Abstract
Pseudomonas cichorii D-tagatose 3-epimerase (PcDTE), which has a broad substrate specificity, efficiently catalyzes the epimerization of not only D-tagatose to D-sorbose but also D-fructose to D-psicose (D-allulose) and also recognizes the deoxy sugars as substrates. In an attempt to elucidate the substrate recognition and catalytic reaction mechanisms of PcDTE for deoxy sugars, the X-ray structures of the PcDTE mutant form with the replacement of Cys66 by Ser (PcDTE_C66S) in complexes with deoxy sugars were determined. These X-ray structures showed that substrate recognition by the enzyme at the 1-, 2-, and 3-positions is responsible for enzymatic activity and that substrate-enzyme interactions at the 4-, 5-, and 6-positions are not essential for the catalytic reaction of the enzyme leading to the broad substrate specificity of PcDTE. They also showed that the epimerization site of 1-deoxy 3-keto D-galactitol is shifted from C3 to C4 and that 1-deoxy sugars may bind to the catalytic site in the inhibitor-binding mode. The hydrophobic groove that acts as an accessible surface for substrate binding is formed through the dimerization of PcDTE. In PcDTE_C66S/deoxy sugar complex structures, bound ligand molecules in both the linear and ring forms were detected in the hydrophobic groove, while bound ligand molecules in the catalytic site were in the linear form. This result suggests that the sugar-ring opening of a substrate may occur in the hydrophobic groove and also that the narrow channel of the passageway to the catalytic site allows a substrate in the linear form to pass through.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
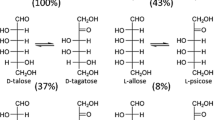
D-Tagatose 3-epimerase (DTE) catalyzes epimerization between D-tagatose and D-sorbose (Fig. 1a), and DTE family enzymes are widely found in various microorganisms (Izumori et al. 1993; Kim et al. 2006a; Zhang et al. 2009; Mu et al. 2011, 2013; Zhu et al. 2012; Zhang et al. 2013a, b, Zhang et al. 2015; Jia et al. 2014). DTE from Pseudomonas cichorii (PcDTE), which was the first to be purified and characterized among DTE family enzymes, has a broad substrate specificity and efficiently catalyzes the epimerization of not only D-tagatose to D-sorbose but also D-fructose to D-psicose (D-allulose) (Fig. 1b) (Izumori et al. 1993; Itoh et al. 1994, 1995). PcDTE is used industrially to produce D-psicose from the more abundant sugar D-fructose. Hexoses have four (aldose) and three (ketose) chiral centers, giving 16 and eight stereoisomers, respectively, and most of these exist in small amounts in nature as rare sugars. D-Psicose is one of these rare sugars and has attracted much attention due to its physiological functions, particularly its ability to promote human health (Matsuo et al. 2002; Matsuo 2006; Iida et al. 2008; Matsuo and Izumori 2009; Hayashi et al. 2010; Hossain et al. 2011, 2012; Mu et al. 2012; Iida et al. 2013). A low-calorie syrup containing D-psicose has recently become commercially available as a functional food, Rare Sugar Sweet® (Matsutani Chemical Industry Co., Ltd., Hyogo, Japan) (Hayashi et al. 2014). D-Psicose is also generally considered as safe (GRN No. 400) by the US Food and Drug Administration (FDA) and has been categorized as one of the Food Ingredient & Packaging Inventories (Mu et al. 2012). Therefore, PcDTE has become an industrially valuable enzyme.
Chemical reaction catalyzed by PcDTE and ligand molecules in this study. a The epimerization reaction between D-tagatose and D-sorbose is shown. b The epimerization reaction between D-fructose and D-psicose is shown. c The chemical structures of deoxy sugars in this study and L-erythrulose are shown. d The chemical structures of polyalcohols in this study are shown
We previously reported the X-ray structures of PcDTE in complexes with D-tagatose and D-fructose, and proposed a C3–O3 proton-exchange mechanism via a cis-enediolate intermediate as the catalytic reaction mechanism used by PcDTE (Yoshida et al. 2007). In these X-ray structures, the hydroxyl groups (O1 and O3) and ketone group (O2) at the 1-, 2-, and 3-positions of the substrate were strictly recognized by the enzyme with a large number of interactions, while the enzyme-substrate interactions at the 4-, 5-, and 6- positions were unclear due to the poor electron density for this region of the substrate (Yoshida et al. 2007). Since Cys66 may have formed a weak hydrogen bond with O6 of the substrate, we prepared a PcDTE mutant form with the replacement of Cys66 by Ser (PcDTE_C66S), in order to enhance hydrophilic interactions at the 4-, 5-, and 6- positions of the substrate. Unexpectedly, PcDTE_C66S did not provide the improved electron density for the bound D-tagatose molecules (unpublished results); however, crystals of PcDTE_C66S diffracted to a higher resolution (1.59 Å∼) than wild-type PcDTE crystals (1.79 Å∼), without significant differences in their structures.
We subsequently noted that three hydrophobic residues (Trp15, Leu108, and Trp113) were proximal to the 4-, 5-, and 6-positions of the substrate and that the hydrophobic deoxy sugars may be easily recognized in a hydrophobic environment by these hydrophobic residues. This was also supported by the findings of Gullapalli et al. who showed that PcDTE had the ability to recognize various deoxy sugars, including deoxy rare sugars, as substrates (Gullapalli et al. 2010). Although there are few reports of the physiological functions of deoxy rare sugars, they are expected to have potential as bioactive compounds, as 2-deoxy-L-ribose was reported to suppress tumor growth (Nakajima et al. 2004). The production of various deoxy sugars is desired to find their physiological activities. Several synthesis methods for deoxy rare sugars have been developed (Shompoosang et al. 2014, 2016), and PcDTE also has potential as key enzyme for the bioproduction of these sugars (Gullapalli et al. 2010). It is very important to understand the substrate recognition and catalytic reaction mechanisms used by PcDTE for deoxy sugars based on three-dimensional structures. In order to obtain crystals of the enzyme/deoxy sugar complexes, PcDTE_C66S was used, because crystals of PcDTE_C66S/deoxy sugar complexes were expected to diffract well than those of wild-type PcDTE. We herein report the X-ray structures of PcDTE_C66S in complexes with four deoxy rare sugars (6-deoxy L-psicose, 1-deoxy 3-keto D-galactitol, 1-deoxy D-tagatose, and 1-deoxy L-tagatose), and L-erythrulose without groups at the 5- and 6-positions, with enzymatic activity measurements (Fig. 1c). An unusual inhibitor-binding mode was found in some of these structures, which suggested that the polyalcohol binds to the enzyme as an inhibitor; therefore, the X-ray structures of the PcDTE_C66S complexes with two polyalcohols (glycerol and D-talitol) were also determined (Fig. 1d).
