Abstract
Yellowing, stunting, and seedling death associated with cold stress is a common problem in many Asian countries for winter rice cultivation. Improvement of cultivars through marker-assisted selection of QTLs for cold tolerance at seedling stage from locally adapted germplasm/cultivar is the most effective and sustainable strategy to resolve this problem. A study was undertaken to map QTLs from 151 F 2:3 progenies of a cross between a cold susceptible variety, BR1 and a locally adapted traditional indica cultivar, Hbj.BVI. A total of six significant QTLs were identified for two cold tolerance indices—cold-induced leaf discoloration and survival rate after a recovery period of seven days on chromosomes 6, 8, 11, and 12. Among these QTLs, qCTSL-8-1 and qCTSS-8-1 being co-localized into RM7027–RM339 on chromosome 8 and qCTSL-12-1 and qCTSS-12-1 into RM247–RM2529 on chromosome 12 showed 12.78 and 14.96% contribution, respectively, to the total phenotypic variation for cold tolerance. Validation of QTL effect in BC1F3 population derived a cross between a cold susceptible BRRI dhan28 and Hbj.BVI showed dominating effect of qCTSL-12-1 on cold tolerance at seedling stage and it became stronger when one or more other QTLs were co-segregated with it. These results suggest that the QTLs identified in this study are stable and effective on other genetic background also, which warrant the use of these QTLs for further study aiming to cultivar development for seedling stage cold tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Low temperature is a major abiotic stress affecting growth and development of rice plants and constraints sustainable rice production in many countries of the world (Maclean et al. 2002). Cold stress at early vegetative stage of rice plants causes stunted growth and increased seedling mortality that eventually lead to uneven seedling stand establishment (Nakagahra et al. 1997). It can lead to leaf discoloration or yellowing, leaf rolling or wilting, stunted growth, delayed heading, and subsequently decreased yield of rice (Kaneda and Beachell 1973; Yoshida et al. 1996; Andaya and Mackill 2003; Fujino et al. 2004; Suh et al. 2010). Therefore, improvement of cold tolerance at seedling stage of rice has been an important issue to guarantee fast recovery and uniform crop stand (Krishnasamy and Seshu 1989) and ultimately higher yield, particularly in the rice crops which are grown in the winter season. However, breeding for cold tolerance at seedling stage has been difficult due to its polygenic nature and inadequate knowledge on genetic architecture.
With the advent of molecular biology, several reports on QTLs for cold tolerance at seedling stage in rice are available. Until now, more than 80 QTLs have been reported (Singh et al. 2016; Qian et al. 2000; Andaya and Mackill 2003; Zhan et al. 2005; Han et al. 2004, 2007; Lou et al. 2007; Jiang et al. 2008; Koseki et al. 2010; Suh et al., 2012) for cold tolerance at seedling stage in rice. In summary, Andaya and Mackill (2003) mapped a major QTL associated with cold-induced necrosis and wilting tolerance on chromosome 12 from a RIL population of M202 × IR50, which was fine mapped later on by Andaya and Tai (2006). Lou et al. (2007) mapped five main effect QTLs for survival percentage after cold treatment on chromosomes 1, 2, and 8 from a double haploid population of AAV002863 × Zhenshan97B. Koseki et al. (2010) identified and fine mapped a major QTL on chromosome 11 from Oryza rufipogon. Han et al. (2004) identified three QTLs on chromosomes 1, 5, and 9 from a cross between Milyang23 × Jileng. Han et al. (2007) mapped 12 QTLs on chromosomes 1, 2 7, 8, and 12 from a F 2:3 population of Milyang 23 × Jileng. Ji et al. (2010) reported five QTLs for seedling stage cold tolerance on chromosomes 1, 2, 4, 10, and 11 from a double haploid population of TN1 × Chunjiang 06. Park et al. (2013) mapped four QTLs on chromosomes 2 and 5 from a cross between Milyang 23 × Hapcheonaengmi3. Suh et al. (2012) identified two QTLs on long and short arms of chromosome 4 from a RIL population of Guemobyeo and IR66160-121-4-4-2. Kim et al. (2014) mapped six QTLs associated with cold tolerance at seedling stage from a RIL population of BR29 × Jinbubyeo and developed gene-based markers for two candidate genes that are located on chromosomes 1 and 11. From an RIL population of Lijiangxintuanheigu × Sanhuangzhan-2, Zhang et al. (2014) mapped four QTLs on chromosomes 1, 6, 9, and 12 for seedling yellowing and percent seedling survival after cold water irrigation and five QTLs on chromosomes 7, 8, 9, 11, and 12 responsible for leaf rolling under low-temperature exposure in phytotron. It is noticeable that all of these studies identified QTLs from Japonica germplasm; however, there only few reports available that exploited indica germplasms (Wang et al. 2016) for cold tolerance at seedling stage. Identification and use of QTLs from indica germplasm in the breeding program might be helpful and low cost for the improvement of cold tolerance in indica rice avoiding undesirable linkage drag from japonica donors. Thus, a study was undertaken to map QTLs from Hbj.BVI, a traditional indica cultivar, which showed stronger cold tolerance at seedling stage than other japonica donor parents (BRRI 2013; Khatun and Biswas 2015; Kundu 2015).
