Abstract
Microbial contamination during fish flesh spoilage process can easily induce food-borne outbreaks and consumer health problems. Hyperspectral imaging in the spectral range of 400–1000 nm was developed to measure the Escherichia coli (E. coli) loads in grass carp fish for evaluation and visualization of microbial spoilage. Partial least square regression (PLSR) model was conducted to build prediction models between the spectral data and the reference E. coli loads estimated by classical microbiological plating method. The PLSR model based on full wavelengths showed good performance on predicting E. coli loads with the residual predictive deviation (RPD) of 5.47 and determination coefficient of R 2 P = 0.880. Six characteristic wavelengths were selected by the weighted regression coefficients from PLSR analysis and used to simplify the models. The simplified PLSR and multiple linear regression (MLR) models also presented good prediction capability. The better simplified MLR model (RPD = 5.22 and R 2 P = 0.870) was used to transfer each pixel in the image for visualizing the spatial distribution of E. coli loads. The results demonstrated that hyperspectral imaging technique with multivariate analysis has the potential to rapidly and non-invasively quantify and visualize the E. coli loads in grass carp fish flesh during the spoilage process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, food quality and safety control has received special emphasis and great social concern from the government and the public. Many effective measures and techniques such as drying (Sun and Byrne 1998; Sun and Woods 1997; Delgado and Sun 2002a, b), refrigeration (Sun 1997; Sun et al. 1996; McDonald and Sun 2001; Kiani and Sun, 2011) and edible coating (Xu et al. 2001) have been taken ensure food quality and safety. On the other hand, with the rapid development of camera technology and the processing power of computer hardware, imaging techniques such as computer vision (Jackman et al. 2008, 2009; Sun 2004; Valous et al. 2009; Wang and Sun 2002) have particular advantages in rapid, non-contact and non-destructive detection of food quality and safety. Hyperspectral imaging (HSI) originated from remote sensing, as a promising imaging technique shows its superiority and has recently emerged as a powerful analytical tool for rapid and non-destructive quality and safety analysis and evaluation of food (Feng and Sun 2012; Gowen et al. 2007; Sun 2010), fruit and vegetables (Lorente et al. 2012), meat (Barbin et al. 2012; Elmasry et al. 2011a, b, 2012a, b; Kamruzzaman et al. 2011, 2012), agriculture and agro-food product (Liu et al. 2013) and fish and seafood (Cheng and Sun 2014; Menesatti et al. 2010). HSI integrates conventional imaging and spectroscopy technology into one system to achieve both spatial and spectral information from an object. The obtained hyperspectral images normally called hypercubes (x, y and λ) characterize three-dimensional (3-D) data cubes, which are composed of hundreds of contiguous wavebands for each spatial position of a target studied (Sun 2010). Accordingly, the spatial-feature enables characterization of complex heterogeneous samples and image texture, while the spectral-feature allows for the identification of internal chemical information (Gowen et al. 2007).
It is well-known that fish is a kind of muscle food vulnerable to microbial contamination. Microbial contamination can easily cause food-borne outbreaks and consumer health problems (Siripatrawan et al. 2011). Escherichia coli (E. coli) is a common bacterium with the characteristics of rod-shaped, Gram-negative, facultatively anaerobic and non-spore forming (Cassin et al. 1998). E. coli O157:H7 is an enteric bacterium that has been implicated in food- and water-borne human illnesses worldwide, including bloody diarrhoea, hemolytic uremic syndrome and hemorrhagic colitis (Cassin et al. 1998). The current microbial detection methods commonly include the culture-based (Yeni et al. 2014), immunology-based (Iqbal et al. 2000) and polymerase chain reaction-based method (Nugen and Baeumner 2008). However, these techniques are generally time-consuming, laborious, destructive and invasive and require complicated sample preprocessing, which can lead to great difficulty for real-time and on-line monitoring in food manufacturing. HSI is capable of solving the problems mentioned above and has been proven to be feasible and successful for quality and safety evaluation of fish such as grass carp (Ctenopharyngodon idella) depending on some significant parameters mainly related to colour (Cheng et al. 2014a), textural firmness (Cheng et al. 2014b), total volatile basic nitrogen (TVB-N) value (Cheng et al. 2014c) and freshness (Cheng et al. 2013). On the other hand, some studies were also conducted on the potential of using hyperspectral imaging technique for evaluating E. coli contamination in pork meat (Tao et al. 2012; Tao and Peng 2014) and packaged fresh spinach (Siripatrawan et al. 2011), detecting Shiga toxin-producing E. coli serogroups on rainbow agar (Windham et al. 2013) and differentiating colonies of non-O157 Shiga-toxin producing E. coli serogroups on spread plates (Yoon et al. 2013) and on agar media (Windham et al. 2012). Although the results available in the above-mentioned studies proved that HSI has the potential for the detection of E. coli loads, the models established cannot be used to predict the bacterial contamination in fish due to the fact that fish flesh has its own distinct connective tissue and chemical components such as the special protein, fatty acids and astaxanthin contents, which to some extent affect the absorbance of substances and the selection of characteristic wavelengths. Further study is thus needed for detecting and quantifying E. coli loads in fish flesh using HSI technique.
