Abstract
Purpose
Thyroid cancer and thyroid nodules are the most prevalent form of thyroid endocrine disorder. The balance of gut microbiome is highly crucial for a healthy human body, especially for the immune and endocrine system. However, the relationship between gut microbiome and the thyroid endocrine disorders such as thyroid cancer and thyroid nodules has not been reported yet.
Methods
A cohort of 74 patients was recruited for this study. Among them, 20 patients had thyroid cancer, 18 patients had thyroid nodules, and 36 were matched healthy controls. Gut microbiome composition was analyzed by 16S rRNA (16S ribosomal RNA) gene-based sequencing protocol.
Results
We compared the gut microbiome results of 74 subjects and established the correlation between gut microbiome and thyroid endocrine function for both thyroid cancer and thyroid nodules. The results inferred that alpha and beta diversity were different for patients with thyroid tumor than the healthy controls (p < 0.01). In comparison to healthy controls, the relative abundance of Neisseria (p < 0.001) and Streptococcus (p < 0.001) was significantly higher for thyroid cancer and thyroid nodules. Butyricimonas (p < 0.001) and Lactobacillus (p < 0.001) displayed notably lower relative abundance for thyroid cancer and thyroid nodules, respectively. It was also found that the clinical indexes were correlated with gut microbiome.
Conclusion
Our results indicate that both thyroid cancer and thyroid nodules are associated with the composition of gut microbiome. These results may support further clinical diagnosis to a great extent and help in developing potential probiotics to facilitate the treatment of thyroid cancer and thyroid nodules.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With technological advancement, the relationship between the composition of human microbiome such as the gut microbiome and corresponding diseases is being explored gradually and studied extensively [1]. The intestinal tract, which is the biggest digestive organ in a human body, harbors trillions of bacteria whose genomes are 100 times that of humans [2]. Human intestinal tract fosters a number of phyla including Firmicutes, Bacteroidetes, Actinobacteria and Proteobacteria [3]. Recent studies have reported that the microbiome being an immunity influencer, contributes to the immune system control or escape of distant metastasis, such as liver cancer and pancreatic cancer [4,5,6]. The variation of gut microbiome was connected with endocrinopathy, like Polycystic Ovarian Syndrome (PCOD/PCOS), obesity, type 1 and 2 diabetes [7,8,9,10], etc. Thyroid cancer is the most common endocrine malignant tumor, which is divided into 5 types [11]. Papillary thyroid cancer is another very common thyroid cancer, accounting for approximately 80–85% thyroid cancer patients in developed countries [12]. In the year 2012, about 230,000 new cases of thyroid cancer were estimated among women and men with an age-standardized (world population) rate of 6.10/100,000 for women and 1.90/100,000 for men [13]. In recent years, the patient pool suffering from thyroid disorders has seen a rapid increment worldwide, especially in women [14]. If similar trends are maintained, thyroid cancer may become the fourth most common cancer by 2030 in the United States [15]. The increasing incidence of thyroid cancer in many European countries exhibits a parallel trend as that of the United States followed by China [16]. Thyroid nodules, a form of thyroid endocrine disease, is more prevalent clinically and accounts approximately up to 4 to 7% of the population [17]. 19 to 68% of individual thyroid nodules can be detected with high-resolution ultrasound, especially in women and the elderly [18]. There are many factors that may lead to a certain thyroid disease, including ethnicity, estrogen, BMI (Body Mass Index), radioactive radiation and abnormal iodine intake [19,20,21,22,23]. It is very well-known that any thyroid disease is very closely related to thyroid hormone levels or the hormone functioning, which could influence the composition of gut microbiome [24]. At the same time, the gut microbiome could affect the integration of peripheral and central immune, metabolic and endocrine signals by gut–brain axis [25]. Functional thyroid disorders were associated with excessive bacterial growth and a different microbial composition [24]. Several studies have demonstrated that both the Graves’ disease and Hashimoto’s thyroiditis are connected with gut microbiome [26,27,28]. However, no reports of the relationship between gut microbiome, thyroid cancer and thyroid nodules have been validated.
Therefore, in this study, we analyzed the gut microbiome of thyroid cancer, thyroid nodules and healthy controls by sequencing the 16S ribosomal RNA (rRNA) gene. The results discovered effective biomarkers of these diseases while revealing significant information about the variation of the gut microbiome based on structure, composition and function. Several specific microbial biomarkers of thyroid cancer and thyroid nodules were evaluated and correlated with clinical indices.
Methods
Study subjects
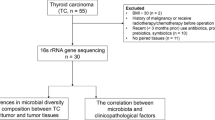
Following the approval by the Institutional Review Boards of the Affiliated Central Hospital of Qingdao University (2016-06-2101), we recruited a cohort of 74 subjects at the Affiliated Central Hospital of Qingdao University, of which 20 patients were suffering from thyroid cancer, 18 patients had thyroid nodules and there were 36 healthy controls. All the subjects signed informed consent forms, and belonged to the same geographical area. Data collected was in the format of a standardized questionnaire that included basic information, medical history and examination results (Table 1). 1) Thyroid nodule: Nodular tumors formed by benign hyperplasia of thyroid follicular epithelium; 2) Thyroid cancer: Thyroid malignant epithelial tumor characterized by follicular epithelial differentiation; 3) All the subjects had not taken antibiotics for the prior three months. All study procedures were approved by the Medical Ethics Committee of the Affiliated Central Hospital of Qingdao University.
