Abstract
The reductase (PgCR) from recombinant Escherichia coli CCZU-Y10 displayed high reductase activity and excellent stereoselectivity for the reduction of ethyl 4-chloro-3-oxobutanoate (COBE) into ethyl (S)-4-chloro-3-hydroxybutanoate ((S)-CHBE). To efficiently synthesize (S)-CHBE (>99 % enantiomeric excess (ee)), the highly stereoselective bioreduction of COBE into (S)-CHBE with the whole cells of E. coli CCZU-Y10 was successfully demonstrated in a dibutyl phthalate-water biphasic system. The appropriate ratio of the organic phase to water phase was 1:1 (v/v). The optimum reaction temperature, reaction pH, cosubstrate, NAD+, and cell dosage of the biotransformation of 100 mM COBE in this biphasic system were 30 °C, 7.0, mannitol (2.5 mmol/mmol COBE), 0.1 μmol/(mmol COBE), and 0.1 g (wet weight)/mL, respectively. Moreover, COBE at a high concentration of (1,000 mM) could be asymmetrically reduced to (S)-CHBE in a high yield (99.0 %) and high enantiometric excess value (>99 % ee). Significantly, E. coli CCZU-Y10 shows high potential in the industrial production of (S)-CHBE (>99 % ee).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
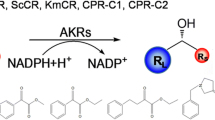
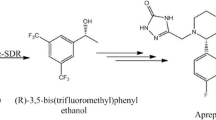
Optically active alcohols are one kind of key chiral building blocks for synthesizing many bioactive agricultural and pharmaceutical products [1, 2]. Ethyl (S)-4-chloro-3-hydroxybutanoate [(S)-CHBE] is an important precursor for the production of chiral drugs, including cholesterol-lowering HMG-CoA reductase inhibitors such as Lipitor [3, 4]. Lipitor is one of best selling prescription medicines worldwide with annual sales of over $10 billion [5]. Therefore, effective methods that can be economically used for synthesizing highly optically active (S)-CHBE (>99 % enantiomeric excess (ee)) are of great interest.
Asymmetric bioreduction of prochiral ketone, ethyl 4-chloro-3-oxobutanoate (COBE), is a more practical way to synthesize highly optically active (S)-CHBE [6]. However, carbonyl reduction enzymes for asymmetrically reducing prochiral ketone into chiral alcohol often require cofactor NADH or NADPH as an electron donor [7, 8]. To avoid using these expensive cofactors, it is necessary to establish a highly efficient and cost-effective cofactor regeneration system (e.g., enzyme-coupled system or substrate-coupled system) [8–14]. Formate dehydrogenase or glucose dehydrogenase could be used as an enzyme-coupled system for recycling NAD+ or NADP+ [9–13]; glucose could be chosen as an appropriate cosubstrate for improving the yield of (S)-CHBE [8, 14]. Therefore, searching for new carbonyl reductases and improving their application performances are of great interest.
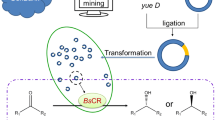
In post-genomic era, genetic data resources in various databases are quite rich. Besides, over 99 % of microbes in the environment are unculturable [15, 16]. Genome data mining is thus becoming a burgeoning area and provides an unprecedented opportunity for searching more novel enzymes due to the abundance of already existing but unexplored gene resources. Recently, it has been employed for the screening of L-aminoacylase, cytochrome P450 monooxygenase, endoglucanases, laccase, nitrilase, reductase, xylanase, etc. [8, 17, 18]. Compared with the laborious traditional screening from soil samples, genome data mining is a more effective approach which involves searching gene databases for sequences similar to those of known enzymes [7].
This study is the first to report that an NADH-dependent reductase (PgCR) from Pichia guilliermondii CGMCC 2.1801 was discovered by genome data mining. After PgCR was overexpressed in Escherichia coli BL21, a high activity of reductase-producing strain, recombinant E. coli CCZU-Y10, was employed for the efficient synthesis of ethyl (S)-CHBE (>99 % ee) from the reduction of COBE. It is worthy that no overexpression operation of coenzyme-recycling enzyme was carried out. Using mannitol as a cosubstrate, NADH or/and NADPH need not be extra added into reaction media. To increase the yield of (S)-CHBE, various parameters (organic solvent, volumetric phase ratio, reaction temperature, reaction pH, cell dosage, and substrate loading) on the asymmetric reduction were investigated in the aqueous-organic system using mannitol as a cosubstrate. Subsequently, asymmetric reduction of 1,000 mM COBE by recombinant E. coli CCZU-Y10 in the organic-aqueous biphasic system was successfully demonstrated.
