Abstract
Expansins are plant cell-wall loosening proteins involved in cell enlargement, adaptive responses to environmental stimuli, and various developmental processes. Although expansins have been characterized in many plant species, little is reported on this family in watermelon. In this study, 30 expansin genes in the watermelon genome (ClEXPs) were identified. These genes which were divided into four subfamilies (7 ClEXLAs, 2 ClEXLBs, 18 ClEXPAs, and 3 ClEXPBs) are unevenly distribute on 10 of 11 watermelon chromosomes. Chromosome mapping suggested that tandem duplication events may have played important roles in the expanding of watermelon expansins. Gene structure and motif identification revealed that same subfamily and subgroup have conserved gene structure and motif. Detection of cis-acting elements revealed that ClEXPs gene promoter regions were enriched with light-responsive elements, hormone-responsive, environmental stress-related, and development-related elements. Expression patterns of ClEXPs were investigated by qRT-PCR. The results showed that expression patterns of 15 ClEXP genes differed in three tissues. Through our own and public RNA-seq analysis, we found that ClEXPs had different expression patterns in fruit flesh, fruit rind, and seed at various developmental stages, and most of ClEXPs were highly responsive to abiotic and biotic stresses. Remarkably, 7 ClEXPs (ClEXLA1, ClEXLA6, ClEXLB1, ClEXLB2, ClEXPA5, ClEXPA10, and ClEXPA16) exhibited positive response to at least three kinds of stresses, suggesting that they might play important roles in the crosstalk of stress signal pathways. The results of this study provide useful insights for the functional identification of expansin gene family in watermelon.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant cell walls are dynamic structures that determine and maintain the size and the shape of the cells and serve as a protective barrier. The cell walls are highly complex structures composed mainly of polysaccharides that vary in structure, function, and abundance (Santiago et al. 2018). Expansin gene family is important regulators of extension and relaxation of cells in growing tissues through a non-enzymatic activity (Cosgrove 2015). They bind to glucan-coated cellulose in the cell wall, causing a reversible disruption of hydrogen bonds between cellulose microfibrils and the glucan matrix to loosen the cell wall (Sampedro and Cosgrove 2005). Typical expansins in plants have 250–275 amino acids and comprise of two domains preceded by a signal peptide. Domain 1 is homologous to the catalytic domain of proteins in the glycoside hydrolase family 45 (GH45); domain 2 is homologous to group-2 grass pollen allergens, which bind to cell-wall polysaccharides. Expansins in plants is divided into four families, namely α-expansin (EXPA), β-expansin (EXPB), expansin-like A (EXLA), and expansin-like B (EXLB) (Kende et al. 2004). EXPA and EXPB families are the two major subfamilies. EXPAs were identified as mediators of acid-induced wall loosening, while EXPBs include a subset of well-studied proteins known as group-1 grass pollen allergens as well as a larger set of proteins about which we know very little (Sampedro et al. 2015). EXLA and EXLB families are identified only by their conserved amino acid sequences, with far less known about their precise functions. They play important roles in many plant growth and developmental processes by modifying and elongating the cell-wall structure, such as seed development and germination (Chen and Bradford 2000; Chen et al. 2001), root elongation and growth (Lee et al. 2003; Che et al. 2016), stem growth and internode elongation (Takada et al. 1997; Wang et al. 2011), leaf formation and development (Reinhardt et al. 1998; Kuluev et al. 2017), flower development, opening and fertilization (Harada et al. 2011; Castillo et al. 2018), fruit development, ripening, firmness, and weight (Dotto et al. 2006; Xie et al. 2009). These genes could also been associated with nutrient uptake and efficiency (Zhou et al. 2014), abiotic stress (Noh et al. 2009; Zhou et al. 2011), and biotic stress tolerance (Song et al. 2017). Expansins in plant breeding programs present an opportunity to improve crops in various aspects. These include but are not limited to improving germination, leaf size, fruit growth, and ripening and tolerance to abiotic and biotic stresses (Marowa et al. 2016).
Some studies characterized expansins in plant genomes of angiosperms (Arabidopsis, poplar, grape, soybean, apple, Chinese cabbage, rice, and maize) and nonflowering plants (Selaginella moellendorffii and Physcomitrella patens) (Cosgrove 2015). However, systematic analysis of expansins in watermelon has not been reported. Watermelon is an important cucurbit crop grown throughout the world, and the annual world production of watermelon is about 118 million tons (http://faostat.fao.org/). In the Cucurbitaceae family, the genome of watermelon has been sequenced after cucumber and melon (Wechter et al. 2008). The availability of these sequences and large-scale transcriptome data provide an excellent opportunity to investigate watermelon in the field of molecular biology. In this study, we identified the expansin genes in the watermelon genome, analyzed gene structures and phylogenetic relationships, and detected the cis-acting elements in ClEXP gene promoter regions. Publically available and our own RNA-seq dataset were employed to study the expression of the ClEXP genes response to fruit and seed development, and abiotic and biotic stresses. QRT-PCR was used to study the ClEXPs’ tissue expression patterns. Our findings should pave a way for further functional researches on expansin gene family in watermelon.
Materials and methods
Genome-wide identification of expansin genes in watermelon (ClEXPs)
First, expansin protein sequences in Arabidopsis (AtEXPs) (http://www.arabidopsis.org/) were used as tblastn (E values ≤ 1 × 10−5) against the watermelon reference genome sequence (ftp://cucurbitgenomics.org/pub/cucurbit/genome/watermelon/97103/v1/watermelon_v1.genome.gz). The highly matched sequences were reorganized and merged. Then, all protein sequences of putative expansin genes were scanned for conserved domain by InterProScan (http://www.ebi.ac.uk/interpro/). The sequences lacking expansin domains were rejected. Finally, we further verified these sequences using EST database in NCBI (http://www.ncbi.nlm.nih.gov). Identification method of expansin genes in cucumber (CsEXPs) and melon (CmEXPs) was similar to that in watermelon.
