Abstract
The frequency of soybean white mold (Sclerotinia sclerotiorum, SWM) outbreaks has increased since 1978, and this disease is currently considered to be the second most important cause of soybean yield loss worldwide. We have studied SWM in stems of soybean cultivar Maple Arrow, which shows partial resistance to SWM, in an attempt to identify the quantitative trait loci (QTLs) underlying soluble pigment(s) content, which is associated with SWM resistance. A SWM-susceptible cultivar, Hefeng 25, was crossed with Maple Arrow, and 149 F5:6 recombinant inbred lines were subsequently advanced through single-seed-descent. A total of 109 simple sequence repeat (SSR) markers were used to construct a genetic linkage map. Three QTLs for soluble pigment content in soybean stems associated with the resistance to SWM, namely, Qsp-1 (Satt502-Sat_159), Qsp-2 (Sat_156-Satt251), and Qsp-3 (Satt525-Satt233), were identified in 2007 and 2008 and located onto linkage groups D1a+q, B1 and A2, respectively. The phenotypic variation (R 2) explained by these QTLs ranged from 6.29 to 15.37%. These three QTLs were not significantly related to known QTLs associated with escape resistant mechanisms. The use of these QTLs in marker-assisted selection may contribute to improved soybean resistance to SWM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean white mold (Sclerotinia sclerotiorum, SWM), also known as Sclerotinia stem rot, was first identified in Ontario, Canada in 1946 (Koch and Hildebrand 1946; Hildebrand 1948). The frequency of SWM outbreaks has increased since 1978, spreading over the entire world (Hine and Wheeler 1970; Signoret et al. 1975; Thompson and van der Westhuizen 1979; Grau et al. 1982; Cline and Jacobsen 1983; Boland and Hall 1986). This widespread occurrence has been attributed to the rotation of soybean in fields with a history of SWM of other crops (Grau et al. 1982; Phipps and Porter 1982), the favorable climate in soybean growing regions (higher humidity, cooler temperatures), and changes in cultural practices, such as decreased row spacing and/or the use of crop irrigation (Grau and Radke 1984). SWM is currently considered to be the second most important cause of soybean yield loss, surpassed only by soybean cyst nematode (Wrather et al. 1997). Current strategies, such as increased row spacing and reduced irrigation before and during crop flowering (Grau and Radke 1984) and biocontrol (Fernando et al. 2005), have not been effective in controlling this disease. A biotechnological approach consisting of the transformation of soybean with the germin gene, encoding the oligomeric protein oxalate oxide (OxO), which oxidizes oxalic acid to carbon dioxide and hydrogen peroxide (H2O2), does provide an effective means of controlling this disease (Donaldson et al. 2001). However, the legal obstacles to commercially growing transgenic soybean in most countries have limited this application. Hence, the selection of endogenously resistant cultivars and germplasm may be an alternative method to control SWM (Steadman 1979; Grau 1988).
To date, there is no known soybean genotype that is completely resistant to SWM (Boland and Hall 1987; Chun et al. 1987; Nelson et al. 1991; Gondran and Leclercq 1993; Kim et al. 1999). The genetics of differential resistance among soybean cultivars suggests a multi-locus model (Kim and Diers 2000), and both physiological resistance and escape mechanisms have been found to contribute to partial resistance in field trials. Escape mechanisms include early flowering and maturity, less lodging, and an upright, open canopy, and one or more of these mechanisms have been shown to be significantly associated with control of this disease (Boland and Hall 1987; Nelson et al. 1991; Kim et al. 1999; Kim and Diers 2000). Partial resistance can provide economically feasible disease control, and is one of breeding objectives in many soybean improvement programs.
Breeding for SWM resistance is difficult partly because of the low association between field and laboratory tests of resistance (Boland and Hall 1987; Wegulo et al. 1998; Kim and Diers 2000) and because resistance is often due to disease-avoidance traits rather than physiological resistance (Sutton and Deverall 1984; Boland and Hall 1987; Kim et al. 1999; Kim and Diers 2000). Molecular markers offer a faster and more accurate approach to breeding programs, since selection can be based on genotype rather than only on phenotype. The use of molecular markers for indirectly selecting important agronomic traits, or marker-assisted selection (MAS), can improve the efficiency of conventional plant breeding programs. Cregan et al. (1999) and Song et al. (2004) developed an integrated genetic linkage map of soybean that contains 1849 markers in one or more of five different populations and aligns the molecular linkage groups (MLGs) into a consensus map of 20 MLGs. These 20 MLGs correspond to the 20 pairs of soybean chromosomes (Zou et al. 2003), and this information had greatly facilitated MAS in soybean breeding.
