Abstract
Recently, herbal medicinal plants have gained more attention worldwide to assimilate in an innovative nanomaterial’s fabrication. Therefore, this is the first report devoted to emphasizing the biogenic synthesis of palladium nanoparticles using Madhuca longifolia or Mahua leaves extract (MLE@PdNPs) for their biomedical applications. In this, we have used the herbal plant M. longifolia leaves as a bioreductant and capping agent. The structural and morphological features of synthesized MLE@PdNPs have been studied using different analytical techniques. Triangular sheet-like stable MLE@PdNPs have an average crystallite size of 16.22 nm with a face center cubic (fcc) structure which has been observed. The biological screening of MLE@PdNPs has been performed against the human lung cancer cells A549 and bacterial strains S. aureus, K. pneumonia, Salmonella, and E. coli. The minimal inhibitory concentration (MIC or IC-50) value against lung cancer cells A549 has been calculated 29.20 μg/ml. The IC-50 values have been calculated 26.01, 30.11, 32.12, and 35.16 μg/mL against S. aureus, K. pneumonia, Salmonella, and E. coli bacterial strains, respectively. The findings of the present work will become beneficial to expand the studies on the therapeutic potential of MLE@PdNPs in the future. Enthusiastically, MLE@PdNPs can be a promising material for formulating nanomedicine after subsequent clinical experiments.
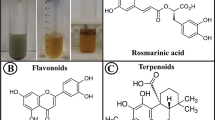
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The growing edge of science and technology mainly dedicated to developed nanomaterials of the dimension lesser than a hundred nanometers, i.e., nanoparticles (NPs), as they are handy contribution toward smarter, less heavy, eco-friendly, and stronger surfaces systems (Baig et al. 2021). Nanomaterials, especially based on noble metals, gain immense attention due to their surprising physicochemical characteristics. Nanoparticles’ physical and chemical properties are different and unpredictable from those formed in corresponding bulk material (Gangwar et al. 2022; Yaseen et al. 2023). In addition, these characteristics are enormously dependent on their shape and size. Also, the shape and size of nanomaterials may be tuned depending upon the synthesis route, surfactant, or stabilizer surrounding the nanomaterial to provide them stability, etc. (Gangwar et al. 2021). Due to the high demand for noble metal-based nanomaterials, researchers continuously try to serve new and intelligent materials with unique features. Although many studies had reported different methods for synthesis of metal such as silver (Ag), platinum (Pt), palladium (Pd), etc., based nanomaterials to successfully employed in different areas (Abdel-Rahman et al. 2022; MubarakAli et al. 2022; Muñiz-Diaz et al. 2022; Prema et al. 2022). Still, some new methodologies are required to fulfill the requirements of green chemistry worldwide. In the present study, we have tried to develop a sustainable approach to synthesize the palladium nanoparticles (PdNPs) as they are potentially utilized in different areas (MubarakAli et al. 2022; Siddiqi & Husen 2016; Vinodhini et al. 2022a). Environmentally benign or biogenic synthesis of noble metal-based nanomaterials is preferentially followed in this work because this leads to the elimination of the formation of dangerous or unwanted products and the involvement of non-toxic solvents or surfactants (I. Kumar et al. 2021a, b; Noah & Ndangili 2022). Since plants are the source of a variety of chemicals that can act as a bioreductant as well as a stabilizer, hence, here plants are preferred for biogenic synthesis.
The benefit of using plants for biogenic synthesis is that the experiment can be successfully achieved at its natural pH and physiological condition of temperature and pressure. Nevertheless, at the same time, it has been found that the morphologies, dimensions, and stability of nanomaterials are remarkably influenced by temperature, pH, concentration of plant extract/metal ions, and incubation time (Siddiqi & Husen 2016). Therefore, the present study deals with Madhuca longifolia leaves aqueous extract to synthesize MLE@PdNPs. M. longifolia is a herbal plant, commonly known as Mahua, and also known as the Moah tree, Moah wood, Mowra butter, Butternut tree, and Moa tree (Verma et al. 2014). It belongs to the family of Sapotaceae. It is distributed in the northern, central, and southern regions of the Indian Peninsula, Nepal, Sri Lanka, and Myanmar. It is a large-sized multipurpose tropical tree and flourishes in dry environments. It is mainly harvested for its oilseeds and edible flowers (Al Qubeissi et al. 2023; Y. Kumar et al. 2017; Singh et al. 2022). As per Ayurveda, Mahua is a medicinal plant and is well known for numerous phytochemicals, e.g., vitamin A and vitamin C in flowers, α- and β- amyrin acetates in fruits, and arachidic, linolenic, oleic, myrisic, palmitic, stearic acid, α-alanine, aspartic acid, cysteine, glycine, isoleucine, and leucine, lysine, methionine, proline, serine, threonine, myricetin, quercetin, and misaponin are present in seeds (Kumar et al. 2017; Reddy 2022). However, Annalakshmi et al. reported that there were 20 phytochemicals, including hydrocarbons, ketones, aldehydes, phenolic acids, carbohydrates, and heterocyclic compounds present in Mahua leaves. Also, a highly markable amount of quercetin was found during high-performance thin-layer chromatography (HPTLC) (Annalakshmi et al. 2013; Jha & Mazumder 2018; Reddy 2022).
Besides, it has been found to be effective in curing eczema, rheumatism, and hemorrhoids. One of the research reports mentioned that Mahua leaves extract in methanol was found to be responsible for the enhanced release of a neuroleptic agent, i.e., GABA, which is agitable to the neuronal activity in epilepsy patients (Jodh et al. 2022; Sinha et al. 2017). Consequently, it not only delays the onset of seizures but also diminishes their duration. Its leaves are also used to provide symptomatic relief to the skin affected by eczema. It is an extraordinary remedy for parasitic infections and shows potent anti-helminthic activity in the presence of another plant. Moreover, it shows good antioxidant activity (Kurhade et al. 2022). After studying the numerous medicinal utilities of the Mahua plant, we have planned to incorporate its aqueous leaf extract as a bioreductant and stabilizing agent for the synthesis of MLE@PdNPs to employ them in biomedical applications.
