including three color-indices: CIVE, ExG, and ExGR with Otsu, one with mean threshold, and the fifth with EASA (Tian and Slaughter 1998). The results demonstrated that AP-HI achieved high performance in the segmentation of the maize in various illimitation circumstances. It scored 96.68%, while EASA registered 93.2%. The advantage of AP-HI is the robustness to illumination changes. The disadvantage of such a method is that it shows low results in classifying some highlighted leaves’ patches. Geometrical distribution was utilized by K-Means to structure the similar pixels with common features into groups (Behmann et al. 2015). Kumar and Prema (2016) used k-means to separate the background from the foreground and filter the image. The problem with K-means is that it requires a known number of clusters, which is difficult when there are unspecific numbers of plant species to be grouped in a cluster. To solve this problem, Hall et al. (2017) proposed the Dirichlet Process Gaussian Mixture Model (DPGMM), which does not require prior knowledge of cluster accounts (Hall et al. 2017; Wang et al. 2019a).
Deep learning-based segmentation models
The researchers scrambled forward to develop their network depending on convolution, and deconvolution layers such as UNet, SegNet, reSeg multiscale FCN, and DeepLabV3 (Chen et al. 2017). Each of these segmentation models has its own powerful strength if it is applied to the correct dataset (Zhang et al. 2018). Long et al. (2015) used a fully convolutional network (FCN) based on deep learning for image segmentation. Their method employed convolutional layers to increase the ability to detect accurate features better than the features extracted by traditional machine learning methods. Volpi and Tuia (2016) employed a Fully Convolutional Network (FCN) for pixel-wise segmentation to classify a vast area of vegetation. Multi-architecture versions of FCN are developed for pixel-based high-fidelity UAV remote sensing imagery in an end-to-end style. The first three generations included FCN-8s, FCN-16s, and FCN-32s. Lottes et al. (2016) employed a fully connected network (FCN) for semantic segmentation. The experimental test exploits robust results with data that has never been applied before, so that it can generalize to work with real data.
Furthermore, it handled the data with various growth stages. The problem with their approach is that it addressed the crop or weed individually on an obvious background represented by the soil. In this case, the presence of overlapping issues is relatively low. Their approach evaluated the performance’s accuracy on recall to report 94% and 91% for crops and weeds, respectively. Mortensen et al. (2016) explored the adapted VGG-16 for pixel-wise classification or semantic segmentation of RGB multi-species imagery. Their proposed approach adapted the depth of the original VGG-16 to fit the number of classes. Furthermore, the fully connected layers of the convolutional connected layer were modified. In addition, 32-strides were added to the deconvolutional layer to make the output layer's size match the input layer's original size. They used the data augmentation technique in their approach to increase the dataset artificially.
Transfer learning was utilized in such an approach by fine-tuning the pre-trained weights to reduce the time-consuming process of building from scratch. The problem with such an approach is that it is time-consuming to address the online conditions and the low accuracy in detecting small objects with fine details. The reported results of this network were 79% for pixel accuracy of multi-class images. Di Cicco et al. (2017) concluded that SegNet-based crop-weed segmentation can learn from data generated artificially. Transfer learning (life-long learning) technique was defined as the method of reusing or recycling knowledge gained from solving a certain problem. With a small number of changes in the original weights for the previous pre-trained model, the new transferred knowledge is ready to tackle a new separate task. This technique was employed to overcome the limitations of deep learning for the identification of high-resolution images. Another benefit of the transfer learning method is that the information in each image is recognized using features transferred from a previously trained CNN model (Kounalakis et al. 2019). Furthermore, the transfer learning approach is used to reduce the time-consuming for images labeled by the experts, which are then transferred into the synthetic world (Di Cicco et al. 2017; Pan and Yang 2009). Furthermore, transfer learning was utilized to overcome the overfitting issue (Espejo-Garcia et al. 2020). The disadvantage of their technique is that when the pre-trained data is smaller than the new data for achieving one task at a specific time. In this regard, multitask learning was utilized to reduce the time by achieving multitasks simultaneously instead of performing one task sequentially as in transfer learning; as a result, it enhances the generalization performance (Caruana 1997).
A novelty of the CNN-based approach proposed by Badrinarayanan et al. (2017) is the SegNet convolutional network for semantic pixel-wise segmentation. Some advantages of SegNet are the low consuming time, good utilizing memory, low number of parameters compared with other relevant networks, and finally, the ability to segment the overlapped plants. The author produced two versions of the SegNet network, the old small network consisting of only eight layers, while the bigger version consists of 26 layers to handle the multi-classes. Whereas the problem with a new big version network is overfitting when addressing a small number of classes. Another CNN architecture for semantic segmentation was conducted by Milioto et al. (2018) to identify the sugar beets, weed, and background of the RGB images in real-time. They extracted extra information from 14 channels using color-indices without the need for NIR-infrared information to reduce the cost. Furthermore, their method addressed the heavily overlapping, various growth stages, and illumination. A new FCN model was proposed by Mohammadimanesh et al. (2019) to classify multi-classes in wetlands employing polarimetric (PolarSAR) imagery. The architecture of their CNN model consists of two stages; the encoder stage and the decoder stage. The first one is to extract the high-level features, while the second one is to up-sample into the spatial resolution of the original input size using the output data of the encoder stage as input data to feed it. Their proposed model used escape connections between the encoder and decoder layers. These connections are useful in transferring information between these two layers to enhance the extraction of spatial features. The results of the proposed model achieved 93% classification accuracy, which is higher than the FCN-8s and FCN-32s. The problem with their CNN model is ground truth images' limitations for remote sensing applications. Table 4 depicts the previous summary work.
Zhao et al. (2019) utilized UNet, or U-shaped network for segmentation, especially semantic segmentation tasks, which achieved high performance in medical images. It consists of two parts. The first one is the encoder, which shrinks or contracts the input image. The second one is a decoder, which expands the image to recover its original size. The encoder part contains max-pooling layers. The duty of the decoder part is to up-sample the low-level features to fit the original input size. The two parts are connected via a bottleneck. The encoder was created to extract low-level features, whereas the decoder was created to extract high-level features. Direct concatenation was utilized to merge the decoder and encoder parts. Despite the high performance of UNet with 3D architecture to benefit from the extra features, it still suffered from consuming memory and calculation complexity issues due to the vast number of parameters. Noori et al. (2019) used 2D architecture to overcome the memory issue in 3D architecture for brain tumor segmentation. One of the advantages of UNet convolutional layers is using the skip connection between the low-level characteristics from the encoder layers, and the high-level characteristics from the decoder layers using a residual unit.
However, Mahdianpari et al. (2018) claimed that using the skip connection not only facilitates the propagation of information in two directions, including forward propagation and backward propagation for computations but also assists in provide a powerful design network. Some studies used UNet for crop or weed detection and as one of the classification approaches for woody vegetation on satellite imagery.
Feature extraction and classification phase
In agriculture, there are four groups of descriptive features: visual textures, spatial contexts, spectral features, and biological morphological features (Slaughter 2014).
Visual texture feature
Humans can discern various aspects of texture through their senses, such as recognizing soft or hard, coarse or fine, and smooth or rough. In vision-based methods, the texture of an image is represented by calculating the clustered pixels’ intensity in the spatial. The Grey Level Co-occurrence Matrices (GLCM) method was used for extracting the texture feature. Texture analysis is one of the most vital features utilized in identifying plant species or focal areas to extract extra useful information (Bakhshipour et al. 2017; Kadir 2014; Wang et al. 2019a).