Protein properties, chromosomal location, and gene structure analysis
ExPaSy (http://expasy.org/) was used to compute the molecular weights (MWs) and isoelectric point (pI). All expansin genes’ chromosomal locations were obtained from Cucurit Genomics Database (CuGenDB, http://cucurbitgenomics.org/) and then were mapped to the chromosomes by MapInspect Version 1.0 (http://www.softsea.com/review/MapInspect.html) software. The exon/intron structures of ClEXPs were generated using GSDS (http://gsds.cbi.pku.edu.cn) through aligning the coding domain sequences (CDS) and DNA sequences of expansin genes. The conserved motifs were identified using the MEME program (http://meme.sdsc.edu/meme/). The maximum motif search value was set at 15 and an optimum motif width of 10–100 amino acid residues. Other parameters are default.
Sequence alignment, and phylogenetic and homology analysis
The full-length amino acid sequences were aligned with ClustalW (http://www.ebi.ac.uk/Tools/msa/clustalw2/) and a phylogenetic tree was constructed using the neighbour-joining method with 1000 bootstrap replicates by MEGA 7.0 software (Kumar et al. 2016). The expansin genes in watermelon and Arabidopsis were clustered using the entire protein sequences by OrthoMCL software (http://orthomcl.org/orthomcl/; E values < 1 × 10−5) and the homologous relationships were drawn using Circos software (http://circos.ca/).
Analysis of promoter regions cis-acting elements
The promoter regions of the watermelon expansin genes were identified by searching the watermelon genome database, and were analyzed using PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) to predict cis-acting elements.
Expression patterns analysis of watermelon expansin gene response to low light
We study watermelon expansin gene expression pattern response to low light stress through RNA-seq technology. The study was conducted in plastic greenhouse at Luhe production base at Jiangsu Academy of Agricultural Sciences. Watermelon variety ‘Sumi No.8’ (Institute of Vegetable Crops, Jiangsu Academy of Agricultural Sciences, Nanjing, China) was as the plant materials. Low light (yin) treatment was carried out 7 days before flowering. The light intensity under the sunshade net was 50% less than that under natural conditions (CK). Flesh of watermelon fruit center under yin and CK at 0, 3, 9, and 15 days after pollination (DAP) was harvested for RNA-seq. The 24 cDNA library was sequenced on the Illumina sequencing platform (Illumina HiSeqTM2500) using the paired-end technology. The raw sequence data (in FastQ format) have been submitted to the National Center for Biotechnology Information (NCBI) Sequence Read Archive databases with the study accession (SRP243255). The levels of gene expression were estimated by FPKM values. The raw FPKM values were first normalized by logarithmic method and a heat map was constructed to show the different expression profiles by the Heml software (version 1.0, http://hemi.biocuckoo.org/).
Expression patterns analysis of watermelon expansin gene response to watermelon fruit and seed development and other stress
Cucurbit Genomics Database (CuGenDB, http://cucurbitgenomics.org/) have collected all available RNA-Seq data from NCBI Sequence Read Archive (SRA, https://www.ncbi.nlm.nih.gov/sra) for the cucurbit species that have available reference genome sequences. In addition, a unified pipeline for RNA-Seq data processing and analysis has been applied to these RNA-Seq datasets. Raw RNA-Seq reads are processed to remove adaptor and low-quality sequences, etc. Raw counts are derived for each predicted gene model and normalized to FPKM. Gene expression profiles for a specific gene can be accessed under the ‘Gene Expression’ section of the gene feature page, where after selecting an RNA-Seq project. The average expression values and the standard deviation from biological replicates are calculated for each gene and loaded into CuGenDB using the ‘RNA-Seq’ extension module (Zheng et al. 2018). Therefore, transcriptome sequencing data for watermelon fruit development (SRP012849), seed development (PRJNA319011), root or leaf response to osmotic stress (PRJNA326331), cold stress (PRJNA328189) and drought stress (PRJNA454040), and fruit flesh response to cucumber green mottle mosaic virus (CGMMV) (PRJNA389184) were obtained from CuGenDB using the identified ClEXPs ID.
The levels of gene expression were estimated by FPKM values. The FPKM values were first normalized by logarithmic method and a heat map was constructed to show the different expression profiles by the Heml software (version 1.0, http://hemi.biocuckoo.org/).
Plant materials, RNA isolation, and qRT-PCR analysis
The watermelon variety ‘Sumi No.8’ (Institute of Vegetable Crops, Jiangsu Academy of Agricultural Sciences, Nanjing, China) was used as the plant materials to investigate the expression profiles of expansin genes in different tissues. Root, stem, and leaf samples were collected 1 month after planting and stored at − 80 °C.
RNA was extracted from triplicated biological replicates of the aforementioned samples using RNAprep Pure Plant Kit (TIAGEN, China), and treated with DNaseI (Transgen). The quality of RNAs was examined by agarose gel electrophoresis. The RNA samples were reverse transcribed to cDNAs with TranScript® One-step gDNA Removal and cDNA Synthesis SuperMix (Transgen). The cDNA were then diluted ten times and stored at − 20 °C for the subsequent qRT-PCR.