Materials and methods
Plant materials
A set of 151 F 2:3 lines derived from a cross between a cold susceptible high yielding variety, BR1, and a cold tolerant cultivar, Hbj.BVI were used in this study. Hbj.BVI is a pure line selection of Poshusail, a local variety, which is still grown in some parts of Northeastern districts of Bangladesh. On the other hand, BR1 was released in Bangladesh in 1973 through introduction of IR532-1-176 (developed from a cross between IR262-24-3 and TKM6) from IRRI, but later, studies found it highly susceptible to cold stress at seedling stage (Biswas et al. 2012). Japonica variety Jinbubyeo, Guemmbyeo, and M202 were used as cold tolerant check varieties, and IR50 and Milyang23 were used as susceptible check varieties in this study. For validation of QTL effect, BC1F3 lines derived from a cross between BRRI dhan28 (a high yielding but susceptible to cold stress at both seedling and reproductive stage) and Hbj.BVI were used. BC1F3 lines of BRRI dhan28 × Hbj.BVI were developed through the conventional backcrossing method. The F1 plants were identified comparing morphological traits of the both parents. BRRI dhan28 was used as recurrent parent in making BC1F1 which followed two successive selfing generations to make BC1F3.
Evaluation of cold tolerance at seedling stage
Parents, check varieties, and F 2:3 progenies were evaluated under artificial cold stress condition using cold water irrigation in cold water tank in 2014 following the protocol described in Khatun et al. (2016) with slight modification. Briefly, 30 seedlings of each of parents, check varieties, and F3 families were raised in plastic trays filled with gravel free fertilized soil. Each tray accommodated 30 lines including two parents at either side. The seedlings were allowed to grow until 3-leaf stage, and then, the trays were placed in the cold water tank preset at a constant temperature of 10 °C (Fig. 1). Cold water stress was withdrawn at 7 days after treatment or until the susceptible check variety dies. The trays were pulled out from cold water and allowed them to settle in ambient temperature. Leaf discoloration (LD) score was recorded immediately based on an arbitrary scale of 1 (green plants) to 9 (almost dead plants) following standard evaluation system (SES) for rice (IRRI 2013), as shown in Fig. 1. Survivability was recorded at 7 days of recovery period after withdrawal of cold treatment as the percentage of green plants to the total plants tested per family. The experiment was conducted in three replicates. One-way analysis of variance (ANOVA) procedures in Statistix 10.0 was used to analyze variance and co-efficient of correlation for the cold tolerance indices of the parents and 151 F 2:3 progenies. Heritability in broad sense was estimated from the genetic components obtained from the ANOVA. Mean differences between parents were compared by Student’s t test at 5 and 1% level of significance.
Genotyping of parents and F 2 population
Genomic DNA was extracted from leaf tissues of parents and 151 F 2 plants. Total DNA was extracted following modified miniprep CTAB method as described in Das and Biswas (2017). Six hundred and twenty SSR markers distributed over 12 chromosomes were analyzed following the protocol described in Syed et al. (2016) to identify polymorphic markers between BR1 and Hbj.BVI. Briefly, PCR reaction was performed with a 10 μl reaction mixture containing 2 μl of DNA template of 25 ng/μl conc., 4.25 μl sterile water, 1 μl 10× TB buffer (200 mMTris–HCl pH 8.3, 500 mMKCl), 1.35 μl of 25 mM MgCl2, 0.2 μl of 10 mM dNTPs, 0.5 μl each of 10 μM forward and reverse primers, and 0.20 μl of Taq DNA polymerase (5 U/μl). Gel electrophoresis of the amplified PCR products was performed in 6% polyacrylamide gel containing 0.5X TBE buffer at 100 V for 1.5–2.5 h along with a known DNA ladder. F 2 plants were genotyped with 127 polymorphic SSR markers. Allele scoring of the F 2 plant was performed based on the relative positions of DNA bands comparative to the parental bands.
Construction of linkage map, detection of QTLs, and candidate gene analysis
A linkage map was constructed based on Kosambi function using QTL mapping tool QTL IciMapping version 4.0 (http://www.isbreeding.net/software/?type=detail&id=14). A total of 127 SSR markers were used in construction of linkage map. A Chi-square test (P < 0.01) was used to identify markers with distorted segregation. An LOD value of 3.0 was used for estimation of map distance. QTL analysis was performed using composite interval mapping option in QTL IciMapping. The QTLs were declared with threshold LOD at 5% level of significance in permutation test with 1000 iterations. QTLs were named following the nomenclature system proposed by McCouch et al. (1997), and McCouch and CGSNL (2008). The marker intervals of all the significant QTLs were analyzed in silico for the presence of candidate genes. The physical distance of the QTLs was obtained from Gramene annotated Nipponbare sequence map (http://www.gamene.org) based on the position of the flanking markers. BAC clones and candidate gene information were obtained from Rice Genome Annotation Project (http://rice.plantbiology.msu.edu) and annotation database of NCBI (https://www.ncbi.nlm.nih.gov).
Validation of QTL in BC1F3 lines of BRRI dhan28XHbj.BVI
A set of 87 BC1F3 lines derived from a cross between BRRI dhan28 and Hbj.BVI was tested for seedling stage cold tolerance. Selected tolerant and susceptible lines were genotyped using flanking markers of the QTLs identified in this study. Efficacy and relative competency of expressing cold tolerance by the QTLs either singly or in combination were assessed on BRRI dhan28 background.
Results
Phenotypic evaluation of parents and mapping population
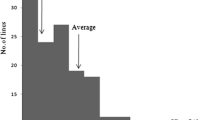
Significant differences were observed between the parents in LD and % survivability (Table 1). Under cold stress, BR1 showed very high average LD value (8.7 ± 0.33) and very low survivability rate (4.6 ± 2.39%), while low average LD (2.4 ± 0.76) and very high survivability (79.5 ± 3.88%) rate were observed with Hbj.BVI, which indicated that Hbj.BVI was highly tolerant to cold stress. Phenotypic evaluation of indica and japonica parents under cold water irrigation also showed that Hbj.BVI had stronger cold tolerance than the japonica cold tolerant varieties (Fig. 2). The 151 F 2:3 progenies showed significant variation in both of the cold tolerant indices and no significant variation was observed in any indices between the replicates. The LD values ranged from 1.8 to 9.0, while % survivability varied from 0 to 100%. A continuous and transgressive variation with bimodal frequency distribution was observed for LD, while % survivability showed a discrete but still bimodal distribution with transgressive variations (Fig. 3). However, at least 50% increase in cold tolerance over susceptible parent BR1 was observed in 46.4% families for LD and 60.3% families for survivability among F 2:3 progenies (Table 1). The heritability estimates of LD and % survivability were also observed quite high. LD value was found significant and negatively correlated with % survivability.
