Abstract
Enantiopure alcohols have received much attention due to their widespread use as pharmaceutical intermediates. In the asymmetric biosynthesis of enantiopure alcohols, the excellent performance of carbonyl reductase makes it be the best choice as the biocatalysts. In this work, an alkali-tolerant carbonyl reductase (BsCR, encoded by yueD) from Bacillus subtilis (strain 168) was obtained through gene mining, and successfully heterologously expressed in Escherichia coli with pET-32a. BsCR showed excellent alkali resistance and even can keep more than 70% of its peak activity after incubation in Tris–HCl buffer at pH 9.0 for 40 h. The Michaelis constants and maximal velocity of the BsCR to NADPH (A) and ethyl 4-chloroacetoacetate (B) are \(K_{m}^{A}\) = 5.390 × 10−2 mmol/L, \(K_{m}^{B}\) = 1.855 mmol/L, and \(V_{max}\) = 147.3 μmol·min−1·mg−1, respectively. Applying the E. coli BL21(DE3)/pET-32a-yueD to catalyze asymmetric reduction of ethyl 4-chloroacetoacetate and acetophenone, the yield of S-CHBE reached 89.9% and S-1-phenyl ethanol reached 66.7%, and e.e. of both products reached more than 99%. This work provides a novel CR for asymmetric reduction.
Graphic Abstract
A carbonyl reductase (BsCR) and its gene were identified through gene mining, and overexpressed in Escherichia coli BL21(DE3) for whole-cell biocatalytic asymmetric reduction

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Chiral alcohols are important pharmaceutical intermediates in various multi-step asymmetric synthesis due to their unique structural properties [1,2,3]. In general, the two enantiomers of the chiral compound tend to have different biological activities and opposite pharmacological activities. It is particularly important to obtain enantiomerically pure chiral alcohols [4]. Chiral alcohols can be synthesized from ketones by chemical or biological catalysis routes. In order to obtain highly stereoselective alcohols, chemical methods often require toxic metals, expensive hydrides, and stringent conditions compared to biological methods [5,6,7]. Thanks to the advantages of low pollution, remarkable enantioselectivity, regioselectivity and mild reaction conditions, carbonyl reductases (CRs) are widely applied to catalyze the reduction of various ketones to the corresponding chiral alcohols making considerable contributions to the industrial production of valuable chiral pharmaceutical intermediates including anticancer drugs, β-lactam antibiotics and other important drugs [8,9,10,11,12]. Because of the irreplaceable application prospect of CRs in asymmetric synthesis, they have attracted more and more attention and favor in the field of biocatalysts. To obtain enantiopure chiral alcohols, Wei et al. isolated a strain Acetobacter sp. CCTCC M209061 which can specifically catalyze ethyl 4-chloroacetoacetate (COBE) to chiral alcohols with anti-prelog stereospecificity [13]. Liu et al. reported a newly cloned CR expressed in recombinant Escherichia coli that catalyzes the asymmetric reduction of ET-4 to (S)-ET-5 [14]. Wang et al. cloned an NADH-dependent KlAKR and an NADPH-dependent KmAKR to synthesize the t–butyl 6–cyano–(3R,5R)–dihydroxyhexanoate [15, 16]. These developed enzymes can effectively reduce the cost for the chiral building blocks production. Nevertheless, due to its high specificity, many enzymes are difficult to adapt to the practical conditions, such as high-temperature environments [17], high substrate tolerance [18] and highly stereoselective product [19]. Therefore, there is a great interest to develop new CRs with high enantioselectivity and excellent application properties.
Bacillus subtilis is generally regarded as safe (GRAS) bacterium because of its free endotoxins, and it’s genetic, physiology are well characterized [20]. The enzymes from B. subtilis such as YqiG [21], BDH/AR [20] and P5C reductase [22] perform excellent catalytic ability. Especially, the GDHs (glucose dehydrogenase) from B. subtilis are often used for coenzyme regeneration in recombinant cells [20, 23, 24]. However, as laboratory strains, only a few CRs from B. subtilis were discovered. In recent years, with the rapid development of high-throughput gene sequencing technology, the whole DNA genome sequence of B. subtilis has been sequenced. On this basis, it is very economical and convenient to discover potential industrial enzymes in this bacterium. In the current study, we mined an NADPH-dependent CR from B. subtilis (strain 168), which has not been reported before. The corresponding gene yueD was cloned, and expressed in E. coli BL21 (DE3) cells with pET-32a vector. Its enzymatic properties were detailly investigated. Furthermore, its application as a catalyst for the asymmetric synthesis of chiral alcohols was preliminarily studied.
