Abstract
Plant cell walls primarily comprise lignin, which performs functions of mechanical support, water transport, and stress responses. Lignin biosynthesis pathway proceeds through metabolic grid featuring complexity and diversity in enzymatic reaction. Cinnamate-4-hydroxylase (C4H, EC 1.14.13.11) is the gene encoding enzyme that catalyzes the second step of phenylpropanoid pathway responsible for biosynthesis of lignin. A full-length cDNA of C4H (designated as GbC4H), which spanned 1816-bp with a 1518-bp open reading frame encoding a 505-amino-acid protein, was cloned from Ginkgo biloba. A GbC4H genomic DNA fragment, spanning 3249-bp, was cloned and found to contain two exons and one intron. GbC4H protein showed high similarities with other plant C4Hs to include conserved domains of cytochrome P450 family. GT-1, W-box, and Myb/Myc recognition sites involved in stress response were detected in a 1265-bp upstream promoter region of GbC4H. Phylogenetic analysis suggested the common evolutionary ancestor shared by plant C4Hs including the gymnosperm enzyme. pET-28a-GbC4H plasmid was constructed and expressed in Escherichia coli strain BL21. Enzymatic assay revealed that recombinant GbC4H protein catalyzes conversion of trans-cinnamic acid to p-coumaric acid. Expression analyses in different organs showed high expression of GbC4H in stems and roots, whereas low expressions was found in fruits, carpopodium, and petioles. Further analysis indicated linear correlation of lignin contents with transcript levels of GbC4H among different tissues. GbC4H transcription was increased by treatments with UV-B, cold, salicylic acid, and abscisic acid, indicating the possible role of GbC4H in response to stresses and hormonal signal. Understanding of GbC4H function could benefit molecular breeding and reinforcement of defense mechanisms in Ginkgo.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
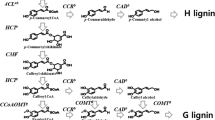
Lignin is the most complicated polyphenolic compounds on earth, and the second largest biopolymer of vascular plants (Sarkanen and Ludwig 1971; Boerjan et al. 2003; Boudet et al. 2003; Himmel 2008). Primary function of lignin is to provide structure to cell walls and barrier to defense stresses (Ferrer et al. 2008; Weng and Chapple 2010; Sykes et al. 2015), executing fundamental biological functions, such as mechanical support, unpermeability, and resistance to biodegradation, in addition to crucial roles in defense mechanisms (Bhuiyan et al. 2009; Tao et al. 2009; Xu et al. 2009). Lignin biosynthesis is regulated by enzymes and their corresponding genes that participate in formation, transportation, and polymerization of the precursor compound (Anterola and Lewis 2002). Lignin is synthesized from three hydroxycinnamyl alcohols through combined free radical coupling reactions (Ralph et al. 2004; Zeng et al. 2014). These hydroxycinnamyl alcohols are p-coumaryl, coniferyl, and sinapyl alcohols, which, respectively, produce p-hydroxyphenyl (H), guaiacyl (G), and syringyl (S) type of lignin (Sarkanen and Ludwig 1971; Terashima et al. 2009). Lignin biopolymer in dicots mainly contain G and S units with minor amounts of H units, whereas the main components of monocot lignin are G and S units. Gymnosperm lignin is almost entirely consisted of G units, with trace amounts of H units (Gross 1981; Boerjan et al. 2003). The lignin contents and composition change widely within and between species. Furthermore, organs from the same plant in different developmental stages and under different conditions also exhibit the difference (Plomion et al. 2001; Bhuiyan et al. 2009).
The number of sequenced plant genomes has increased extensively in the past few years, providing opportunities to trace evolutionary origins of biosynthetic and signaling pathways of plant lignins. Lignin biosynthesis process and its regulatory mechanisms have been investigated in depth (Chiang 2006; Vanholme et al. 2008; Weng et al. 2008). Phenylpropanoid pathway is the primary metabolic process responsible for the synthesis of a number of secondary metabolites (Vogt 2010; Tohge et al. 2013). The metabolites of phenylpropanoid origin include phenolic compounds, flavonoids, anthocyanins, phytoalexins, and lignin (Dixon and Paiva 1995; Weisshaar and Jenkins 1998). Lignin biosynthesis in plants begins with three consecutive reactions usually considered as the general phenylpropanoid pathway (Dixon et al. 1996; Schilmiller et al. 2009). The core three reactions are each catalyzed by phenylalanine ammonia lyase (PAL), cinnamate-4-hydroxylase (C4H), and 4-coumarate coenzyme A ligase (4CL) (Boerjan et al. 2003; Vogt 2010; Naoumkina et al. 2010). C4H operating in the second step of phenylpropanoid pathway catalyzes conversion of trans-cinnamic acid to p-coumaric acid, which is in turn derived from phenylalanine by PAL (Hahlbrock and Scheel 1989; Schilmiller et al. 2009; Tohge et al. 2013). As a common cytochrome P450 monooxygenase-type hydroxylase in higher plants, C4H is a member of the CYP73 family often involved in biosynthesis of diverse metabolites (Kochs and Grisebach 1989; Chapple 1998; Achnine et al. 2004; Singh et al. 2009). Being a P450 protein, C4H catalyzes irreversible reactions (Ehlting et al. 2006). C4H activity is regulated by several factors and positively correlates with lignin metabolism (Tabata 1996). C4H also controls the carbon fluxes leading to many phytoalexins, which are synthesized by plants on attack by bacterium, virus or other microorganism (Teutsch et al. 1993; Ro and Douglas 2004). Therefore, molecular cloning and functional characterization of C4H gene could be a crucial step to understand lignin pathway of a particular plant in question. Various C4H genes have been cloned from plants: Helianthus tuberosus (Werck-Reichhart et al. 1993), Catharanthus roseus (Hotze et al. 1995), Arabidopsis thaliana (Mizutani et al. 1997), P. trichocarpa × P. deltoids (Ro et al. 2001), Oryza sativa (Yang et al. 2005), Populus tremuloides (Lu et al. 2006), Salvia miltiorrhiza (Huang et al. 2008), Brassica napus (Chen et al. 2007), Parthenocissus henryana (Liu et al. 2009), Scutellaria baicalensis (Xu et al. 2010), Camellia sinensis (Rani et al. 2012), Leucaena leucocephala (Kumar et al. 2013), and Ornithogalum saundersiae (Kong et al. 2014). However, only limited information has been reported about the Ginkgo C4H gene in public databases.
Plants have been developing a broad range of complex defense mechanisms to cope with pathogenic infections, because they are confined to the place where they grow. Ginkgo biloba is an important ornamental and, more importantly, medicinal plant. Extracts of Ginkgo leaves contain bioactive flavonoids and terpene lactones, which have been used to improve blood circulation, to treat cardiovascular diseases, and to protect liver function (Smith and Luo 2004; van Beek 2002). Both flavonoid and lignin biosyntheses share core phenylpropanoid metabolism. Cheng et al. (2013a, b) showed that lignin synthesis can affect the accumulation of Ginkgo flavonoids. Therefore, cloning and characterization of the genes involved in phenylpropanoid pathway in Ginkgo can extend our knowledge in lignin biosynthesis of the plant. Deep understanding of the pathway-related enzymes is also necessary to identify the goal of biotechnological operation for modulating the lignin accumulation.