SSR analysis and linkage map construction
Out of 620 SSRs analyzed, 226 markers were found polymorphic between BR1 and Hbj.BVI. Maximum polymorphism rate (51.2%) was observed on chromosome 8 and minimum (18.6%) on chromosome 2 (Supplementary Table 1). Among the polymorphic markers, many of them were very close to each other, and in some other cases, the intervals between adjacent markers were comparatively higher (up to 36.8 cM) (Supplementary Fig. 1). The linkage map was constructed with 127 polymorphic SSR markers showing no distorted segregation and more or less evenly distributed over 12 chromosomes (Supplementary Fig. 2). Out of 127 SSRs, 15 markers were distributed on chromosome 1; 12 markers on each of chromosomes 2 and 3; 10 markers on each of chromosomes 4, 7, and 11; nine markers on each of chromosomes 5 and 9; 13 markers on chromosome 8 and eight markers on each of chromosomes 9 and 12. The map covered 1407.2 cM of the rice genome with an average interval of 10.99 cM. There were still five marker intervals bigger than 20 cM—one on each of chromosomes 3, 4, 5, 7, and 12.
QTL analysis
A total of six significant QTLs were localized for two cold tolerance indices on chromosomes 6, 8, 11, and 12. These QTLs and their chromosomal position, linked makers, additive effects, and contributions in explaining phenotypic variations were given in Table 2. Among six QTLs, qCTSL-6-1, qCTSL-8-1, and qCTSL-12-1 for LD were located on chromosomes 6, 8, and 12 with bordering markers RM19996–RM3, RM7027–RM339, and RM247–RM2529, respectively (Fig. 4). The phenotypic contributions of these QTLs were 7.29, 12.78, and 14.96%, respectively. While, three QTLs—qCTSS-8-1, qCTSS-11-1, and qCTSS-12-1 with flanking markers, RM7027–RM339, RM26324–RM7283, and RM247–RM2529, respectively, were mapped on chromosomes 8, 11, and 12 for % survivability. The qCTSS-8-1, qCTSS-11-1, and qCTSS-12-1 had phenotypic variance of 12.79, 8.70, and 13.06%, respectively. The additive effects of the QTLs for LD were negative in direction, while positive additive effects were observed for the QTLs governing % survivability.
Candidate gene analysis
Among six QTLs, qCTSL-6-1, qCTSL-8-1, qCTSS-11-1, and qCTSL-12-1 were analyzed for underlying candidate genes. The annotation information of the candidate genes is given in Table 3. In total, 27 putative candidate genes, responsible cold tolerance, were identified from 1638 ORFs present in the marker intervals of all four QTLs. For qCTSL-6-1, four loci LOC_Os06g30780, LOC_Os06g30690, LOC_Os06g30310, and LOC_Os06g26340 encoding, respectively—acyl-desaturase (chloroplast precursor), alcohol oxidase related protein, alpha-glucan water dikinase, and CGMC_MAPKCMGC_2.10—CGMC includes CDA, MAPK, GSK3, and CLKC kinases were found to be related with cold acclimatization. LOC_Os08g27170 for calmodulin-binding protein, LOC_Os08g27860 and LOC_Os08g27870 for early flowering protein, LOC_Os08g27850 for endothelial differentiation-related factor 1, LOC_Os08g29100 for SAM domain family protein, and LOC_Os08g29110 for Thioredoxin were identified in the marker interval of qCTSL-8-1. On the other hand, LOC_Os11g14040, LOC_Os11g14050, and LOC_Os11g14900 encoding glutathione S-transferase, N-terminal domain containing protein, leucine-rich repeat family protein, and thiol protease SEN102 precursor, respectively, were identified as putative candidate genes of qCTSS-11-1. A total of 14 ORFs including LOC_Os12g07230 and LOC_Os12g12860 encoding CAMK_CAMK_like.45—CAMK includes calcium/calmodulin-dependent protein kinases, LOC_Os12g07610 and LOC_Os12g07640 encoding MYB family transcription factor, LOC_Os12g10720 and LOC_Os12g10730 encoding Glutathione S-transferase and LOC_Os12g12730 encoding OsCML28—calmodulin-related calcium sensor protein were identified as putative candidate genes in the marker interval of qCTSL-12-1.
Validation of QTL in BC1F3 lines of BRRI dhan28 × Hbj.BVI
BC1F3 lines of BRRI dhan28 × Hbj.BVI showed wide range of variation in LD values under cold water treatment at seedling stage. Out of 87 lines tested, we found 53 lines showing tolerant (LD score: 1–3) to moderately tolerant (LD score: 5) reaction (Fig. 5). Genotyping of 60 BC1F3 plants including five susceptible plants and two parents with bordering SSR markers of qCTSL-6-1, qCTSL-8-1, qCTSS-11-1 and qCTSL-12-1 showed that 49 lines had positive results for at least one flanking markers for one or more QTLs. We further scrutinized the lines for the presence of positive results (homozygous Hbj.BVI alleles) at both flanking markers for the QTLs. Out of 49, seven lines showing tolerance reaction to cold stress (Fig. 6) had both flanking markers homozygous for Hbj.BVI alleles for at least two QTLs including qCTSL-12-1 except two lines (F and G). The line F and G possessed combination of two QTLs, qCTSL-6-1 + qCTSL-8-1 and qCTSL-8-1 + qCTSS-11-1, respectively. On the other hand, the lines carrying single QTL, qCTSL-6-1 (line K), or qCTSS-11-1 (line M) showed susceptible reaction except line H, which contained qCTSL-12-1 and showed tolerant reaction. However, the line (I) containing both qCTSL-6-1 and qCTSS-11-1 showed moderate tolerance to cold stress.
