Abstract
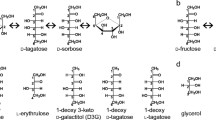
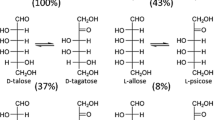
d-Psicose 3-epimerase (DPEase) is demonstrated to be useful in the bioproduction of d-psicose, a rare hexose sugar, from d-fructose, found plenty in nature. Clostridium cellulolyticum H10 has recently been identified as a DPEase that can epimerize d-fructose to yield d-psicose with a much higher conversion rate when compared with the conventionally used DTEase. In this study, the crystal structure of the C. cellulolyticum DPEase was determined. The enzyme assembles into a tetramer and each subunit shows a (β/α)8 TIM barrel fold with a Mn2+ metal ion in the active site. Additional crystal structures of the enzyme in complex with substrates/products (d-psicose, d-fructose, d-tagatose and d-sorbose) were also determined. From the complex structures of C. cellulolyticum DPEase with d-psicose and d-fructose, the enzyme has much more interactions with d-psicose than d-fructose by forming more hydrogen bonds between the substrate and the active site residues. Accordingly, based on these ketohexose-bound complex structures, a C3-O3 proton-exchange mechanism for the conversion between d-psicose and d-fructose is proposed here. These results provide a clear idea for the deprotonation/protonation roles of E150 and E244 in catalysis.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Brünger, A.T., Adams, P.D., Clore, G.M., DeLano, W.L., Gros, P., Grosse-Kunstleve, R.W., Jiang, J.-S., Kuszewski, J., Nilges, M., Pannu, N.S., et al. (1998). Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr D Biol Crystallogr 54, 905–921.
Granström, T.B., Takata, G., Tokuda, M., and Izumori, K. (2004). Izumoring: a novel and complete strategy for bioproduction of rare sugars. J Biosci Bioeng 97, 89–94.
Itoh, H., Khan, A.R., Tajima, S., Hayakawa, S., and Izumori, K. (1994). Purification and characterization of d-tagatose 3-epimerase from Pseudomonas sp.ST-24. Biosci Biotechnol Biochem 57, 1037–1039.
Izumori, K. (1995). Preparation of d-psicose from d-fructose by immobilized d-tagagtose-3-epimerase. J Ferment Bioeng 80, 101–103.
Izumori, K. (2002). Bioproduction strategies for rare hexose sugars. Naturwissenschaften 89, 120–124.
Kim, H.-J., Hyun, E.-K., Kim, Y.-S., Lee, Y.-J., and Oh, D.-K. (2006a). Characterization of an Agrobacterium tumefaciens d-psicose 3-epimerase that converts d-fructose to d-psicose. Appl Environ Microbiol 72, 981–985.
Kim, H.-J., Lim, B.-C., Yeom, S.-J., Kim, Y.-S., Kim, D., and Oh, D.-K. (2010a). Roles of Ile66 and Ala107 of d-psicose 3-epimerase from Agrobacterium tumefaciens in binding O6 of its substrate, d-fructose. Biotechnol Lett 32, 113–118.
Kim, H.-J., Yeom, S.-J., Kim, K., Rhee, S., Kim, D., and Oh, D.-K. (2010b). Mutational analysis of the active site residues of a D: -psicose 3-epimerase from Agrobacterium tumefaciens. Biotechnol Lett 32, 261–268.
Kim, K., Kim, H.-J., Oh, D.-K., Cha, S.-S., and Rhee, S. (2006b). Crystal structure of d-psicose 3-epimerase from Agrobacterium tumefaciens and its complex with true substrate d-fructose: a pivotal role of metal in catalysis, an active site for the non-phosphorylated substrate, and its conformational changes. J Mol Biol 361, 920–931.
Kim, N.-H., Kim, H.-J., Kang, D.-I., Jeong, K.-W., Lee, J.-K., Kim, Y., and Oh, D.-K. (2008). Conversion shift of d-fructose to d-psicose for enzyme-catalyzed epimerization by addition of borate. Appl Environ Microbiol 74, 3008–3013.
Matsuo, T., Baba, Y., Hashiguchi, M., Takeshita, K., Izumori, K., and Suzuki, H. (2001). Dietary d-psicose, a C-3 epimer of d-fructose, suppresses the activity of hepatic lipogenic enzymes in rats. Asia Pac J Clin Nutr 10, 233–237.
Matsuo, T., and Izumori, K. (2006). Effects of dietary d-psicose on diurnal variation in plasma glucose and insulin concentrations of rats. Biosci Biotechnol Biochem 70, 2081–2085.
Matsuo, T., Suzuki, H., Hashiguchi, M., and Izumori, K. (2002). d-psicose is a rare sugar that provides no energy to growing rats. J Nutr Sci Vitaminol (Tokyo) 48, 77–80.
McCoy, A.J., Grosse-Kunstleve, R.W., Adams, P.D., Winn, M.D., Storoni, L.C., and Read, R.J. (2007). Phaser crystallographic software. J Appl Crystallogr 40, 658–674.
McRee, D.E. (2004). Differential evolution for protein crystallographic optimizations. Acta Crystallogr D Biol Crystallogr 60, 2276–2279.
Mu, W., Chu, F., Xing, Q., Yu, S., Zhou, L., and Jiang, B. (2011). Cloning, expression, and characterization of a d-psicose 3-epimerase from Clostridium cellulolyticum H10. J Agric Food Chem 59, 7785–7792.
Oh, D.-K. (2007). Tagatose: properties, applications, and biotechnological processes. Appl Microbiol Biotechnol 76, 1–8.
Otwinowski, Z., and Minor, W. (1997). Processing of X-ray diffraction data collected in oscillation mode. In: Methods in enzymology, macromolecular crystallography, part A. Carter C.W., Jr. and Sweet R. M. eds. New York: Academic Press, 307–326.
Emsley, P., and Cowtan, C. (2004). Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60, 2126–2132.
Potterton, E., Briggs, P., Turkenburg, M., and Dodson, E. (2003). A graphical user interface to the CCP4 program suite. Acta Crystallogr D Biol Crystallogr 59, 1131–1137.
Samuel, J., and Tanner, M.E. (2002). Mechanistic aspects of enzymatic carbohydrate epimerization. Nat Prod Rep 19, 261–277.
Yoshida, H., Yamada, M., Nishitani, T., Takada, G., Izumori, K., and Kamitori, S. (2007). Crystal structures of d-tagatose 3-epimerase from Pseudomonas cichorii and its complexes with d-tagatose and d-fructose. J Mol Biol 374, 443–453.
Zhang, L., Mu, W., Jiang, B., and Zhang, T. (2009). Characterization of d-tagatose-3-epimerase from Rhodobacter sphaeroides that converts d-fructose into d-psicose. Biotechnol Lett 31, 857–862.
Author information
Authors and Affiliations
Corresponding authors
Additional information
These authors contributed equally to the work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Chan, HC., Zhu, Y., Hu, Y. et al. Crystal structures of d-psicose 3-epimerase from Clostridium cellulolyticum H10 and its complex with ketohexose sugars. Protein Cell 3, 123–131 (2012). https://doi.org/10.1007/s13238-012-2026-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13238-012-2026-5