To the best of our knowledge, until today, no articles report the synthesis of palladium nanoparticles by using M. Longifolia or Mahua leaves extract (MLE@PdNPs) at 80 ℃. The confirmed synthesis of palladium nanoparticles was investigated by using various characterization techniques. The biological screening of the MLE@PdNPs has been performed against the lung cancer cells A549 and four bacterial strains, namely S. aureus, K. pneumonia, Salmonella, and E. coli. The minimal inhibitory concentration (MIC) for lung cancer cells A549 has been evaluated by using 96 well plates experiment. However, the antibacterial efficacy of MLE@PdNPs has been evaluated by MIC against normal and multiple drug-resistant (MDR) strains of S. aureus, K. pneumoniae, Salmonella, and E. coli.
Experimental
Materials
PdCl2 salt was purchased from Sisco research laboratories private limited and was used during the synthesis without further purification. All other chemicals were of analytical grade (AG) and procured from Sigma and Hi-Media, Mumbai, India. Note: A stock solution of 0.1 mM has been prepared by first dissolving PdCl2 salt into the required volume of 1% mineral acid solution and then adding the double-distilled water to make up the volume of the graduated flask 100 ml. For buffer and media preparation, ultrapure milli-Q water was used.
Methods
Collection and extract preparation of mahua leaves
10 g of fresh mature (wet) leaves of Mahua was collected from the area of SGPGI, Lucknow, Uttar Pradesh (satellite imagery is shown in Fig. 1). The leaves were washed twice with distilled water and once double-distilled (dd) water. Finally, the rinsed leaves were chopped into small pieces, soaked in a beaker containing 100 ml dd water, and gently heated for an hour at 60 ℃. The filtrate so obtained is mahua leaves extract (MLE).
Synthesis of MLE@PdNPs
A simple one-pot sequential procedure was involved in synthesizing MLE@PdNPs. A round-bottom flask containing 20 ml MLE and 75 ml dd water was left for an hour in a thermostat to maintain its temperature at 80 ℃. Then, the thermally equilibrated PdCl2 solution (5 ml, 0.01 M) was added dropwise into the round-bottom flask and fitted it with a water condenser to avoid evaporation of the reaction mixture. The overall reaction mixture was allowed to stir on a magnetic stirrer at 3000 rounds per minute (rpm) at 80 ℃ for 1 h. The change in reaction mixture color from blood red to brownish black was observed, which confirms the formation of MLE@PdNPs colloid. Then, this MLE@PdNPs colloid was centrifuged for about 20 min to obtain the powder MLE@PdNPs. This powder MLE@PdNPs was washed with dd water followed by ethyl alcohol and dried in an oven for 5 h at 100 ℃ (as shown in Fig. 2).
Instrumentation
A double-beam spectrophotometer of LAB UV Next Generation scanned in the wavelength range of 200–700 nm with a least count of 2 nm was used to record absorption spectra. Fourier transformed infrared spectrum (FTIR) spectrum was recorded on the Shimadzu company’s IR Affinity-1S double-beam spectrophotometer. The spectrum was recorded in the range of wavenumber from 3600–400 cm−1 with a resolution of 2 nm. It helps determine the functional groups surrounding the synthesized MLE@PdNPs compared to M. Longifolia leaves. It involves the KBr pellet method; the pellet of prepared MLE@PdNPs was made by assorting with KBr in 1:1000. The powder X-ray diffraction was done on a Rigaku diffractometer built with a 9000 W rotating anode X-ray generator with a NaI scintillation counter-detection system. The copper (Cu) anode of the wavelength of 0.154 nm was used to target the material at an angle (2θ) range between 10° and 80°. It helps in predicting the lattice parameter and average crystallite size of MLE@PdNPs. Field emission scanning electron microscopy (FESEM) verified the morphology of the synthesized MLE@PdNPs. To capture the images, the field emission scanning electron microscope of JSM-6490 LV, JEOL, Tokyo, Japan, operating at a voltage of 5.0 kV was used. Transmission electron microscopy (TEM) was used to examine the size, morphology, or topology of the synthesized MLE@PdNPs. The cryo-TEM (Talos) transmission electron microscope was used in the present investigation. Dynamic light scattering (DLS) is used to determine the electrokinetic potential (ξ) of the synthesized MLE@PdNPs. The Nano Zetasizer, Malvern Instruments, Worcestershire, UK, was used to record the intensity and number distribution plot.
Evaluation of cytotoxicity of MLE@PdNPs
The 96-well plates method has been used to grow 5 × 103 A549 cells and incubated in an incubator for a day at 37 °C in a humidified 5% CO2 atmosphere. After that, the cells have been subjected to different concentrations of MLE@PdNPs, i.e., 10, 20, 30, 40, 50, 60, and 70 µg/ml. The triplicate runs have been performed with incubation for an extra day. Afterward, into each well, 10 µL MTT was added. Then, the plates were incubated for 2 h in the same incubator. Additionally, formazan crystals (FC) were dissolved in DMSO (100 µL) to make a clear solution and subjected to record the absorbance at a wavelength (λ) of 570 nm using an ELISA reader (Microplate Reader BIORAD-680).
Study of cytomorphological changes of MLE@PdNPs-treated A549 cells
The A549 cells were pre-treated with IC-50 concentrations of MLE@PdNPs and incubated in a CO2 (5%) atmosphere for a day at 37 °C. An inverted phase-contrast microscope (Nikon ECLIPSE Ti-S, Nikon Corporation, Tokyo, Japan) was used to study the net morphological variations in treated cells.
Quantification and detection of production of reactive oxygen species (ROS)
The generation of ROS in MLE@PdNPs-treated A549 cells has been determined under the fluorescence microscope. The observation was based on the following factors such as fluorescence of a fluorogenic marker, 5-(6)-carboxy-2’,7’-dichlorodihydrofluorescein diacetate (carboxy-H2DCFDA) in viable cells. The cells (5 × 103 in each well) were seeded into 24-well culture plates and incubated for 24 h inside the CO2 (5%) incubator at 37 °C. Later, the cells were incubated with MLE@PdNPs for 24 h. Ultimately, cells were incubated with H2DCFDA (10 µM) for 30 min at 37 °C, and other cells were supplemented with 200 µL of phosphate buffer saline (PBS) in every well. These plates were kept in a shaker for the next 10 min at room temperature. The fluorescence of cells was determined by using a magnifying lens (fluorescence microscope evosFLc). Quantification of cellular fluorescence from fluorescence microscopy images was performed by ImageJ software.