QTL effect between tolerant and susceptible BC1F3 families, the bar divided by different colors indicates four QTLs associated with phenotypic variance for cold tolerance of each QTL. The letter A, B, C, D, E, F, G, H, I, J, K, and L indicated BC1F3 families, BC1F3-14-2, BC1F3-14-9, BC1F3-26-7, BC1F3-26-12, BC1F3-26-22, BC1F3-29-3, BC1F3-29-5, BC1F3-29-10, BC1F3-132-1, BC1F3-132-6, BC1F3-132-11, and BC1F3-132-18, respectively. T, MT, and S indicated tolerant, moderately tolerant, and susceptible, respectively
Discussion
In general, rice genotypes belonging to japonica subspecies represents higher cold tolerance than indica, but there is variability within the subspecies. Rice crop in tropical and sub-tropical regions where indica rice is commonly cultivated is usually affected by low temperature at early vegetative stage limiting growth, delaying flowering that eventually results into low yield. However, most of the breeding programs and QTL mapping works across the world used japonica donors for seedling stage cold tolerance (Qian et al. 2000; Andaya and Mackill 2003; Zhan et al. 2005; Han et al. 2004, 2007; Lou et al. 2007; Jiang et al. 2008; Koseki et al. 2010; Ji et al. 2010; Suh et al. 2012; Park et al. 2013; Yang et al. 2013; Kim et al. 2014; Ranawake et al. 2014; Zhang et al. 2014). A very few work using indica donors has been reported so far. Wang et al. (2016) used a rice diversity panel comprising japonica, indica, and aus cultivars to map QTLs for seedling stage cold tolerance. In a previous study, we identified Hbj.BVI as a potential indica donor for cold tolerance at seedling stage (BRRI 2013; Khatun and Biswas 2015). From a haplotype analysis study with marker information of the reference QTLs identified from different japonica donors, Kundu (2015) reported that Hbj.BVI showing better tolerance to cold stress at seedling stage was grouped into a different cluster from the other japonica donors such as, M202, Jinbubyeo, IR66160-121-4-4-2, Jinbubyeo, and Jinbubyeo derived lines. Phenotypic evaluation with japonica cultivars as cold tolerant check varieties also showed strong evidence that Hbj.BVI had higher cold tolerance at seedling stage (Fig. 2). These results clearly indicated that Hbj.BVI might be a potential indica donor. Mapping of QTLs from Hbj.BVI, thus, might be helpful in developing high yielding cold tolerant indica rice. Although high-throughput and robust genotyping techniques are available for maker-assisted breeding, small-scale breeding program of public sectors can reliably use indica donor in the breeding program without shouldering robust budget to minimize undesired linkage drags from Japonica donors into the introgression lines.
The most crucial step of a QTL mapping work is phenotyping, particularly for seedling stage cold tolerance. Plants’ responses to cold stress are a complex phenomenon and are highly influenced by environment. Thus, the extent of low temperature and duration of exposure are extremely important for effective evaluation of cold tolerance. In different previous studies, different temperature regimes and durations of exposure were used to test cold tolerance of rice at seedling stage. Some studies used constant temperature varying from 4 to 12 °C (Zhan et al. 2005; Andaya and Mackill 2003; Qu et al. 2003; Han et al. 2005a, b) and some others used variable temperature over day to night (20/13 °C by Park et al. 2013, 10/6 °C by Qian et al. 2000). Khatun et al. (2016) reported that cold water irrigation with constant 13 °C in cold water tank at 3-leaf stage-seedlings of rice for 7 days is sufficiently enough to discriminate cold tolerant varieties from intolerant ones, and it is as good as constant 10 °C treatment. Yoshida (1981) reported 10 °C as the minimum critical value, below which rice seedlings get yellowing and stunted growth and loses potency to survive (Yoshida 1981). In our study, we used constant 10 °C cold water irrigation for cold screening at seedling stage. Usually cold stress at seedling stage of rice manifests different symptoms of cold response, such as leaf rolling, yellowing, wilting, seedling growth retardation, seedling death, and so on. Some indices reflecting these phenomena are used to evaluate cold response. Cold-induced leaf discoloration (Andaya and Mackill 2003; Han et al. 2005a, b), seedling survivability (Qian et al. 2000; Chen et al. 2002; Qu et al. 2003; Lou et al. 2007; Zhang et al. 2014), seedling vigor under cold stress (Zhang et al. 2005), and germination rate (Lee et al. 2001; Sheng et al. 2001; Chen et al. 2006; Dashtmian et al. 2013) are widely used indices of assessing cold response. Leaf yellowing or discoloration and survivability rate under low-temperature stress are directly related to seedling mortality in the seedbed and in the main field immediately after transplanting. In fact, seedling mortality in the seedbed and in the main field is very common in rice crop that is grown in winter season in South Asian countries including Bangladesh. Considering this fact, in our study, we phenotyped an F 2:3 population of 151 individuals and their parents for cold-induced LD and % survivability after a recovery period of seven days from cold stress withdrawal. The F 2:3 progenies showed significant and wide variation in both LD and % survivability ranging from 1.8–9.0 to 0–100%, respectively. While, the parental values for LD were 8.7 ± 0.33 and 2.4 ± 2.39 and 4.6 ± 0.76 and 79.5 ± 3.88 for % survivability. It is clearly evident that the ranges of variation in both of the indices of the mapping population exceeded the parental range (Table 1) indicating the presence of transgressive segregants. Syed et al. (2016) observed similar results for arsenic phyto-toxicity tolerance, Mohiuddin (2016) for grain zinc content, Swamy et al. (2011, 2014) for yield and yield related traits, Zhang et al. (2014) for cold-induced yellowing tolerance and wilting tolerance, and Park et al. (2013) for SPAD value of rice seedlings under cold stress. It is worthy to note that at least 50% increase of cold tolerance over susceptible parent BR1 was observed in 46.4% families for LD and 60.3% families for survivability among the F 2:3 progenies. However, the variation was continuous and bimodal for LD, and for % survivability, it was discrete but still bimodal among the F 2:3 progenies. Bimodal distribution was reported by Swamy et al. (2012) in amylose content of rice, Kuroki et al. (2009) for heading date under cold stress in rice, and Andaya and Tai (2006; 2007) for CT score at seedling stage. The above findings indicated that both major and minor QTLs for LD and major QTLs for % survivability might involve in the mapping population of this study.
