Abstract
Higher crop productivity in a sustainable manner is being perceived as one of the most crucial factors to ensure food security in light of the shrinking arable land resources and limiting fresh water resources. Soil salinity is the chief climatic constraint to agricultural productivity, restricting the suitability of agricultural land and affecting both biomass and grain yields in crops, including rice. The objective of this study was to screen a collection of rice mutants (O. sativa L. cv IR64) produced by γ-ray irradiation, for their higher yield and biomass under saline conditions. The initial screening was carried out at seedling stage employing hydroponics system, followed by screening in saline microplots under standard agronomic practices. Mircoplots were maintained at three different saline levels i.e., low salinity (EC ∼ 6 dS/m), moderate salinity (EC ∼ 10 dS/m) and high salinity (EC ∼ 14 dS/m). Based on screening carried out for various yield and related parameters i.e., plant height, tiller number, shoot and root weight, total biomass, panicle length, and harvest index, four ideotypes were observed to be performing significantly better than the wild type plants. Most importantly, selected mutants, such as D100-211 and D100-209 showed an increase of 18 and 34% in yield as compared to WT plants under moderate saline conditions (EC = 10 dS/m). These results suggest that regulated genetic modulation can improve crops to get optimum biomass and yield despite environmental vagaries. Further, detailed genetic and molecular characterization of these mutants will help to identify and characterize the key genes regulating the traits for high biomass and yield under salinity stress in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
By 2050, the world’s population will reach 9.1 billion. To keep pace with this situation, it is a great challenge to increase the crop productivity by up to 70% (Joshi et al. 2016a). Globally, more than 80 Mha of agricultural land is already salt affected to varying degrees, and 1.5 Mha of arable land is expected to be further lost each year as a result of salinization due to climate change and irrigation practices (Kushwaha et al. 2012; Kumar et al. 2015; Joshi et al. 2016a). Soil salinity can impair cellular structures and disrupt various physiological processes, thereby preventing plant growth and development. In addition, osmotic stress leads to inhibition of photosynthesis, metabolic dysfunction and turgor loss resulting in membrane disorganization, growth retardation, reduced fertility, premature senescence, severe penalty of yield and harvestable biomass (Joshi et al. 2014; Soda et al. 2016). Salinity tolerance is a quantitative trait governed by a complex gene network regulating various physiological and biochemical processes (Zheng et al. 2015). Plants adapt several strategies to survive under salt stress i.e., salt ion exclusion from the cytoplasm to extracellular space, inhibition of ion uptake by the roots and their active transport into the leaves, vacuolar compartmentalization of toxic ions, disruption of cellular membrane, proline accumulation and enhanced antioxidative enzyme activity (Song et al. 2012; Soni et al. 2013).
Increasing population and fossil fuel-based energy consumption have contributed to a great extent in environmental pollution in developing countries (Ullah et al. 2015). However, to maintain uninterrupted energy demand for sustainable development, renewable energy like biomass is the need of the hour as the only greener sources of energy (Ahmed et al. 2016; Mazzola et al. 2016). There is currently an increasing demand in biomass based biofuels as it is essentially carbon neutral and provides a convenient way for energy storage (Chandra et al. 2012). In contrast to first generation biofuels, second generation biofuels, solves food versus fuel competition by processing lignocellulosic biomass, which is a cheap, abundant, renewable and non-edible residue of food crops (Ullah et al. 2015).
Rice is one of the major food crops of India in terms of area and productivity and 15 Mt of rice straw is burnt alone in Punjab every year (Singh et al. 2016a). In addition, rice is a valuable source of feedstock for bio-refinery since it can produce a lot of biomass (Kondhia et al. 2015). Thus, to prevent environmental pollution and proper utilization of renewable resources, it is desirable to ensure large scale production of second generation bio-fuel to meet the global energy demands by identifying high yielding rice varieties outperforming under saline conditions. In addition, development of salt tolerant lines can also provide a suitable way to meet the growing food demand, even in unfavorable areas (Song et al. 2012; Kondhia et al. 2015).