2 Materials and Methods
2.1 Chemicals, Plasmids and Strains
The enzymes for gene manipulation were purchased from New England Biolabs (Beijing) Ltd. The DNA markers were purchased from Vazyme Biotech (Nanjing) Co., Ltd. 2×Taq PCR Master Mix, Plasmid Miniprep Kit, Gel Extraction Kit and PCR Purification Kit were purchased from TIANGEN Biotech (Beijing) Co., Ltd. The primers were synthesized by Quintiles Biosciences (Wuhan) Co., Ltd. Other chemicals were purchased from Sigma-Aldrich and Aladdin.
All strains and plasmids used in this study are listed in Table 1. E. coli DH5α and E. coli BL21(DE3) were used for plasmid amplification and expression of a host cell, respectively. The pET-32a plasmid with ampicillin resistance was used as the cloning and expression vector. All strains were cultured in LB medium (10 g/L tryptone, 10 g/L NaCl, 5 g/L yeast extract, 1 L deionized H2O), and the antibiotics ampicillin, if needed, were added at a concentration of 100 μg/mL.
2.2 Sequence Analysis and Molecular Docking
Potential CRs were screened from the UniProt knowledgebase and BLAST tool at NCBI was used to check the gene of the CRs. The amino acid sequence and structure alignment of CR were analyzed on the website of SWISS-MODEL (https://www.swissmodel.expasy.org/), CLUSTALW (https://www.genome.jp/tools-bin/clustalw) and Espript 3.0 (http://espript.ibcp.fr/ESPript/ESPript/index.php). Molecular docking between substrate and CR was conducted by the AutoDock 4.2.
2.3 Construction of the Plasmid pET-32a-yueD and E. coli BL21(DE3)/pET-32a-yueD
The gene yueD was obtained by PCR amplification using B. subtilis 168 chromosomal DNA as the template [reaction conditions: initial denaturation: 95 °C, 2 min; 34×(denaturation: 95 °C, 10 s; annealing: 63 °C, 20 s; extension: 72 °C, 30 s) final extension: 72 °C, 10 min]. The forward primer is: 5′-CGCG/GATCCATGGAACTTTATATCATCACCGGAG-3′ (with BamHI site, G/GATCC), and the reverse primer is: 5′-CCGC/TCGAGCAAAAACTCTTTAATATCATAAATGCG-3′ (with XhoI site, C/TCGAG). As shown in Supplementary Scheme 1, the pET-32a-yueD vector was constructed by ligation reaction with T4 ligase between the digestion product of PCR and pET-32a digested by BamHI and XhoI, which was the expression vector for the target protein BsCR. After the recombinant plasmids were transformed E. coli DH5α by electroporation for plasmid amplification, the positive transformant was identified with colony PCR and verified with double restriction endonuclease digestion. Further, the recombinant plasmid was sequenced to confirm that the yueD gene was correctly ligated into the plasmid vector. To express the yueD gene, the vector pET-32a-yueD was anew transformed E. coli BL21(DE3) by electroporation. The transformant E. coli BL21(DE3)/pET-32a-yueD was selected with LB medium agar plate with ampicillin, then the recombinant E. coli BL21(DE3)/pET-32a-yueD was obtained.
2.4 Expression and Purification of BsCR
E. coli BL21(DE3)/pET-32a-yueD was cultivated in LB medium containing 100 μg/mL ampicillin at 37 °C with 220 rpm in a constant temperature shaker. When the optical density at 600 nm (OD600) of the culture reached 0.6–0.8, IPTG (Isopropyl β-D-Thiogalactoside, 0.4 mM) was added to induce BsCR expression at 24 °C for 12 h. The cells were collected by centrifuge (10,000×g for 10 min) at 4 °C and then washed twice with 0.1 mol/L phosphate buffer (PBS, pH 7.0). To release the expression product, the cells were lysed with sonication. The supernatant was collected with centrifuge (10,000×g for 10 min) at 4 °C, which was BsCR crude enzyme solution. BsCR was further purified by affinity chromatography with Ni2+ affinity resin (Profinity IMAC Resins, Bio-rad Shanghai, China). The expressed and purified BsCR were detected by SDS-PAGE.