Few recent studies have attempted to map quantitative trait loci (QTLs) associated with SWM resistance. Delaney et al. (1997) used a F2 population of Williams 82 × Corsoy 79 to analyze SWM resistance using randomly amplified polymorphic DNA (RAPD) markers and showed that one QTL on linkage group (LG) E was associated with SWM resistance. Kim and Diers (2000) reported three QTLs on LGs C2, K, and M associated with SWM resistance in a Williams 82 × S19-90 F3-derived population. Of these, two QTLs were also significantly associated with disease escape mechanisms, such as plant height, lodging, and date of flowering. Arahana et al. (2001) identified 28 putative QTLs for SWM resistance on 15 different LGs in five recombinant inbred line (RIL) populations by crossing Williams 82, a susceptible cultivar, with each of five cultivars that exhibited partial resistance: Corsoy 79, Dassel, DSR173, S19-90, and Vinton 81. Guo et al. (2008) analyzed the QTL for SWM resistance using two backcross populations, showing that four genome regions (Satt411-Satt369 on MLG E, Satt269-AW186493 on MLG F, Satt463-Satt323 on MLG M, and Satt581-Satt153 on MLG O) were associated with SWM resistance.
It is important to have stable phenotypic data for QTL mapping studies (Beavis 1998). However, the phenotypic data reported in many earlier studies often varies dues to different inoculation methods and different experimental conditions (controlled environment vs. field conditions (Wegulo et al. 1998). The method for measuring soluble pigment(s) content in the stem, based on the oxalic acid reaction, was developed by Wegulo et al. (1998) and produces consistent results in both controlled environmental and field conditions. Although soluble pigment(s) content in the stem has been found to be representative of a component of resistance to SWM, the effectiveness of this method using soluble pigment(s) content has proven useful in screening SWM resistance in soybean germplasm (Wegulo et al. 1998) and common bean (Kolkman and Kelly 2000).
In the study reported here, phenotypic data on soluble pigment(s) content in the stem, as reflected by the plant absorbance (PA) value, which is based on the OXO reaction, was used to identify the QTLs associated with SWM resistance in F5:6 and F5:7 RIL populations from a cross between soybean cvs. Hefeng 25 × Maple Arrow.
Materials and methods
Plant material
The mapping population, consisting of 149 F5:6 RILs, was advanced by single-seed-descent (SSD) from the cross of Hefeng 25 and Maple Arrow. Soybean cv. Hefeng 25 (the reference PA value for soluble pigment content was 0.004), developed by the Agricultural Academic Institute of Heilongjiang Province, China, is susceptible to SWM, and cv. Maple Arrow (reference PA value is 0.100) has partial resistance to SWM (Boland and Hall 1987).
Evaluation of disease symptoms and agronomic traits
The results of a preliminary experiment showed that the amount of a pink pigment dissolved in OxO from cv. Maple Arrow stems was greater than that in cv. Hefeng 25 stems (data not shown). In 2007 and 2008, the RILs and their parents were grown in the greenhouse in 50-cm-diameter buckets (six plants of one genotype or RIL per bucket; three buckets per replication) with three replications, in a randomized complete block design. The PA value was evaluated using the method described by Wegulo et al. (1998). At the V1 or V2 growth stage, the plant stem was severed approximately 0.5 cm above soil level, completely defoliated, and placed immediately in 13 × 100-mm test tubes containing 5 ml 40 mM oxalic acid (one stem per test tube). The PA value was measured for five of the plants in each bucket within each replicate of RILs. Test tubes containing soybean stems were arranged vertically in test tube racks and incubated at 20°C in under a 12/12-h ( light/dark) photoregime. After 48 h, 4 ml of the OxO in each test tube was transferred to a 5-ml cup, and the PA value was determined in a Perkin Lambda 35 spectrophotometer at 518 nm (Perkin–Elmer, Foster City, CA).