Study of variations in nuclear morphology of cells
The fluorescent nuclear dye DAPI has been used to estimate the effect of MLE@PdNPs on the genetic materials of A549 cells. The grown cells were washed with PBS buffer and then incubated for 10 min with 4% paraformaldehyde (PFA). Finally, these treated cells were permeabilized with 3% paraformaldehyde and 0.5% Triton X-100, i.e., the permeabilizing buffer, and stained by DAPI (fluorescence dye). The images of stained cells were captured using the fluorescence microscope. The cells with fragmented and condensed nuclei had been considered apoptotic cells. ImageJ software has been used to quantify cellular fluorescence in cells from fluorescence microscopy images.
Antimicrobial activity
MLE@PdNPs that were synthesized have been examined for their antibacterial susceptibility against pathogenic and multidrug-resistant (MDR) bacteria. The antibacterial effectiveness of MLE@PdNPs was evaluated using the agar well diffusion method. To produce a uniform microbial growth plate, bacteria were sub-cultured in 0.03% Luria–Bertani (LB) broth medium and incubated at 37 °C for 24 h. Then, overnight cultures were taken and spread on the agar plates. The bacterial strains were (A) Staphylococcus aureus (MDR), (B) Klebsiella pneumonia, (C) Salmonella, (D) Escherichia coli (MDR). The positive control for the investigation of antibacterial activity was distilled water (DW), and the negative control was considered from previously reported studies (Baker et al. 2021). The petri dishes were then incubated for further 24 h at 37 °C. The diameter of the inhibition zone was measured and contrasted with the control groups to assess the antibacterial activity of MLE@PdNPs.
Determination of minimal inhibitory concentration (MIC)
The antibacterial efficacy of MLE@PdNPs has been evaluated by minimal inhibitory concentration (MIC) against normal and multiple drug-resistant (MDR) strains of S. aureus, K. pneumoniae, Salmonella, and E. coli. Bacterial strains in a mid-logarithmic phase were harvested and diluted to 2 × 105 colony-forming units per mL (CFU/mL) in PBS containing 0.03% Luria–Bertani (LB) broth. The MLE@PdNPs were serially diluted from 2 to 320 μg/ml in 100 μL of LB medium in 96-well plates according to the method described elsewhere (Baker et al. 2021). The MIC was considered the lowest concentration of MLE@PdNPs at which microbial growth was inhibited. Autoclaved water was used as a negative control for each experiment.
Result and discussions
A simple one-step process has been used to synthesize the MLE@PdNPs. In this, palladium chloride was used as a precursor for the synthesis of MLE@PdNPs. After executing experiments with different concentrations of Pd2+(aq.) solution, the best result for fabricating MLE@PdNPs from the MLE was 2 × 10–3 M Pd2+(aq.) solution concentration in mixture at 80 °C. The intensified brownish black color was observed as time passed and consequently proved the formation of MLE@PdNPs by reducing the wine-red-colored Pd2+(aq.) solution. This color change was observed due to stimulation of the surface plasmonic resonance (SPR) band of MLE@PdNPs (Gangwar et al. 2023; S. Vinodhini et al. 2022a, b). Since MLE constitutes many phytochemicals in their leaves, it can be an excellent reducing agent for the fortunate synthesis of MLE@PdNPs (Bains et al. 2020; Dambhare et al. 2020). However, not all phytochemicals are involved in the reduction reaction; few have an excellent ability to act as a stabilizing agent. A most plausible mechanism (Fig. 3) based on the significant amount of phytochemical, i.e., quercetin (Annalakshmi et al. 2013; Dalvi et al. 2022; Khare et al. 2018), was proposed to explain the synthesis of MLE@PdNPs colloid. The quinone form of quercetin accompanied the formation of MLE@PdNPs. It leads to generating hydrogen ions and electrons. Pd2+ ions accept these electrons to reduce themselves into MLE@PdNPs. However, the other constituents of MLE stabilized the formed MLE@PdNPs by forming the layer over their surface. Further, the adsorption of other phytochemicals was also confirmed by FTIR and EDS. Therefore, the biogenic or green synthesis can be a sustainable approach for stabilized black-colored MLE@PdNPs formation. The identification of MLE@PdNPs was done by several analytical techniques discussed below.
Powder X-ray diffraction study
The diffraction pattern of MLE@PdNPs is shown in Fig. 4. Three peaks at 2θ values 39.98°, 46.48°, and 67.96° were observed analogous to the (111), (200), and (200) planes, respectively. It shows face center cubic (fcc) structure according to JCPDS card numbers 87-0643 and 89-4897 (Fahmy et al. 2020; Gangwar et al. 2023; Venkatesham et al. 2015). Notably, a broad peak between 2θ (in degree) equals 20°, and 30° is observed due to the adsorption of phytochemicals on the surface of PdNPs to stabilize them (Seku et al. 2022). Additionally, the PXRD pattern reveals the highly intense peak for the (111) Bragg’s plane. It suggests that the growth of MLE@PdNPs was preferred along the (111) plane. The average crystallite size (D) of the synthesized MLE@PdNPs was estimated using Debye Scherrer equation and calculated value of D was 16.22 nm.
EDS analysis
Figure 5 represents the EDS profile of MLE@PdNPs. A sharp peak ~ 1.5 keV energy was also there in the EDS profile due to the drop-casting of MLE@PdNPs onto silicon/silicon oxide substrate. A prominent peak between 2.7 and 4.0 keV energy for palladium (Pd) was detected, verifying the formation of MLE@PdNPs. It shows 59.83% weight composition of palladium in MLE@PdNPs. Noticeably, the presence of carbon (C) and oxygen (O) with % weight composition of 1.04 and 39.13 was observed due to phytochemicals’ adsorption onto the surface to stabilize MLE@PdNPs (Gangwar et al. 2022).
Electron microscopy analysis (FESEM & TEM)
The MLE@PdNPs synthesized from MLE were seen under the field emission scanning electron microscope (FESEM) and transmission electron microscope (TEM). Figure 6a, b shows images captured from FESEM at a 100 nm scale having resolutions of 30 and 100 k. It reveals that MLE@PdNPs are agglomerated to form a spherical cluster. The maximum height of the spherical cluster perimeter was observed to be 80 nm. Therefore, the average diameter of the cluster was found to be 25.47 nm. Furthermore, Fig. 6c, d represents the topology of MLE@PdNPs at a scale of 50 and 100 nm. Our examination showed that the synthesized MLE@PdNPs are of triangular topology with an average area of 400 nm2.