Therefore, the major objective of this study was to investigate the potentiality and suitability of HSI in the spectral range of 400–1000 nm for the determination and visualization of E. coli loads in fish flesh during the spoilage process with multivariate analysis.
Material and Methods
Fish Samples Preparation
Fifteen fresh grass carps from the same batch with similar age of three months, approximately weight of 1.5 kg, and similar feeding environment from the same freshwater aquaculture ponds were purchased from a local aquatic products market in Guangzhou, China, and directly transported to the laboratory alive in water within 15 min. Upon arrival, the fish samples were stunned by a sharp blow to the head with a wooden stick and then gill cutting with a knife. The internal organs were removed along with bloodletting from the belly location of grass carp. Afterwards, they were instantly beheaded, filleted, skinned and washed with cold water. Thirty fish fillets each with similar size and weight were obtained. In order to acquire more fish samples for further building robust and reliable prediction models, the fresh fillets were immediately subsampled into a rectangular shape with similar size of 3.0 × 3.0 × 1.0 cm (length × width × thickness). Consequently, a total of 150 subsamples of fish fillets were obtained from different locations of the fish fillets. For the purpose of assembling a practical range of E. coli loads for indication of bacterial contamination degree from freshness to spoilage (inedible or unacceptable), all the subsamples were labelled and packaged into the sealed plastic bags and randomly divided into four groups. Among them, three groups named G1, G2 and G3 had 40 subsamples, respectively, and the fourth group (G4) had 30 subsamples. Then the four groups were sequentially subjected to postmortem spoilage during cold storage at 4 ± 1 °C for 0, 3, 6 and 9 days in a lab refrigerator (Haier Company, Qingdao, China) for further acquisition of hyperspectral images. The measurement of E. coli loads by traditional standard plate count method was conducted according to the previous study reported by Tao et al. (2012). Among the 150 subsamples, two thirds of the samples (100 subsamples) including 27G1, 27G2, 26G3 and 20G4 were used as the calibration set and the remaining one third samples (50 subsamples) consisting of 13G1, 13G2, 14G3 and 10G4 were utilized as the prediction set.
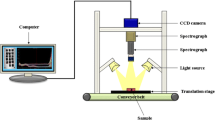
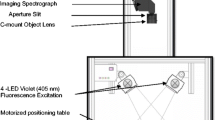
Hyperspectral Imaging System and Image Acquisition
A laboratory HSI system in a reflectance model was assembled to acquire hyperspectral images of grass carp fillets. The system consisted of an imaging spectrograph, a charge-coupled device (CCD) camera, an illumination system and a computer control system, and the detailed description of the system is available in the literature (Cheng et al. 2014b).
For image acquisition, based upon the cold storage conditions, at 3-day intervals, each group of subsamples were taken from the lab refrigerator and placed on the moving platform and then transfered to the field of the view of the camera to be scanned line by line for acquisition of hyperspectral images. Accordingly, a total of 150 three-dimensional (3-D) hyperspectral images were collected, documented and stored in a Band Interleaved by Line (BIL) format. In order to decrease the effects of illumination and detector sensitivity as well as the differences in camera and physical configuration of the imaging system, the raw acquired hyperspectral images (R 0) needed to be calibrated into the reflectance mode with two extra images for standard white (W) and black (B) reference images. The white reference image was acquired using a uniform Teflon white calibration tile (∼100 % reflectance). The black reference image (∼0 % reflectance) was obtained by fully covering the camera lens with its black cap. The calibrated image (R C) was then calculated by the following equation:
After acquisition and calibration of the hyperspectral images, the regions of interests (ROIs) were isolated from the fish subsamples, where E. coli loads were determined by the reference plate count method. The average spectral data within ROIs were manually extracted using the software ENVI version 4.8 (ITT Visual Information Solutions, Boulder, CO, USA). Then, the extracted spectral information and the corresponding traditional measured E. coli loads were used to conduct the quantitative analysis.
Multivariate Data Analysis
PLSR Analysis
The large spectral data extracted from the hyperspectral images normally include amounts of effective and valuable information and unavoidably some redundant and interferential information that affects the prediction performance. In order to improve the predictive robustness and reliability of models and reduce the variability between samples due to scattering and optical interference possibly caused by water movement during cold storage, in this study, the common spectral preprocessing method of multiplicative scatter correction (MSC) was used to remove the undesirable scatter effect from the data matrix prior to data modelling (Jin et al. 2011). After spectral preprocessing, PLSR as one of the most widely used algorithms was employed to establish the quantitive analysis for spectral data modelling. This regression analysis is useful to solve the colinearity problem and difficulty due to the number of variables being more than the number of samples, and the detailed description of PLSR was reported in the previous study (Mehmood et al. 2012). In this study, PLSR found a set of independent variables (wavelengths), called the X-matrix (100 × 381 and 50 × 381) in calibration and prediction model, and the dependent variable (E. coli loads), named the corresponding Y-matrix (100 × 1 and 50 × 1).