Van Evert et al. (2009) used two-dimensional Fourier analysis to detect Rumex obtusifolius, a broad-leaved weed, in a grass background. In the case of an image of R. obtusifolius, Fourier analysis revealed a significant contribution of high-frequency basis functions to the overall signal. The presence or absence of R. obtusifolius is determined by comparing the relative contributions of various basis functions to the original signal. The following Equation was used to convert photos from color to gray scale:
Spatial contexts feature
Principally, this feature is considered one of the oldest features, which was used to identify the plant cultivar using its leaves. In one of the first trials, Guyer et al. (1986) depended on the accounted number of leaves. Besides, McCarthy et al. (2010) used the shape and length of each leaf as a spatial feature to recognize the little corn plants. In their method, elongatedness, index, moment, and central moment were used to determine the leaf's shape. Although the shape played an important role in identifying the leaf, it was still insufficient without employing other properties (Guyer et al. 1986). The curvature method is one of the earliest methods of detecting partially occluded leaves, proposed by Franz et al. (1991) to depict the edges of both fully and not fully occluded leaves. The author elucidated the impact that a not fully occluded leaf was recognized by aligning the resampled curvatures for every species. The problem with their method was that curvature alone was not sufficient to detect different aspects of serration.
According to Tian (1995), any plant's location is determined using the spatial image features of certain cotyledons. Their method employed the stem center during the seeding stage to specify the information plant location. Woebbecke et al. (1995) suggested a technique for distinguishing between dicot and monocot weeds, which represent two types of weed plants found in the western USA. The researcher suggested that the best time to handle these weeds must be considered between 14th and 23rd days. This young age is considered the ideal age for these weeds because their shape remains stable during this period. Lin (2010) claimed that shape-based computer vision approaches are ineffective for handling the occlusion issue.
Hall et al. (2015) evaluated hand-crafted methods against self-extracted methods to depict the performance of feature extraction. To increase the challenge, the researcher created artificial images such as rotation, scaling, illumination, and occlusion to simulate the real-world conditions for the application on the Flavia dataset. The classification accuracy results demonstrated that the combination of both approaches achieved approximately 97% with a 0.6% error rate, while the machine learning method achieved 92% with a 1.6% error rate. However, the classification accuracy for ideal conditions outperformed 5.7% as an average. Weed segmentation is regularly an issue for all weed classification systems that exploit visual shapes to classify plants (Hall 2018). He used AlexNet and GoogLeNet to extract the boundary of the object or contour in a segmented plant image.
Spectral feature
Generally, most plants tend to be green regardless of their differences in size, direction, and occlusion under fixed illumination, which facilitates the segmentation process of vegetation. The worst part of vegetation segmentation occurred when some undesirable damages, occlusions, diseases, or shadows resulted in green color variation (Bai et al. 2014).
Biological morphology features
In agricultural fields, any plant or any of their parts have five biological morphology features represented by shape, structure, size, pattern, and color. Shape features play an essential role in both identifying plant species by human experts and image analysis for weed recognition (Slaughter et al. 2008; Slaughter 2014; Woebbecke et al. 1995). Some undesirable materials, such as clay or dead leaves, cause the occlusion and physical appearance changes of the leaf (Bai et al. 2014). Sarkar and Wolfe (1985) investigated the eight-neighbor code of the eight directions N, NE, NW, S, SE, SW, W, and E as a feature to identify the shape. This code represents the low degree of curvature of the chain-coded tomato boundary. The author showed that the natural shape has a high curvature, whereas the unnatural shape has a minimum angle in the abnormal area. Their algorithm was designed to work under ideal conditions such as illumination and non-occlusion leaves. Initially, many techniques based on biological morphology were conducted to detect plants’ multi-species (Franz et al. 1991; Guyer et al. 1986; Woebbecke et al. 1995). These techniques examined the shape feature of the leaf’s parts and the complete leaf in an extensive range. The shape feature of the leaf represented by curvature is a crucial tool in detecting the plant species, while the shape of the whole leaf represented by height and width was noteworthy for distinguishing the occluded and overlapped issues. The importance of these techniques is to achieve high accuracy and precision in the detection of biological morphology features within unreal-world conditions when the leaves are well-separated (Lee et al. 1999).
Søgaard (2005) utilized an active shape-based approach to handle the large-scale various plant shapes to classify the weeds. Shape or template techniques were employed to detect the leaf shape and the complete plant structure during growth stages. Notably, the previous template technique was designed to acquire an image of a seedling via a digital camera fixed on the top of the covered cylinder. The cylinder is covered with a cloth material to diffuse and decrease the amount of unrequired sunlight, or shadows. His approach applied to 19 various types of plants that were located on a Danish farm. Its usage was restricted to being applied only for the non-real computational time of weed maps, which was considered a robust processing speed compared with real-time images. Furthermore, Persson and Åstrand (2008) used active shape approaches with 19 to 53% of occluded plants in 2008 for variant training images and variant levels of description. The author concluded that the accuracy of recognizing the occluded weeds was improved by 83% after using the proper training images.
Deep learning-based feature extraction approaches are considered self-extracting features, which are the most innovative methods that are employed in this manner. The significant advantage of deep learning is that it incorporates self-learning characteristics without the need to extract these characteristics manually as in conventional machine learning methods. Table 5 depicts the features categories in vegetation.
Deep learning (DL) uses multiple convolutions to create various hierarchical representations of the input. This provides significant learning abilities as well as increases performance and precision. It is considered a multi-neural network architecture, including three models: CNN, Artificial Neural Network (ANN), and Recurrent Neural Network (RNN). Recently, CNN methods have gained popularity as an effective extractor and recognition method (Oquab et al. 2014). These features incorporate a fundamental process for the classification phase to obtain effective features using specific algorithms for the extraction process. Concretely, the main drives of these algorithms are to dispel unnecessary features and concentrate on valuable properties (Toğaçar et al. 2020). The classification stage is one of the crucial sections in any computer vision application related to images. Initially, the performance of classification depends on the features that were extracted from the previous stage. These features contain useful information, playing a crucial role in grading and determining the predicted image types.
The following section examines the most relevant works for classifying various weed species. There are two types of classification methods. The first one works under the concept of machine learning, while the second one uses deep learning convolutional architecture to classify the more convoluted issues.
Machine learning-based weed classification methods
According to the traditional methods of machine learning, the features depend on humans to extract them manually. This process is time-consuming and requires changes whenever the dataset changes (Kamilaris and Prenafeta-Boldú 2018). As a result, these characteristics are deemed highly effort-demanding, requiring well-known well-knowledge with limited generalization (Amara et al. 2017). The classification accuracy (CA) technique is used to define the relationship between the number of correctly classified patterns and the total number of patterns. It can be defined as the ratio of the sum of true positives (TP) and true negatives (TN) to the total number of trials, which is the sum of TP, false positives (FP), false negatives (FN), and TN.
Some conventional machine learning performance evaluation tools, such as SVM and Random Forest (RF), tackle the shortcomings of conventional methods. Such methods have succeeded in resolving several pixel-based classification problems (Lardeux et al. 2009). Espejo-Garcia et al. (2020) focused on two species of crops, including tomato and cotton, and two species of weeds, including velvetleaf and black nightshade, to differentiate between these two categories, which were generated manually. The outcome of this approach showed a high accuracy of the classified crop/weed plants when combined with one of the deep learning models named Densenet with one of the traditional machine learning approaches represented by support vector machine (SVM) to extract features and classify these species, respectively. Even though this approach achieved high results in classification, the occluded leaf issue was not presented fairly. There are two groups of classification methods; the first one is supervised learning, such as SVM, RF, and ANN. Table 6 summarized some related methods for plants classification.
