Abstract
Purpose
Pea hulls have a significant contribution to agricultural waste and are left unattended. It has a rich nutrient media and may have application in waste utilization. In this investigation, the potential of pea hulls for the production of cellulase has been explored.
Methods
The effect of H2O2 concentration, agitation speed and harvesting time on cellulase production by Trichoderma reesei QM9414 under submerged fermentation at constant pH of 5.0 and temperature 30 ± 0.1 °C were studied.
Results
The maximum filter paper (FP) cellulase activity of 0.372 ± 0.019 U/mL was obtained at 91 h incubation period and 120 rpm agitation speed. Based on the optimized results of fermentation parameters, 120 rpm agitation speed, 5% H2O2 concentration and 91 h harvesting time was recommended for efficient cellulase production. The effect of harvesting time on protein, reducing sugar and cellulase activity was pre-dominant. The purified cellulase enzyme specific activity was recorded as 13.8 U/mL.
Conclusions
The results indicate that the production of cellulase from green pea hulls may provide a novel and economical solution for industrial waste disposal.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Increasing concern for pollution that occurs from agricultural and industrial wastes has stimulated interest in converting waste materials into commercially valuable products. Agricultural wastes are widely available in the form of crop residues (residual stalks, straw, leaves, roots, husks, sheaths, bagasse etc.) and are easily renewable. The food industry produces large volumes of wastes (both solids and liquids) which have potential for recycling and conversion into different products [1]. Agriculture waste being organic in nature is easily assimilated by microorganisms mainly fungi which make it appropriate for enzyme production. Lignocellulosic biomass such as carrot peel, onion peel, potato peel and wheat straw among others are known to be good source for microbial enzyme production. Pea hulls are another such contributor to agricultural waste, the potential of which attracts attention. Pea hulls are waste, generated after separation of pea (Pisum sativum) seeds and represents up to 60% of total pea. These hulls are rich in dietary fiber, twice as much as wheat bran [2] and are mostly produced by frozen food processing industries. It is a rich source of nutrition with 3.8% protein, 41.1% glucose, 1.7% ash, 1.0% acetic acid and 0.5% methanol in its dry matter composition [3]. Furthermore, they are lightly colored and tasteless which makes them a good source of fiber and carbon with 82.30% total dietary fiber, 62.3% cellulose and 8.2% hemicellulose [4]. Pea hulls may therefore be utilized for manufacturing of ethanol, starch, dietary fiber, enzymes and other value added products after fermentation. Since, the ultimate goal is to develop a commercial low cost technique for processing pea hull waste; submerged fermentation may be preferred over solid state fermentation. In submerged fermentation, substrate is used in a slurry form making it economical due to its lower capital and operating expenses and higher productivity [5, 6]. Industrially important enzymes have traditionally been obtained from submerged fermentation (SmF) because of the ease of handling and greater control of environmental factors such as temperature and pH. The production of cellulase can be enhanced by choosing an appropriate fungal strain complimentary to the substrate. Pea hull could be a better carbon source for cellulase enzyme using Trichoderma reesei. Cellulase produced by T. reesei hydrolyzes 91% of the cellulose in pretreated wheat straw within 24 h [7] showing good capability for enzyme production.
The use of rice straw [8, 9], rice bran [10, 11], wheat bran [12,13,14] olive processing residue [15], onion peel, potato peel, pineapple waste [16], sugarcane bagasse [17], sugar beet pulp [18] as a substrate for T. reesei has been reported in literatures. However, the use of pea hull as a sole source of carbon for this microorganism remains unexplored. The objective of this study therefore, is to examine the effect of fermenting parameters on cellulase yield from peahull using T. reseei under submerged fermentation. This study attempts to examine and optimize a process which could make pea production economically more viable, minimizing the problems of pollution and waste disposal in food industries.