Materials and methods
Protein preparation
The expression vector of His-tagged PcDTE was constructed by subcloning. The DNA fragment containing the structural gene of PcDTE digested with NcoI and BamHI from pIK-01 (Ishida et al. 1997) was inserted into the NcoI/BamHI sites of the plasmid pQE60 (Qiagen, Hilden, Germany). The resulting plasmid was named pQE60DTE and the recombinant protein was produced as C-terminal His-tagged PcDTE. PcDTE_C66S was constructed by site-directed mutagenesis using the Quick Change II Site-Directed Mutagenesis Kit (Agilent Technology, CA, USA), the plasmid pQE60DTE, and the primers 5′-ggtgatgtgcTCTatcggactgaag-3′ (sense, mutation underlined) and 5′-cttcagtccgatAGAgcacatcacc-3′ (antisense). Recombinant C-terminal His-tagged PcDTE_C66S was expressed in Escherichia coli JM109, and cells were grown at 37 °C in 2 × YT medium containing 100 μg/ml ampicillin until the culture reached an optical density of 0.4–0.5 at 600 nm. After the addition of 0.3 mM β-D-isopropylthiogalactopyranoside and 1 mM MnCl2, cells were cultivated at 25 °C overnight. Cells were then harvested and sonicated in buffer solution (50 mM NaH2PO4, and 300 mM NaCl, pH 8.0). The sonicated sample was centrifuged (20,400×g at 4 °C for 30 min), and the resultant cell-free extract was applied to a HisTrap™ HP column 5 ml (GE Healthcare Bio-Sciences Corp., Piscataway, NJ, USA) equilibrated with buffer solution (50 mM NaH2PO4, and 300 mM NaCl, pH 8.0). After washing with buffer solution (50 mM NaH2PO4, 300 mM NaCl, and 20 mM imidazole pH 8.0), the protein was eluted with a 20–450 mM imidazole gradient. After dialyzing the purified protein with buffer solution (5 mM Tris-HCl pH 8.0), the protein solution was concentrated to 6–7 mg/ml using an Amicon Ultra-4 10 kDa Ultracel (Millipore, Billerica, MA, USA).
Enzyme assay
In order to remove bound metal ions from the enzyme, the purified enzyme was dialyzed two times against buffer containing 10 mM EDTA and 20 mM glycine-NaOH, pH 9.0, and then dialyzed three times against the same buffer without EDTA. The EDTA-treated enzyme was incubated with the same volume of substrate solution (50 mM substrate and 50 mM Tris-HCl, pH 9.0) including 1 mM CoCl2 at 60 °C for 30 min, followed by a heat treatment at 100 °C for 5 min. The reaction sample was assayed by a HPLC system using a Shimadzu RID-6A refractive index detector (Kyoto, Japan) and Hitachi HPLC column GL-C611 (Tokyo, Japan). Chromatography was carried out at 60 °C using 2 mM NaOH solution at a flow rate of 1.0 ml/min, and the ratio of the substrate and product after epimerization was quantitated by integration of the peak area in the HPLC analysis (Supplemental Fig. S1). The products were identified by an NMR spectrophotometer (ECA-600 spectrometer, FEOL, Tokyo) and/or polarimetry measurements using a Jasco R1030 polarimeter (Na lamp; Jasco, Tokyo). One unit of enzymatic activity was defined as the amount of the enzyme that catalyzes the formation of 1 μmol of the product per minute (Itoh et al. 1994).
X-ray crystallography
Using the hanging drop vapor diffusion method, crystals of PcDTE_C66S were grown in a droplet containing 2 μl of protein solution (6–7 mg/ml in 5 mM Tris-HCl, pH 8.0) and 2 μl of reservoir solution (6.0–11.0 % (w/v) PEG 4000 and 100 mM CH3COONa, pH 4.6) against 450 μl of reservoir solution. We also attempted to obtain crystals of PcDTE_C66S grown in microgravity by utilizing the crystallization facilities of the JAPAN Aerospace Exploration Agency (JAXA) on board the Japanese Experiment Module Kibo in the International Space Station (ISS) under a high-quality protein crystal growth project (JAXA-PCG). In order to obtain crystals grown in a microgravity environment in the ISS, crystallization was established using a counter-diffusion method (Takahashi et al. 2013) in solution containing 15 % (w/v) PEG 4000 and 100 mM CH3COONa, pH 4.6.
A crystal of PcDTE_C66S mounted in a cryoloop was soaked in solution containing 25–50 % (w/v) ligand molecules and then flash-cooled in a gas-stream of liquid nitrogen at 100 K. X-ray diffraction data were collected using the ADSC Quantum 315r, 210r, and 270 CCD detector system on the PF BL5A, PF-AR NW12A, NE3A beam lines in the KEK (Tsukuba, Japan), and a Rigaku R-AXIS VII imaging system on a Rigaku RA-Micro7HF rotating anode (CuKα) X-ray generator with ValiMax optics (40 kV, 30 mA). Diffraction data were processed using HKL2000 (Otwinowski and Minor 1997), the CrystalClear system (Rigaku Corp. Tokyo, Japan), and CCP4 program suite (Winn et al. 2011). The initial phases of PcDTE_C66S were obtained by molecular replacement using the program MOLREP (Vagin and Teplyakov 1997) with the structure of PcDTE (PDB code: 1QUL) as a probe model. The following model building for protein and ligand molecules was performed with the programs Coot (Emsley et al. 2010) and X-fit (McRee 1999). The structure was refined using the programs Refmac5 (Murshudov et al. 1997) and CNS (Brunger 1993), and water molecules were introduced if peaks greater than 3.0 σ in the (F o − F c) electron density map were in the range of a hydrogen bond. Structure validation was performed using the program PROCHECK (Laskowski et al. 1992). Data collection and refinement statistics are listed in Table 1. Figures 2, 3, and 4 were drawn using the program PyMol (Schrödinger, LLC, NY, USA).