The second group is unsupervised learning, such as K-means clustering (Kumar and Prema 2016), and practical swarm optimization (PSO) based upon K-means clustering (Bai et al. 2014). In classical CNN, a random initial value is set to pre-train the model, which has great effects on increasing the error rate from one layer to another (Wang et al. 2019a). Tang et al. (2017) used a combination of the K-means algorithm and the CNN model to identify soybeans from three species of weeds. This method handles the shortcoming in traditional machine learning by fine-tuning the weights of the initial value rather than using random weights. This method achieved 92.8% accuracy, exceedingly 1.8% higher than using the randomizing weights method, and 6% higher than the traditional CNN of two layers without fine-tuning parameters. The drawback of such a method is that it cannot be employed in real-time applications due to the time-consuming computation of a vast number of parameters. Generally, the most popular classification methods in vegetation based upon machine learning are SVM, RF, and CNN (Di Cicco et al. 2017).
Weed Detection Phase
Dyrmann et al. (2017) and Yu et al. (2019) exploited DetectNet to differentiate between crop and weed plants like wheat in real farm conditions. DetectNet is a Deep Convolutional Neural Network (DCNN) model based upon the GoogLeNet structure. It demonstrated promising results for detecting well-occluded leaves by dividing the data into two sets of training and validation with about 17,000 labeled weeds. The approach faced difficulties in locating the tiny weeds, overlapped leaves, and crops. In addition, it cannot generate a precise bounding box that fits plants in their late growth stage. The performance’s accuracy was reported at 46.3% and 86.6% on recall and precision measures, respectively. Hall (2018) applied five different approaches. The first one was a hyper-method of traditional hand-crafted features (HCF) with a deep convolutional neural network (DCNN). The other methods include DCNN, HCF, HCF-ScaleRobust, and histogram of curvature over scale (HoCS). Their results were 96.6, 95.4, 90.5, 89.3, and 72.0%, respectively. However, the hyper-method reported higher accuracy than other methods, except that it provided a consistent performance of about (± 1.3%), which was less than the (± 1%) of the DCNN approach. Otherwise, it is considered a superior consistent performance compared with the reset methods. The experimental results investigated that the accuracy of all his implementation works demonstrated low performance after adding a few extra occluded images. The disadvantages of their method are the limited training dataset and generalization (Wang et al. 2019b). VGGNet (Yu et al. 2019) outperformed GoogLeNet for detecting narrow and broadleaf weed plants in grassland. It exhibited high performance with various levels of mowing and various circumstances. Jiang et al. (2020) proposed the Graph Convolutional Network (GCN)-ResNet-101 method to recognize 6000 images of weeds and mixed crops such as lettuce, corn, and radish. The authors implemented ResNet-101 for feature extraction and then incorporated it with semi-supervised GCN for weed and crop recognition. The GCN builds the graph depending on the extracted features and establishes their Euclidean distances. The key idea in their proposed method is to find the relationships between limited labeled features from the CNN model and then measure the distance between each entity (i.e., feature) using Euclidean distances. After which, a propagation process is performed over the graph to test the samples for attaining useful features of unlabeled information from the extracted labeled features. Their GCN-ResNet-101 proposed method outperformed the compared CNN models, including AlexNet, VGG-16, and ResNet-101. It scored various accuracy levels, ranging from 96 to 99%, in recognizing four weed species.
Overfitting and underfitting are the two common causes of poor performance in convolutional networks using CNN, or RNN during training data. Overfitting means that the model starts to learn useless features (Espejo-Garcia et al. 2020), especially when the model tries to predict a trend in noisy data, caused by the vast number of parameters that can affect the performance’s accuracy. Another reason was monitored by (Kounalakis et al. 2019) related to using the data without any constraint in the training time considering that the background dominates the plant. Furthermore, the author observed that when the amount of training data is small compared to a high training rate, overfitting might occur (Kounalakis et al. 2019; Lin et al. 2020; Rangarajan and Purushothaman 2020). The model must predict any future data that was unseen before by the model to achieve the machine learning goal. Overfitting is observed when the model has well-performance in the trained data and low performance in the tested data. To avoid overfitting, there are several techniques such as dropout, augmentation, fine-tuning of the transferred learning, and background removal. Espejo-Garcia et al. (2020) showed that overfitting is a sophisticated case impacted by many factors. These factors cannot be measured using a single metric. Dropout is a robust regularization method for convolutional neural network models employed in Python with Keras.
Dropout is utilized to avoid overfitting by adding dropout layers in the backward pass training. In most cases, dropout using ignore technique of some nodes and learning the reset nodes all the possible features to enhance the learning (You et al. 2020) to generate more robust features by reducing the dependency between neurons (Dyrmann et al. 2016). Krizhevsky et al. (2012) used the dropout technique to avoid overfitting. They set zero for every single hidden node with a probability equal to 0.5. In this case, all these nodes with zero values are dropout from the participation in both foreground and backward propagation. The benefit of this technique is that it mitigates the complexity of the architecture by eliminating these nodes every time while the weights remain fixed. Furthermore, this technique assists the network in learning robust features as the nodes change their connections during the pass. The last technique in DL is image augmentation, which was applied to enhance the performance and mitigate overfitting (Lin et al. 2020) by increasing the training samples. Different transformation ways of single processing or combination of multiple processing for augmentation, include scaling (Li et al. 2018), changing the pixel intensities (Lin et al. 2020), random rotation, flips horizontally, or vertically (Noori et al. 2019), shifts, shear (Rangarajan and Purushothaman 2020), salt-pepper noise, blurring, and Gaussian noise (Espejo-Garcia et al. 2020). Furthermore, data augmentation was used to improve generalization (Espejo-Garcia et al. 2020).
To enhance the classification accuracy for a large-scale image, as with ImageNet, Simonyan and Zisserman (2014) proposed a convolutional network (ConvNet) model with more convolutional layers than the 19 weighted layers. McCool et al. (2017) presented an adapted deep convolutional neural network (DNN) called Adapted-IV3 to detect weeds in a crop field. This approach used two features: pixel statistics and shape features. Despite the ability of this method to segment the partially occluded leaves accurately, it has high computations, time-consuming for processing, and requires memory space due to the enormous numbers of parameters. These limitations restrict this method from being applied in the real-time system. Deep learning models performed with higher accuracy in classification than machine learning methods. Kamilaris and Prenafeta-Boldú (2018) showed that CNN achieved 1–8% higher CA in comparison to SVM. In addition, CNN performed more accurately than unsupervised learning, with more than 10% using the same measurement (i.e., CA) (Luus et al. 2015). Flood et al. (2019) employed U-Net to classify and map the presence and non-presence of woody trees using a very high resolution of RGB imagery. The UNet network was affected by the texture and shape of the small areas consisting of various scales. The median accuracy of this method is 90% for classifying three woody and non-woody classes. The limitation of this model is the per-pixel classification, which is quite a restricted implementation to work only with satellite imagery. Residual nets such as ResNet-50 and ResNet-101 extractors are employed by He et al. (2016) on the ImageNet dataset. They used the faster RCNN as a detector model. Both extractors have similar performance. Olsen et al. (2019) adapted the faster RCNN to handle the time-consuming issue of small objects remote sensing. They adapted the network architecture of RPN, top-down, and skip connection. A simple sampling strategy is used to accelerate the unbalanced number of classes. Their results showed an improvement in the average precision for detecting small objects in remote sensing images.
Olsen et al. (2019) implemented two CNN-based models, including Inception-v3 and ResNet-50, on multi-species of weeds from eight various regions. The total number of images in the dataset is 17509 images, which are freely available in DeepWeeds datasets. ResNet-50 showed slightly higher performance in accuracy than Inception-v3 overall. This high performance of ResNet-50 is due to the complexity of its architecture, which has a vast number of parameters compared with Inception-V3. Both of these models require a powerful GPU card for processing the complex architecture in a real-time application. Rangarajan and Purushothaman (2020) tested the most popular classifiers of deep learning networks, including ResNet-101, GoogLeNet (22 layers), AlexNet, and DensNet-201. The results of the tested networks showed that ResNet-101 gained higher accuracy than other networks. For broad-leafed Docks (i.e., Rumex), Kounalakis et al. (2019) approved that ResNet-50 outperformed ResNet-101 in weed recognition.