With several years of considerable efforts, almost 2000 trait genes including both single Mendelian loci/genes and quantitative trait loci (QTLs) have been characterized (Kurata et al. 2005). During the last decade, extensive progress made in the rice genome mapping and sequencing has lend great support to map-based cloning of novel trait genes (Soda et al. 2013). Analysis of mutations is the most effective strategy to investigate gene function(s). Several mutant libraries have been produced by using T-DNA insertions, transposon or retrotransposon tagging, and chemical/irradiation mutagens in various model species (Wu et al. 2005; Kuromori et al. 2009). Gamma rays and fast neutrons can generate several allelic series containing small, as well as large (70–500 kb) deletions in genomes in a very short time (Wang et al. 2013). A twenty-one lesion mimic mutants were isolated earlier from IR64 rice mutant populations to comprehensively study disease resistance pathways (Wu et al. 2008). Taken together, these studies show the usefulness of forward genetics approach to not only obtain desirable phenotypes, but also provide a powerful tool to identify and isolate useful genes (s).
Screening under field conditions is important for identifying tolerant genotypes. However, the true expression of genotypes is hindered due to soil heterogeneity and inherent spatial variability of salt concentration under natural conditions. To avoid this limitation, specially designed microplots are crucial for screening under controlled conditions (Krishnamurthy et al. 2016). The present study, for the first time, endeavored to study the simultaneous performance of mutant lines of IR64 genotype under precisely controlled non-stress and saline conditions, so that adapted ideotypes having higher biomass and yield even under unfavorable areas could be identified and characterized. In addition, this study can generate basic information on high biomass IR64 mutants which are important for the future bioenergy related research activities.
Materials and methods
Plant materials
The high quality seeds of Oryza sativa L. cv. ‘IR64’ that demonstrated high yield traits, along with moderate salt tolerance were irradiated at Central Instrumentation Facility, School of Life Sciences, Jawaharlal Nehru University, with gamma-rays that were generated using a 60Co gamma-irradiator (240 TBq 60Co Model 4000 A). A total of 1500 seeds were irradiated in three doses i.e., 50 Gy, 75 Gy and 100 Gy to produce M1 seeds. These M1 mutant seeds and the controls were sown under field condition at ICAR-Central Soil Salinity Research Institute (CSSRI), Karnal. The M1 plants were harvested individually to obtain 1166 M2 generation mutant lines. These M2 generation lines were further multiplied under field conditions at CSSRI, Karnal to obtain M3 generation seeds. The genetically fixed mutant lines with excellent agricultural characteristics were further used to screen for salt tolerance (Fig. 1).
Schematic representation of the development and screening of IR64 mutant lines. The mutants were generated through three doses of gamma irradiation. The developed M1 population was multipled further to obtain M2 and M3 generation seeds. The genetically fixed mutant population was screened in hydroponics to standardize 50% lethal dose under Ec = 10dS/m salt condition for 4 days. The screened M3 population seeds were further screened in micro-plot/lysimeters at CSSRI, Karnal under three doses of salinity stress (EC-6 dS/m, EC-10 dS/m and EC-14 dS/m) to obtain M4 generation mutants and various physiological parameters, biomass content and yield traits were calculated. FW fresh weight, DW dry weight, FB fresh biomass, DB dry biomass
Screening for salinity tolerance in hydroponics
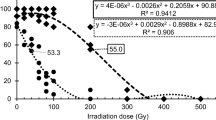
In order to determine the concentration of the 50% lethal dose (LD50) under the salt concentration, the hulled seeds of M3 generation mutant population were sterilized with 5% sodium hypochlorite and germinated in a hydroponic system for 14 days (28 ± 1 °C; 12 h day/night) in half strength Yoshida medium. Seedlings were exposed to salinity stress (EC-10 dS/m; NaCl) in half strength Yoshida medium for 4 days. After four days seedlings were recovered in half strength Yoshida medium for 7 days. Selected healthy mutant seedlings were further grown under field conditions to obtain M4 generation seeds.
Testing mutant lines in saline microplots
IR-64 is salt-sensitive during the seedling stage and moderately tolerant at maturity stage. Therefore, 18 selected salt tolerant lines (based on hydroponics) were screened for different levels of salt tolerance in micro-plots (size 6 × 3 × 1 m under rainout shelter) at CSSRI, Karnal, India (26°47′ 58″N, 80°46′ 24″E, 120 m above MSL) in the kharif season (Fig. 2). The seeds of selected 18 lines of M4 generation were initially germinated in a nursery bed under control conditions. After 20 days, the seedlings were transplanted in three replications in a randomized block design (RBD) in non-stress conditions (Singh et al. 2016b). Each ideotype was planted as a single row of 3.0 m length following row × plant spacing of 20 × 15 cm. Fertilizer rate at 120–60–40–25 kg N–P2O5–K2O–ZnSO4·7H2O ha−1 was applied uniformly. Half N and full P2O5, K2O and ZnSO4·7H2O were broadcasted as basal after puddling and the remaining N was applied in equal splits at active tillering (30 days after transplanting) and panicle initiation (60 days after transplanting) (Sharma et al. 2006). Three doses of salinity stress (EC-6 dS/m, EC-10 dS/m and EC-14 dS/m) were given after one month of transplanting and maintained. Plants were brought to maturity under these stress conditions until harvested for the analysis.