Materials and Methods
Chemicals
COBE (95 % purity), (S)-CHBE (96 % purity), and (R)-CHBE (95 % purity) were obtained from Aladdin Chemistry Co., Ltd. (Shanghai, China). All other chemicals were also from local commercial sources and of analytical grade.
Microorganism
Genome data mining is an effective approach which involves searching gene databases for sequences similar to those of known enzymes [8]. After a BLAST search on the webnet (http://www.ncbi.nlm.nih.gov), the reductase from P. guilliermondii ATCC 6260 (GenBank accession no. XM_001482396) has a high identity (64 %) with the carbonyl reductase S1 from Candida magnoliae (GenBank accession no. AB036927.1) [19]. In this study, genomic DNA was extracted from P. guilliermondii CGMCC 2.1801 which was purchased from the China General Microbiological Culture Collection Center, using a TIANamp Bacteria DNA Kit from Tiangen (Shanghai). Oligonucleotide primers (PgCR-F: 5′-CATGCTAGCATGAAGTCTATGATCAATGAAAAC-3′ and PgCR-R: 5′-CCGCTCGAGTTATGGCGCACAGTAGCC-3′) with NdeI and XhoI restriction sites were designed according to the PgCR gene sequence (GenBank accession no. XM_001482396). The DNA fragment of PgCR gene (831 bp) was amplified (Fig. 1a, lane 2) and double-digested with NdeI and XhoI and then inserted into the expression vector pET-28a (Novagen, Shanghai). The resulting plasmid, pET-28a-PgCR, was transformed into E. coli BL21 (DE3) cells. This strain named recombinant E. coli CCZU-Y10 was preserved in our laboratory. The expression of PgCR was about 33.5 kDa on SDS–PAGE (Fig. 1b, lane 2).
Growth Conditions
Recombinant E. coli CCZU-Y10 was cultivated at 37 °C in LB medium containing 50 μg/mL of kanamycin. When the optical density at 600 nm (OD600) of the culture reached 0.60, IPTG as an inducer was added to a final concentration of 0.50 mM, and cultivation was continued at 37 °C for an additional 5 h. The recombinant E. coli CCZU-Y10 cells were harvested by centrifugation (8,000 × g, 10 min) at 4 °C and washed twice with potassium phosphate buffer (100 mM, pH 7.0).
Optimization of Bioconversion Process
To efficiently synthesize (S)-CHBE, various parameters (e.g., cosubstrate, organic solvent, reaction temperature, reaction pH, cell dosage, and substrate concentration) on the effects of bioreduction reaction were investigated. The effects of cosubstrate on the reductase activity were carried out by adding 0.16 g E. coli CCZU-Y10 wet cells into 2 mL potassium phosphate buffer (100 mM, pH 7.0) containing 0.2 mmol COBE, 0.2 mmol potential cosubstrate (mannose, fructose, glucose, sucrose, glycerol, mannitol, ethanol, or isopropanol), and 0.02 μmol NAD+. To investigate the effect of cosubstrate concentration, bioconversion was carried out with 0.2 mmol COBE, 0.02 μmol NAD+, and certain concentration of cosubstrate (0.05–2.5 mmol). To investigate the effects of organic solvent on the asymmetric reduction of COBE to (S)-CHBE, bioconversion was performed at 30 °C and 160 rpm by adding 0.16 g E. coli CCZU-Y10 wet cells into the biphasic system containing 1.6 mL potassium phosphate buffer (100 mM, pH 7.0), 0.4 mL test organic solvent (dibutyl phthalate, ethyl acetate, n-hexane, n-heptane, n-octane, or toluene), 0.2 mmol COBE, 0.5 mmol mannitol, and 0.02 μmol NAD+. To investigate the effect of phase ratio on the reduction, biotransformation of 0.2 mmol COBE was performed by adding 0.16 g E. coli CCZU-Y10 wet cells, 0.5 mmol mannitol, and 0.02 μmol NAD+ into a 2-mL biphasic system containing a certain volume (0.2–1.8 mL) of potassium phosphate buffer (100 mM, pH 7.0) and a certain volume (0.2–1.8 mL) of dibutyl phthalate. To investigate the effect of reaction temperature and pH on the reduction, bioconversions were performed at various temperature (25–50 °C) and various pH (citrate buffer solution, pH 3–5; phosphate buffer solution, pH 6–8; carbonate buffer solution, pH 9–10) by adding 0.16 g E. coli CCZU-Y10 wet cells, 0.2 mmol COBE, 0.5 mmol mannitol, and 0.02 μmol NAD+ into the biphasic media containing 1 mL potassium phosphate buffer and 1 mL dibutyl phthalate. To investigate the effects of cell dosage on the