MLR Analysis
MLR is another method to establish the quantitative relationship between two or more explanatory independent variables and one dependent variable by fitting a linear equation to the observed data (Wu et al. 2012). The approach is competent when the number of samples is more than the number of variables. In this study, the number of variables was much greater than the number of samples (381 vs. 100 or 50). Therefore, after selection of the most important wavelengths, the application of MLR algorithm would be useful to establish a better model. The analyses of PLSR and MLR were carried out by the Unscrambler chemometric software (Unscrambler version 9.7, CAMO, Trondheim, Norway).
Characteristic Wavelengths Selection
Based on the above multivariate data analysis using the full spectral range of 400–1000 nm, multicolinearity among contiguous wavebands and high dimensionality of hyperspectral images can easily make data processing time-consuming with low computation speed. Variable selection can improve model performance and characteristics and facilitate the establishment of consistent hyperspectral imaging systems with simple structure, short acquisition time and low cost for real-time applications (Liu et al. 2014). Thus, it is interesting to allocate a set of optimal wavelengths that carry the most valuable information and may be equally or more efficient than the full wavelength range for providing satisfactory prediction results. Some frequently used variable selection methods such as genetic algorithm (Arakawa et al. 2011), PLS regression coefficients and stepwise regression (Mehmood et al. 2012), successive projections algorithm (Ghasemi-Varnamkhasti et al. 2012) and uninformative variable elimination (Balabin and Smirnov 2011) have been developed. In this study, the most sensitive wavelengths indicating E. coli contamination were identified and selected by calculating the weighted regression coefficients (WRC) method also called β-coefficients (B W ) from PLSR analysis with the full range of spectra. The wavelengths located at the highest and the lowest values of weighted regression coefficients were affirmed as the optimal wavelengths for further prediction of E. coli loads (Kamruzzaman et al. 2012). On the basis of the selected characteristic wavelengths, the simplified PLSR and MLR models also named WRC-PLSR and WRC-MLR models were generated and compared. The implementation procedure for variable selection was carried out in the Unscrambler chemometric software (Unscrambler version 9.7, CAMO, Trondheim, Norway).
Model Validation and Evaluation
Model validation is important for weighing the calibration models in multivariate data analysis. Validation refers to comparing the model predictions with a real-world dataset, for evaluation of its prediction accuracy. In this study, full cross-validation also called leave-one-out cross-validation was used to validate the established calibration models. The process of this technique was conducted by removing one sample or a subset of samples from the calibration data set and a new PLSR model was then built based on the remaining calibration samples (ElMasry and Wold 2008). In addition, the optimal number of latent variables (LV) from PLSR analysis was determined by the minimum value of predicted residual error sum of squares. The performance of the established models was commonly evaluated by calculating the residual predictive deviation (RPD), the coefficients of determination (R 2) and root mean square errors in calibration (R 2 C, RMSEC), cross-validation (R 2 CV, RMSECV) and prediction (R 2 P, RMSEP), respectively. Generally, an admirable and comparable model should have higher values of RPD, R 2 C, R 2 CV and R 2 P and lower values of RMSEC, RMSECV and RMSEP as well as a small difference between them. It is always expected to acquire RMSEs as close as zero and R 2 as close as one. According to Williams (2001), specifically, the value of R 2 of more than 0.90 shows excellent performance and lower than 0.82 means poor performance. As to RPD, RPD lower than 1.5 indicates the model established is not acceptable and larger than 3 means the model is satisfactory.
Visualization of Bacterial Distribution
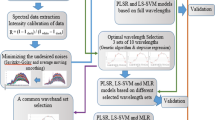
In order to clearly observe the degree of bacterial contamination in the fish flesh from sample to sample at different spoilage stages, visualization of E. coli loads distribution map is required instead of the measurement of the E. coli loads for the whole fish fillet. In the visualization process, each pixel in the images has a spectral profile with its spatial position. This was carried out by calculating the dot product between spectrum of each pixel in the image and the regression coefficients achieved from the simplified model, which was used to transfer and visualize every pixel of the hyperspectral images into the chemical images for the exhibition of E. coli loads distribution of the tested fish fillets. The visualization procedure was programmed in the software Matlab version 2010a (The Mathworks Inc., MA, USA). Figure 1 shows the main steps of quantification analysis of E. coli loads and visualization of bacterial contamination in grass carp fish fillets by hyperspectral imaging technique.
