Abstract
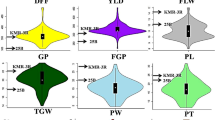
Two soybean recombinant inbred line populations, Jinpumkong 2 × SS2-2 (J × S) and Iksannamulkong × SS2-2 (I x S) showed population-specific quantitative trait loci (QTLs) for days to flowering (DF) and days to maturity (DM) and these were closely correlated within population. In the present study, we identified QTLs for six yield-related traits with simple sequence repeat markers, and biological correlations between flowering traits and yield-related traits. The yield-related traits included plant height (PH), node numbers of main stem (NNMS), pod numbers per plant (PNPP), seed numbers per pod (SNPP), 100-seed weight (SW), and seed yield per plant (SYPP). Eighteen QTLs for six yield-related traits were detected on nine chromosomes (Chrs), containing four QTLs for PH, two for NNMS, two for PNPP, three for SNPP, five for SW, and two for SYPP. Two highly significant QTLs for PH and NNMS were identified on Chr 6 (LG C2) in both populations where the major flowering gene, E1, and two DF and DM QTLs were located. One other PNPP QTL was also located on this region, explaining 12.9% of phenotypic variation. Other QTLs for yield-related traits showed population-specificity. Two significant SYPP QTLs potentially related with QTLs for SNPP and PNPP were found on the same loci of Chrs 8 (Satt390) and 10 (Sat_108). Also, highly significant positive phenotypic correlations (P < 0.01) were found between DF with PH, NNMS, PNPP, and SYPP in both populations, while flowering was negatively correlated with SNPP and SW in the J × S (P < 0.05) and I × S (P < 0.01) populations. Similar results were also shown between DM and yield-related traits, except for one SW. These QTLs identified may be useful for marker-assisted selection by soybean breeders.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ariyo OJ. 1995. Correlation and path-coefficient analysis of components of yield in soybean. Afr. Crop Sci. J. 3: 29–33
Arshad M, Ali N, Ghafoor A. 2006. Character correlation and path coefficient in soybean Glycine max (L.) Merrill. Pak. J. Bot. 38: 121–130
Brondani C, Rangel P, Brondani R, Ferreira M. 2002. QTL map ping and introgression of yield-related traits from Oryza glumaepatula to cultivated rice (Oryza sativa) using microsatellite markers. Theor. Appl. Genet. 104: 1192–1203
Chapman A, Pantalone V, Ustun A, Allen F, Landau-Ellis D, Trigiano R, Gresshoff P. 2003. Quantitative trait loci for agronomic and seed quality traits in an F2 and F4:6 soybean population. Euphytica 129: 387–393
Chen G, Geng JF, Rahman M, Liu XP, Tu JX, Fu TD, Li GY, McVetty PBE, Tahir M. 2010. Identification of QTL for oil content, seed yield, and flowering timein oilseed rape (Brassica napus). Euphytica 175: 161–174
Choi IY, Hyten DL, Matukumalli LK, Song Q, Chaky JM, Quigley CV, Chase K, Lark KG, Reiter RS, Yoon M-S, Hwang E-Y et al. 2007. A soybean transcript map: gene distribution, haplotype and single-nucleotide polymorphism analysis. Genetics 176: 685–696
Cober E, Molnar S, Charette M, Voldeng H. 2010. A new locus for early maturity in soybean. Crop Sci. 50: 524–527
Cobos M, Rubio J, Fernandez Romero M, Garza R, Moreno M, Millan T, Gil J. 2007. Genetic analysis of seed size, yield and days to flowering in a chickpea recombinant inbred line pop ulation derived from a Kabuli x Desi cross. Ann. Appl. Biol. 151: 33–42
Funatsuki H, Kawaguchi K, Matsuba S, Sato Y, Ishimoto M. 2005. Mapping of QTL associated with chilling tolerance during reproductive growth in soybean. Theor. Appl. Genet. 111: 851–861
Hoeck J, Fehr W, Shoemaker R, Welke G, Johnson S, Cianzio S. 2003. Molecular marker analysis of seed size in soybean. Crop Sci. 43: 68–74
Hyten D, Pantalone V, Sams C, Saxton A, Landau-Ellis D, Stefaniak T, Schmidt M. 2004. Seed quality QTL in a prominent soybean population. Theor. Appl. Genet. 109: 552–561
Keim P, Diers BW, Olson TC, Shoemaker RC. 1990. RFLP mapping in soybean: association between marker loci and variation in quantitative traits. Genetics 126: 735–742
Kim MY, Van K, Lestari P, Moon JK, Lee S-H. 2005. SNP identification and SNAP marker development for a GmNARK gene controlling supernodulation in soybean. Theor. Appl. Genet. 110: 1003–1010
Kim SD, Kim YH, Park KY, Yun HT, Lee YH, Lee SH, Seong YK, Kim HS, Hong EH, Kim YS. 1997. A new beany taste less soybean variety ‘Jinpumkong 2’ with good seed quality. RDA J. Agri. Sci. 39: 112–115
Lander ES, Daly MJ, Lincoln SE. 1993. Constructing genetic linkage maps with MAPMAKER/EXP Version 3.0: A tutorial and reference manual, A Whitehead Institute for Biomedical Research Technical Report, 3rd Edition, January.