The watermelon actin gene (Cla007792) was used as an internal control. The primers for 15 randomly selected expansin genes in watermelon were designed by Primer Premier 5.0 software. Primer sequences were listed in Table S1. qRT-PCR assays were performed with three replicates. The SYBR Premix Ex Taq II reagent (Takara, Japan) with SYBR Green I as the fluorescent dye was used for qPCR on an ABI 7300 real-time PCR system (Applied Biosystems, Foster City, CA, USA). Each reaction contained 10 μL 2 × SYBR Premix Ex TaqII Reagent, 2.0 μL cDNA sample, and 500 nM gene-specific primers in a final volume of 20 μL. QPCR conditions were set at 95 °C pre-denaturation for 3 min and followed by 40 cycles of 95 °C denaturation for 15 s, 60 °C annealing for 30 s, and 72 °C extension for 15 s. All the data were analyzed using the \(2^{{ - \Delta \Delta C_{\text{t}} }}\) method, where Ct is the cycle threshold measured according to a previous method (Livak and Schmittgen 2001).
Results
Identification of expansin gene family in watermelon
After filtering and removing the sequences lacking expansin conserved domain, 30 expansin genes with two conserved domains in their protein sequences were identified in the watermelon genome. In accordance with the nomenclature methods for members of expansin superfamily proposed by Kende et al. (2004), expansins of watermelon, melon, and cucumber were sequentially named on the basis of family identity and chromosomal position (Table S2). As shown in Table S3, the identified ClEXP genes encoded proteins ranging from 237 (ClEXLA1) to 434 (ClEXPA11) amino acids (aa) and had predicted MWs of 26.39–45.54 kDa. The majority members of ClEXPA (except ClEXPA18), ClEXPB (except ClEXPB2), and ClEXLA (except ClEXLA7) subfamilies had pI values above 7.0, whereas pI values of all members of ClEXLB were below 7.0.
The chromosomal distribution map of ClEXPs is presented in Fig. 1. All ClEXPs were successfully mapped to the 10 watermelon chromosomes. Chr2 contained the maximum number (10) of ClEXPs genes and all ClEXLA genes members are distributed on this chromosome. The two ClEXLB genes are both located on the Chr9, and two ClEXPBs were distributed on Chr2 and Ch5, respectively. ClEXPAs were unevenly distributed on ten watermelon chromosomes.
Sequence alignment and phylogenetic analysis
To obtain detailed information regarding each ClEXPs gene and superfamily, we completed multiple alignments of the deduced protein sequences of 30 ClEXPs genes (Fig. S1). Most of ClEXPs protein consisted of three parts, namely, a 20–30 amino acid signal peptide, domains 1 and domain 2. Only ClEXPA7 lack the signal peptide region. The amino acid sequence of domain 1 was more conserved than that of domain 2. As reported by Li et al. (2002), the EXPA and EXPB subfamilies could be easily distinguished based on two types of insertions. The α-insertions existed only in the EXPA subfamily and consisted of about 14 residues, with most insertions containing four highly conserved residues at the 3′ end, WCNP. The β-insertions were present among EXPB, EXLA, and EXLB members. All EXPBs and the majority of EXPA members (excluding ClEXPA3, ClEXPA4) contained a conserved HFD motif in domain 1. Among the nine expansin-like protein sequences with an incompletely conserved HFD motif, seven were classified into the EXLA subfamily based on the presence of a characteristic EXLA extension at the C-terminus; two were classified into EXLB subfamily.
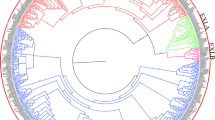
To evaluate evolutionary relationships of expansin proteins, complete protein sequences of expansins from Arabidopsis, cucumber, melon, and watermelon, were subjected to phylogenetic analysis. As shown in Fig. 2, all expansin genes were divided into four major subfamilies: EXPA, EXPB, EXLA, and EXLB. EXPA was the largest subfamily, and EXLB was the smallest. The four expansin gene subfamilies were further divided into 17 subgroups according to the grouping rules used for Arabidopsis expansin gene families (Sampedro and Cosgrove 2005). EXPA-V and EXPA-VI subgroups did not contain any watermelon expansin gene. We found that EXLA-I subgroup, containing 23 members, constituted the largest subgroup, while the smallest subgroup, EXPA-VI, comprised only one gene.
Phylogenetic analysis of AtEXPs, CsEXPs, CmEXPs, and ClEXP. The full-length amino acid sequences were aligned with ClustalW and a phylogenetic tree was constructed using the neighbour-joining method with 1000 bootstrap replicates by MEGA 7.0 software. The subgroups are marked by color bars. Red, green, purple, and mazarine represent melon, watermelon, cucumber, and Arabidopsis expansins, respectively. The phylogenetic tree was generated using the maximum likelihood method in MEGA 7.0. Arabidopsis expansins sequences were retrieved from the databases of Arabidopsis Information Resource (http://www.arabidopsis.org/) and the Expansin Central website (http://www.personal.psu.edu/fsl/ExpCentral/index.html)
Gene structure and putative motif identification
A structural analysis can provide valuable information concerning duplication events, so we analyzed the exon–intron structures (Fig. 3). The number of exons in the ClEXPs family varied from two to five, with members within each subgroup or subfamily having similar exon–intron structures. In the EXPA subfamily, 13 members (72.22%) had three exons; three members (16.67%) had two exons, and one member had four (ClEXPA4) or five (ClEXPA11). The EXPB subfamily had four exons except of ClEXPB2. Members of the ClEXLA and ClEXLB subfamily possessed five exons with the exception of ClEXLA2, ClEXLA4, and ClEXLB2, which contained four.
