Abstract
This study builds upon prior work showing that methane (CH4) could be utilized as the sole electron donor and carbon source in a membrane biofilm reactor (MBfR) for complete perchlorate (ClO4 −) and nitrate (NO3 −) removal. Here, we further investigated the effects of salinity on the simultaneous removal of the two contaminants in the reactor. By testing ClO4 − and NO3 − at different salinities, we found that the reactor performance was very sensitive to salinity. While 0.2 % salinity did not significantly affect the hydrogen-based MBfR for ClO4 − and NO3 − removals, 1 % salinity completely inhibited ClO4 − reduction and significantly lowered NO3 − reduction in the CH4-based MBfR. In salinity-free conditions, NO3 − and ClO4 − removal fluxes were 0.171 g N/m2-day and 0.091 g/m2-day, respectively, but NO3 − removal fluxes dropped to 0.0085 g N/m2-day and ClO4 − reduction was completely inhibited when the medium changed to 1 % salinity. Scanning electron microscopy (SEM) showed that the salinity dramatically changed the microbial morphology, which led to the development of wire-like cell structures. Quantitative real-time PCR (qPCR) indicated that the total number of microorganisms and abundances of functional genes significantly declined in the presence of NaCl. The relative abundances of Methylomonas (methanogens) decreased from 31.3 to 5.9 % and Denitratisoma (denitrifiers) decreased from 10.6 to 4.4 % when 1 % salinity was introduced.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Perchlorate (ClO4 −) is widely used in rocket fuel, fireworks, and explosives due to its high energetic value as a strong oxidant and its chemical stability (USEPA 2005). Its release into water resources causes contamination. Perchlorate can interfere with the synthesis and secretion of thyroid hormone, which affects human metabolism and hinders human growth and development (USEPA 2008). Though the US Environmental Protection Agency (USEPA) has not yet established a nationwide maximum contaminant level (MCL) for ClO4 −, some states have set up cleanup target levels from 2 to 18 μg/L in drinking water (Gu and Coates 2006). Nitrate (NO3 −) is a co-contaminant with ClO4 − in groundwater (USEPA 2001). Nitrate has been regulated by the USEPA at 10 mg N/L in drinking water (USEPA 2009).
Commonly used technologies for removing ClO4 − and NO3 − from water include membrane filtration, ion exchange, chemical reduction, and biological reduction (Coates and Achenbach 2004; Matos et al. 2008; Zhao et al. 2014). Membrane filtration and resin regeneration in ion exchange produce highly saline ClO4 − and NO3 − brines that need to be further treated or disposed of. Biological reduction can be used to directly treat ClO4 −- and NO3 −-contaminated water or brines produced during membrane filtration or ion exchange. Compared to chemical reduction, biological reduction consumes less energy and requires less chemical addition because enzymes in microbes are used as the catalysts (Ye et al. 2012). During biological reduction, the perchlorate reductase (pcrABCD) reduces ClO4 − to chlorite (ClO2 −) (Kengen et al. 1999), and chlorite dismutase (cld) catalyzes the disproportionation of chlorite (ClO2 −) to chloride (Cl−) and oxygen (O2) (Van Ginkel et al. 1996), which is subsequently reduced to H2O. During the NO3 − reduction, reductions of nitrate to nitrite (narGHJI), nitrite to nitric oxide (nirSJFD/GH/L), NO to N2O (norZ), and N2O to N2 (nosZDFY) are performed by various reductases (Ettwig et al. 2010).
Perchlorate-reducing bacteria (PRB) and denitrifiers can use a variety of electron donors to gain energy (Miller and Logan 2000; Sahu et al. 2009; Son et al. 2006; Yu et al. 2006). When the electron donor is a gas, such as hydrogen and methane, membrane biofilm reactors (MBfRs) are one of the most widely used reactors because they have close to 100 % gas utilization efficiency and high NO3 − and ClO4 − removal rates (Zhao et al. 2013; Tang et al. 2012a, b; Luo et al. 2015; Chen et al. 2016).