In this study, a full-length cDNA of C4H gene (designated as GbC4H) was isolated from G. biloba. To understand the structure and the regulation of GbC4H gene in lignin biosynthesis, its coding region and promoter sequences were cloned. The combination of biochemical and gene expression analyses suggested that GbC4H is a key enzyme related to lignification and defense processes in Ginkgo. We also correlated the locus of gene expression with distribution of lignin among plant tissues. The full-length coding region of GbC4H was sub-cloned into E. coli expression vector pET-28a and enzyme activity was determined to confirm the function of the recombinant GbC4H protein.
Materials and methods
Plant materials and treatments
For this study, biennial, triennial, 4-year, and 5-year Ginkgo seedlings at six-leaf stage were raised in plastic pots (45 cm × 35 cm, peat:humus soil:perlite (1:1:1) mixture) and housed in a plant growth chamber (temperature, 25 °C ± 1 °C; light intensity, 300 µmol m−2 s−2, 16 h/8 h light/dark cycle; relative humidity, 70%), in Yangtze University, Jingzhou, China (30.35°N, 112.14°E). For gene cloning and the determination of transcripts among organs, young leaves, stems, roots, fruits, carpopodium, and petiole were sampled (Xu et al. 2008). The collected samples were immediately frozen in liquid nitrogen and stored at − 80 °C until use. All the samples were harvested in early April.
To investigate expression patterns of GbC4H in different stress (UV-B and cold) and hormonal treatments [salicylic acid (SA) and abscisic acid (ABA)], a total of 240 3-year-old Ginkgo seedlings at six-leaf stage were planted in 48 plastic pots (five seedlings in one plastic pot). Among them, 210 seedlings (42 plastic pots) with consistent growth and size were selected and divided uniformly into seven groups (six pots in one group), with four groups being used for different treatments and another three groups as control (SA and ABA treatments used same control). For SA and ABA treatments, plants were mist-sprayed with SA (500 µM) or ABA (50 µM) solution to both sides of leaves until liquids dripped. Double-distilled water (ddH2O) served as control. SA-treated plants were collected at 0, 2, 4, 8, 24, and 48 h, whereas ABA-treated ones were collected at 0, 4, 8, 16, 24, and 48 h. For UV treatment, Ginkgo seedlings were placed in a closed chamber with UV-B illumination at 1500 µJ/m2, whereas control seedlings were left in the dark. Then, the leaves were harvested at 0, 1, 2, 4, 8, and 24 h. For cold treatment, seedlings were placed in the growth room at 4 °C, whereas control seedlings were treated at 20 °C. Treated leaves were harvested at 0, 1, 2, 4, 8, and 24 h. Preliminary experiment demonstrated no visible damage on plants with the above-mentioned treatments. Two uppermost leaves from the stem tip were taken from each seedling at prescribed time point to make one biological sample. One data point was obtained from five such biological replicates.
Identification of cDNA and genomic DNA sequences coding for GbC4H by polymerase chain reaction (PCR)
First-strand cDNA was synthesized using PrimeScript™ 1st Strand cDNA Synthesis Kit (TaKaRa, Dalian, China) using 1 µg RNA as template according to manufacturer’s instruction. Nucleotide sequence of annotated GbC4H candidate was downloaded from the National Center for Biotechnology Information (NCBI) database (GenBank accession no. AY748324). A pair of specific primers, GbC4H-F and GbC4H-R (Table 1), were designed for PCR amplification to obtain full-length cDNA of GbC4H. Amplified PCR products were purified and ligated into pMD18-T vector (TaKaRa, Dalian, China), and then cloned into the E. coli strain DH5α followed. To detect any introns within GbC4H, PCR amplification was carried out using the same reaction system as the full-length cDNA clone, except that the template was 1.5 µg of total genomic DNA. PCR products were purified, ligated into pMD18-T vectors, and sequenced.
Bioinformatics analysis
Sequence alignments and homology sequence searches were performed on the NCBI BLAST webpage (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Molecular mass and isoelectric point of deduced GbC4H protein were predicted using ExPASy Proteomics Server (http://www.expasy.ch/). Multiple sequence alignments were performed using Vector NTI 11.5 program. Phylogenetic relationship among sequences was detected with CLUSTAL X2 and MEGA 6.0 software programs. The neighbor-joining (NJ) method was used to construct a guided tree, which was supported by bootstrapping based on 1000 replicates.
Cloning of promoter of GbC4H
Promoter region fragment of Ginkgo GbC4H gene was isolated using the TaKaRa Genome Walking Kit (Dalian, China). Two specific primers, GbC4H-1 and GbC4H-2 (Table 1), and two degenerated primers, AP1 and AP2, were designed for nested PCR amplification using the kit. PCR amplification was performed at a final volume of 50 µL containing 1.0 µL DNA, 1.0 µL of 10 mM dNTP mixture, 5.0 µL of 10× Advantage 2 PCR Buffer, 1.0 µL of 10 µM degenerate primer, 1.0 µL of 10 µM specific primer, 1.0 µL Advantage 2 Polymerase Mix (50×), and 40.0 µL ddH2O. Both first and nested PCR reactions were carried out under the same amplification condition: 7 cycles at 94 °C for 25 s and 72 °C for 3 min, with 32 cycles at 94 °C for 25 s and 67 °C for 3 min, and a final reaction at 67 °C for 7 min. First PCR products were diluted tenfold and used as templates in nested PCR amplification. Nested PCR products were analyzed on 1% agarose gel. Major DNA bands were isolated from the gel using DNA Gel Extraction Kit (TaKaRa, Dalian, China), ligated into pMD18-T vectors, and sequenced. Putative cis-elements in the promoter of GbC4H gene were analyzed using online software PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html) and PLACE (http://www.dna.affrc.go.jp/PLACE/signalup.html).
Quantitative real-time PCR (qRT-PCR) analysis
qRT-PCR was performed to detect the GbC4H transcription levels in different Ginkgo organs and treated leaf samples. A total of 500 ng RNA in a 10 µL reaction was used with TaKaRa PrimeScript™ RT Reagent Kit (Perfect Real Time). cDNA was diluted to 100 µL and stored at − 20 °C. Gene quantification was carried out using SYBR® Premix Ex Taq™ Kit (TaKaRa, Dalian, China) according to manufacturer’s instructions. Each 25 µL reaction was run in triplicate. The above mixture consisted of 2 µL diluted cDNA, 0.5 µL of each primer, and 0.5 µL of 1× SYBR® Premix Ex Taq™ II. PCR amplification was performed as follows: 95 °C for 30 s, 40 cycles of 95 °C for 5 s and 60 °C for 30 s. The housekeeping gene Gb18S (GenBank accession no. D16448) was used to evaluate qRT-PCR assays as described by Xu et al. (2014). Fluorescence data were collected after each extension step. Reading temperature at 82 °C was determined by melting curve analysis. Relative expression level was calculated by a formula using the relative \(2^{{ - \Delta \Delta C_{\text{t}} }}\) method (Schmittgen and Livak 2008). To compare expression levels of GbC4H among different organs or treatments, relative expression of GbC4H was obtained by correcting with Gb18S level. All experiments were conducted in quintuplicate using five biological replicates. All primers used in this study are listed in Table 1.