Phylogenetic relationships and structural analysis of ClEXPs. Introns and exons are represented by gray and green boxes, respectively. Lengths of introns and exons of each expansin gene are displayed proportionally. Gene structures were generated with GSDS 2.0 (http://gsds.cbi.pku.edu.cn/)
We detected conserved motifs in ClEXPs family and found 15 distinct motifs (Fig. 4). Schematic diagrams of these ClEXPs motifs are given in Table 1. Motifs 2 and 10 specified the well-conserved N-terminal domain 1, and motifs 3, 6, 9, and 11 consisted of the well-conserved C-terminal domain 2. At least two of these six motifs were shared among the ClEXPs (Table 1). Genes from the same subfamily, especially within subgroups, were generally characterized by a similar motif type and distribution. Motif 2, 5, and 6 were detected in all ClEXPs, and Motifs 1 and 3 were only detected in the ClEXPA subfamily, but Motif 9, 11 and 15 were found in all ClEXPs, except ClEXPA. These results indicate that the ClEXPA subfamily differs significantly from the other three subfamilies in terms of gene structure and motif composition. Others motifs were unevenly distributed in different clades.
Homology analysis and detection of cis-acting elements in ClEXPs gene promoter regions
Because Arabidopsis is the most important model plant and many of its expansin genes function have been well characterized, we performed a homology analysis of watermelon and Arabidopsis expansin genes to predict the function of ClEXPs (Fig. 5). The analysis revealed the presence of 34 homologous gene pairs between watermelon and Arabidopsis as well as 11 gene pairs in watermelon and 15 in Arabidopsis. All ClEXP genes showed duplication events, and 13 tandem duplications between expansin genes were determined in watermelon with the percentage of 43.33%. These tandem duplications located on chromosome 2 (ClEXLA1, ClEXLA2, ClEXLA3, ClEXLA4, ClEXLA5, ClEXLA6, and ClEXLA7), chromosome 4 (ClEXPA6 and ClEXPA7), chromosome 7 (ClEXPA12 and ClEXPA13), and chromosome 9 (ClEXLB1 and ClEXLB2) (Table S4). However, there have no segmental duplications which were determined.
As detailed in Table 2 and Supplementary Tables S5, cis-acting elements in the promoter regions of ClEXPs are extremely diverse. We detected 91 types of cis-acting elements, 34 with unknown functions. The most abundant elements were light-responsive elements, which contained 24 types (Table S5). Each promoter possessed 3–19 types, which suggests that ClEXPs are differentially regulated by light (Table 2). Twelve types of hormone-responsive elements were detected. The CGTCA-motif and TGACG-motif, which both involved in methyl jasmonate (MeJA) response, were also the most abundant hormone-responsive elements in the 19 ClEXPs gene promoters. One AuxRR-core, one AuxRE, and one TGA box were found in the promoter regions of ClEXPA, ClEXLA, and ClEXLB genes, respectively, which indicate that auxin may regulate these genes. Moreover, six types of environmental stress-related elements were identified in ClEXPs promoters. ARE and WUN-motif elements were present in over half of the identified 30 watermelon expansin genes. The fourth important type of cis-acting elements in the promoter regions of ClEXPs was development-related elements. The O2 site was the most abundant regulatory element of the 7 types. ClEXLA1, ClEXPA9, and ClEXPB1 contained MSA-like involved in cell cycle regulation, which suggests that these genes have important function in cell division and cell enlargement. (Table S5).
Expression profiles of ClEXPs in fruit and seed developmental stages
We examine the expression profiles of ClEXPs genes in fruit and seed at multiple developmental stages (Fig. 6). 22 ClEXPs were detected in fruit and seed developmental stages, which were divided into three subgroups. Group I showed relatively higher expression levels in fruit flesh and rind at the early developmental stage. Moreover, ClEXPA4 had higher expression levels in all seed developmental stage, ClEXPA10 and ClEXPB1 had higher expression in 31 DAP. Group II have highest expression in seed early developmental stage. Group III have lower expression level in all seed development stages, but higher levels in early stage of fruit flesh and rind development.
Expression profiles of ClEXPs genes in different watermelon fruit (cultivars 97103; SBA number SRP012849) and seed developmental stages (SBA number PRJNA319011). Subdivided into groups (labelled I–III) based on the transcriptome data. The legend represents the logarithmic normalized FPKM. DAP days after pollination, FF fruit flesh, FR fruit rind, WmSeed watermelon seed. Fruit flesh and rind was used as RNA-seq material at 10, 18, 26, and 34 days after pollination (DAP) during watermelon fruit development. Watermelon seed tissue was taken out at 25, 31, 37, and 49 days after pollination for RNA-seq
Expression profiles of ClEXPs under abiotic and biotic stresses
According to our RNA-seq results of watermelon fruit response to low light stress (Table S6), the 27 ClEXPs could be divided into five major clusters (Fig. 7). It is noteworthy that the expression of ClEXPA4, ClEXPB1, ClEXPA16, and ClEXPA12 in group I was significantly induced by low light stress at 0 DAP and then rapidly declined. Gene expression level of group II was significantly induced at 3 DAP, while group III had highest expression level at 9 DAP except for ClEXPA3, other genes in group III were significantly downregulated by low light at 9 DAP. Gene expression levels in group IV and V were repressed by low light stress.
Expression profiles of ClEXPs genes response to low light stress in watermelon (cultivars: Sumi No.8; tissue: fruit flesh). Subdivided into groups (labelled I–V) based on the transcriptome data. The legend represents the logarithmic normalized FPKM. DAP days after pollination. Low light (yin) treatment was carried out 7 days before flowering. The light intensity under the sunshade net was 50% less than that under natural conditions (CK). Flesh of watermelon fruit center under yin and CK at 0, 3, 9, and 15 days after pollination (DAP) were harvested for RNA-seq
After cold stress, 23 ClEXPs divided into seven major clusters (Fig. 8). It is worth noting that the expression level of ClEXPs in group VII (ClEXLB1, ClEXPA16, ClEXPA5, ClEXLA1, ClEXLB2) was significantly induced by cold stress. In group III (ClEXLA7), melatonin treatment can significantly increase the gene expression level under cold stress and normal temperature (CK). Except this two groups, expression of ClEXPs in other groups showed significant down-regulation or no significant change in response to cold stress.