Materials and Methods
Fungal Strain and Inoculum
Different strains have different tolerance limit for utilization of cellulose and sugar concentration as carbon source. For the present study, T. reesei QM9414 was used for the production of cellulase enzyme. Trichoderma reesei QM9414 strain was obtained from IMTECH, Chandigarh on the basis of their enzyme production properties. For experimental purposes, spore suspension was prepared by incubating the cultures on potato dextrose agar (PDA) plates at 30 °C for 5–6 days until sufficient sporulation was observed. The spores were harvested using distilled water and the count of approximately 108 spores/mL was used for inoculation.
Experimental Design
Agitation speed (93, 100, 110, 120, 127 rpm), H2O2 pre-treatment concentration (3.64, 5, 7, 9, 10.36%) and harvesting time (31, 48, 72, 96, 112 h) were identified as the most important process variables as per literature review. Some parameters for the experiments were kept constant as described in Table 1. Central composite rotatable design (CCRD) was adopted for a three variable-5 level (− a, − 1, 0, + 1, + a) problem computing to a total of 20 experiments with six set of replicates. The terminal levels (− a and + a) were calculated as a = 20.25v, where v is the number of variables. Coded values (Cv) of levels were converted to actual values (Av) using Eq. (1), where H and L represent Av corresponding to + 1 and − 1. The experimental design matrix has been shown in Table 2.
Substrate Procurement and Pre-treatment
Pea pods were procured from the local market of Pantnagar, as per requirement and separation of peas from pea pods was done manually. Slurry was made by grinding the peahulls in a food processor (Sujata-make, 810 W motor; 500 mL capacity). No additional water was added, as sufficient amount of moisture was present in the peahulls. The sample were taken in conical flash, tightly sealed with cotton plug and placed in an autoclave for 10 min to avoid solubilization. During thermal treatment, the disruption of cell wall, cell membrane and bond hydrolization occurs by the phenomenon of blanching, resulting in greater availability of cellulosic as well as antioxidant and phenolic constituents. Under these critical situations, T. reesei has the ability to grow well. Literature suggests, T. reesei has the tolerance to grow in the phenolic environment up to a certain level [19]. After thermal treatments the slurry was cooled to 25 °C. Different concentrations of H2O2 (3.64, 5, 7, 9, 10.36% v/v) was then added to 100 g of slurry and taken in a 250 mL conical flask for alkaline treatment. Alkaline treatment results in the disruption of the lignin seal to increase enzyme access to holocellulose (residue after delignification), reduction of cellulose crystallinity, increase in surface area and porosity of the substrate and also the hydrolysis rate. The pH of the substrate was adjusted to 5.0 by using HCl from an initial pH of 7.23.
Estimation of Reducing Sugar
Sugar was estimated by DNS method [20]. DNS reagent was prepared and standard protocol was followed [21]. Fermented samples were withdrawn and centrifuged at 9000 rpm for 20 min. One milliliters of the centrifuged sample was taken to which 3 mL of freshly prepared DNS reagent was added. The contents were thoroughly mixed and placed in a water bath at 90 °C for 10 min. The tubes were taken out and cooled immediately and the contents of the tube were diluted if required. The absorbance was measured using a UV–Vis spectrophotometer. A reference blank was prepared containing 1 mL distilled water in place of sample extract. The concentration of sugar in the sample was estimated by computing the absorbance at 540 nm against the standard curve of glucose.
Estimation of Protein
Protein was estimated by Lowry’s method and standard reagents A (copper–tartarate carbonate complex) and B (Folin–Ciocalteu reagent) were prepared [22]. The samples were withdrawn after fermentation and centrifuged at 9000 rpm for 20 min. Five milliliters of reagent A was then added to 1 mL of the supernatant collected from the centrifuged sample. The tubes were shaken vigorously and allowed to stand for 10 min following which 0.5 mL of reagent B was added and mixed well. The final solutions were incubated at room temperature under dark conditions for 30 min till the development of blue color due to the formation of heteropolymolybdenum blue complex. Absorbance measurements were taken at 660 nm against blank using UV–Vis spectrophotometer.