Results and Discussion
Spectra of Fish Fillets
The average reflectance spectral information of the tested grass carp fillets with three different E. coli loads was obtained and is shown in Fig. 2. In this study, the measured E. coli loads of grass carp fillet varied from 4.11 to 10.02 log10 CFU/g, providing a reasonable contamination range of fish flesh from freshness to spoilage. As can be seen in Fig. 2, the spectral information obtained with the bacterial loads of 4.24 log10 CFU/g and 6.08 log10 CFU/g showed similar trends and minor fluctuations in the spectral range of 400–1000 nm. Compared with the former spectral information, the spectra obtained with the bacterial loads of 8.48 log10 CFU/g showed great difference on the spectral longitudinal shift. It has been demonstrated that the increase of E. coli loads to some extent affected the spectral information of fish flesh. This phenomenon was probably ascribed to variations of chemical components of fish flesh induced by bacterial activities during cold storage.
From another perspective, the overtone and combination vibrations of the molecular chemical bonds related to O–H, C–H, C–O, N–H and others are commonly used to elucidate the variations of the spectra. As shown in Fig. 2, a conspicuous and significant absorption peak was located at about 550 nm, possibly associated with the absorption of pigments such as astaxanthin and canthaxanthin in fish muscle (Kimiya et al. 2013). Another absorption peak located near to 970 nm was mainly related to the second overtone stretching of O–H by water (Cheng et al. 2014a).
PLSR Analysis Based on Full Wavelengths
The performance of PLSR models established in the calibration, cross-validation and prediction processes based on the full spectral range of 400–1000 nm was obtained as shown in Table 1. As can be seen in Table 1, regardless of the spectral data being preprocessed, the PLSR models showed a good performance with R 2 > 0.87 and RPD > 5.00. In addition, the PLSR model established using the spectral data preprocessed by MSC method presented a little better performance than that obtained using the raw spectra with an increase by 1.7, 0.6 and 0.9 % in R 2 C, R 2 CV and R 2 P and a decrease by 7.1, 2.6 and 1.5 % in RMSEs, respectively. Also, the value of RPD was increased from 5.38 to 5.47, which meant that using the preprocessing method of MSC to some extent improved the model performance. Figure 3 shows the prediction capability (R 2 P = 0.880, RMSEP = 0.262 log10 CFU/g and RPD = 5.47) of MSC-PLSR model between the actual measured and predicted values of E. coli loads, which demonstrated that the PLSR model was satisfactory for predicting E. coli loads. Also, it was confirmed that the HSI using full spectral range (400–1000 nm) was suitable for use in determining and quantifying the E. coli contamination of grass carp fillet during cold storage in a rapid and non-invasive way. Similarly, Tao et al. (2012) used the hyperspectral scattering technique in the spectral range of 400–1100 nm with Lorentzian distribution function for predicting E. coli contamination of pork meat, but poor validation result (R 2 CV = 0.707) was acquired. Afterwards, in order to improve the prediction capability, Tao and Peng (2014) used the same technique with Gompertz function for determining pork meat E. coli contamination, and an increase of R 2 CV by 0.174 was obtained. Another study was reported by Siripatrawan et al. (2011) who utilized the hyperspectral reflectance technique in the spectral range of 400–1000 nm with principal component analysis and artificial neural network analysis for rapid detection of E. coli contamination in packaged fresh spinach, and excellent performance was obtained with R 2 P = 0.97. Likewise, HSI technique with multivariate analysis has been successfully developed to evaluate microbial contaminations. For example, Feng and Sun (2013a) used the near-infrared HSI (910–1700 nm) for the determination of total viable count (TVC) in chicken breast fillets. The PLSR model established using the absorbance spectral data yielded a good performance with RPD of 2.60, R 2 CV of 0.865 and RMSECV of 0.57 log10 CFU/g. Later, Feng and Sun (2013b) used the same technique with PLSR analysis for the determination of Pseudomonas loads in chicken fillets, but a relatively poor prediction result was obtained with R 2 P of 0.656 and RMSEP of 0.80 log10 CFU/g, respectively. In another study, the potential of time series-hyperspectral imaging in visible and near infrared region (400–1700 nm) was used for the determination of surface total viable count (TVC) of salmon flesh during spoilage process. The least-squares support vector machines (LS-SVM) model showed an excellent performance with RPD of 5.09, R 2 P of 0.961 and RMSEP of 0.290 log10 CFU/g (Wu and Sun 2013). Compared with the current study, although all of these investigations have proved the potentiality of hyperspectral imaging technique for determining bacterial loads, the prediction capabilities are different mainly due to the used spectral range and multivariable analysis methods used. Therefore, in order to acquire a better prediction performance, more efforts should be made on applying different spectral region and developing effective analysis algorithms.