Heterologous expression of GbC4H in E. coli
To construct a GbC4H prokaryotic expression vector, a pair of primers (GbC4H-EF and GbC4H-ER) with a restriction enzyme site (EcoRI) was designed, and used for amplification of the coding region. After digestion with EcoRI, the full-length coding sequence was sub-cloned into pET-28a vector. Recombinant vectors (pET28a-GbC4H) were then introduced into E. coli BL21 (DE3) and confirmed by PCR and sequencing. Transformants harboring recombinant plasmid were cultured at 37 °C in LB liquid medium containing kanamycin (100 mg/L) until optical density at 600 nm reached 0.6, induced with 0.2 mM isopropyl-β-d-thiogalactoside (IPTG), and incubated overnight. Bacterial cells were collected by centrifugation at 5000 rpm for 1 min, and pellets were resuspended in extraction buffer (100 mM Tris–HCl, pH 7.5; 10 mM MgCl2; 2% glycerol; and 1 mM) and disrupted by sonication. Then, suspension was boiled and iced at − 20 °C for 5 min, and lysate was centrifuged at 12,000 rpm for 15 min at 4 °C. Subsequently, 10 µL supernatant was analyzed by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) with 12% (v/v) polyacrylamide separation gel and 5% (v/v) stacking gel. Proteins were visualized by Coomassie Brilliant Blue G-250 staining. Recombinant GbC4H proteins extracted from induced cells were purified by nickel-CL agarose affinity chromatography (Bangalore Genei, India) and used for enzymatic assays.
Enzymatic assay of GbC4H
GbC4H activity was determined according to Salvador et al. (2013) with minor modifications. A mixture of 50 mM phosphate buffer (pH 7.6), 500 µM trans-cinnamic acid, 300 µM NADPH was added with an appropriate amount of enzyme extracts in a final volume of 200 µL. The mixture without the substrate was used as control. Reaction mixture was incubated at 35 °C on a gyratory shaker at 200 rpm for 40 min. The reaction was stopped by addition of 75 µL of 5 M HCl. Denatured proteins were pelleted by centrifuging at 17,000g for 3 min. After the removal of denatured proteins, p-coumaric acid in solution was detected by high-performance liquid chromatography (HPLC) equipped with a ZORBAX Eclipse Plus C18 column (250 mm × 4.6 mm, Agilent Technologies, Santa Clara, CA, USA) at 1 mL min−1, with column temperature set at 35 °C. trans-Cinnamic acid and p-coumaric acid were identified by comparing its retention time with the standard compounds.
Lignin determination and histochemical lignin staining
Lignin contents were estimated by the Klason method with slight modification (Kirk and Obst 1988; Kumar et al. 2012). Briefly, tissues were chopped and air-dried. Air-dried organs were ground to fine powder and oven-dried at 105 °C to constant weight. About 200 mg of accurately weighed tissue powder was continuously extracted in acetone:water (10:1, v/v) for 48 h at 55 °C. Cell wall residue (CWR) was obtained by drying tissue powders to constant weight at 105 °C. CWRs were acid-hydrolyzed (72% H2SO4) for 3 h at 25 °C, diluted with 190 mL H2O, and then autoclaved for 1 h. After cooling, samples were filtered and washed with hot water to remove acids and dried to constant weight at 105 °C to obtain acid-insoluble or Klason lignin (KL), which was expressed as a percentage of CWR. Filtrate was diluted to 500 mL and used for spectrophotometric determination of acid-soluble lignin (ASL) at 205 nm (Yeh et al. 2005). Lignin contents were determined from three biological replicates and estimated as weight percent of dry extract-free tissues.
For lignin staining, hand-cut sections were made from various organs of Ginkgo plant. Stems and taproots about 3 cm from collar were used. Lignin histochemistry was observed according to Pomar et al. (2004). Hand-cut sections of different tissues were placed in a phloroglucinol solution (2% in 95% ethanol) for 10 min, and then treated with 18% HCl for 5 min. The stained sections were observed on a microscope (Leica 165C). Image was taken with Canon EOS500D.
Results
Isolation and characterization of cDNA and genomic DNA of GbC4H
GbC4H was amplified using specific primers designed from sequence information available at NCBI database. Full-length GbC4H cDNA was 1816-bp long containing 1518-bp ORF, encoding a 505-amino acid protein. The ORF was flanked by a 138-bp 5′ noncoding sequence and 160-bp 3′ untranslated region with a canonical polyadenylation signal sequence (AATAAA) and a poly(A) tail (GenBank Accession No. MF680550, Fig. S1). The theoretical molecular weight and isoelectric point of the deduced GbC4H protein were 57.9 kDa and 8.47, respectively. Full-length genomic DNA of GbC4H gene, consisted of 3249-bp (GenBank Accession No. MF680551, Fig. S1), had full consistency with cDNA sequence. Exons 1 (788-bp) and 2 (730-bp) were separated by a 1433-bp long intron 1 (Fig. 1).
Multiple alignment of GbC4H protein with other C4H homologues was shown (Fig. 2). Amino acid sequence of GbC4H showed 89% identity with that of Pinus taeda; 83% with those of Nelumbo nucifera, Jatropha curcas, and Camellia sinensis; and 82% with those of Camptotheca acuminata, Gossypium hirsutum, Theobroma cacao, and Humulus lupulus. Database search (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) of the conserved domain indicated that GbC4H belongs to cytochrome P450 superfamily. A proline-rich region was present in the N-terminal, and a conserved heme-binding motif was also discovered near the C-terminal. Some conserved general topography and structural folds, including I helix, K helix, and K′ helix structures, were found (Fig. 2). Three C4H-conserved residues, N302, I371, and K484, were also identified in deduced GbC4H protein sequence, which are related to substrate discrimination and direction (Schoch et al. 2003, Fig. 2).
Alignments of amino acid sequences of GbC4H and other plant C4Hs. Amino acid sequences used in the analysis are from Pinus taeda (PtC4H, AAD23378.1), Camptotheca acuminate (CaC4H, ANR76395.1), Gossypium hirsutum (GhC4H, XP_016677259.1), Theobroma cacao (TcC4H, EOY20175.1), Nelumbo nucifera (NnC4H, XP_010253046.1), Jatropha curcas (JcC4H, XP_012078176.1), Camellia sinensis (CsC4H, ALD83478.1), and Humulus lupulus (HlC4H, ACM69364.1). Identical amino acid residues in this alignment are shaded in black. The putative conserved Proline-rich region, heme-binding motif, I helix, K helix, and K′ helix are indicated by colorful boxes. The conserved residues are indicated by asterisks and red color
To investigate the evolutionary relationship among different plant C4Hs, we obtained plant C4Hs and putative C4H sequences from NCBI database. Subsequently, a phylogenetic tree was conducted with full-length amino acid sequences. The tree clearly showed evolutionary branching of gymnosperm from monocots and dicots, and of 12 families in angiosperm clade (Fig. 3). GbC4H belongs to gymnosperms and, bears a close relationship to P. taeda C4H. In addition to that, C4Hs from same plant family showed normally higher sequence similarity, and closest evolutionary relationship.