Biospecimen collection, DNA extraction and sequencing
Fecal specimens were collected in sterilized 2-ml tubes containing pure ethanol on ice, immediately frozen (within 30 min) and stored at –80 °C until analysis. Genomic DNA was extracted using CTAB (cetyl trimethylammonium bromide) method [29]. We used Nanodrop 2000 (Thermo Scientific) spectrophotometer to determine the concentration of the extracted DNA. The V1-V2 regions of the 16S rRNA gene were amplified and sequenced on an Illumina HiSeq 2500 system. The PCR was conducted using the bacterial universal primers 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) and 338R-I (5′-GCWGCCTCCCGTAGGAGT-3′) and 338R-II (5′-GCWGCCACCCGTAGGTGT-3′) [30]. Follow-up experiments were carried out according to the sequencing manual.
16S rRNA gene sequence analysis
The 16S sequence paired-end data set was joined and quality filtered using Laser FLASH method, described by Magoč and Salzberg [31]. All sequences were analyzed using the Quantitative Insights into Microbial Ecology (QIIME, version 1.9.1) software suite [32], according to the QIIME tutorial (http://qiime.org/) with a few modified methods. Chimeric sequences, where a single organism has distinct genotypes, were removed using Metagenomics tool—usearch61 [33] with de novo models. Sequences were clustered against the 2013 Greengenes (13_5 release) ribosomal database’s 97% reference data set (http://greengenes.secondgenome.com/downloads). Sequences that remained unmatched with any of the entries in this reference were subsequently clustered into de novo OTUs at 97% similarity with UCLUST algorithm. Taxonomy was assigned to all OTUs using the RDP classifier [34] within QIIME and the Greengenes reference data set. Rarefaction and rank abundance curves were calculated from OTU tables using alpha diversity and rank abundance scripts within the QIIME pipeline. The hierarchical clustering based on population profiles of most common and abundant taxa was performed using UPGMA clustering (Unweighted Pair Group Method with Arithmetic mean, also known as average linkage) on the distance matrix of OTU abundance. This resulted in a Newick-formatted tree, which was obtained utilizing the QIIME package. Furthermore, QIIME was used to analyze Alpha diversity (Shannon, ACE), beta diversity (weighted UniFrac, Principal Coordinate Analysis (PCoA)), Linear discriminant analysis (LDA) and Effect Size (LEfSe). The Venn diagram using the taxa of the average relative abundance > 10^–5 was derived using the ‘jvenn website’ (http://jvenn.toulouse.inra.fr/app/example.html).
Statistical analysis
The clinical characteristics of the subjects are represented as the mean ± SD, which were determined using the Mann-Whitney U test. The diversity categorization of alpha and beta diversity was defined in the OTU table to a sequencing depth of 20,000 per sample. Moreover, Alpha diversity was determined using Mann-Whitney U test and beta diversity was acquired by ANOSIM- Analysis of Similarities. LEfSe combines Kruskal-Wallis test or pair wise Wilcoxon ranksum test with linear discriminant analysis (LDA), whose threshold value on the logarithmic LDA score equals to 2.0. Spearman’s rank correlation method was used to analyze the relationship between the microbiome and clinical index. Analyses were performed using the SPSS statistical package (version 17.0) and R software (version 3.4.4). p values < 0.05 were considered statistically significant.
Results
To establish the gut microbiome characteristics of patients with abnormal thyroid endocrine function, we conducted 16S rDNA gene sequencing to analyze 74 fecal samples from 74 individuals (20 patients with thyroid cancer, 18 patients with thyroid nodules and 36 healthy controls). The clinical characteristics of subjects are summarized in Table 1. No significant differences were noticed in either age or gender among the thyroid cancer, thyroid nodules and healthy control groups. Through pre-processing of the sequencing data, 4,153,725 high-quality sequences (Phred ≥ Q30) with an average of 56,131 sequences per sample were obtained.
Gut microbiome composition of thyroid cancer, thyroid nodules and healthy control
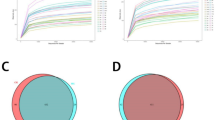
To characterize the community attributes of gut microbiome of individuals with thyroid disease, relative taxon abundances of thyroid cancer, thyroid nodules and healthy control groups were compared. At the phylum level, three prominent phyla were found to be Bacteroidetes (50 ± 12%), Firmicutes (35 ± 12%) and Proteobacteria (11 ± 6%), which represented over 90% of the total phyla in the three groups (Fig. 1a). At the genus level, genus of Prevotella demonstrated significant difference among the three groups (p = 0.0005, Fig. 1b). Venn diagrams distinguished between gut microbiome of the three groups. Within the healthy control group, 10 unique species were found, such as Proteus, Acinetobacter, Erwinia, Psychrobacter and Sneathia. Similarly, 35 unique species like Thiobacillus, Rhodobacter, Rheinheimera, Mycobacterium and Anaerotruncus were found in the thyroid cancer group; while 36 unique species, including Rubellimicrobium, Propionibacterium, Peptostreptococcus, and Parvimonas were found in the thyroid nodules group.