Compared with CH4-based MBfRs, H2-based MBfRs have been extensively studied and used at the field scale for NO3 − and ClO4 − removal. The effects of salinity on nitrate and perchlorate reduction in H2-based MBfRs are well understood. H2-based MBfRs reach simultaneous reduction of nitrate and perchlorate, but the reduction rate decreases as the salt concentration increases. Chung et al. (2007) reported that nitrate and perchlorate reduction was not affected by 2 % salinity (20 g/L NaCl) in a H2-based MBfR, while 4 % salinity (40 g/L NaCl) decreased reduction by 40 %. Van Ginkel et al. (2008) investigated the kinetics of ClO4 − and NO3 − reduction in a H2-based MBfR fed with brines. They found that the NO3 − removal fluxes and H2 pressure were roughly first order but roughly zero order with NO3 − concentration in H2-based MBfR. NO3 − and ClO4 − were reduced simultaneously in a CH4-based MBfR (Luo et al. 2015), but the effects of salinity on NO3 − and ClO4 − reduction in CH4-based MBfRs are unknown.
CH4-based MBfRs have been much less studied compared to their H2-based counterparts. Previous research has demonstrated the potential of using CH4-based MBfRs to simultaneously remove NO3 − and ClO4 − from water (Luo et al. 2015; Chen et al. 2016); however, these studies were all conducted using synthetic fresh waters. Therefore, it is of interest to investigate how salinity affects biological perchlorate and nitrate reduction in a CH4-based MBfR. The overall objective of this work is to investigate the effects of salinity on the simultaneous reduction of NO3 − and ClO4 − in a CH4-based MBfR.
Materials and methods
MBfR setup
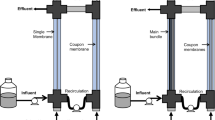
A two-column MBfR system similar to Zhao et al. (2011) was used in this study. The MBfR had composite hollow fibers (hydrophobic microporous polyethylene fiber, 280-μm o.d., 180-μm i.d., and 0.1–0.15-μm pore size) manufactured by Mitsubishi Rayon (Model MHF-200TL, Mitsubishi, Ltd., Japan). One column contained 32 fibers, and the other contained 20 fibers as “coupon fibers” for sampling purposes. Each fiber ran all the way through the column, with one end glued into a CH4 supply manifold and the other end sealed to avoid gas leakage. The total volume of the MBfR was 65 mL, and the total membrane surface area was 90.24 cm2. The MBfR was completely mixed via recirculation with a peristaltic pump (Longer Pump, Model 1515X, Longer Precision Pump Co, Ltd., China) at 100 mL/min.
Inoculum source and culture medium
We inoculated the MBfR with a 10-mL culture from a CH4-based MBfR (Luo et al. 2015). The medium contained the following mineral salts (analytical grade or purer) per liter of deionized water: 1 mg CaCl2, 0.3 g NaHCO3, 5 mg MgSO4∙7H2O, 0.2 g KH2PO4, 0.4 g NaHPO4∙12H2O, 1-mL acid trace element solution (100 mM HCL, 2.085 g FeSO4·7H2O, 68 mg ZnSO4·7H2O, 14 mg H3BO3, 120 mg CoCl2·6H2O, 500 mg MnCl2·4H2O, 320 mg CuSO4, 95 mg NiCl2·6H2O per liter), and 1-mL alkaline trace element solution (10 mM NaOH, 67 mg SeO2, 50 mg Na2WO4·2H2O, 242 mg Na2MoO4·2H2O per liter). The medium was degassed with N2 for 15 min to maintain anaerobic conditions. The medium pH was adjusted to 7.5 ± 0.2 with hydrochloric acid. The influent feeding rate was 0.5 mL/min (hydraulic retention time (HRT) of 130 min), the CH4 pressure was 10 psig (0.69 bar), and the temperature was 29 ± 1 °C for all experiments.
Start up and continuous operation of the MBfR
After inoculation, the MBfR was fed with NO3 − and ClO4 − at ∼15 and ∼ 10 mg/L, respectively, and maintained for more than 20 days to allow sufficient biomass to accumulate. After the startup stage, we fed the reactor with 2 mg/L of perchlorate from stage 1 to stage 5. To investigate the effects of salinity on ClO4 − and/or NO3 − reduction, we set up salinity at 1 % (10 g/L of NaCl) in stage 2 and 0.2 % (2 g/L of NaCl) in stages 5 and 6. To further investigate if nitrate inhibition plays a role in the inhibition mechanisms, we varied the nitrate concentrations in different stages: 10 mg NO3 −/L in stages 1, 2, and 3 and 5 mg NO3 −/L in stage 6. We allowed each stage to reach steady state, which was defined as when effluent concentrations were stable (<10 % variation) at least three times the HRT.