Overall structure of PcDTE_C66S. a The structure of a subunit of PcDTE_C66S is shown with the bound metal ion (blue sphere) and the amino acid residues involved in the metal coordination. Twelve α-helices (H1–H12) and eight β-strands (B1–B8) are labelled. The replaced Ser66 is indicated by magenta. b The surface model of four molecules (Mol-A, Mol-B, Mol-C, and Mol-D) in an asymmetric unit is shown. c Ribbon and surface models of the dimer (Mol-A and Mol-B) are shown. The hydrophobic residues forming the hydrophobic groove at the dimer interface are indicated by gray
Structure of the catalytic site of PcDTE_C66S. a–h The catalytic site is illustrated with the bound ligand molecules: 6-deoxy L-psicose (a, green), L-erythrulose (b, salmon pink), 1-deoxy 3-keto D-galactitol (D3G, c, yellow), 1-deoxy D-tagatose in form A (d, cyan), 1-deoxy D-tagatose in form B (e, cyan), 1-deoxy L-tagatose (f, blue), glycerol (g, magenta), and D-talitol (h, orange). A metal ion and water molecules are shown in blue and red spheres, respectively. Selected interactions among amino acid residues, substrates, water molecules, and a metal ion are indicated by dotted lines. i Schematic drawings of the enzyme-substrate interactions for 6-deoxy L-psicose (left) and 1-deoxy D-tagatose in form B (right, inhibitor-binding mode) are shown
Structure of bound D3G molecules in the hydrophobic groove. a The interaction between amino acid residues and bound D3G molecules is illustrated with hydrogen bonds (dotted lines). D3G molecules in the ring and linear forms are indicated by cyan and yellow, respectively. b The sphere model of D3G molecules on the surface of the enzyme is illustrated. The hydrophobic groove is shown in gray. The positions of the catalytic sites are indicated by the labels of D3G_1A and D3G_1B with red ellipses. c A close-up view of the channel of the passageway to the catalytic site from the hydrophobic groove is illustrated. Amino acid residues interacting with the D3G molecule are shown with hydrogen bonds (dotted lines)
Results
Enzyme activity
Since the highest activity of PcDTE_C66S was detected in the presence of Co2+ (118 % relative to that with Mn2+ as 100 %), the enzyme assay of PcDTE_C66S was performed using the reaction buffer containing CoCl2. The enzymatic activities of PcDTE_C66S for the various substrates are listed in Table 2. PcDTE_C66S exhibited enzymatic activity of 108 U/mg to D-tagatose, which was similar to that of wild-type PcDTE (92 U/mg in the presence of Mn2+) (Ishida et al. 1997). PcDTE_C66S exhibited enzymatic activities of 16.9 U/mg for 6-deoxy L-psicose (15.7 % of D-tagatose) and 7.6 U/mg for 1-deoxy 3-keto D-galactitol (D3G) (7.0 %). Using the enzyme solution (1.46 mg/ml), enzymatic activities for 1-deoxy D-tagatose and 1-deoxy L-tagatose were detected by HPLC analyses (Supplemental Fig. S1) and showed low activities 0.127 U/mg (0.12 %) and 0.15 U/mg (0.14 %), respectively. It was not possible to measure the enzymatic activity for L-erythrulose by a HPLC analysis due to the same retention times of the HPLC column for L-erythrulose (substrate) and D-erythrulose (product). The formation of D-erythrulose was confirmed by measuring the specific rotation of reactants (Supplemental Fig. S1f).
X-ray structure determination
After many attempts to prepare crystals of PcDTE_C66S in complexes with various ligand molecules, we successfully determined the X-ray structures of PcDTE_C66S in complexes with seven ligand molecules: 6-deoxy L-psicose, L-erythrulose, 1-deoxy 3-keto-D-galactitol (D3G), 1-deoxy D-tagatose, 1-deoxy L-tagatose, glycerol, and D-talitol (Fig. 1c, d). The structure of the PcDTE_C66S/1-deoxy L-tagatose complex was determined using crystals grown on the ground and in microgravity, and a crystal grown in microgravity gave higher resolution data (Table 1). Refinement statistics showed that the refined structures were reliable with good chemical geometries (Table 1). Since PcDTE_C66S was expressed in cells in medium containing 1 mM MnCl2, the bound metal ion at the catalytic site was refined as Mn2+, giving moderate B-factors (Table 1). The final (2F o – F c) electron density maps showed that most atoms of protein molecules, metal ions, and solvent molecules fit well. Poor and invisible electron densities were detected in several ligand molecules and His-tag residues.
Overall structure of PcDTE_C66S
The overall structure of PcDTE_C66S was equivalent to the previously reported structure of wild-type PcDTE (PDB code: 1QUL). Briefly, the monomeric structure of PcDTE_C66S has 12 α-helices (H1–H12) and eight β-strands (B1–B8), forming a (β/α)8 barrel fold with four additional short α-helices: H2, H4, H8, and H11 (Fig. 2a). A metal ion binds to the center of the barrel, which is coordinated by Glu152, Asp185, His211, and Glu246 to form the catalytic site. The replaced amino acid residue of Ser66 is on B3, to direct its side chain group to the catalytic site. There are four molecules (Mol-A, Mol-B, Mol-C, and Mol-D) in an asymmetric unit (Fig. 2b). The PISA server (Krissinel and Henrick 2007) showed the formation of two homo-dimers with a two-fold symmetry (Mol-A/Mol-B and/or Mol-C/Mol-D) in a crystal (Fig. 2b, c). Mol-A and Mol-B (or Mol-C and Mol-D) make contact with each other through additional α-helices (H8 and H11) and loop regions, in order to form a stable dimer with an interface area of 1439 Å2 (11.5 % of the monomer surface). The catalytic sites of Mol-A and Mol-B are located on the same side of the dimer. Through dimerization, the hydrophobic groove is formed by Trp15, Ile67, Trp113, Pro114, Pro117, Leu119, Phe157, Trp160, Ala258, and Trp262 in Mol-A and Mol-B, as shown in the gray surface in Fig. 2c. This hydrophobic groove reaching the catalytic sites may act as a favorable accessible surface for substrate binding.