Determination of Enzyme Activity
Cellulase activity was measured using the filter paper activity (FPA) assay, expressed in filter paper units (FPU) as described by [23]. This method measures the release of reducing sugar produced in 60 min from a mixture of enzyme solution (1 mL) and of citrate buffer (0.05 M, pH 4.8) in the presence of 50 mg Whatman No.1 filter paper (1 × 6 cm strip) and incubated at 50 °C. Sample withdrawn after fermentation was centrifuged at 9000 rpm for 20 min. The supernatant (solution) collected from the centrifuged sample was treated as enzyme source. The reaction was terminated by adding DNS reagent (3 mL). A reagent blank (1.5 mL citrate buffer), enzyme blank (1.0 mL citrate buffer + 1.0 mL enzyme dilution) and substrate blank (1.5 mL citrate buffer + filter-paper strip) were prepared and DNS reagent (3 mL) was added to them. All the samples were boiled for 5 min in a water bath and then cooled at room temperature for colour stabilization. The tube contents were then diluted with distilled water (15 mL), mixed thoroughly and allowed to stand for 20 min until the pulp was settled. The absorbance was recorded at 540 nm and values were quantified using the standard curve calibration after subtraction of enzyme and substrate blank. The reducing sugars released were determined using the 3, 5-dinitrosalicylic acid (DNS) assay with glucose as a standard [20].
Enzyme Purification
All procedures of cellulase purification were carried out at 4 °C. The culture supernatant was separated by centrifugation at 9500 rpm for 15 min. After obtaining maximum clarity, solid crystals of ammonium sulphate were added to the crude enzyme extract until 60% saturation and kept for over 12 h. The resulting precipitate was collected by centrifugation at 9500 rpm for 15 min. After centrifugation, the supernatant was separated and the sediments were dissolved in 0.05 M citrate buffer (pH 4.8). Ten mL solution was kept in a dialysis bag which was then sealed securely and dialyzed against citrate buffer. The buffer was changed every 4 h for a period of 12 h. The specific activity of partially purified cellulase was determined before and after dialysis. The final sample was lyophilized at a temperature of − 40 °C and 0.25 mbar pressure before storage at 4 °C for further use.
Statistical Analysis
Experiments were conducted randomly. The experimental design chosen (Table 2) consisted of six replicates from which the standard deviation (SD) was computed. The same SD was assumed for all experiments due to design symmetry. Mean values for all parameters were examined for significance by analysis of variance (ANOVA). The statistical data (significant model fits, simulated graphs) was generated using Design Expert software v.9.0.6.2 (Stat-Ease) at 1, 5 and 10% level of significance. The response was expressed as a second-order polynomial as described by Eq. (2). Mathematical expressions for the coefficient of determination (R2) and adjusted R2 or R2adj have been shown in Eqs. (3) and (4). The use of R2adj may be justified due to non-linearity in data which may not be accurately determined by the value of R2.
Results and Discussion
Reducing sugar, protein content and cellulase activity were estimated using analytical methods described in Materials and Methods (Estimation of reducing sugar, Estimation of protein and Determination of enzyme activity respectively). The responses obtained for different sets of process conditions have been shown in Table 3. The significance of the effect of individual parameters on the responses has been shown in Table 4. Cellulase produced amounted to 0.172 mg from 10 mL of partial purified sample of fermented peahull slurry (100 g). The purified enzyme had specific cellulase activity 13.8 U/mL. These results were higher or in close agreement with reported findings [16, 24,25,26].