Gao et al. (2020) employed a combination of artificial and real data to detect the weed in the sugar beet field. Their proposed method developed a CNN-based tiny YOLO-v3 (You Only Look Once) model to detect sepium weed on a sugar beet farm. The number of generated training datasets for the artificial data was 2271, and the number for the real data was 452. K-means clustering was employed to calculate the size of the anchor box. The results of their method revealed that combining synthetic data with real data improved performance by 7% when compared to using only real data. Unfortunately, because their real-world data did not include an overlapping case, their method cannot be generalized to real-time conditions. Nevertheless, their proposed method reported high performance in terms of accuracy and speed compared with YOLO-v3 and tiny YOLO. Mask R-CNN (Region-based Convolutional Neural Network) is a robust model to obtain individual segmentation, which is known as instance segmentation. Osorio et al. (2020) employed this model to extract multi-objects from an image. Mask R-CNN provides a wealth of information regarding each detected object. The main issue with most object detection techniques is that they are concerned with locating objects using only the bounding box. The good idea with Mask R-CNN is not only to detect the location of each object but also to mask the outline of the object boundary. The cons of this model are the loss of its consistency and insufficient morphological features such as shape, structure, pattern, and size. In addition, Ren et al. (2016) claimed that Mask R-CNN is computationally intensive. Table 7 summarizes the most recent cutting-edge work on vegetation categorization.
Weed detection challenges
All the discussed methods that are aforementioned above for weed detecting and classifying, encounter difficulties. As illustrated in Fig. 5, these problems are mostly relevant to lighting, overlapping, occlusion, development stage, inadequate data, time-consuming, and low precision. To some extent, the approaches mentioned in the previous section all encounter and overcome these problems. Various kinds of difficulties are stated as follows:
-
Uncontrolled lighting conditions increase the amount of noise in an image by affecting contrast, brightness, saturation, reflections, and shadows. Thus, it minimizes the image quality and necessitates robust processing to avoid it. The normalization algorithm was used to eliminate the effects of the various illumination conditions, such as light and shadow of the color channels. However, this review shows that color transformation and threshold approaches produce low performance in the presence of high/over or low/insufficient illumination variations. Deep learning-based, on the other hand, provides the promise of outcomes in dealing with illumination difficulties. As a result, illumination causes low accuracy in the segmentation and classification results.
-
Leaves overlapping, caused by leaves stacking on top of each other to form an indistinguishable object that can be segmented as one object, is considered another serious issue. It affects vegetation processing. It decreases the performance of the weed detection approaches for crop plants and weeds. Observably, numerous studies showed that the most overlapped cases occurred in the late growth period rather than in the early stage of plant growth (Bakhshipour et al. 2017).
-
The influence of occlusion in leaves is one of the most challenging for weed detection. Some researchers depend on plant height as a solution for occlusion to recognize the weed from the crop, as the weed grows more rapidly than the crop. The spectral feature is robust enough to detect the partial occlusion issue.
-
The impact of various growth stage development plays a crucial role in changing the spectral, texture, and morphological features such as shape, leaf size, and structure. The growth stage affects classification accuracy and detection performance. In agriculture, each plant has multiple growth stages. In RGB images, the early growth stage is an arduous task for detection approaches to recognize crops from weeds due to the similarity of features at this period. Thus, to solve the growth stage issues, a sufficient number of samples is required to model the learning features of each growth stage.
Discussion and future trends
Several agricultural studies have been conducted and published, concentrating on weed plant detection. There are still some open research areas where a few have been conducted. As shown in Fig. 5, these open research areas have distinctive challenges that can be addressed in future work. One of these issues is the lack of standardized benchmarks and assessment metrics in this field. Some of these challenges, such as illumination and various growth stages, have been solved using deep learning, but it requires numerous labeled images, an accurate model with reasonable structure, and powerful GPU hardware. The majority of these researches were conducted under synthetic conditions in terms of lighting and grass density. Many methods have so far been proposed for vegetation analysis. One of these methods used an automatic model to generate the artificial dataset via a robot recognizing the crop and weed on a virtual farm. This method reduces the time-consuming annotation by the human expert (Di Cicco et al. 2017).
However, it is designed to work under unreal circumstances that have separated plants without any occlusion, as illustrated in Fig. 6. The problem with this method is that it cannot be generalized to work in real-world conditions (Ahmed et al. 2012).
McCool et al. (2017) proposed an adapted deep convolutional neural network (DNN) for weed detection. Despite the high-performance accuracy of this approach, it is still not feasible to utilize in real-time applications due to its complexity in terms of speed and memory space. Gao et al. (2020) used a combination of synthetic and real data. According to their real dataset section, there was no overlapping issue, while the artificial data contained only two simple overlapped areas. These areas are not considered complicated enough to be detected. This method achieved high performance in terms of efficiency and accuracy. Dyrmann et al. (2017) used a fully connected network of the CNN-based DetectNet model to detect well-occluded weed leaves on a wheat farm during the winter season. This method works under real conditions, except that it suffers from generalization. The accuracy of this method can be enhanced using segmentation methods. Due to the complexities in the rangeland area, spectral-based approaches are considered challenging to apply in this environment. On the other hand, image-based approaches are utilized to address the real-time images captured under various illumination conditions collected using UAVs. This challenge can be solved in computer vision by identifying the plant species through its leaves (Olsen et al. 2019). However, in occlusion, the leaf shape is not displayed accurately, and the variance in the appearance is increased. For these reasons, there is insufficient information to learn the detector and classifier how to extract useful features (Zhou and Yuan 2019). Despite advanced weed detection approaches based on machine learning in the last decades, several challenges are faced with the real applications in the pasture including leaves occlusion, varying illuminations, and various growth stages (Wang et al. 2019b). Thus, these limitations motivate most machine learning techniques to work less sensibly (Slaughter et al. 2008). In this regard, further investigation researches were conducted in computer vision applications to handle these shortcomings (Lee et al. 1999). Self-learning approaches, such as CNN models, represent the most state-of-the-art approaches compared with hand-crafted methods like SVM, since they have less effect on these shortages (Espejo-Garcia et al. 2020). In addition, CNN has more accurate results than the support vector machine (SVM) and artificial neural network (ANN), and it reduces the problem of feature extraction selection (Osorio et al. 2020).
Additionally, this review highlighted the four common procedures for processing weed detection in agriculture. Some approaches have been tested and evaluated using images that were taken under controlled conditions (Di Cicco et al. 2017; Haug et al. 2014; Montalvo et al. 2012). Other works implemented their approaches using images that were taken in real-world conditions (Koščević et al. 2020; Kounalakis et al. 2019; Kumar and Prema 2016; Liu et al. 2016) while others used mixed of artificial and real data (Gao et al. 2020). From literature analysis, weed detection has various tasks. Several factors that affect detection performance are shown in Fig. 5.
-
Some weed species share similarities in their features with crops. This will increase the challenge of the detection scenario.
-
Most challenging part of weed detection is the classification of two green plants, represented by weed and grass, using real-world applications. The illumination issue affects the image contrast, while occlusion and overlapping issues impact the extracted features and then result in low classification accuracy.
-
ML-based methods are not suitable to solve sophisticated problems like occlusion (Adhikari et al. 2019; Slaughter et al. 2008), which are overlapped due to their shallow layer to extract sufficient information. In the last decades, the trends to employ DL architecture networks relying upon CNN models (Dyrmann et al. 2017; Espejo-Garcia et al. 2020; Gao et al. 2020; Osorio et al. 2020) are dramatically increased by the researchers to overcome the knotted issue of ML methods as depicted in Table 8.
-
A combination of hand-crafted and deep learning models introduces the opportunity for a more robust model that is able to detect the partially occluded leaves so that the classification accuracy is enhanced. This combination overcomes the limitation of manually-designed extracting features in the shallow layer and overcomes the overfitting case in deep learning architecture.