Rice mutant lines were grown in micro-plot/lysimeters at ICAR-CSSRI, Karnal in the kharif season under control and three saline regimes till maturity. A control, B low saline (EC = 6 dS/m), C moderate saline (EC = 10 dS/m), and D high saline (EC = 14 dS/m). The plants grown under normal conditions were treated as control. The experiment was designed in RBD condition in three replicates, EC electric conductivity
Physiological parameters, biomass and yield attribute determination
To evaluate the agronomic characteristics of these mutant lines for salt tolerance, the WT IR-64 plants were employed as controls. The physiological parameters i.e., plant height, tiller number per plant, total cholorophyll content, K+/Na+ ratio, proline content and electrolyte leakage were calculated as described earlier (Joshi et al. 2014; Kumar et al. 2013). Tiller count was gathered by counting the tillers including the newly emerged tillers with one fully expanded leaf. Free proline concentration was measured using 100 mg of fresh leaf tissues of non-stressed and salt stressed plants using samples from three independent replicates and expressed as µmol g−1 FW. Similarly, 100 mg young and fully expanded leaf material from WT and mutant lines under control and salt stress was used for pigment extraction in dimethyl sulfoxide and the chlorophyll concentration was measured at A663 and A645 nm. For determination of endogenous Na+ and K+ contents, 100 mg of leaf tissue (control and salinity stressed) were taken and digested in 0.1% HNO3. Ions were extracted in distilled H2O by boiling for 30 min twice and the filtrate was used to measure specific ions in flame photometer.
For biomass analysis, shoot fresh weight, shoot dry weight, root fresh weight, root dry weight, total fresh weight and total dry weight were observed (Kondhia et al. 2015). The shoot and root weights were collected by weighing the upper part of the plant and root (including the node where the upper most roots originated) after carefully removing soil. These samples were air dried for 30 days to obtain the shoot and root dry weights. To calculate yield parameters i.e. number of panicles per plant, panicle length, filled grain number per panicle, unfilled grain number per panicle, 1000 grain weight, and harvest index at maturity (for all mutant lines, and the WT IR64) were calculated (Tripathi et al. 2016).
Statistical analysis
All the collected data were subjected to an analysis of variance (ANOVA) (Gomez and Gomez 1984). The least significant difference (LSD) at 5% probability was used to compare treatment means (Steel and Torri 1980). Data was presented as means across experiments for all parameters and main effects were reported as there are no significant interactions between plant spacing and number of seedlings hill−1.
Results and discussion
Induced mutants are an important resource to study specific agronomic traits. We have carried out an initial screening of 1500 M3 mutant lines at seedling stage under 200 mM NaCl salt stress condition. Based on the germination rate, survival rate, and seedling growth, we have selected 18 salt tolerant mutant lines (data not shown). These 18 selected lines were further grown under saline conditions in microplots at CSSRI, Karnal. The present study was performed in order to investigate the physiological responses of the mutant lines under salt stress.
Physiological analysis of selected mutant lines in field under saline conditions
Rice is very salt-sensitive during the seedling and flowering stages. However, the salt tolerance in the early growth stages does not always correlate with that at maturity level. Therefore, we have screened 18 selected salt tolerant mutant lines of IR64 for salt tolerance at different levels of salinity stress throughout their development under field conditions. The physiological characteristics of all the 18 mutant lines were evaluated under saline conditions in microplots taking WT IR64 plants as control. The plant height, chlorophyll content, proline content, K+/Na+ ratio, electrolyte leakage and tiller number per plant were recorded for 18 mutant lines, and the WT IR-64 plants under control and saline conditions as given in Fig. 3. These physiological changes in salt-stressed plants are directly co-related with their overall growth (Khan and Panda 2008). A reduction in plant height was observed with the increase in salt concentration. Under control conditions, mutant lines D100-211, D100-204 and D75-16 attained more height than IR64 WT plants. In contrast, D100-200, D100-51, D50-76, D50-33 and D50-11 attained less height than IR64 WT plants. However, under saline conditions, the plant height showed reduction drastically in WT (82%) as well as all the mutant lines, but tolerant lines showed least decrease in the plant height (45–56%; Fig. 3A). Under stress conditions, mutant lines D100-211, D100-209, D100-204 and D50-92 (these ideotypes were termed as ‘tolerant’ in the following text) attain more height than IR64 WT plants. In contrast, D100-51, D50-76, D50-33 and D50-11 (these ideotypes were termed as ‘sensitive’ in the following text) attain less height than IR64 WT plants. Plant height is a critical component in rice plants to avoid lodging that may lead to lower biomass harvest. However, reduction in plant height under salinity stress may be due to osmotic stress which confines cell elongation resulting in retardation of internodal length, number of nodes and eventually plant height (Joshi et al. 2016a; Kondhia et al. 2015).