Plant height (PH), node number on the main stem (NN), number of pods per plant (PN), seed number per plant (SN), seed weight per plant (SW), maturity (MA), flowering time (FT), protein content (PC), and oil content (OC) were also measured. The PC and OC were measured using an quality analysis system instrument (Inframatic 8620; Perten, Segeltorp, Sweden). Each trait was measured on five plants in each bucket within each replicate of RIL.
Simple sequence repeat marker analysis
Total DNA of plants was isolated from frozen-dried leaf tissue using the CTAB method (Doyle and Doyle 1990). The simple sequence repeat (SSR) analysis was performed with primers developed by Cregan et al. (1999). The PCR analysis was performed in 20-μl reaction volumes containing 2 μl genomic DNA (25 ng/μl), 1.5 μl MgCl2 (25 mM), 0.3 μl dNTP mixtures (10 mM), 2 μl 10× PCR buffer, 2 μl SSR primer (2 μM), 0.2 μl Taq polymerase enzyme (10 U/μl), and 12 μl double distilled water. The amplification profile was 2 min at 94°C, followed by 35 cycles of 30 s at 94°C, 30 s at 47°C, 30 s at 72°C, then 5 min at 72°C. After amplification, the PCR product was mixed with loading buffer [2.5 mg/ml bromophenol blue, 2.5 mg/ml diphenylamine blue, 10 mM EDTA, 95% (v/v) formamide], denatured for 5 min at 94°C, and then put on ice for 5 min. The denatured PCR products were separated on a 6% (w/v) denaturing polyacrylamide gel and visualized by silver staining (Trigiano and Caetano-Anolles 1998).
Data analysis
Broad-sense heritability of PA was computed as \( h^{2} = h^{2}_{g} /\left( {h^{2}_{g} + h^{2}_{e} /n} \right) \) where \( h^{2}_{g} \) and \( h^{2}_{e} \) are the estimates of genetic and residual variance, respectively, derived from the expected mean squares of the variance, and n is the number of replications (Blum et al. 2001). The correlations between the PA value and the other agronomic traits in F5:6 RIL population of Hefeng 25 × Maple Arrow were analyzed using the SAS procedure PROC.COR.SAS (SAS Institute, Cary, NC), and the frequency distribution of PA in RILs was analyzed using the SAS procedure PROC.Shapiro-Wilk.SAS.
Mapmaker/EXP ver. 3.0b (Lander et al. 1987) was used for the genetic linkage analysis. The genetic linkage map was constructed using Mapchart ver. 2.1 (Voorrips 2002). The QTLs were analyzed using QTL Cartographer ver. 2.0 (Zeng 1993) with the composite interval mapping (CIM) module (Basten et al. 1996). The window size was 5 cM (Haldane units), and the walk speed was 1 cM. The threshold of the LOD score for evaluating the statistical significance of QTL effects was determined by 1000 permutations using the Zmapqtl program in QTL Cartographer (Churchill and Doerge 1994). A LOD value corresponding to an experiment-wise threshold of a = 0.05 was used to declare a QTL as significant. The estimate of the QTL position was the point of maximum LOD score in the region under consideration.
Results
Phenotypic analysis of SWM resistance
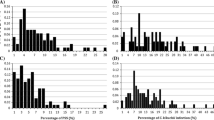
The resistance to SWM in Hefeng 25 × Maple Arrow RILs showed a continuous frequency distribution, ranging in PA value from a low of 0.001 to a high of 0.143 in 2007 and from a low of 0.001 to a high of 0.137 in 2008 (Fig. 1, Table 1). Cultivars Hefeng 25 and Maple Arrow differed significantly in terms of PA values (0.004 in 2007 and 0.005 in 2008 for Hefeng 25; 0.096 in 2007 and 0.100 in 2008 for Maple Arrow) (Table 1). Both the skewness and kurtosis values of the PA value were less than 1.0, suggesting that the segregation of this trait fits a normal distribution model (Table 1). The PA value between the 2 years was significantly correlated at the 0.05 level (R = 0.9426). The other agronomic traits also showed normal distributions with limited skewness and kurtosis (data not shown). The broad-sense heritability of PA value was 0.42 in 2007 and 0.44 in 2008.
Correlation analysis between the PA value and associated agronomic traits
Correlations among all the traits indicated that SWM resistance, as measured by PA value, was positively correlated with MA and FT at P < 0.01 and negatively correlated with SW per plant and SN per plant at P < 0.01 (Table 2).