UV–visible studies of MLE@PdNPs
The absorption spectra of MLE and pure palladium chloride solution are shown in Fig. 7a, b, respectively. Figure 7a shows an absorption peak at λ equals 272 nm corresponding to carbonyl group of the constitutes present in M. longifolia, while the absorption spectra of palladium chloride solution (as shown in Fig. 7b) show absorption maxima at λ equals 442 nm due to the surface plasmonic resonance excitation (Kuniyil et al. 2019; Tanreh et al. 2018). From the individual spectral study of MLE and pure palladium chloride solution, it has been found that no interruption caused due to MLE while studying MLE@PdNPs absorption spectra. This finding reveals that the sharp peak at 442 nm of palladium chloride changes into a broad spectrum with the formation of MLE@PdNPs (as shown in Fig. 7c, d). Different experiments have been performed to achieve a standard condition for MLE@PdNPs synthesis. The standard condition was finalized by monitoring the absorption spectra of each set. In the experiment’s first batch, the palladium chloride solution concentration varied from 0.5 to 2.0 × 10–3 M, keeping a fixed volume of MLE (1 ml), and their corresponding absorption spectra are shown in Fig. 7c. Likewise, in the experiment’s second batch, the MLE volume was changed from 1 to 5 ml, keeping a fixed concentration of palladium chloride solution (0.5 × 10–3 M), and their corresponding absorption spectra are shown in Fig. 7d. Notably, the formation of MLE@PdNPs was also observed by the change in a color of the palladium chloride solution from blood red to brownish black (Gangwar et al. 2023; Raut et al. 2013), and the intensity of the color of MLE@PdNPs was found to be strongly dependent upon the concentration of palladium chloride solution (as shown in Fig. 8). Figure 7c inset graph shows that the absorbance at 442 nm is increased with increasing palladium chloride concentration. Conclusively, it reveals a reciprocal relation between palladium chloride concentration and the rate of formation of MLE@PdNPs at a fixed volume of MLE, i.e., 1 ml, and it may be suggested that 0.5 × 10–3 M is the optimum concentration for the formation of MLE@PdNPs. The formation rate of MLE@PdNPs is first order concerning palladium chloride solution (Gangwar et al. 2023). Figure 6d inset shows a complied absorption spectra corresponding to each variation in MLE and at 0.5 × 10–3 M palladium chloride solution. From Fig. 7d inset graph, it is visualized that on increasing the volume of MLE, the absorbance of the colloidal MLE@PdNPs first decreases until it reaches the optimum volume, i.e., 2 ml, and then starts increasing. It suggests that the rate of formation of MLE@PdNPs first increases and then decreases. This might happen due to the competition between different organic moieties present in MLE to behave as a bioreductant.
a Absorption spectra of M. longifolia, b absorption spectra of palladium chloride solution, c absorption spectra of MLE@PdNPs different palladium chloride concentration at a fixed (1 ml) MLE volume, d absorption spectra of MLE@PdNPs for different volume of MLE at fixed (2.0 × 10–3 M) palladium chloride concentration
FTIR analysis
FTIR spectrum of pure MLE and MLE@PdNPs is shown in Fig. 9a, b. The presence of different valleys corresponding to different functional groups in the spectrum is evidence for the confirmatory synthesis of MLE@PdNPs. The detailed information of functional group present in pure MLE is summarized in Table 1, while several bands at 3599.17, 3057.17, 2331.93, 1710.85, 1514.12, 1381.03, 1201.65 & 1018.41, 891.11 cm−1 are observed in the FTIR spectrum of MLE@PdNPs (as shown in Fig. 9b) corresponding to N–H group; sp3–CH group of aldehyde or alkane; sp-hybrid carbon–carbon bond; > C = O group of aldehyde; sp2-hybrid carbon–carbon double bond of aromatic ring; C–N bond of amide or amine group; C–O bond of ester, alcohol, aldehyde, phenol; bending vibration of C–H bond of aromatic ring, respectively (Gangwar et al. 2023; Patil et al. 2018). The band below 500 cm−1 is due the interplanar vibration of MLE@PdNPs. It reveals that MLE@PdNPs are stabilized by several functional groups of phytochemicals present in MLE and prevent the formation of clustered MLE@PdNPs in a liquid state (Gangwar et al. 2023; Gopalkrishnan & Shimpi 2012; Patil et al. 2018; Yoshikawa et al. 2000).
Dynamic light scattering (DLS) study
DLS is used to study the overall surface charge of the synthesized MLE@PdNPs. Notably, dd water has been used as a dispersion medium for MLE@PdNPs. Figure 10 shows the electrokinetic potential distribution plot with a potential of − 18.40 mV, which inferred negative charge distribution over the surface of the MLE@PdNPs colloid having sufficient stability (Gangwar et al. 2023, 2022). Overall negative charge on the surface might be due to the presence of other phytochemicals which stabilized the synthesized MLE@PdNPs (Patil et al. 2018).
Cytotoxicity of MLE@PdNPs-treated A549 cells and their cytomorphological changes
The MLE@PdNPs were highly active against the lung cancer cells (cell line A549). The cytotoxicity of MLE@PdNPs was found to be dose-dependent and increased with increased concentration of MLE@PdNPs (as shown in Fig. 11). From Fig. 11, the IC-50 value of MLE@PdNPs on A549 cells has been evaluated 29.2 µg/ml. The phase-contrast microscopy images (shown in Fig. 12a, b) clearly show the changes in the morphology of untreated cells and A549 cells that were seen for IC-50 concentrations of MLE@PdNPs after the incubation for a specific time. Noticeably, no remarkable and distinct changes in the morphology of the control cells after the incubating for a day were observed with the uniform distribution or diffusion of cells (shown in Fig. 12a). Nonetheless, irregular, necrotic, shrunken, and detached morphology from the well surface was observed (Baker et al. 2021; Gangwar et al. 2023). Also, it has been observed that some of the cells were kept intact with the plasma membrane, illustrating apoptosis had begun in A549 cells treated with MLE@PdNPs (as shown in Fig. 12b).