2.5 Enzymatic Properties of BsCR
The following enzymatic properties of BsCR expressed by the E. coli BL21(DE3)/pET-32a-yueD were evaluated: the optimum pH, temperature and effects of metal ions (include EDTA, a metal chelating agent). The temperature range was from 25 to 65 °C. The pH range was from 5.0 to 9.0, the buffering systems at different pH were: citric acid buffer for pH 5.0–6.0, sodium phosphate buffer for pH 6.0–8.0, Tris–HCl buffer for pH 8.0–9.0. The concentration of metal ions was 1 mmol/L. The thermal and pH stability were also evaluated. Furthermore, its kinetic parameters were assessed. All the above experiments were completed in triplicate, the average values were calculated based on the three independent experiments, and the standard deviation was calculated.
2.6 Asymmetric Reduction Reactions of Prochiral Ketones by Whole-cell
To assess the application prospects of BsCR, the E. coli BL21(DE3)/pET-32a-yueD was applied to catalyze asymmetric reduction reactions of COBE and acetophenone to the corresponding chiral alcohols (CHBE and 1-phenyl ethanol). The general procedure as follows: 10 mL PBS buffer (100 mM, pH 6.5) containing 40 mM substrate, 500 mM glucose and 100 mg/mL wet-cells was added to a 25 mL stoppered flask capped with a septum. The reaction mixture was shaken in an air-bath shaker at 250 rpm, 37 °C and samples (800 μL) were taken regularly. The product and the residual substrate were extracted from the sample with 2 × 800 μL of ethyl acetate containing 40 mM benzaldehyde (as an internal standard) and dried over anhydrous Na2SO4. Finally, the concentrations of the product and substrate were analyzed by GC (Gas chromatography) and HPLC (High-performance liquid chromatography), the chemical yield and enantioselectivity were then evaluated.
2.7 Analytical Methods
2.7.1 Enzyme Activity Assay
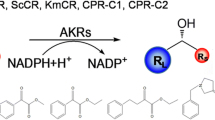
The molecular mass of BsCR was measured by SDS-PAGE with 12% separating gel. According to the catalytic reaction Eq. (1), the activity was assayed at 30 °C by monitoring the decrease in absorbance of NADPH at 340 nm using prochiral ketones as the substrate ([S] represents the substrates: COBE, ethyl acetoacetate, acetone, 2-octanone, acetophenone, α-chloroacetophenone, 2′-fluoroacetophenone, 3′-fluoroacetophenone, 4′-fluoroacetophenone and 2,2,2-trifluoroacetophenone. [P] represents the corresponding catalytic products).
The reaction mixture consisted of 100 mM of sodium phosphate buffer (pH 6.5), 5 mM of substrate, 0.5 mM of NADPH, and 50 μL of BsCR solution in a total volume of 200 μL. One unit of enzyme activity is defined as the amount of enzyme that catalyzes the oxidation of 1 μmol of NADPH per min. The potential of NADH and NADPH to act as electron donors was tested by replacing NADPH with NADH during the reaction.
2.7.2 Kinetic Analysis
A double substrate Michaelis–Menten equation was applied as BsCR kinetic equation. The kinetic parameters of BsCR were assessed according to the initial-velocity double-reciprocal plot by using COBE as the substrate at varying concentrations (2.0, 3.0, 4.0, and 5.0 mM), and the NADPH concentrations for 0.10, 0.15, 0.20, 0.25 and 0.3 mM. BsCR catalyzed-reaction followed the ordered sequential reaction mechanism.
The double substrate Michaelis–Menten equation is presented as follows:
where [A] represents the concentration of NADPH, [B] represents the concentration of COBE, Vmax is the maximal velocity, \(K_{{m}}^{{A}}\) and \(K_{m}^{B}\) are the Michaelis constants of the purified BsCR for NADPH and COBE. \(K_{s}^{A}\) is the dissociation constant for the binding of BsCR and NADPH.
