Abstract
The characterization of bacterial enzymatic pathways of phenol metabolism is important to better understand phenol biodegradation. Phenol hydroxylase is the first enzyme involved in the oxidative metabolism of phenol, followed by further degradation via either meta- or ortho-pathways. In this study, the first known instance of phenol degradation via the meta-pathway by a member of the genus Acinetobacter (Acinetobacter sp. strain AQ5NOL 1) is reported. Phenol hydroxylase converts phenol to catechol, which is then converted via the meta-pathway to 2-hydroxymuconic semialdehyde by the catechol 2,3-dioxygenase enzyme. Phenol hydroxylase extracted from strain AQ5NOL 1 was fully purified using DEAE-Sepharose®, DEAE-Sephadex®, Q-Sepharose® and Zorbax® Bioseries GF-250 gel filtration and was demonstrated by SDS-PAGE to have a molecular weight of 50 kDa. The phenol hydroxylase was purified to about 210.51 fold. The optimum pH and temperature for enzyme activities are 20 °C and 7–7.5, respectively. The apparent K m and V max values of phenol hydroxylase with phenol as the substrate were 13.4 µM and 2.5 µmol min−1 mg−1, respectively. The enzyme was stable at −20 °C for 36 days.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Phenol and its derivatives are commonly found in the aqueous environment, land and in the biosphere. They have a wide number of applications in various industries, and many of the industrial effluents with high phenol level come from oil refineries, phenolic resin production, plastic, pesticide, textile and coke oven industries (González et al. 2001; Hori et al. 2006). The environmental-friendly technologies of bioremediation are gaining an increasing prominence for their potential in the safe remediation of phenolic compounds. Microorganisms such as bacteria, fungi and yeast have evolved various means of extracting essential nutrients from the environment and can be used to degrade or treat phenolic pollutants (Carvalho et al. 2009; Yotova et al. 2009; Chakraborty et al. 2010; Arif et al. 2013; Suhaila et al. 2013; Norazah et al. 2015; Nawawi et al. 2016). In previous works, Gram-negative bacteria which belong to the genera Pseudomonas and Acinetobacter have shown the ability to degrade phenolic compounds such as phenol (Hannaford and Kuek 1999; Dong et al. 2008), chlorophenol (Farrell and Quilty 2002; Hao et al. 2002), pentachlorophenol (Cassidy et al. 1997; Sharma et al. 2009), catechol (Briganti et al. 1997; Kagle and Hay 2006) and hydrocarbon (Salleh et al. 2003: Etoumi et al. 2008).
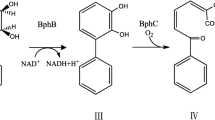
Bacteria have the ability to convert phenol into non-toxic intermediates either anaerobically, via a single pathway or aerobically, via ortho- or meta-pathway (Basha et al. 2010). In aerobic metabolism, the first step for both routes is the monohydroxylation of the aromatic ring’s ortho position. Phenol hydroxylase (EC 1.14.13.7) is responsible for converting phenol into catechol, which is the initial and rate-limiting step in phenol degradation pathways (van Schie and Young 2000; Tuan et al. 2011). Next, catechol is further degraded by catechol dioxygenase via meta- or ortho-cleavage pathways. Catechol 1,2-dioxygenase (EC 1.13.11.1) metabolizes catechol to form cis, cis-muconate by ortho-cleavage pathway. Catechol 2,3-dioxygenase (EC 1.13.11.2) metabolizes catechol to form 2-hydroxymuconic semialdehyde by meta-cleavage pathway (Basha et al. 2010). A number of phenol hydroxylase have been purified from a wide variety of microorganisms, such as Acinetobacter radioresistens (Divari et al. 2003), Acinetobacter calcoacetius (Paller et al. 1995), Acinetobacter sp. (Dong et al. 2008), Alcaligenes faecalis (Zhu et al. 2008), Bacillus thermoglucosidasius (Kirchner et al. 2003), Candida albicans (Tsai et al. 2005), Pseudomonas putida (Viggor et al. 2008), Pseudomonas sp. (Kagle and Hay 2006), Rhodococcus erythropolis (Saa et al. 2009), Pseudomonas fluorescens (Viggor et al. 2008) and Trametes versicolor (Yemendzhiev et al. 2008).