Many works on detecting various species of weed plants in agriculture using machine learning and deep learning were discussed. In traditional machine learning, various color space transformations and segmentation approaches were utilized to separate the foreground from the background (Chen et al. 2017; Milioto et al. 2018). The obtained results are satisfactory in terms of efficiency. For more complicated issues such as real-world data, deep learning-based models produce promising performance in terms of accuracy (Di Cicco et al. 2017; Tang et al. 2017). In vegetation, the weed and crop leaves share similar features, especially in color and shape, which raises the difficulty of recognizing them.
In general, this study summarizes the features of leaves into four categories: biological morphology features, visual texture features, spectral features, and spatial contextual features. In deep learning, a tremendous number of possible features can be extracted to boost the performance of prediction results. For classification, single or multi-approaches can be performed by hybrid machine learning methods with deep learning to produce one robust detection model. This model can be utilized in a robotic weeding system to perform precision farming or automated precision weeding. Various growth stages, diseases, illumination, overlapping, and occlusion are the most sophisticated issues of real-world data that face computer vision techniques in nature. The dedicated methods to overcome these issues need to be enhanced in terms of efficiency and effectiveness. A vast amount of data is considered a further issue facing researchers when they apply deep learning models to avoid overfitting and to generalize the model in reality.
Several future suggestions have been proposed. Firstly, the selected dataset should be sufficient and comprehensive to cover various real-world conditions for a robust model. Secondly, the intensive problem requires a more convoluted model to address it. For this reason, each scenario must be analyzed and studied thoroughly to trade-off the complexity and efficiency. Therefore, developing an automated weed detection application to detect multi-species of weeds under real-world conditions remains an open challenge in agriculture. To our best knowledge, one key point to be taken into account is utilizing the high dimensional resolution of multispectral images with CNN models to tackle the detection of weeds under real-world conditions. In the future, thorough investigations into new robotic machines that improve weed detection by reducing the effects of real-world data in terms of efficiency and complexity are being considered.
Data availability
Data availability data and materials are made available upon request.
Code availability
Codes, algorithm, and trained model are made available upon request.
References
Abouzahir S, Sadik M, Sabir E (2018) Enhanced approach for weeds species detection using machine vision. In: 2018 international conference on electronics, control, optimization and computer science (ICECOCS). IEEE, pp 1–6. https://doi.org/10.1109/icecocs.2018.8610505
Adams J, Qiu Y, Xu Y, Schnable JC (2020) Plant segmentation by supervised machine learning methods. Plant Phenom J 3(1):e20001. https://doi.org/10.1002/ppj2.20001
Adhikari SP, Yang H, Kim H (2019) Learning semantic graphics using convolutional encoder–decoder network for autonomous weeding in paddy. Front Plant Sci 10:1404. https://doi.org/10.3389/fpls.2019.01404
Ahmed F, Al-Mamun HA, Bari AH, Hossain E, Kwan P (2012) Classification of crops and weeds from digital images: a support vector machine approach. Crop Prot 40:98–104. https://doi.org/10.1016/j.cropro.2012.04.024
Alam M, Alam MS, Roman M, Tufail M, Khan MU, Khan MT (2020) Real-time machine-learning based crop/weed detection and classification for variable-rate spraying in precision agriculture. In: 2020 7th international conference on electrical and electronics engineering (ICEEE). IEEE, pp 273–280. https://doi.org/10.1109/ICEEE49618.2020.9102505
Ali H, Lali M, Nawaz MZ, Sharif M, Saleem B (2017) Symptom based automated detection of citrus diseases using color histogram and textural descriptors. Comput Electron Agric 138:92–104. https://doi.org/10.1016/j.compag.2017.04.008
Amara J, Bouaziz B, Algergawy A (2017) A deep learning-based approach for banana leaf diseases classification. Datenbanksysteme für Business, Technologie und Web (BTW 2017)-Workshopband, pp 79–88. https://dl.gi.de/handle/20.500.12116/944
Badrinarayanan V, Kendall A, Cipolla R (2017) Segnet: a deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans Pattern Anal Mach Intell 39(12):2481–2495. https://doi.org/10.1109/TPAMI.2016.2644615
Bai X, Cao Z, Wang Y, Yu Z, Hu Z, Zhang X, Li C (2014) Vegetation segmentation robust to illumination variations based on clustering and morphology modelling. Biosys Eng 125:80–97. https://doi.org/10.1016/j.biosystemseng.2014.06.015
Bakhshipour A, Jafari A (2018) Evaluation of support vector machine and artificial neural networks in weed detection using shape features. Comput Electron Agric 145:153–160. https://doi.org/10.1016/j.compag.2017.12.032
Bakhshipour A, Jafari A, Nassiri SM, Zare D (2017) Weed segmentation using texture features extracted from wavelet sub-images. Biosys Eng 157:1–12. https://doi.org/10.1016/j.biosystemseng.2017.02.002
Behmann J, Mahlein A-K, Rumpf T, Römer C, Plümer L (2015) A review of advanced machine learning methods for the detection of biotic stress in precision crop protection. Precis Agric 16(3):239–260. https://doi.org/10.1007/s11119-014-9372-7
Bergasa LM, Mazo M, Gardel A, Sotelo M, Boquete L (2000) Unsupervised and adaptive Gaussian skin-color model. Image vis Comput 18(12):987–1003. https://doi.org/10.1016/S0262-8856(00)00042-1
Binch A, Fox C (2017) Controlled comparison of machine vision algorithms for Rumex and Urtica detection in grassland. Comput Electron Agric 140:123–138. https://doi.org/10.1016/j.compag.2017.05.018
Brinkhoff J, Vardanega J, Robson AJ (2020) Land cover classification of nine perennial crops using sentinel-1 and-2 data. Remote Sens 12(1):96. https://doi.org/10.3390/rs12010096
Burgos-Artizzu XP, Ribeiro A, Tellaeche A, Pajares G, Fernández-Quintanilla C (2009) Improving weed pressure assessment using digital images from an experience-based reasoning approach. Comput Electron Agric 65(2):176–185. https://doi.org/10.1016/j.compag.2008.09.001
Caruana R (1997) Multitask learning. Mach Learn 28(1):41–75. https://doi.org/10.1023/A:1007379606734
Chen Y, He X, Wang J, Xiao R (2014) The influence of polarimetric parameters and an object-based approach on land cover classification in coastal wetlands. Remote Sens 6(12):12575–12592. https://doi.org/10.3390/rs61212575
Chen SW, Shivakumar SS, Dcunha S, Das J, Okon E, Qu C, Taylor CJ, Kumar V (2017) Counting apples and oranges with deep learning: a data-driven approach. IEEE Robot Autom Lett 2(2):781–788. https://doi.org/10.1109/LRA.2017.2651944
Di Cicco M, Potena C, Grisetti G, Pretto A (2017) Automatic model based dataset generation for fast and accurate crop and weeds detection. In 2017 IEEE/RSJ international conference on intelligent robots and systems (IROS), Vancouver, BC, Canada, 24–28 Sept. 2017. IEEE, pp 5188–5195. https://doi.org/10.1109/IROS.2017.8206408