The slopes of a series of 1/v versus 1/[A] are calculated to be:
The vertical intercepts of straight lines are calculated to be:
2.7.3 Asymmetric Reduction Reaction Assay
The concentrations of product and substrate were quantitatively analyzed by GC (GC-2010 plus, Shimadzu) equipped with an RT-βDEXm chiral column (Restek, America, 30 m × 0.32 mmID × 0.25 μm df) and FID detector. The configurations of phenyl ethanol were determined by GC and the configurations of ethyl S-4-chloro-3-hydroxybutyrate (CHBE) were determined by HPLC (UltiMate3000, Thermo Fisher Scientific). The HPLC equipped with a CHIRALCEL OB column (4.6 × 250 mm) (Daicel Chemical Industries, Japan), and conditions were hexane/2-propanol (85/15, v/v) as mobile phase, flow rate of 0.8 mL/min, ambient column temperature, and ultraviolet (UV) detection set at 220 nm. The analysis method was the same as our previous work [25, 26].
The extent of reaction and enantioselectivity are indicated by yield (chemical yield, %) and e.e. (enantiomeric excess, %), respectively, defined as follows:
where C[S] is the initial substrate concentration, C[P] is the final product concentration, CS is the final S-configuration product concentration, and CR is the final R-configuration product concentration.
3 Results and Discussion
3.1 Gene Mining Result
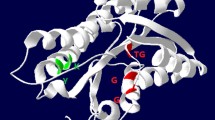
Previously, a carbonyl reductase (BYueD) from Bacillus sp. ECU0013 has been reported. As a versatile reductase, the enzyme has a broad substrate and exhibits excellent diastereoselectivity [27]. In this work, the B. subtilis (strain 168) was chosen as the source strain for mining the target CR. A CR was obtained by the similarity of the amino acid sequences between B. subtilis (strain 168) and BYueD. This CR was encoded by the gene yueD (Gene ID: 936558) and named BsCR. The sequence of BsCR contains 243 amino acids and has 40.24% identity to the yueD protein isolated from Bacillus cereus by Maruyama et al. [28]. The amino acid sequence analysis results show that BsCR contains the active center conserved region Tyr-X-X-X-Lys (position 147–151) and the cofactor binding site conserved region Thr-Gly-X-X-X-Gly-X-Gly (position 7–14), which belongs to the short-chain dehydrogenases/reductases family (SDRs) [27]. The alignment of multiple amino acid sequences of BsCR with other three SDRs is shown in Fig. 1. The structure of BsCR was built based on the protein PDB: 4H15 as the model and the substrate molecule COBE which was docked into the BsCR with program AutoDock 4.2. The protein was set to be rigid and the small molecule to be a flexible residue during the docking process. As shown in Fig. 2a, the protein structure of BsCR has four subunits. Combining with the results of the sequence analysis, the cofactor binding site, the active center conserved region and the substrate binding site have been labeled in the figure. Further, the molecular docking results are shown in Fig. 2b. The results indicate that this CR has the potential ability to reduce carbonyl compounds.
Alignment of multiple amino acid sequences of BsCR and other three SDRs. A conserved coenzyme-binding motif Thr-Gly-X-X-X-Gly-X-Gly (position 7–14) was found in four sequences, which means both of them preferred NADPH to NADH. The conserved motif Tyr-X-X-X-Lys (position 147–151) was regarded as the active center. The Asn (position 80), Gly (position 82) and Thr (position 184) were regarded as the substrate binding site
3.2 Construction of the Plasmid pET-32a-yueD and E. coli BL21(DE3)/pET-32a-yueD
In order to explore the properties of BsCR, an efficient expression vector pET-32a-yueD containing the yueD gene was constructed to express the BsCR in E. coli. Supplementary Figs. 1 and 2 were the agarose gel electrophoresis results of yueD PCR product and double enzymes (BamHI and XhoI) digestion product of pET-32a-yueD. First, the size of the PCR product was about 730 bp, which was consistent with the 723 bp of the gene shown in NCBI. Second, the double enzymes digestion product shows that there were two bands: one was about 5900 bp for the plasmid pET-32a, and the other was about 723 bp for yueD gene. Also, the cloned yueD was sequenced and aligned with the DNA sequence of Gene ID: 936558. The result was presented in Supplementary Fig. 3. All of the results evidenced that the expression vector pET-32a-yueD was successfully constructed.