Analysis of GbC4H promoter region
To further study the function of GbC4H, we isolated a 1265-bp fragment upstream of start codon (MF680551), including the 139-bp 5´UTR which corresponds to the putative promoter region of GbC4H, via genome walking methodology. The promoter sequence features a typically high A + T content (65%) generally discovered in other plant promoters. Further analysis on putative cis-elements was carried out using PlantCARE and PLACE tools (Fig. 4). The analysis revealed that putative transcription start site (TSS) was located 139-bp upstream of the ATG and was defined as + 1. Table 2S summarizes functions of in silico-identified putative cis-elements. A typical TATA box was found at position 30 (−) relative to TSS. Another typical cis-acting element known as CAAT box was also identified at positions 91 (+), 49 (−), 183 (−), 322 (−), 350 (−), 411 (−), 468 (−), 475 (−), 532 (−), 558 (−), 664 (−), 970 (−), and 1077 (−).
The putative cis-acting elements presenting in the promoter region of GbC4H gene. Nucleotides were numbered relative to A (transcription start site, + 1). The putative TATA-box, CAAT-box, I-box, GATA-box, and CAAT-boxes were shown and labeled in rectangular boxes with blue or pink. The transcription start site was marked by green
Several stress-related cis-acting elements were also found. Two Myb-binding sites [positions 317 (−) and 830 (−)] and one Myc recognition site [position 826 (−)] were found. All Myb and Myc protein family members play key roles in plant responses to stresses, such as pathogen, cold, and drought. Three types of cis-acting elements involved in light responsiveness were also found; these elements included three GATA boxes [positions 389 (−), 899 (−), and 1101 (−)] (Reyes et al. 2004), two GT-1 motifs [843 (−), 855 (−)], and one I box [1100 (−)]. Several important phytohormone responsive elements were predicted within the GbC4H promoter region. These hormone responsive elements, including the W-box involved in SA responsiveness (Redman et al. 2002), TGAC motif in response to gibberellic acid, and Myc-binding site, are not only involved in light responsiveness but also in response to ABA. However, specific functions of these putative cis-elements in Ginkgo remain undetermined calling for further study.
In vitro activity of GbC4H
To explore characteristics of GbC4H protein, GbC4H coding sequence (1515-bp) was expressed in E. coli cells. IPTG treatment induced the expected protein band of about 58 kDa in SDS PAGE of total cellular protein (Fig. 5, lane 2). This size was consistent with GbC4H protein as predicted (57.9 kDa). This band was absent in negative control (Fig. 5, lane 1). Purified recombinant proteins again exhibited the predicted size (Fig. 5, lane 3). Purified GbC4H protein was thus obtained.
SDS –PAGE analysis of GbC4H expressed in E. coli BL21 (DE3). After IPTG induction, E. coli BL21 cells containing pET28a-GbC4H were grown at 30 °C for 3 h. M, molecular marker; lane 1, proteins of total cells without IPTG induction; lane 2, proteins of total cells with 0.2 mM IPTG induction for 4 h; lane 3, purified recombinant GbC4H protein with Nickel-CL agarose affinity chromatography and used for enzyme activity assay
To investigate whether GbC4H possessed catalytic activity, trans-cinnamic acid was used as substrate in the reaction with purified GbC4H protein. HPLC analysis of the reaction mixture showed a new peak at 19.15 min corresponding to standard p-coumaric acid (Fig. 6a, b), whereas no such peak was observed in the negative controls (Fig. 6c). The above result demonstrated that GbC4H catalyzed conversion of trans-cinnamic acid into p-coumaric acid.
HPLC profiles of GbC4H reaction product. The activity of recombinant GbC4H were assayed using trans-cinnamic acid as substrate. The recombinant protein of empty vector was assayed as control. a Authentic standards of p-coumaric acid and trans-cinnamic acid. b Reaction product of recombinant GbC4H protein. c Empty vector control
Lignin contents in Ginkgo organs
Fresh stem, root, fruit, carpopodium, and petiole sections from Ginkgo plants were stained using phloroglucinol, which typically stains lignin with pink color (Fig. 7a). Stem and root sections were stained pink, whereas fruit, carpopodium, and petiole sections were stained with brown color. Xylem and periderm were stained in stems and 2-year-old taproot (A–E) and sclerenchymatous and epidermic cells were stained in 5-year-old taproot section. In the case of fruit, only xylem was stained, and xylem and sclerenchymatous cells were stained in carpopodium. Petiole was stained in xylem and sclerenchymatous and epidermic cells (Fig. 7a). Figure 7b indicated abundant lignin contents in stems and roots, whereas low lignin contents in fruit, carpopodium, and petiole as indicated by the staining.
The section stainings, lignin contents, and expression levels of GbC4H in different ginkgo organs. a Lignin staining. A biennial stem; B triennial stem; C 4-year-old stem; D 5-year-old stem; E taproot from 2-year-old seedling; F taproot from 5-year-old seedling; G the horizontal cutting of fruit; H carpopodium; I petiole. The arrow indicates histo-chemical lignin staining. b Lignin content of various tissue samples as determined by modified Kalson method. c The relative levels of GbC4H mRNA copy number measured by qRT-PCR. The labels, A through I, are same as above. Data are mean values of triplicate tests ± SD of three biological replicates. Different lower case letter indicates values are significantly different at p < 0.05
Transcript analysis of the GbC4H gene in different organs
To investigate whether GbC4H is involved in lignin biosynthesis in Ginkgo, transcript profile of GbC4H gene was examined by RT-PCR. We observed that GbC4H transcription level varied depending on organs and their developmental stages (Fig. 7c). Among the tissues, GbC4H was most predominantly transcribed in stems and roots, followed by fruits, carpopodium, and petiole. In addition, the highest transcription levels were detected in 5-year-old stems compared to those in 2-year-old stems. Linear regression analysis indicated that lignin content correlated linearly with transcription level of GbC4H. Concomitant probability value was p = 0.0017, and correlation coefficient was R 2 = 0.7762.
GbC4H transcription in leaves in response to various abiotic stresses and hormonal treatments
GbC4H transcription in response to various stress and hormonal treatment was assessed in 3-year-old Ginkgo seedling. GbC4H transcripts significantly changed after all abiotic treatments such as cold and UV-B (Fig. 8). Treatment of SA at 500 µM induced GbC4H transcription at 2 h after treatment, with the highest induction being attained at 8 h, followed by a rapid decline (Fig. 8a). GbC4H transcription was also induced by ABA treatment and peaked at 24 h before decreasing gradually (Fig. 8b). However, in the case of cold treatment, onset of GbC4H transcription was observed in 1 h after treatment and reached maximum at 24 h (Fig. 8c). Finally, UV-B irradiation caused rather delayed response with discernable increase in GbC4H transcription being observed at 4 h after treatment. The level of transcript gradually increased until 24 h after UV-B treatment (Fig. 8d).