Strain AQ5NOL 1 Acinetobacter sp. has a high ability to degrade phenol up to 1100 and 1900 mg L−1 by free and immobilised cells in gellan gum beads, respectively (Ahmad et al. 2011, 2012). Moreover, this strain is able to tolerate the presence of 100 ppm of pesticides such as carbofuran, paraquat dichloride and atrazine that did not cause any inhibition to the degradation of phenol (Ahmad et al. 2015). The present work is the first known report of phenol degradation via the meta-pathway by a member of the genus Acinetobacter and describes purification and characterization of a new low molecular weight phenol hydroxylase form Acinetobacter sp. strain AQ5NOL 1.
2 Materials and methods
2.1 Microorganism and culture condition
The phenol-degrading Acinetobacter sp. strain AQ5NOL 1 used in this study was isolated from the pesticide-polluted site at Johor, Malaysia as described previously by Ahmad et al. (2011). The bacterial strain was cultured in mineral salt medium (MSM) containing (g L−1): K2HPO4, 0.4; KH2PO4, 0.2; NaCl, 0.1; MgSO4, 0.1; MnSO4·H20, 0.01; Fe2(SO4)·H2O, 0.01; NaMoO4·2H2O, 0.01; (NH4)2SO4, 0.4 at pH 7.5. The MSM was supplemented with 0.5 g L−1 phenol as the carbon source.
2.2 Preparation of crude extracts
Acinetobacter sp. strain AQ5NOL 1 culture was grown in 15 L of MSM containing 0.5 g L−1 phenol at room temperature. After 24 h of incubation time, the culture was centrifuged at 10,000g for 10 min at 4 °C. Pellet was suspended in 50 mM phosphate buffer, pH 7.5 containing 0.5 mM DTT and 0.1 mM PMSF as an anti-protease. The cell suspension was sonicated at 4 °C for a total duration of 15 min using a Branson Sonifier 450. The suspension was then ultracentrifuged for 90 min at 35,000 rpm or 105,000g at 4 °C.
2.3 Determination of phenol-degrading pathway
The strain AQ5NOL 1 was incubated for different duration of times (0, 12, 24, 36, 48, 60 and 72 h) before the crude was extracted. The assay was carried out according to the method developed by Zaki (2006). Catechol 2,3-dioxygenase (EC 1.13.11.2) and catechol 1,2-dioxygenase (EC 1.13.11.1) were detected at 375 and 260 nm using vis/UV spectrophotometer, respectively.
2.4 Purification of phenol hydroxylase
The purification procedure was carried out using 50 mM phosphate buffer, pH 7.5 at 4 °C unless stated otherwise. The clear supernatant (crude) obtained after ultracentrifugation was directly applied onto three different continuous ion exchange columns (dimensions in brackets): DEAE-Sepharose® (8 × 1.6 cm), DEAE-Sephadex® (14 × 1.6 cm) and Q-Sepharose® (7 × 1.6 cm). The Q-Sepharose® fractions with phenol hydroxylase activity were then ran through a Zorbax® Bioseries GF-250 gel filtration (2.50 × 0.94 cm) column which was connected to a HPLC system. The enzyme was eluted at the flow rate of 1 mL min−1 with 0.02 M sodium phosphate buffer (pH 7.5) containing 0.2 M NaCl. Phenol hydroxylase (EC 1.14.13.7) activity was assayed at room temperature based on the oxidation of NADH at 340 nm with an extinction coefficient for NADH of 6220 M−1 cm−1. Control experiment where phenol was omitted as a substrate showed minimal NADH oxidation activity. Protein was assayed using the Coomassie dye-binding method (Bradford 1976) using crystalline BSA as the standard. The absorbance readings were taken at 595 nm. In this study, SD2S-PAGE was used to characterize the proteins from each purification step.