Assessment of physiological parameters of field-grown IR64 mutants. Bar graph depicting the comparison of plant height (cm) (A), total chlorophyll content calculated as µg gm−1 fresh weight (FW) of the leaves (B), proline content, calculated as µmol gm−1 fresh weight (FW) of the leaves (C), K+/Na+ ratio (D), and electrolyte leakage (%) (E) between the wild-type IR64 and eighteen mutant lines grown under non-stress, low saline (EC = 6 dS/m), moderate saline (EC = 10 dS/m), and high saline (EC = 14 dS/m) environments till maturity. Values are the mean ± standard deviation (n = 3). ‘*’ shows the promising salt tolerant mutant lines and ‘#’ shows salt sensitive mutant lines
The total chlorophyll content was found to be significantly higher in the tolerant lines (Fig. 3B). However the chlorophyll content gradually decreased in WT (94%) as well as mutant lines after the salt treatment, but the tolerant mutant lines showed least decrease (68–76%) under salinity stress in comparison to other lines. Reduction of chlorophyll content due to salinity stress is a common phenomenon in salt-sensitive plant species because of salt toxicity which mostly causes burning of leaves or other succulent parts and degradation of other pigments too (Munns et al. 2006; Stępień and Kłbus 2006). Salinity can also affect photosynthesis by decreasing intercellular CO2 availability caused by confinement of diffusion due to stomatal closure (Flexas et al. 2007; Stoeva and Kaymakanova 2008). But tolerant species can protect themselves from such deterioration of salinity stress (Alam et al. 2015). Plant responses to salt stress are also related to metabolic regulation, including proline accumulation, which may counter the osmotic stress caused by salt stress (Joshi et al. 2016b; Soda et al. 2016). In the present study, salt stress resulted in an increase in the proline concentration in leaves; however, tolerant mutant lines accumulated drastically higher levels of proline than the WT IR64 plants (Fig. 3C).
The maintenance of the ratios of Na+ and K+ is considered as a vital determinant of stress tolerance. Plants have to either sequester or extrude excess Na+ ions and thereby maintain higher K+ ions in order to survive under higher concentrations of NaCl (Zhang et al. 2008; Kumari et al. 2015). Estimation of K+ and Na+ in leaves of WT IR64 and mutant lines indicated that WT IR64 plants were able to maintain its ratio under control conditions indicating their unstressed physiological status. However, tolerant mutant lines showed much higher K+/Na+ ratio under control as well as stress conditions (Fig. 3D). In contrast, sensitive mutant lines showed significantly lower K+/Na+ ratio in comparison to IR64 WT under control as well as stress conditions. Electrolyte leakage is considered to be an indicator of cell membrane stability and integrity under abiotic stress conditions (Kumari et al. 2015). The electrolyte leakage of all the mutant lines and the IR64 WT plants were evaluated in order to study the degree of membrane injury caused by salt stress. All the tolerant mutant lines showed less decrease in electrolyte leakage under salt stress conditions (17–22%) as compared to IR64 WT plants (Fig. 3E) indicating higher cellular damage in WT IR64 plants under high salinity levels.