The cytotoxicity of MLE@PdNPs and their hybrid nanomaterials may rest on the cell types. The cell propagation at the subG1 cell cycle was stopped upon the action of MLE@PdNPs (Al-Sheddi et al. 2018). The activated signaling pathways responsible for apoptosis were observed due to improved generation of reactive oxygen species (ROS) by exposure of MLE@PdNPs. Also, it is responsible for mitochondrial membrane damage and further leads to irrevocable cell destruction, although the lysosome degradation during autophagy and increasing programmed cancer cell death can clearly explain the apoptosis process (Gurunathan et al. 2009). Due to the apoptosis process, cell death was noticed in the cells treated with MLE@PdNPs and ultimately displayed the activation of caspase-3 and fragmentation/ condensation of chromatin (Mollick et al. 2019). Conclusively, due to the low toxicity of MLE@PdNPs and considerably high anti-carcinogenic properties, the prepared MLE@PdNPs are an excellent anti-carcinogenic agent. It may be successfully utilized for treating lung cancer cells (A549 cell lines) after executing the required clinical trials or experiments (Gangwar et al. 2023).
Estimation of reactive oxygen species (ROS) of MLE@PdNPs-treated A549 cells
To study the production of ROS in A549 cells 5- and 6-carboxy-2’,7’-dichlorodihydrofluorescein diacetate (DCFHDA) was used. The images captured upon the projection of DCFHDA on the control cells A549 and cells interact with MLE@PdNPs are shown in Fig. 13a, b, respectively. The fluorescence intensity has been observed relative to the ROS generated in the A549 cells. The present study reveals that the A549 cells treated with MLE@PdNPs (as shown in Fig. 12b) indicate the generation of higher intensity fluorescence than that in controls (Fig. 13a). The biogenic MLE@PdNPs emitted brilliant fluorescence with mutilated morphological structure due to stress disrupting impact in the compactness of plasma membrane caused by ROS produced. Interestingly, the untreated A549 cells did not showed any considerable fluorescence and held their native morphology. However, for quantification ImageJ software has been used to measure the fluorescence produced by ROS; MLE@PdNPs-treated A549 cells generated greater fluorescence than the untreated control cells (Fig. 13c).
Effect of MLE@PdNPs on nuclear morphology of A549
The fluorescent dye, i.e., 4’,6-diamidino-2-phenylindole (DAPI), has been used to examine the interaction of MLE@PdNPs with nuclear materials (as shown in Fig. 14a, b). Therefore, it helps to prove the internalization of particles into the nucleus. Previous studies predicted that the nanoparticles having anionic surface charge density followed the caveolae-mediated endocytosis and got to the nucleus without interacting with lysosomes. The A549 cells treated with MLE@PdNPs (IC-50 value equal to 29.2 µg/ml) were incubated in an incubator for a day at 37 °C and stained with DAPI. The apoptotic impact on A549 cells treated with MLE@PdNPs (Fig. 14a) was more prominent than on untreated A549 cells (Fig. 14b). To quantify treated and control cells, the ImageJ software was used (as shown in Fig. 14c). Also, it is clear from Fig. 13b that more fluorescence was obtained for the cells treated with MLE@PdNPs than the untreated cells.
Mechanism for cytotoxic action
Usually, nanoparticles having negative surface charge density internalize into the cells via caveolae-mediated endocytosis apart from diffusion (Chithrani et al. 2006; Valko et al. 2007). After the dissolution of caveosomes in the cells, the MLE@PdNPs are released into the cytoplasm. Because of the high surface energy and exchanging non-specifically with biomolecules, the free MLE@PdNPs cause cytotoxicity by generating ROS (O2−, H2O2, OH) (Gangwar et al. 2023; Valko et al. 2007). Living cells tend to neutralize intrinsic ROS, but a high level of ROS developed after the internalization of MLE@PdNPs causes severe damage to the cells. After internalization, the internalized MLE@PdNPs can disturb the mitochondrial membrane and cause damage to the key enzymes with the simultaneous release of caspase-3. This caspase-3 elicits intrinsic apoptosis with the simultaneous internalization of MLE@PdNPs into the nucleus and finally causes cell death (Baker et al. 2021; Yan et al. 2013). The cell death can be further confirmed by using DAPI. It was also evaluated that internalized MLE@PdNPs induced intracellular ROS generation in A549 cells and elicited apoptosis. The DCFDA method has been employed for evaluating the intracellular ROS production in MLE@PdNPs-treated cells, and it has been observed that it increases with increasing concentration of MLE@PdNPs compared to control. However, an overall increase in ROS production in mitochondria is responsible for the damage in the inner mitochondrial membrane-inducing disruption.
Antimicrobial activity
The antibacterial activity of MLE@PdNPs was studied against pathogenic bacterial strains of gram-negative E. coli, K. pneumoniae; Salmonella; and gram-positive S. aureus using the well diffusion method. The results of the antibacterial activity showed that MLE@PdNPs had efficient antibacterial activity against all gram-positive and negative bacterial strains. The inhibition zone of MLE@PdNPs against E. coli, K. pneumonia, Salmonella, and S. aureus has been 6.5, 6.2, 5.5, and 7.1 mm, respectively. Figure 15 a, b, c, and d shows the zone inhibition against S. aureus, K. pneumoniae, Salmonella, E. coli., respectively.
Minimum inhibition concentration
The antimicrobial activity of MLE@PdNPs was established by its MIC (Mohana & Sumathi 2020). The inoculated plates were examined for 24–48 h after incubation at 37 °C. The MIC value of the MLE@PdNPs was found to be around 30.85 ± 4.31 μg/mL. The IC-50 values have been found to be 26.01, 30.11, 32.12, and 35.16 μg/mL against S. aureus, K. pneumonia, Salmonella, and E. coli, respectively (as shown in Fig. 16). The results are in good resemblance with already reported studied against different pathogens (Chlumsky et al. 2021; Manikandan et al. 2016; Patil et al. 2018; Vinodhini et al. 2022a).