Ligand-binding structure at the catalytic site
6-deoxy L-psicose and L-erythrulose
The catalytic site structure with the bound 6-deoxy L-psicose is shown in Fig. 3a. The electron density of the simulated annealing omit map clearly indicates that the bound 6-deoxy L-psicose is in the linear form (Supplemental Fig. S2a). O2 and O3 coordinate to the metal ion and form hydrogen bonds with His188 and Glu152, respectively. Glu246 directs its OE2 atom to the hydrogen atom attached to C3 with a distance of 2.6 Å, suggesting that Glu246 removes a proton at C3 of the substrate in the epimerization reaction, as described later. O1 forms hydrogen bonds with Glu158, His188, and Arg217 in order to fix the correct metal coordination by O2 and O3 of the substrate (Fig. 3i, left). The enzyme forms strong interactions with the 1-, 2-, and 3-positions of the substrate, while a relatively weak interaction occurs at the 4-, 5-, and 6-positions of the substrate. O4 does not form a hydrogen bond with the enzyme, and O5 interacts with Glu35 by a water-mediated hydrogen bond. C6 forms hydrophobic interactions with Trp15 and Trp113. The binding mode of 6-deoxy L-psicose in PcDTE_C66S is similar to that of the bound D-tagatose in PcDTE/D-tagatose complex (Yoshida et al. 2007) and PcDTE_C66S/D-tagatose complex (unpublished results), in which the positions of O5 and C6 are interchanged, because l-psicose is a C5 epimer of D-tagatose. The structure of the PcDTE_C66S/6-deoxy L-psicose complex shows that interactions between the enzyme and substrate at the 4-, 5, and 6-positions are not essential for substrate recognition. In practice, a small ketose, L-erythurulose without groups at the 5- and 6-positions, binds to the catalytic site, forming the same interactions between the substrate at the 1-, 2-, and 3-positions with the enzyme (Fig. 3b). Another L-erythulose molecule proximal to the catalytic site makes hydrophobic contacts with Trp15. This binding site may act as the substrate sub-binding site, as described later.
1-Deoxy 3-keto-D-galactitol (D3G)
The catalytic site structure with the bound 1-deoxy 3-keto-D-galactitol (D3G) is shown in Fig. 3c. O3 (ketone group) and O4 (hydroxyl group) of D3G coordinate to the metal ion, forming hydrogen bonds with His188 and Glu152. Glu246 directs its OE2 atom to the hydrogen atom attached to C4 with a distance of 2.3 Å. O2 forms hydrogen bonds with Glu158, His188, and Arg217. C1 (methyl group) is located in the hydrophobic pocket formed by Trp113, Phe248, and Val259. O5 does not form a hydrogen bond with the enzyme, and O6 interacts with Ser66 (Cys66 in wild-type PcDTE) by a water-mediated hydrogen bond. This structure shows that the epimerization reaction occurs at the 4-position of D3G to give 1-deoxy 3-keto d-allitol as a product (Supplemental Fig. S1c). The position of a ketone group coordinating to the metal ion (O3 of D3G or O2 of 6-deoxy L-psicose) is strictly fixed by interactions between the hydroxyl group (O2 of D3G or O1 of 6-deoxy L-psicose) and enzyme. Another D3G molecule is found at the substrate sub-binding site, interacting with Trp15.
1-Deoxy D-tagatose and 1-deoxy L-tagatose
The electron density of the bound 1-deoxy D-tagatose at the catalytic site indicated that the bound substrate adopts two alternative conformations, form A and form B, as shown in Fig. 3d, e. In form A, O2 and O3 coordinate to the metal ion, and O5 and O6 interact with Ser66 directly or via a water molecule. In form B, the direction of the bound substrate molecule is opposite to that of form A, in which O6 interacts with Glu158, His188, and Arg217 (Fig. 3i, right). O5 and O4 coordinate to the metal ion. O2 forms a water-mediated hydrogen bond with Ser66. The ketone group of the substrate is required to coordinate to the metal ion for the epimerization reaction. The binding mode of form A is suitable for the epimerization reaction, while that of form B cannot lead to the epimerization reaction, which is assumed to be the inhibitor-binding mode. The molecule of 1-deoxy L-tagatose is found to bind to the catalytic site in the inhibitor-binding mode (Fig. 3f), as found in form B of 1-deoxy D-tagatose in which O6 interacts with Glu158, His188, and Arg217, O5 and O4 coordinate to the metal ion, and O2 forms a hydrogen bond with Glu152. Another 1-deoxy L-tagatose molecule is found at the substrate sub-binding site.
Glycerol and D-talitol
The inhibitor-binding mode is found in the PcDTE_C66S complex structures with 1-deoxy D-tagatose and 1-deoxy L-tagatose, in which a ketone group does not coordinate to the metal ion, suggesting that polyalcohols (sugar alcohols) bind to the catalytic site of DTE to act as an inhibitor. The PcDTE C66S complex structures with two polyalcohols, glycerol and D-talitol, were determined, and bound polyalcohol molecules at the catalytic site were detected (Fig. 3g, h). In both structures, two hydroxyl groups, O2 and O3, coordinate to the metal ion, and O1 forms interactions with Glu158, His188, and Arg217. O6 of d-talitol forms a hydrogen bond with Ser66. Another glycerol molecule binds to the substrate sub-binding site.
Ligand binding to the hydrophobic groove
Ligand molecules were found to bind on the surface of the enzyme beside the catalytic site in the linear and/or furanose ring forms (Supplemental Fig. S3). Most of these molecules are in the hydrophobic groove formed by the dimerization of the enzyme, suggesting that it efficiently acts as the substrate accessible surface.