Cellulase Activity
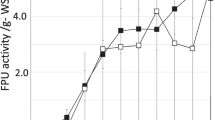
Cellulase production was compared under various conditions with T. reesei. Maximum cellulase activity of 0.315 ± 0.019 U/mL was obtained at 110 rpm, 7% H2O2 concentration and 72 h harvesting time (Table 3) which was comparable to the findings of Bhavna [21] and Nema [27] but lower than Mrudula [28] and Tallapragada [29]. The reported difference in cellulase activity under submerged fermentation may vary due to variation in substrate considered. The results show that the production of enzyme is slow at the beginning of fermentation and then increased with fermentation. As the alkaline concentration increased upto a particular range, the growth of T. reesei also increased. This could be attributed to the increase in crystallinity of cellulose and removal of lignin and other inhibitors. Thereby, cells utilized cellulose as a carbon source and increased the production of cellulase. At lower harvesting time, the cellulase activity was reduced. This indicated that short reaction times may not be sufficient for the degradation of cellulose or crystalline fraction of cellulose. Thus, the hydrolysis time should be sufficient for the degradation of cellulose to increase the enzyme production [30,31,32]. The effect of harvesting time on cellulase activity was observed to be statistically significant (p < 0.01). The interaction effect of variables were found to be non-significant (p > 0.05) which shows no synergy between variables which could contribute to increase in cellulase activity. However, minor interaction between H2O2 concentration and harvesting time (p < 0.1) was observed (Table 4) which indicate that substrate pre-treatment may alter enzyme production and activity. A linear model (Fig. 1) showed better fit (92.87%) than a non-linear model (86%).
Reducing Sugar
Results of reducing sugar of different experimental runs are shown in Table 3. During fermentation, T. reesei utilized reducing sugar for their growth. Utilization of sugar is an important consideration during fermentation, as it is linked to cellulase yield. Reports show that the accumulation of reducing sugar represses the production of cellulase [23]. The reducing sugar ranged from 1.03 to 3.55 ± 0.15 mg/mL which was in accordance with the findings of John [33] and Wang [34]. The maximum reducing sugar was observed for the samples having 31 h harvesting time, H2O2 conc. of 7% and 110 rpm agitation speed. The variation in reducing sugar depended on the cell growth as suggested by Raimbault [35]. The utilization of reducing sugar increased with fermentation and harvesting time to increase the desired production. The growth of T. reesei lowered the pH which complimented the utilization of reducing sugar rapidly thereby increasing the cellulase production [36]. Significant decrease (p < 0.01) in reducing sugar was observed with increase in harvesting time. As no interaction terms were found significant (Table 4), a linear model was found best suited (Fig. 2) to fit the data (97.72%). Comparable values of R2adj and R2 further pointed towards non-linearity in data.
Protein Content
Results of different experimental conditions and observed values of protein are shown in Table 3. The protein content ranged from 0.89 to 1.78 ± 0.07 mg/mL which was higher than that reported by Ghosh [37]. Maximum protein for the fermented peahull slurry was observed for an agitation speed of 110 rpm, H2O2 conc 7% and harvesting time 112 h, while minimum protein was observed for experimental condition with agitation speed 110 rpm, H2O2 conc 7% and harvesting time 31 h. As per observations, the protein content of the biomass increased with harvesting time. These finding are in accordance with [36] who reported that the maximum protein production was observed on the fifth day of fermentation by Aspergillus niger (KA-06) and thereafter the protein yield declined. The protein content increased with increase in agitation speed upto a threshold level beyond which it decreased gradually (Fig. 3a). This could be explained due to increase in shear stress on higher speeds which could damage the fungal hyphae thereby affecting the overall protein content [38]. All process variables affected the protein content significantly (Table 5) with higher influence of harvesting time (p < 0.01) and agitation speed (p < 0.05) than H2O2 concentration. Figures 3a and b shows the variation in protein content with X1 and X3 respectively. The interaction effects were non-significant (p > 0.1). H2O2 concentration contributed to declining non-linear variations in data significantly (p < 0.01) followed by agitation speed (p < 0.05). This showed that lower pre-treatment concentrations and agitation speed are beneficial for higher enzyme production. A non-linear polynomial is therefore suggested in prediction of protein content (Fig. 3a).