2.5 Phenol hydroxylase kinetic studies
In this enzyme kinetic study, phenol was used as a substrate. The K m and V max values were analyzed using Graphpad Prism™ version 4.0 software. The temperatures and pH were varied accordingly in the optimization experiments: temperature range of 5–40 °C and pH range of 6.0–9.0 in 50 mM citrate, phosphate and tris–HCl buffers, respectively.
2.6 Temperature stability
Extracted phenol hydroxylase was incubated at −20, 4, 0 and room temperature (23 °C). Samples (100 µL) were assayed every 24 h for 40 days. Boiled enzyme extract served as control.
2.7 Statistical analysis
The data obtained were analyzed statistically using one-way ANOVA. p < 0.05 is deemed statistically significant.
3 Results and discussion
3.1 Phenol-degrading pathway determination
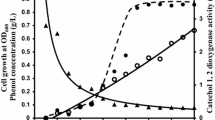
From the results obtained (Fig. 1), Acinetobacter sp. Strain AQ5NOL 1 metabolised phenol via the meta-pathways. At zero time, catechol 2,3-dioxygenase (375 nm) and catechol 1,2-dioxygenase (260 nm) activities were not detected. When Acinetobacter sp. strain AQ5NOL 1 started to degrade phenol, catechol 2,3-dioxygenase activity was detected. When phenol hydroxylase started converting phenol to catechol, catechol 2,3-dioxygenase simultaneously converted catechol to 2-hydroxymuconic semialdehyde. Then, after the phenol was completely degraded, the absorbance of catechol 2,3-dioxygenase decreased slowly. The activity of catechol 2,3-dioxygenase had decreased because some of the catechol had been converted to 2-hydroxymuconic semialdehyde. In the meta-pathway, the subsequent metabolism of catechol yields 2-hydroxymuconic semialdehyde. Further metabolisms via the meta-pathway lead to the compounds acetaldehyde, pyruvic acid, acetyl-CoA and succinate that may enter the Krebs cycle (Veenagayathri and Vasudevan 2011). Acinetobacter sp. strain AQ5NOL 1 is the first isolate from the genus Acinetobacter to metabolize phenol via the meta-pathway. Previous studies have shown that most of the Acinetobacter metabolized phenol via ortho-pathway such as Acinetobacter calcoacetius (Paller et al. 1995), Acinetobacter radioresistens (Briganti et al. 1997), Acinetobacter sp. strain W-17, RD12, DF4, and PD4 (El-Haleem et al. 2003; Wang et al. 2007; Zaki 2006; Dong et al. 2008). In some cases, both pathways can be used to degrade phenol; for example, Bacillus stearothermophilus IC3 (Adams and Ribbons 1998) and Alcaligenes eutrophus JMP 134 (Muller and Babel 1996), which can degrade phenol in both meta- and the ortho-pathways. In terms of biodegradation efficiency, phenol degradation by the meta-pathway is higher in efficiency than through the ortho-pathway (Yang and Lee 2007). The meta-pathway is strongly induced and favors high phenol concentrations that will give an advantage for the bacterial survival (Jiang et al. 2006). Anabolic and catabolic regulations via the meta-pathway produce energy for synthesis of proteins and enzyme. At higher phenol concentrations, the microbial population towards the metabolic pathway, which can cope with energy demand related to non-growth related activities, are needed to counter the inhibition of cellular activity. Some of this non-growth related energy is also used to maintain cell membrane integrity and active transport of substrates into the cell. This energy requirement is expected to be relatively high due to the higher phenol concentration (Jiang et al. 2004, 2006). In this study, it is shown that Acinetobacter sp. strain AQ5NOL 1 metabolized phenol using the meta-pathway.