Comparison of OTUs and relative taxa abundance among thyroid cancer, thyroid nodules, and healthy controls. a Comparison of relative abundance among thyroid cancer, thyroid nodules and healthy controls at phylum level. b Comparison of relative abundance among thyroid cancer, thyroid nodules and healthy controls at genus level; c Venn diagram
Variations of the gut microbiome among the three groups
We analyzed the alpha and beta diversity to explore the variations in the gut microbiome among the three groups. Although no significant difference was found among the healthy control, thyroid cancer and thyroid nodules groups with respect to microbial diversity (Shannon index, p > 0.05), microbial abundance (ACE index) was higher in both thyroid cancer and nodules than in healthy control group (p < 0.05, Fig. 2). In addition, we calculated the unweighted and weighted UniFrac distance to testify the correlation of gut microbiome between the thyroid disease and healthy control group. These calculations revealed that the healthy control group was significantly different from the thyroid cancer group (r = 0.598, p= 0.001, weighted UniFrac; r = 0.972, p = 0.001, unweighted UniFrac) and thyroid nodules group (r = 0.596, p = 0.001, weighted UniFrac; r = 0.966, p = 0.001, unweighted UniFrac) based on ANOSIM method (Fig. 3). Adding the results, they indicated a significant variation of gut microbiome between thyroid disease and healthy control groups.
Bacterial taxonomic differences among the three groups
In order to select biomarkers from diseased group, we used linear discriminant effect size (LEfSe) to analyze and distinguish the composition of gut microbiome between thyroid disease and healthy control groups. The gut microbiome of the thyroid cancer group was characterized by a dominance of Prevotella, Roseburia, Coprococcus, Anaerostipes, Ruminococcus, Neisseria, Streptococcus and Porphyromonas, whereas the microbiome in healthy control group were dominated by genera of Bacteroides, Sutterella, and Butyricimonas (p < 0.05, Fig. 4). The thyroid nodules group’s microbiome were characterized by a dominance of Roseburia, Neisseria, Streptococcus, Anaerostipes and Porphyromonas, whereas the microbiome in healthy control group displayed a dominance of Bacteroides, Sutterella, Pseudomonas, Lactobacillus and Oscillospira (p < 0.05, Fig. 5).
Characteristics of microbial community composition in thyroid cancer and healthy control groups. a The most differentially abundant taxa between thyroid cancer and healthy control (LDA score above 2), which was generated from LEfSe analysis. b The enriched taxa in thyroid cancer and healthy control gut microbiome is represented in the Cladogram. The central point represents the root of the tree (Bacteria) and each ring represents the next lower taxonomic level (phylum to genus: p phylum, c class, o order, f family, g genus). The diameter of each circle represents the relative abundance of the taxon. c–f Relative abundance of taxonomy between thyroid cancer and healthy control was compared (Wilcoxon Signed Rank Test)
Characteristics of microbial community composition in thyroid cancer and healthy control groups. a The most differentially abundant taxa between thyroid nodule and healthy control (LDA score above 2), which was generated from LEfSe analysis. b The enriched taxa in thyroid cancer and healthy control gut microbiome represented in the Cladogram. The central point represents the root of the tree (Bacteria), and each ring represents the next lower taxonomic level (phylum to genus: p phylum, c class, o order, f family, g genus). The diameter of each circle represents the relative abundance of the taxon. c–e Relative abundance of taxonomy between thyroid nodule and healthy control was compared (Wilcoxon Signed Rank Test)
The gut microbiome of thyroid disease was associated with clinical indices
Spearman’s rank correlation coefficient method was used to evaluate the correlation between each subject’s gut microbiome and clinical indices parameters including FT3 (Freetriiodothyronine), FT4 (Freetetraiodothyronine), TSH (Thyroid stimulating hormone), Tg (Thyroglobulin), TgAb (Thyroglobulin antibody), TRAb (Thyrotropin receptor antibody) and TPOA (Thyroid peroxidase antibody) (Fig. 6). Strong correlations (correction r > 0.40 or r < 0.40, p < 0.05, n = 20) were found among 23 taxa and the 7 clinical indices in thyroid nodules. TG exhibited a significant negative correlation with the family Lactobacillaceae (r = −0.73, p < 0.01) and the genus Bacteroides (r = −0.62, p < 0.01). At the same time, FT3 was negatively correlated with the genera Bacteroides (r = −0.47, p < 0.05) and Lactobacillus (r = −0.47, p < 0.05). Additionally, a distinctive positive association was observed between TSH and the genera Porphyromonas (r = 0.55, p < 0.05) and Roseburia (r = 0.47, p < 0.05). Similarly, the relationship between the biomarkers (22 taxa) and clinical indices (7 indices) in thyroid cancer was established. TSH exhibited a significant positive connection with the genus Porphyromonas (r = 0.57, p < 0.01). At the same time, FT3 was positively correlated with the genera Streptococcus (r = 0.47, p < 0.05). In addition, a significant negative association can be observed between the TRAb and the families Clostridiaceae (r = −0.54, p < 0.05) and Lachnospiraceae (r = −0.53, p < 0.05). Furthermore, TPOAB was negatively correlated with the genera Ruminococcus (r = −0.53, p < 0.05).
Discussion
In this study, analysis of gut microbiome composition was performed for stool samples from subjects affected by thyroid cancer, thyroid nodules, and matched healthy controls. It was found that the gut microbiome of patients both with thyroid cancer and thyroid nodules demonstrated higher microbial richness as well as distinct composition compared to the healthy control group. It was further revealed that gut microbiome might be associated with multiple clinical indices. Our pilot study reported that the composition of gut microbiome was correlated with thyroid cancer and thyroid nodules.