Dos Santos FA, Freitas DM, Da Silva GG, Pistori H, Folhes MT (2019) Unsupervised deep learning and semi-automatic data labeling in weed discrimination. Comput Electron Agric 165:104963. https://doi.org/10.1016/j.compag.2019.104963
Dyrmann M, Karstoft H, Midtiby HS (2016) Plant species classification using deep convolutional neural network. Biosys Eng 151:72–80. https://doi.org/10.1016/j.biosystemseng.2016.08.024
Dyrmann M, Jørgensen RN, Midtiby HS (2017) RoboWeedSupport-Detection of weed locations in leaf occluded cereal crops using a fully convolutional neural network. Adv Anim Biosci 8(2):842–847. https://doi.org/10.1017/S2040470017000206
Espejo-Garcia B, Mylonas N, Athanasakos L, Fountas S, Vasilakoglou I (2020) Towards weeds identification assistance through transfer learning. Comput Electron Agric 171:105306. https://doi.org/10.1016/j.compag.2020.105306
Etienne A (2019) Automated weed detection using machine learning techniques on uas-acquired imagery. Dissertation, Purdue University Graduate School, Lafayette
Fernández-Quintanilla C, Peña J, Andújar D, Dorado J, Ribeiro A, López-Granados F (2018) Is the current state of the art of weed monitoring suitable for site-specific weed management in arable crops? Weed Res 58(4):259–272. https://doi.org/10.1111/wre.12307
Finlay S (2012) Data transformation (pre-processing). In: Credit scoring, response modeling, and insurance rating: a practical guide to forecasting consumer behavior. Palgrave Macmillan, London, pp 144–164
Flood N, Watson F, Collett L (2019) Using a U-net convolutional neural network to map woody vegetation extent from high resolution satellite imagery across Queensland, Australia. Int J Appl Earth Obs Geoinf 82:101897. https://doi.org/10.1016/j.jag.2019.101897
Franz E, Gebhardt M, Unklesbay K (1991) Shape description of completely visible and partially occluded leaves for identifying plants in digital images. Trans ASAE 34(2):673–681. https://doi.org/10.13031/2013.31716
Gao J, Fan W, Jiang J, Han J (2008) Knowledge transfer via multiple model local structure mapping. In: Proceedings of the 14th ACM SIGKDD international conference on Knowledge discovery and data mining, pp 283–291. https://doi.org/10.1145/1401890.1401928
Gao J, Nuyttens D, Lootens P, He Y, Pieters JG (2018) Recognising weeds in a maize crop using a random forest machine-learning algorithm and near-infrared snapshot mosaic hyperspectral imagery. Biosys Eng 170:39–50. https://doi.org/10.1016/j.biosystemseng.2018.03.006
Gao J, French AP, Pound MP, He Y, Pridmore TP, Pieters JG (2020) Deep convolutional neural networks for image-based Convolvulus sepium detection in sugar beet fields. Plant Methods 16(1):1–12. https://doi.org/10.1186/s13007-020-00570-z
Guo W, Rage UK, Ninomiya S (2013) Illumination invariant segmentation of vegetation for time series wheat images based on decision tree model. Comput Electron Agric 96:58–66. https://doi.org/10.1016/j.compag.2013.04.010
Guyer DE, Miles G, Schreiber M, Mitchell O, Vanderbilt V (1986) Machine vision and image processing for plant identification. Trans ASAE 29(6):1500–1507. https://doi.org/10.13031/2013.30344
Hall DR (2018) A rapidly deployable approach for automated visual weed classification without prior species knowledge. PhD thesis, Queensland University of Technology, Brisbane
Hall D, Mccool C, Dayoub F, Sunderhauf N, Upcroft B (2015) Evaluation of features for leaf classification in challenging conditions. In: 2015 IEEE winter conference on applications of computer vision, IEEE, pp 797–804. https://doi.org/10.1109/WACV.2015.111
Hall D, Dayoub F, Kulk J, Mccool C (2017) Towards unsupervised weed scouting for agricultural robotics. In: 2017 IEEE international conference on robotics and automation (ICRA). IEEE, pp 5223–5230. https://doi.org/10.1109/ICRA.2017.7989612
Hamuda E, Glavin M, Jones E (2016) A survey of image processing techniques for plant extraction and segmentation in the field. Comput Electron Agric 125:184–199. https://doi.org/10.1016/j.compag.2016.04.024
Hamuda E, Mc Ginley B, Glavin M, Jones E (2017) Automatic crop detection under field conditions using the HSV colour space and morphological operations. Comput Electron Agric 133:97–107. https://doi.org/10.1016/j.compag.2016.11.021
Hamuda E, Mc Ginley B, Glavin M, Jones E (2018) Improved image processing-based crop detection using Kalman filtering and the Hungarian algorithm. Comput Electron Agric 148:37–44. https://doi.org/10.1016/j.compag.2018.02.027
Hari SS, Sivakumar M, Renuga P, Suriya S (2019) Detection of plant disease by leaf image using convolutional neural network. In: 2019 international conference on vision towards emerging trends in communication and networking (ViTECoN), Vellore, India. IEEE, pp 1–5. https://doi.org/10.1109/vitecon.2019.8899748
Haug S, Michaels A, Biber P, Ostermann J (2014) Plant classification system for crop/weed discrimination without segmentation. In: IEEE winter conference on applications of computer vision, Steamboat Springs, CO, USA. IEEE, pp 1142–1149. https://doi.org/10.1109/wacv.2014.6835733
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 770–778. https://doi.org/10.1109/cvpr.2016.90
Herrmann I, Shapira U, Kinast S, Karnieli A, Bonfil D (2013) Ground-level hyperspectral imagery for detecting weeds in wheat fields. Precis Agric 14(6):637–659. https://doi.org/10.1007/s11119-013-9321-x
Husham A, Hazim Alkawaz M, Saba T, Rehman A, Saleh Alghamdi J (2016) Automated nuclei segmentation of malignant using level sets. Microsc Res Tech 79(10):993–997. https://doi.org/10.1002/jemt.22733
Jiang H, Zhang C, Qiao Y, Zhang Z, Zhang W, Song C (2020) CNN feature based graph convolutional network for weed and crop recognition in smart farming. Comput Electron Agric 174:105450. https://doi.org/10.1016/j.compag.2020.105450
Kadir A (2014) A model of plant identification system using glcm, lacunarity and shen features. arXiv preprint arXiv:1410.0969. https://doi.org/10.48550/arXiv.1410.0969
Kamilaris A, Prenafeta-Boldú FX (2018) Deep learning in agriculture: a survey. Comput Electron Agric 147:70–90. https://doi.org/10.1016/j.compag.2018.02.016
Kazmi W, Garcia-Ruiz FJ, Nielsen J, Rasmussen J, Andersen HJ (2015) Detecting creeping thistle in sugar beet fields using vegetation indices. Comput Electron Agric 112:10–19. https://doi.org/10.1016/j.compag.2015.01.008
Koščević K, Subašić M, Lončarić S (2020) Deep learning-based illumination estimation using light source classification. IEEE Access 8:84239–84247. https://doi.org/10.1109/ACCESS.2020.2992121
Kounalakis T, Triantafyllidis GA, Nalpantidis L (2016) Weed recognition framework for robotic precision farming. In: 2016 IEEE international conference on imaging systems and techniques (IST). IEEE, pp 466–471. https://doi.org/10.1109/IST.2016.7738271
Kounalakis T, Triantafyllidis GA, Nalpantidis L (2018) Image-based recognition framework for robotic weed control systems. Multimedia Tools Appl 77(8):9567–9594. https://doi.org/10.1007/s11042-017-5337-y
Kounalakis T, Triantafyllidis GA, Nalpantidis L (2019) Deep learning-based visual recognition of rumex for robotic precision farming. Comput Electron Agric 165:104973. https://doi.org/10.1016/j.compag.2019.104973