Analyses
Liquid samples were taken from the MBfR with 5-mL syringes and filtered immediately through 0.2-μm membrane filters (LC + PVDF membrane, Shanghai Xinya, China). NO3 − and NO2 − were assayed using ion chromatography (Metrohm 833 Basic IC plus, Switzerland) with an A-Supply-5 column and an eluent containing 3.2 mM NaHCO3, 1.0 mM Na2CO3, and 5 % acetone at a flow rate of 1 mL/min. ClO4 − was measured using ion chromatography (DIONEX ICS-1000, USA) with an AS 16 column and AG 16 precolumn, an eluent concentration of 35 mM KOH, and a 1.0 mL/min flow rate. The pH values of the influent and effluent were measured using a pH meter (Seven Easy, Mettler Toledo, Switzerland) and were between 7.5 and 7.7 for all stages. The bacterial morphology on the surface of membrane at the end of each stage was analyzed using scanning electron microscopy (SEM).
Biofilm sampling and DNA extraction
When the reactor reached a steady state, biofilm was collected. N2 gas was sparged at the sampling point to preclude O2 exposure. We cut off one 5-cm-long section from a coupon fiber and then sealed the remaining fiber by tying the end into a knot. The DNA was extracted using a DNeasy Blood & Tissue Kit (Qiagen, USA) as previously described by Zhao et al. (2011).
Quantification of bacterial 16S rRNA and functional genes
Plasmids containing target fragments were used as positive controls and to produce calibration curves (Zhao et al. 2011). The primers and qPCR conditions were the same as previously described: pcrA for ClO4 − reductase (Nozawa-Inoue et al. 2008), nirS for NO2 − reductase (Throbaeck et al. 2004), mcrA for the formation of methane by most of methanogens and reverse methanogenesis (Steinberg and Regan 2008), pMMO for CH4 monooxygenase (Paszczynski et al. 2011), and 16S ribosomal RNA (rRNA) gene for bacteria (Maeda et al. 2003). We used the SYBR Premix Ex Taq Kit (Takara Bio Inc., Japan) and performed qPCR as previously described by Zhao et al. (2011). The copy numbers of each gene were calculated by comparison to standard curves. Negative controls included water instead of template DNA in the PCR reaction mix. We performed triplicate PCR reactions for all samples and negative controls. The functional genes (pcrA, nirS, pMMO, mcrA) and 16S rRNA gene abundances in different stages are shown in Fig. 3.
Results and discussion
Perchlorate and nitrate reduction
CH4 pressure was set at 10 psig throughout the experiment, which made CH4 available in all stages (Table S1). The influent and effluent concentrations of ClO4 − and NO3 − for the MBfR are plotted in Fig. 1. The reactor was operated for 23 days in batch mode before stage 1 to accumulate sufficient biomass to reduce up to 5 mg/L of perchlorate and 15 mg/L of nitrate.
The NO3 − and ClO4 − surface loadings in stage 1 were 0.171 g N/m2-day and 0.091 g/m2-day, respectively. Without salinity, the nitrate was fully removed and 57.2 % of the ClO4 − was reduced after the reactor reached steady state. When 1 % salinity was applied in stage 2, the NO3 − removal percentage dropped to ∼50 %, and ClO4 − reduction was completely inhibited. Obviously, the perchlorate and nitrate reductions were both severely affected by the salinity in the CH4-based MBfR. Van Ginkel et al. (2008) reported that in a H2-based MBfR, NO3 − removal was not affected at NaCl concentrations less than 20 g/L (i.e., 2 %). Chung et al. (2007) demonstrated that both nitrate and perchlorate were reduced in the presence of NaCl ranging from 10 to 40 g/L (i.e., 1–4 %) in a H2-based MBfR. By comparison, microorganisms using methane as the electron donor are thus more sensitive to salinity than the microorganisms using H2.