reduction, bioconversion was performed at 30 °C and 160 rpm by adding E. coli CCZU-Y10 wet cells (0.02–0.40 g), 0.2 mmol COBE, 0.5 mmol mannitol, and 0.02 μmol NAD+ into the biphasic system containing 1 mL potassium phosphate buffer (100 mM, pH 7.0) and 1 mL dibutyl phthalate. To investigate the effects of substrate concentration on the reduction, biotransformations were carried out in monophasic aqueous and aqueous-organic system, respectively. The substrate concentration in the monophasic aqueous system was 50–200 mM, while the substrate concentration in the biphasic system was 200–1,000 mM. The 2-mL reaction mixture, comprising 0.1 μmol NAD+/mM COBE and a certain amount of wet cells and cosubstrate in 100 mM buffer was added into a 10-mL Erlenmeyer flask capped with a septum. After preincubation in a 160-rpm rotary shaker at selected temperature for 15 min, a certain volume of organic solvent containing 20 μL dodecane (internal standard) and a certain amount of COBE were added and then the incubation was continued. Samples were withdrawn periodically for the assay.
Analytical Methods
Unless otherwise specified, the reported substrate COBE and product CHBE concentration refer only to the concentration in the reaction media based on the organic phase. The concentrations of COBE and CHBE were assayed by GC (Chirasil-DEX CB, Varian, USA) equipped with a flame ionization detector and a CP WAX 52CB capillary column (30 m, i.d. 0.25 mm, film thickness 0.25 mm) [20]. The ee value of (S)-CHBE was assayed by HPLC equipped with a CHIRACEL OD-H column (Daicel Co., Japan, 250 mm × 4.6 mm) [8], which was eluted with hexane/isopropanol (90:10, v/v, 0.8 mL/min) and detected at 254 nm.
Yield of CHBE in the monophasic system was calculated as follows:
Yield of CHBE in the biphasic system was calculated as follows:
Results and Discussion
Asymmetric Reduction of COBE in the Monophasic Aqueous System
To employ reductase for the asymmetric reduction of prochiral ketones, an economical and efficient way for recycling NAD(P)H is needed. The oxidation of the cosubstrate could regenerate the required cofactor. Additionally, the cosubstrate might increase the conversion of bioreduction reaction [19–22]. Thus, eight potential cosubstrates (ethanol, fructose, glycerol, glucose, isopropanol, mannitol, mannose, and sucrose) were investigated in the monophasic aqueous system. As shown in Fig. 2a, all the tested cosubstrates could increase the reductase activity. Clearly, glucose and mannitol as cosubstrate could improve the reductase activity greatly. Using mannitol as a cosubstrate, however, the reductase activity was increased 3.7-fold compared to that when no cosubstrate was added. It was reported that mannitol was chosen as the appropriate cosubstrate for the reductase of Rhodotorula sp. AS2.2241, while other compounds such as glucose, ethanol, and 2-propanol had no obvious effect on the bioreduction reaction [21]. A substrate-coupled biocatalytic process was developed based on the reactions catalyzed by an NADPH-dependent reductase from Candida albicans, and the sorbitol/mannitol driven system produced more (S)-CHBE than the xylitol cosubstrate system [22]. In this study, no overexpression operation of coenzyme-recycling enzyme was performed. Clearly, the recombinant E. coli CCZU-Y10 had a substrate-coupled system. It was reported that the substrate-coupled system had been employed for the efficient and cost-effective cofactor-recycling systems [7, 8, 14]. Thereby, mannitol was employed as the optimum cosubstrate in this study, and a substrate-coupled bioprocess was demonstrated using mannitol as a cosubstrate. Moreover, the effects of cosubstrate concentration (0.25–12.5 mM mannitol/mM COBE) were also investigated on the reductase activity. As shown in Fig. 2b, when the cosubstrate concentration was 2.5 mM mannitol/mM COBE, the reductase activity was increased significantly. When the cosubstrate concentration was increased from 2.5 to 7.5 mM mannitol/mM COBE, the reductase activity had no significant increase. Therefore, the optimum amount of cosubstrate was 2.5 mM mannitol/mM COBE.