Using LD scores and survivability rate as cold tolerance indices and genotyping data with 127 polymorphic SSR markers distributed at an average interval of 10.99 cM over 12 chromosomes, we mapped three QTLs—one at each of chromosomes 6, 8, and 12 for cold-induced LD and three QTLs—one at each of chromosomes 8, 11, and 12 for % survivability accounting 35.03 and 34.55% cumulative contribution to the total phenotypic variation, respectively. Among these QTLs, qCTSL-8-1 for LD and qCTSS-8-1 for % survivability were located into the same marker interval (RM7027–RM339) on chromosome 8. In addition, the qCTSL-12-1 for LD and qCTSS-12-1 for % survivability were co-located into the same RM247–RM2529 marker interval on chromosome 12. Very high and significant correlation among the indices also supports of co-localization of one or more QTLs conferring these traits. The co-localized QTLs might be the same QTLs or overlapped QTLs affecting two or more traits. Co-localized QTLs were also reported earlier by Wang et al. (2012) for salinity tolerance in rice at seedling stage, Andaya and Mackill (2003), Han et al. (2004, 2007), Zhang et al. (2014) for cold tolerance at seedling stage in rice. These co-localized QTLs might be useful in simultaneous improvement of one or more traits as the desirable alleles of these QTLs were contributed by a single parent. The positive and negative additive effects of all the significant QTLs for % survivability and LD, respectively, indicated that increasing frequency of favorable alleles from Hbj.BVI has resulted into increased in survival rate and decreased in LD values, i.e., tolerance to cold stress at seedling stage in terms of survivability and LD score in the F 2:3 population.
The cross validation with the previous studies for the physical position in the rice genome showed that the newly identified QTLs from this study were located at the proximity of the QTLs reported in other previous studies (Table 4). The qCTSL-6-1, which was mapped at 14.4–19.5 Mb on chromosome 6, was located near the QTLs identified by Liu et al. (2013), Zhang et al. (2014) and Andaya and Mackill (2003). The qCTS6 identified by Liu et al. (2013) at 20.9–28.6 Mb, was just at 1.4 Mb upstream of qCTSL-6-1, while, the qCTS-6 identified by Zhang et al. (2014) at 19.5–28.4 Mb and qCTS6-2 reported by Andaya and Mackill (2003) at 19.4–25.8 Mb were located at tandem with qCTSL-6-1. The qCTSS-11-1, which was mapped at 7.4–9.1 Mb on chromosome 11, was found into the marker interval of qGAS11 reported by Han et al. (2004) at 4.0–16.8 Mb. Ranawake et al. (2014) also mapped qCTS11 (1)-1 at marker interval of 4.0–9.0 Mb in which we mapped qCTSS-11-1. However, the marker intervals of qGAS11 and qCTS11 (1)-1 were much bigger (10.1 and 2.3 Mb, respectively) than that of qCTSS-11-1. Furthermore, Zhang et al. (2005) mapped a major QTL (qSCT-11) for seedling stage cold tolerance, which is tandemly located with qCTSS-11-1 at 9.0–17.8 Mb. On the other hand, qCTSL-8-1 and qCTSS-8-1, which were co-localized at 15.8–17.9 Mb on chromosome 8, were mapped at 3.2 Mb upstream of the qCTS8-1 reported by Yang et al. (2013). However, Andaya and Mackill (2003) mapped three QTLs (qCTS8-1, qCTS8-2a and qCTS8-2a) at 2.7–3.2 Mb downstream of qCTSL-8-1. Besides, the other two co-localized QTLs (qCTSL-12-1 and qCTSS-12-1), which were mapped at 3.2–7.5 Mb on chromosome 12, in this study, were found in the vicinity the QTLs reported by Zhang et al. (2014) and Han et al. (2007). However, marker interval of the QTLs reported by Zhang et al. (2014) and Han et al. (2007) was, respectively, 2.3 Mb smaller and 11.1 Mb bigger than of that of the QTLs identified in this study. Interestingly, Andaya and Mackill (2003) mapped a major QTL (qCTS12) at 1.3 Mb upstream of our QTLs on chromosome 12. The above findings indicated that the QTLs identified in this study are reliable and stable in different genetic background and we mapped QTLs with relatively smaller marker interval in the proximity of previously identified QTLs except few cases. When we analyzed the marker intervals of the QTLs for the presence of candidate gene in silico, we found at total of 27 putative candidate genes, of which 4 were for qCTSL-6-1, 6 for qCTSL-8-1, 3 for qCTSS-11-1, and 14 for qCTSL-12-1. These results clearly indicated that seedling stage cold tolerance in this study was governed by many genes situated at different QTL regions with preponderance share of qCTSL-12-1. Among the putative candidate genes, LOC_Os11g14040 in qCTSS-11-1 and, LOC_Os12g10720 and LOC_Os12g10730 in qCTSL-12-1 encoding glutathione S-transferase and LOC_Os12g07230 and LOC_Os12g12860 encoding CAMK_CAMK_like.45—CAMK includes calcium/calmodulin-dependent protein kinases and LOC_Os12g12730 encoding OsCML28—calmodulin-related calcium sensor protein in qCTSL-12-1 are considered as important factor for cold tolerance (Doherty et al. 2009). Calmodulin-binding transcription activator governing cold tolerance pathway in rice was also reported earlier by Kim et al. (2014) in japonica germplasm Jinbubyieo. We located LOC_Os12g07610 and LOC_Os12g07640 in qCTSL-12-1, which encode calcium-dependent MYB family transcription factor that mediates cold signaling pathways. Su et al. (2010) reported that over-expressing MYBS3 transgenic rice lines showed tolerance to 4 °C for at least 1 week. The above findings suggest that the QTLs identified in this study have much worth in transferring from Hbj.BVI into other genetic background to develop high yielding cultivars tolerant to cold stress at seedling stage.