To investigate if the biofilm could be recovered after salinity shock, we removed salinity in stage 3 so that the operating conditions in stage 3 were the same as in stage 1. To our surprise, the perchlorate and nitrate removal reached only 28.3 and 64.4 %, respectively. This removal is slightly higher than that observed in stage 2 but much lower than that in stage 1. The biofilm was possibly irreversibly damaged by the high salinity. Park et al. (2001) reported that the activity of denitrifying bacteria decreased after adding salt to the system. Gingras and Batista (2002) reported that the ClO4 − reduction rate declined to 10 % when 1–1.5 % salt concentrations were present in culture.
To investigate if NO3 − contributed to the inhibition of ClO4 − reduction, we removed NO3 − from the influent in stage 4. Without feeding NO3 −, the ClO4 − bioreduction improved to around 70 %. Therefore, it is possible that, in stage 2, the dual inhibitions of nitrate and salinity caused the absence of ClO4 − reduction. In a model, Tang et al. (2012a, b) explained that in a H2-MBfR simultaneously reducing NO3 − and ClO4 −, a high load of NO3 − (>0.6 g N/m2-day) strongly inhibits ClO4 − reduction.
To test the perchlorate or nitrate reduction under lower salinity, we reduced the salinity to 0.2 % NaCl in stages 5 and 6. ClO4 − was the only electron acceptor in stage 5, and NO3 − was the only electron acceptor in stage 6. The ClO4 − and NO3 − removals were 53.3 and 59.2 %, respectively. This performance is much worse than that in stage 1, which can be explained by the 0.2 % salinity inhibition and the irreversible inhibition in stage 2. The inhibition mechanisms will be further explored in “Microbial morphology” section.
Microbial morphology
To investigate the bacterial morphological change, we analyzed biomass samples from each stage at steady state using SEM. Figure 2 compares the microbial morphologies at stages 1 (Fig. 2a), 2 (Fig. 2b), 3 (Fig. 2c), and 6 (Fig. 2d). Stage 2 had the highest salinity (1 %). Correspondingly, it has the most wire-like structures and the cell surfaces had the most wrinkles (Fig. 2b). These bacterial filaments may be a way for the microorganisms to accommodate the change in osmotic pressure and protect the cells from salt shock. The damaging effect of salinity on bacteria leads to cell content emission and the formation of extracellular polymeric substances (EPS)/soluble microbial products (SMP) (Di Bella et al. 2013; Jang et al. 2013; Reid et al. 2006). Some studies argue that the accumulation of cell contents and intermediate products and bacterial autolysis are the responses to salt toxicity (Sharghi et al. 2014; Pendashteh et al. 2011; Sun et al. 2010). However, stage 6 had a medium salinity (0.2 %) and the biofilm also had some wire-like structures. Similarly, Jin et al. (2007) observed that under higher osmotic stress, the structure of a nitrifying bacterial community was disrupted and microbial diversity decreased. Therefore, salinity inhibition likely played a role in the decrease of nitrate removal in stage 6.
The SEM images of biomass samples from different stages. a Stage 1 at 10,000 magnification (10 mg/L NO3 −, 2 mg/L ClO4 −). b Stage 2 at 10,000 magnification (10 mg/L NO3 −, 2 mg/L ClO4 −, and 10 g/L NaCl). c Stage 3 at 10,000 magnification (10 mg/L NO3 −, 2 mg/L ClO4 −). d Stage 6 at 10,000 magnification (5 mg/L NO3 −, 2 g/L NaCl)
Functional community structure
Figure 3 shows the functional and 16S rRNA gene copy numbers of the microbial community at all stages. The bacterial community was very dense in stage 1, illustrated by the highest copy numbers of the 16S rRNA gene (2.84 × 1012 copies/m2 fiber). The functional respiration genes such as pcrA and nirS and methane oxidation gene pMMO reached the highest in stage 1, indicating a very good correlation between methane oxidation and nitrate/perchlorate reduction. When 1 % salinity was added in stage 2, the abundances of all tested genes dropped by three orders of magnitude and reached their lowest points, which was consistent with the drop in nitrate and perchlorate bioreduction. All the functional genes increased in stages 3 and 4, concurrent with the removal of NaCl, and decreased in stages 5 and 6 when 0.2 % salinity was added to the influent. Throughout the whole experiment, the functional gene abundance correlated very well with the salinity.