Among the traits significantly correlated with PA value, MA and FT were also positively correlated with NN of the main stem, PN, SN per plant, and SW per plant (SW) at P < 0.01. Seed weight per plant and PC were negatively correlated with OC (Table 2).
Linkage analysis
A total of 600 SSR markers were used to screen polymorphisms between the two parents. One hundred and nine polymorphic SSR markers (18.5%) were mapped onto 19 LGs according to Cregan et al. (1999) and Song et al. (2004). The developed map encompassed 1848.34 cM, with an average distance of 16.96 cM between markers (data not shown).
Quantitative trait locus analysis of the PA value
Three QTLs associated with soluble pigment content in the stems, Qsp-1 (Satt502-Sat_159), Qsp-2 (Sat_156-Satt251), and Qsp-3 (Satt525-Satt233), were identified and located onto MLG D1a+q, B1, and A2, respectively (Fig. 2). Qsp-1 explained 12.55 and 10.39% of the phenotypic variation in 2007 and 2008, respectively and Qsp-2 explained 6.29 and 8.94%, respectively. Qsp-3 explained 7.99% of the phenotypic variations in 2008.
Quantitative trait locus analysis of related agronomic traits
A total of 19 QTLs associated with ten related traits of soluble pigment content, including PH, NN of main stem, PN, SN per plant, SW per plant, PC, OC, MA, and FT, were identified and anchored on eight linkage groups (MLG D1b+w, MLG F, MLG D2, MLG C2, MLG I, MLG C1, MLG M, MLG O) (Table 3).
Two QTLs associated with PH were identified on MLG D1b+w (Satt579-Satt282) and MLG I (Satt419-Satt127) and found to explain 8.09 and 9.82% of the phenotypic variability (Table 3). Three QTLs associated with NN were identified on MLG D2 (Satt458-Satt154), MLG C2 (Satt376-Satt286), and MLG I (Satt419-Satt127) and explained 6.13–22.57% of the phenotypic variability. Four QTLs associated with PN were identified on MLG D2 (Satt256-Satt458), MLG I (Satt292-Satt354), MLG N (Satt460-Satt079), and MLG C1 (Satt565-Satt180) and explained 2.49–11% of the phenotypic variability. Two QTLs associated with SN were identified on MLG D1b+W (Satt189-Satt350) and MLG C1 (Satt180-Satt524) and explained 5.99 and 10.28% of the phenotypic variability. Three QTLs associated with SW were identified on MLG M (Satt150-Satt201), MLG D2 (Satt256-Satt458), and MLG C1 (Satt565-Satt180) and explained 7.08–20.29% of the phenotypic variability. QTLs associated with MA and FT were found in the same regions between Satt072 and Sat_069 of MLG D1b+W, with the one for MA explaining 13.57 and 21.66% of the associated trait phenotypic variability in 2007 and 2008, respectively, and that for FT explaining 10.47% in 2008 (Table 3). The QTL associated with PC and OC is listed in Table 3.
Discussion
A number of studies have demonstrated that SWM can cause yield losses equal to those caused by the soybean cyst nematode, phytophthora root rot, and stem rot under the appropriate environmental conditions (Arahana et al. 2001; Grau et al. 2004). With each 10% increase in disease incidence, yield reductions have been found to average between 170 and 330 kg ha−1 (Hoffman et al. 1998). Partial resistance to these disease organisms can provide economically feasible disease control and is one of breeding objectives in soybean resistance improvement programs, given that there is at yet no soybean cultivar that demonstrates complete resistance to SWM (Kim and Diers 2000). The partial SWM resistance of soybean cv. Maple Arrow has been extensively reported (Boland and Hall 1987), and this cultivar has attracted the attention of plant breeders as the means to gaining an understanding of the resistance mechanisms and for transferring this resistance into commercial cultivars in northeastern China. Because resistance against SWM in soybean is difficult to be evaluated based on phenotype alone, we believe that increasing the selection intensity by the MAS of genotypes carrying the trait of soluble pigment content in stems (referred to as PA value) will lead to improved selection gain. To the best of our knowledge, little information is currently available on the QTL analysis of SWM resistance using PA methodology.