Expression profiles of ClEXPs genes response to cold stress in watermelon (cultivars: Y134; tissue: Leaf; SBA number: PRJNA328189). Subdivided into groups (labelled I–VII) based on the transcriptome data. The legend represents the logarithmic normalized FPKM. MT melatonin treatment, Cold cold stress, Wmleaf watermelon leaf. Y134. Normal temperature was 25/18 °C (day/night); cold stress was 4 °C. Seedlings at the four-leaf stage were sprayed with 150 mM melatonin solution for 3 days, with distilled water used as the control
Under drought stress, the 20 ClEXPs could be divided into nine major clusters (Fig. 9). Expression of group V (ClEXPA4, ClEXPA12) was up-regulated by drought stress in drought-resistant variety M20 and drought-sensitive varieties Y34, while expression pattern in group III was contrary. In group VI, gene expression in M20 was lower under stress treatment, while that in Y34 drought stress can significantly induced the gene expression. ClEXPs in group I were induced after 4 days and declined rapidly after 8 days under drought stress in M20, while gene expression patterns in Y34 were opposite. Gene expression levels in group II and IV were lower and not affected by drought stress in M20, while expression level in Y34 was highest at 4 days and drought stress can significantly decrease expression level.
Expression profiles of ClEXPs’ genes response to drought stress in watermelon (cultivars: Y34 and M20; tissue: Leaf; SBA number: PRJNA454040). Subdivided into groups (labelled I–VI) based on the transcriptome data. The legend represents the logarithmic normalized FPKM. Y34 watermelon Y34, M20 watermelon M20, d days after drought stress or control treatment. Y34 was M20 was drought tolerance material and Y34 was drought-sensitive material
The 29 ClEXPs genes in response to osmotic stress were divided into two major clusters (Fig. 10). Group I have 10 ClEXPs and expression of these genes showed increasing pattern upon osmotic stress. The gene expression patterns of group II were opposite, which decreased significantly after osmotic stress.
Expression profiles of ClEXPs genes response to osmotic stress in watermelon (cultivars: M08; tissue: root; SBA number: PRJNA326331). Subdivided into groups (labelled I–II) based on the transcriptome data. The legend represents the logarithmic normalized FPKM. OS osmotic. M08 was relatively high tolerance to water deficits. The root samples were harvested at 6 h after 20% polyethylene glycol (PEG) 6000 treatment and untreated samples were used as controls
Through comparative analyses of ClEXPs expression profiles of watermelon fruits inoculated with cucumber green mottle mosaic virus (CGMMV) or non-inoculated, we found 19 ClEXPs which can be divided into two groups (Fig. 11). Group I has 5 ClEXPs and showed rapid and negative response, while group II showed rapid and positive response.
Expression profiles of ClEXPs genes response to CGMMV stress in watermelon (cultivars: Jingxin No.3; tissue: Fruit flesh; SBA number: PRJNA389184). Subdivided into groups (labelled I–II) based on the transcriptome data. The legend represents the logarithmic normalized FPKM. CGMMV cucumber green mottle mosaic virus
Expression profiles of expansin genes in different watermelon tissues analyzed by qRT-PCR
To comprehensively understand the physiological functions of watermelon expansin genes, we analyzed the expressions of 15 randomly selected expansin genes in watermelon under normal growth conditions in three different tissues by qRT-PCR. As shown in Fig. 12, expression levels of the 15 ClEXP genes differed in three tissues, indicating their involvement in the development of the three tissues. ClEXLA7 was dominantly expressed in root, and ClEXLA1 and ClEXLA5 have higher expression level in root than in leaf and stem, but ClEXLA2 and ClEXLA6 showed high expression in leaf. ClEXLB2 was notably high in root. ClEXPA5, ClEXPA6, and ClEXPA11 were notably high in root, but ClEXPA4, ClEXPA9, ClEXPA12, ClEXPA15, ClEXPA16, and ClEXPA18 showed high expressions in leaf. These results further highlighted that expansin genes were involved in watermelon plant growth.