Relative quantities of the GbC4H mRNA at various time points post-treatment with SA (a), ABA (b), cold (c) and UV-B (d). Each sample was individually assayed in quintuplicate. Data are mean values of quintuplicate tests ± SD of five biological replicates. Means with different letters from each time of post-treatment are significantly different at p < 0.05
Discussion
GbC4H gene is a member of C4H gene family
In the present study, full-length cDNA and genomic DNA of GbC4H were cloned from Ginkgo. Genomic DNA of GbC4H was 3249-bp long with a 1518-bp ORF, which was interrupted by two exons and one intron (Supplementary Fig. S1). Multiple alignments showed that GbC4H belongs to the cytochrome P450 superfamily, which contains heme-thiolate proteins involved in oxidative degradation of xenobiotic chemicals such as environmental toxins and mutagens. Although the similarity of amino acid sequences among protein members of the superfamily is a little low, their common morphology and structural fold are known to be highly conserved. GbC4H contained a proline-rich region (PPGPFALP) in the N-terminal, which is consistent with the consensus sequence ((P/I)PGPX(G/P)XP (Szczesnaskorupa et al. 1993; Yamazaki et al. 1993) that is suggested to play a central role in cytochrome P450 superfamily. An absolutely conserved heme-binding motif PPGVGRRSCPG corresponding to consensus sequence PFGXGRRXCXG (Durst and Nelson 1995) was identified near the C-terminal. Conserved general topography and structural folds, including I helix (AAIETT), K helix (ETLR), and K′ helix (AWWLANN) (Kong et al. 2014), were also found in GbC4H (Fig. 2). Three conserved residues, N302, I371, and K484, which are important in substrate identification and direction (Schoch et al. 2003) were also found (Fig. 2). N302 and I371 are suggested to be pivotal determining factors of substrate recognition and orientation, whereas K484 was found not to participate in the initial substrate binding, but plays an important role in catalysis through contributing to substrate re-orientation during the catalytic reaction (Schoch et al. 2003).
Sequence comparison among C4H members confirmed previous classification of plant C4Hs into angiosperm classes I, IIA and B, and gymnosperm groups (Nedelkina et al. 1999; Schoch et al. 2003). As shown in Fig. 3, GbC4H belongs to gymnosperm groups, and has a closest relationship with PtC4H. Phylogenetic tree analysis showed sequence similarities within each plant family from different plant species are commonly higher than those between different family C4Hs. Taken together, these results indicated that much of plant P450 diversity existed prior to the divergence of dicot and monocot plants, estimated to have occurred 200 million years ago; and all of plant P450 family members existed before gymnosperms branched from angiosperms, estimated to have occurred 360 million years ago (Nelson et al. 2004).
Properties of recombinant GbC4H protein
GbC4H protein shared high sequence similarity to C4H from P. taeda. C4H catalyzes first oxygenation step of common phenylpropanoid pathway, conversion of trans-cinnamic acid to p-coumaric acid (Hahlbrock and Scheel 1989; Schilmiller et al. 2009; Tohge et al. 2013). To confirm catalytic activity of GbC4H, trans-cinnamic acid and the recombinant GbC4H from E. coli were reacted in vitro. The enzyme assay resulted in hydroxylation of the substrate into p-coumaric acid (Fig. 6). C4H, a subfamily of cytochrome P450 monooxygenase superfamily, is one of the first P450 identified in plants (Ehlting et al. 2006). C4H activity and gene expression level are regulated by a variety of determinants, including illumination, wounding, and pathogen infection, which demonstrates diverse roles of C4H in phenylpropanoid pathway (Chapple 1998). Schilmiller et al. (2009) found that decrease in C4H activity leads to a strong reduction in phenylpropanoids metabolites and severely influenced the Arabidopsis growth and development. Tabata (1996) observed that activity of C4H positively correlates with lignin metabolism. Therefore, improving activity of C4H may be a way to obtain higher lignin content. In this study, GbC4H was confirmed to possess expected catalytic activity, indicating that this protein is involved in lignin biosynthesis in Ginkgo.
The relationship between GbC4H transcription and lignin contents
Because P450 enzymes are catalytically slow, so they probably catalyze the rate limiting step in phenylpropanoid pathway. In addition to that, C4H is a key enzyme involved in lignin biosynthesis. Thus, lignin contents in different Ginkgo organs could be influenced by C4H expression level. To confirm the relationship between GbC4H and lignin, GbC4H transcription level and lignin contents were determined in the Ginkgo organs of different developmental stage (Fig. 7).
Fresh stem, root, fruit, carpopodium, and petiole sections from Ginkgo plants were subjected to phloroglucinol staining, and stained sections showed obvious pink color due to reaction with aldehyde end group, for example, of coniferaldehyde of G units. Gymnosperm lignin is consisted of almost entirely G units (Lewis and Yamamoto 1990), with small amounts of H units (Gross 1981; Boerjan et al. 2003). Lignin containing a large number of G units possesses more resistant linkages than that comprising S units because of greater number of carbon–carbon bonds (Kim et al. 2013). In this study, stems and roots sections were stained fresh pink color, whereas fruit, carpopodium, and petiole were stained deep purple color; in addition to that, the stained secondary structures were different among various plant organs (Fig. 7a, A–I). These results indicated that secondary lignification occurred in different tissues of the Ginkgo plants with the tree age. These staining characteristics prompted us to analyse lignin contents in different Ginkgo tissues. Lignin contents were detected in CWRs by the improved Klason method. As expected, lignin contents of stems and roots were higher than those of other tissues (Fig. 7b). Meanwhile, insoluble lignin contents increased with tree age. The lignin contents and composition change widely in organs from the same plant in different developmental stages, and in response to different environmental changes (Plomion et al. 2001; Bhuiyan et al. 2009).
So far, transcript profiling of tree C4H in various tissues at different growth phases has not been reported in detail. In the present study, high transcription levels of GbC4H were detected in stems and roots, followed by fruits, carpopodium, and petiole tissues, these results were consistent with pattern of lignin contents (Fig. 7b, c). In addition, GbC4H transcription levels in stems of different developmental stages also changed significantly. The highest transcription levels were detected in 5-year-old stems, whereas lower transcription levels were observed in 2-year-old stems (Fig. 7c), suggesting positive relationship of GbC4H transcription level with tree age. Finally, lignin content had high correlation with the transcription level of GbC4H. Taken together, these results indicated that lignin contents in different Ginkgo organs were influenced by GbC4H expression level. Further in this study, enzyme is necessary to deepen our understanding of Ginkgo lignification in time- and tissue-dependent regulation of C4H expression during plant development.
UV-B, cold conditions, and hormones regulate the GbC4H transcription levels
Plant defense can be reinforced by stress-induced syntheses of defense-related enzymes and lignin (Fujita et al. 2006; Hano et al. 2006; Desender et al. 2007; Hamann et al. 2009). Plant hormones, including ABA, SA, jasmonic acid, and ethylene, can be induced by stresses. These plant hormones may regulate the expressions of different genes that related to lignin synthesis. The initial signal in the gene expression can be reinforced by the plant hormones that produce a second round signal transduction (Mahajan and Tuteja 2005; Shao et al. 2007). Study on signal molecules regulating plant stress response is a crucial step in better understanding mechanisms of plant acclimatization to adverse environments.
SA is known as a signal molecule or a phytohormone synthesized for the activation of plant defense mechanisms (Khan et al. 2015). Studies have confirmed that exogenous application of SA results in significantly influences on plant growth and development, flowering, ion uptake, stress response, and photosynthesis (Khan et al. 2003; Popova et al. 2009; Kadioglu et al. 2011; Saharkhiz et al. 2011). In the present study, with treatment of SA, a slight induction of GbC4H gene was observed at 2 h after treatment, with the highest level being achieved at 8 h followed by a rapid decline (Fig. 8a). SA performs an important role in defense responses against pathogens and abiotic stress, and higher SA contents can induce more strongly defense mechanisms. Therefore, exogenous SA can induce GbC4H expression, which may lead to higher supply of pivotal precursors for defense compounds, including lignin. In this study, SA treatment significantly enhanced transcription level of GbC4H, which may be involved in lignin biosynthesis in Ginkgo (Fig. 8a). This result was consistent with the fact that GbC4H promoter region contained GT-1 and W-box motifs (Table 2S) which are known SA-responsive cis-elements (Buchel et al. 1999; Redman et al. 2002). The signal transduction mechanism of SA to induce GbC4H expression will be studied.