Several studies have demonstrated that the dysbiosis of human microbiome could be primarily caused by inflammatory disorders and diverse cancer types throughout the body [4,5,6, 35, 36]. In our study, we observed that the microbial richness was dominantly higher in both the thyroid cancer group and thyroid nodules group than in the healthy control group. A number of studies suggested that the microbial diversity was significantly higher in cancer cases [37, 38]. The assembled results indicate that the opportunistic pathogen may colonize in patients suffering from thyroid endocrine disease.
With respect to gut microbiome composition, the relative abundance of Clostridiaceae (p = 0.045), Neisseria (p < 0.001) and Streptococcus (p < 0.001) was significantly higher in the thyroid cancer group, while the abundance of Neisseria (p < 0.001) and Streptococcus (p < 0.001) was notably increased in the thyroid nodules group. Clostridium subterminale from gut microbiome was detected in the blood of a metastatic gastrointestinal adenocarcinoma patient, which indicated that a few species of Clostridiaceae have carcinogenic effects [39]. Neisseria levels were found to be elevated in the setting of esophageal inflammation, which was correlated to inflammatory disorders [40,41,42]. Meanwhile, it was also associated with pancreatic disease [43]. Several studies also demonstrated that the increment of both adenomas and carcinomas was associated with the presence of Streptococcus [44,45,46]. Our results showed an increment in these three genera of bacteria, which indicates that these bacteria could be a factor that affects thyroid disease or thyroid functioning.
Overall, the relative abundance of Butyricimonas (p < 0.001) and Lactobacillus (p < 0.001) was significantly lower in the thyroid cancer group and thyroid nodules group, respectively. As we know, Butyricimonas is strictly anaerobic gram-negative bacteria, which could produce SCFAs (Short Chain Fatty Acids) to maintain the health of intestinal tract [47, 48]. Butyrate (BT) is one of the SCFAs, which could regulate innate and adaptive immune cell function. For instance, BT develops an anti-inflammatory effect by inhibiting the recruitment and pro-inflammatory activity of immune cell, such as neutrophils. And it could activate the differentiation of colonic Treg cells to suppress inflammation [49]. Lactobacillus is an important genus in the human gut which could improve the concentration of various trace elements in human cells, such as selenium [50]. Selenium is crucial for the transformation of thyroid hormone activation and protects the thyroid gland from oxidative damage during hormone synthesis [51].
The gut microbiome plays a critical role in substance metabolism and influence the essential diagnosis and treatment for various pivotal diseases such as cancer, diabetes and melanoma [52,53,54]. In this study, our results indicated that gut microbiome was associated with the disease index. TSH was found to have a significant positive association with the Porphyromonas (p < 0.01) and FT3 was positively connected with the Streptococcus in thyroid cancer cases. It is known that TSH and FT3 levels are higher in thyroid cancer cases [55]. In the previous description, Porphyromonas and Streptococcus were conditional pathogens [56]. Meanwhile, we also found TSH was positively associated with the Porphyromonas (p < 0.01) in thyroid nodules, which indicated gut microbiome possessed some of the same mechanisms for affecting thyroid cancer and thyroid nodules. In addition, FT3 was negatively connected with Lactobacillus and its relative abundance was slightly lower than that of the healthy controls. Lactobacillus could reduce inflammation by producing SCFAs [47, 48]. In summary, our results indicated Porphyromonas, Streptococcus and Lactobacillus could serve as essential biomarkers having potentials in detecting thyroid cancer or thyroid nodules.
Conclusion
This study indicates that the variation of gut microbiome is associated with thyroid cancer and thyroid nodules. It also suggests the potential pathogens active in thyroid cancer and thyroid nodules. These results may be essential for clinical diagnosis and could be further used to develop potential probiotics that could facilitate the treatment of thyroid cancer and thyroid nodules. However, it still remains unknown if thyroid disorders cause dysbiosis or vice-versa. Further studies need to scrutinize and explore the mechanisms between the gut microbiome and thyroid cancer or nodules using modeled organisms.