Microbial community structure
We analyzed the diversity and structure of the MBfR microbial communities using pyrosequencing of the 16S rRNA gene from the MBfR biofilm samples. Figure 4 shows the relative abundance of dominant microbial phylotypes at the genus level, while Fig. S1 shows the class level. Methylomonas was the dominant (31.3 %) bacterium in stage 1. Its abundance sharply decreased to 5.9 % in stage 2 when 1 % salinity was introduced and remained at low levels in the latter stages. Methylomonas are known methanotrophs and sensitive to salinity. For example, Sherry et al. (2016) reported the disappearance of Methylomonas when NaCl was higher than 1 g/L. Cunliffel et al. (2008) analyzed samples from surface (lower salinity) and subsurface (higher salinity) water of the Blyth estuary and found that Methylomonas was present in the surface samples but not in subsurface samples. In our MBfR, Methylococcus emerged in stage 2 (1.1 %), and its abundance continued to increase in the rest of stages. Methylococcus grows on CH4 under low O2 concentrations (Ward et al. 2004). Some studies show that Methylococcus is highly abundant in coastal saline soil (Mishra et al. 2012; Yousuf et al. 2012). Another methanotroph, Methylocystis, was present at a low abundance (2.1 %) in stage 1 but increased to ∼10 % in stage 2 at 1 % salinity and further increased to 21.3 % in stage 3 when the salinity was removed. Its abundance decreased to ∼10 % in stage 4 when nitrate was absent and perchlorate was the only electron acceptor. Methylomonas and Methylocystis reduce NO3 − using O2 to activate CH4 (Dam et al. 2013; Kits et al. 2015).
Methylophilus (Betaproteobacteria) abundance reached ∼9 % in stage 2 but remained at a low abundance in other stages. Giri et al. (2013) reported that Methylophilus sp. grow well at a NaCl concentration ≤200 mM. Methylophilus is phylogenetically close to known methanol assimilation denitrifiers. Figure 5 shows that Methylophilus is phylogenetically close to Dechloromonas agitata, Dechloromonas aromatica, and Azospira restricta, which are all known PRB. Therefore, Methylophilus probably played an important role in the reduction of NO3 − and ClO4 − under salinity stress. Denitratisoma abundance remained stable (∼10 %) through all stages. It is phylogenetically close to the known denitrifier Denitratisoma oestradiolicum, which reduces NO3 − to a mixture of N2O and N2 (Fahrbach et al. 2006). Unclassified Xanthomonadaceae, phylogenetically close to Thermomonas haemolytica, existed in all the stages and was significantly enriched in stages 2 and 3. It is possibly a denitrifier that can live under saline conditions. Mergaert et al. (2003) isolated two species of Thermomonas from a denitrifying reactor and reported that they could reduce NO3 − and NO2 −. Schmalenberger et al. (2013) reported the existence of Xanthomonadaceae in high salinity.
Figure S2 shows the PICRUSt analysis results of the functional genes. Genes encoding transporters (particularly ATP-binding cassette (ABC) transporters) and bacterial motility proteins were the most highly abundant. The abundance of these dominant genes decreased when the salinity increased in stage 2. Glycine betaine maintains osmotic pressure balance and bacterial morphology in salt environments, and ABC transporters are a member of the major betaine transporter family (Youssef et al. 2014; Silke et al. 2007). When NaCl appeared in stage 2, genes predicted to encode ABC transporters dropped to the lowest point. It is possible that the high salinity inhibited betaine absorption and caused changes in the microbial morphology and death, which explained the decreased NO3 − and ClO4 − reduction in stage 2.
Conclusions
In summary, salinity significantly affected the simultaneous ClO4 − and NO3 − reduction in a methane-based MBfR. Salinity at 1 % completely inhibited ClO4 − reduction and significantly reduced NO3 − reduction. Under saline conditions, the number of functional bacteria and total microorganisms decreased, wire-like cell structures became prevalent, and the cell surfaces developed wrinkles. In addition, the diversity of methanotrophs and denitrifying bacteria decreased significantly, and unclassified Xanthomonadaceae became more dominant when the salinity increased to 1 %. The saline environment changed cell osmotic pressure, which influenced the cell membranes and enzymes.