Under abiotic stress conditions, ABA biosynthesis is rapidly induced to initiate various molecular responses. Among these responses, expression of stress-related genes leading to adaptation to stress conditions is well-known (Cutler et al. 2010; Lee and Luan 2012; Peirats-Llobet et al. 2016). Although genes in ABA biosynthesis pathway are induced by a certain stress, they do not respond to exogenous ABA (Zhu 2002). These findings imply that there are two pathways of ABA signal transduction, ABA-independent and -dependent pathways. C4H gene in kenaf is up-regulated in response to ABA treatment, implying that HcC4H may be classified into an ABA-dependent type (Kim et al. 2013). In this study, GbC4H transcription levels were significantly increased after ABA treatment and peaked at 24 h before decreasing gradually (Fig. 8b). In addition to that, Myb/Myc recognition sites were identified in GbC4H promoter region. Both sites are known to play crucial roles in plant response to pathogens. Therefore, GbC4H is regulated in ABA-dependent manner in Ginkgo.
The function of lignin in adaptation to cold remains unclear, although accumulating evidence shows that cold condition may lead to changes in lignin content (Moura et al. 2010). Induction of genes related to lignin biosynthesis at low temperature, for example, Rhododendron catawbiense RcC3H, Ipomoea batatas IbCAD1, Glycine max GmPAL, Hibiscus cannabinus HcC4H, and PAL, 4CL, and CAD in winter barley (Wei et al. 2006; Kim et al. 2010; Moura et al. 2010; Janská et al. 2011; Kim et al. 2013), has been demonstrated. As shown in Fig. 8c, GbC4H expression significantly increased at 4 °C and reached maximum at 24 h post-treatment. Up-regulation of GbC4H by cold temperature was predicted by Myb/Myc recognition sites related to low temperature response, found in the promoter of GbC4H gene (Table 2S, Fig. 4).
Intensity of UV-B irradiation affects plants’ defense and repair mechanisms. Biosynthesis of UV-absorbing compounds, primarily flavonoids and other metabolites of phenylpropanoid pathway, is the primary protection mechanism in response to potentially nocuous irradiation (Hahlbrock and Scheel 1989). Previous researches indicated that UV-absorbing compounds, notably accumulate after UV-B irradiation, are the result of the increased expression of phenylpropanoid pathway genes (Jaakola et al. 2004). As shown in Fig. 8d, GbC4H transcripts increased markedly at 4 h after UV-B irradiation and reached a maximum after 24 h of UV-B treatment. Up-regulation of GbC4H by UV-B may be due to mobilization of the gene by light-responsive cis-elements identified in the GbC4H promoter region (Table 2S, Fig. 4).
Conclusion
Lignin biosynthesis is regulated by enzymes and their corresponding genes, which participate in the formation, transportation, and polymerization of lignin. C4H, a key enzyme of lignin biosynthesis pathway, being positioned at the second step of the phenylpropanoid pathway, catalyzes conversion of trans-cinnamic acid to p-coumaric acid, the intermediate of lignin synthesis. In the present study, a full-length C4H gene (GbC4H) putatively encoding cinnamate 4-hydroxylase was isolated from G. biloba. This gene contained common conserved domains of cytochrome P450 monooxygenase family and shared the highest identity with C4H from other gymnosperms. Spatial–temporal C4H transcript levels, histochemical lignin staining, and lignin content analysis suggested that C4H was active in Ginkgo with lignin being accumulated in stems and roots. Interestingly, we discovered a strong correlation between expression level of C4H and lignin accumulation, which supported the essential role of GbC4H in lignification in Ginkgo. Furthermore, phenylpropanoid pathway in Ginkgo was activated immediately after treatment with SA, ABA, UV-B, and cold. GbC4H gene may play a crucial role in phenylpropanoid pathway by responding to increase of SA and ABA to enhance defense mechanisms. Therefore, GbC4H led us to understand organ-specific lignification and defense mechanisms. The findings of the present study could not only promote further study on the enzymes in lignin biosynthesis pathway in G. biloba but also stimulate effort to increase lignin content by the metabolic engineering.
Author contribution statement
Shuiyuan Cheng, Jiaping Yan and Feng Xu conceived and designed the experiments and drafted the manuscript. Shuiyuan Cheng, Jiaping Yan, and Xiangxiang Meng performed the experiments. Weiwei Zhang, Yongling Liao and Jiabao Ye analyzed the data. All authors read and approved the manuscript.
References
Achnine L, Blancaflor EB, Rasmussen S et al (2004) Colocalization of l-phenylalanine ammonia-lyase and cinnamate 4-hydroxylase for metabolic channeling in phenylpropanoid biosynthesis. Plant Cell 16:3098–3109
Anterola AM, Lewis NG (2002) Trends in lignin modification: a comprehensive analysis of the effects of genetic manipulations/mutations on lignification and vascular integrity. Phytochemistry 61:221–294
Bhuiyan NH, Selvaraj G, Wei Y et al (2009) Role of lignification in plant defense. Plant Signal Behav 4:158–159
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Boudet AM, Kajita S, Grima-Pettenati J et al (2003) Lignins and lignocellulosics: a better control of synthesis for new and improved uses. Trends Plant Sci 8:576–581
Buchel AS, Brederode F, Bol JF et al (1999) Mutation of GT-1 binding sites in the Pr-1A promoter influences the level of inducible gene expression in vivo. Plant Mol Biol 40:387–396
Chapple C (1998) Molecular-genetic analysis of plant cytochrome P450-dependent monooxygenases. Annu Rev Plant Biol 49:311–343
Chen AH, Chai YR, Li JN et al (2007) Molecular cloning of two genes encoding cinnamate 4-hydroxylase (C4H) from oilseed rape (Brassica napus). J Biochem Mol Biol 40:247–260
Cheng H, Li LL, Xu F et al (2013a) Expression patterns of an isoflavone reductase-like gene and its possible roles in secondary metabolism in Ginkgo biloba. Plant Cell Rep 32:637–650
Cheng H, Li LL, Xu F et al (2013b) Expression patterns of a cinnamyl alcohol dehydrogenase gene involved in lignin biosynthesis and environmental stress in Ginkgo biloba. Mol Biol Rep 40:707–721
Chiang VL (2006) Monolignol biosynthesis and genetic engineering of lignin in trees, a review. Environ Chem Lett 4:143–146
Cutler SR, Rodriguez PL, Finkelstein RR et al (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679
Desender S, Andrivon D, Val F (2007) Activation of defence reactions in Solanaceae: where is the specificity. Cell Microbiol 9:21–30