Mechanism of antibacterial action
The bactericidal effect of MLE@PdNPs is due to their adherence to bacterial cell walls surface and interactions with sulfur containing. It leads to irreversible change in sulfur-containing proteins (Abbaszadegan et al. 2015); reduce the compactness of lipid bilayer; and alter the permeability of the cell membrane, which leads to leakage of cellular contents, including ions, sugars, proteins, and the cellular energy reservoir (Lu et al. 2013). MLE@PdNPs were found active against a panel of G+ and G− bacteria due to the presence of antibacterial secondary metabolites, including dibenzopyranones and its derivatives, tenuazonic acid, altechromone A, altenusin, alternariol, alternariol monomethylether, altertoxin I, altertoxin II, and alterperylenol (Qader et al. 2021). Furthermore, MLE@PdNPs act against bacteria through excessive generation of reactive oxygen species H2O2, O2−, OH, etc. which interfere with respiration and membrane transport, release potassium ions from these cells, and interfere with cellular growth (Gomaa 2017). They also cause apoptosis-like responses; lipid peroxidation; depletion of antioxidant enzymes, such as GSH; and DNA damage (Korshed et al. 2016). These particles and palladium ions can damage cellular structures (e.g., ribosomes), and biomolecules such as proteins, lipids, and DNA, by interacting with them and hence disturb their maintenance and activities.
Conclusion
This study provides a sustainable method for synthesizing palladium nanoparticles (MLE@PdNPs) to obey the green chemistry principles. Here, Madhuca longifolia leaf extract in an aqueous medium was used as a bioreductant and stabilizing agent. Incorporating the plant-based extract is an efficient alternative methodology for the high-yielding fast synthesis of MLE@PdNPs, which becomes useful for large-scale synthesis and hence becomes an environmentally benign method. The advantages of the method we have used are the elimination of toxic chemicals, surfactants etc., easy work-up process, high yield, high tolerance ability for different functional groups, and a high percentage of reusability of the nanoparticles in the different areas. The results reveal that the triangular sheet-like stable MLE@PdNPs with an average crystallite size of 16.22 nm having a face center cubic (fcc) structure were obtained. The biological screening of MLE@PdNPs has been performed against the human lung cancer cells A549 and bacterial strains S. aureus, K. pneumonia, Salmonella, and E.coli which showed that MLE@PdNPs can be a promising material for formulating nanomedicine after subsequent clinical experiments. This study will become beneficial to expand the studies on the therapeutics potential of MLE@PdNPs in the future.
References
Abbaszadegan A, Ghahramani Y, Gholami A, Hemmateenejad B, Dorostkar S, Nabavizadeh M, Sharghi H (2015) The Effect of charge at the surface of silver nanoparticles on antimicrobial activity against gram-positive and gram-negative bacteria: a preliminary study. J Nanomater. https://doi.org/10.1155/2015/720654
Abdel-Rahman LH, Al-Farhan BS, El-ezz A, Sayed AE, Zikry MM, Abu-Dief AM (2022) Green biogenic synthesis of silver nanoparticles using aqueous extract of moringa oleifera: access to a powerful antimicrobial, anticancer, pesticidal and catalytic agents. J Inorg Organomet Polym Mater 32(4):1422–1435. https://doi.org/10.1007/s10904-021-02186-9
Al Qubeissi M, Mahmoud A, Al-Damook M, Almshahy A, Khatir Z, Soyhan HS, Raja Ahsan Shah RM (2023) Comparative analysis of battery thermal management system using biodiesel fuels. Energies. https://doi.org/10.3390/en16010565
Al-Sheddi ES, Farshori NN, Al-Oqail MM, Al-Massarani SM, Saquib Q, Wahab R, Siddiqui MA (2018) Anticancer potential of green synthesized silver nanoparticles using extract of nepeta deflersiana against human cervical cancer cells (HeLA). Bioinorg Chem Appl. https://doi.org/10.1155/2018/9390784
Annalakshmi R, Mahalakshmi S, Charles A, Sahayam CS (2013) GC–MS and HPTLC analysis of leaf extract of Madhuca longifolia (Koenig) Linn. Drug Invent Today 5(2):76–80. https://doi.org/10.1016/j.dit.2013.05.004
Baig N, Kammakakam I, Falath W (2021) Nanomaterials: a review of synthesis methods, properties, recent progress, and challenges. Mater Adv 2(6):1821–1871. https://doi.org/10.1039/D0MA00807A
Bains S, Kaur R, Sethi M (2020) Phytochemical analysis of medicinal plants and their antimicrobial activity. Agric Res J 57(3):444–448
Baker A, Iram S, Syed A, Elgorban AM, Al-Falih AM, Bahkali AH, Kim J (2021) Potentially bioactive fungus mediated silver nanoparticles. Nanomaterials (basel). https://doi.org/10.3390/nano11123227
Chithrani BD, Ghazani AA, Chan WCW (2006) Determining the size and shape dependence of gold nanoparticle uptake into mammalian cells. Nano Lett 6(4):662–668. https://doi.org/10.1021/nl052396o
Chlumsky O, Purkrtova S, Michova H, Sykorova H, Slepicka P, Fajstavr D, Demnerova K (2021) Antimicrobial properties of palladium and platinum nanoparticles: a new tool for combating food-borne pathogens. Int J Mol Sci. https://doi.org/10.3390/ijms22157892
Dalvi TS, Kumbhar UJ, Shah N (2022) Madhuca longifolia: ethanobotanical, phytochemical studies, pharmacological aspects with future prospects. Interdiscip J Appl Basic Sub 2(7):01–09
Dambhare AV, Patil PS, Khetade RH, Umekar MJ (2020) A review on: phytochemical screening and pharmacological activity on Madhuca longifolia. J Med Plants 8(2):54–60