Discussion
In this study, a total of 30 watermelon expansin genes containing full-length ORFs and two conserved domains, DPBB_1 and Pollen_allerg_1, were first identified in watermelon genome through genome-wide analysis. Similar to other plants, the 30 watermelon expansins were divided into four subfamilies, ClEXPA, ClEXPB, ClEXLA, and ClEXLB. They clustered together with genes of the same subfamily from other plants rather than genes of the same species from different subfamilies, suggesting that their ancestors differentiated before the divergence of different plant species. Seader et al. (2016) estimated that there were 12–13 EXPA genes, 2 EXPB genes, 1 EXLA gene, and 2 EXLB genes in the last common ancestor of all angiosperms. In our study, the four watermelon expansin subfamilies ClEXPA, ClEXPB, ClEXLA, and ClEXLB were accordingly divided into 9, 2, 1, and 2 subgroups with each group of genes deriving from a common ancestor by frequent gene duplication. The missing genes belonged to the other three subgroups in the ClEXPA subfamily might span different genome sequences that have not been scaffolded and, therefore, cannot be identified or be removed in our sequence analysis, and the results was similar to other studies (Ding et al. 2016). The disappearance of descendents from these ancestors could also attribute to gene deaths which were observed in Arabidopsis and rice as well (Sampedro and Cosgrove 2005). Through analysis and comparison of the sizes of expansin subfamilies in Arabidopsis, melon, cucumber, and watermelon, we found an uneven distribution of each gene subfamily among species. For example, Arabidopsis has 26 EXPA genes, 6 EXPB genes, 3 EXLA genes, and 1 EXLB genes, which contain more EXPA and EXPB members than Cucurbitaceae family (i.e., watermelon, melon, and cucumber). In addition, these four eudicots species have less EXPB members than a monocot rice, which correlate to the previous studies (Li et al. 2016; Hou et al. 2018) and possibly due to the differences in their cell-wall composition and their evolutionary differences (Cosgrove et al. 2002; Dal Santo et al. 2013). Watermelon, melon, and cucumber have more than five members in EXLA subfamily, but Arabidopsis have only two members in EXLA subfamily. Plants retaining certain kind of genes with larger subfamily size during the long evolutionary time were to increase its adaption to certain functions and environments (Ding et al. 2016). Therefore, the additional ClEXLA genes might have special functions in watermelon, which need to be validated. We also found that the size of EXPA was the biggest and the size of EXPB and EXLB were equal in Cucurbitaceae, which was in accordance with the observation in Solanaceae (Lu et al. 2016). Gene duplication events are considered to be one of the evolutionary forces in genome evolution, providing materials for generation of new genes and development of new functions (Kong et al. 2007). Segmental and tandem duplications are considered to be the two main causes of gene family expansion in plants (Cannon et al. 2004), the impacts of which on the expansion of watermelon expansin gene family were focused in our study. Our study showed that 43.33% ClEXPs have tandem duplication events. Therefore, our results indicate that tandem duplication events may be one of the main reasons for ClEXP gene family expansion. Phylogenetic analysis revealed that the majority of watermelon expansin genes show more closely related to melon and cucumber expansin genes than those of Arabidopsis. The reason was that there was a set of ancestral expansin genes corresponding to each subfamily predating the divergence of monocot and eudicot (Valdivia et al. 2007). Within the same phylogenetic subfamily and even subgroup, members mostly have a conserved gene structure, thus confirming their close evolutionary relationship and classification (Zhang et al. 2014).
Gene expression pattern can provide important clues for gene function, which are believed to be associated with the divergence of the promoter regions (Ding et al. 2016; Hou et al. 2018), so we used the PlantCARE program to analyze cis-acting elements in all ClEXP promoter regions. The four classes of cis-acting elements, namely, development-related elements, environmental stress-related elements, hormone-responsive elements, and light-responsive elements, are significantly abundant in ClEXPs. In plants, both environmental and internal factors can affect the expression patterns of expansin genes, which enable them to participate in various developmental processes conjectured to be regulated by the corresponding cis-acting elements contained within them (Hou et al. 2018).
Plant hormones and environmental stress can regulate expansin gene expression levels, which have been reported in many plants, such as TaEXPB23 in wheat, was induced by exogenous MeJA and salt stress, but suppressed by exogenous gibberellins, ethylene, indole-3-acetic acid, and α-naphthlcetic acid (Han et al. 2012). Variations in number and type of cis-elements associated with each ClEXPs indicate that the functions of these genes are differentially regulated by these signals and have varied functional roles in many developmental process related to cell-wall modification.
According to the results of RNA-seq data and qRT-PCR analyses, ClEXPs are expressed in various tissues and at many developmental processes, which indicated their possible involvement in certain tissues and developmental stages. The expression levels of ClEXLB2, ClEXLB1, and ClEXPA4 were relatively higher in fruit flesh, fruit rind, and seed at all stages, respectively, which suggested that these genes might be important for fruit flesh, fruit rind, and seed development. Watermelon fruits develop and mature rapidly, and early fruit development involves rapid cell division, followed by a long phase of cell expansion to form large vacuolated cells that make up the flesh of watermelon fruits (Wechter et al. 2008). Most ClEXPs have higher expression levels in early stages of fruit flesh, fruit rind, and seed shows that ClEXPs play important role in fruit and seed early developmental stage. And different ClEXPs show that different tissue expression patterns, such as ClEXLA7, were dominantly expressed in root, while ClEXLA2 and ClEXLA6 showed high expression in leaf. The results show that different ClEXPs play important roles in the growth and development of certain tissues. Expansins are likely to be involved in physiological adaptation in response to abiotic and biotic stresses (Kuluev et al. 2016; Song et al. 2017). Our results of the expression patterns of ClEXPs response to abiotic and biotic stresses are also similar to the previous studies. Notably, seven ClEXPs (ClEXLA1, ClEXLA6, ClEXLB1, ClEXLB2, ClEXPA5, ClEXPA10, and ClEXPA16) exhibited positive response to at least three kinds of stresses, implying their involvement in the crosstalk of stress signal pathways.
Comprehensive analysis of phylogenetic, homologous relationship and gene expression profiles can provide important clues for gene function. Based on our phylogenetic tree, ClEXPA9 and AtEXPA10 belong to the EXPA-I subgroup, and our homology analysis revealed that ClEXPA9 is homologous to AtEXPA10. AtEXPA10 influences the sizes of vegetative organs (Kuluev et al. 2012), our transcriptome and qRT-PCR analysis also showed ClEXPA9 have higher expression level in the early developmental stage of fruit flesh, fruit rind, seed, and in leaf.