3.2 Phenol hydroxylase purification
The phenol hydroxylase was found to be present in the soluble fraction of the crude cells extract of Acinetobacter sp. strain AQ5NOL 1. The elution profiles of the DEAE-Sepharose®, DEAE-Sephadex®, Q-Sepharose® and Zorbax® Bioseries GF-250 gel filtration columns are shown in Figs. 2, 3, 4, and 5, respectively. Elution using a linear gradient produced a sharp peak with high phenol hydroxylase activity at the NaCl concentrations from 0.36 to 0.51 M, 0.51 to 0.6 M, and 0.50 to 0.53 M for DEAE-Sepharose®, DEAE-Sephadex® and Q-Sepharose® (Figs. 2, 3 and 4), respectively. A single activity peak with the retention time of 10 min was obtained on the Zorbax gel filtration column (Fig. 5). The purification scheme of phenol hydroxylase is summarized in Table 1. The phenol hydroxylase was purified about 210-fold to homogeneity with a yield of 0.30 %. The results obtained show that the phenol hydroxylase from Acinetobacter sp. strain AQ5NOL 1 is very sensitive to temperature and environmental conditions, which resulted in low enzyme yields after each chromatographic step. The purification fold increased when gel filtration step was employed. The purification fold increased as the purification progressed, with the highest purification occurring after the gel filtration chromatography. Only one band was visualized on the gel filtration fraction at 50 kDa using the SDS-PAGE (Fig. 6). This suggests that the enzyme was purified to homogeneity, and indicates that the enzyme is a monomer. In a previous report, phenol hydroxylase from Acinetobacter radioresistens S13 that is purified using anion exchange De52-cellulose (2.6 x 20 cm), Q-Sepharose FF (Pharmacia) (1.3 × 26 cm) and Phenyl-Sepharose 6FF (Pharmacia) (1.3 × 13 cm) column is shown to be a dimer of 206 kDa in size.
SDS polyacrylamide gel analysis of purified phenol hydroxylase. Lane M molecular mass marker in dalton (Prestained SDS-PAGE Standards Broad Range); lane 1 crude extract of phenol hydroxylase after ultracentrifugation; 2 purified phenol hydroxylase after anion exchange chromatography (DEAE-Sepharose®); 3 purified phenol hydroxylase after anion exchange chromatography (DEAE-Sephadex®); 4 purified phenol hydroxylase after gel filtration. 5 purified phenol hydroxylase after anion exchange chromatography Q-Sepharose®
3.3 Phenol hydroxylase kinetic studies
In this study, the K m and V max values of the extracted phenol hydroxylase with phenol substrate have been determined using a Michaelis–Menten plot (Fig. 7). The correlation coefficient of 0.956 for the Michaelis–Menten plot suggested a good fit for the model. The Michaelis–Menten plot shows the K m and V max to be 13.4 µM and 2.5 µmol min−1 mg−1, respectively. A previous study showed that the K m of phenol hydroxylase from Pseudomonas fluorescens B PC18 and Pseudomonas mendcina PC1 is 140 and 17 µM, respectively (Viggor et al. 2008). Thus, our results show that phenol hydroxylase from Acinetobacter sp. strain AQ5NOL 1 has a slightly higher catalytic efficiency than P. fluorescens B PC18 and P. mendcina PC1.
The extracted phenol hydroxylase exhibited optimum activity between the temperature range of 15–25 °C with the maximum activity occurring at 20 °C (Fig. 8). Phenol hydroxylase from the local Acinetobacter sp. strain AQ5NOL 1 is a mesophilic enzyme because it has the capacity to function at moderate temperatures. In Malaysia, the annual average temperature is between 25 and 28 °C in the lowlands, while the mean minimum temperature seldom falls below 22 °C. From the result obtained, phenol hydroxylase from the local Acinetobacter sp. strain AQ5NOL 1 is suitable for bioremediation in Malaysia. Previous study showed that the optimum temperatures for phenol hydroxylase (crude extracts) from Rhodococcus sp. P1, Pseudomonas sp. CF600, Acinetobacter radioresistens S13 are 20, 22 and 24 °C, respectively (Straube 1987; Kagle and Hay 2006; Divari et al. 2003).