PLSR and MLR Analysis Based on Selected Wavelengths
On the basis of the full wavelengths in the spectral range of 400–1000 nm, although it has been proven the feasibility of the HSI system for potential determination of E. coli loads in fish flesh, it is a little difficult to develop the real-time and on-line detection system for such an application in the industry due to the huge data analysis required and computer hardware limitations. In order to solve the problems and increase the computing speed for optimizing the structure of imaging detection system and satisfy the real-time inspection, the WRC from PLSR model analysis in this study was used to select the optimal wavelengths for simplifying the original obtained models. As a result, six optimal wavelengths including 424, 451, 545, 567, 585, and 610 nm were obtained as shown in Fig. 4. These wavelengths recognized as the effective wavelengths were used to replace the full wavelengths for further predicting E. coli loads in fish flesh. It is interesting to find that these optimal wavelengths fell in the visible range, possibly due to the fact that the astaxanthin content and the special protein showed some influence on microbial activity. The performances of simplified models named WRC-PLSR and WRC-MLR for prediction of E. coli loads in grass carp fillet are shown in Table 1. It can be noticed that the WRC-PLSR model with four latent variables showed comparable and equivalent performance with the models developed using the full wavelengths. The number of variables was reduced from 381 to six variables, which helped to develop a simple PLSR model and saved the computing time of 98.4 %. It can thus be concluded that the HSI technique using the only selected six most informative wavelengths is also suitable for prediction and quantification of E. coli loads in grass carp flesh. Meanwhile, it was interesting to discover that the most sensitive and valuable wavelengths for indicating the E. coli contamination were concentrated in the visible region, which also facilitated the original HSI system into a new one in the spectral range of 400–700 nm. In addition, as illustrated in Table 1, compared with the WRC-PLSR model, the simplified WRC-MLR model demonstrated better effectiveness and robustness in predicting E. coli loads with the value of R 2 P of 0.870, RPD of 5.22 and RMSEP of 0.274 log10 CFU/g, which confirmed that MLR is more advantageous than PLSR when the number of variables was much less than the number of samples (6 vs. 100 or 50). On the basis of the better WRC-MLR model for prediction of E. coli loads in grass carp fillets, the quantitative regression equation for the detection of E. coli contamination was obtained and is presented below:
where X i nm is the reflectance spectral value at the wavelength of i nm and Y is the predicted E. coli loads. Although it has been proven that using six optimal wavelengths replacing the full wavelengths for developing a multispectral imaging system in industrial on-line application is potential and suitable, the reliability and applicability is still a little lower. Thus, more samples should be required in the calibration process to reduce the variability of E. coli measurement.
Visualization of E. coli Contamination
The great advantage of HSI against the conventional spectroscopy is its capability of visualizing the distribution map of the prediction values in a pixel-wise manner. Therefore, the final MLR model obtained from the effective wavelengths was used to transfer each pixel of the image to predict E. coli loads in all spots of the sample. After multiplying the regression coefficients of the MLR model by the spectrum of each pixel in the image, a prediction image was generated for showing the distribution of E. coli within the fish flesh. A linear colour scale was created with the different E. coli loads from small to large presented by different colours from blue to red. It means that pixels having similar spectral features offered the same predicted values of E. coli, which were then visualized in a similar colour in the image. Different colours in the final distribution map represented different values of E. coli in the image in proportion to the spectral differences of the corresponding pixels (ElMasry et al. 2012a). Figure 5 shows examples of distribution maps of E. coli contamination in some tested grass carp fillets with different E. coli loads. As can be seen in Fig. 5, the distribution maps indicated how the level of E. coli contamination varied from sample to sample and even from pixel to pixel within the same sample. Moreover, there was a general trend of increase of colour intensity from blue to red. As the E. coli loads increased, the colours of the images were gradually shifting from blue to reddish, which obviously reflected the growth of bacteria and the presentation of E. coli contamination status during the spoilage process. For example, Fig. 5a shows mostly the blue colour distribution with the low E. coli value (N = 4.115 log10 CFU/g) of fresh fish flesh, which indicated that the fish sample was at the early stage of spoilage process. The distributions of E. coli loads illustrated in Fig. 5b (N = 6.209 log10 CFU/g) and Fig. 5c (N = 8.185 log10 CFU/g) were fairly non-uniform along with different locations of fish fillet samples. This phenomenon was mainly associated with the uneven distribution of nutrients in fish flesh that promoted the growth of bacteria. Figure 5d (N = 9.788 log10 CFU/g) indicates the homogenous distribution of E. coli loads with the same red colour, implying that a great level of fish spoilage occurred and finally induced severe freshness loss in fish flesh. These phenomena are impossible to be observed by the naked eyes, thus, it is very useful and meaningful for the better understanding of the dynamic changes of E. coli loads in fish flesh during storage and is also helpful and important for the fishery industry to directly judge and evaluate the fish quality and safety and for further improving fish safety assurance.