Furthermore, time courses for the reduction of COBE with the resting cells were investigated in the monophasic aqueous system (Fig. 3). Biotransformation of 50 mM COBE for 8 h, (S)-CHBE (>99 % ee) was obtained in a high yield (>99.0 %). Moreover, 80 mM COBE could be biotransformed into (S)-CHBE with the yield of 98.0 % after 24 h. However, COBE at 100 mM could be reduced into (S)-CHBE with the yield of 70.1 % after 24 h. Prolonging the reaction time for another 24 h, the yield had no significant increase. Therefore, it is necessary to enhance the biocatalytic efficiency for biotransforming the high concentration of COBE.
Establishment and Optimization of the Aqueous-Solvent Biphasic System
To improve the biocatalytic efficiency, a suitable aqueous-solvent biphasic system could be employed for the synthesis of key intermediates [23–25]. In the aqueous-solvent biphasic system, a potential biocompatible organic solvent could improve the biocatalytic activity [8, 23, 25]. Notably, the selection of the biocompatible organic solvent as the carrier for substrate and product is based on its biocompatibility towards the biocatalyst and adequate partition of both substrate and product to provide a pool for these chemicals [20]. In this study, six potential organic solvents (dibutyl phthalate, ethyl acetate, n-hexane, n-heptane, n-octane, and toluene) were tested on the reductase activity. All the ee values of (S)-CHBE were over 99 % (data not shown). As shown in Table 1, the reductase activities were well correlated with solvent log P values. The reductase activity of COBE increased with increases in the log P value of the solvent. Significantly, dibutyl phthalate (log P = 5.4) was found to be of higher biocompatibility than other organic solvents (log P < 5.4) for use as organic phase because of the reductase activity rate and the optical purity of product (S)-CHBE. Although both COBE and (S)-CHBE could be efficiently extracted into the organic layer using ethyl acetate (log P = 0.68), the lower reductase activity was observed because of its possible lower biocompatibility. Using n-hexane (log P = 3.5) as an organic phase, the reductase activity was not so ideal. This might be due to the unfavorable partition, resulting in the inhibitory effect of COBE and/or (S)-CHBE on the biocatalyst. Consequently, dibutyl phthalate (log P = 5.4) was chosen as an appropriate organic phase for the asymmetric reduction of COBE to (S)-CHBE. In an aqueous-solvent biphasic system, the volumetric phase ratio not only influenced the phase interfacial area but also affected bioconversion rates [20–25]. In the dibutyl phthalate-water biphasic system, the volumetric phase ratio had a significant influence on the initial reaction rate and conversion (Fig. 4), but had no effect on the optical purity of the product (S)-CHBE (data not shown). Significantly, the optimum volumetric phase ratio (organic phase versus water phase) was considered to be 1:1, under which a higher yield (81.6 %) with an excellent optical purity of (S)-CHBE (>99 % ee) was obtained.
During the biotransformation, reaction temperature, reaction pH, and cell dosage could significantly affect the biocatalytic activity [25–30]. As shown in Fig. 5a, in the dibutyl phthalate-water (1:1, v/v) biphasic system, it was found that the reductase activity significantly increased with the reaction temperature rising up to 30 °C. When the temperature further increased, the reductase activity dropped quickly. At temperatures above 30 °C, however, the reductase activity decreased considerably, possibly due to the thermal deactivation of reductase in the cells of E. coli CCZU-Y10 during the reductase reaction. Moreover, the effects of different reaction pH values (3–10) on the reductase activity were also investigated in the dibutyl phthalate-water (1:1, v/v) biphasic system. As shown in Fig. 5b, the optimum reaction pH was found to be 7.0. It was reported that the maximum activity of reductase from Streptomyces coelicolor was observed at pH 6.5 and more than 80 % of the maximum activity was retained between pH 6.0 and 7.5 [8]. Furthermore, the amounts of cell dosage added into the biphasic reaction media were investigated. As shown in Fig. 6, the formation of the product (S)-CHBE was considerably increased with increased cell dosage. Both the initial reaction rate and high conversion were obtained at 0.1 g (wet weight)/mL. When the cell concentration of E. coli CCZU-Y10 was above 0.1 g (wet weight)/mL, the viscosity of the aqueous phase increased so that the low dissolved oxygen (DO) in the aqueous phase might be prominent, and the initial reaction rate decreased significantly.
