Abstract
The aim of this study was to isolate and characterize bacteria from the compost of fruit and vegetable waste (FVW) for plant growth-promoting (PGP) activities and investigate the pro-active influence of bacterial isolates on wheat growth. Fourteen bacterial strains (RHC-1 to RHC-14) were isolated and purified in tryptic soya agar (TSA). In addition to being biochemically characterized, these bacterial strains were also tested for their PGP traits, such as phosphate (P)-solubilization, nifH gene amplification, indole-3-acetic acid (IAA) quantification and the production of ammonia, oxidase and catalase. Based on 16S rRNA gene sequencing, these bacterial strains were identified as belonging to species of Bacillus, Lysinibacillus, Lysobacter, Staphylococcus, Enterobacter, Pseudomonas and Serratia. All bacterial strains solubilized tri-calcium phosphate and produced IAA. Two bacterial strains RHC-8 (Enterobacter sp.) and RHC-13 (Pseudomonas sp.) solubilized the maximum amount of tri-calcium phosphate, i.e. 486 and 464 μg/ml, respectively. P-solubilization was associated with a significant drop in the pH of the broth culture from an initial pH of 7 to pH 4.43. In addition to P-solubilization and IAA production, six bacterial strains also carried the nifH gene and were further evaluated for their effect on wheat (Triticum aestivum) growth under controlled conditions. All six bacterial strains enhanced wheat growth as compared to uninoculated control plants. Two of the bacterial strains, RHC-8 and RHC-13, identified as Enterobacter aerogenes and Pseudomonas brenneri, respectively, were assessed as potential PGP rhizobacteria due to exhibiting characteristics of four or more PGP traits and enhancing wheat growth though their specific mechanism of action.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
The soils of Pakistan are nutrient deficient, and farmers are dependent on the use of inorganic fertilizers for crop production, which are both expensive and environmentally damaging. However, organic and biofertilizers, based on beneficial soil bacteria, are an alternative option which could, if better understood, result in decreased NH4NO3 fertilizer usage and improved crop production, especially in resource-poor farming systems such as those found in Pakistan. Soil organic matter level can be maintained and improved though the use of crop residues, farm yard manures, green manuring and other organic wastes (Manici et al. 2004). Organic wastes can be recycled and converted into nutrient-rich amorphous humus substance by applying composting technology.
Bacteria beneficial to plant growth, usually referred to as plant growth-promoting rhizobacteria (PGPR), are capable of promoting plant growth by colonizing the plant root. PGPR, also termed plant health-promoting rhizobacteria (PHPR) or nodule-promoting rhizobacteria (NPR) are attached to the root rhizosphere, which is the soil ecological environment for plant–microbe interactions (Hayat et al. 2010). PGPR have the potential to contribute to sustainable plant growth promotion though direct and indirect mechanisms. Indirect mechanisms include the production of metal-binding siderophores, antibiotics, hydrogen cyanide, chitinase and ß-1,3-glucanase, and direct plant growth promotion activities include the production of auxins, cytokinins, gibberellins, ethylene, abscisic acid, indole-3-ethanol or indole-3-acetic acid (IAA). PGPR also help to solubilize mineral phosphates (P-solubilization) and fix nitrogen (N2-fixation). PGPR retain more soil organic N and other nutrients in the plant–soil system, thus enhancing the release of nutrients and reducing the need for chemical fertilizers (Hayat et al. 2010). Different genera of bacteria, including Bacillus, Pseudomonas, Enterobacter, Rhizobium, Azospirillum, Azotobacter and Serratia, possess a range of properties, including P-solubilization, N2-fixation and the ability to produce cytokinins, antibiotics and hydrolytic enzymes, have been shown to increase yield in wheat, rice, maize, sugar beet and canola (Hayat et al. 2012).
Enrichment of compost with PGPR and plant growth regulators has a synergistic effect on crop growth (Akhtar et al. 2009; Hameeda et al. 2006). The aim of our study was to isolate and characterize bacteria from composting fruit and vegetable wastes (FVW) for their PGP traits and then investigate the pro-active effect of these bacterial isolates on wheat growth.