References
Chen R, Luo YH, Chen JX, Zhang Y, Wen LL, Shi LD, Zhao HP, Tang YN, Rittmann BE, Zheng P (2016) Evolution of the microbial community of the biofilm in a methane-based membrane biofilm reactor reducing multiple electron acceptors. Environ Sci Poll Res 23:9540–9548
Chung J, Nerenberg R, Rittmann BE (2007) Evaluation for biological reduction of nitrate and perchlorate in brine water using the hydrogen-based membrane biofilm reactor. J Environ Eng 133:157–164
Coates JD, Achenbach LA (2004) Microbial perchlorate reduction: rocket-fuelled metabolism. Nat Rev 2:569–580
Cunliffel M, Scha¨fer H, Harrison E, Cleave S, Goddard RU, Murrell JC (2008) Phylogenetic and functional gene analysis of the bacterial and archaeal communities associated with the surface microlayer of an estuary. The ISME Journal 2:776–789
Dam B, Dam S, Blom J, Liesack W (2013) Genome analysis coupled with physiological studies reveals a diverse nitrogen metabolism in Methylocystis sp. strain SC2. PLoS One. doi:10.1371/journal.pone.0074767
Di Bella G, Di Trapani D, Torregrossa M, Viviani G (2013) Performance of a MBR pilot plant treating high strength wastewater subject to salinity increase: analysis of biomass activity and fouling behavior. Bioresour Technol 147:614–618
Ettwig KF, Butler MK, Le Paslier D, Pelletier E, Mangenot S, Kuypers MMM, Schreiber F, Dutilh BE, Zedelius J, de Beer D, Gloerich J, Wessels HJCT, van Alen T, Luesken F, Wu ML, van de Pas-Schoonen KT, Op den Camp HJM, Janssen-Megens EM, Francoijs KJ, Stunnenberg H, Weissenbach J, Jetten MSM, Strous M (2010) Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464:544–548
Fahrbach M, Kuever J, Meinke R, Kampfer P, Hollender J (2006) Denitratisoma oestradiolicum gen. nov., sp. nov., a 17b-oestradiol-degrading, denitrifying betaproteobacterium. Int J Syst Evol Microbiol 56:1547–1552
Gingras TM, Batista JR (2002) Biological reduction of perchlorate in ion exchange regenerant solutions containing high salinity and ammonium levels. J Environ Monit 4:96–101
Giri DD, Kumar A, Shukla PN, Singh R, Singh PK, Pandey KD (2013) Salt stress tolerance of methylotrophic bacteria Methylophilus sp and Methylobacterium sp isolated from coal mine spoils. Pol J Microbiol 62:273–280
Gu B, Coates JD (2006) Perchlorate environmental occurrence: interactions and treatment. Springer, Boston
Jang D, Hwang Y, Shin H, Lee W (2013) Effects of salinity on the characteristics of biomass and membrane fouling in membrane bioreactors. Bioresour Technol 141:50–56
Jin RC, Zheng P, Mahmood Q, Hu BL (2007) Osmotic stress on nitrification in an airlift bioreactor. J Hazard Mater 146:148–154
Kengen SWM, Rikken GB, Hagen WR, Van Ginkel CG, Stams AJM (1999) Purification and characterization of perchlorate reductase from the chlorate-respiring strain GR-1. J Bacteriol 181:6706–6711
Kits DK, Klotz MG, Stein LY (2015) Methane oxidation coupled to nitrate reduction under hypoxia by the Gammaproteobacteria Methylomonas denitrificans sp. nov., type strain FJG1. Environ Microbiol. doi:10.1111/1462-2920.12772
Luo YH, Chen R, Wen LL, Meng F, Zhang Y, Lai CY, Rittmann BE, Zhao HP, Zheng P (2015) Complete perchlorate reduction using methane as the sole electron donor and carbon source. Environ. Sci. & Technol. 49:2341–2349
Maeda H, Fujimoto C, Haruki Y, Maeda T, Kokeguchi S, Petelin M, Arai H, Tanimoto I, Nishimura F, Takashiba S (2003) Quantitative real-time PCR using TaqMan and SYBR Green for Actinobacillus actinomycetemcomitans, Porphyromonas gingivalis, Prevotella intermedia, tetQ gene and total bacteria. FEMS Immunol Med Microbiol 39:81–86
Matos CT, Velizarov S, Reis MA, Crespo JG (2008) Removal of bromate from drinking water using the ion exchange membrane bioreactor concept. Environ. Sci. Technol. 42:7702–7708