The D3G molecules in the hydrophobic groove are shown in Fig. 4a, b. Two D3G molecules (D3G_4A and D3G_3B) in the furanose ring form bind at both ends of the hydrophobic groove, interacting with Trp11 and Arg25. Two D3G molecules (D3G_3A and D3G_2B) in the linear form bind at the center of the hydrophobic groove, surrounded by Pro114, Pro117, Phe157, and Trp262 of Mol-A and Mol-B. The side chain groups of Ser116 of Mol-A and Mol-B protrude on the surface to form hydrogen bonds with D3G_3A and D3G_2B. The channel of the passageway to the catalytic site from the hydrophobic groove was found and formed by Trp15, Ser37, Ile67, Gly68, Trp113, and Ala258 (Fig. 4c). D3G_2A is located in this channel, forming hydrophobic interactions with the hydrophobic residues and hydrogen bonds with Ser37, which is described above as substrate sub-binding (Figs. 3b–g and 4c). The diameter of this channel is 4–5 Å, which may allow a substrate in the linear form to pass through (Fig. 4c). All of the ligand molecules binding in the channel are in the linear form (Fig. 3b–g).
Discussion
PcDTE catalyzes the epimerization of ketohexoses at the 3-position adjacent to the ketone group at the 2-position (Fig. 1a, b). The ketone group (O2) and hydroxyl group at the 3-position (O3) are required to coordinate to the metal ion at the catalytic site of the enzyme for the epimerization reaction, and this correct metal coordination by the substrate is performed by the recognition of a hydroxyl group at the 1-position (O1) with Glu158, His188, and Arg217. Thus, PcDTE strictly recognizes the substrate at the 1-, 2-, and 3-positions. On the other hand, PcDTE loosely recognizes the substrate at the 4-, 5-, and 6-positions with a few specific interactions between the substrate and enzyme. The substrate recognition at the 1-, 2-, and 3-positions is responsible for enzymatic activity, and substrate-enzyme interactions at the 4-, 5-, and 6-positions are not essential for the catalytic reaction of the enzyme; they merely adjust substrate to the proper orientation. This may be the reason why PcDTE efficiently catalyzes the epimerization of various ketoses including deoxy sugars as substrates (Itoh et al. 1994; Gullapalli et al. 2010).
The 6-deoxy L-psicose molecule binds to the catalytic site of the enzyme with the correct metal coordination, and the enzyme exhibits similar enzymatic activity on 6-deoxy L-psicose as on D-tagatose. L-Erythrulose without groups at the 5- and 6- positions also binds to the catalytic site of the enzyme with the correct metal coordination, thereby supporting substrate-enzyme interactions at the 4-, 5-, and 6-positions not being essential for the catalytic reaction.
The recognizing position of the D3G molecule is shifted by one carbon, because C1 of D3G (methyl group) can localize in the hydrophobic pocket formed by Trp113, Phe246, and Val259, and also because O2 of D3G is recognized by Glu158, His188, and Arg217 in order to make the correct metal coordination by the ketone group (O3) and hydroxyl group (O4). Thus, the enzyme exhibits similar enzymatic activity for D3G to give 1-deoxy 3-keto D-allitol as the product (Supplemental Fig. S1).
Two alternative conformations (form A and form B) are found in the structures of the bound 1-deoxy D-tagatose molecule. Form A forms the correct metal coordination by O2 and O3 and has no hydroxyl group at the 1-position. Form B forms the incorrect metal coordination by two hydroxyl groups, O5 and O4, and has the hydroxyl group, O6, recognized by Glu158, His188, and Arg217. This suggests that forms A and B compete with each other for binding to the catalytic site of the enzyme, giving two conformers in a crystal. The bound 1-deoxy L-tagatose molecule is found in the binding mode of form B, the inhibitor-binding mode (Fig. 3f). 1-Deoxy ligand molecules may predominantly bind to the catalytic site of the enzyme in the inhibitor-binding mode and may partially act as inhibitors, giving low enzymatic activities for 1-deoxy ligand molecules.
Glycerol and D-talitol molecules bind to the catalytic site in the inhibitor-binding mode with two hydroxyl groups, O2 and O3, coordinating to the metal ion. Although these polyalcohol molecules appear to act as competitive inhibitors, they exhibit neither competitive inhibition nor inhibitive effects in the presence of D-tagatose (data not shown). The ligand-enzyme interactions of polyalcohol molecules may be weaker than those of D-tagatose, which can correctly coordinate the metal ions, and can favorably interact with the enzyme.
Based on the X-ray structure of PcDTE in complex with D-tagatose, we previously proposed the C3–O3 proton-exchange mechanism via a cis-enediolate intermediate for the catalytic reaction mechanism used by PcDTE (Yoshida et al. 2007). Since monosaccharides exist in furanose and/or pyranose ring forms in solution, the enzyme is required to open the sugar ring for the epimerization reaction (Fig. 1a). The sugar-ring opening mechanism used by DTE is currently unknown. The X-ray structures of the sugar epimerases and sugar isomerases in complexes with monosaccharides in the ring form contributed to elucidating the sugar-ring opening mechanisms used by enzymes (Carrell et al. 1989, 1994; Collyer et al. 1990; Whitlow et al. 1991; Fenn et al. 2004; Kovalevsky et al. 2010; Yoshida et al. 2010, 2012, 2014; Munshi et al. 2014; Langan et al. 2014; Terami et al. 2015). In most cases, the substrate in the ring form binds to the catalytic site of the enzyme, in which the amino acid residues or a water molecule act as an acid/base catalyst for ring opening. In the PcDTE/D3G complex, five of the bound D3G molecules (two in the ring form and three in the linear form) are located in the hydrophobic groove (Fig. 4a, b). The channel of the passageway to the catalytic site is too narrow for a substrate in the ring form to pass, but it may allow a substrate in the linear form to pass through, suggesting that the sugar-ring opening of a substrate may occur in the hydrophobic groove.