The effect of pH on phenol hydroxylase activity shows a typical bell-shaped profile (Fig. 9). Phenol hydroxylase from Acinetobacter sp. strain AQ5NOL 1 had an optimum pH of from 6.5 to 7.5 using phosphate buffer and from pH 7–7.5 using Tris–HCl buffer (p < 0.05). Scopes (1998) reported that pH 7.5 for enzyme activity is common for most enzymes since this is the physiological pH. Report from previous studies shows that phenol hydroxylases from Pseudomonas sp. CF600 and Acinetobacter radioresistens S13 show optimum pH of 7.4 and 7.5, respectively (Kagle and Hay 2006; Divari et al. 2003).
3.4 Temperature stability
Figure 10 shows that phenol hydroxylase from Acinetobacter sp. strain AQ5NOL 1 was stable at −20 °C for 36 days (p < 0.05). At 4 and 23 °C, phenol hydroxylase activity was reduced by 90 and 100 %, respectively, in just 1 day. The instability of enzymes at room temperature (23 °C) is caused by several factors including tertiary and quaternary protein denaturation through thermal vibrations leading to loss of cofactors, accelerated activity of bacterial and contaminating protease (from handling) at higher temperatures, and accelerated oxidation of sulfhydryl groups at higher temperatures (Scopes 1998). These denaturing reactions have standard free energies of activation such that, above a critical temperature, there is a rapid rate of loss of activity (Scopes 1998).
4 Conclusions
Acinetobacter sp. strain AQ5NOL 1 degrades phenol at high concentration and the elucidation of its phenol degradation pathway and purification of the phenol-degrading enzyme will help in further studies for understanding its degradation mechanism and future optimization of phenol degradation. This study is the first to report phenol degradation via the meta-pathway by a member of the genus Acinetobacter. The purification of phenol hydroxylase from this bacterium indicates its unique monomeric property of the enzyme. Characterization and stability studies have shed some light on the enzyme properties. Our current study includes the cloning of the phenol-degrading gene and sequencing of the enzyme.
References
Adams D, Ribbons DW (1998) The metabolism of aromatic ring fission products by baciluus stearothermophilus IC3. J Gen Microbiol 134:3179–3185
Ahmad SA, Shamaan NA, Arif NM, Shukor MYA, Syed MA (2011) Identification and characterization of a phenol degrading Acinetobacter sp. strain AQ5NOL 1. Aust J Basic Appl Sci 5:1035–1045
Ahmad SA, Shamaan NA, Arif NM, Koon GB, Shukor MYA, Syed MA (2012) Enhancement of biodegradation of phenol by immobilized cells of Acinetobacter sp. strain AQ5NOL 1. World J Microbiol Biotech 28:347–352
Ahmad SA, Shukor MY, Shamaan NA, Rahman NAA, Dahalan FA, Khalil KA, Syed MA (2015) Effects of pesticides and respiratory inhibitors on phenol degradation by Acinetobacter sp. strain AQ5NOL 1 immobilized in gellan gum. J Pure Appl Microbiol 9:489–495
Arif NM, Ahmad SA, Syed MA, Shukor MY (2013) Isolation and characterization of a phenol-degrading Rhodococcus sp. strain AQ5NOL 2 KCTC 11961BP. J Basic Microbiol 53:9–19
Basha KM, Rajendran A, Thangavelu V (2010) Recent advances in the biodegradation of phenol: a review. Asian J Exp Biol Sci 1:219–234
Bradford M (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–252
Briganti F, Pessione E, Giunta C, Scozzafava A (1997) Purification, biochemical properties and substrate specificity of a catechol 1,2-dioxygenase from phenol degrading Acinetobacter radioresistens. FEBS Lett 416:61–64