3.3 Expression and Purification of BsCR
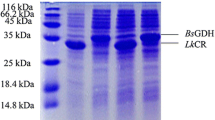
For expression of yueD gene, the vector pET-32a-yueD was transformed into E. coli BL21(DE3) to obtain the heterologous overexpression system, E. coli BL2 l (DE3)/pET-32a-yueD. The yueD was expressed by E. coli BL21(DE3)/pET-32a-yueD in LB medium with low-temperature induction after IPTG was added. The proteins were extracted by the afore mentioned method and detected by SDS-PAGE. The results, shown in Fig. 3a, indicated that the BsCR was successfully expressed with a molecular weight of 31 kDa, which is consistent with the value predicted based on the amino acid sequence. Based on the presence of a 6×His tag at the C terminus, the affinity chromatography with Ni2+ affinity resin was used to successfully purify the target protein. The BsCR was collected from the eluent. Figure 3b is the SDS-PAGE for the purified product.
SDS-PAGE for the BsCR expressed in E. coli BL21(DE3). a Total proteins in E. coli BL21(DE3), M: Protein Marker; Lane 1: Proteins from E. coli BL21(DE3)/pET-32a; Lane 2: Proteins from E. coli BL21(DE3)/pET-32a-yueD. b Purified BsCR overexpressed in E. coli BL21(DE3), M: Protein Marker; Lane 1: flow-through of the crude extract; Lane 2: purified BsCR
3.4 Enzymatic Properties of BsCR
BsCR showed activity only when NADPH was used as the coenzyme, this indicated that it is an NADPH depend CR, which is consistent with amino acid sequence analysis results. To investigate the substrate specificity of BsCR, various ketones were examined as the substrates for this CR. The results shown in Table 2 demonstrated that BsCR exhibit wide substrate spectrum. Specially, it shows superior catalytic ability to halogenated carbonyl compounds and long-chain carbonyl compounds.
The effect of metal ions and EDTA on BsCR activity was shown in Table 3. The activity of BsCR was slightly inhibited by 1 mM Mg2+ and Ba2+ while enhanced by the other metal ions to a varying extent. EDTA had a negligible effect on BsCR activity, this indicated that BsCR was a nonmetallic dependent enzyme, which followed the SDRs family feature.
The effect of pH on BsCR activity was studied at 30 °C, in different buffer systems (200 mM). As shown in Fig. 4a, the optimum reaction pH for the BsCR is about 6.3. Compared to the activity in sodium phosphate buffer, BsCR showed better activity in citric acid buffer at the same pH values. The pH stability of BsCR was assessed by detecting the residual activity in various buffers with pH from 5.0 to 9.0 at 4 °C for up to 40 h. Samples were withdrawn at an appropriate time and the residual activity was detected under the standard assay conditions with the unincubated BsCR as the control. The results, as shown in Fig. 4b, demonstrated that BsCR was quite stable at pH 7.0, and can keep more than 70% of its peak activity after being incubated in pH 9.0 buffer for 40 h. This means that BsCR is more advantageous for storage in alkaline environments than most CRs such as PsCRII [30], LcSDR [31], and ScCR [32].
The effect of temperature on BsCR activity was investigated at temperatures ranging from 25 to 65 °C. As shown in Fig. 4c, within a temperature range of 25–45 °C, BsCR activity increased with temperature and peaked at 45 °C. When the temperature was above 45 °C, BsCR activity decreased rapidly due to the thermal inactivation. Therefore, we can deduce that the optimum temperature is around 45 °C. To evaluate its thermo-stability, BsCR was incubated at 25, 35, 45 and 55 °C. As presented in Fig. 4d, its half-life (t1/2) is 18.57, 12.99, 0.76 and 0.06 h at 25, 35, 45 and 55 °C respectively.