Conclusions
In the current study, we presented a genome-wide analysis of expansin gene family in the watermelon genome and identified 18 ClEXPAs, 3 ClEXPBs, 7 ClEXLAs, and 2 ClEXLBs, respectively. The four subfamilies were further grouped into 9, 2, 1, and 2 subgroups which were derived from 14 common ancestors by gene duplications. The ClEXPs exhibited specific characteristics in terms of exon–intron structural, amino acid sequences, or protein motif composition within subfamilies or subgroups. Tandem duplication events might have contributed to the expansion of ClEXPs. Each expansin gene contained a number of cis-acting elements in its 1.5 kb upstream region, suggesting that its expression was regulated by various internal or environmental factors, such as plant hormones, and light and environmental stresses, thus participating in watermelon development and resistance to stresses. Here, we showed a global expression landscape of ClEXPs in response to development and various stress. The expression patterns of the ClEXPs were also studied using our and public RNA-seq and qRT-PCR analysis, which revealed that most ClEXPs are expressed in various tissues and at many developmental processes, which indicated their possible involvement in certain tissues and developmental stages. In addition, here are seven ClEXPs (ClEXLA1, ClEXLA6, ClEXLB1, ClEXLB2, ClEXPA5, ClEXPA10, and ClEXPA16), exhibited positive response to at least three kinds of stresses, implying their involvement in the crosstalk of stress signal pathways. Our results provide valuable information for further elucidation of the evolution and divergence of plant expansin genes. Results of this study may also aid understanding of the biological function of ClEXPs and molecular basis of many watermelon important agriculturally traits.
References
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10
Castillo FM, Canales J, Claude A, Calderini DF (2018) Expansin genes expression in growing ovaries and grains of sunflower are tissue-specific and associate with final grain weight. BMC Plant Biol. https://doi.org/10.1186/s12870-018-1535-7
Che J, Yamaji N, Shen RF, Ma JF (2016) An Al-inducible expansin gene, OsEXPA10 is involved in root cell elongation of rice. Plant J 88:132–142. https://doi.org/10.1111/tpj.13237
Chen F, Bradford KJ (2000) Expression of an expansin is associated with endosperm weakening during tomato seed germination. Plant Physiol 124:1265–1274. https://doi.org/10.1104/pp.124.3.1265
Chen F, Dahal P, Bradford KJ (2001) Two tomato expansin genes show divergent expression and localization in embryos during seed development and germination. Plant Physiol 127:928–936. https://doi.org/10.1104/pp.127.3.928
Cosgrove DJ (2015) Plant expansins: diversity and interactions with plant cell walls. Curr Opin Plant Biol 25:162–172
Cosgrove DJ, Li LC, Cho HT, Hoffmann-Benning S, Moore RC, Blecker D (2002) The growing world of expansins. Plant Cell Physiol 43:1436–1444. https://doi.org/10.1093/pcp/pcf180
Dal Santo S, Vannozzi A, Tornielli GB, Fasoli M, Venturini L, Pezzotti M, Zenoni S (2013) Genome-wide analysis of the expansin gene superfamily reveals grapevine-specific structural and functional characteristics. PLoS ONE 8:e62206. https://doi.org/10.1371/journal.pone.0062206
Ding AM, Marowa P, Kong YZ (2016) Genome-wide identification of the expansin gene family in tobacco (Nicotiana tabacum). Mol Genet Genomics 291:1891–1907. https://doi.org/10.1007/s00438-016-1226-8
Dotto MC, Martinez GA, Civello PM (2006) Expression of expansin genes in strawberry varieties with contrasting fruit firmness. Plant Physiol Biochem 44:301–307. https://doi.org/10.1016/j.plaphy.2006.06.008
Han YY, Li AX, Li F, Zhao MR, Wang W (2012) Characterization of a wheat (Triticum aestivum L.) expansin gene, TaEXPB23, involved in the abiotic stress response and phytohormone regulation. Plant Physiol Biochem 54:49–58. https://doi.org/10.1016/j.plaphy.2012.02.007
Harada T, Torii Y, Morita S, Onodera R, Hara Y, Yokoyama R, Nishitani K, Satoh S (2011) Cloning, characterization, and expression of xyloglucan endotransglucosylase/hydrolase and expansin genes associated with petal growth and development during carnation flower opening. J Exp Bot 62:815–823. https://doi.org/10.1093/jxb/erq319
Hou L, Zhang Z, Dou S, Zhang Y, Pang X, Li Y (2018) Genome-wide identification, characterization, and expression analysis of the expansin gene family in Chinese jujube (Ziziphus jujuba Mill.). Planta 249:815–829. https://doi.org/10.1007/s00425-018-3020-9
Kende H, Bradford KJ, Brummell DA, Cho HT, Cosgrove DJ, Fleming AJ, Gehring C, Lee Y, McQueen-Mason S, Rose JKC, Voesenek LACJ (2004) Nomenclature for members of the expansin superfamily of genes and proteins. Plant Mol Biol 55:311–314. https://doi.org/10.1007/s11103-004-0158-6
Kong H, Landherr LL, Frohlich MW, Leebens-Mack J, Ma H, dePamphilis CW (2007) Patterns of gene duplication in the plant SKP1 gene family in angiosperms: evidence for multiple mechanisms of rapid gene birth. Plant J 50:873–885. https://doi.org/10.1111/j.1365-313X.2007.03097.x
Kuluev BR, Knyazev AB, Lebedev YP, Chemeris AV (2012) Morphological and physiological characteristics of transgenic tobacco plants expressing expansin genes: AtEXP10 from Arabidopsis and PnEXPA1 from poplar. Russ J Plant Physiol 59:97–104. https://doi.org/10.1134/s1021443712010128
Kuluev B, Avalbaev A, Mikhaylova E, Nikonorov Y, Berezhneva Z, Chemeris A (2016) Expression profiles and hormonal regulation of tobacco expansin genes and their involvement in abiotic stress response. J Plant Physiol 206:1–12. https://doi.org/10.1016/j.jplph.2016.09.001