We validated the QTLs mapped in this study in a BC1F3 population derived from a cross between BRRI dhan28 and Hbj.BVI. Phenotyping under cold water irrigation followed by genotyping of selected 53 tolerant to moderately tolerant lines and five susceptible lines with their parents using flanking SSRs of qCTSL-6-1, qCTSL-8-1, qCTSS-11-1, and CTSL-12-1 showed that the lines having two or more QTLs including qCTSL-12-1 were tolerant to cold stress and lines with single QTL except qCTSL-12-1 were susceptible. The line with single qCTSL-12-1 showed tolerant reaction. On the other hand, the lines which contained combination of two QTLs other than qCTSL-12-1 also showed tolerance reaction. These indicated that qCTSL-12-1 has dominating effect on cold tolerance at seedling stage and it became stronger when one or more other QTLs were co-segregated with it. These results suggest that the QTLs identified from Hbj.BVI are stable in different genetic backgrounds, which warrant the use of these QTLs for further study aiming to cultivar development for seedling stage cold tolerance. In addition, MAS with linked markers of qCTSL-12-1 in combination with one or more of qCTSL-6-1, qCTSL-8-1, and qCTSS-11-1 might increase cold tolerance in the pyramided lines.
References
Andaya VC, Mackill DJ (2003) Mapping of QTLs associated with cold tolerance during the vegetative stage in rice. J Exp Bot 54:2579–2585
Andaya VC, Tai TH (2006) Fine mapping of the qCTS12 locus, a major QTL for seedling cold tolerance in rice. Theor Appl Genet 113:467–475
Andaya VC, Tai TH (2007) Fine mapping of the qCTS4 locus associated with seedling cold tolerance in rice (Oryza sativa L.). Mol Breed 20:349–358. doi:10.1007/s11032-007-9096-8
Biswas PS, Khatun H, Anisuzzaman M, Hwang HG (2012) Study on the cold tolerant rice varieties for Bangladesh. Korea International Cooperation Agency (KOICA), The Republic of Korea
BRRI (2013) BRRI Annual Report 2012–13. Bangladesh Rice Research Institute Gazipur-1701, Bangladesh, p 334. http://brri.portal.gov.bd/sites/default/files/files/brri.portal.gov.bd/page/6d86d72e_953c_4f74_9157_3cfeaa80c2b1/BRRI_A_Report_2011-12.pdf
Chen DZ, Zhong PA, Xiao YQ, Huang YJ, Xie JK (2002) Identification of QTL for cold tolerance at seedling stage in Dongxiang wild rice (Oryza rufipogon Griff) by SSR markers. Acta Agriculturae Universitatis Jiangxiensis 24(6):753–756 (in Chinese with English abstract)
Chen L, Lou Q, Sun Z, Xing Y, Yu X, Luo L (2006) QTL mapping of low temperature on germination rate of rice. Rice Sci 13:93–98
Das N, Biswas PS (2017) Molecular characterization of parental lines of rice aiming to address high yield and nutritional quality under drought and cold stress condition. Curr Res Agril Sci 4:51–60. doi:10.18488/journal.68.2017.42.51.60
Dashtmian FP, Hosseini MK, Esfahani M (2013) Methods for rice genotypes cold tolerance evaluation at germination stage. Intl J Agric Crop Sci 5:2111–2116
Doherty CJ, Van Buskirk HA, Myers SJ, Thomashow MF (2009) Roles for arabidopsis CAMTA transcription factors in cold-regulated gene expression and freezing tolerance. Plant Cell 21(3):972–984
Fujino K, Sekiguchi H, Sato T, Kiuchi H, Nonoue Y, Takeuchi Y, Ando T, Lin SY, Yano M (2004) Mapping of quantitative trait loci controlling low-temperature germinability in rice (Oryza sativa L.). Theor Appl Genet 108:794–799
Han L, Qiao Y, Cao G, Zhang Y, An Y, Ye J, Koh H (2004) QTLs analysis of cold tolerance during early growth period for rice. Rice Sci 11:245–250
Han LZ, Zhang SY, Qiao YL, Zhang JG, Ruan RC, Cao GL, Koh HJ (2005a) QTL analysis of root traits at the seedling stage in rice under cold irrigation. Acta Agron Sin 31:1415–1421
Han LZ, Qiao YL, Cao GL, Zhang YY, An YP, Ye JD, Koh HJ (2005b) QTL analysis on cold tolerance during early growth period in rice. Chin J Rice Sci 19:122–126. http://caod.oriprobe.com/orderhtm?id=626355&ftext=base (in Chinese with English abstract)
Han L, Qiao Y, Zhang S, Zhang Y, Cao G, Kim J, Lee K, Koh H (2007) Identification of quantitative trait loci for cold response of seedling vigor traits in rice. J Genet Genomics 34:239–246
IRRI (2013) Standard evaluation system of rice, 5th edn. International Rice Research Institute, Philippines, p 55p
Ji ZJ, Zeng YX, Zeng DL, Ma LY, Li XM, Liu BX, Yang CD (2010) Identification and fine mapping of a major quantitative trait locus originating from wild rice controlling cold tolerance at the seedling stage. Mol Genet Genom 284:45–54