Optimization of Fermenting Parameters
The objective of the study was to optimize the process parameters to get the best possible combination of independent variables that would result in a cellulase production with better efficiency. Design Expert software 9.0.6.2 was used to analyze data to obtain the optimized solution for cellulase production. The optimized conditions could be a single point or a range of points in which all the possible combinations would yield good results. While using any optimization technique, some constraints set the guideline to get the desired results. The response values and analysis of models give the valuable information in deciding the constraints for independent variables and responses. In the present study, experiments were carried out using CCRD in realistic vicinity to locate the true optimal values of multiple independent variables. Table 5 represents the goals which were fixed for all independent variables and dependent variable as per the objective of the study.
Optimization of the process variables yielded 58 solution sets out of which a single set most suited to the criteria was selected. The optimum level of independent variables in coded and actual form is given in Table 6.
The optimized set of independent variables (agitation speed, H2O2 conc and harvesting time) obtained from Design-Expert software, was used to verify the results. The values of the responses (reducing sugar, protein and cellulase activity) were compared with the values given by the software after optimization. The results revealed that the actual values of all attributes were close to the predicted value (Table 7) and ANOVA showed that variation between predicted and actual score was highly non-significant (Ftab > Fcal). Thus, the production of cellulase enzyme from pre-treated peahulls at 120 rpm agitation speed, 5% H2O2 concentration and 91 h harvesting time may be considered optimum for cellulase production. The final cellulase obtained after dialysis and lyophilization is shown in Fig. 4.
Conclusions
A novel, environment friendly and low-cost approach to pea hull waste utilization for enzyme production has been discussed. Optimized fermentation parameters were identified for enhanced production of cellulase from pre-treated pea hulls. The described work may aid to provide relevant information to food process industries for solid and liquid waste management while enhancing their profit margin through value-added products. Moreover, the cellulase produced could have numerous applications in textile, detergent, paper and food industries. Application of recombinant cultures or phenol degrading microbes along with cellulase producing cultures could be investigated as another effective technique for utilization of lignin enriched waste.
Abbreviations
- β 0, β i, β ij, β ii :
-
Model coefficients
- p:
-
Number of explanatory variables (excluding constants)
- y i :
-
Observed data
- \({\bar {y}_l}\) :
-
Mean of observed data
- \({\hat {y}_l}\) :
-
Predicted data
- N:
-
Sample size
- X i,j :
-
Independent variables
- Y :
-
Dependent variable
References
Rashad, M.M., Nooman, M.U.: Production, purification and characterization of extracellular invertase from Saccharomyses Cerevisiae NRRL Y- 12632 by solid-state fermentation of red carrot residue. Aus. J. Basic and Appl. Sci. 3(3), 1910–1919 (2009)
Arrigoni, E., Caprez, A., Amado, R., Neukom, H.: Food hydrocolloids. Food Biochem. 1, 57–64 (1986)
Ralet, M.C., Valle, D.G., Thibault, J.F.: Raw and extruded fiber from pea hulls. I. Composition and physico-chemical properties. Carbohydr. Polym. 20, 17–23 (1993)