Carvalho MB, Martins I, Leitão MC, Garcia H, Rodrigues C, San Romão V, McLellan I, Hursthouse A, Silva Pereira C (2009) Screening pentachlorophenol degradation ability by environmental fungal strains belonging to the phyla Ascomycota and Zygomycota. J Ind Microbiol Biotechnol 36:1249–1256
Cassidy MB, Mullineers H, Lee H, Trevors JT (1997) Mineralization of pentachlorophenol in a contaminated soil by Pseudomonassp UG30 cells encapsulated in κ-carrageenan. J Ind Microbiol Biotechnol 19:43–48
Chakraborty S, Bhattacharya T, Patel TN, Tiwari KK (2010) Biodegradation of phenol by native microorganisms isolated from coke processing wastewater. J Environ Biol 31:293–296
Divari S, Valetti F, Caposia P, Pessione E, Cavaletto M, Griva E, Gribaudo G, Gilardi G, Giunta G (2003) The oxygenase component of phenol hydroxylase from Acinetobacter radioresistens S13. Eur J Biochem 270:2244–2253
Dong X, Hong Q, He L, Jiang X, Li S (2008) Characterization of phenol-degrading bacterial strains isolated from natural soil. Int Biodeterior Biodeg 62:257–262
El-Haleem D, Beshay U, Abdelhamid A, Moawad H, Zaki S (2003) Effect of mixed nitroghen sources on biodegradtion of phenol by immobilized Acinetobacter: sp strain W-17. Afr J Biotechnol 2:8–12
Etoumi A, El Musrati I, El Gammoudi B, El Behlil M (2008) The reduction of wax precipitation in waxy crude oils by Pseudomonas species. J Ind Microbiol Biotechnol 35:1241–1245
Farrell A, Quilty B (2002) Substrate-dependent autoaggregation of Pseudomonas putida CP1 during the degradation of mono-chlorophenols and phenol. J Ind Microbiol Biotechnol 28:316–324
González G, Herrera G, García T, Peña M (2001) Biodegradation of phenolic industrial wastewater in a fluidized bed bioreactor with immobilized cells of Pseudomonas putida. Biores Technol 80:137–142
Hannaford AM, Kuek C (1999) Aerobic batch degradation of phenol using immobilized Pseudomonas putida. J Ind Microbiol Biotechnol 22:121–126
Hao OJ, Kim MH, Seagren EA, Kim H (2002) Kinetics of phenol and chlorophenol utilization by Acinetobacter species. Chemosphere 46:797–807
Hori TSF, Avilez IM, Inoue LK, Moraes G (2006) Metabolical changes induced by chronic phenol exposure in matrinxã Brycon cephalus (Teleostei: Characidae) juveniles. Comp Biochem Physiol Part C 143:67–72
Jiang HL, Tay JH, Tay STL (2004) Changes in structure, activity and metabolism of erobic granules as a microbial response to high phenol loading. Appl Microbiol Biotechnol 63:602–608
Jiang HL, Tay STL, Maszenan AM, Tay JH (2006) Physiol traits bacterial strains isolated phenol degrading aerobic granules. FEMS Microbiol Ecol 57:182–191
Kagle J, Hay AG (2006) Phenylacetylene reversibly inhibits the phenol hydroxylase of Pseudomonas sp. CF600 at high concentrations but is oxidized at lower concentrations. Appl Microbiol Biotechnol 72:306–315
Kirchner U, Westphal AH, Muller R, Berkel WJHV (2003) Phenol hydroxylase from Bacillus thermoglucosidasius A7, a two-protein component monooxygenase with a dual roel for FAD. J Biol Chem 278:47545–47553
Muller RH, Babel W (1996) Growth rate-dependent expression of phenolassimilation pathways in Alcaligenes eutrophus JMP 134- the influence of formate as an auxiliary energy sources on phenol conversion characteristics. Appl Microbiol Biotechnol 46:156–162
Nawawi NM, Ahmad SA, Shukor MY, Syed MA, Khalil KA, Ab Rahman NA, Dahalan FA, Ibrahim AL (2016) Statistical optimization for improvement of phenol degradation by Rhodococcus sp. NAM 81. J Envion Biol 37:443–451
Norazah MN, Jayasree N, Ahmad SA, Shukor MY, Abdul Latif I (2015) Disrupting Rhodococcus sp: a competent method for genomics and proteomics. J Chem Pharm Sci 8:336–341