Dixon RA, Paiva NL (1995) Stress-induced phenylpropanoid metabolism. Plant Cell 7:1085
Dixon RA, Lamb CJ, Masoud S et al (1996) Metabolic engineering: prospects for crop improvement through the genetic manipulation of phenylpropanoid biosynthesis and defense responses—a review. Gene 179:61–71
Durst F, Nelson DR (1995) Diversity and evolution of plant P450 and P450-reductases. Drug Metab Drug Interact 12:189–206
Ehlting J, Hamberger B, Million-Rousseau R et al (2006) Cytochromes P450 in phenolic metabolism. Phytochem Rev 5:239–270
Ferrer JL, Austin MB, Stewart C et al (2008) Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant Physiol Biochem 46:356–370
Fujita K, Komatsu K, Tanaka K et al (2006) An in vitro model for studying vascular injury after laser microdissection. Histochem Cell Biol 125:509–514
Gross GG (1981) The biochemistry of lignification. Adv Bot Res 8:25–63
Hahlbrock K, Scheel D (1989) Physiology and molecular biology of phenylpropanoid metabolism. Annu Rev Plant Biol 40:347–369
Hamann T, Bennett M, Mansfield J et al (2009) Identification of cell-wall stress as a hexose-dependent and osmosensitive regulator of plant responses. Plant J 57:1015–1026
Hano C, Addi M, Bensaddek L, Crônier D et al (2006) Differential accumulation of monolignol-derived compounds in elicited flax (Linum usitatissimum) cell suspension cultures. Planta 223:975–989
Himmel ME (2008) Biomass recalcitrance: deconstructing the plant cell wall for bioenergy. Blackwell, Oxford, pp 1–6
Hotze M, Schröder G, Schröder J (1995) Cinnamate 4-hydroxylase from Catharanthus roseus and a strategy for the functional expression of plant cytochrome P450 proteins as translational fusions with P450 reductase in Escherichia coli. FEBS Lett 374:345–350
Huang B, Duan Y, Yi B et al (2008) Characterization and expression profiling of cinnamate 4-hydroxylase gene from Salvia miltiorrhiza in rosmarinic acid biosynthesis pathway. Russ J Plant Physiol 55:390–399
Jaakola L, Määttä-Riihinen K, Kärenlampi S et al (2004) Activation of flavonoid biosynthesis by solar radiation in bilberry (Vaccinium myrtillus L.) leaves. Planta 218:721–728
Janská A, Aprile A, Zámečník J et al (2011) Transcriptional responses of winter barley to cold indicate nucleosome remodelling as a specific feature of crown tissues. Funct Integr Genom 11:307–325
Kadioglu A, Saruhan N, Sağlam A et al (2011) Exogenous salicylic acid alleviates effects of long term drought stress and delays leaf rolling by inducing antioxidant system. Plant Growth Regul 64:27–37
Khan W, Prithiviraj B, Smith DL (2003) Photosynthetic responses of corn and soybean to foliar application of salicylates. J Plant Physiol 160:485–492
Khan MIR, Fatma M, Per TS et al (2015) Salicylic acid-induced abiotic stress tolerance and underlying mechanisms in plants. Front Plant Sci 6:462
Kim YH, Bae JM, Huh GH (2010) Transcriptional regulation of the cinnamyl alcohol dehydrogenase gene from sweetpotato in response to plant developmental stage and environmental stress. Plant Cell Rep 29:779–791
Kim J, Choi B, Natarajan S et al (2013) Expression analysis of kenaf cinnamate 4-hydroxylase (C4H) ortholog during developmental and stress responses. Plant Omics 6:65–72
Kirk TK, Obst JR (1988) Lignin determination. Method Enzymol 161:87–101
Kochs G, Grisebach H (1989) Phytoalexin synthesis in soybean: purification and reconstitution of cytochrome P450 3, 9-dihydroxypterocarpan 6a-hydroxylase and separation from cytochrome P450 cinnamate 4-hydroxylase. Arch Biochem Biophys 273:543–553
Kong JQ, Lu D, Wang ZB (2014) Molecular cloning and yeast expression of cinnamate 4-hydroxylase from Ornithogalum saundersiae baker. Molecules 19:1608–1621
Kumar S, Omer S, Chitransh S et al (2012) Cinnamate 4-hydroxylase downregulation in transgenic tobacco alters transcript level of other phenylpropanoid pathway genes. Int J Adv Biotechnol Res 3:545–557
Kumar S, Omer S, Patel K et al (2013) Cinnamate 4-hydroxylase (C4H) genes from Leucaena leucocephala: a pulp yielding leguminous tree. Mol Biol Rep 40:1265–1274
Lee SC, Luan S (2012) ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ 35:53–60
Lewis NG, Yamamoto E (1990) Lignin: occurrence, biogenesis and biodegradation. Annu Rev Plant Biol 41:455–496
Liu S, Hu Y, Wang X et al (2009) Isolation and characterization of a gene encoding cinnamate 4-hydroxylase from Parthenocissus henryana. Mol Biol Rep 36:1605–1610
Lu S, Zhou Y, Li L et al (2006) Distinct roles of cinnamate 4-hydroxylase genes in Populus. Plant Cell Physiol 47:905–914
Mahajan S, Tuteja N (2005) Cold, salinity and drought stresses: an overview. Arch Biochem Biophys 444:139–158
Mizutani M, Ohta D, Sato R (1997) Isolation of a cDNA and a genomic clone encoding cinnamate 4-hydroxylase from Arabidopsis and its expression manner in planta. Plant Physiol 113:755–763
Moura JCMS, Bonine CAV, De Oliveira Fernandes Viana J (2010) Abiotic and biotic stresses and changes in the lignin content and composition in plants. J Integr Plant Biol 52:360–376
Naoumkina MA, Zhao Q, Gallegogiraldo L et al (2010) Genome-wide analysis of phenylpropanoid defence pathways. Mol Plant Pathol 11:829–846
Nedelkina S, Jupe SC, Blee KA et al (1999) Novel characteristics and regulation of a divergent cinnamate 4-hydroxylase (CYP73A15) from French bean: engineering expression in yeast. Plant Mol Biol 39:1079–1090
Nelson DR, Schuler MA, Paquette SM et al (2004) Comparative genomics of rice and Arabidopsis. Analysis of 727 cytochrome P450 genes and pseudogenes from a monocot and a dicot. Plant Physiol 135:756–772
Peirats-Llobet M, Han SK, Gonzalez-Guzman M et al (2016) A direct link between abscisic acid sensing and the chromatin-remodeling ATPase BRAHMA via core ABA signaling pathway components. Mol Plant 9:136–147
Plomion C, Leprovost G, Stokes A (2001) Wood formation in trees. Plant Physiol 127:1513–1523
Pomar F, Novo M, Bernal MA et al (2004) Changes in stem lignins (monomer composition and crosslinking) and peroxidase are related with the maintenance of leaf photosynthetic integrity during Verticillium wilt in Capsicum annuum. New Phytol 163:111–123
Popova LP, Maslenkova LT, Yordanova RY et al (2009) Exogenous treatment with salicylic acid attenuates cadmium toxicity in pea seedlings. Plant Physiol Biochem 47:224–231
Ralph J, Lundquist K, Brunow G et al (2004) Lignins: natural polymers from oxidative coupling of 4-hydroxyphenyl-propanoids. Phytochem Rev 3:29–60
Rani A, Singh K, Ahuja PS (2012) Molecular regulation of catechins biosynthesis in tea [Camellia sinensis (L.) O. Kuntze]. Gene 495:205–210