Evaluation of biomass in selected mutant lines in field under saline conditions
Dry matter content is a functional parameter used to assess the plant strategy for resource acquisition (Duru et al. 2009). The interaction of genotypes with environment is significant for plant biomass determination indicating that genotypes vary significantly depending upon the severity of stress conditions (Tripathi et al. 2016). It has already been well established that taller plant should have strong tillers to produce higher biomass besides minimize lodging. The mutant lines grown under control conditions had significantly faster tiller emergence and the rate of increase in plant height than those grown under saline conditions. In all the mutant lines and IR64 WT, the tiller number per plant significantly decreased under salinity stress (Fig. 4A). It was reported earlier that plant height and tiller number per plant were reduced under salinity stress conditions because plants are unable to absorb soil water resulting in less availability of essential elements (Bunnag and Pongthai 2013). Both tiller number per plant and spikelet number per panicle were most significantly affected by salinity stress (Joshi et al. 2016a). Moreover, salinity, during tillering, reduced shoot dry weight twice as much as that during mature stages under field conditions (Zeng et al. 2001). The fresh and dry weights of shoot, root and total biomass were significantly higher in all the mutant lines under control conditions than those obtained under saline conditions (Fig. 4B–E). Among the 18 mutant lines selected, tolerant lines were shown to have the highest fresh and dry biomass under control conditions, whereas, sensitive lines have the lowest fresh and dry biomass under control conditions (Fig. 4F, G). The tolerant lines produced 37–63% higher biomass under control conditions, where as sensitive lines showed 26–36% less biomass under control conditions. Similarly, tolerant mutant lines were shown to have the highest fresh and dry biomass under saline conditions, whereas, sensitive lines have the lowest fresh and dry biomass under saline conditions (Fig. 4F, G). Under severe stress the tolerant lines showed 53–69% decline in biomass in comparison to control conditions, where as sensitive lines showed 66–97% decline in total biomass in comparison to control conditions. The decrease in biomass under salinity stress is either because of limited assimilates produced due to inhibition of photosynthesis (Joshi et al. 2016b) or it might be due to the inhibition in hydrolysis of reserved foods and their translocation to the growing shoots (Alam et al. 2015).
Evaluation of biomass of field-grown IR64 mutants. Bar graph depicting the means of A tiller number per plant, B shoot fresh weight, C shoot dry weight, D root fresh weight, E root dry weight, F total fresh biomass, and G total dry biomass across IR64 WT and 18 mutant lines grown under non-stress, low saline (EC = 6 dS/m), moderate saline (EC = 10 dS/m), or high saline (EC = 14 dS/m) environments till maturity. Secondary axis of the graph shows % decrease in the respective parameter under salinity stress. Values are the mean ± standard deviation (n = 3). ‘*’ shows the promising salt tolerant mutant lines and ‘#’ shows salt sensitive mutant lines
Yield analysis of selected mutant lines in field under saline conditions
Soil salinity is a major environmental constraint for crop productivity (Joshi et al. 2016a). Higher productivity and grain weight under stress and control conditions are considered as qualitative trait for crop improvement. QTL mapping and genetic engineering significantly contributed in identification of genes for yield enhancement (Huang et al. 2013; Li et al. 2013). Rice is a global food crop as well as model cereal along with an excellent system for studying stress signaling pathways for improved growth and yield under saline conditions (Joshi et al. 2014). In our study, number of panicles per plant substantially decreased in IR64 WT and all the mutant lines under salinity stress (Fig. 5A). The tolerant lines produced 16–39% more panicle under saline conditions than IR64, where as sensitive lines showed 44–58% reduction in panicle number under saline conditions. However, panicle length was found to be similar in IR64 WT and mutant lines, but it also reduces significantly under salinity stress (Fig. 5B). The tolerant lines produced 8–35% longer panicles under saline conditions than IR64, where as sensitive lines showed 18% shorter panicles under similar conditions. In addition, filled grain number per panicle was also decreased significantly under salinity stress in IR64 WT (48%) as well as mutant lines i.e., 27–30% in tolerant lines whereas, up to 62% in sensitive lines (Fig. 5C). However, unfilled grain number per panicle increased significantly under salt stress in comparison to the control (Fig. 5D). In contrast, 1000 grain weight of the IR64 WT and mutant lines does not show any significant variation under salinity stress as well as in control conditions when compared to IR64 WT plants (Fig. 5E). The tolerant lines showed 2–7% higher harvest index under saline conditions (Fig. 5F). The grain yield or economic yield is referred to as a complex character resulting from multiplicative interactions of several yield contributing components. Sink capacity is the capability of plants to transport and utilize photosynthates for growth and biomass production (Ainsworth et al. 2004). Under salinity stress, biological yield and harvest index showed significant positive association with grain yield (Krishnamurthy et al. 2016). Across all salt stress environments, it was observed that biological yield and harvest index were generally important traits for salinity conditions. In the present analysis both high yielding tolerant mutants and low yielding sensitive mutants were scored (Fig. 5G). Two very promising mutants viz. D100-211 and D100-209 were found to show an average of 18% and 34% increase in yield respectively over WT plants under moderate saline conditions (EC = 10 dS/m). These results clearly indicate that among all the traits recorded, taller plants with profuse tillering and more productive tillers and higher grain yield coupled with more harvest index appeared to be potential traits for the selection in non-stress and salt stress conditions to identify salt tolerant mutant lines. Similarly, several other screening methods have been designed to identify genotypes with high resilience and productivity under various abiotic stresses (Thiry et al. 2016).