Conclusions
This study was conducted to investigate the potentiality and suitability of visible and near infrared HSI technique (400–1000 nm) for quantifying and visualizing E. coli contamination in grass carp flesh during spoilage process at 4 °C. The results demonstrated that this emerging technique was feasible for rapid and non-invasive prediction and detection of E. coli loads. On the basis of the full wavelengths, the quantitative PLSR model established between the traditional measured E. coli loads and the spectral data preprocessed by MSC method showed a good performance with the value of RPD of 5.47, R 2 P of 0.880 and RMSEP of 0.262 log10 CFU/g. Six characteristic wavelengths including 424, 451, 545, 567, 585, and 610 nm were selected via the weighted regression coefficients from PLSR analysis. The simplified PLSR and MLR models established using the six selected wavelengths also presented an equivalent performance to the original models using the full wavelengths. Compared with the new PLSR model, the simplified MLR model yielded a better predictability with the value of RPD of 5.22, R 2 P of 0.870 and RMSEP of 0.274 log10 CFU/g, which was thus used to transfer each pixel of the image into its corresponding E. coli loads for visualizing E. coli contamination distribution using image processing algorithms. The distribution maps of bacterial loads were of great importance to provide more detailed information of postmortem spoilage development in grass carp flesh. In view of fish safety evaluation, these results verified this technique to be an admirable alternative to the time-consuming and conventional methods. As the first research on rapid and non-destructive prediction and quantification of E. coli loads in grass carp fish flesh, the whole results are potential and promising and will be helpful to make more efforts on the HSI technique for on-line applications and evaluation of bacterial contamination of grass carp fillet and other aquatic products during cold storage.
References
Arakawa, M., Yamashita, Y., & Funatsu, K. (2011). Genetic algorithm‐based wavelength selection method for spectral calibration. Journal of Chemometrics, 25(1), 10–19.
Balabin, R. M., & Smirnov, S. V. (2011). Variable selection in near-infrared spectroscopy: benchmarking of feature selection methods on biodiesel data. Analytica Ahimica Acta, 692(1), 63–72.
Barbin, D. F., ElMasry, G., Sun, D.-W., et al. (2012). Predicting quality and sensory attributes of pork using near-infrared hyperspectral imaging. Analytica Chimica Acta, 719, 30–42. Published: MAR 16 2012.
Cassin, M. H., Lammerding, A. M., Todd, E. C., Ross, W., & McColl, R. S. (1998). Quantitative risk assessment for Escherichia coli O157: H7 in ground beef hamburgers. International Journal of Food Microbiology, 41(1), 21–44.
Cheng, J.-H., & Sun, D.-W. (2014). Hyperspectral imaging as an effective tool for quality analysis and control of fish and other seafoods: current research and potential applications. Trends in Food Science & Technology, 37(2), 78–91.
Cheng, J., Sun, D.-W., Zeng, X.-A., & Liu, D. (2013). Recent advances in methods and techniques for freshness quality determination and evaluation of fish and fish fillets: a review. Critical Reviews in Food Science and Nutrition. doi:10.1080/10408398.2013.769934.
Cheng, J.-H., Qu, J.-H., Sun, D.-W., & Zeng, X.-A. (2014a). Visible/near-infrared hyperspectral imaging prediction of textural firmness of grass carp (Ctenopharyngodon idella) as affected by frozen storage. Food Research International, 56, 190–198.
Cheng, J.-H., Sun, D.-W., Pu, H., & Zeng, X.-A. (2014b). Comparison of visible and long-wave near-infrared hyperspectral imaging for colour measurement of grass carp (Ctenopharyngodon idella). Food and Bioprocess Technology, 7, 3109–3120.
Cheng, J.-H., Sun, D.-W., Zeng, X.-A., & Pu, H.-B. (2014c). Non-destructive and rapid determination of TVB-N content for freshness evaluation of grass carp (Ctenopharyngodon idella) by hyperspectral imaging. Innovative Food Science & Emerging Technologies, 21, 179–187.
Delgado, A. E., Sun, D.-W. (2002a). Desorption isotherms for cooked and cured beef and pork. Journal of Food Engineering, 51(2), 163-170
Delgado, A. E., Sun, D.-W. (2002b). Desorption isotherms and glass transition temperature for chicken meat. Journal of Food Engineering, 55(1), 1–8.
ElMasry, G., & Wold, J. P. (2008). High-speed assessment of fat and water content distribution in fish fillets using online imaging spectroscopy. Journal of Agricultural and Food Chemistry, 56(17), 7672–7677.
ElMasry, G., Sun, D.-W., & Allen, P. (2011a). Non-destructive determination of water-holding capacity in fresh beef by using NIR hyperspectral imaging. Food Research International, 44(9), 2624–2633. Published: NOV 2011.
ElMasry, G., Iqbal, A., Sun, D.-W., et al. (2011b). Quality classification of cooked, sliced turkey hams using NIR hyperspectral imaging system. Journal of Food Engineering, 103(3), 333–344. Published: APR 2011.
ElMasry, G., Barbin, D. F., Sun, D.-W., & Allen, P. (2012a). Meat quality evaluation by hyperspectral imaging technique: an overview. Critical Reviews in Food Science and Nutrition, 52(8), 689–711.
ElMasry, G., Sun, D.-W., & Allen, P. (2012b). Near-infrared hyperspectral imaging for predicting colour, pH and tenderness of fresh beef. Journal of Food Engineering, 110(1), 127–140. Published: MAY 2012.
Feng, Y.-Z., & Sun, D.-W. (2012). Application of hyperspectral imaging in food safety inspection and control: a review. Critical Reviews in Food Science and Nutrition, 52(11), 1039–1058.