The proposed catalytic reaction mechanism including sugar-ring opening of PcDTE for D3G as a substrate is shown in Fig. 5. D3G in the furanose ring form approaches the hydrophobic groove of the dimeric enzyme (Fig. 5a and D3G_4A and D3G_3B in Fig. 4b). A water molecule in the hydrophobic groove may transfer a proton from O3 to O6 in order to open the furanose ring (Fig. 5a, b). In the hydrophobic groove, the position of equilibrium between the linear form and ring form may move to the linear form, because the hydrophobic environment is favorable for a substrate in the linear form, which has the ability to change its conformation to form hydrophobic interactions with the enzyme. The narrow channel formed by Trp15, Ser37, Ile67, Gly68, Trp113, and Ala258 allows the D3G molecule in the linear form to pass through to the catalytic site, acting as a substrate sub-binding site (Fig. 5b). In the catalytic site, D3G forms the metal coordination with O3 and O4. His188 gives a proton to O3, and Glu152 and Glu256 remove protons from O4 and C4, respectively, to produce a cis-enediolate intermediate with a planar structure of O3–C3–C4–O4 that may be stabilized by the metal ion (Fig. 5c, d). Proton transfer then occurs from Glu246 to O4 and from Glu152 to C4, giving the product (Fig. 5d, e). Consequently, the protons of C4 and O4 are exchanged through the catalytic reaction. Since Fig. 5 shows the catalytic reaction for D3G as a substrate, proton exchange occurs between C4 and O4. This catalytic reaction mechanism may be applied to other substrates as the C3–O3 proton-exchange mechanism proposed in PcDTE/D-tagatose complex structure (Yoshida et al. 2007), in which His188 gives a proton to O2, and Glu152 and Glu256 remove protons from O3 and C3, respectively.
The X-ray structure of Agrobacterium tumefaciens D-psicose 3-epimerase (AtDPE), one of DTE family enzymes, was reported, and its catalytic reaction mechanism was proposed (Kim et al. 2006b). In AtDPE, there was no wide hydrophobic groove acting as a favorable accessible surface for substrate binding, and the channel of the passageway to the catalytic site changed its size from open (6–7 Å) to close (4–5 Å) conformations depending on substrate binding; Trp112 (Trp113 in PcDTE) approached to the substrate by 5.6 Å (Supplemental Fig. S4). In PcDTE, such conformational change was not found, and the channel to the catalytic site has the same size (4–5 Å) through the substrate binding (Yoshida et al. 2007). Thus, the hydrophobic groove and the narrow channel are unique to PcDTE, suggesting that sugar-ring opening mechanism may be different between PcDTE and AtDPE. Another difference is found in the ionized states of the catalytic residues. In AtDPE, Glu244 (Glu246 in PcDTE) in ionized form removes a proton from C3 to produce a cis-enediolate intermediate and subsequently Glu150 (Glu152) in unionized form protonates C3 on the opposite side. However, it is likely that both Glu residues (Glu152 and Glu246) are in the negatively ionized form stabilized by the positive charge of Mn2+, as indicated in Fig. 5c, e.
The X-ray structures of PcDTE_C66S with deoxy sugars could successfully explain the relationship between substrate-enzyme interactions and enzymatic activities and also provide insights into the catalytic reaction mechanism including sugar-ring opening of PcDTE. We obtained the preliminary experimental results that the enzymatic activities drastically decrease by the amino acid replacements of Trp15 and/or Trp113, which form the narrow channel for the substrate, partially supporting the proposed sugar-ring opening mechanism of PcDTE. The further study is now in progress.
References
Brunger AT (1993) X-PLOR 3.1: a system for X-ray crystallography and NMR. Yale University Press, New Haven and London
Carrell HL, Glusker JP, Burger V, Manfre F, Tritsch D, Biellmann JF (1989) X-ray analysis of D-xylose isomerase at 1.9 A: native enzyme in complex with substrate and with a mechanism-designed inactivator. Proc Natl AcadSci USA 86:4440–4444
Carrell HL, Hoier H, Glusker JP (1994) Modes of binding substrates and their analogues to the enzyme D-xylose isomerase. Acta Crystallogr Sect D 50:113–123. doi:10.1107/S0907444993009345
Collyer CA, Henrick K, Blow DM (1990) Mechanism for aldose-ketose interconversion by D-xylose isomerase involving ring opening followed by a 1,2-hydride shift. J Mol Biol 212:211–235. doi:10.1016/0022-2836(90)90316-E
Emsley P, Lohkamp B, Scott W, Cowtan K (2010) Features and development of Coot. Acta Crystallogr D Biol Crystallogr 66:486–501. doi:10.1107/S0907444910007493
Fenn TD, Ringe D, Petsko GA (2004) Xylose isomerase in substrate and inhibitor michaelis states: atomic resolution studies of a metal-mediated hydride shift. Biochemistry 43:6464–6474. doi:10.1021/bi049812o
Gullapalli P, Yoshihara A, Morimoto K, Rao D, Akimitsu K, Jenkinson SF, Fleet GWJ, Izumori K (2010) Conversion of l-rhamnose into ten of the sixteen 1- and 6-deoxyketohexoses in water with three reagents: d-tagatose-3-epimerase equilibrates C3 epimers of deoxyketoses. Tetrahedron Lett 51(6):895–898. doi:10.1016/j.tetlet.2009.12.024
Hayashi N, Iida T, Yamada T, Okuma K, Takehara I, Yamamoto T, Yamada K, Tokuda M (2010) Study on the postprandial blood glucose suppression effect of D-psicose in borderline diabetes and the safety of long-term ingestion by normal human subjects. Biosci Biotechnol Biochem 74:510–519. doi:10.1271/bbb.90707
Hayashi N, Yamada T, Takamine S, Iida T, Okuma K, Tokuda M (2014) Weight reducing effect and safety evaluation of rare sugar syrup by a randomized double-blind, parallel-group study in human. J Funct Foods 11:152–159. doi:10.1016/j.jff.2014.09.020