Abbreviations
- Tg:
-
Thyroglobulin
- TgAb:
-
Thyroglobulin antibody
- TSH:
-
Thyroid stimulating hormone
- FT3:
-
Free triiodothyronine
- FT4:
-
Free tetraiodothyronine
- TRAb:
-
Thyrotropin receptor antibody
- TPOAb:
-
Thyroid peroxidase antibody
References
J.A. Gilbert, M.J. Blaser, J.G. Caporaso, J.K. Jansson, S.V. Lynch, R. Knight, Current understanding of the human microbiome. Nat. Med. 24(4), 392–400 (2018). https://doi.org/10.1038/nm.4517
C. De Filippo, D. Cavalieri, M. Di Paola, M. Ramazzotti, J.B. Poullet, S. Massart, S. Collini, G. Pieraccini, P. Lionetti. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl Acad. Sci. USA. 107(33), 14691-14696 (2010). https://doi.org/10.1073/pnas.1005963107
L. Dethlefsen, M. McFall-Ngai, D.A. Relman, An ecological and evolutionary perspective on human-microbe mutualism and disease. Nature 449(7164), 811–818 (2007). https://doi.org/10.1038/nature06245
O. Huck, J. Al-Hashemi, L. Poidevin, O. Poch, J.L. Davideau, H. Tenenbaum, S. Amar. identification and characterization of microrna differentially expressed in macrophages exposed to porphyromonas gingivalis infection. Infect. Immunity 85(3), (2017). https://doi.org/10.1128/IAI.00771-16
C. Pere-Vedrenne, M. Prochazkova-Carlotti, B. Rousseau, W. He, L. Chambonnier, E. Sifre, A. Buissonniere, P. Dubus, F. Megraud, C. Varon, A. Menard, The cytolethal distending toxin subunit cdtb of helicobacter hepaticus promotes senescence and endoreplication in xenograft mouse models of hepatic and intestinal cell lines. Front. Cell. Infect. Microbiol. 7, 268 (2017). https://doi.org/10.3389/fcimb.2017.00268
L. Zitvogel, M. Ayyoub, B. Routy, G. Kroemer, Microbiome and anticancer immunosurveillance. Cell 165(2), 276–287 (2016). https://doi.org/10.1016/j.cell.2016.03.001
Y. Guo, Y. Qi, X. Yang, L. Zhao, S. Wen, Y. Liu, L. Tang, Association between Polycystic Ovary Syndrome and Gut Microbiota. PloS One 11(4), e0153196 (2016). https://doi.org/10.1371/journal.pone.0153196
R.J. Perry, L. Peng, N.A. Barry, G.W. Cline, D. Zhang, R.L. Cardone, K.F. Petersen, R.G. Kibbey, A.L. Goodman, G.I. Shulman, Acetate mediates a microbiome-brain-beta-cell axis to promote metabolic syndrome. Nature 534(7606), 213–217 (2016). https://doi.org/10.1038/nature18309
T. Vatanen, E.A. Franzosa, R. Schwager, S. Tripathi, T.D. Arthur, K. Vehik, A. Lernmark, W.A. Hagopian, M.J. Rewers, J.X. She, J. Toppari, A.G. Ziegler, B. Akolkar, J.P. Krischer, C.J. Stewart, N.J. Ajami, J.F. Petrosino, D. Gevers, H. Lahdesmaki, H. Vlamakis, C. Huttenhower, R.J. Xavier, The human gut microbiome in early-onset type 1 diabetes from the TEDDY study. Nature 562(7728), 589–594 (2018). https://doi.org/10.1038/s41586-018-0620-2
K. Forslund, F. Hildebrand, T. Nielsen, G. Falony, E. Le Chatelier, S. Sunagawa, E. Prifti, S. Vieira-Silva, V. Gudmundsdottir, H.K. Pedersen, M. Arumugam, K. Kristiansen, A.Y. Voigt, H. Vestergaard, R. Hercog, P.I. Costea, J.R. Kultima, J. Li, T. Jorgensen, F. Levenez, J. Dore, H.I.Tc Meta, H.B. Nielsen, S. Brunak, J. Raes, T. Hansen, J. Wang, S.D. Ehrlich, P. Bork, O. Pedersen, Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528(7581), 262–266 (2015). https://doi.org/10.1038/nature15766
C.F.L. Goncalves, M.L. de Freitas, A.C.F. Ferreira. Flavonoids, thyroid iodide uptake and thyroid cancer-a review. Int. J. Mol. Sci. 18(6), (2017). https://doi.org/10.3390/ijms18061247
T. Carling, R. Udelsman, Thyroid cancer. Annu. Rev. Med. 65, 125–137 (2014). https://doi.org/10.1146/annurev-med-061512-105739
J. Ferlay, I. Soerjomataram, R. Dikshit, S. Eser, C. Mathers, M. Rebelo, D.M. Parkin, D. Forman, F. Bray, Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136(5), E359–386 (2015). https://doi.org/10.1002/ijc.29210
G. Pellegriti, F. Frasca, C. Regalbuto, S. Squatrito, R. Vigneri, Worldwide increasing incidence of thyroid cancer: update on epidemiology and risk factors. J. Cancer Epidemiol. 2013, 965212 (2013). https://doi.org/10.1155/2013/965212
L. Rahib, B.D. Smith, R. Aizenberg, A.B. Rosenzweig, J.M. Fleshman, L.M. Matrisian, Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 74(11), 2913–2921 (2014). https://doi.org/10.1158/0008-5472.CAN-14-0155
D.M. Parkin, J. Ferlay, M.P. Curado, F. Bray, B. Edwards, H.R. Shin, D. Forman, Fifty years of cancer incidence: CI5 I-IX. Int. J. Cancer 127(12), 2918–2927 (2010). https://doi.org/10.1002/ijc.25517