Materials and methods
Isolation and biochemical characterization of compost bacteria
Bacterial strains were isolated from an aerobically prepared compost of FVW though the dilution plate technique using phosphate buffered saline (PBS, 1×) as the saline solution. These strains were grown in tryptic soya agar medium [TSA; (in g/l): pancreatic digest of casein, 15; papaic digest of soybean, 5; NaCl, 5; agar, 15; pH 7.3 ± 0.2; Difco Co, BD Diagnostic Systems, Detroit, MI] in sterilized petri plates at 28 °C for at least 72 h to allow bacterial growth. Individual colonies were then picked and streaked on plates containing TSA medium for purification and screening under sterilized conditions. Single colonies were re-streaked on TSA medium repeatedly until purified cultures were obtained. Fourteen bacterial strains were screened and stored in glycerol (35 %, w/v) stock at −80 °C for further characterization. The ability of the isolates to utilize a specific substrate was assessed using the AP-20E (bioMérieux, Marcy l'Etoile, France) micro-test galleries. Cells were grown in TSA medium and inoculated AP strips were incubated at 28–30 °C for 6–7 h. Initial readings were taken after every 24 h. Similarly, oxidase activity was measured using the Oxidase kit (bioMérieux), and catalase activity was determined following Cowan and Steel (2004). All of these commercial kits were used according to the manufacturer’s protocol. Ammonia production by bacterial strains was tested in peptone water. Each bacterial strain was inoculated in 10 ml peptone water and incubated at 28 °C for 48 h. Nessler’s reagents (0.5 ml) were added to each tube, and the subsequent appearance of a brown to yellow color indicated the culture tested positive for ammonia production (Cappuccino and Sherman 2005).
PGP activities of bacterial strains
Bacterial strains were evaluated for their capacity to produce indole acetic acid (IAA), solubilize phosphorous and fix nitrogen (nifH gene amplification). For IAA production, bacterial cultures were grown for 48 h in tryptic soya broth (TSB) at 28 ± 2 °C. A bacterial suspension (100 μl each) of fully grown bacterial culture was inoculated in 5 ml Luria Broth (LB) medium either supplemented or not with 500 μg/ml tryptophan and, placed for 48 h in an incubating shaker at 28 ± 2 °C. Fully grown cultures were centrifuged at 10,000 rpm for 10 min, and two drops of orthophosphoric acid and 4 ml of Salkowski reagents (50 ml, 35 % of perchloric acid, 1 ml 0.5 M FeCl3 solution) were added to 2 ml supernatant solution. The development of a pink color indicated IAA production, and the optical density (O.D, 530 nm) was read using a spectrophotometer. The concentration of IAA produced by the cultures was measured using a standard curve graph and a range of standards up to 10 μg/ml (Brick et al. 1991). A qualitative analysis of phosphorus solubility by the bacterial strains was also performed by measuring the halo zone around bacterial growth on Pikovskaya agar medium (Gaur 1990; Pikovskaya 1948). To determine the quantitative phosphorus-solubilizing capacity of bacteria, TSB or liquid PKV medium was prepared and 100 ml was dispensed into each 250-ml conical flask. To each flask, 5 g/l insoluble phosphate in the form of tri-calcium phosphate was added. The pH of media was recorded, and the flasks with media were sterilized at 121 °C for 15 min. The flasks were then inoculated with 500 μl bacterial suspension after cooling and placed on a shaker for 8 days at 30 °C. The pH of the media was recorded after 8 days, and then each culture was centrifuged at 8,500 rpm for 25 min, following which the supernatant of each culture was collected in 15-ml centrifuge tubes. An extract of each solution was prepared (Soultanpour and Workman 1979), and the available phosphorus in each broth culture was determined by Watanabe and Olsen (1965). Optical density readings were made at 700 nm, and the concentration of phosphate solubilized by strains was measured using a standard curve graph and standards ranging up to 1 μg/ml. The nifH gene was amplified to check the nitrogen-fixing ability of the bacterial strains after PCR amplification of the genes, as described by Katsivela et al. (1999), using universal forward and reverse primers (PolFb: TGC GAY CCS AAR GCB GAC TC; PolRb: ATS GCC ATC ATY TCR CCG GA).