Based on the above results, the optimum reaction medium was the dibutyl phthalate-water (1:1, v/v) biphasic system. After the optimization in this biphasic system, the optimum reaction conditions were obtained: NAD+ 0.1 μmol/(mmol COBE), cosubstrate mannitol 2.5 mmol/(mmol COBE), reaction temperature 30 °C, reaction pH 7.0, and cell dosage 0.1 g (wet weight)/mL. With the biotransformation of 100 mM COBE for 12 h, the yield of (S)-CHBE was 92.2 % (Fig. 6).
Asymmetric Reduction of COBE in the Dibutyl Phthalate-Water Biphasic System
To effectively synthesize (S)-CHBE (>99 % ee), asymmetric bioreductions of COBE (200–1,000 mM) by recombinant E. coli CCZU-Y10 were carried out in the dibutyl phthalate-water biphasic system. As shown in Fig. 7, the yield of (S)-CHBE in the dibutyl phthalate-water biphasic system was higher than that in the monophasic aqueous system. The capacity for biotransforming COBE into (S)-CHBE was enhanced more greatly in the dibutyl phthalate-water biphasic system as compared with that in the aqueous phase (1,000 mM COBE in the dibutyl phthalate-water biphasic system versus 200 mM COBE in the monophasic aqueous system). With the biotransformation of 1,000 mM COBE in the biphasic system for 48 h, a high yield (99.0 %) was obtained. However, with the biotransformation of 200 mM COBE for 48 h in the monophasic aqueous system, the yield of (S)-CHBE was only 38.5 %. Significantly, the dibutyl phthalate-water biphasic system had more advantages than the monophasic aqueous phase for the asymmetric reduction of COBE catalyzed by the recombinant E. coli CCZU-Y10 cells. Simultaneously, the optical purity of the product (S)-CHBE (>99 % ee) had no significant change by introducing dibutyl phthalate as the organic phase (data not shown). Therefore, a highly stereoselective bioreduction of 1,000 mM COBE into (S)-CHBE by E. coli CCZU-Y10 was successfully demonstrated in the dibutyl phthalate-water biphasic system.
Conclusions
By genome data mining, an NADH-dependent reductase (PgCR) from P. guilliermondii CGMCC 2.1801 was discovered. After PgCR was overexpressed in E. coli BL21, the recombinant E. coli CCZU-Y10 with high biocatalytic activity and excellent stereoselectivity was employed for the reduction of COBE into (S)-CHBE (>99 % ee) using mannitol as a cosubstrate. The good performance was obtained in the dibutyl phthalate-water biphasic system. After the reaction optimization, the optimum reaction temperature, reaction pH, cosubstrate, NAD+, and cell dosage were 30 °C, 7.0, mannitol (2.5 mmol/mmol COBE), 0.1 μmol/(mmol COBE), and 0.1 g (wet weight)/mL, respectively. Furthermore, a high concentration of COBE (1,000 mM) could be asymmetrically reduced to (S)-CHBE with high yield (99.0 %) and excellent enantiometric excess value (>99 % ee).
References
Andres, I., Ana, C., Lorena, W., & Guillermo, R. C. (2012). Bioresource Technology, 115, 48–57.
Ni, Y., Zhou, J. Y., & Sun, Z. H. (2012). Process Biochemistry, 47, 1042–1048.
Asako, H., Shimizu, M., & Itoh, N. (2009). Applied Microbiology Biotechnology, 84, 397–405.
Ma, S. K., Gruber, J., Davis, C., Newman, L., Gray, D., Wang, A., Grate, J., Huisman, G. W., & Sheldon, R. A. (2010). Green Chemistry, 12, 81–86.
Barrios-González, J., & Miranda, R. U. (2010). Applied Microbiology Biotechnology, 85, 869–883.
Ye, Q., Cao, H., Yan, M., Cao, F., Zhang, Y. Y., Li, X. M., Xu, L., Chen, Y., Xiong, J., Ouyang, P. K., & Ying, H. J. (2010). Bioresource Technology, 101, 6761–6767.
Cao, H., Mi, L., Ye, Q., Zhang, G. L., Yan, M., Wang, Y., Zhang, Y. Y., Li, X. M., Xu, L., Xiong, J., Ouyang, P. K., & Ying, H. J. (2011). Bioresource Technology, 102, 1733–1739.
Wang, L. J., Li, C. X., Ni, Y., Zhang, J., Liu, X., & Xu, J. H. (2011). Bioresource Technology, 102, 7023–7028.