Sosulski, F.W., Wu, K.K.: High-fiber breads containing field pea hulls, wheat, corn, and wild oat brans. Cereal Chem. 65(3), 186–191 (1988)
Gao, J.: Production and characterization of cellulolytic enzymes from the thermoacidophilic fungal Aspergillus terreus M11 under solid state cultivation of corn stover. Biores. Technol. 99, 3634–3637 (2008)
Mushimiyimana, I., Tallapragada, P.: Agro wastes residues as strategy to produce cellulase. Int. J. Chem.Tech. Res. 8(1), 89–97 (2015)
El-Shishtawy, R.M., Mohamed, S.A., Asiri, A.M., Gomaa, A.M., Ibrahim, H.I., Tahhi, A.H.: Saccharification and hydrolytic enzyme production of alkali pre-treated wheat bran by Trichoderma virens under solid state fermentation. BMC Biotechnol. 15, 37 (2015)
Muthuvelayudham, R., Viruthagiri, T.: Fermentative production and kinetics of cellulase protein on Trichoderma reesei using sugarcane bagasse and rice straw. Afr. J. Biotechnol. 5(20), 1873–1881 (2006)
Fatima, G., Satinder, K.B., Tyagi, R.D., Verma, M., Surampalli, R.Y.: Screening of agro-industrial wastes to produce ligninolytic enzymes by Phanerochaete chrysosporium. Biochem. Eng. J. 49, 388–394 (2010)
Esfahani Hamidi, Z., Salimi Rocky, K.: Evaluation of the effect of particle size, aeration rate and harvest time on the production of cellulase by Trichoderma reesei QM9414 using response surface methodology. Food Bioprod. Process. 88, 61–66 (2010)
Verma, N., Bansal, C., Mukesh Kumar, V.: Pea peel waste: a lignocellulosic waste and its utility in cellulase production by Trichoderma reesei under solid state cultivation. Bio Resour. 6(2), 1505–1519 (2011)
Kang, S.W., Park, Y.S., Lee, J.S., Hong, S.I., Kim, S.W.: Production of cellulases and hemicellulases by A.niger KK2 from lignocellulosic biomass. Biores. Technol. 91, 153–156 (2004)
Javed, I.M.M., Khant, T.S.: An innovative approach for hyperproduction of cellulolytic and hemicellulolytic enzymes by consortium of Aspergillus niger MSK-7 and Trichoderma viride MSK-10. Afr. J. Biotechnol. 5(8), 609–614 (2006)
Dhillon, G.S., Oberoi, H.S., Kaur, S., Bansal, S., Brar, S.K.: Value-addition of agricultural wastes for augmented cellulase and xylanase production through solid-state tray fermentation employing mixed-culture of fungi. Ind. Crops Prod. 34, 1160–1167 (2011)
Damato, G., Vivona, G., Stoller, M., Bubbico, R., Bravi, M.: Cellulase production from olive processing residues. Chem. Eng. Trans. 20, 978–988 (2010)
Saravanan, P., Muthuvelayudham, R., Rajesh, K.R., Viruthagiri, T.: Optimization of cellulase production using Trichoderma reesei by RSM and comparison with genetic algorithm. Front. Chem. Sci. Eng. 6(4), 443–452 (2012)
Sukumaran, R.K., Singhania, R.R., Mathew, G.M., Pandey, A.: Cellulase production using biomass feedstock and its application in lignocellulose saccharification for bio-ethanol production. Renew. Energy. 34(2), 421–424 (2006)
Moosavi-Nasab, M., Majdi-Nasab, M.: Cellulase production by Trichoderma reesei using sugar beet pulp. Iran Agri. Res. 25(1–2), 107–116 (2007)
Selvakumar, G., Saha, S., Kundu, S.: Inhibitory activity of pine needle extracts on some agriculturally resourceful microbes. Ind. J. Microbiol. 47(3), 267–270 (2007)
Miller, G.L.: Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 31, 426–428 (1959)
Bhavna, M.V., Magar, J.G.: Use of agricultural wastes for cellulases production by Aspergillus niger with submerged and solid state fermentation. Bionano Front. 3, 189–192 (2010)
Lowry, O.H., Rosebrough, N.J., Farr, A.L., Randall, R.J.: Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193, 265–275 (1951)
Ghose, T.K.: Measurements of cellulase activities. Pure Appl. Chem. 59, 257–268 (1987)