Identification of bacterial strains using 16SrRNA gene sequencing
The DNA template was prepared by picking an individual colony of each bacterial strain, and amplification of the 16S rRNA gene was carried out by the PCR. PCR amplification of DNA was performed following Katsivela et al. (1999) using universal primers (9F: 5′́-GAGTTTGATCCTGGCTCAG-3′; 1510R: 5′́-GGCTACCTTGTTACGA-3′́) in a reaction mixture (25 μl). The amplification program for the full-length 16S rRNA gene consisted of an initial denaturion at 94 °C for 2 min, followed by 30 cycles of denaturation at 94 °C for 2 min, primer annealing at 55 °C for 1 min and primer extension at 72 °C for 2 min, followed by a final extension at 72 °C for 10 min, in a thermocycler. Amplified PCR products of the 16S ribosomal gene were separated on 1 % agarose gel in 0.5× TE (Tris-EDTA) buffer containing 2 μl ethidium bromide (20 mg/ml). The λ HindIII ladder were used as a size marker. The gel was viewed under UV light and photographed using a gel documentation system. Amplified PCR products of the full-length 16S rRNA gene were purified using PCR purification kit (Qiagen, Venlo, The Netherlands) according to the standard protocol recommended by the manufacturer and earlier followed by Ahmed et al. (2007). The purified PCR product samples were sent to MACROGEN (Seoul, Korea) for sequencing using universal 16S rRNA sequencing primers. The sequence results were obtained from a BLAST search of the EzTaxon server (Chun et al. 2007), and the sequences of all the related species were retrieved to determine the exact nomenclature of the isolates. Phylogenetic analyses were performed using bioinformatics software MEGA-4 (Tamura et al. 2007). CLUSTAL X and BioEdit were used for sequence alignment and comparison, respectively. The DNA accession numbers of each strain were obtained from the DNA Data Bank of Japan (DDBJ).
Wheat inoculation
A pot experiment was carried out in the glasshouse to investigate the beneficial effects of six potential bacterial strains based on their PGP activities on wheat (Triticum aestivum) growth during the winter season of 2010. Prior to sowing, the pots were filled with 0.5 kg of autoclaved soil. Four seeds of wheat crop were sown in each pot, but after germination, two seedlings were removed. The seeds were treated with the inoculums of six different strains. The experiment was laid down in a completely randomized design with five replications. To evaluate the response of bacterial strains, shoot and root length, shoot and root fresh weight and shoot and root dry weight were observed after 2 months. Analysis of variance (ANOVA) was performed on each set of data, and means were compared by the least significant difference (LSD) at the α = 0.05 level (Steel et al. 1997).
Results
Isolation and biochemical characterization of compost bacteria
Fourteen bacterial strains, designated RHC-1 to RHC-14, differing in colony morphology in terms of color, shape and elevation were isolated from FVW compost. Both Gram-positive and -negative strains were present. Almost all strains exhibited the catalase-positive reaction except for RHC-10. Details on the biochemical characterizations of all bacterial strains are given in Table 1. Some strains produced relatively fast-growing colonies within 36 h (28 °C) while others grew after 2–3 days.
Plant growth-promoting activities
The results of PGP activities of all bacterial strains are shown in Table 2. All bacterial strains produced IAA in both the presence and absence of tryptophan, but the quantity produced by each strain was lower in the absence of tryptophan. The amount of IAA produced by a specific strain varied greatly, ranging from 0.74 (absence of tryptophan) to 16.24 μg/ml (presence of tryptophan). RHC-13 (Pseudomonas brenneri) produced the maximum amount of IAA followed by RHC-8 (Enterobacter aerogenes) with tryptophan supplemented to the broth culture. The bacterial strains identified as Bacillus and Lycinibacillus produced small quantities of IAA (0.74–1.20 μg/ml), while Pseudomonas, Enterobacter and Serratia produced substantial quantities (up to 16.24 μg/ml). The inorganic P-solubilization by the bacterial strains ranged from 37.19 to 486.28 μg/ml, and there was a variation among the different strains. A significant drop in the pH of the broth medium was observed during P-solubilization by the different bacterial strains. All bacterial strains solubilized substantial quantity of inorganic phosphorus, however maximum P-solubilization (486.28 μg/ml) was observed with RHC-8 followed by RHC-13 (464.49 μg/ml). This solubilization of phosphates by bacterial strains in broth culture dropped the pH (4.43 and 4.64, respectively) significantly from an initial pH level of 7.0 during 8 days of incubation. The maximum drop in pH (4.43) was observed for strain RHC-8, which also showed the highest amount of tri-calcium P-solubilization. These results indicate that some bacterial strains, such as RHC-3, RHC-4, RHC-8, RHC-12, RHC-13 and RHC-14, possess PGP activities, such as solubilization of insoluble tri-calcium phosphate, production of auxin-IAA, and nifH positivity. Based on nearly 400 bp of nifH gene amplification, we found that only the strains showing these activities carried the nifH gene, while the nifH gene was absent in all others strains. Consequently, we selected the bacterial strains RHC-3, RHC-4, RHC-8, RHC-12, RHC-13 and RHC-14 as potential PGPR for further crop tests.