Yield analysis of field-grown IR64 mutants. Web diagram depicting the means of A Panicle number per plant, B panicle length, C filled grain number per panicle, D unfilled grain number per panicle, E thousand grain weight, F harvest index and G bar diagram depicting % change in yield per plant across IR64 WT and 18 mutant lines grown under non-stress, low saline (EC = 6 dS/m), moderate saline (EC = 10 dS/m), or high saline (EC = 14 dS/m) environments till maturity
In the present study, it is likely that the WT plant did not perform well under severe salt stress. Few reports are available on selection of stable mutant populations for salt tolerance at different growth stages and trait correlation analysis. Our study indicates that selection based on salt stress indices is likely to identify mutant ideotypes with better performance across varying salt stress environments. Based on the overall analysis, the tolerant lines (i.e., D100-211, D100-209, D100-204 and D50-92) have been found to be promising high-yielding and high-biomass containing mutant salt-tolerant lines for a range of salt stress conditions, molecular basis of which is under investigation.
References
Ahmed, S., Mahmood, A., Hasan, A., Sidhu, G. A., & Butt, M. F. (2016). A comparative review of China, India and Pakistan renewable energy sectors and sharing opportunities. Renewable and Sustainable Energy Reviews, 57, 216–225.
Ainsworth, E. A., Rogers, A., Nelson, R., & Long, S. P. (2004). Testing the “source sink” hypothesis of down-regulation of photosynthesis in elevated [CO2] in the field with single gene substitutions in Glycine max. Agricultural Meteorology, 122, 85–94.
Alam, M. A., Juraimi, A. S., Rafii, M. Y., & Abdul Hamid, A. (2015). Effect of salinity on biomass yield and physiological and stem-root anatomical characteristics of purslane (Portulaca oleracea L.) accessions. BioMed research international, 23, 2015.
Bunnag, S., & Pongthai, P. (2013). Selection of rice (Oryza sativa L.) cultivars tolerant to drought stress at the vegetative stage under field conditions. American. Journal of Plant Sciences, 4, 1701.
Chandra, R., Takeuchi, H., & Hasegawa, T. (2012). Methane production from lignocellulosic agricultural crop wastes: A review in context to second generation of biofuel production. Renewable and Sustainable Energy Reviews, 16, 1462–1476.
Duru, M., Khaled, R. A., Ducourtieux, C., Theau, J. P., de Quadros, F. L., & Cruz, P. (2009). Do plant functional types based on leaf dry matter content allow characterizing native grass species and grasslands for herbage growth pattern? Plant Ecology, 201, 421–433.
Flexas, J., Diaz-Espejo, A. N., Galmes, J., Kaldenhoff, R., Medrano, H., & Ribas-Carbo, M. I. (2007). Rapid variations of mesophyll conductance in response to changes in CO2 concentration around leaves. Plant, Cell and Environment, 30, 1284–1298.
Gomez, K. A., & Gomez, A. A. (1984). Statistical Procedures for Agricultural Research (2nd ed.). New York: John Wiley and Sons.
Huang, R., Jiang, L., Zheng, J., Wang, T., Wang, H., Huang, Y., et al. (2013). Genetic bases of rice grain shape: so many genes, so little known. Trends Plant Science, 18, 218–226.
Joshi, R., Karan, R., Singla-Pareek, S. L., & Pareek, A. (2016a). Ectopic expression of Pokkali phosphoglycerate kinase-2 (OsPGK2-P) improves yield in tobacco plants under salinity stress. Plant Cell Reports, 35, 27–41.