Mergaert J, Cnockaert MC, Swings J (2003) Thermomonas fusca sp. nov. and Thermomonas brevis sp. nov., two mesophilic species isolated from a denitrification reactor with poly(e-caprolactone) plastic granules as fixed bed, and emended description of the genus Thermomonas. Int J Syst Evol Microbiol 53:1961–1966
Miller JP, Logan BE (2000) Sustained perchlorate degradation in an autotrophic, gas-phase, packed-bed bioreactor. Environ. Sci. Technol. 34:3018–3022
Mishra RR, Swain MR, Dangar TK, Thatoi H (2012) Diversity and seasonal fluctuation of predominant microbial communities in Bhitarkanika, a tropical mangrove ecosystem in India. Revista de Biologia Tropical 60:909–924
Nozawa-Inoue M, Jien M, Hamilton NS, Stewart V, Scow KM, Hristova KR (2008) Quantitative detection of perchlorate-reducing bacteria by real-time PCR targeting the perchlorate reductase gene. Appl Environ Microbiol 74:1941–1944
Park EJ, Seo JK, Kim MR, Jung IH, Kim JY, Kim SK (2001) Salinity acclimation of immobilized freshwater denitrifier. Aquac Eng 24:169–180
Paszczynski AJ, Paidisetti R, Johnson AK, Crawford RL, Colwell FS, Green T, Delwiche M, Lee H, Newby D, Brodie EL, Conrad M (2011) Proteomic and targeted qPCR analyses of subsurface microbial communities for presence of methane monooxygenase. Biodegradation 22:1045–1059
Pendashteh AR, Fakhru’l-Razi A, Madaeni SS, Abdullah LC, Abidin ZZ, Biak DRA (2011) Membrane foulants characterization in a membrane bioreactor (MBR) treating hypersaline oily wastewater. Chem Eng J 168:140–150
Reid E, Liu X, Judd SJ (2006) Effect of high salinity on activated sludge characteristics and membrane permeability in an immersed membrane bioreactor. J Membr Sci 283:164–171
Sahu AK, Conneely T, Nusslein KR, Ergas SJ (2009) Biological perchlorate reduction in packed bed reactors using elemental sulfur. Environ. Sci. Technol. 43:4466–4471
Schmalenberger A, O’ Sullivan O, Gahan J, Cotter PD, Courtney R (2013) Bacterial communities established in bauxite residues with different restoration histories. Environ. Sci. Technol. 47:7110–7119
Sharghi EA, Bonakdarpour B, Pakzadeh M (2014) Treatment of hypersaline produced water employing a moderately halophilic bacterial consortium in a membrane bioreactor: effect of salt concentration on organic removal performance, mixed liquor characteristics and membrane fouling. Bioresour Technol 164:203–213
Sherry A, Osborne KA, Sidgwick FR, Gray ND, Talbot HMA (2016) Temperate river estuary is a sink for methanotrophs adapted to extremes of pH, temperature and salinity. Environ Microbiol Rep 8:112–131
Silke S, Katharina P, Stephanie K, Regina S, Volker M (2007) The salt-induced ABC transporter Ota of the methanogenic archaeon Methanosarcina mazei Go1 is a glycine betaine transporter. FEMS Microbiol Lett 277:44–49
Son A, Lee J, Chiu PC, Kim BJ, Cha DK (2006) Microbial reduction of perchlorate with zero-valent iron. Water Res 40:2027–2032
Steinberg LM, Regan JM (2008) Phylogenetic comparison of the methanogenic communities from an acidic, oligotrophic fen and an anaerobic digester treating municipal wastewater sludge. Appl Environ Microbiol 74:6663–6671
Sun C, Leiknes T, Weitzenbock J, Thorstensen B (2010) Salinity effect on a biofilm-MBR process for shipboard wastewater treatment. Sep Sci Technol 72:380–387
Tang Y, Zhao HP, Marcus A, Krajmalnik-Brown R, Rittmann BE (2012a) A steady-state-biofilm model for simultaneous reduction of nitrate and perchlorate part 2: parameter optimization and results and discussion. Environ. Sci. Technol. 46:1608–1615
Tang YN, Zhou C, Van Ginkel S, Ontiveros-Valencia A, Shin JH, Rittmann BE (2012b) Hydrogen-permeation coefficients of the fibers used in H2-based membrane biofilm reactors. J Membr Sci 407:176–183