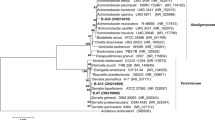
Molecular identification based on 16S rRNA gene sequence
The identification of bacterial strains based on 16S rRNA gene sequences is presented in Table 3. A BLAST search of nucleotide sequences (1,400–1,500 bp) through the EzTaxon server showed that two of the isolated bacterial strains had the highest (98–99 %) similarity with genus Bacillus (RHC-1: 99.693 % with B. anthacis ATCC 14578T AB190217; RHC-11: 98.324 % with B. mycoides DSM 2048T ACMU01000002), and three isolated strains had the highest homology with genus Pseudomonas (RHC-2: 99.435 % with Pseudomonas beteli ATCC 19861T AB021406; RHC-5: 99.313 % with Pseudomonas stutzeri CCUG 11256T U26262; RHC-13: 98.031 % with Pseudomonas brenneri CFML 97–391T AF268968). Closely related species of RHC-3, RHC-4 and RHC-9 were identified by the BLAST search as Lysinibacillus xylanilyticus XDB9T FJ477040. Similarly, the sequences of RHC-6 and RHC-8 showed the highest similarity with Lysobacter defluvii IMMIB APB-9T AM283465 and Enterobacter aerogenes NCTC 10006T AJ251468, respectively. The two bacterial strains designated as RHC-7 and RHC-10 were identified as Staphylococcus spp., while the sequence of other two strains, RHC-12 and RHC-14, was similar to Serratia glossinae C1T FJ790328. The gene sequences of all bacterial strains were submitted to the DDBJ GenBank for accession numbers (Table 3).
Effect of inoculation with potential PGPR on wheat growth
Six bacterial stains were screened as potential PGPR based on their PGP activities and used as bio-inoculants in pots filled with sterilized soil under controlled conditions. All six strains increased wheat root and shoot length, root shoot fresh and dry biomass per plant as compared with the un-inoculated media (TSB) and un-inoculated control soil (Table 4). Two bacterial strains, RHC-8 and RHC-13, identified as Enterobacter sp. and Pseudomonas sp., respectively, were the best potential PGPR in terms of wheat root shoot length and biomass production. Maximum shoot (41.90 cm) and root (37.02 cm) length was observed with the inoculation of RHC-8 (Enterobacter aerogenes) followed by RHC-13 (Pseudomonas brenneri).
Discussion
Bacterial strains were characterized biochemically using commercial kits, and variable reactions were observed for Gram staining, catalase and oxidase activities. Some of the strains tested displayed known PGP activities, i.e. P-solubilization and IAA production, and possessed the nifH gene (+). PGPR mediate P-solubilization through the biosynthesis of organic acids, creating acidified conditions in the media (Goldstein 2007). PGPR solubilize more inorganic phosphate than their non-PGPR counterparts though gene modification followed by expression of the modified gene (Rodríguez et al. 2006). Phytase genes have been cloned from a number of PGPR (Tye et al. 2002). The level of P-solubilization depends on the bacterial species, host crop and environmental conditions (Çakmakçi et al. 2006). In our study, we also observed a significant drop in the pH of the broth culture after 8 days of incubation, and the maximum drop was associated with the highest P-solubilization (486 μg/ml). A similar pH decrease in broth media by different PGPR has been reported by other researchers (Illmer and Schinner 1992; Yu et al. 2011). In our study, RHC-8 (Enterobacter sp.) and RHC-13 (Pseudomonas sp.) solubilized the maximum amount of tri-calcium phosphate, i.e. 486.28 and 464.49 μg/ml, respectively. Plant available phosphorus is increased by the activity of PGPR, especially those belonging to the genera Bacillus, Pseudomonas and Enterobacter (Hayat et al. 2012), and various Pseudomonas species have been shown to be the most powerful P-solubilizing bacteria (Banerjee et al. 2006). Plant physiological processes, starting from root initiation and extending to phototropism, can be controlled by the auxin IAA, which is synthesized from tryptophan (Khan et al. 2009). In our study, the ability of PGPR such as Lysinibacillus xylanilyticus, Enterobacter aerogenes, Pseudomonas brenneri and Serratia glossinae to solubilize tri-calcium phosphate and produce IAA and their possession of the nifH gene suggests their potential for use as biofertilizer for crop production.