Ema, T., Yagasaki, H., Okita, N., Takeda, M., & Sakai, T. (2006). Tetrahedron, 62, 6143–6149.
Ema, T., Ide, S., Okita, N., & Sakai, T. (2008). Advanced Synthesis and Catalysis, 350, 2039–2044.
Yamamoto, H., Mitsuhashi, K., Kimoto, N., Esaki, N., & Kobayshi, Y. (2004). Bioscience, Biotechnology, and Biochemistry, 68, 638–649.
Goldberg, K., Schroer, K., Lütz, S., & Liese, A. (2007). Applied Microbiology Biotechnology, 76, 237–248.
Kizaki, N., Yasohara, Y., Hasegawa, J., Wada, M., Kataoka, M., & Shimizu, S. (2001). Applied Microbiology Biotechnology, 55, 590–595.
Xu, G. C., Yu, H. L., Zhang, X. Y., & Xu, J. H. (2012). ACS Catalysis, 2, 2566–2571.
Kimura, N. (2006). Microbe Environment, 21, 201–215.
Vartoukian, S. R., Palmer, R. M., & Wade, W. G. (2010). FEMS Microbiology Letters, 309, 1–7.
Gong, J. S., Lu, Z. M., Li, H., Zhou, Z. M., Shi, J. S., & Xu, Z. H. (2013). Applied Microbiology and Biotechnology, 97, 6603–6611.
Luo, X. J., Yu, H. L., & Xu, J. H. (2012). Enzyme Engineering, 1, 104–108.
Yasohara, Y., Kizaki, N., Hasegawa, J., Wada, M., Kataoka, M., & Shimizu, S. (2000). Bioscience, Biotechnology, and Biochemistry, 64, 1430–1436.
He, J. Y., Sun, Z. H., Ruan, W. J., & Xu, Y. (2006). Process Biochemistry, 41, 244–249.
Yang, W., Xu, J. H., Pan, J., Xu, Y., & Wang, Z. L. (2008). Biochemical Engineering Journal, 42, 1–5.
Cai, P., An, M. D., Xu, L., Xu, S., Hao, N., Li, Y., Guo, K., & Yan, M. (2012). Biotechnology Letters, 34, 2223–2227.
He, Y. C., Zhou, Q., Ma, C. L., Cai, Z. Q., Wang, L. Q., Zhao, X. Y., Chen, Q., Gao, D. Z., Zheng, M., Wang, X. D., & Sun, Q. (2012). Bioresource Technology, 115, 88–95.
Zhang, Z. J., Pan, J., Liu, J. F., Xu, J. H., He, Y. C., & Liu, Y. Y. (2011). Journal of Biotechnology, 152, 24–29.
He, Y. C., Ma, C. L., Yang, Z. X., Zhou, M., Xing, Z., Ma, J. T., & Yu, H. L. (2013). Applied Microbiology Biotechnology, 97, 10329–10337.
He, Y. C., Xu, J. H., & Su, J. H. (2010). Applied Biochemistry and Biotechnology, 160, 1428–1440.
Jiang, Y. J., Wang, Q., He, Y., Zhou, L. Y., & Gao, J. (2014). Applied Biochemistry and Biotechnology, 172, 2496–2506.
He, Y. C., Liu, Y. Y., Ma, C. L., & Xu, J. H. (2011). Biotechnology and Bioprocess Engineering, 16, 901–907.
Wang, D., Sun, J., Yu, H. L., Li, C. X., Bao, J., & Xu, J. H. (2012). Applied Biochemistry and Biotechnology, 166, 176–186.
Ji, Q. C., Xiao, S. J., He, B. F., & Liu, X. N. (2010). Journal of Molecular Catalysis B: Enzymatic, 66, 264–269.
Acknowledgments
All authors gratefully acknowledge support from the National Natural Science Foundation of China (no. 21102011), the Natural Science Foundation of Jiangsu Province, and the Open Project Program of the State Key Laboratory of Bioreactor Engineering (Shanghai, China).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
He, YC., Yang, ZX., Zhang, DP. et al. Biosynthesis of Ethyl (S)-4-Chloro-3-Hydroxybutanoate by NADH-Dependent Reductase from E. coli CCZU-Y10 Discovered by Genome Data Mining Using Mannitol as Cosubstrate. Appl Biochem Biotechnol 173, 2042–2053 (2014). https://doi.org/10.1007/s12010-014-1001-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-014-1001-4