Throbaeck IN, Enwall K, Jarvis A, Hallin S (2004) Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol Ecol 49:401–417
United States Environmental Protection Agency (2001). Record of decision of the western groundwater operable unit OU-3, US EPA Region 9, San Francisco, CA. http://www.epa.gov/superfund/sites/rods/fulltext/r0901535.pdf
United States Environmental Protection Agency (2005). Perchlorate in the Pacific West. http://www.epa.gov/region9/toxic/ perchlorate/index.html
United States Environmental Protection Agency (2008). Federal facilities restoration and reuse, perchlorate. Available from http://www.epa.gov/swerffrr/documents/perchlorate/htm
United States Environmental Protection Agency (2009). National Primary Drinking Water Regulations. http://water.epa.gov/drink/contaminants/upload/mcl-2.pdf
Van Ginkel CG, Rikken GB, Kroon AGM, Kengen SWM (1996) Purification and characterization of chlorite dismutase: a novel oxygen-generating enzyme. Arch Microbiol 166:321–326
Van Ginkel SW, Ahn CH, Badruzzaman M, Roberts DJ, Lehman SG, Adham SS, Rittmann BE (2008) Kinetics of nitrate and perchlorate reduction in ion exchange brine using the membrane biofilm reactor (MBfR). Wat Res 42:4197–4205
Ward N, Larsen Ø, Sakwa J, Bruseth L, Khouri H, Durkin AS, Dimitrov G (2004) Genomic insights into methanotrophy: the complete genome sequence of Methylococcus capsulatus (bath). PLoS Biol 2:1616–1628
Ye L, You H, Yao J, Su H (2012) Water treatment technologies for perchlorate: a review. Desalination 298:1–12
Youssef NH, Savage-Ashlock KN, McCully AL, Luedtke B, Shaw EI, Hoff WD, Elshahed MS (2014) Trehalose/2-sulfotrehalose biosynthesis and glycine-betaine uptake are widely spread mechanisms for osmoadaptation in the Halobacteriales. The ISME Journal 8:636–649
Yousuf B, Sanadhya P, Keshri J, Jha B (2012) Comparative molecular analysis of chemolithoautotrophic bacterial diversity and community structure from coastal saline soils, Gujarat, India. BMC Microbiol 12:150 1471-2180
Yu XY, Amrhein C, Deshusses MA, Matsumoto MR (2006) Perchlorate reduction by autotrophic bacteria in the presence of zero-valent iron. Environ Sci Technol 40:1328–1334
Zhao HP, Van Ginkel S, Tang Y, Kang D-W, Rittmann BE, Krajmalnik-Brown R (2011) Interactions between perchlorate and nitrate reductions in the biofilm of a hydrogen-based membrane biofilm reactor. Environ. Sci. Technol. 45:10155–10162
Zhao HP, Ontiveros-Valencia A, Tang Y, Kim B, Krajmalnik-Brown R, Rittmann BE (2013) Using a two-stage hydrogen-based membrane biofilm reactor (MBfR) to achieve complete perchlorate reduction in the presence of nitrate and sulfate. Environ Sci & Technol 47(3):1565–1572
Zhao HP, Ontiveros-Valencia A, Tang Y, Kim B, Van-Ginkel S, Friese D, Overstreet R, Smith J, Evens P, Krajmalnik-Brown R, Rittmann BE (2014) Removal of multiple electron acceptors by pilot-scale, two-stage membrane biofilm reactors. Water Res 54:115–122
Acknowledgments
The authors greatly thank the “National Natural Science Foundation of China (Grant Nos. 21377109 and 21577123),” the “Fundamental Research Funds for the Central Universities (2016QNA6007),” and “Public Welfare Project of the Science and Technology Department of Zhejiang Province (2015C33016)” for their financial support. We thank Dr. Erica Hartmann at Northwestern University who helped us review the paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Bingcai Pan
Electronic supplementary material
ESM 1
(DOCX 1580 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Chen, JX., Wen, LL. et al. Effects of salinity on simultaneous reduction of perchlorate and nitrate in a methane-based membrane biofilm reactor. Environ Sci Pollut Res 23, 24248–24255 (2016). https://doi.org/10.1007/s11356-016-7678-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-016-7678-x