PGPR promote crop growth by facilitating nutrient uptake though P-solubilization and N2-fixation as well by producing growth-promoting hormones such as IAA (Hayat et al. 2010; Vessey 2003). This increase in crop growth indicates the plant growth- and plant health-promoting traits of these bacterial strains. Enterobacter, Pseudomonas and Bacillus spp. are used as PGPR (Çakmakçi et al. 2007; Vessey 2003). These PGPR have the potential to fix N2, solubilize tri-calcium phosphates and produce phytohormones and when used as bioinoculants increase crop yield (Khan et al. 2009). These PGPR are also widely used as PHPR to reduce plant diseases and produce antibiotics (Herman et al. 2008). Along possessing PGP traits, many species of Serratia have antifungal characters and have been shown to enhance the growth and yield of maize, rape seed, legumes and sorghum (Kishore et al. 2005). Similarly, Enterobacter, when used as biofertilizer, can enhance growth in rice and maize (Kim et al. 1998). Co-inoculation of P-solubilizing bacteria and PGPR can reduce the application of phosphorus fertilizers by 50 % without affecting crop yield (Yazdani et al. 2009).
In conclusion, we isolated 14 bacterial strains from the compost of FVW and purified, characterized and identified these strains by 16S rRNA gene sequencing. These bacterial strains were identified as belonging to the genera Bacillus, Lysinibacillus, Lysobacter, Staphylococcus, Enterobacter, Pseudomonas and Serratia. Although all of the bacterial strains isolated possessed PGP traits, the Pseudomonas and Enterobacter strains appear to be potential PGPR based on their maximum tri-calcium phosphate solubilization and IAA production, as well as the presence of the nifH gene, which codes for the nitrogenase reductase enzyme involved in N2-fixation. Higher wheat growth was observed after wheat seeds were treated with these PGPR. Based on these results, we suggest that these PGPR are potential bio-alternatives to inorganic fertilizers and could serve as bio-fertilizer.
References
Ahmed I, Yokota A, Fujiwara T (2007) A novel highly boron tolerant bacterium, Bacillus boronophilus sp. Nov., isolated from soil that require boron for its growth. Extremophiles 11:217–224
Akhtar MJ, Asghar HM, Shahzad K, Arshad M (2009) Role of plant growth promoting rhizobacteria applied in combination with compost and mineral fertilizers to improve growth and yield of wheat (Triticum aestivum). Pak J Bot 41:381–390
Banerjee MR, Yesmin L, Vessey JK (2006) Plant growth promoting rhizobacteria as biofertilizers and biopesticides. In: Rai MK (ed) Handbook of microbial biofertilizers. Food Products Press, New York, pp 137–181
Brick JM, Bostock RM, Silverstone SE (1991) Rapid in situ assay for indole acetic acid production by bacteria immobilized on nitrocellulose membrane. Appl Environ Microbiol 57:535–538
Çakmakçi R, Donmez F, Aydin A, Sahin F (2006) Growth promotion of plants by plant growth promoting rhizobacteria under greenhouse and two different field soil conditions. Soil Biol Biochem 38:1482–1487
Çakmakçi R, Erat M, Erdoğan ÜG, Dönmez MF (2007) The influence of PGPR on growth parameters, antioxidant and pentose phosphate oxidative cycle enzymes in wheat and spinach plants. J Plant Nutr Soil Sci 170:288–295
Cappuccino JC, Sherman N (2005) Microbiology. A laboratory manual. Wesley Publ. Co., New York
Chun J, Lee JH, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Cowan ST, Steel KJ (2004) Manual for the identification of medical bacteria, 3rd edn. Cambridge University Press, London
Gaur AC (1990) Physiological functions of phosphate solubilizing micro-organisms. In: Gaur AC (ed) Phosphate solubilizing micro-organisms as biofertilizers. Omega Scientific Publishers, New Delhi, pp 16–72
Goldstein AH (2007) Future trends in research on microbial phosphate solubilization: one hundred years of insolubility. In: Velázquez E, Rodríguez-Barrueco C (eds) First Int Meeting on Microbial Phosphate Solubilization. Springer, Dordrecht, pp 91–96
Hameeda B, Rupela OP, Reddy G, Satyavani K (2006) Application of plant growth-promoting bacteria associated with composts and macrofauna for growth promotion of Pearl millet (Pennisetum glaucum L.). Biol Fertil Soils 43:221–227