Feng, Y.-Z., & Sun, D.-W. (2013a). Determination of total viable count (TVC) in chicken breast fillets by near-infrared hyperspectral imaging and spectroscopic transforms. Talanta, 105, 244–249.
Feng, Y.-Z., & Sun, D.-W. (2013b). Near-infrared hyperspectral imaging in tandem with partial least squares regression and genetic algorithm for non-destructive determination and visualization of Pseudomonas loads in chicken fillets. Talanta, 109, 74–83.
Ghasemi-Varnamkhasti, M., Mohtasebi, S. S., Rodriguez-Mendez, M. L., Gomes, A. A., Araújo, M. C. U., & Galvão, R. K. (2012). Screening analysis of beer ageing using near infrared spectroscopy and the successive projections algorithm for variable selection. Talanta, 89, 286–291.
Gowen, A., O’Donnell, C., Cullen, P., Downey, G., & Frias, J. (2007). Hyperspectral imaging—an emerging process analytical tool for food quality and safety control. Trends in Food Science & Technology, 18(12), 590–598.
Iqbal, S. S., Mayo, M. W., Bruno, J. G., Bronk, B. V., Batt, C. A., & Chambers, J. P. (2000). A review of molecular recognition technologies for detection of biological threat agents. Biosensors and Bioelectronics, 15(11), 549–578.
Jackman, P., Sun, D.-W., Du, C.-J., et al. (2008). Prediction of beef eating quality from colour, marbling and wavelet texture features. Meat Science, 80(4), 1273–1281. Published: DEC 2008.
Jackman, P., Sun, D.-W., Du, C.-J., et al. (2009). Prediction of beef eating qualities from colour, marbling and wavelet surface texture features using homogenous carcass treatment. Pattern Recognition, 42(5), 751–763. Published: MAY 2009.
Jin, J.-W., Chen, Z.-P., Li, L.-M., Steponavicius, R., Thennadil, S. N., Yang, J., & Yu, R.-Q. (2011). Quantitative spectroscopic analysis of heterogeneous mixtures: the correction of multiplicative effects caused by variations in physical properties of samples. Analytical Chemistry, 84(1), 320–326.
Kamruzzaman, M., ElMasry, G., Sun, D.-W., et al. (2011). Application of NIR hyperspectral imaging for discrimination of lamb muscles. Journal of Food Engineering, 104(3), 332–340. Published: JUN 2011.
Kamruzzaman, M., ElMasry, G., Sun, D.-W., et al. (2012). Prediction of some quality attributes of lamb meat using near-infrared hyperspectral imaging and multivariate analysis. Analytica Chimica Acta, 714, 57–67. Published: FEB 10 2012.
Kiani, H., Sun, D.-W. (2011). Water crystallization and its importance to freezing of foods: A review. Trends in Food Science & Technology, 22(8), 407–426.
Kimiya, T., Sivertsen, A. H., & Heia, K. (2013). VIS/NIR spectroscopy for non-destructive freshness assessment of Atlantic salmon (Salmo salar L.) fillets. Journal of Food Engineering, 116(3), 758–764.
Liu, D., Zeng, X.-A., & Sun, D.-W. (2013). Recent developments and applications of hyperspectral imaging for quality evaluation of agricultural products: a review. Critical Reviews in Food Science and Nutrition. doi:10.1080/10408398.2013.777020.
Liu, D., Sun, D.-W., & Zeng, X.-A. (2014). Recent advances in wavelength selection techniques for hyperspectral image processing in the food industry. Food and Bioprocess Technology, 7(2), 307–323.
Lorente, D., Aleixos, N., Gómez-Sanchis, J., Cubero, S., García-Navarrete, O. L., & Blasco, J. (2012). Recent advances and applications of hyperspectral imaging for fruit and vegetable quality assessment. Food and Bioprocess Technology, 5(4), 1121–1142.
McDonald, K., Sun, D.-W. (2001). The formation of pores and their effects in a cooked beef product on the efficiency of vacuum cooling. Journal of Food Engineering, 47(3), 175–183.
Mehmood, T., Liland, K. H., Snipen, L., & Sæbø, S. (2012). A review of variable selection methods in partial least squares regression. Chemometrics and Intelligent Laboratory Systems, 118, 62–69.
Menesatti, P., Costa, C., & Aguzzi, J. (2010). Quality evaluation of fish by hyperspectral imaging. In D.-W. Sun (Ed.), Hyperspectral imaging for food quality analysis and control (pp. 273–294). California: Academic Press/Elsevier.
Nugen, S. R., & Baeumner, A. (2008). Trends and opportunities in food pathogen detection. Analytical and Bioanalytical Chemistry, 391(2), 451–454.
Siripatrawan, U., Makino, Y., Kawagoe, Y., & Oshita, S. (2011). Rapid detection of Escherichia coli contamination in packaged fresh spinach using hyperspectral imaging. Talanta, 85(1), 276–281.