The levels of soluble pigments in soybean stems have generally been found to be inversely related to disease incidence in the field (Wegulo et al. 1998). Although the soluble pigment has not yet been isolated and characterized, the presence of anthocyanins in it is highly likely based on the similarity in color between the observed pigment and a number of anthocyanin pigments that have been isolated and characterized (Harborne 1965, 1967; Hrazdina 1982). Anthocyanins, a group of pigments widely distributed in plants and responsible for the pink, red, scarlet, mauve, violet, and blue colors (Harborne 1965, 1967; Hrazdina 1982), are products of the flavonoid biosynthesis pathway (Harborne 1965, 1967, 1976; Hrazdina 1982; Deikman and Hammer 1995). They have also been reported to be involved in disease resistance (Biehn et al. 1968a, b; Harborne 1976; Hammerschmidt and Nicholson 1977; Kraft 1977; Muehlbauer and Kraft 1978; Kumar et al. 1991). In an earlier study carried out by our group, 21 germin-like genes, encoding the oligomeric protein OxO, which can oxidize OxO to carbon dioxide and H2O2 and confer SWM resistance, were cloned from cv. Maple Arrow (data not shown). The association between germin or germin-like genes and soluble pigment in resistant cv. Maple Arrow conditioning on OxO is being presently investigated.
We identified three QTLs (Qsp-1, Qsp-2, and Qsp-3) that significantly affected the resistance to SWM based on the PA value. The phenotypic variation contributed by these QTLs was 12.55 and 6.29% in 2007 and 10.39, 8.94, and 15.37% in 2008, respectively. QTL Qsp-1 has not been reported earlier, while QTLs Qsp-2 and Qsp-3 fell into the same location as a QTL identified in a study that used other inoculation methods (Arahana et al. 2001; Guo et al. 2008). Selecting a resistance cultivar to SWM was difficult because of the interaction between escape mechanisms and physiological resistance (Arahana et al. 2001). Escape mechanisms, including flowering date, lodging, canopy architecture, and maturity, have all been shown to be significantly associated with disease severity (Boland and Hall 1987; Nelson et al. 1991; Kim et al. 1999), similar to the results of this study (Table 2). These escape mechanisms are influenced by environmental conditions, which makes the resistance of this cultivar variable. Hence, it was important to ascertain the proportion of physiological resistance and escape mechanisms in SWM resistance for selecting the resistance cultivar. The correlation between PA and the main agronomic traits was significant at the 0.01 levels in this study (Table 2). The three QTLs associated with SWM resistance in this study were not significantly associated with escape mechanisms (Fig. 2, Table 3), indicating that these three QTL affect resistance directly, rather than through escape mechanisms.
The environmental variability associated with field studies makes it difficult to obtain good estimates of resistance due to failure of the disease to develop, especially when the weather conditions are hot and dry. Markers tightly linked to resistance QTLs would facilitate plant breeders in their search to identify resistant soybean lines on the basis of genotype, maximizing the effectiveness of selection. Although the phenotypic variance explained by these resistance QTLs in this study was relatively small, combining these resistance QTLs in one cultivar could also be an important step towards improving soybean resistance to SWM. Resistance QTLs whose alleles exert smaller effects on phenotype may also be manipulated more effectively (Young 1996). Hence, both the resistance source Maple Arrow and the QTLs identified in this study can aid in improving resistance against SWM.