Hossain A, Yamaguchi F, Matsunaga T, Hirata Y, Kamitori K, Dong Y, Sui L, Tsukamoto I, Ueno M, Tokuda M (2012) Rare sugar D-psicose protects pancreas β-islets and thus improves insulin resistance in OLETF rats. Biochem Biophys Res Commun 425:717–723. doi:10.1016/j.bbrc.2012.07.135
Hossain MA, Kitagaki S, N akano D, Nishiyama A, Funamoto Y, Matsunaga T, Tsukamoto I, Yamaguchi F, Kamitori K, Dong Y, Hirata Y, Murao K, Toyoda Y, Tokuda M (2011) Rare sugar D-psicose improves insulin sensitivity and glucose tolerance in type 2 diabetes Otsuka Long-Evans Tokushima Fatty (OLETF) rats. Biochem Biophys Res Commun 405:7–12. doi:10.1016/j.bbrc.2010.12.091
Iida T, Kishimoto Y, Yoshikawa Y, Hayashi N, Okuma K, Tohi M, Yagi K, Matsuo T, Izumori K (2008) Acute D-psicose administration decreases the glycemic responses to an oral maltodextrin tolerance test in normal adults. J Nutr Sci Vitaminol (Tokyo) 54:511–514. doi:10.3177/jnsv.54.511
Iida T, Yamada T, Hayashi N, Okuma K, Izumori K, Ishii R, Matsuo T (2013) Reduction of abdominal fat accumulation in rats by 8-week ingestion of a newly developed sweetener made from high fructose corn syrup. Food Chem 138(2–3):781–785. doi:10.1016/j.foodchem.2012.11.017
Ishida Y, Kamiya T, Izumori K (1997) Production of d-tagatose 3-epimerase of Pseudomonas cichorii ST-24 using recombinant Escherichia coli. J Ferment Bioeng 84:348–350. doi:10.1016/S0922-338X(97)89257-4
Itoh H, Okaya H, Khan AR, Tajima S, Hayakawa S, Izumori K (1994) Purification and characterization of D-tagatose 3-epimerase from Pseudomonas sp. ST-24. Biosci Biotechnol Biochem 58:2168–2171. doi:10.1271/bbb.58.2168
Itoh H, Sato T, Izumori K (1995) Preparation of D-psicose from D-fructose by immobilized D-tagatose 3-epimerase. J Ferment Bioeng 80:101–103. doi:10.1016/0922-338X(95)98186-O
Izumori K, Khan AR, Okaya H, Tsumura T (1993) A new enzyme, D-ketohexose 3-epimerase, from Pseudomonas sp. ST-24. Biosci Biotechnol Biochem 57:1037–1039. doi:10.1271/bbb.57.1037
Jia M, Mu WM, Chu FF, Zhang XM, Jiang B, Zhou LL, Zhang T (2014) A D-psicose 3-epimerase with neutral pH optimum from Clostridium bolteae for D-psicose production: cloning, expression, purification, and characterization. Appl Microbiol Biotechnol 98:717–725. doi:10.1007/s00253-013-4924-8
Kim HJ, Hyun EK, Kim YS, Lee YJ, Oh DK (2006a) Characterization of an Agrobacterium tumefaciens D-psicose 3-epimerase that converts D-fructose to D-psicose. Appl Environ Microbiol 72:981–985. doi:10.1128/AEM.72.2.981-985.2006
Kim K, Kim HJ, Oh DK, Cha SS, Rhee S (2006b) Crystal structure of D-psicose 3-epimerase from Agrobacterium tumefaciens and its complex with true substrate D-fructose: a pivotal role of metal in catalysis, an active site for the non-phosphorylated substrate, and its conformational changes. J Mol Biol 361:920–931. doi:10.1016/j.jmb.2006.06.069
Kovalevsky AY, Hanson L, Fisher SZ, Mustyakimov M, Mason SA, Forsyth VT, Blakeley MP, Keen DA, Wagner T, Carrell HL, Katz AK, Glusker JP, Langan P (2010) Metal ion roles and the movement of hydrogen during reaction catalyzed by D-xylose isomerase: a joint x-ray and neutron diffraction study. Structure 18:688–699. doi:10.1016/j.str.2010.03.011
Krissinel E, Henrick K (2007) Inference of macromolecular assemblies from crystalline state. J Mol Biol 372:774–797. doi:10.1016/j.jmb.2007.05.022
Langan P, Sangha AK, Wymore T, Parks JM, Yang ZK, Hanson BL, Fisher Z, Mason SA, Blakeley MP, Forsyth VT, Glusker JP, Carrell HL, Smith JC, Keen DA, Graham DE, Kovalevsky A (2014) L-Arabinose binding, isomerization, and epimerization by D-xylose isomerase: X-ray/neutron crystallographic and molecular simulation study. Structure 22:1287–1300. doi:10.1016/j.str.2014.07.002
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1992) PROCHECK v.2: programs to check the stereochemical quality of protein structures. Oxford Molecular Ltd, Oxford, England
Matsuo T, Izumori K (2009) D-Psicose inhibits intestinal alpha glucosidase and suppresses the glycemic response after ingestion of carbohydrates in rats. J Clin Biochem Nutr 45:202–206. doi:10.3164/jcbn.09-36
Matsuo T, Suzuki H, Hashiguchi M, Izumori K (2002) D-Psicose is a rare sugar that provides no energy to growing rats. J Nutr Sci Vitaminol 48:77–80. doi:10.3177/jnsv.48.77
Matsuo T (2006) Inhibitory effects of D-psicose on glycemic responses after oral carbohydrate tolerance test in rats. J Jpn Soc Nutr Food Sci 59:119–121. doi:10.4327/jsnfs.59.119
McRee DE (1999) XtalView/Xfit: a versatile program for manipulating atomic coordinate and electron density. J Struct Biol 125:156–165. doi:10.1006/jsbi.1999.4094
Mu W, Zhang W, Feng Y, Jiang B, Zhou L (2012) Recent advances on applications and biotechnological production of D-psicose. Appl Microbiol Biotechnol 94:1461–1467. doi:10.1007/s00253-012-4093-1
Mu WM, Chu FF, Xing QC, Yu SH, Zhou L, Jiang B (2011) Cloning, expression, and characterization of a D-psicose 3-epimerase from Clostridium cellulolyticum H10. J Agric Food Chem 59:7785–7792. doi:10.1021/jf201356q
Mu WM, Zhang WL, Fang D, Zhou L, Jiang B, Zhang T (2013) Characterization of a D-psicose-producing enzyme, D-psicose 3-epimerase, from Clostridium sp. Biotechnol Lett 35:1481–1486. doi:10.1007/s10529-013-1230-6
Munshi P, Snell EH, van der Woerd MJ, Judge RA, Myles DA, Ren Z, Meilleur F (2014) Neutron structure of the cyclic glucose-bound xylose isomerase E186Q mutant. Acta Crystallogr D Biol Crystallogr 70:414–420. doi:10.1107/S1399004713029684
Murshudov GN, Vagin AA, Dodson EJ (1997) Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr 53:240–255. doi:10.1107/S0907444996012255