Hayat R, Ali S, Amara U, Khalid R, Ahmed I (2010) Soil beneficial bacteria and their role in plant growth promotion: a review. Ann Microbiol 60:579–598
Hayat R, Ahmed I, Sheirdil RI (2012) An overview of plant growth promoting rhizobacteria (PGPR) for sustainable agriculture. In: Ashraf M, Öztürk M, Ahmad MSA, Aksoy A (eds) Crop Production for Agricultural Improvement, part 3. Springer, Dordrecht, pp 557–579
Herman MAB, Nault BA, Smart CD (2008) Effects of plant growth promoting rhizobacteria on bell pepper production and green peach aphid infestation in New York. Crop Prot 27:996–1002
Illmer P, Schinner F (1992) Solubilization of inorganic phosphates by microorganisms isolated from forest soil. Soil Biol Biochem 24:389–395
Katsivela E, Bonse D, Kruger A, Strompl C, Livingston A, Ittich RM (1999) An extractive membrane biofilm reactor for degradation of 1,3-dichloropropene in industrial wastewater. Appl Microbiol Biotechnol 52:853–862
Khan MS, Zaidi A, Wani PA, Ahemad M, Oves M (2009) Functional diversity among plant growth-promoting rhizobacteria. In: Khan MS, Zaidi A, Musarrat J (eds) Microbial strategies for crop improvement. Springer, Berlin Heidelberg, pp 105–132
Kim KY, Jordan D, McDonald GA (1998) Enterobacter agglomerans, phosphate solubilizing bacteria, and microbial activity in soil: effect of carbon sources. Soil Biol Biochem 30:995–1003
Kishore GK, Pande S, Podile AR (2005) Biological control of late leaf spot of peanut (Arachis hypogaea L.) with chitinolytic bacteria. Phytopathology 95:1157–1165
Manici LM, Caputo F, Babibi V (2004) Effect of green manure on Pythium spp. population and microbial communities in intensive cropping systems. Plant Soil 263:133–142
Pikovskaya RI (1948) Mobilization of phosphorus in soil connection with the vital activity of some microbial species. Microbiolgiya 17:362–370
Rodríguez H, Fraga R, Gonzalez T, Bashan T (2006) Genetics of phosphate solubilization and its potential applications for improving plant growth-promoting bacteria. Plant Soil 287:15–21
Soultanpour PN, Workman SM (1979) Modification of the NH4HCO3-DTPA soil test to omit carbon black. Commun Soil Sci Plant Anal 10:1411–1420
Steel RGD, Torrie JH, Boston MA (1997) Principles and procedures of statistics: a biometrical approach. McGraw–Hill, New York
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tye AJ, Siu FK, Leung TY, Lim BL (2002) Molecular cloning and the biochemical characterization of two novel phytases from Bacillus subtilis 168 and Bacillus licheniformis. Appl Microbiol Biotechnol 59:190–197
Vessey JK (2003) Plant growth promoting rhizobacteria as biofertilizers. Plant Soil 255:571–586
Watanabe FS, Olsen SR (1965) Test of an ascorbic acid method for determining phosphorus in water and NaHCO3 extracts from soil. Soil Sci Soc Am Proc 29:677–678
Yazdani M, Bahmanyar MA, Pirdashti H, Esmaili MA (2009) Effect of Phosphate solubilization microorganisms (PSM) and plant growth promoting rhizobacteria (PGPR) on yield and yield components of Corn (Zea mays L.). Proc World Acad Sci Eng Tech 37:90–92
Yu X, Liu X, Zhu TH, Liu GH, Mao C (2011) Isolation and characterization of phosphate-solubilizing bacteria from walnut and their effect on growth and phosphorus mobilization. Biol Fert Soils 47:437–446
Acknowledgments
Financial assistance from PMAS-Arid Agriculture University, Rawalpindi under the project entitled Preparation of Biofertilizer for Improving Legume N2-Fixation and Soil Health is highly acknowledged. DNA studies were carried out at the Plant Biotechnology Program, National Institute for Genomics and Advanced Biotechnology (NIGAB), National Agricultural Research Center (NARC), Islamabad, Pakistan. Bacterial strains were commercially sequenced with funding from the Pakistan Science Foundation project no. PSF-UAAR/Agr-374.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hayat, R., Sheirdil, R.A., Iftikhar-ul-Hassan, M. et al. Characterization and identification of compost bacteria based on 16S rRNA gene sequencing. Ann Microbiol 63, 905–912 (2013). https://doi.org/10.1007/s13213-012-0542-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-012-0542-4