Jiang L, Xun MM, Wang JK, Wan JM (2008) QTL analysis of cold tolerance at seedling stage in rice (Oryza sativa L.) using recombination inbred lines. J Cereal Sci 48:173–179
Kaneda C, Beachell HM (1973) Response of indica-japonica rice hybrids to low temperatures. SABRAO J 6:17–32
Khatun H, Biswas PS (2015) BR18 and HbjBVI: Potential donors for QTLs/genes conferring seedling stage cold tolerance in rice. In: Saleque MA, Kashem MA, Ali MA, Kabir MS (eds) Bangladesh Rice Research Abstracts 2014. Bangladesh Rice Research Institute, Gazipur-1701, Bangladesh. http://brriportalgovbd/sites/default/files/files/brriportalgovbd/page/b3bb0382_9b99_43f3_8490_20455d2ec5e4/Bangladesh%20Rice%20Research%20Abstract%202014pdf
Khatun H, Biswas PS, Hwang HG, Kim KM (2016) A quick and simple in-house screening protocol for cold tolerance at seedling stage in rice. Plant Breed Biotech 4:373–378. doi:10.9787/PBB.2016.4.3.373
Kim SM, Suh JP, Lee CK, Le JH, Kim YG, Jena KK (2014) QTL mapping and development of candidate gene-derived DNA markers associated with seedling cold tolerance in rice (Oryza sativa L.). Mol Genet Genomics 289:333–343
Koseki M, Kitazawa N, Yonebayashi S, Maehara Y, Wang ZX, Minobe Y (2010) Identification and fine mapping of a major quantitative trait locus originating from wild rice, controlling cold tolerance at the seedling stage. Mol Genet Genomics 284:45–54
Krishnasamy V, Seshu DV (1989) Seed germination rate and associated characters in rice. Crop Sci 29:904–908
Kundu A (2015) Haplotype diversity analysis in cold tolerant rice. MS thesis, Bangabandhu Sheikh Mujibur Rahman Agricultural University, Salna, Gazipur-1703, Bangladesh
Kuroki M, Saito K, Matsuba S, Yokogami N, Shimizu H, Ando I, Sato Y (2009) Quantitative trait locus analysis for cold tolerance at the booting stage in a rice cultivar, Hatsushizuku. JARQ 43 (2):115–121. http://www.jircas.affrc.go.jp
Lee MH, Fukai S, Basnayake J (2001) Increased lowland rice production in the Mekong Region. In: Proceedings of an international workshop, Vientiane, Laos, 30th October to 2nd November, 2000 at Australian Center for International Agricultural Research, Canberra, Australia on Low temperature tolerance in rice: The Korean Experience, pp 109–117
Liu FX, Xu WY, Song Q, Tan LB, Liu JY, Zhu ZF, Fu YC, Su Z, Sun CQ (2013) Microarray-assisted fine-mapping of quantitative trait loci for cold tolerance in rice. Mol Plant 6:757–767. doi:10.1093/mp/sss161
Lou Q, Chen L, Sun Z, Xing Y, Li J, Xu X, Mei H, Luo L (2007) A major QTL associated with cold tolerance at seedling stage in rice (Oryza sativa L.). Euphytica 158:87–97
Maclean JL, Dawe DC, Hardy B, Hettel GP (2002) Rice almanac: source book for the most important economic activity on earth, 3rd edn. CABI Publishing, UK
McCouch SR, CGSNL (Committee on Gene Symbolization, Nomenclature and Linkage, Rice Genetics Cooperative) (2008) Gene nomenclature system for rice. Rice 1:72–84
McCouch SR, Chen X, Panaud O, Temnykh S, Xu Y, Cho YG, Huang N, Ishii T, Blair M (1997) Microsatellite marker development, mapping and applications in rice genetics and breeding. Plant Mol Biol 35:88–99
Mohiuddin SJ (2016) Molecular mapping of quantitative trait loci conferring grain zinc content in rice (Oryza sativa L). MS Thesis, Bangabandhu Sheikh Mujibur Rahman Agricultural University, Salna, Gazipur, Bangladesh
Nakagahra M, Okuno K, Vaughan D (1997) Rice genetic resources: history, conservation, investigative characterization and use in Japan. Plant Mol Biol 35:69–77
Park IK, Oh CS, Kim DM, Yeo SM, Ahn SN (2013) QTL Mapping for cold tolerance at the seedling stage using introgression lines derived from an inter-subspecific cross in Rice. Plant Breed Biotech 1:1–8. doi:10.9787/PBB.2013.1.1.001
Qian Q, Zeng DL, He P, Zheng X, Chen Y, Lihuang Z (2000) QTL analysis of the rice seedling cold tolerance in a double haploid population derived from anther culture of a hybrid between indica and japonica rice. Chin Sci Bull 145(5):448–453
Qu TT, Chen LY, Zhang ZH, Hu ZH, Liping ZH, Lihuang ZY (2003) Molecular mapping of genes conferring cold tolerance at seedling stage using doubled haploid lines from an indica × japonica cross in rice. J Wuhan Bot Res 21:385–389
Ranawake AL, Manangkilb OE, Yoshidac S, Ishiid T, Morie N, Nakamura C (2014) Mapping QTLs for cold tolerance at germination and the early seedling stage in rice (Oryza sativa L.). Biotechnol Biotechnol Equip 28:989–998. doi:10.1080/13102818.2014.978539
Sheng T, Dali Z, Qian Q, Yasufumi K, Danian H, Lihuang Z (2001) QTL analysis of rice low temperature germinability. Chin Sci Bull 46:1800–1803