References
Arahana VS, Graef GL, Specht JE, Steadman JR, Eskridge KM (2001) Identification of QTLs for resistance to Sclerotinia sclerotiorum in soybean. Crop Sci 41:180–188
Basten CJ, Weir BS, Zeng ZB (1996) QTL cartographer. North Carolina State University, Raleigh
Beavis WD (1998) QTL analyses: power, precision, and accuracy. In: Paterson AH (ed) Molecular dissection of complex traits. CRC Press, Boca Raton, pp 145–162
Biehn WL, Kuc J, Williams EB (1968a) Accumulation of phenols in resistant plant-fungi interactions. Phytopathology 58:1255–1260
Biehn WL, Williams EB, Kuc J (1968b) Fungitoxicity of phenols accumulating in Glycine max–fungi interactions. Phytopathology 58:1261–1264
Blum A, Klueva N, Nguven HT (2001) Wheat cellular thermo tolerance is related to yield under heat stress. Euphytica 117:117–123
Boland GJ, Hall R (1986) Growth room evaluation of soybean cultivars for resistance to Sclerotinta sclerotiorum. Can J Plant Sci 66:559–564
Boland GJ, Hall R (1987) Evaluating soybean cultivars for resistance to Sclerotinia sclerotiorum under field conditions. Plant Dis 71:934–936
Chun D, Kao LB, Lockwood JL, Isleib TG (1987) Laboratory and field assessment of resistance in soybean to stem rot caused by Sclerotinia sclerotiorum. Plant Dis 71:811–815
Churchill RW, Doerge GA (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Cline MN, Jacobsen BJ (1983) Methods for evaluating soybean cultivars for resistance to Sclerotinia sclerotiorum. Plant Dis 67:784–786
Cregan PB, Jarvik T, Bush AL, Shoemaker RC, Lara KG, Kahler AL, Kaya N, VanToai TT, Lohnes DG, Chung J, Specht JE (1999) An integrated genetic linkage map of the soybean genome. Crop Sci 39:1464–1490
Deikman J, Hammer PE (1995) Induction of anthocyanin accumulation by cytokinins in Arabidopsis thaliana. Plant Physiol 108:47–57
Delaney DE, Graef GL, Rutledge S, Power K, Higgins B, Steadman JR, Diers B (1997) Mapping QTL associated with Sclerotinia stem rot resistance in soybean (abstract). Poster presented at Plant and Animal V, San Diego
Donaldson PA, Anderson T, Lane BG, Davidson A, Simmonds DH (2001) Soybean plants expressing an active oligomeric oxalate oxidase from the whieat gf-2.8(germin) gene are resistant to the oxalate-secreting pathogen Sclerotina sclerotiorum. Physiol Mol Plant P 59:297–307
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Fernando WGD, Ramarathnama R, Krishnamoorthyb AS, Savchuka SC (2005) Identification and use of potential bacterial organic antifungal volatiles in biocontrol. Soil Biol Biochem 37:955–964
Gondran J, Leclercq P (1993) Genetic variability of soybean for resistance to Sclerotinia sclerotiorum (Lib) de Bary. Statistical relationship with some traits of the crop. Agronomy 13(2):85–94
Grau CR (1988) Sclerotinia stem rot of soybean. In: Wyllie D, Scott DH (eds) Soybean diseases of the North Central region. American Phytopathological Society, St. Paul, pp 56–149
Grau CR, Radke VL (1984) Effects of cultivars and cultural practices on Sclerotinia stem rot of soybean. Plant Dis 68:56–58
Grau CR, Radke VL, Gillespie FL (1982) Resistance of soybean cultivars to Sclerotinia sclerotiorum. Plant Dis 66:506–508
Grau CR, Dorrance AE, Bond J, Russin J (2004) Fungal diseases. In: Boerma HR, Specht JE (eds) Soybeans: improvement, production and uses, 3rd edn. Agronomy Mongraph 16. ASA, CSSA, and SSSA, Madison, pp 679–763
Guo XM, Wang DC, Gordon SG, Helliwell E, Smith T, Berry SA, Martin SK, Dorrance AE (2008) Genetic mapping of QTLs underlying partial resistance to Sclerotinia sclerotiorum in soybean PI 391589A and PI 391589B. Crop Sci 48:1129–1139
Hammerschmidt R, Nicholson RL (1977) Resistance of maize to anthracnose: changes in host phenols and pigments. Phytopathology 67:251–258
Harborne JB (1965) Flavonoid pigments. In: Boner J, Varner JE (eds) Plant biochemistry. Academic Press, New York, pp 618–640
Harborne JB (1967) Comparative biochemistry of the flavonoids. Academic Press, London
Harborne JB (1976) Functions of flavonoids in plants. In: Goodwin TW (ed) Chemistry and biochemistry of plant pigments. Academic Press, London, pp 736–778
Hildebrand AA (1948) Soybean diseases in Ontario. Soybean Digest 10:16–17
Hine RB, Wheeler JE (1970) The occurrence of some previously unreported diseases in Arizona. Plant Dis Rep 54:179–180