Kuluev BR, Knyazev AV, Mikhaylova EV, Chemeris AV (2017) The role of expansin genes PtrEXPA3 and PnEXPA3 in the regulation of leaf growth in poplar. Russ J Genet 53:651–660. https://doi.org/10.1134/s1022795417060084
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lee DK, Ahn JH, Song SK, Choi YD, Lee JS (2003) Expression of an expansin gene is correlated with root elongation in soybean. Plant Physiol 131:985–997. https://doi.org/10.1104/pp.009902
Li Y, Darley CP, Ongaro V, Fleming A, Schipper O, Baldauf SL, McQueen-Mason SJ (2002) Plant expansins are a complex multigene family with an ancient evolutionary origin. Plant Physiol 128:854–864. https://doi.org/10.1104/pp.010658
Li NN, Pu YY, Gong YC, Yu YL, Ding HF (2016) Genomic location and expression analysis of expansin gene family reveals the evolutionary and functional significance in Triticum aestivum. Genes Genomics 38:1021–1030. https://doi.org/10.1007/s13258-016-0446-y
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lu YG, Liu LF, Wang X, Han ZH, Ouyang B, Zhang JH, Li HX (2016) Genome-wide identification and expression analysis of the expansin gene family in tomato. Mol Genet Genomics 291:597–608. https://doi.org/10.1007/s00438-015-1133-4
Marowa P, Ding AM, Kong YZ (2016) Expansins: roles in plant growth and potential applications in crop improvement. Plant Cell Rep 35:949–965
Noh SA, Park SH, Huh GH, Paek KH, Shin JS, Bae JM (2009) Growth retardation and differential regulation of expansin genes in chilling-stressed sweetpotato. Plant Biotechnol Rep 3:75–85. https://doi.org/10.1007/s11816-008-0077-0
Reinhardt D, Wittwer F, Mandel T, Kuhlemeier C (1998) Localized upregulation of a new expansin gene predicts the site of leaf formation in the tomato meristem. Plant Cell 10:1427–1437
Sampedro J, Cosgrove DJ (2005) The expansin superfamily. Genome Biol. https://doi.org/10.1186/gb-2005-6-12-242
Sampedro J, Guttman M, Li LC, Cosgrove DJ (2015) Evolutionary divergence of beta-expansin structure and function in grasses parallels emergence of distinctive primary cell wall traits. Plant J 81:108–120. https://doi.org/10.1111/tpj.12715
Santiago TR et al (2018) Genome-wide identification, characterization and expression profile analysis of expansins gene family in sugarcane (Saccharum spp.). PLos One. https://doi.org/10.1371/journal.pone.0191081
Seader VH, Thornsberry JM, Carey RE (2016) Utility of the Amborella trichopoda expansin superfamily in elucidating the history of angiosperm expansins. J Plant Res 129:199–207
Song YL et al (2017) Effect of endophytic fungi on the host plant growth, expression of expansin gene and flavonoid content in Tetrastigma hemsleyanum. Diels Gilg ex Diels Plant Soil 417:393–402. https://doi.org/10.1007/s11104-017-3266-1
Takada T, Sakai F, Hayashi T (1997) Expression of gene encoding endo-1,4-beta-glucanase, endoxyloglucan transferase and expansin in elongating pea stems. Plant Physiol 114:336
Valdivia ER, Sampedro J, Lamb JC, Chopra S, Cosgrove DJ (2007) Recent proliferation and translocation of pollen group 1 allergen genes in the maize genome. Plant Physiol 143(3):1269–1281
Wang GF, Gao Y, Wang JJ, Yang LW, Song RT, Li XR, Shi JS (2011) Overexpression of two cambium-abundant Chinese fir (Cunninghamia lanceolata) alpha- expansin genes ClEXPA1 and ClEXPA2 affect growth and development in transgenic tobacco and increase the amount of cellulose in stem cell walls. Plant Biotechnol J 9:486–502. https://doi.org/10.1111/j.1467-7652.2010.00569.x
Wechter WP et al (2008) Gene expression in developing watermelon fruit. BMC Genomics. https://doi.org/10.1186/1471-2164-9-275
Xie H et al (2009) Differential expression and regulation of expansin gene family members during fruit growth and development of ‘Shijia’ longan fruit. Plant Growth Regul 58:225–233. https://doi.org/10.1007/s10725-009-9370-3
Zhang S, Xu R, Gao Z, Chen C, Jiang Z, Shu H (2014) A genomewide analysis of the expansin genes in Malus × Domestica. Mol Genet Genomics 289(2):225–236
Zheng Y et al (2018) Cucurbit Genomics Database (CuGenDB): a central portal for comparative and functional genomics of cucurbit crops. Nucleic Acids Res 47:D1128–D1136. https://doi.org/10.1093/nar/gky944
Zhou P, Zhu Q, Xu JC, Huang BR (2011) Cloning and characterization of a gene, AsEXP1, encoding expansin proteins inducible by heat stress and hormones in creeping. Bentgrass Crop Sci 51:333–341. https://doi.org/10.2135/cropsci2010.07.0391
Zhou J, Xie JN, Liao H, Wang XR (2014) Overexpression of β-expansin gene GmEXPB2 improves phosphorus efficiency in soybean. Physiol Plant 150:194–204
Acknowledgements
This study was funded by grants from the National Natural Science Foundation of China (no. 31601795), and The National Key Research and Development Program of China (2016YFD0201000). The authors would like to thank Jian Zhang, Ph.D. (China National Rice Research Institute) for revising the manuscript and thank the reviewers for their valuable suggestions during the revision of the early manuscripts. We would also like to thank Guangzhou Gene Denovo Biotechnology Co., Ltd for assisting in bioinformatics analysis.
Author information
Authors and Affiliations
Contributions
XW and GX guided the design of the experiment. WG conducted data analysis and manuscript writing. DL, YS, BH, and XF contributed to the data analysis and performed the experiment. XW revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest in the publication.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gao, W., Li, D., Fan, X. et al. Genome-wide identification, characterization, and expression analysis of the expansin gene family in watermelon (Citrullus lanatus). 3 Biotech 10, 302 (2020). https://doi.org/10.1007/s13205-020-02293-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-020-02293-3