3.5 Kinetic Analysis
The enzymatic properties are the fundamental enzymology data to enzyme research and its application. The kinetic parameters of the BsCR expressed by E. coli BL21(DE3)/pET-32a-yueD were evaluated based on the double substrate Michaelis–Menten model. From the data in Fig. 5a, it is apparent that these straight lines intersected at the same point (− 1.1195, 0.0078). In addition, the slope and intercept of each line were plotted as shown in Fig. 5b, c. From the results of the correlational analysis, the values of \(V_{max}\), \(K_{m}^{A}\), and \(K_{m}^{B}\) were 147.3 μmol·min−1·mg−1, 5.390 × 10−2 mM and 1.855 mM, respectively. Compared with the reported SDR, such as BcCR (0.22 μmol·min−1·mg−1) [29] and LcSDR (133.9 μmol·min−1·mg−1) [31], the BsCR has a higher \(V_{max}\), which means greater prospects in industrial applications.
3.6 Asymmetric Reduction Reactions of Prochiral Ketones by Whole-cell
To investigate the catalysis capability of the recombinant E. coli BL21(DE3)/pET-32a-yueD, the asymmetric reduction reactions of prochiral ketones are carried out with whole-cell. Due to the product, chiralCHBE and 1-phenyl ethanol are key important chiral building blocks and which were widely researched, COBE and acetophenone were chosen as the model substrates for β-oxo ester and aromatic ketone respectively. Results were given in Fig. 6. After investigated, the optical purity of CHBE and phenyl ethanol were over 99% e.e. for S-configuration. As shown in Fig. 6a, when the reaction was carried out for 4 h, the yield of S-CHBE reached 89.9%. In Fig. 6b, with the increase of reaction time, the yield of S-1-phenyl ethanol increased from 22.2% at 0.5 h to 66.7% at 12 h. These indicate that E. coli BL21(DE3)/pET-32a-yueD show good catalysis capability to β-oxo ester and aromatic ketone and display potential application prospects. For example, the CR from Streptomyces coelicolor can only catalyze the conversion of acetophenone in 55% yield but get a great application in water/toluene biphasic system [32]. This also provides a direction for our further researches.
4 Conclusion
With the increasing availability of public genome information, many putative CRs can be obtained from GenBank. Compared to traditional CR screening, the use of gene mining technology can not only improve efficiency but also save costs. In this work, an alkali-tolerant CR (BsCR) was discovered from B. subtilis (strain 168) through gene mining. The gene of BsCR was cloned and successfully expressed in E. coli BL21 (DE3) with pET-32a-yueD vector. Further, the enzymatic properties of BsCR were studied. The results showed that the optimal reaction pH and temperature for the CR are 6.3 and 45 °C respectively. The most unexpected result to emerge from the data is that BsCR exhibits excellent alkali resistance, and there are literature reported describing the characteristics of an alkali-resistant CR. The Michaelis constants and maximal velocity of BsCR to NADPH and COBE are \(K_{m}^{A}\) = 5.390 × 10−2 mM, \(K_{m}^{B}\) = 1.855 mM, and \(V_{max}\) = 147.3 μmol·min−1·mg−1. The asymmetric reduction reactions of prochiral ketones catalyzed by the E. coli BL21(DE3)/pET32a-yueD was investigated. Without additional NADPH addition, the yield of S-CHBE reached 89.9% at 4 h, and the yield of S-1-phenyl ethanol reached 66.7% at 12 h. The reaction show excellent enantioselectivity. This work not only indicates a detailed understanding of the properties and applications of BsCR, but further demonstrates that gene mining technology provides an efficient route to discover new CRs from existing strains.