Joshi, R., Ramanarao, M. V., Lee, S., Kato, N., & Baisakh, N. (2014). Ectopic expression of ADP ribosylation factor 1 (SaARF1) from smooth cordgrass (Spartina alterniflora Loisel) confers drought and salt tolerance in transgenic rice and Arabidopsis. Plant Cell, Tissue and Organ Culture, 117, 17–30.
Joshi, R., Singh, B., Bohra, A., & Chinnusamy, V. (2016b). Salt stress signalling pathways: specificity and crosstalk (pp. 51–78). Managing Salinity Tolerance in Plants: Molecular and Genomic Perspectives.
Khan, M. H., & Panda, S. K. (2008). Alterations in root lipid peroxidation and antioxidative responses in two rice cultivars under NaCl-salinity stress. Acta Physiologiae Plantarum, 30, 81–89.
Kondhia, A., Tabien, R. E., & Ibrahim, A. (2015). Evaluation and selection of high biomass rice (Oryza sativa L.) for drought tolerance. American. Journal of Plant Sciences, 6, 1962.
Krishnamurthy, S. L., Gautam, R. K., Sharma, P. C., & Sharma, D. K. (2016). Effect of different salt stresses on agro-morphological traits and utilisation of salt stress indices for reproductive stage salt tolerance in rice. Field Crops Research, 190, 26–33.
Kumar, M., Choi, J. Y., Kumari, N., Pareek, A., & Kim, S. R. (2015). Molecular breeding in Brassica for salt tolerance: importance of microsatellite (SSR) markers for molecular breeding in Brassica. Frontiers in Plant Science, 6, 688.
Kumar, P., Pal, M., Joshi, R., & Sairam, R. K. (2013). Yield, growth and physiological responses of mung bean [Vigna radiata (L.) Wilczek] genotypes to waterlogging at vegetative stage. Physiology and Molecular Biology of Plants, 19, 209–220.
Kumari, S., Joshi, R., Singh, K., Roy, S., Tripathi, A. K., Singh, P., et al. (2015). Expression of a cyclophilin OsCyp2-P isolated from a salt-tolerant landrace of rice in tobacco alleviates stress via ion homeostasis and limiting ROS accumulation. Functional & Integrative Genomics, 15, 395–412.
Kurata, N., Miyoshi, K., Nonomura, K. I., Yamazaki, Y., & Ito, Y. (2005). Rice mutants and genes related to organ development, morphogenesis and physiological traits. Plant and Cell Physiology, 46, 48–62.
Kuromori, T., Takahashi, S., Kondou, Y., Shinozaki, K., & Matsui, M. (2009). Phenome analysis in plant species using loss-of-function and gain-of-function mutants. Plant and Cell Physiology, 50, 1215–1231.
Kushwaha, H. R., Singla-Pareek, S. L., Sopory, S. K., Pareek, A., Tuteja, N., Gill, S. S., et al. (2012). Understanding the “commoneome” operative in plants in response to various abiotic stresses. Improving Crop Resistance to Abiotic Stress, 1 & 2, 3–25.
Li, S., Zhao, B., Yuan, D., Duan, M., Qian, Q., Tang, L., et al. (2013). Rice zinc finger protein DST enhances grain production through controlling Gn1a/OsCKX2 expression. Proceedings of the National Academy of Sciences, USA, 110, 3167–3172.
Mazzola, S., Astolfi, M., & Macchi, E. (2016). The potential role of solid biomass for rural electrification: A techno economic analysis for a hybrid microgrid in India. Applied Energy, 169, 370–383.
Munns, R., James, R. A., & Läuchli, A. (2006). Approaches to increasing the salt tolerance of wheat and other cereals. Journal of Experimental Botany, 57, 1025–1043.
Sharma, R. C., Singh, R., Singh, Y. P., & Singh, G. (2006). Sodic soils of Shivri experimental farm: site characterization, reclamability and use potential for different land uses. CSSRI Technical Bulletins, 1, 36.
Singh, R., Srivastava, M., & Shukla, A. (2016a). Environmental sustainability of bioethanol production from rice straw in India: A review. Renewable and Sustainable Energy Reviews, 54, 202–216.
Singh, Y. P., Mishra, V. K., Singh, S., Sharma, D. K., Singh, D., Singh, U. S., et al. (2016b). Productivity of sodic soils can be enhanced through the use of salt tolerant rice varieties and proper agronomic practices. Field Crops Research, 190, 82–90.
Soda, N., Kushwaha, H. R., Soni, P., Singla-Pareek, S. L., & Pareek, A. (2013). A suite of new genes defining salinity stress tolerance in seedlings of contrasting rice genotypes. Functional & Integrative Genomics, 13, 351–365.