K.D. Burman, L. Wartofsky, Clinical practice. thyroid nodules. New Engl. J. Med. 373(24), 2347–2356 (2015). https://doi.org/10.1056/NEJMcp1415786
S. Guth, U. Theune, J. Aberle, A. Galach, C.M. Bamberger, Very high prevalence of thyroid nodules detected by high frequency (13 MHz) ultrasound examination. Eur. J. Clin. Investig. 39(8), 699–706 (2009). https://doi.org/10.1111/j.1365-2362.2009.02162.x
F. Clavel-Chapelon, G. Guillas, L. Tondeur, C. Kernaleguen, M.C. Boutron-Ruault, Risk of differentiated thyroid cancer in relation to adult weight, height and body shape over life: the French E3N cohort. Int. J. Cancer 126(12), 2984–2990 (2010). https://doi.org/10.1002/ijc.25066
A. Magreni, D.V. Bann, J.R. Schubart, D. Goldenberg, The effects of race and ethnicity on thyroid cancer incidence. JAMA Otolaryngol. Head. Neck Surg. 141(4), 319–323 (2015). https://doi.org/10.1001/jamaoto.2014.3740
M. Sokouti, V. Montazeri, A. Fakhrjou, S. Samankan, M. Goldust, Thyroid cancer, clinical and hystopathological study on patients under 25 years in Tabriz, Iran (2000-2012). Pak. J. Biol. Sci.: PJBS 16(24), 2003–2008 (2013)
L.H. Veiga, E. Holmberg, H. Anderson, L. Pottern, S. Sadetzki, M.J. Adams, R. Sakata, A.B. Schneider, P. Inskip, P. Bhatti, R. Johansson, G. Neta, R. Shore, F. de Vathaire, L. Damber, R. Kleinerman, M.M. Hawkins, M. Tucker, M. Lundell, J.H. Lubin, Thyroid cancer after childhood exposure to external radiation: an updated pooled analysis of 12 studies. Radiat. Res. 185(5), 473–484 (2016). https://doi.org/10.1667/RR14213.1
M. Zane, C. Parello, G. Pennelli, D.M. Townsend, S. Merigliano, M. Boscaro, A. Toniato, G. Baggio, M.R. Pelizzo, D. Rubello, I.M. Boschin, Estrogen and thyroid cancer is a stem affair: a preliminary study. Biomed. Pharmacother. 85, 399–411 (2017). https://doi.org/10.1016/j.biopha.2016.11.043
C. Virili, M. Centanni, “With a little help from my friends” - The role of microbiota in thyroid hormone metabolism and enterohepatic recycling. Mol. Cell. Endocrinol. 458, 39–43 (2017). https://doi.org/10.1016/j.mce.2017.01.053
E. Jasarevic, K.E. Morrison, T.L. Bale, Sex differences in the gut microbiome-brain axis across the lifespan. Philosophical transactions of the Royal Society of London. Ser. B, Biol. Sci. 371(1688), 20150122 (2016). https://doi.org/10.1098/rstb.2015.0122
D. Covelli, M. Ludgate, The thyroid, the eyes and the gut: a possible connection. J. Endocrinol. Investig. 40(6), 567–576 (2017). https://doi.org/10.1007/s40618-016-0594-6
H.M. Ishaq, I.S. Mohammad, H. Guo, M. Shahzad, Y.J. Hou, C. Ma, Z. Naseem, X. Wu, P. Shi, J. Xu, Molecular estimation of alteration in intestinal microbial composition in Hashimoto’s thyroiditis patients. Biomed. Pharmacother. 95, 865–874 (2017). https://doi.org/10.1016/j.biopha.2017.08.101
F. Zhao, J. Feng, J. Li, L. Zhao, Y. Liu, H. Chen, Y. Jin, B. Zhu, Y. Wei, Alterations of the gut microbiota in hashimoto’s thyroiditis patients. Thyroid 28(2), 175–186 (2018). https://doi.org/10.1089/thy.2017.0395
X. Wang, L. Zhang, Y. Wang, X. Liu, H. Zhang, Y. Liu, N. Shen, J. Yang, Z. Gai, Gut microbiota dysbiosis is associated with Henoch-Schonlein Purpura in children. Int. Immunopharmacol. 58, 1–8 (2018). https://doi.org/10.1016/j.intimp.2018.03.003
K. Findley, J. Oh, J. Yang, S. Conlan, C. Deming, J.A. Meyer, D. Schoenfeld, E. Nomicos, M. Park; Program, N.I.H.I.S.C.C.S., H.H. Kong, J.A. Segre, Topographic diversity of fungal and bacterial communities in human skin. Nature 498(7454), 367–370 (2013). https://doi.org/10.1038/nature12171
T. Magoc, S.L. Salzberg, FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21), 2957–2963 (2011). https://doi.org/10.1093/bioinformatics/btr507
J.G. Caporaso, J. Kuczynski, J. Stombaugh, K. Bittinger, F.D. Bushman, E.K. Costello, N. Fierer, A.G. Pena, J.K. Goodrich, J.I. Gordon, G.A. Huttley, S.T. Kelley, D. Knights, J.E. Koenig, R.E. Ley, C.A. Lozupone, D. McDonald, B.D. Muegge, M. Pirrung, J. Reeder, J.R. Sevinsky, P.J. Turnbaugh, W.A. Walters, J. Widmann, T. Yatsunenko, J. Zaneveld, R. Knight, QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7(5), 335–336 (2010). https://doi.org/10.1038/nmeth.f.303
R.C. Edgar, Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19), 2460 (2010)
J.R. Cole, Q. Wang, E. Cardenas, J. Fish, B. Chai, R.J. Farris, A.S. Kulam-Syed-Mohideen, D.M. McGarrell, T. Marsh, G.M. Garrity, J.M. Tiedje, The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 37(Database issue), D141–145 (2009). https://doi.org/10.1093/nar/gkn879