Fahmy SA, Preis E, Bakowsky U, Azzazy HME (2020) Palladium nanoparticles fabricated by green chemistry: promising chemotherapeutic. Antioxid Antimicrob Agents Mater (basel) 13(17):3661. https://doi.org/10.3390/ma13173661
Gangwar C, Yaseen B, Kumar I, Singh NK, Naik RM (2021) Growth kinetic study of tannic acid mediated monodispersed silver nanoparticles synthesized by chemical reduction method and its characterization. ACS Omega 6(34):22344–22356. https://doi.org/10.1021/acsomega.1c03100
Gangwar C, Yaseen B, Nayak R, Praveen S, Kumar Singh N, Sarkar J, Mohan Naik R (2022) Silver nanoparticles fabricated by tannic acid for their antimicrobial and anticancerous activity. Inorg Chem Commun. https://doi.org/10.1016/j.inoche.2022.109532
Gangwar C, Yaseen B, Kumar I, Nayak R, Sarkar J, Baker A, Mohan Naik R (2023) Nano palladium/palladium oxide formulation using Ricinus communis plant leaves for antioxidant and cytotoxic activities. Inorg Chem Commun. https://doi.org/10.1016/j.inoche.2023.110417
Gomaa EZ (2017) Silver nanoparticles as an antimicrobial agent: a case study on Staphylococcus aureus and Escherichia coli as models for gram-positive and gram-negative bacteria. J Gen Appl Microbiol 63(1):36–43. https://doi.org/10.2323/jgam.2016.07.004
Gopalkrishnan B, Shimpi SN (2012) Pharmacognostical studies on stem bark of Madhuca longifolia (Koen.)Macbr. var. latifolia (Roxb.) A. Cheval. Indian J Nat Prod Resour 3:232–236
Gudala M, Banerjee S, Kumar R, Mandal A, Naiya T (2018) Experimental investigation on hydrodynamics of two-phase crude oil flow in horizontal pipe with novel surfactant. J Fluids Eng. https://doi.org/10.1115/1.4039130
Gurunathan S, Lee K-J, Kalishwaralal K, Sheikpranbabu S, Vaidyanathan R, Eom SH (2009) Antiangiogenic properties of silver nanoparticles. Biomaterials 30(31):6341–6350. https://doi.org/10.1016/j.biomaterials.2009.08.008
Jha D, Mazumder P (2018) Biological, chemical and pharmacological aspects of Madhuca longifolia. Asian Pac J Trop Med 11:9. https://doi.org/10.4103/1995-7645.223528
Jodh R, Tawar M, Kachewar A, Mahanur V, Sureka Y, Atole V (2022) Pharmacological review on Madhuca longifolia. Asian J Res Pharm Sci. https://doi.org/10.52711/2231-5659.2022.00006
Khare P, Kishore K, Sharma DK (2018) Medicinal uses, phytochemistry and pharmacological profile of Madhuca longifolia. Asian J Pharm Pharmacol 4(5):570–581
Korshed P, Li L, Liu Z, Wang T (2016) The molecular mechanisms of the antibacterial effect of picosecond laser generated silver nanoparticles and their toxicity to human cells. PLoS ONE. https://doi.org/10.1371/journal.pone.0160078
Kumar Y, Kumar B, Chandraker S, Padwar G, Dubey A, Thakur T, Sahu ML (2017) Mahua (Madhuca indica) (Koenig) J.F. Macribide) a nature, reward to tribal ecosystem of central india. Int J Curr Microbiol App Sci 6:1519–1526. https://doi.org/10.20546/ijcmas.2017.604.186
Kumar I, Yaseen B, Gangwar C, Mishra SK, Mohan Naik R (2021a) Environmental benign synthesis and characterization of nickel oxide nanoparticles using chicken egg white as template and evaluations of their antibacterial/antifungal activities. Mater Today Proc 46:2272–2276. https://doi.org/10.1016/j.matpr.2021.03.735
Kumar I, Yaseen B, Gangwar C, Yadav R, Mishra SK, Mohan Naik R (2021b) Ovalbumin mediated eco-friendly synthesis of silver oxide nanoparticles and their antibacterial and antifungal studies. Mater Today Proc 46:2330–2334. https://doi.org/10.1016/j.matpr.2021.04.403
Kuniyil M, Kumar JVS, Adil SF, Shaik MR, Khan M, Assal ME, Al-Warthan A (2019) One-pot synthesized Pd@N-doped graphene: an efficient catalyst for suzuki-miyaura couplings. Catalysts. https://doi.org/10.3390/catal9050469
Kurhade P, Kodape S, Junghare K, Bansod PG, Bhutada D (2022) Development of MgO nanoparticles via green synthesis at varying concentrations of precursor and mahua flower extract. Inorg Nano Met Chem 52:1–12. https://doi.org/10.1080/24701556.2022.2068581
Lu Z, Rong K, Li J, Yang H, Chen R (2013) Size-dependent antibacterial activities of silver nanoparticles against oral anaerobic pathogenic bacteria. J Mater Sci 24(6):1465–1471. https://doi.org/10.1007/s10856-013-4894-5
Manikandan V, Velmurugan P, Park J-H, Lovanh N, Seo S-K, Jayanthi P, Oh B-T (2016) Synthesis and antimicrobial activity of palladium nanoparticles from Prunus × yedoensis leaf extract. Mater Lett 185:335–338. https://doi.org/10.1016/j.matlet.2016.08.120
Mohana S, Sumathi S (2020) Multi-functional biological effects of palladium nanoparticles synthesized using agaricus bisporus. J Cluster Sci 31(2):391–400. https://doi.org/10.1007/s10876-019-01652-2
Mollick MMR, Rana D, Dash SK, Chattopadhyay S, Bhowmick B, Maity D, Chattopadhyay D (2019) Studies on green synthesized silver nanoparticles using abelmoschus esculentus (L.) pulp extract having anticancer (in vitro) and antimicrobial applications. Arabian J Chem. 12(8):2572–2584. https://doi.org/10.1016/j.arabjc.2015.04.033
MubarakAli D, Kim H, Venkatesh PS, Kim J-W, Lee S-Y (2022) A systemic review on the synthesis, characterization, and applications of palladium nanoparticles in biomedicine. Appl Biochem Biotechnol 194:1–20. https://doi.org/10.1007/s12010-022-03840-9
Muñiz-Diaz R, Gutiérrez de la Rosa SY, Gutiérrez Coronado Ó, Patakfalvi R (2022) Biogenic synthesis of platinum nanoparticles. Chem Pap 76:1–22. https://doi.org/10.1007/s11696-021-01970-8
Noah NM, Ndangili PM (2022) Green synthesis of nanomaterials from sustainable materials for biosensors and drug delivery. Sensors Int. https://doi.org/10.1016/j.sintl.2022.100166