Singhania, R.R., Sukumaran, R.K., Patel, A.K., Larroche, C., Pandey, A.: Advancement and comparative profiles in the production technologies using solid-state and submerged fermentation for microbial cellulases. Enzyme Microb. Tech. 46, 541–549 (2010)
Jhadhav, A.R., Girde, A.V., More, S.M., More, S.B., Khan, S.: Cellulase production by utilizing agriculture wastes. Res. J. Agri. For. Sci. 1(7), 6–9 (2013)
Devi, M.C., Kumar, M.S.: Production, Optimization and partial purification of cellulase by Aspergillus niger fermented with paper and timber sawmill industrial wastes. J. Microbiol. Biotechnol. Res. 2, 120–128 (2012)
Nema, N., Alamir, L., Mohammad, M.: Production of cellulase from Bacillus cereus by submerged fermentation using corn husks as substrates. Int. Food Res. J. 22, 1831–1836 (2015)
Mrudula, S., Murugammal, R.: Production of cellulase by Aspergillus niger under submerged and solid state fermentation using coir waste as a substrate. Braz. J. Microbiol. 42, 1119–1127 (2011)
Tallapragada, P., Venkatesh, K.: : Isolation, identification and optimization of xylanase enzyme produced by aspergillus niger under submerged fermentation. J. Microbiol. Biotechnol. Res. 1, 137–147 (2011)
Shen, D., Xiao, R., Gu, S., Luo, K.: The pyrolytic behavior of cellulose in lignocellulosic biomass: a review. RSC Adv. 1, 1641–1660 (2011)
Zhang, P.Y.H., Himmel, E., Michael, M.R.J.: Outlook for cellulase improvement: screening and selection strategies. Biotechnol. Adv. 24, 452–481 (2006)
Rabelo, S.C., Carrere, H., Filno, M.R., Costa, A.C.: Production of bioethanol, methane and heat from sugarcane bagasse in a biorefinery concept. Biores. Technol. 102, 7887–7895 (2011)
John, R.P., Nampoothiri, M., Pandey, A.: Solid-state fermentation for l-lactic acid production from agro wastes using Lactobacillus delbrueckii. Process Biochem. 41, 759–763 (2006)
Wang, Q., Wang, X., Wang, X., Ma, H.: Glucoamylase production from food waste by Aspergillus niger under submerged fermentation. Process Biochem. 43, 280–286 (2008)
Raimbault, M.: Solid State Fermentation: Growth Filamentous Fungi on Starch Substrate, pp. 127–291. ORSTOM, Paris (1981)
Santhanam, A., Yalentesfa, B., Alemu, T.: Solid substrate fermentation and conversion of orange waste into fungal biomass using Aspergillus niger KA-06 and Chaetomium Spp KC-06. Afr. J. Microbiol. Res. 4(12), 1275–1281 (2010)
Ghosh, P., Ghosh, U.: Statistical optimization of laccase production by Aspergillus flavus PUF5 through submerged fermentation using agro-waste as cheap substrate. Acta Biol. Szeged. 61, 25–33 (2017)
Fratebianchi, D., Crespo, J.M., Tari, C., Cavalitto, S.: Control of agitation rate and aeration for enhanced polygalacturonase production in submerged fermentation by Aspergillus sojae using agro-industrial wastes. J. Chem. Technol. Biotechnol. 92, 305–310 (2016)
Acknowledgements
This work was submitted as thesis in partial fulfillment for the requirement of the degree of Master of Technology of the first author. The authors thankfully acknowledge the financial assistance received under University funded project, G. B. Pant University of Agriculture & Technology, Pantnagar. We extend our gratitude to all the colleagues of the department for helping us in the manual separation of pea from pea pods.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sirohi, R., Singh, A., Tarafdar, A. et al. Cellulase Production from Pre-treated Pea Hulls Using Trichoderma reesei Under Submerged Fermentation. Waste Biomass Valor 10, 2651–2659 (2019). https://doi.org/10.1007/s12649-018-0271-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12649-018-0271-4