S.V. Rajagopala, S. Vashee, L.M. Oldfield, Y. Suzuki, J.C. Venter, A. Telenti, K.E. Nelson, The human microbiome and cancer. Cancer Prev. Res. 10(4), 226–234 (2017). https://doi.org/10.1158/1940-6207.CAPR-16-0249
S. Meng, B. Chen, J. Yang, J. Wang, D. Zhu, Q. Meng, L. Zhang, Study of microbiomes in aseptically collected samples of human breast tissue using needle biopsy and the potential role of in situ tissue microbiomes for promoting malignancy. Front Oncol. 8, 318 (2018). https://doi.org/10.3389/fonc.2018.00318
X. Wang, T. Ye, W.J. Chen, Y. Lv, Z. Hao, J. Chen, J.Y. Zhao, H.P. Wang, Y.K. Cai, Structural shift of gut microbiota during chemo-preventive effects of epigallocatechin gallate on colorectal carcinogenesis in mice. World J. Gastroenterol. 23(46), 8128–8139 (2017). https://doi.org/10.3748/wjg.v23.i46.8128
J. Zhu, M. Liao, Z. Yao, W. Liang, Q. Li, J. Liu, H. Yang, Y. Ji, W. Wei, A. Tan, S. Liang, Y. Chen, H. Lin, X. Zhu, S. Huang, J. Tian, R. Tang, Q. Wang, Z. Mo, Breast cancer in postmenopausal women is associated with an altered gut metagenome. Microbiome 6(1), 136 (2018). https://doi.org/10.1186/s40168-018-0515-3
K.M. Trapani, L.J. Boghossian, E. Caskey, Clostridium subterminale Septicemia in a patient with metastatic gastrointestinal adenocarcinoma. Case Rep. Infect. Dis. 2018, 6031510 (2018). https://doi.org/10.1155/2018/6031510
A.J. Benitez, C. Hoffmann, A.B. Muir, K.K. Dods, J.M. Spergel, F.D. Bushman, M.L. Wang, Inflammation-associated microbiota in pediatric eosinophilic esophagitis. Microbiome 3, 23 (2015). https://doi.org/10.1186/s40168-015-0085-6
S. Macfarlane, E. Furrie, G.T. Macfarlane, J.F. Dillon, Microbial colonization of the upper gastrointestinal tract in patients with Barrett’s esophagus. Clin. Infect. Dis. 45(1), 29–38 (2007). https://doi.org/10.1086/518578
J. Si, C. Lee, G. Ko, Oral microbiota: microbial biomarkers of metabolic syndrome independent of host genetic factors. Front Cell Infect. Microbiol 7, 516 (2017). https://doi.org/10.3389/fcimb.2017.00516
R. Memba, S.N. Duggan, H.M. Ni Chonchubhair, O.M. Griffin, Y. Bashir, D.B. O’Connor, A. Murphy, J. McMahon, Y. Volcov, B.M. Ryan, K.C. Conlon, The potential role of gut microbiota in pancreatic disease: a systematic review. Pancreatology 17(6), 867–874 (2017). https://doi.org/10.1016/j.pan.2017.09.002
J.S. Gold, S. Bayar, R.R. Salem, Association of Streptococcus bovis bacteremia with colonic neoplasia and extracolonic malignancy. Arch. Surg. 139(7), 760–765 (2004). https://doi.org/10.1001/archsurg.139.7.760
A.S. Abdulamir, R.R. Hafidh, F. Abu Bakar, The association of Streptococcus bovis/gallolyticus with colorectal tumors: the nature and the underlying mechanisms of its etiological role. J. Exp. Clin. Cancer Res 30, 11 (2011). https://doi.org/10.1186/1756-9966-30-11
J.D. Dahmus, D.L. Kotler, D.M. Kastenberg, C.A. Kistler, The gut microbiome and colorectal cancer: a review of bacterial pathogenesis. J. Gastrointest. Oncol. 9(4), 769–777 (2018). https://doi.org/10.21037/jgo.2018.04.07
M. Sakamoto, A. Takagaki, K. Matsumoto, Y. Kato, K. Goto, Y. Benno, Butyricimonas synergistica gen. nov., sp. nov. and Butyricimonas virosa sp. nov., butyric acid-producing bacteria in the family ‘Porphyromonadaceae’ isolated from rat faeces. Int. J. Syst. Evolut. Microbiol. 59(Pt 7), 1748–1753 (2009). https://doi.org/10.1099/ijs.0.007674-0
J. Zhao, L. Nian, L.Y. Kwok, T. Sun, J. Zhao, Reduction in fecal microbiota diversity and short-chain fatty acid producers in Methicillin-resistant Staphylococcus aureus infected individuals as revealed by PacBio single molecule, real-time sequencing technology. Eur. J. Clin. Microbiol. Infect. Dis. 36(8), 1463–1472 (2017). https://doi.org/10.1007/s10096-017-2955-2
Y. Furusawa, Y. Obata, S. Fukuda, T.A. Endo, G. Nakato, D. Takahashi, Y. Nakanishi, C. Uetake, K. Kato, T. Kato, M. Takahashi, N.N. Fukuda, S. Murakami, E. Miyauchi, S. Hino, K. Atarashi, S. Onawa, Y. Fujimura, T. Lockett, J.M. Clarke, D.L. Topping, M. Tomita, S. Hori, O. Ohara, T. Morita, H. Koseki, J. Kikuchi, K. Honda, K. Hase, H. Ohno, Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504(7480), 446–450 (2013). https://doi.org/10.1038/nature12721
E. Pessione, Lactic acid bacteria contribution to gut microbiota complexity: lights and shadows. Front. Cell. Infect. Microbiol. 2, 86 (2012). https://doi.org/10.3389/fcimb.2012.00086