Krizhevsky A, Sutskever I, Hinton GE (2012) Imagenet classification with deep convolutional neural networks. In: Advances in neural information processing systems, pp 1097–1105. https://doi.org/10.1145/3065386
Kumar DA, Prema P (2016) A novel wrapping curvelet transformation based angular texture pattern (WCTATP) extraction method for weed identification. ICTACT J Image Video Process. https://doi.org/10.21917/ijivp.2016.0174
Lardeux C, Frison P-L, Tison C, Souyris J-C, Stoll B, Fruneau B, Rudant J-P (2009) Support vector machine for multifrequency SAR polarimetric data classification. IEEE Trans Geosci Remote Sens 47(12):4143–4152. https://doi.org/10.1109/TGRS.2009.2023908
Lee WS, Slaughter D, Giles D (1999) Robotic weed control system for tomatoes. Precis Agric 1(1):95–113. https://doi.org/10.1023/A:1009977903204
Lee SH, Chan CS, Wilkin P, Remagnino P (2015) Deep-plant: plant identification with convolutional neural networks. In: 2015 IEEE international conference on image processing (ICIP). IEEE, pp 452–456. https://doi.org/10.1109/ICIP.2015.7350839
Li N, Grift TE, Yuan T, Zhang C, Momin MA, Li W (2016) Image processing for crop/weed discrimination in fields with high weed pressure. In: 2016 ASABE annual international meeting, American Society of Agricultural and Biological Engineers, p 1. https://doi.org/10.13031/aim.20162460475
Li X, Chen H, Qi X, Dou Q, Fu C-W, Heng P-A (2018) H-DenseUNet: hybrid densely connected UNet for liver and tumor segmentation from CT volumes. IEEE Trans Med Imaging 37(12):2663–2674. https://doi.org/10.1109/TMI.2018.2845918
Liakos KG, Busato P, Moshou D, Pearson S, Bochtis D (2018) Machine learning in agriculture: a review. Sensors 18(8):2674. https://doi.org/10.3390/s18082674
Lin C (2010) A support vector machine embedded weed identification system. Master Thesis, University of Illinois at Urbana-Champaign, Urbana
Lin A, Wu J, Yang X (2020) A data augmentation approach to train fully convolutional networks for left ventricle segmentation. Magn Reson Imaging 66:152–164. https://doi.org/10.1016/j.mri.2019.08.004
Liu T, Wu W, Chen W, Sun C, Zhu X, Guo W (2016) Automated image-processing for counting seedlings in a wheat field. Precis Agric 17(4):392–406. https://doi.org/10.1007/s11119-015-9425-6
Long J, Shelhamer E, Darrell T (2015) Fully convolutional networks for semantic segmentation. In: Proceedings of the IEEE conference on computer vision and pattern recognition, Boston, MA, USA, 7–12 June 2015. IEEE, pp 3431–3440. https://doi.org/10.1109/cvpr.2015.7298965
Lottes P, Hoeferlin M, Sander S, Müter M, Schulze P, Stachniss LC (2016) An effective classification system for separating sugar beets and weeds for precision farming applications. In: 2016 IEEE international conference on robotics and automation (ICRA), Stockholm, Sweden, 16–21 May 2016. IEEE, pp 5157–5163. https://doi.org/10.1109/icra.2016.7487720
Luus FP, Salmon BP, Van Den Bergh F, Maharaj BTJ (2015) Multiview deep learning for land-use classification. IEEE Geosci Remote Sens Lett 12(12):2448–2452. https://doi.org/10.1109/LGRS.2015.2483680
Mahdianpari M, Salehi B, Rezaee M, Mohammadimanesh F, Zhang Y (2018) Very deep convolutional neural networks for complex land cover mapping using multispectral remote sensing imagery. Remote Sens 10(7):1119. https://doi.org/10.3390/rs10071119
Mccarthy CL, Hancock NH, Raine SR (2010) Applied machine vision of plants: a review with implications for field deployment in automated farming operations. Intel Serv Robot 3(4):209–217. https://doi.org/10.1007/s11370-010-0075-2
Mccool C, Perez T, Upcroft B (2017) Mixtures of lightweight deep convolutional neural networks: applied to agricultural robotics. IEEE Robot Autom Lett 2(3):1344–1351. https://doi.org/10.1109/LRA.2017.2667039
Meyer GE, Neto JC (2008) Verification of color vegetation indices for automated crop imaging applications. Comput Electron Agric 63(2):282–293. https://doi.org/10.1016/j.compag.2008.03.009
Meyer GE, Neto JC, Jones DD, Hindman TW (2004) Intensified fuzzy clusters for classifying plant, soil, and residue regions of interest from color images. Comput Electron Agric 42(3):161–180. https://doi.org/10.1016/j.compag.2003.08.002
Milioto A, Lottes P, Stachniss C (2018) Real-time semantic segmentation of crop and weed for precision agriculture robots leveraging background knowledge in CNNs. In: 2018 IEEE international conference on robotics and automation (ICRA), Brisbane, QLD, Australia, 21–25 May 2018. IEEE, pp 2229–2235. https://doi.org/10.1109/icra.2018.8460962
Mohammadimanesh F, Salehi B, Mahdianpari M, Gill E, Molinier M (2019) A new fully convolutional neural network for semantic segmentation of polarimetric SAR imagery in complex land cover ecosystem. ISPRS J Photogram Remote Sens 151:223–236. https://doi.org/10.1016/j.isprsjprs.2019.03.015
Montalvo M, Pajares G, Guerrero JM, Romeo J, Guijarro M, Ribeiro A, Ruz JJ, Cruz J (2012) Automatic detection of crop rows in maize fields with high weeds pressure. Expert Syst Appl 39(15):11889–11897. https://doi.org/10.1016/j.eswa.2012.02.117
Mortensen AK, Dyrmann M, Karstoft H, Jørgensen RN, Gislum R (2016) Semantic segmentation of mixed crops using deep convolutional neural network. In: International conference on agricultural engineering. Aarhus University, p 6
Namin ST, Esmaeilzadeh M, Najafi M, Brown TB, Borevitz JO (2018) Deep phenotyping: deep learning for temporal phenotype/genotype classification. Plant Methods 14(1):66. https://doi.org/10.1186/s13007-018-0333-4
Noori M, Bahri A, Mohammadi K (2019) Attention-guided version of 2D UNet for automatic brain tumor segmentation. In: 2019 9th international conference on computer and knowledge engineering (ICCKE). IEEE, pp 269–275. https://doi.org/10.1109/ICCKE48569.2019.8964956
Olsen A, Konovalov DA, Philippa B, Ridd P, Wood JC, Johns J, Banks W, Girgenti B, Kenny O, Whinney J (2019) DeepWeeds: a multiclass weed species image dataset for deep learning. Sci Rep 9(1):1–12. https://doi.org/10.1038/s41598-018-38343-3
Oquab M, Bottou L, Laptev I, Sivic J (2014) Learning and transferring mid-level image representations using convolutional neural networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 1717–1724. https://doi.org/10.1109/cvpr.2014.222
Osorio K, Puerto A, Pedraza C, Jamaica D, Rodríguez L (2020) A deep learning approach for weed detection in lettuce crops using multispectral images. AgriEngineering 2(3):471–488. https://doi.org/10.3390/agriengineering2030032
Otsu N (1979) A threshold selection method from gray-level histograms. IEEE Trans Syst Man Cybern 9(1):62–66. https://doi.org/10.1109/CAR.2010.5456687
Pan SJ, Yang Q (2009) A survey on transfer learning. IEEE Trans Knowl Data Eng 22(10):1345–1359. https://doi.org/10.1109/TKDE.2009.191
Persson M, Åstrand B (2008) Classification of crops and weeds extracted by active shape models. Biosys Eng 100(4):484–497. https://doi.org/10.1016/j.biosystemseng.2008.05.003
Prasanna Mohanty S, Hughes D, Salathe M (2016) Using deep learning for image-based plant disease detection. Front Plant Sci 7:1419. https://doi.org/10.3389/fpls.2016.01419
Rangarajan AK, Purushothaman R (2020) Disease classification in eggplant using pre-trained VGG16 and MSVM. Sci Rep 10(1):1–11. https://doi.org/10.1038/s41598-020-59108-x