Soda, N., Sharan, A., Gupta, B. K., Singla-Pareek, S. L., & Pareek, A. (2016). Evidence for nuclear interaction of a cytoskeleton protein (OsIFL) with metallothionein and its role in salinity stress tolerance. Scientific Reports, 6, 34762.
Song, J. Y., Kim, D. S., Lee, M. C., Lee, K. J., Kim, J. B., Kim, S. H., et al. (2012). Physiological characterization of gamma-ray induced salt tolerant rice mutants. Australian Journal of Crop Science, 6, 421.
Soni, P., Kumar, G., Soda, N., Singla-Pareek, S. L., & Pareek, A. (2013). Salt overly sensitive pathway members are influenced by diurnal rhythm in rice. Plant Signaling & Behavior, 8, e24738.
Steel, R. G. D., & Torri, J. H. (1980). Principal and Procedures of Statistics. New York: McGraw HillBook Co.Inc.
Stępień, P., & Kłbus, G. (2006). Water relations and photosynthesis in Cucumis sativus L. leaves under salt stress. Biologia Plantarum, 50, 610–616.
Stoeva, N., & Kaymakanova, M. (2008). Effect of salt stress on the growth and photosynthesis rate of bean plants (Phaseolus vulgaris L.). Journal of Central European Agriculture, 9, 385–392.
Thiry, A. A., Chavez Dulanto, P. N., Reynolds, M. P., & Davies, W. J. (2016). How can we improve crop genotypes to increase stress resilience and productivity in a future climate? A new crop screening method based on productivity and resistance to abiotic stress. Journal of Experimental Botany, 16, 5593–5603.
Tripathi, A. K., Pareek, A., & Singla-Pareek, S. L. (2016). A NAP-Family histone chaperone functions in abiotic stress response and adaptation. Plant Physiology, 171, 2854–2868.
Ullah, K., Sharma, V. K., Dhingra, S., Braccio, G., Ahmad, M., & Sofia, S. (2015). Assessing the lignocellulosic biomass resources potential in developing countries: A critical review. Renewable and Sustainable Energy Reviews, 51, 682–698.
Wang, N., Long, T., Yao, W., Xiong, L., Zhang, Q., & Wu, C. (2013). Mutant resources for the functional analysis of the rice genome. Molecular Plant, 6, 596–604.
Wu, C., Bordeos, A., Madamba, M. R., Baraoidan, M., Ramos, M., Wang, G. L., et al. (2008). Rice lesion mimic mutants with enhanced resistance to diseases. Molecular Genetics and Genomics, 279, 605–619.
Wu, J. L., Wu, C., Lei, C., Baraoidan, M., Bordeos, A., Madamba, M. R., et al. (2005). Chemical-and irradiation-induced mutants of indica rice IR64 for forward and reverse genetics. Plant Molecular Biology, 59, 85–97.
Zeng, L., Shannon, M. C., & Lesch, S. M. (2001). Timing of salinity stress affects rice growth and yield components. Agricultural Water Management, 48, 191–206.
Zhang, X., Wollenweber, B., Jiang, D., Liu, F., & Zhao, J. (2008). Water deficits and heat shock effectson photosynthesis of a transgenic Arabidopsis thaliana constitutively expressingABP9, a bZIP transcription factor. Journal of Experimental Botany, 59, 839–848.
Zheng, H., Zhao, H., Liu, H., Wang, J., & Zou, D. (2015). QTL analysis of Na+ and K+ concentrations in shoots and roots under NaCl stress based on linkage and association analysis in japonica rice. Euphytica, 201(1), 109–121.
Acknowledgements
RJ acknowledges Dr D S Kothari Postdoctoral Fellowship from University Grant Commission, Government of India. SLS-P would like to acknowledge funds received from Council of Scientific and Industrial Research (CSIR), Government of India. AP would like to acknowledge the financial support received from Jawaharlal Nehru University, JNU (India), International Atomic Energy Agency (Vienna) and Indo US Science and Technology Forum (IUSSTF), New, Delhi.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Joshi, R., Rama Prashat, Sharma, P.C. et al. Physiological characterization of gamma-ray induced mutant population of rice to facilitate biomass and yield improvement under salinity stress. Ind J Plant Physiol. 21, 545–555 (2016). https://doi.org/10.1007/s40502-016-0264-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40502-016-0264-x