Lee SH, Bailey MA, Mian MAR, Shipe ER, Ashley DA, Parrott WA, Hussey RS, Boerma HR. 1996. Identification of quantitative trait loci for plant height, lodging, and maturity in a soybean population segregating for growth habit. Theor. Appl. Genet. 92: 516–523
Lee SH, Park KY, Lee HS, Park EH, Boerma HR. 2001. Genetic mapping of QTLs conditioning soybean sprout yield and quality. Theor. Appl. Genet. 103: 702–709
Li DM, Sun MM, Han YP, Teng WL, Li WB. 2010. Identification of QTL underlying soluble pigment content in soybean stems related to resistance to soybean white mold (Sclerotinia sclerotiorum). Euphytica 172: 49–57
Liu WX, Kim MY, Van K, Sun SL, Lee SH. 2010. Identification of population-specific QTLs for flowering in soybean. J. Crop Sci. Biotech. 13: 127–136
Malik M, Ashraf M, Qureshi A, Ghafoor A. 2007. Assessment of genetic variability, correlation and path analyses for yield and its components in soybean. Pak. J. Bot. 39: 405–413
Mansur LM, Lark KG, Kross H, Oliveira A. 1993. Interval map ping of quantitative trait loci for reproductive, morphological, and seed traits of soybean (Glycine max. L.). Theor. Appl. Genet. 86: 907–913
Mansur LM, Orf JH, Chase K, Jarvik T, Cregan PB, Lark KG. 1996. Genetic mapping of agronomic traits using recombi nant inbred lines of soybean. Crop Sci. 36: 1327–1336
Maughan P, Maroof M, Buss G. 1996. Molecular-marker analysis of seed-weight: genomic locations, gene action, and evidence for orthologous evolution among three legume species. Theor. Appl. Genet. 93: 574–579
Orf J, Chase K, Jarvik T, Mansur L, Cregan P, Adler F, Lark K. 1999. Genetics of soybean agronomic traits: I. Comparison of three related recombinant inbred populations. Crop Sci. 39: 1642–1651
Panthee D, Pantalone V, Saxton A, West D, Sams C. 2007. Quantitative traitloci for agronomic traits in soybean. Plant Breed. 126: 51–57
Paterson AH, Tanksley SD. 1991. DNA markers in plant improvement. Adv. Agron. 46: 39–89
Patterson DT, Peet MM, Bunce JA. 1977. Effect of photoperiod and size at flowering on vegetative growth and seed yield in soybean. Agron. J. 69: 631–635 SAS Institute Inc. 2002–2003. SAS User’s Guide, Version 9.1, SAS Institute Inc., Cary, NC
Sebolt A, Shoemaker R, Diers B. 2000. Analysis of a quantitative trait locus allele from wild soybean that increases seed protein concentration in soybean. Crop Sci. 40: 1438–1444
Sharma SL, Juneja SL. 1971. Correlation studies for yield and other characters in soyabeans (Glycine max. (L.) Merr.). Indian J. Agril. Res. 1: 40–45
Showkat M, Tyagi SD. 2010. Correlation and path coefficient analysis of some quantitative traits in soybean. Res. J. Agric. Sci. 1: 102–106
Shure M, Wessler S, Fedoroff N. 1983. Molecular identification and isolation of the waxy locus in maize. Cell 35: 225–233
Specht J, Chase K, Macrander M, Graef G, Chung J, Markwell J, Germann M, Orf J, Lark K. 2001. Soybean response to water: A QTL analysis of drought tolerance. Crop Sci. 41: 493
Suh SK, Kim HS, Oh YJ, Lee JJ, Lee SY, Kim SD, Jang YS, Cho SY. 1997. A disease resistant and high yielding soybean variety “Iksannamulkong” for sprouting. RDA J. Crop Sci. 39: 123–126
Tasma IM, Lorenzen L, Green D, Shoemaker RC. 2001. Mapping genetic loci for flowering time, maturity, and photoperiod insensitivity in soybean. Mol. Breed. 8: 25–35
Wang D, Graef G, Procopiuk A, Diers B. 2004. Identification of putative QTL that underlie yield in interspecific soybean backcross populations. Theor. Appl. Genet. 108: 458–467
Yamanaka N, Ninomiya S, Hoshi M, Tsubokura Y, Yano M, Nagamura Y, Sasaki T, Harada K. 2001. An informative link age map of soybean reveals QTLs for flowering time, leaflet morphology and regions of segregation distortion. DNA Res. 8: 61–72
Zhang D, Cheng H, Wang H, Zhang HY, Liu CY, Yu DY. 2010a. Identification of genomic regions determining flower and pod numbers development in soybean (Glycine max. L.). J. Genet. Genomics 37: 545–556
Zhang LY, Liu DC, Guo XL, Yang WL, Sun JZ, Wang DW, Zhang AM. 2010b. Genomic distribution of quantitative trait loci for yield and yield related traits in common wheat. J. Int. Plant Biol. 52: 996–1007
Zhang W, Wang Y, Luo G, Zhang J, He C, Wu X, Gai J, Chen S. 2004. QTL mapping of ten agronomic traits on the soy bean (Glycine max (L.) Merr.) genetic map and their association with EST markers. Theor. Appl. Genet. 108: 1131–1139
Zhao GY, Wang J, Han YP, Teng WL, Sun GL, Li WB. 2008. Identification of QTL underlying the resistance of soybean to pod borer, Leguminivora glycinivorella (Mats.) obraztsov, and correlations with plant, pod and seed traits. Euphytica 164: 275–282
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, W., Kim, M.Y., Van, K. et al. QTL identification of yield-related traits and their association with flowering and maturity in soybean. J. Crop Sci. Biotechnol. 14, 65–70 (2011). https://doi.org/10.1007/s12892-010-0115-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12892-010-0115-7