Nakajima Y, Gotanda T, Uchiyama H, Furukawa T, Haraguchi M, Ikeda R, Sumizawa T, Yoshida H, Akiyama S (2004) Inhibition of metastasis of tumor cells overexpressing thymidine phosphorylase by 2-deoxy-L-ribose. Cancer Res 64:1794–1801
Otwinowski Z, Minor W (1997) Processing of X-ray diffraction data collected in oscillation mode. Method in enzymology 276: macromolecular crystallography, part A, 307–326. doi:10.1016/S0076–6879(97)76066-X
Shompoosang S, Yoshihara A, Uechi K, Asada Y, Morimoto K (2014) Enzymatic production of three 6-deoxy-aldohexoses from L-rhamnose. Biosci Biotechnol Biochem 78:317–325. doi:10.1080/09168451.2014.878217
Shompoosang S, Yoshihara A, Uechi K, Asada Y, Morimoto K (2016) Novel process for producing 6-deoxy monosaccharides from l-fucose by coupling and sequential enzymatic method. J Biosci Bioeng 121:1–6. doi:10.1016/j.jbiosc.2015.04.017
Takahashi S, Ohta K, Furubayashi N, Yan B, Koga M, Wada Y, Yamada M, Inaka K, Tanaka H, Miyoshi H, Kobayashi T, Kamigaichi S (2013) JAXA protein crystallization in space: ongoing improvements for growing high-quality crystals. J Synchrotron Radiat 20(Pt 6):968–973. doi:10.1107/S0909049513021596
Terami Y, Yoshida H, Uechi K, Morimoto K, Takata G, Kamitori S (2015) Essentiality of tetramer formation of Cellulomonas parahominis L-ribose isomerase involved in novel L-ribose metabolic pathway. Appl Microbiol Biotechnol 99:6303–6313. doi:10.1007/s00253-015-6417-4
Vagin A, Teplyakov A (1997) MOLREP: an automated program for molecular replacement. J Appl Crystallogr 30:1022–1025. doi:10.1107/S0021889897006766
Whitlow M, Howard AJ, Finzel BC, Poulos TL, Winborne E, Gilliland GL (1991) A metal-mediated hydride shift mechanism for xylose isomerase based on the 1.6 Å Streptomyces rubiginosus structures with xylitol and D-xylose. Proteins 9:153–173. doi:10.1002/prot.340090302
Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AGW, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS (2011) Overview of the CCP4 suite and current developments. Acta Crystallogr D Biol Crystallogr 67:235–242. doi:10.1107/S0907444910045749
Yoshida H, Yamada M, Nishitani T, Takada G, Izumori K, Kamitori S (2007) Crystal structures of D-tagatose 3-epimerase from Pseudomonas cichorii and its complexes with D-tagatose and D-fructose. J Mol Biol 374(2):443–453. doi:10.1016/j.jmb.2007.09.033
Yoshida H, Yamaji M, Ishii T, Izumori K, Kamitori S (2010) Catalytic reaction mechanism of Pseudomonas stutzeri L-rhamnose isomerase deduced from X-ray structures. FEBS J 277:1045–1057. doi:10.1111/j.1742-4658.2009.07548.x
Yoshida H, Yoshihara A, Teraoka M, Terami Y, Takata G, Izumori K, Kamitori S (2014) X-ray structure of a novel L-ribose isomerase acting on a non-natural sugar L-ribose as its ideal substrate. FEBS J 281:3150–3164. doi:10.1111/febs.12850
Yoshida H, Yoshihara A, Teraoka M, Yamashita S, Izumori K, Kamitori S (2012) Structure of L-rhamnose isomerase in complex with L-rhamnopyranose demonstrates the sugar-ring opening mechanism and the role of a substrate sub-binding site. FEBS Open Bio 3:35–40. doi:10.1016/j.fob.2012.11.008
Zhang L, Mu W, Jiang B, Zhang T (2009) Characterization of D-tagatose-3-epimerase from Rhodobacter sphaeroides that converts D-fructose into D-psicose. Biotechnol Lett 31:857–862. doi:10.1007/s10529-009-9942-3
Zhang W, Fang D, Xing Q, Zhou L, Jiang B, Mu WM (2013b) Characterization of a novel metal-dependent D-psicose 3-epimerase from Clostridium scindens 35704. PLoS One 8:e62987. doi:10.1371/journal.pone.0062987
Zhang W, Fang D, Zhang T, Zhou L, Jiang B, Mu W (2013a) Characterization of a metal-dependent D-psicose 3-epimerase from a novel strain, Desmospora sp. 8437. J Agric Food Chem 61:11468–11476. doi:10.1021/jf4035817
Zhang W, Zhang T, Jiang B, Mu W (2015) Biochemical characterization of a d-psicose 3-epimerase from Treponema primitia ZAS-1 and its application on enzymatic production of d-psicose. J Sci Food Agric 96:49–56. doi:10.1002/jsfa.7187
Zhu YM, Men Y, Bai W, Li XB, Zhang LL, Sun YX, Ma Y (2012) Overexpression of D-psicose 3-epimerase from Ruminococcus sp. in Escherichia coli and its potential application in D-psicose production. Biotechnol Lett 34:1901–1906. doi:10.1007/s10529-012-0986-4
Acknowledgments
We thank Mr. Y. Tahara, Ms. S. Kayahara, and Mr. S. Ohga for assisting with protein purification; Mr. K. Yube for his technical assistance with DNA sequencing; and Dr. K. Inaka, Dr. N. Furubayashi, Dr. M. Yamada, and Dr. K. Ohta for assisting with crystallizations under microgravity. This research was performed with the approval of the Photon Factory Advisory Committee and National Laboratory for High Energy Physics, Japan. This study is contributed by a part of “High-Quality Protein Crystal Growth Experiment on KIBO” promoted by JAXA (Japan Aerospace Exploration Agency). Russian spacecraft “Progress” and/or “Soyuz” provided by the Russian Federal Space Agency were used for space transportation. A part of space crystallization technology had been developed by the European Space Agency and University of Granada.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded in part by Grants-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (25440028, 23770122).
Conflict interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Yoshida, H., Yoshihara, A., Ishii, T. et al. X-ray structures of the Pseudomonas cichorii D-tagatose 3-epimerase mutant form C66S recognizing deoxy sugars as substrates. Appl Microbiol Biotechnol 100, 10403–10415 (2016). https://doi.org/10.1007/s00253-016-7673-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7673-7