Ren S, He K, Girshick R, Sun J (2016) Faster R-CNN: towards real-time object detection with region proposal networks. IEEE Trans Pattern Anal Mach Intell 39(6):1137–1149. https://doi.org/10.1109/tpami.2016.2577031
Ruiz-Ruiz G, Gómez-Gil J, Navas-Gracia L (2009) Testing different color spaces based on hue for the environmentally adaptive segmentation algorithm (EASA). Comput Electron Agric 68(1):88–96. https://doi.org/10.1016/j.compag.2009.04.009
Sarkar N, Wolfe R (1985) Feature extraction techniques for sorting tomatoes by computer vision. Trans ASAE 28(3):970–0974. https://doi.org/10.13031/2013.32371
Sathesh A, Rasitha K (2010) A nonlinear adaptive median filtering based noise removal algorithm. In: Proceedings of 1st international conference on modeling, control, automation and communication (ICMCAC-2010), pp 108–113
Šeatović D, Grüninger R, Anken T, Holpp M (2008) 3D object recognition, localization and treatment of rumex obtusifolius in its natural environment. In: 1st international conference on machine control & guidance, June 24, pp 24–26
Sharma A, Jain A, Gupta P, Chowdary V (2020) Machine learning applications for precision agriculture: a comprehensive review. IEEE Access. https://doi.org/10.1109/ACCESS.2020.3048415
Simonyan K, Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556. https://doi.org/10.48550/arXiv.1409.1556
Sladojevic S, Arsenovic M, Anderla A, Culibrk D, Stefanovic D (2016) Deep neural networks based recognition of plant diseases by leaf image classification. Comput Intell Neurosci. https://doi.org/10.1155/2016/3289801
Slaughter DC (2014) The biological engineer: sensing the difference between crops and weeds. In: Automation: the future of weed control in cropping systems. Springer, pp 71–95
Slaughter D, Giles D, Downey D (2008) Autonomous robotic weed control systems: a review. Comput Electron Agric 61(1):63–78. https://doi.org/10.1016/j.compag.2007.05.008
Søgaard HT (2005) Weed classification by active shape models. Biosys Eng 91(3):271–281. https://doi.org/10.1016/j.biosystemseng.2005.04.011
Sørensen RA, Rasmussen J, Nielsen J, Jørgensen RN (2017) Thistle detection using convolutional neural networks. In: 2017 Efita WCCA Congress, p 161
Tang J, Wang D, Zhang Z, He L, Xin J, Xu Y (2017) Weed identification based on K-means feature learning combined with convolutional neural network. Comput Electron Agric 135:63–70. https://doi.org/10.1016/j.compag.2017.01.001
Tian L (1995) Knowledge-based machine vision system for outdoor plant identification. Doctoral dissertation, University of California, Davis
Tian LF, Slaughter DC (1998) Environmentally adaptive segmentation algorithm for outdoor image segmentation. Comput Electron Agric 21(3):153–168. https://doi.org/10.1016/S0168-1699(98)00037-4
Tian H, Wang T, Liu Y, Qiao X, Li Y (2020) Computer vision technology in agricultural automation—a review. Inf Process Agric 7(1):1–19. https://doi.org/10.1016/j.inpa.2019.09.006
Toğaçar M, Ergen B, Cömert Z (2020) Classification of flower species by using features extracted from the intersection of feature selection methods in convolutional neural network models. Measurement 158:107703. https://doi.org/10.1016/j.measurement.2020.107703
Van Evert F, Polder G, Van Der Heijden G, Kempenaar C, Lotz L (2009) Real-time vision-based detection of Rumex obtusifolius in grassland. Weed Res 49(2):164–174. https://doi.org/10.1111/j.1365-3180.2008.00682.x
Vijayalakshmi B, Mohan V (2016) Kernel-based PSO and FRVM: an automatic plant leaf type detection using texture, shape, and color features. Comput Electron Agric 125:99–112. https://doi.org/10.1016/j.compag.2016.04.033
Volpi M, Tuia D (2016) Dense semantic labeling of subdecimeter resolution images with convolutional neural networks. IEEE Trans Geosci Remote Sens 55(2):881–893. https://doi.org/10.1109/TGRS.2016.2616585
Wang A, Zhang W, Wei X (2019a) A review on weed detection using ground-based machine vision and image processing techniques. Comput Electron Agric 158:226–240. https://doi.org/10.1016/j.compag.2019.02.005
Wang C, Peng X, Liu M, Xing Z, Bai X, Xie B, Wang T (2019b) A learning-based approach for automatic construction of domain glossary from source code and documentation. In: Proceedings of the 2019b 27th ACM joint meeting on European software engineering conference and symposium on the foundations of software engineering, August 2019b, pp 97–108. https://doi.org/10.1145/3338906.3338963
Weis M, Sökefeld M (2010) Detection and identification of weeds. Precis Crop Protect—Chall Use Heterogen. https://doi.org/10.1007/978-90-481-9277-9_8
Weis M, Gutjahr C, Ayala VR, Gerhards R, Ritter C, Schölderle F (2008) Precision farming for weed management: techniques. Gesunde Pflanzen 60(4):171–181. https://doi.org/10.1007/s10343-008-0195-1
Woebbecke D, Meyer G, Von Bargen K, Mortensen D (1995) Shape features for identifying young weeds using image analysis. Trans ASAE 38(1):271–281. https://doi.org/10.13031/2013.27839
You J, Liu W, Lee J (2020) A DNN-based semantic segmentation for detecting weed and crop. Comput Electron Agric 178:105750. https://doi.org/10.1016/j.compag.2020.105750
Yu Z, Cao Z, Wu X, Bai X, Qin Y, Zhuo W, Xiao Y, Zhang X, Xue H (2013) Automatic image-based detection technology for two critical growth stages of maize: emergence and three-leaf stage. Agric for Meteorol 174:65–84. https://doi.org/10.1016/j.agrformet.2013.02.011
Yu J, Sharpe SM, Schumann AW, Boyd NS (2019) Deep learning for image-based weed detection in turfgrass. Eur J Agron 104:78–84. https://doi.org/10.1016/j.eja.2019.01.004
Zhang W, Hansen MF, Volonakis TN, Smith M, Smith L, Wilson J, Ralston G, Broadbent L, Wright G (2018) Broad-leaf weed detection in pasture. In: 2018 IEEE 3rd international conference on image, vision and computing (ICIVC), Chongqing, China, 27–29 June 2018. IEEE, pp 101–105. https://doi.org/10.1109/ICIVC.2018.8492831
Zhang S, Guo J, Wang Z (2019) Combing K-means clustering and local weighted maximum discriminant projections for weed species recognition. Front Comput Sci 1:4. https://doi.org/10.3389/fcomp.2019.00004
Zhao X, Yuan Y, Song M, Ding Y, Lin F, Liang D, Zhang D (2019) Use of unmanned aerial vehicle imagery and deep learning unet to extract rice lodging. Sensors 19(18):3859. https://doi.org/10.3390/s19183859
Zheng L, Zhang J, Wang Q (2009) Mean-shift-based color segmentation of images containing green vegetation. Comput Electron Agric 65(1):93–98. https://doi.org/10.1016/j.compag.2008.08.002
Zheng L, Shi D, Zhang J (2010) Segmentation of green vegetation of crop canopy images based on mean shift and fisher linear discriminant. Pattern Recogn Lett 31(9):920–925. https://doi.org/10.1016/j.patrec.2010.01.016
Zhou C, Yuan J (2019) Occlusion pattern discovery for object detection and occlusion reasoning. IEEE Trans Circuits Syst Video Technol 30(7):2067–2080. https://doi.org/10.1109/TCSVT.2019.2909982
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Al-Badri, A.H., Ismail, N. ., Al-Dulaimi, K. et al. Classification of weed using machine learning techniques: a review—challenges, current and future potential techniques. J Plant Dis Prot 129, 745–768 (2022). https://doi.org/10.1007/s41348-022-00612-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41348-022-00612-9