Paller G, Hommel RK, Kleber HP (1995) Phenol degradation by Acinetobacter calcoaceticus NCIB 8250. J Basic Microbiol 35:325–335
Saa L, Jaureguibeitia A, Largo E, Llama MJ, Serra JL (2009) Cloning, purification and characterization of two components of phenol hydroxylase from Rhodococcus erythropolis UPV-1. Appl Microbiol Biotechnol 86:201–211
Salleh AB, Ghazali FM, Rahman RNZA, Basri M (2003) Bioremediation of petroleum hydrocarbon pollution. Afr J Biotechnol 2:411–425
Scopes RK (1998) Protein purification, principles and practice. Springer-Verlag, New York
Sharma A, Thakur IS, Dureja P (2009) Enrichment, isolation and characterization of pentachlorophenol degrading bacterium Acinetobacter sp. ISTPCP-3 from effluent discharge site. Biodegradation 20:643–650
Straube G (1987) Phenol hydroxylase from Rhodococcus sp. P1. J Basic Microbiol 27:229–232
Suhaila YN, Rosfarizan M, Ahmad SA, Latif IA, Ariff AB (2013) Nutrients and culture conditions requirements for the degradation of phenol by Rhodococcus UKMP-5M. J Environ Biol 34:635–643
Tsai SC, Tsai LD, Li YK (2005) An isolated candida albicans TL3 capable of degrading phenol at large concentration. Biosci Biotechnol Biochem 69:2358–2367
Tuan NN, Hsieh HC, Lin YW, Huang SY (2011) Analysis of bacterial degradation pathways for long-chain alkylphenols involving phenol hydroxylase, alkylphenol monooxygenase and catechol dioxygenase genes. Biores Technol 102:4232–4240
van Schie PM, Young LY (2000) Biodegradation of phenol mechanisms and applications. Bioremed J 4:1–18
Veenagayathri K, Vasudevan N (2011) Ortho and mete cleavage dioxygenase detected during the degradation of phenolic compounds by a moderately halophilic bacterial consortium. Int Res J Microbiol 2:406–414
Viggor S, Heinaru E, Kuennapas A, Heinaru A (2008) Evaluation of different phenol hydroxylase-possessing phenol-degrading pseudomonas by kinetic parameters. Biodegradation 19:759–769
Wang Y, Tian Y, Han B, Zhao HB, Bi JN, Cai BL (2007) Biodegradation of phenol by free and immobilized Acinetobacter sp. strain PD12. J Environ Sci 19:222–225
Yang CF, Lee CM (2007) Enrichment, isolation, and characterization of phenol-degrading Pseudomonas resinovorans strain P-1 and Brevibacillus sp. strain P-6. Int Biodeterior Biodeg 59:206–210
Yemendzhiev H, Gerginova M, Krastanov A, Stoilova I, Alexieva Z (2008) Growth of Trametes versicolor on phenol. J Ind Microbiol Biotechnol 35:1309–1312
Yotova L, Tzibranska I, Tileva F, Markx GH, Georgieva N (2009) Kinetics of the biodegradation of phenol in wastewaters from the chemical industry by covalently immobilized Trichosporon cutaneum cells. J Ind Microbiol Biotechnol 36:367–372
Zaki S (2006) Detection of meta- and ortho-cleavage dioxygenases in bacterial phenol-degraders. J Appl Sci Environ Manag 10:75–81
Zhu C, Zhang L, Zhao L (2008) Molecular cloning, genetic organization of gene cluster encoding phenol hydroxylase and catechol 2,3-dioxygenase in Alcaligenes faecalis IS-46. World J Microbiol Biotechnol 24:1687–1695
Acknowledgments
This work was supported by the Research Grant Scheme (RUGS) 2009, Universiti Putra Malaysia (Vote No. 91851).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ahmad, S.A., Shamaan, N.A., Syed, M.A. et al. Meta-cleavage pathway of phenol degradation by Acinetobacter sp. strain AQ5NOL 1. Rend. Fis. Acc. Lincei 28, 1–9 (2017). https://doi.org/10.1007/s12210-016-0554-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12210-016-0554-2