Sun, D.-W. (1997). Thermodynamic design data and optimum design maps for absorption refrigeration systems. Applied Thermal Engineering, 17(3), 211–221.
Sun, D.-W. (2004). Computer vision - An objective, rapid and non-contact quality evaluation tool for the food industry. Journal of Food Engineering, 61(1), 1–2. Published: JAN 2004.
Sun, D.-W. (2010). Hyperspectral imaging for food quality analysis and control: Academic Press/Elsevier, San Diego, California, USA, 496 pp., ISBN: 978-0-12-374753-2 (2010).
Sun, D.-W., Byrne, C. (1998). Selection of EMC/ERH isotherm equations for rapeseed. Journal of agricultural engineering research, 69(4), 307–315.
Sun, D.-W., Woods, J. L. (1997). Simulation of the heat and moisture transfer process during drying in deep grain beds. Drying Technology, 15(10), 2479–2508.
Sun, D.-W., Eames, I.W., Aphornratana, S. (1996). Evaluation of a novel combined ejector-absorption refrigeration cycle .1. Computer simulation. International Journal of Refrigeration-Revue Internationale Du Froid, 19(3), 172–180.
Tao, F., & Peng, Y. (2014). A method for nondestructive prediction of pork meat quality and safety attributes by hyperspectral imaging technique. Journal of Food Engineering, 2014(126), 98–106.
Tao, F., Peng, Y., Li, Y., Chao, K., & Dhakal, S. (2012). Simultaneous determination of tenderness and Escherichia coli contamination of pork using hyperspectral scattering technique. Meat Science, 90(3), 851–857.
Valous, N. A., Mendoza, F., Sun, D.-W., et al. (2009). Colour calibration of a laboratory computer vision system for quality evaluation of pre-sliced hams. Meat Science, 81(1), 132–141. Published: JAN. 2009.
Wang, H. H., & Sun, D.-W. (2002). Melting characteristics of cheese: analysis of effect of cheese dimensions using computer vision techniques. Journal of Food Engineering, 52(3), 279–284. Published: MAY 2002.
Williams, P. C. (2001). Implementation of near-infrared technology. Near-infrared Technology in the Agricultural and Food Industries, 2, 143.
Windham, W. W., Yoon, S.-C., Ladely, S. R., Heitschmidt, J. W., Lawrence, K. C., Park, B., Narrang, N., & Cray, W. C. (2012). The effect of regions of interest and spectral pre-processing on the detection of non-0157 Shiga-toxin producing Escherichia coli serogroups on agar media by hyperspectral imaging. Journal of Near Infrared Spectroscopy, 20(5), 547–558.
Windham, W. R., Yoon, S.-C., Ladely, S. R., Haley, J. A., Heitschmidt, J. W., Lawrence, K. C., Park, B., Narrang, N., & Cray, W. C. (2013). Detection by hyperspectral imaging of Shiga toxin-producing Escherichia coli serogroups O26, O45, O103, O111, O121, and O145 on rainbow agar. Journal of Food Protection, 76(7), 1129–1136.
Wu, D., & Sun, D.-W. (2013). Potential of time series-hyperspectral imaging (TS-HSI) for non-invasive determination of microbial spoilage of salmon flesh. Talanta, 111, 39–46.
Wu, D., Shi, H., Wang, S., He, Y., Bao, Y., & Liu, K. (2012). Rapid prediction of moisture content of dehydrated prawns using online hyperspectral imaging system. Analytica Ahimica Acta, 726, 57–66.
Xu, S. Y., Chen, X. F., Sun, D.-W. (2001). Preservation of kiwifruit coated with an edible film at ambient temperature. Journal of Food Engineering, 50(4), 211–216.
Yeni, F., Acar, S., Polat, Ö. G., Soyer, Y., & Alpas, H. (2014). Rapid and standardized methods for detection of foodborne pathogens and mycotoxins on fresh produce. Food Control, 40, 359–367.
Yoon, S. C., Windham, W. R., Ladely, S., Heitschmidt, G. W., Lawrence, K. C., Park, B., Narang, N., & Cray, W. C. (2013). Differentiation of big-six non-O157 Shiga-toxin producing Escherichia coli (STEC) on spread plates of mixed cultures using hyperspectral imaging. Journal of Food Measurement and Characterization, 7(2), 47–59.
Acknowledgments
The authors were grateful to the Guangdong Province Government (China) for its support through the program of “Leading Talent of Guangdong Province (Da-Wen Sun)”. This research was also supported by the National Key Technologies R&D Program (2014BAD08B09) and The International S&T Cooperation Projects of Guangdong Province (2013B051000010).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cheng, JH., Sun, DW. Rapid Quantification Analysis and Visualization of Escherichia coli Loads in Grass Carp Fish Flesh by Hyperspectral Imaging Method. Food Bioprocess Technol 8, 951–959 (2015). https://doi.org/10.1007/s11947-014-1457-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11947-014-1457-9