Singh BK, Singh AK, Ngasepam TM, Mukharjee A, Mandal N (2016) QTL mapping for cold tolerance at the seedling stage in rice. Int J Biores Stress Mgt 7:1214–1224
Su CF, Wang YC, Hsieh TH, Lu CA, Tseng TH, Yu SM (2010) A novel MYBS3-dependent pathways confers cold tolerance in rice. Plant Physol 153:145–158
Suh JP, Jeung JU, Lee JI, Choi YH, Yea JD, Virk PS, Mackill DJ, Jena KK (2010) Identification and analysis of QTLs controlling cold tolerance at the reproductive stage and validation of effective QTLs in cold-tolerance genotypes of rice (Oryza sativa L.). Theor Appl Genet 120:985–995
Suh JP, Lee CK, Lee JH, Kim JJ, Kim SM, Cho YC, Park SH, Shin JC, Kim YG, Jena KK (2012) Identification of quantitative trait loci for seedling cold tolerance using RILs derived from a cross between japonica and tropical japonica rice cultivars. Euphytica 184:101–108
Swamy BPM, Kaladhar K, Ramesha MS, Viraktamath BC, Sarla N (2011) Molecular mapping of QTLs for yield and related traits in Oryza sativa cv Swarna × O. nivara (IRGC81848) backcross population. Rice Sci 18:178–186
Swamy BPM, Kaladhar K, Rani SN, Prasad GSV, Viraktamath BC, Reddy GA, Sarla N (2012) QTL analysis for grain quality traits in 2 BC2F2 Populations Derived from Crosses between Oryza sativa cv Swarna and 2 Accessions of O. nivara. J Heredity 103:442–452
Swamy M, Kaladhar K, Reddy GA, Viraktamath BC, Sarla N (2014) Mapping and introgression QTLs for yield and related traits in two backcross populations derived from O. sativa cv Swarna and two accessions of O. nivara. J Genet 93:643–654
Syed MA, Iftekharuddaula MK, Mian MAK, Rasul MG, Rahmam GKMM, Panaullah GM, Lauren JG, Duxbury JM, Biswas PS (2016) Main effect QTLs associated with arsenic phyto-toxicity tolerance at seedling stage in rice (Oryza sativa L.). Euphytica 209:805–814. doi:10.1007/s10681-016-1683-5
Wang Z, Cheneg J, Chen Z, Huang J, Boo Y, Wang J, Zhang H (2012) Identification of QTLs with main, epistatic and QTL × Environment interaction effects for salt tolerance in rice seedlings under different salinity conditions. Theor Appl Genet 125:807–815. doi:10.1007/s00122-012-1873-z
Wang D, Liu J, Li C, Kang H, Wang Y, Tan X, Liu M, Deng Y, Wang Z, Liu Y, Zhang D, Xiao Y, Wang GL (2016) Genome-wide association mapping of cold tolerance genes at seedling stage in rice Rice 9:10–61. doi:10.1186/s12284-016-0133-2
Yang Z, Huang D, Tan W, Zheng Y, Liang K, Cutler AJ, Wu W (2013) Mapping of quantitative trait loci underlying cold tolerance in rice seedlings via high-throughput sequencing of pooled extremes. PLoS One 8(7):e68433. doi:10.1371/journal.pone.0068433
Yoshida S (1981) Fundamentals of rice crop science. International Rice Research Institute, Los Banos, p 269
Yoshida R, Kanno A, Sato T, Kameya T (1996) Cool temperature induced chlorosis in rice plant. Plant Physiol 110:997–1005
Zhan QC, Zhu KY, Chen ZW, Zeng SZ (2005) Studies on the QTL for cold tolerance related characters of rice seedling by molecular markers. Hybrid rice 20(1):50–55 (in Chinese with English abstract)
Zhang ZH, Su L, Li W, Chen W, Zhu YG (2005) A major QTL conferring cold tolerance at the early seedling stage using recombinant inbred lines of rice (Oryza sativa L.). Plant Sci 168:527–534
Zhang S, Zheng J, Liu B, Peng S, Leung H, Zhao J, Wang X, Yang T, Huang Z (2014) Identification of QTLs for cold tolerance at seedling stage in rice (Oryza sativa L.) using two distinct methods of cold treatment. Euphytica 195:95–104. doi:10.1007/s10681-013-0977-0
Acknowledgements
The authors deeply acknowledge the Integrated Agricultural Productivity Project (IAPP)—BRRI part, Bangladesh Rice Research Institute, Gazipur-1701, Bangladesh for financial support for this work. We also acknowledge Dr. Tamal Lata Aditya, Head of Plant Breeding Division, Bangladesh Rice Research Institute for sharing germplasm and allowing us to use laboratory and experiment field. We also acknowledge laboratory and field technicians for their continuous assistance in conducting phenotyping and genotyping works.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by the Integrated Agricultural Productivity Project (IAPP)-BRRI part, Bangladesh Rice Research Institute, Gazipur-1701.
Conflict of interest
There is conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Biswas, P.S., Khatun, H., Das, N. et al. Mapping and validation of QTLs for cold tolerance at seedling stage in rice from an indica cultivar Habiganj Boro VI (Hbj.BVI). 3 Biotech 7, 359 (2017). https://doi.org/10.1007/s13205-017-0993-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-0993-1