References
Xu P, Zheng GW, Du PX, Zong MH, Lou WY (2016) ACS Sustain Chem Eng 4:371–386
Wei P, Liang J, Cheng J, Zong MH, Lou WY (2016) Microb Cell Fact 15:5
Schmid A, Dordick JS, Hauer B, Kiener A, Wubbolts M, Witholt B (2001) Nature 409:258–268
Bian GL, Yang SW, Huang HY, Zong H, Song L (2016) Sens Actuators, B 231:129–134
Ward TJ, Ward KD (2012) Anal Chem 84:626–635
Bredikhina ZA, Kurenkov AV, Krivolapov DB, Bredikhin A (2015) Tetrahedron Asymmetry 26:577–583
Bai DY, He JY, Ouyang B, Huang J, Wang P (2017) Prog Chem 29:491–501
Zheng YG, Yin HH, Yu DF, Chen X, Tang XL, Zhang XJ, Xue YP, Wang YJ, Liu ZQ (2017) Appl Microbiol Biotechnol 101:987–1001
Itoh N (2014) Appl Microbiol Biotechnol 98:3889–3904
Malatkova P, Wsol V (2015) Drug Metab Rev 46:96–123
Shi SM, Di L (2017) Expert Opin Drug Metab Toxicol 13:859–870
Zhang RZ, Xu Y, Xiao R (2015) Biotechnol Adv 33:1671–1684
Wei P, Gao JX, Zheng GW, Wu H, Zong MH, Lou WY (2016) J Biotechnol 230:54–62
Liu ZQ, Dong SC, Yin HH, Xue YP, Tang XL, Zhang XJ, He JY, Zheng YG (2017) Bioresour Technol 229:26–32
Wang YJ, Ying BB, Shen W, Zheng RC, Zheng YG (2017) Enzyme Microb Technol 107:32–40
Wang YJ, Shen W, Luo X, Liu ZQ, Zheng YG (2017) Biotechnol Prog 33:1235–1242
Fukuda Y, Sakuraba H, Araki T, Ohshima T, Kazunari Y (2016) Enzyme Microb Technol 91:17–25
Xie Y, Xu JH, Xu Y (2010) Bioresour Technol 101:1054–1059
Ni Y, Li CX, Ma HM, Zhang J, Xu JH (2011) Appl Microbiol Biotechnol 89:1111–1118
Samuel N, Bao T, Zhang X, Yang T, Xu M, Li X, Komera I, Philibert T, Rao Z (2017) J Chem Technol Biotechnol 92:2477–2487
Sheng X, Yan M, Xu L, Wei M (2016) J Mol Catal B 130:14–18
Forlana G, Nocek B, Chakravarthy S, Joachimiak A (2017) Front Microbiol 8:1442
Cui Z, Zhang J, Fan X, Zheng G, Chang H, Wei W (2017) J Biotechnol 243:1–9
Chen X, Mei T, Cui Y, Chen Q, Liu X, Feng J, Wu Q, Zhu D (2015) ChemistryOpen 4:483–488
Luo W, Deng XX, Gong ZW, Yang ZH (2016) Asia-Pac J Chem Eng 11:2408–2411
Luo W, Deng XX, Huo J, Ruan T, Gong ZW, Yan JB, Yang ZH, Quan C, Cui ZF (2018) Catal Lett 148:1714–1722
Ni Y, Li CX, Wang LJ, Zhang J, Xu JH (2011) Org Biomol Chem 9:5463–5468
Maruyama R, Nishizawa M, Itoi Y, Ito S, Inoue M (2010) Biotechnol Bioeng 75:630–633
Qin YL, Ruan T, Hou HS, Hou YL, Yang ZH, Quan C (2019) Catal Lett 149:610–618
Cao H, Mi L, Ye Q, Zang G, Yan M, Wang Y, Zhang Y, Li X, Xu L, Xiong J, Ouyang P, Ying H (2011) Bioresour Technol 102:1733–1739
Wang YJ, Ying BB, Min C, Shen W, Liu ZQ, Zheng YG (2017) World J Microbiol Biotechnol 33:144
Wang LJ, Li CX, Ni Y, Zhang J, Liu X, Xu JH (2011) Bioresour Technol 102:7023–7028
Acknowledgments
The authors acknowledge all the financial supports for this research by the National Natural Science Foundation of China (Grant No. 21376184), Foundation from the Educational Commission of Hubei Province of China (Grant No. D20121108), the National Key Research and Development Project (2017YFF0205803, Ministry of Science and Technology of China), and the National Institute of Metrology of China (21-AKY1615).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Luo, W., Du, HJ., Bonku, E.M. et al. An Alkali-tolerant Carbonyl Reductase from Bacillus subtilis by Gene Mining: Identification and Application. Catal Lett 149, 2973–2983 (2019). https://doi.org/10.1007/s10562-019-02873-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10562-019-02873-w