Patil MP, Singh RD, Koli PB, Patil KT, Jagdale BS, Tipare AR, Kim G-D (2018) Antibacterial potential of silver nanoparticles synthesized using Madhuca longifolia flower extract as a green resource. Microb Pathog 121:184–189. https://doi.org/10.1016/j.micpath.2018.05.040
Prema P, Boobalan T, Arun A, Rameshkumar K, Babu RS, Veeramanikandan V, Balaji P (2022) Green tea extract mediated biogenic synthesis of gold nanoparticles with potent anti-proliferative effect against PC-3 human prostate cancer cells. Mater Lett. https://doi.org/10.1016/j.matlet.2021.130882
Qader MM, Hamed AA, Soldatou S, Abdelraof M, Elawady ME, Hassane ASI, Rateb ME (2021) Antimicrobial and antibiofilm activities of the fungal metabolites isolated from the marine endophytes epicoccum nigrum M13 and alternaria alternata 13A. Mar Drugs. https://doi.org/10.3390/md19040232
Raut R, Sana A, Malghe Y, Nikam B, Sahebrao K (2013) Rapid biosynthesis of platinum and palladium metal nanoparticles using root extract of asparagus racemosus linn. Adv Mater Lett 4:650–654. https://doi.org/10.5185/amlett.2012.11470
Reddy IS (2022) Madhuca indica: an untapped forest tree for its medicinal uses. Pharma Innov J 11(3):1747–1751
Rehman R, Anwar J, Mahmud T (2012) Thermodynamical and isothermal modeling of methylene blue dye batch biosorption on formalin modified madhuca longifolia leaf powder. Chem Soc Pakistan 34:460–467
Seku K, Bhagavanth Reddy G, Hussaini SS, Pejjai B, Hussain M, Reddy DM, Mangatayaru G (2022) An efficient biosynthesis of palladium nanoparticles using Bael gum and evaluation of their catalytic and antibacterial activity. Int J Biol Macromol 209:912–922. https://doi.org/10.1016/j.ijbiomac.2022.04.070
Siddiqi KS, Husen A (2016) Green synthesis, characterization and uses of palladium/platinum nanoparticles. Nanoscale Res Lett 11(1):482. https://doi.org/10.1186/s11671-016-1695-z
Singh SP, Yadav B, Anupam K (2022) Madhuca longifolia (Mahuwa).In: Herbs shrubs, trees potential medl benefits. pp 461–470
Sinha J, Singh V, Singh J, Rai A (2017) Phytochemistry, ethnomedical uses and future prospects of mahua (madhuca longifolia) as a food: a review. J Nutr Food Sci. https://doi.org/10.4172/2155-9600.1000573
Tanreh S, Hallajian S, Pourdakheli Y, Nazari P, Darvishi K, Hekmati M (2018) Green synthesis of pd nanoparticles mediated by thymbra spicata leaves extract and its application as a recyclable nanocatalyst for reduction of 4-nitrophenol and suzuki reactions. J Inorg Organomet Polym Mater. https://doi.org/10.1007/s10904-017-0775-6
Valko M, Leibfritz D, Moncol J, Cronin MTD, Mazur M, Telser J (2007) Free radicals and antioxidants in normal physiological functions and human disease. Int J Biochem Cell Biol 39(1):44–84. https://doi.org/10.1016/j.biocel.2006.07.001
Venkatesham M, Ayodhya D, Veerabhadram G (2015) Green synthesis, characterization and catalytic activity of palladium nanoparticles by xanthan gum. Appl Nanosci 5(3):315–320. https://doi.org/10.1007/s13204-014-0320-7
Verma N, Jha KK, Kumar U, Deepak K, Singh NK, Singh AK, Sharma R (2014) Biological properties, phytochemistry and traditional uses of Mahua (Madhuca longifolia): a review. Int J Adv Res Innov 2(3):630–638. https://doi.org/10.4103/1995-7645.223528
Vinodhini S, Vithiya BS, Arul Prasad TA (2022a) Green synthesis, characterization and antimicrobial activity of palladium nanoparticles : a review. J Appl Chem Sci Int 13:13–25. https://doi.org/10.56557/jacsi/2022/v13i37532
Vinodhini S, Vithiya BSM, Prasad TAA (2022b) Green synthesis of palladium nanoparticles using aqueous plant extracts and its biomedical applications. J King Saud Univ Sci. 34(4):102017. https://doi.org/10.1016/j.jksus.2022.102017
Yan L, Gu Z, Zhao Y (2013) Chemical mechanisms of the toxicological properties of nanomaterials: generation of intracellular reactive oxygen species. Chem Asian J 8(10):2342–2353. https://doi.org/10.1002/asia.201300542
Yaseen B, Gangwar C, Nayak R, Kumar S, Sarkar J, Banerjee M, Naik RM (2023) Gabapentin loaded silver nanoparticles (GBP@AgNPs) for its promising biomedical application as a nanodrug: anticancer and antimicrobial activities. Inorg Chem Commun 149:110380. https://doi.org/10.1016/j.inoche.2022.110380
Yoshikawa K, Tanaka M, Arihara S, Pal BC, Roy SK, Matsumura E, Katayama S (2000) New oleanene triterpenoid saponins from Madhuca longifolia. J Nat Prod 63(12):1679–1681. https://doi.org/10.1021/np000351r
Acknowledgements
The authors are thankful to the Head, Department of Chemistry, University of Lucknow, Lucknow (UP.), India, for providing the basic infrastructure for experimental work, UV-visible spectrophotometry, and FT-IR spectroscopy facility. The authors are also thankful to the Advanced Materials Research Centre (AMRC), Kamand Campus, I.I.T. Mandi, Himachal Pradesh, for providing an X-ray diffraction facility. The authors are also thankful to the sophisticated analytical instrumentation facility (SAIF), All India Institute of Medical Science (AIIMS), New Delhi, for providing an electron microscopy facility. The authors are thankful to Ms. Isha Gangwar (M.Tech. student, NIT Raipur, Chhattisgarh, India) for providing the satellite imagery of the location.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gangwar, C., Yaseen, B., Nayak, R. et al. Madhuca longifolia leaves-mediated palladium nanoparticles synthesis via a sustainable approach to evaluate its biomedical application. Chem. Pap. 77, 3075–3091 (2023). https://doi.org/10.1007/s11696-023-02688-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11696-023-02688-5