S. Danzi, I. Klein, Thyroid hormone and the cardiovascular system. Med. Clin. North Am. 96(2), 257–268 (2012). https://doi.org/10.1016/j.mcna.2012.01.006
A.P. Garcia-Gonzalez, A.D. Ritter, S. Shrestha, E.C. Andersen, L.S. Yilmaz, A.J.M. Walhout, Bacterial metabolism affects the C. elegans response to cancer chemotherapeutics. Cell 169(3), 431–441 e438 (2017). https://doi.org/10.1016/j.cell.2017.03.046
V. Gopalakrishnan, C.N. Spencer, L. Nezi, A. Reuben, M.C. Andrews, T.V. Karpinets, P.A. Prieto, D. Vicente, K. Hoffman, S.C. Wei, A.P. Cogdill, L. Zhao, C.W. Hudgens, D.S. Hutchinson, T. Manzo, M. Petaccia de Macedo, T. Cotechini, T. Kumar, W.S. Chen, S.M. Reddy, R. Szczepaniak Sloane, J. Galloway-Pena, H. Jiang, P.L. Chen, E.J. Shpall, K. Rezvani, A.M. Alousi, R.F. Chemaly, S. Shelburne, L.M. Vence, P.C. Okhuysen, V.B. Jensen, A.G. Swennes, F. McAllister, E. Marcelo Riquelme Sanchez, Y. Zhang, E. Le Chatelier, L. Zitvogel, N. Pons, J.L. Austin-Breneman, L.E. Haydu, E.M. Burton, J.M. Gardner, E. Sirmans, J. Hu, A.J. Lazar, T. Tsujikawa, A. Diab, H. Tawbi, I.C. Glitza, W.J. Hwu, S.P. Patel, S.E. Woodman, R.N. Amaria, M.A. Davies, J.E. Gershenwald, P. Hwu, J.E. Lee, J. Zhang, L.M. Coussens, Z.A. Cooper, P.A. Futreal, C.R. Daniel, N.J. Ajami, J.F. Petrosino, M.T. Tetzlaff, P. Sharma, J.P. Allison, R.R. Jenq, J.A. Wargo, Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359(6371), 97–103 (2018). https://doi.org/10.1126/science.aan4236
Y. Gu, X. Wang, J. Li, Y. Zhang, H. Zhong, R. Liu, D. Zhang, Q. Feng, X. Xie, J. Hong, H. Ren, W. Liu, J. Ma, Q. Su, H. Zhang, J. Yang, X. Wang, X. Zhao, W. Gu, Y. Bi, Y. Peng, X. Xu, H. Xia, F. Li, X. Xu, H. Yang, G. Xu, L. Madsen, K. Kristiansen, G. Ning, W. Wang, Analyses of gut microbiota and plasma bile acids enable stratification of patients for antidiabetic treatment. Nat. Commun. 8(1), 1785 (2017). https://doi.org/10.1038/s41467-017-01682-2
H. Zhao, H. Li, T. Huang, High urinary iodine, thyroid autoantibodies, and thyroid-stimulating hormone for papillary thyroid cancer risk. Biol. Trace Elem. Res. 184(2), 317–324 (2018). https://doi.org/10.1007/s12011-017-1209-6
R. Guerrero-Preston, F. Godoy-Vitorino, A. Jedlicka, A. Rodriguez-Hilario, H. Gonzalez, J. Bondy, F. Lawson, O. Folawiyo, C. Michailidi, A. Dziedzic, R. Thangavel, T. Hadar, M.G. Noordhuis, W. Westra, W. Koch, D. Sidransky, 16S rRNA amplicon sequencing identifies microbiota associated with oral cancer, human papilloma virus infection and surgical treatment. Oncotarget 7(32), 51320–51334 (2016). https://doi.org/10.18632/oncotarget.9710
Funding
This study was supported by the National Natural Science Foundation of China under contract No.31471202 (Lei Zhang) and No. 81670822 (Ye Wang); the Shandong Provincial Key Research and Development Program under contract No. 2016YYSP009 (Lei Zhang); Weihai Technique Extension Project under contract No. 2016GNS023 (Lei Zhang). Qingdao Key Research Project No. 17-3-3-10-nsh (Chunling Zhang). Lei Zhang was also supported by the Taishan Scholars Program of Shandong Province (No. tshw20120206).
Author Contributions
L.Z., J.Y. and M.Y. designed the study. J.Z., C.L., Y.Y., H.W., Y.S., Y.W., X.M., D.Z., and C.Z. performed measurements and data analysis. C.Z. and Q.X. obtained samples. F.Z., J.Z. and L.Z. wrote the manuscript. All authors have read and critically revised the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. The authors have nothing to disclose.
Ethical approval
All procedures performed in the above studies involving human participants were in accordance with the ethical standards of the institution and/or national research committee and with the 1964 Declaration of Helsinki along with its later amendments or comparable ethical standards.
Informed consent
All the subjects signed informed consent beforehand and belonged to the same geographical area. Data was collected by using a standardized questionnaire including basic information, medical history and examination results.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, J., Zhang, F., Zhao, C. et al. Dysbiosis of the gut microbiome is associated with thyroid cancer and thyroid nodules and correlated with clinical index of thyroid function. Endocrine 64, 564–574 (2019). https://doi.org/10.1007/s12020-018-1831-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12020-018-1831-x