Hoffman DD, Hartman GL, Mueller DS, Leitz RA, Nickell CD, Pedersen WL (1998) Yield and seed quality of soybean cultivars infected with Sclerotinia sclerotiorum. Plant Dis 82:826–829
Hrazdina G (1982) Anthocyanins. In: Harborne JB, Mabry TJ (eds) The flavonoids. Advances in research. Chapman and Hall, London, pp 135–188
Kim HS, Diers BW (2000) Inheritance of partial resistance to Sclerotinia stem rot in soybean. Crop Sci 40:55–61
Kim HS, Sneller CH, Diers BW (1999) Evaluation of soybean cultivars for resistance to Sclerotinia stem rot in field environments. Crop Sci 39:64–68
Koch LM, Hidebrand AA (1946) Soybean diseases in southwestern Ontario in 1946. Can Plant Dis Sur 26:27–28
Kolkman JM, Kelly JD (2000) An indirect test using oxalate to determine physiological resistance to white mold in common bean. Crop Sci 40:281–285
Kraft JM (1977) The role of delphinidin and sugars in the resistance of pea seedlings to Fusarium root rot. Phytopathology 67:1057–1061
Kumar J, Kaiser WJ, Hannan RM (1991) Damping-off resistance in chickpeas. Plant Dis 75:1244–1245
Lander ES, Green P, Abrahamson J, Baarlow A, Daly MJ, Lincoln SE, Newburg L (1987) MapMaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Muehlbauer FJ, Kraft JM (1978) Effect of pea seed genotype on pre-emergence damping off and resistance to Fusarium and Pythium root rot. Crop Sci 32:1–323
Nelson BD, Helms TC, Olson MA (1991) Comparison of laboratory and field evaluations of resistance in soybean to Sclerotinia sclerotiorum. Plant Dis 75:662–665
Phipps PM, Porter DM (1982) Sclerotinia blight of soybean caused by Sclerotinia minor and Scterotinia sclerotiorum. Plant Dis 66:163–165
Signoret PA, Bemaux PC, Poinso B (1975) Soybean diseases in France in 1974. Plant Dis Rep 59:616–617
Song QJ, Marek LF, Shoemaker RC, Lark KG, Concibido VC, Delannay X, Specht JE, Cregan PB (2004) A new integrated genetic linkage map of the soybean. Theor Appl Genet 109:122–128
Steadman JR (1979) Control of plant diseases caused by Sclerotinia species. Phytopathology 69:904–907
Sutton DC, Deverall BJ (1984) Phytoalexin accumulation during infection of bean and soybean by ascospores and mycelium of Sclerotinia sclerotiorum. Plant Pathol 33:337–383
Thompson AH, van der Westhuizen GCA (1979) Scterotinia sclerotiorum (Lib.) de Bary on soybean in South Africa. Phytophylactica 11:145–148
Trigiano RN, Caetano-Anolles G (1998) Laboratory exercises on DNA amplification fingerprinting for evaluating the molecular diversity of horticultural species. Hort Tech 8:413–423
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTL. Heredity 93:77–78
Wegulo SN, Yang XB, Martinson CA (1998) Soybean cultivar responses to Sclerotinia sclerotiorum in field and controlled environment studies. Plant Dis 82:1264–1270
Wrather JA, Anderson TR, Arsyad DM, Gai J, Ploper LD, Porta-Puglia A, Ram HH, Yorinori JT (1997) Soybean disease loss estimates for the top 10 soybean producing countries in 1994. Plant Dis 81:107–110
Young ND (1996) QTL mapping and quantitative disease resistance in plants. Annu Rev Phytopathol 34:479–501
Zeng Z (1993) Theoretical basis of separation of multiple linked gene effects on mapping quantitative trait loci. Proc Natl Acad Sci USA 90:10972–10976
Zou J, Lee J, Singh R, Xu SS, Cregan PB, Hymowitz T (2003) Assignment of molecular linkage groups to the soybean chromosomes by primary trisomics. Theor Appl Genet 107:745–750
Acknowledgments
This study was conducted in the Key Laboratory of Soybean Biology of Chinese Education Ministry and financially supported by National 863 Projects (Contract No. 2006AA10Z1F1 and 2006AA100104-4), 948 project of Agricultural ministry of China (Contract No. 2006-G5) and National International Cooperation Project.
Author information
Authors and Affiliations
Corresponding author
Additional information
Dongmei Li, Mingming Sun and Yingpeng Han have made equal contributions to this work.
Rights and permissions
About this article
Cite this article
Li, D., Sun, M., Han, Y. et al. Identification of QTL underlying soluble pigment content in soybean stems related to resistance to soybean white mold (Sclerotinia sclerotiorum). Euphytica 172, 49–57 (2010). https://doi.org/10.1007/s10681-009-0036-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-009-0036-z