Redman J, Whitcraft J, Johnson C et al (2002) Abiotic and biotic stress differentially stimulate as-1 element activity in Arabidopsis. Plant Cell Rep 21:180–185
Reyes JC, Muro-Pastor MI, Florencio FJ (2004) The GATA family of transcription factors in Arabidopsis and rice. Plant Physiol 134:1718–1732
Ro DK, Douglas CJ (2004) Reconstitution of the entry point of plant phenylpropanoid metabolism in yeast (Saccharomyces cerevisiae) implications for control of metabolic flux into the phenylpropanoid pathway. J Biol Chem 279:2600–2607
Ro DK, Mah N, Ellis BE et al (2001) Functional characterization and subcellular localization of poplar (Populus trichocarpa × Populus deltoides) cinnamate 4-hydroxylase. Plant Physiol 126:317–329
Saharkhiz MJ, Mohammadi S, Javanmardi J (2011) Salicylic acid changes physio-morphological traits and essential oil content of catnip (Nepeta cataria L.). J Med Spice Plants 16:75–77
Salvador VH, Lima RB, dos Santos WD et al (2013) Cinnamic acid increases lignin production and inhibits soybean root growth. PLoS One 8:e69105
Sarkanen KV, Ludwig CH (1971) Lignins. Occurrence, formation, structure, and reactions. Wiley, New York
Schilmiller AL, Stout J, Weng JK (2009) Mutations in the cinnamate 4-hydroxylase gene impact metabolism, growth and development in Arabidopsis. Plant J 60:771–782
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101
Schoch GA, Attias R, Le Ret M et al (2003) Key substrate recognition residues in the active site of a plant cytochrome P450, CYP73A1. FEBS J 270:3684–3695
Shao HB, Guo QJ, Chu LY et al (2007) Understanding molecular mechanism of higher plant plasticity under abiotic stress. Colloid Surface B 54:37–45
Singh K, Kumar S, Rani A et al (2009) Phenylalanine ammonia-lyase (PAL) and cinnamate 4-hydroxylase (C4H) and catechins (flavan-3-ols) accumulation in tea. Funct Integr Genom 9:125–134
Smith JV, Luo Y (2004) Studies on molecular mechanisms of Ginkgo biloba extract. Appl Microbiol Biotechnol 64:465–472
Sykes RW, Gjersing EL, Foutz K et al (2015) Down-regulation of p-coumaroyl quinate/shikimate 3′-hydroxylase (C3′H) and cinnamate 4-hydroxylase (C4H) genes in the lignin biosynthetic pathway of Eucalyptus urophylla × E. grandis leads to improved sugar release. Biotechnol Biofuels 8:128
Szczesnaskorupa E, Straub P, Kemper B (1993) Deletion of a conserved tetrapeptide, PPGP, in P450 2C2 results in loss of enzymatic activity without a change in its cellular location. Arch Biochem Biophys 304:170–175
Tabata M (1996) The mechanism of shikonin biosynthesis in Lithospermum cell cultures. Plant Tissue Culture Lett 13:117–125
Tao S, Khanizadeh S, Zhang H et al (2009) Anatomy, ultrastructure and lignin distribution of stone cells in two Pyrus species. Plant Sci 176:413–419
Terashima N, Kitano K, Kojima M et al (2009) Nanostructural assembly of cellulose, hemicellulose, and lignin in the middle layer of secondary wall of ginkgo tracheid. J Wood Sci 55:409–416
Teutsch HG, Hasenfratz MP, Lesot A et al (1993) Isolation and sequence of a cDNA encoding the Jerusalem artichoke cinnamate 4-hydroxylase, a major plant cytochrome P450 involved in the general phenylpropanoid pathway. Proc Natl Acad Sci USA 90:4102–4106
Tohge T, Watanabe M, Hoefgen R et al (2013) The evolution of phenylpropanoid metabolism in the green lineage. Crit Rev Biochem Mol 48:123–152
Van Beek TA (2002) Chemical analysis of Ginkgo biloba leaves and extracts. J Chromatogr A 967:21–55
Vanholme R, Morreel K, Ralph J et al (2008) Lignin engineering. Curr Opin Plant Biol 11:278–285
Vogt T (2010) Phenylpropanoid biosynthesis. Mol Plant 3:2–20
Wei HUI, Dhanaraj AL, Arora R et al (2006) Identification of cold acclimation-responsive Rhododendron genes for lipid metabolism, membrane transport and lignin biosynthesis: importance of moderately abundant ESTs in genomic studies. Plant Cell Environ 29:558–570
Weisshaar B, Jenkins GI (1998) Phenylpropanoid biosynthesis and its regulation. Curr Opin Plant Biol 1:251–257
Weng JK, Chapple C (2010) The origin and evolution of lignin biosynthesis. New Phytol 187:273–285
Weng JK, Li X, Bonawitz ND (2008) Emerging strategies of lignin engineering and degradation for cellulosic biofuel production. Curr Opin Biotechnol 19:166–172
Werck-Reichhart D, Batard Y, Kochs G (1993) Monospecific polyclonal antibodies directed against purified cinnamate 4-hydroxylase from Helianthus tuberosus (immunopurification, immunoquantitation, and interspecies cross-reactivity). Plant Physiol 102:1291–1298
Xu F, Cheng H, Cai R (2008) Molecular cloning and function analysis of an anthocyanidin synthase gene from Ginkgo biloba, and its expression in abiotic stress responses. Mol Cell 26:536–547
Xu Z, Zhang D, Hu J (2009) Comparative genome analysis of lignin biosynthesis gene families across the plant kingdom. BMC Bioinform 10:S3
Xu H, Park NI, Li X et al (2010) Molecular cloning and characterization of phenylalanine ammonia-lyase, cinnamate 4-hydroxylase and genes involved in flavone biosynthesis in Scutellaria baicalensis. Bioresour Technol 101:9715–9722
Xu F, Ning YJ, Zhang WW et al (2014) An R2R3-MYB transcription factor as a negative regulator of the flavonoid biosynthesis pathway in Ginkgo biloba. Funct Integr Genom 14:177–189
Yamazaki S, Sato K, Suhara K (1993) Importance of the proline-rich region following signal-anchor sequence in the formation of correct conformation of microsomal cytochrome P-450s. J Biochem 114:652–657
Yang DH, Chung BY, Kim JS (2005) cDNA cloning and sequence analysis of the rice cinnamate-4-hydroxylase gene, a cytochrome P450-dependent monooxygenase involved in the general phenylpropanoid pathway. J Plant Biol 48:311–318
Yeh TF, Yamada T, Capanema E (2005) Rapid screening of wood chemical component variations using transmittance near-infrared spectroscopy. J Agric Food Chem 53:3328–3332
Zeng Y, Zhao S, Yang S (2014) Lignin plays a negative role in the biochemical process for producing lignocellulosic biofuels. Curr Opin Biotechnol 27:38–45
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgements
This work was supported by the National Science Foundation of China (No. 31370680).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A Chandra.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cheng, S., Yan, J., Meng, X. et al. Characterization and expression patterns of a cinnamate-4-hydroxylase gene involved in lignin biosynthesis and in response to various stresses and hormonal treatments in Ginkgo biloba . Acta Physiol Plant 40, 7 (2018). https://doi.org/10.1007/s11738-017-2585-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-017-2585-4